Introduction
In the era of big data, various types of data can be
obtained and shared by all users through the internet or social
network systems. One method to efficiently manage enormous amounts
of data is to apply deep learning (1). One characteristic of deep learning is
that this approach does not require features or representations to
be selected during data input. Recently, a number of studies have
focused on image classification by deep learning, and these
technologies are now becoming readily available for use by
corporations and individuals (2–6). For
example, TensorFlow is a Google software library for machine
learning that was released under an open-source license in 2015
(7). Using this technology, the
present study aimed to potentially integrate deep learning into
gynecological clinical practice.
Cervical cancer is a leading cause of death in women
worldwide (8). Although mortality
rates were drastically reduced following the introduction of the
Pap smear test, determining the types of patients who should be
further screened and treated as high-risk remains an important
issue, particularly for avoiding overmedication (9–11). In
daily practice, patient management is determined by the combined
use of cytology, histology, HPV typing and colposcopy results. The
present study investigated whether deep learning with a focus on
colposcopy images as input could predict the postoperative
diagnosis.
Colposcopy is a well-established tool for observing
the cervix at up to ×10 magnification (12). Cervical intraepithelial lesions are
enhanced and easily recognized when treated with acetic acid
solutions. For instance, areas that turn white following acetic
acid treatment (acetowhitening) and/or areas that present abnormal
vascular patterns are considered for biopsy. These effects become
more visible after a green filter is applied (13). Diagnoses are then evaluated by
gynecologists based on the degree of staining and the underlying
vascular patterns. Studies have attempted to classify images from
colposcopy using neural networks (14–16). For
instance, one group investigated whether neural networks could
recognize the dot pattern, which represents a colposcopy finding,
after learning the pattern from samples annotated by the
researchers (16). The present study
is distinct from the aforementioned studies because features or
representations of the images, for example, the presence of this
dot pattern, were not selected during data input.
For deep learning, the Keras neural network and
TensorFlow libraries were used (7,17). In the
present study, the classification accuracy on the validation
dataset reached ~50%. While this result in itself is not
satisfactory, it suggests that deep learning has the potential to
classify images from colposcopy. In addition, the present study
investigated methods to improve the learning rate and avoid
overfitting due to the limitation of insufficient numbers of
obtained images. In the process presented, L2 regularization, L1
regularization and dropout were applied, and the amount of input
data was increased via data augmentation.
In the present study, the intention was not to
stress the accuracy rate itself but rather to demonstrate that
gynecologists, who are not specialists in artificial intelligence
or machine learning, may be able to utilize deep learning in
clinical practice. Furthermore, the present results suggest that
relevant information from clinical practice should be appropriately
stored for future use.
Materials and methods
Patients
The present study was approved by the Institutional
Ethics Committee of Saitama Cancer Center (approval no. 630).
Written informed consent was obtained from all the patients.
Medical records and data from the gynecological oncology database
were retrospectively reviewed. Patients who underwent conization at
Saitama Cancer Centre (Ina, Japan) from January 2014 to December
2015 were enrolled. Conization management at the facility is
determined according to the guidelines of the Japan Society of
Obstetrics and Gynecology. Although each diagnosis was performed in
principle according to the postoperative pathology (conization),
the preoperative pathology (biopsy) was prioritized when the
results were severe and used as the output (‘target’ in deep
learning) for images from colposcopy.
