Introduction
Hepatocellular carcinoma (HCC) is characterized by
high morbidity and mortality, and is one of the most prevalent
malignancies worldwide (1). Although
progress has been made in the diagnosis and treatment of HCC, the
prognosis for patients with HCC remains poor due to resistance to
conventional chemotherapy and radiotherapy (2,3). Following
surgical treatment, >60% of patients experience recurrence and
metastasis within 1 year (4).
Biomarker-based tumor recognition has promise for condition
assessments that guide treatment; however, currently, few
biomarkers have been applied in clinical practice (5). Therefore, it remains important to
identify novel biomarkers for early diagnosis and prognosis
evaluation for patients with HCC.
Long non-coding RNAs (lncRNAs) are >200
nucleotides in length and do not code for, but may interact with,
proteins (6). Although not as
well-characterized as small non-coding RNAs, including microRNAs,
lncRNAs serve important roles in the regulation of a variety of
cellular processes, including stem cell pluripotency, cell growth,
cell proliferation, apoptosis, metabolism and cancer cell migration
(7–12). Functional lncRNAs may be useful in
cancer diagnosis and prognosis and may serve as potential
therapeutic targets. Tang et al (13) reported that three lncRNAs
(RP11-160H22.5, XLOC_014172 and LOC149086) were upregulated in HCC
relative to cancer-free controls. Furthermore, XLOC_014172 and
LOC149086 were confirmed to be markedly increased in metastatic
HCC. In addition, levels of these three lncRNAs were identified to
be decreased following cancer resection in the majority of
patients. Xu et al (14)
identified an lncRNA, LALR1, that was involved in liver
regeneration. Yuan et al (15)
observed that lncRNA-activated by transforming growth factor b
upregulation in HCC, modulated tumorigenesis and progression. Wang
et al (16) identified that
the oncofetal lncRNA PVT1 promoted proliferation and stem cell-like
properties in HCC cells, suggesting that this lncRNA may be
involved in HCC progression.
In the present study, microarray data from the human
lncRNA datasets GSE55191 and GSE5804 were analyzed, and it was
identified that lncRNA LOC728290 expression levels in HCC tissues
were significantly decreased compared with adjacent non-tumor
tissues (P<0.05). Distinct LOC728290 expression levels in HCC
and paired adjacent non-tumor samples were then validated using the
reverse transcription-quantitative polymerase chain reaction
(RT-qPCR). Furthermore, the association between LOC728290
expression levels, clinicopathological characteristics and
recurrence-free survival (RFS) times in patients with HCC were
analyzed to determine whether lncRNA LOC728290 may be a useful
diagnostic and prognostic indicator in HCC.
Materials and methods
Patients and specimens
Data from 65 consecutive patients (51 males and 14
females), who underwent surgery for HCC at 302 Beijing Hospital
(Beijing, China), between August 2013 and April 2016, were accessed
from the records of the Department of Hepatobiliary Surgery. None
of the patients had received preoperative chemotherapy or radiation
therapy. All HCC diagnoses were confirmed histopathologically by a
clinical pathologist. Tumor tissues and adjacent non-tumor tissue
specimens were collected from the patients subsequent to obtaining
written informed consent, in accordance with the institutional
guidelines of 302 Beijing Hospital's Ethics Committee. Resected
tumor tissue and adjacent normal tissue specimens were immediately
snap-frozen in liquid nitrogen and stored in a tissue bank until
use. The experimental operators were blinded to the clinical data.
Patient characteristics are presented in Table I.
 | Table I.Association of long non-coding RNA
LOC728290 expression with clinicopathological characteristics of
patients with hepatocellular carcinoma. |
Table I.