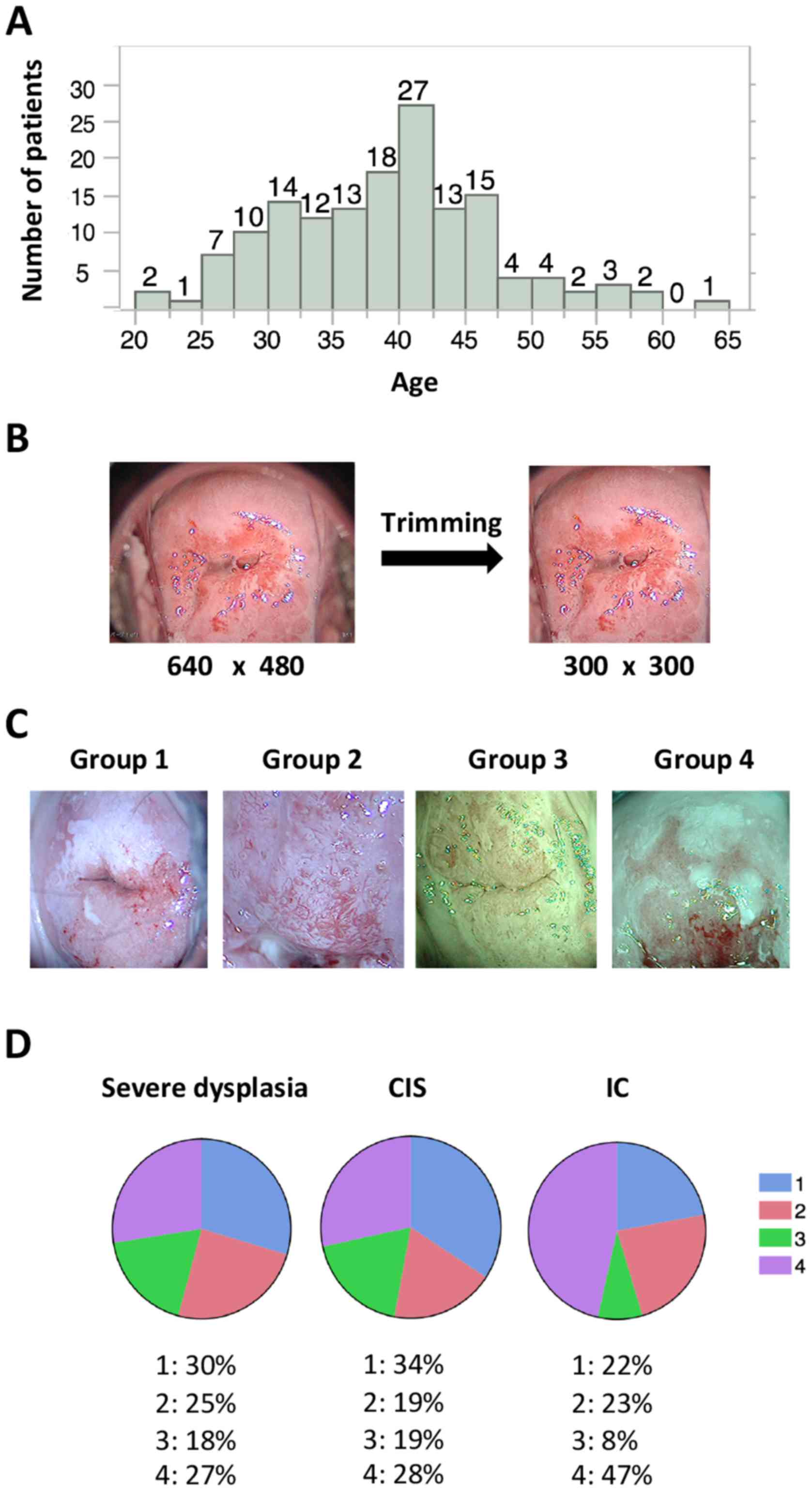
A total of 158 patients were enrolled; their median
age was 39 years (range, 21–63 years; Fig. 1A). The diagnoses and corresponding
patient numbers were as follows: severe dysplasia, 49; carcinoma
in situ (CIS), 78; invasive cancer, (IC) 21; and others
(such as adenocarcinoma in situ and invasive
adenocarcinoma), 10. In the current study, patient classification
was limited to three groups (severe dysplasia, CIS and IC) because
of the limited number of available images.
Images
Preoperative images from colposcopy were used as the
input data for deep learning. Because this investigation was a
retrospective study, there were no criteria for determining the
number and type of colposcopy images to retain. Images following
acetic acid treatment with or without a green filter that
represented areas of biopsy and were used in the diagnoses were
stored. The total number of images was 485, with 142 images for
severe dysplasia (2.9 images/patient), 257 for CIS (3.3
images/patient), and 86 for IC (4.1 images/patient). Of these, 233
images were captured with a green filter, and the remaining 252
were captured without a green filter.