Association of long non-coding RNA
LOC728290 expression with clinicopathological characteristics of
patients with hepatocellular carcinoma.
|
|
| LOC728290
expression |
|
|---|
|
|
|
|
|
|---|
| Characteristic | Total | Low | High | P-value |
|---|
| Sex |
|
|
| 0.203 |
| Male | 51 | 28 | 23 |
|
|
Female | 14 | 5 | 9 |
|
| Age, years |
|
|
| 0.924 |
|
<60 | 41 | 21 | 20 |
|
| ≥60 | 24 | 12 | 12 |
|
| Tumor size, cm |
|
|
| 0.267 |
|
<5 | 28 | 12 | 16 |
|
| ≥5 | 37 | 21 | 16 |
|
| AFP, ng/ml |
|
|
| 0.033a |
|
<20 | 26 | 9 | 17 |
|
| ≥20 | 39 | 24 | 15 |
|
| Histological
grade |
|
|
| 0.136 |
| Well | 2 | 2 | 0 |
|
|
Moderately/poorly | 56 | 26 | 30 |
|
| Clinical stage |
|
|
| 0.236 |
| I and
II | 47 | 26 | 21 |
|
| III and
IV | 18 | 7 | 11 |
|
| Tumor number |
|
|
| 0.087 |
|
Solitary | 54 | 30 | 24 |
|
|
Multiple | 11 | 3 | 8 |
|
| Alcohol
consumption |
|
|
| 0.267 |
|
Yes | 30 | 13 | 17 |
|
| No | 35 | 20 | 15 |
|
| Smoking status |
|
|
| 0.165 |
|
Yes | 23 | 9 | 14 |
|
| No | 42 | 24 | 18 |
|
| HBV |
|
|
| 0.156 |
|
Yes | 39 | 17 | 22 |
|
| No | 26 | 16 | 10 |
|
| Recurrence |
|
|
| 0.337 |
|
Yes | 22 | 13 | 9 |
|
| No | 43 | 20 | 23 |
|
| PVTT |
|
|
| 0.172 |
|
Yes | 33 | 14 | 19 |
|
| No | 32 | 19 | 13 |
|
| Microvascular
invasion |
|
|
| 0.022a |
|
Yes | 47 | 28 | 19 |
|
| No | 18 | 5 | 13 |
|
| Liver
cirrhosis |
|
|
| 0.221 |
|
Absence | 35 | 14 | 21 |
|
|
Presence | 25 | 14 | 11 |
|
RNA preparation, reverse transcription
and RT-qPCR
RNA from frozen HCC tissues and adjacent non-tumor
tissues (n=65) was extracted using TRIzol reagent (Thermo Fisher
Scientific, Inc., Waltham, MA, USA), according to the
manufacturer's protocol. RNA integrity was evaluated using a
NanoDrop ND-1000 spectrophotometer (NanoDrop Technologies; Thermo
Fisher Scientific, Inc.) and cDNA was synthesized from 50 ng total
RNA for each sample. LOC728290 expression levels were quantified
using RT-qPCR performed on an ABI 7500 system (Applied Biosystems;
Thermo Fisher Scientific, Inc.) using Maxima SYBR-Green RT-qPCR
master mix (Thermo Fisher Scientific, Inc.), following the
manufacturers' protocols. GAPDH expression was monitored as the
endogenous control, and all samples were normalized to human GAPDH.
All reactions were run in triplicate, using LOC728290-specific
primers designed and synthesized by Sangon Biotech Co., Ltd.
(Shanghai, China). Primer sequences were as follows: LOC728290
5′-AAAGCACAGGTGACTGTAACAC-3′ (forward) and
5′-TGGGCATTCTCATCGCAGTC-3′ (reverse), and GAP DH
5′-CAGCCTCAAGATCATCAGCA-3′ (forward) and 5′-TGTGGTCATGAGTCCTTCCA-3′
(reverse). The amplification profile was 95°C for 5 min, followed
by 40 cycles of denaturation at 95°C for 15 sec and annealing at
60°C for 30 sec. The median of triplicate reactions was used to
calculate relative lncRNA expression (ΔCq=Cq median lncRNA-Cq
median GAPDH). Expression fold changes were calculated using the
2−ΔΔCq method (17).
Statistical analysis
Statistically significant differences between groups
were determined using two-tailed Student's t-tests. A receiver
operating characteristic (ROC) curve was plotted to determine the
discrimination of the expression level of LOC728290 discriminated
between HCC tissues and adjacent non-tumor tissues. Associations
between LOC728290 expression and clinicopathological
characteristics were analyzed using one-way analysis of variance
with Bonferroni correction. The association between RFS times and
LOC728290 expression levels in patients with HCC was analyzed using
Kaplan-Meier estimator analysis. All statistical analyses were
performed using SPSS for Windows software (version 16.0; SPSS Inc.,
Chicago, IL, USA). P<0.05 was considered to indicate a
statistically significant difference.
Results
lncRNA LOC728290 is downregulated in
HCC tissues relative to adjacent non-tumor tissues
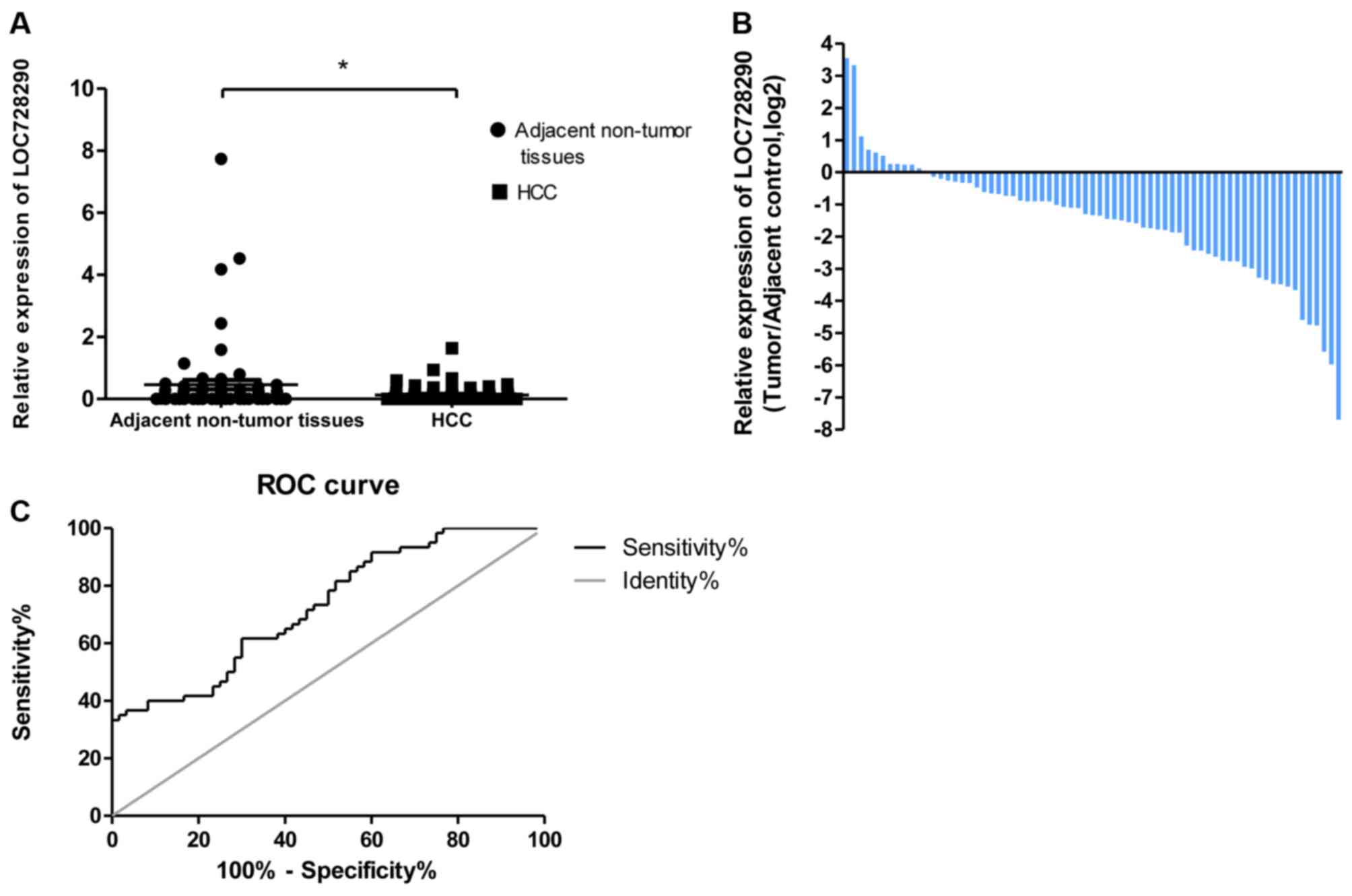
To assess the potential clinical significance of
LOC728290, the expression level in HCC tissues and adjacent
non-tumor tissues was analyzed using RT-qPCR. LOC728290 expression
was significantly downregulated in 83.0% of tumors (54/65; fold
change ≥1.0), relative to adjacent non-tumor tissues (P<0.05;
Fig. 1A and B). A ROC analysis was
performed to evaluate the ability of the LOC728290 expression to
discriminate between the tumor and control samples. The total area
under the curve (AUC) for LOC728290 was 0.728 (Fig. 1C), suggesting that the LOC728290 level
has adequate sensitivity and specificity to discriminate between
HCC tissues and adjacent non-tumor tissues.