Images from colposcopy captured at our facility were
stored in PNG format at a resolution of 640×480 pixels in RGB
3-channel color. These raw images often represented areas
inappropriate and unwanted for deep learning, such as the Cusco
speculum and vaginal wall; therefore, preprocessing was performed
to focus on the cervix by trimming the images to 300×300 pixels
(Fig. 1B). Trimming was performed
with Photoshop CC (Adobe Systems, Inc., San Jose, CA, USA). Images
without a green filter that captured >two thirds of the cervix
or <two thirds of the cervix (magnified images of the lesion)
were assigned to groups 1 and 2, respectively. Images with a green
filter that captured >two thirds of the cervix or <two thirds
of the cervix were assigned to groups 3 and 4, respectively
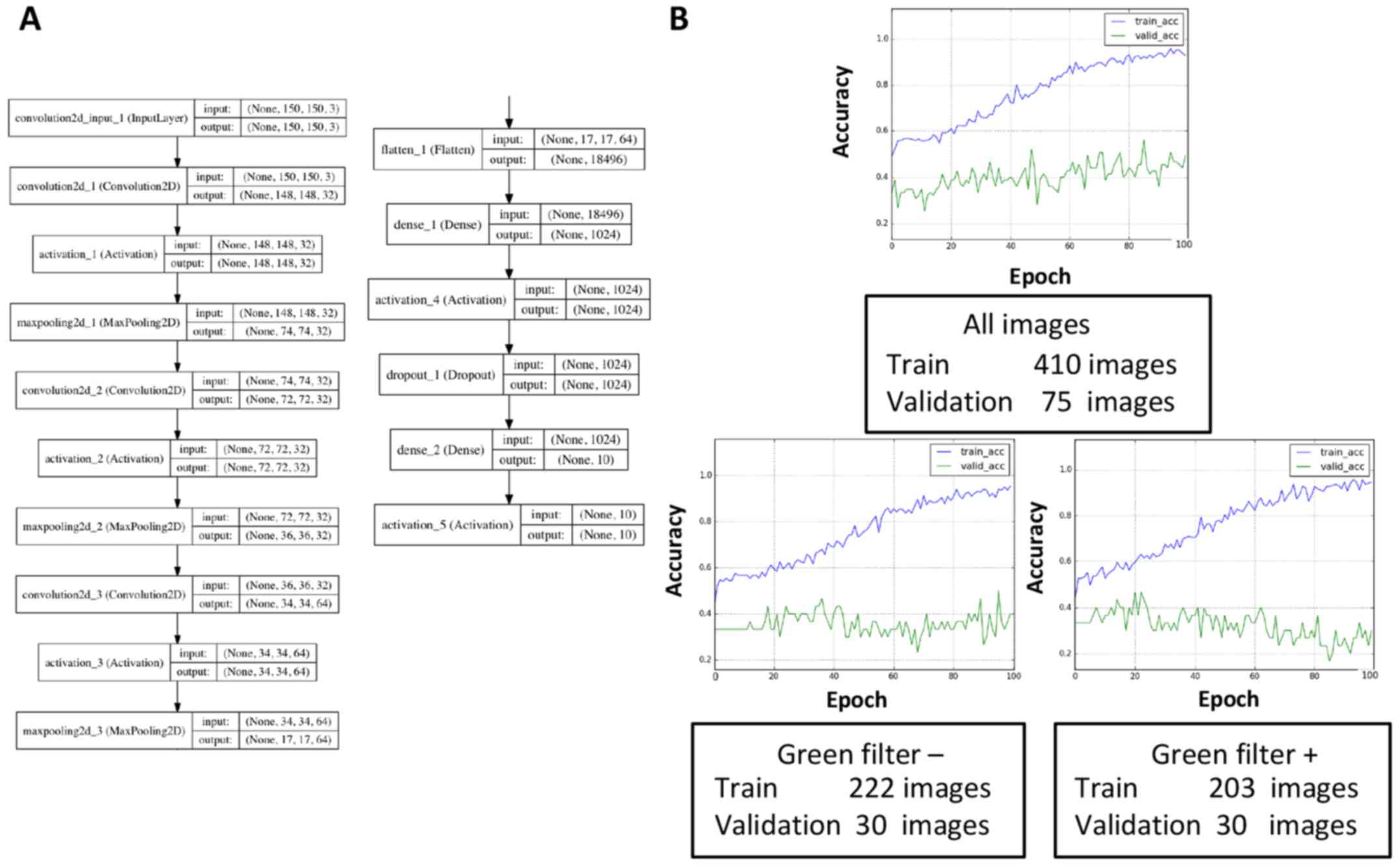
(Fig. 1C). During deep learning,
these images were re-trimmed to 150×150 pixels and then were used
as input (Fig. 2A). The same
procedures were performed with images containing 32×32 pixels or
300×300 pixels; however, images with 150×150 pixels were considered
suitable for learning in terms of the learning efficacy and time
allocated to learning (data not shown), at least in this
small-scale study.