lncRNA LOC728290 expression is
associated with α-fetoprotein (AFP) levels and with microvascular
invasion in patients with HCC
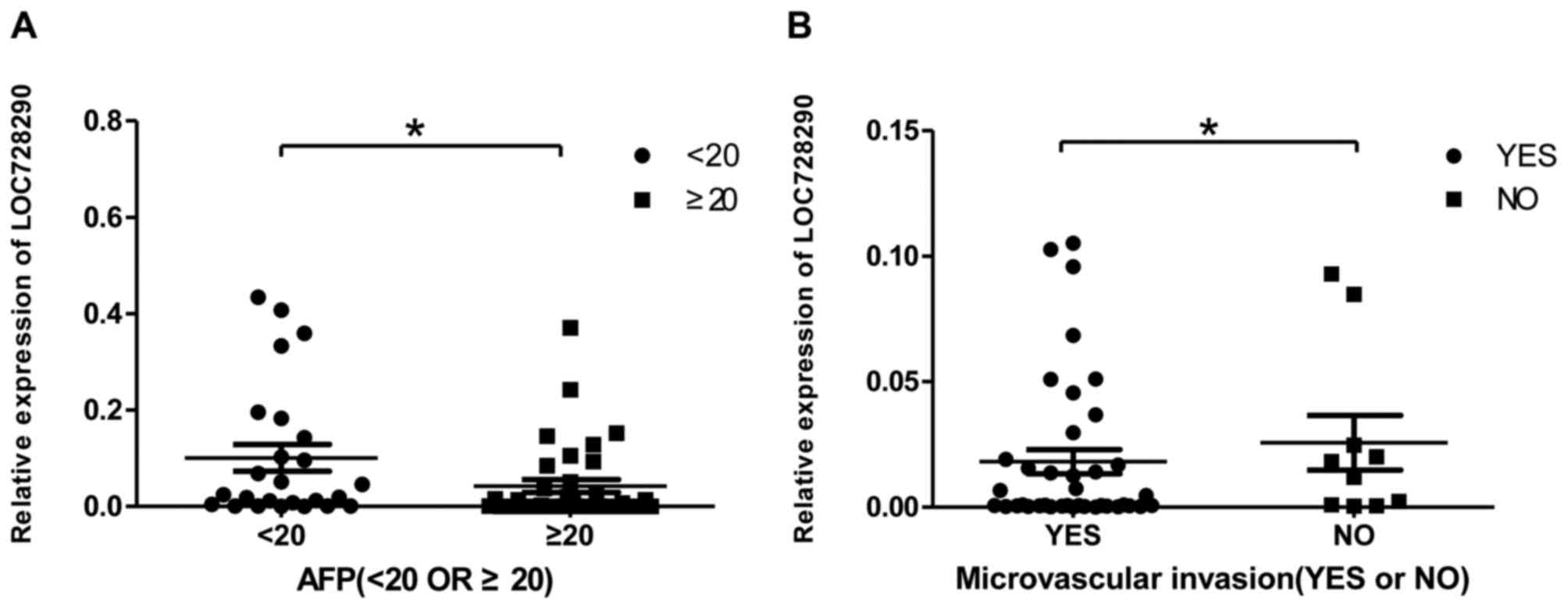
To determine whether LOC728290 expression in HCC
tissue was associated with clinicopathological parameters, AFP
levels and the presence of microvascular invasion in samples from
patients with HCC were examined. The AFP level is a critical tumor
biomarker for patients with HCC. LOC728290 expression was decreased
in tissue samples where serum AFP was ≥20 ng/ml (Fig. 2A). In addition, decreased LOC728290
expression was exhibited in patients with microvascular invasion
compared with those without (Fig.
2B).
Association between lncRNA LOC728290
expression and clinicopathological features of patients with
HCC
To further analyze the association between lncRNA
expression and clinicopathological parameters, the 65 patients with
HCC were divided into high-(n=32) and low-(n=33) LOC728290
expression groups, according to the mean value of LOC728290
expression in their tumor tissues. As presented in Table I, whereas the low-LOC728290 expression
group exhibited increased serum AFP levels (P<0.05) and
microvascular invasion (P<0.01) compared with the high-LOC728290
group, no significant association was identified between LOC728290
expression and other clinicopathological features including age,
sex, tumor size, clinical stage, histological grade, alcohol
consumption, smoking status, hepatitis B virus (HBV), recurrence,
portal vein tumor thrombosis (PVTT) or liver cirrhosis.
LOC728290 expression is associated
with RFS times in HCC
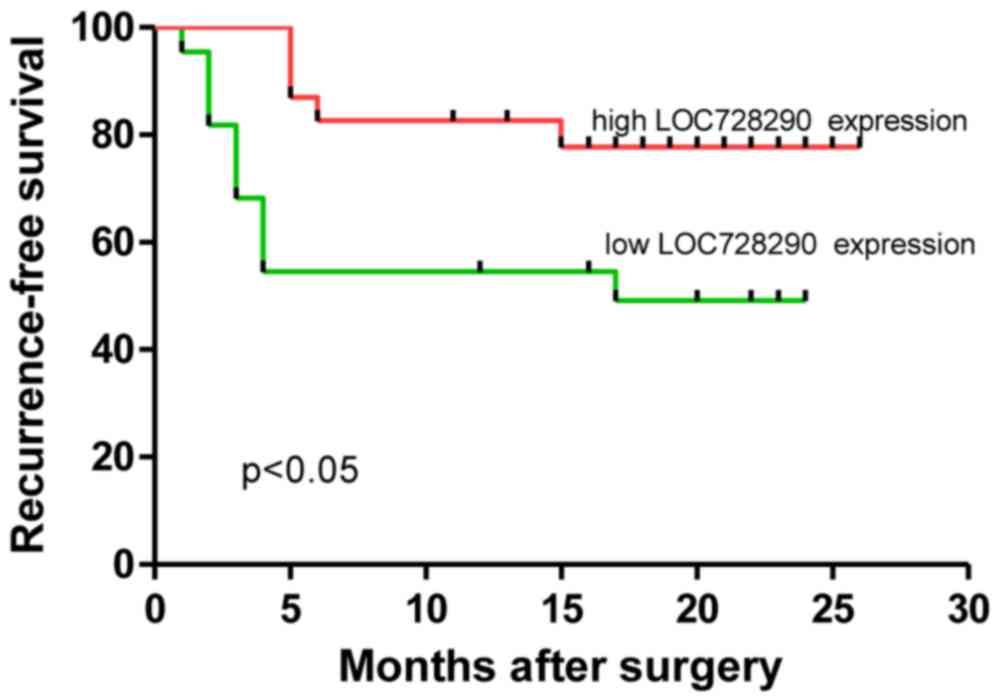
Kaplan-Meier estimator and log-rank analyses were
performed to examine the association between levels of LOC728290
expression and patient survival. As presented in Table II, LOC728290 expression, PVTT, tumor
size and microvascular invasion were significantly associated with
RFS times. In particular, patients with a low level of LOC728290
expression exhibited significantly decreased RFS times (P=0.023;
Fig. 3) compared with patients with
high LOC728290 expression. Significantly decreased RFS times were
also observed in patients with PVTT (P<0.05), tumors ≥5 cm
(P<0.001) or microvascular invasion (P<0.001). Thus, these
data indicate that low expression of LOC728290 may indicate a
poorer prognosis for patients with HCC.
 | Table II.Univariate analysis of RFS times for
the 65 patients with hepatocellular carcinoma studied. |
Table II.