Deep learning
We used the Keras (https://keras.io)
neural network library and the TensorFlow (https://www.tensorflow.org) software library. The code
was frequently referred to in the Keras blog (https://blog.keras.io/building-powerful-image-classification-models-using-very-little-data.html)
and the basic code was adjusted for the learning procedure of the
present study. The validation dataset contained 25 randomly
selected images for each diagnosis (75 images in total), and it was
not used for training in the study unless otherwise mentioned.
Development environment
The development environment used in the present
study was as follows: a Mac running OS X 10.11.3 (Apple, Inc.,
Cupertino, CA, USA); Python language v. 2.7.12; Keras 1.1.0;
TensorFlow 0.8.0; and matplotlib 1.5.3.
Statistical analysis
JMP Pro 11 (SAS Institute, Inc., Cary, NC, USA) was
used for the statistical analysis. One-way analysis of variance was
used for comparing the means. The Tukey-Kramer test was used for
post-hoc analysis. P<0.05 was considered to indicate a
statistically significant difference.
Results
Images
Preoperative images from colposcopy were
retrospectively collected as described in the Methods section.
Statistical analysis suggested that a higher number of images were
stored for more severe lesions (P=0.0085). The % of groups 1–4 were
summarized for each diagnosis (Fig.
1D). Unlike the images of severe dysplasia and CIS, the IC
images tended to include magnified lesions and were usually
captured with a green filter (Fig.
1D).
The total number of images is more important for
avoiding overfitting than dividing the input images according to
the presence or absence of a green filter. A validation set
accuracy of <33% meant that learning did not occur because the
same number of images for each diagnosis was assigned to the
validation dataset. The convolution layers and dense layers were
tuned as described in the Methods section (Fig. 2A). In the present study, dense layers
appeared to affect the learning rates, and a training accuracy that
exceeded 90% in 100 epochs was obtained by tuning the dense layers.
However, the validation accuracy plateaued at ~40–50%, which
suggested that overfitting had occurred. Therefore, methods of
avoiding overfitting to prevent discrepancies between the training
curve and validation curve were explored. First, the set of
collected images included images both with and without a green
filter, and these images were individually used for learning
because of possible learning inefficiencies caused by mixed data
(Fig. 2B). However, the validation
dataset was re-selected (10 images for each diagnosis, 30 in
total), and the results demonstrated that the validation accuracy
was reduced regardless of the presence or absence of a green
filter. This result was likely related to a reduction in the total
number of images. Thus, the total number of images appeared to be
more important for increasing the validation accuracy than the
division of input data according to the presence or absence of
green a filter, at least in the present small-scale study.