Univariate analysis of RFS times for
the 65 patients with hepatocellular carcinoma studied.
| Variable | n | P-value |
|---|
| Sex |
| 0.33 |
|
Male | 51 |
|
|
Female | 14 |
|
| Age, years |
| 0.302 |
|
<60 | 41 |
|
|
≥60 | 24 |
|
| Tumor size, cm |
| 0.0067b |
|
<5 | 28 |
|
| ≥5 | 37 |
|
| AFP, ng/ml |
| 0.1673 |
|
<20 | 26 |
|
|
≥20 | 39 |
|
| Histological
grade |
| 0.5412 |
|
Well | 2 |
|
|
Moderately/poorly | 56 |
|
| Clinical stage |
| 0.7505 |
| I and
II | 47 |
|
| III and
IV | 18 |
|
| Tumor number |
| 0.3714 |
|
Solitary | 54 |
|
|
Multiple | 11 |
|
| Alcohol
consumption |
| 0.7186 |
|
Yes | 30 |
|
| No | 35 |
|
| Smoking status |
| 0.4280 |
|
Yes | 23 |
|
| No | 42 |
|
| HBV |
| 0.4855 |
|
Yes | 39 |
|
| No | 26 |
|
| PVTT |
| 0.0407a |
|
Yes | 33 |
|
| No | 32 |
|
| Microvascular
invasion |
| 0.0261a |
|
Yes | 47 |
|
| No | 18 |
|
| Liver
cirrhosis |
| 0.5881 |
|
Absence | 35 |
|
|
Presence | 25 |
|
| LOC728290
expression |
| 0.047a |
|
Low | 33 |
|
|
High | 32 |
|
Discussion
HCC is one of the most common malignant tumors in
China; its incidence and resulting mortality have increased
annually (18). The mechanisms of
occurrence and development of HCC are complex and involve altered
activity of a number of oncogenes and tumor suppressor genes;
abnormal expression of lncRNAs has been demonstrated to serve an
important role in the processes of invasion and metastasis
(19–21). Previous studies have revealed that the
lncRNAs H19, MALAT1 and HULC are important in HCC progression
(22–24); however, only a limited number studies
have addressed the biological function and clinical significance of
lncRNAs in HCC.
In the present study, differential expression of a
novel lncRNA, LOC728290, between HCC tissue and adjacent non-tumor
tissues from patients with HCC was identified. Results demonstrated
that LOC728290 expression in HCC was significantly decreased
compared with non-tumor tissue. The ROC AUC of LOC728290 was 0.728,
demonstrating specificity and sensitivity in the diagnosis of HCC.
Patients with HCC with low levels of LOC728290 expression exhibited
significantly decreased RFS times (P=0.047) compared with patients
with high expression. Furthermore, patients with HCC with PVTT
(P<0.05), tumor size ≥5 cm (P<0.001) and microvascular
invasion (P<0.001) exhibited significantly decreased RFS times
(P<0.001), respectively, which initially revealed the prognostic
value of LOC728290. Overall, LOC728290 may serve an important role
in the development and progression of HCC.
It has been reported that aberrant lncRNA expression
results in dysregulation of downstream effectors and that lncRNAs
may serve essential roles in numerous biological functions leading
to HCC, as lncRNAs including HOTAIR, H19 and ZEB1-AS1 have been
identified to be potential prognostic indicators in a number of
tumors (25–27).
In the present study, the association between
LOC728290 expression and clinicopathological characteristics of
patients with HCC was analyzed and a significant negative
association was revealed between LOC728290 expression and serum AFP
levels. Determination of serum AFP levels is essential in the
clinical diagnosis of liver cancer (28,29). Thus,
the results of the present study led to the hypothesis that
LOC728290 may be a promising biomarker for HCC. In addition, the
results of the present study revealed that decreased LOC728290
expression was associated with microvascular invasion, which, along
with PVTT, is a primary mechanism of extrahepatic metastasis in HCC
(30,31). Results from the present study
indicated that LOC728290 may serve an important role in the
development of HCC and that an investigation of possible mechanisms
is warranted. However, no significant association was revealed
between LOC728290 expression and other clinicopathological
features, including age, sex, tumor size, clinical stage,
histological grade, alcohol consumption, smoking status, HBV
infection, tumor recurrence, PVTT or liver cirrhosis. Much larger
clinical cohorts and extended follow-up times are required to
confirm the results of the present study. Further in vitro
and in vivo experiments are required to investigate the
function of lncRNA LOC728290 in HCC and to elucidate its role in
the underlying molecular mechanisms of HCC onset and
progression.
To the best of our knowledge, the results of the
present study identified for the first time that lncRNA LOC728290
levels were significantly decreased in HCC tissues and that
downregulation of lncRNA LOC728290 was positively associated with
increased serum AFP levels and microvascular invasion in patients
with HCC. Patients with HCC with low levels of LOC728290 expression
demonstrated significantly decreased RFS times compared with
patients with higher expression. These results demonstrate that
lncRNA LOC728290 may exhibit potential as a diagnostic and
prognostic biomarker for HCC.
References
|
1
|
Tsochatzis EA, Meyer T and Burroughs AK:
Hepatocellular carcinoma. N Engl J Med. 366:92–93. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Llovet JM, Burroughs A and Bruix J:
Hepatocellular carcinoma. Lancet. 362:1907–1917. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Maluccio M and Covey A: Recent progress in
understanding, diagnosing, and treating hepatocellular carcinoma.
CA Cancer J Clin. 62:394–399. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Zimmerman MA, Ghobrial RM, Tong MJ, Hiatt
JR, Cameron AM, Hong J and Busuttil RW: Recurrence of
hepatocellular carcinoma following liver transplantation: A review
of preoperative and postoperative prognostic indicators. Arch Surg.
143:182–188. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Ludwig JA and Weinstein JN: Biomarkers in
cancer staging, prognosis and treatment selection. Nat Rev Cancer.
5:845–856. 2005. View
Article : Google Scholar : PubMed/NCBI
|
|
6
|
Esteller M: Non-coding RNAs in human
disease. Nat Rev Genet. 12:861–874. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Kung JT, Colognori D and Lee JT: Long
noncoding RNAs: Past, present, and future. Genetics. 193:651–669.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Wang J, Xie G, Singh M, Ghanbarian AT,
Raskó T, Szvetnik A, Cai H, Besser D, Prigione A, Fuchs NV, et al:
Primate-specific endogenous retrovirus-driven transcription defines
naive-like stem cells. Nature. 516:405–409. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Wang Y, Wang Y, Li J, Zhang Y, Yin H and
Han B: CRNDE, a long-noncoding RNA, promotes glioma cell growth and
invasion through mTOR signaling. Cancer Lett. 367:122–128. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Leveille N, Melo CA, Rooijers K,
Díaz-Lagares A, Melo SA, Korkmaz G, Lopes R, Moqadam Akbari F, Maia
AR, Wijchers PJ, et al: Genome-wide profiling of p53-regulated
enhancer RNAs uncovers a subset of enhancers controlled by a
lncRNA. Nat Commun. 6:65202015. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Qiu JJ, Wang Y, Ding JX, Jin HY, Yang G
and Hua KQ: The long non-coding RNA HOTAIR promotes the
proliferation of serous ovarian cancer cells through the regulation
of cell cycle arrest and apoptosis. Exp Cell Res. 333:238–248.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Cui M, Xiao Z, Wang Y, Zheng M, Song T,
Cai X, Sun B, Ye L and Zhang X: Long noncoding RNA HULC modulates
abnormal lipid metabolism in hepatoma cells through an
miR-9-mediated RXRA signaling pathway. Cancer Res. 75:846–857.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Tang J, Jiang R, Deng L, Zhang X, Wang K
and Sun B: Circulation long non-coding RNAs act as biomarkers for
predicting tumorigenesis and metastasis in hepatocellular
carcinoma. Oncotarget. 6:4505–4515. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Xu D, Yang F, Yuan JH, Zhang L, Bi HS,
Zhou CC, Liu F, Wang F and Sun SH: Long noncoding RNAs associated
with liver regeneration 1 accelerates hepatocyte proliferation
during liver regeneration by activating Wnt/β-catenin signaling.
Hepatology. 58:739–751. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Yuan JH, Yang F, Wang F, Ma JZ, Guo YJ,
Tao QF, Liu F, Pan W, Wang TT, Zhou CC, et al: A long noncoding RNA
activated by TGF-β promotes the invasion-metastasis cascade in
hepatocellular carcinoma. Cancer Cell. 25:666–681. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Wang F, Yuan JH, Wang SB, Yang F, Yuan SX,
Ye C, Yang N, Zhou WP, Li WL, Li W and Sun SH: Oncofetal long
noncoding RNA PVT1 promotes proliferation and stem cell-like
property of hepatocellular carcinoma cells by stabilizing NOP2.