L2 regularization can improve
overfitting
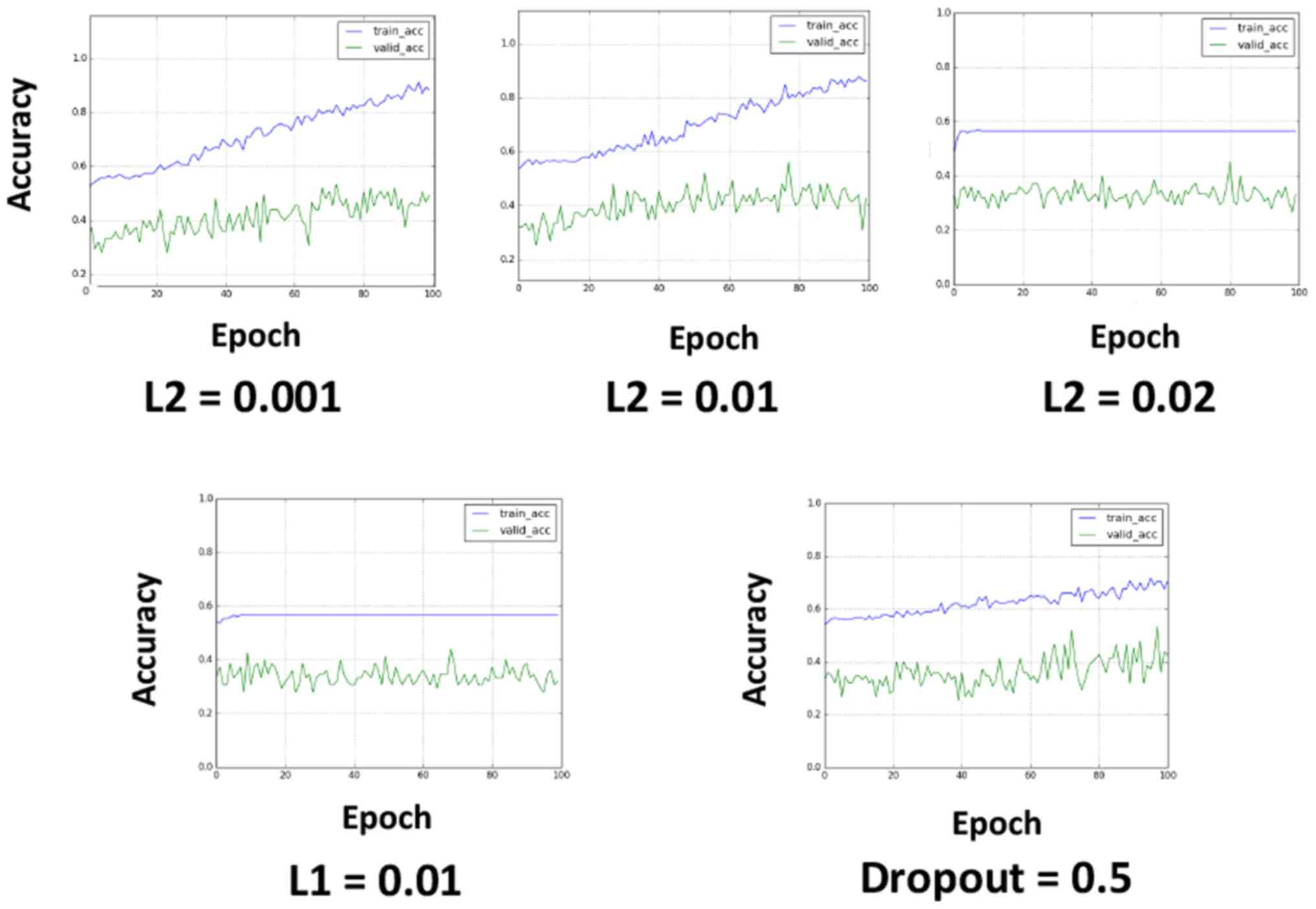
To avoid overfitting, L2 regularization, L1
regularization and dropout were applied (Fig. 3). L2 regularization and L1
regularization were applied in the first input layer, and dropout
was applied to all the layers after max-pooling (the dropout rate
was set to 0.5). L2 regularization appeared to be effective at
avoiding overfitting when properly tuned (Fig. 3). L1 regularization caused learning
failure in the investigated value set, and the application of
dropout reduced the learning efficacy, although this result may
have been related to the relatively short epochs.
Data augmentation slightly improves
the validation accuracy and overfitting
When performing deep learning, the total number of
images is known to be an important factor for improving the
learning accuracy and avoiding overfitting, which is generally
consistent with the aforementioned results (5). Thus, the hypothesis that data
augmentation could improve the accuracy rates was investigated.
When viewing colposcopy images, tilted images were occasionally
encountered because the angles of the cervix relative to the camera
posture varied; however, differences in the angle should not be
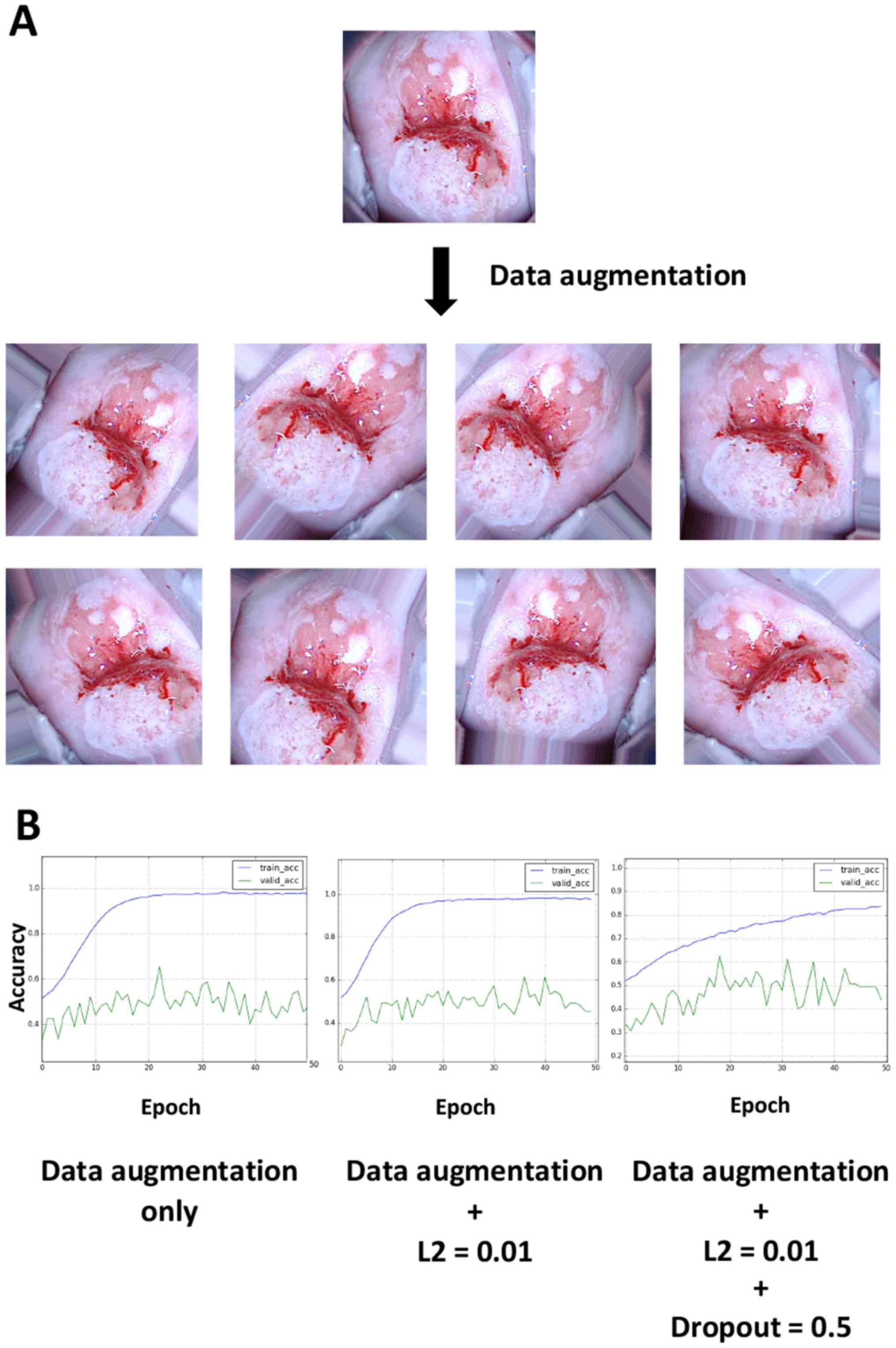
important for recognizing lesions. To test this hypothesis, 20
images were obtained from a single image by randomly rotating it,
applying different zoom magnifications and horizontally flipping
the image and the resulting images were then used as input. Thus,
one image was converted into ~20 images by data augmentation.