Hepatology. 60:1278–1290. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Lechel A and Gougelet A: Early HCC
treatment: A future strategy against interferon/miR-484 axis to
revert precancerous lesions? Gut. 65:1073–1074. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Iguchi T, Uchi R, Nambara S, Saito T,
Komatsu H, Hirata H, Ueda M, Sakimura S, Takano Y, Kurashige J, et
al: A long noncoding RNA, lncRNA-ATB, is involved in the
progression and prognosis of colorectal cancer. Anticancer Res.
35:1385–1388. 2015.PubMed/NCBI
|
|
20
|
Qiu JJ, Lin YY, Ding JX, Feng WW, Jin HY
and Hua KQ: Long non-coding RNA ANRIL predicts poor prognosis and
promotes invasion/metastasis in serous ovarian cancer. Int J Oncol.
46:2497–2505. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Yue B, Qiu S, Zhao S, Liu C, Zhang D, Yu
F, Peng Z and Yan D: LncRNA-ATB mediated E-cadherin repression
promotes the progression of colon cancer and predicts poor
prognosis. J Gastroenterol Hepatol. 31:595–603. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Luo F, Sun B, Li H, Xu Y, Liu Y, Liu X, Lu
L, Li J, Wang Q, Wei S, et al: A MALAT1/HIF-2α feedback loop
contributes to arsenite carcinogenesis. Oncotarget. 7:5769–5787.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Panzitt K, Tschernatsch MM, Guelly C,
Moustafa T, Stradner M, Strohmaier HM, Buck CR, Denk H, Schroeder
R, Trauner M and Zatloukal K: Characterization of HULC, a novel
gene with striking up-regulation in hepatocellular carcinoma, as
noncoding RNA. Gastroenterology. 132:330–342. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Ohrnberger S, Thavamani A, Braeuning A,
Lipka DB, Kirilov M, Geffers R, Autenrieth SE, Römer M, Zell A,
Bonin M, et al: Dysregulated serum response factor triggers
formation of hepatocellular carcinoma. Hepatology. 61:979–989.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Li J, Chen Z, Tian L, Zhou C, He MY, Gao
Y, Wang S, Zhou F, Shi S, Feng X, et al: LncRNA profile study
reveals a three-lncRNA signature associated with the survival of
patients with oesophageal squamous cell carcinoma. Gut.
63:1700–1710. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Thum T: Noncoding RNAs and myocardial
fibrosis. Nat Rev Cardiol. 11:655–663. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Zhang S, Chen S, Yang G, Gu F, Li M, Zhong
B, Hu J, Hoffman A and Chen M: Long noncoding RNA HOTAIR as an
independent prognostic marker in cancer: A meta-analysis. PLoS One.
9:e1055382014. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Carr BI, Guerra V, Giannini EG, Farinati
F, Ciccarese F, Rapaccini GL, Di Marco M, Benvegnù L, Zoli M,
Borzio F, et al: Significance of platelet and AFP levels and liver
function parameters for HCC size and survival. Int J Biol Markers.
29:e215–e223. 2014.PubMed/NCBI
|
|
29
|
Dwyer JP, Hosking P and Lubel J: Multiple
liver lesions in a patient with positive hepatitis C serology and
elevated AFP: Is it HCC? Gastroenterology. 147:e12–e13. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Li T, Xie J, Shen C, Cheng D, Shi Y, Wu Z,
Deng X, Chen H, Shen B, Peng C, et al: Upregulation of long
noncoding RNA ZEB1-AS1 promotes tumor metastasis and predicts poor
prognosis in hepatocellular carcinoma. Oncogene. 35:1575–1584.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Li T, Xie J, Shen C, Cheng D, Shi Y, Wu Z,
Deng X, Chen H, Shen B, Peng C, et al: Amplification of long
noncoding RNA ZFAS1 promotes metastasis in hepatocellular
carcinoma. Cancer Res. 75:3181–3191. 2015. View Article : Google Scholar : PubMed/NCBI
|