Examples of the augmentation results are illustrated in Fig. 4A. Data augmentation appeared to worsen
the overfitting limitation; however, the application of L2
regularization and dropout slightly improved overfitting (Fig. 4B). The validation accuracy ultimately
reached a stable level of ~50%.
Discussion
The present study investigated whether deep learning
could be applied to the classification of images from colposcopy.
Various types of data are increasingly available through the
Internet, and inexpensive high-end smartphones are more readily
available for the general public, facilitating the uploading and
sharing of information, such as pictures. The same is true for data
processing. High-performance personal computers are affordable for
individuals, and statistical analyses or machine learning can be
performed without supercomputers if the information volume is
limited. Furthermore, deep learning technologies are becoming more
accessible for corporations and individuals. For example, the
Google software library for machine learning, TensorFlow, was
released under an open-source license in 2015 (7). Based on these trends, the present study
aimed to apply deep learning to gynecological clinical
practice.
Preoperative images from colposcopy were
retrospectively collected. A total of 485 images were obtained,
with 142 images for severe dysplasia (2.9 images/patient), 257 for
CIS (3.3 images/patient), and 86 for IC (4.1 images/patient). These
results indicate that more images were stored when the lesions were
more severe (P=0.0085), because gynecologists tend to capture a
higher number of images in lesions in order to record important
findings. Accordingly, the IC images tended to include lesions
under greater magnification. Furthermore, these images were more
frequently captured with a green filter compared with the severe
dysplasia and CIS images (Fig.
1D).
One of the greatest challenges associated with
machine learning, including deep learning, is the prevention of
overfitting (5). Overfitting is a
condition in which the model cannot be applied to unknown data
because it has been overly adjusted to the training data. In the
present study, the large discrepancy between the training curve and
validation curve suggests that overfitting occurred (Figs. 2–4),
most likely due to the small number of included images. Ordinarily,
500–1,000 images are prepared for each class during image
classification with deep learning (2). The present study explored methods to
improve the learning rate and avoid overfitting under the
limitation of an insufficient number of included images. In
addition, L2 regularization, L1 regularization and dropout were
applied, and the amount of input data was increased by data
augmentation.
In clinical practice, it would be of interest for
clinicians to distinguish CIN1, CIN2 and CIN3, or to distinguish
CIN1 (or low-grade squamous intraepithelial lesions) from CIN2 (or
high-grade squamous intraepithelial lesions). Furthermore,
classification might not be clinically necessary for severe
dysplasia and carcinoma in situ (CIS), because there is
little difference in diagnosis and treatment between these two
conditions. However, due to technical reasons, the present
preliminary study used images from CIN3 and invasive cancer
patients. In terms of deep learning, the output (i.e., the
ground-truth classification result of an image) is very important.
For instance, what is considered CIN1 may not always be ‘genuine’
CIN1 because only a biopsy is performed in most cases. Ideally,
conization should be performed to provide the true answer. In
addition, providing ground-truth classifications such as ‘white
lesion’ or ‘glandular opening’ would not construct a reliable
model, because those answers are subject to human perception: There
is no strict definition for these terms. As such, for this initial
study, images from CIN3 and invasive cancer patients were used,
because pathological diagnoses of the conization samples were
readily obtained. For clinicians, an improved solution would be to
use the ‘patient's prognosis’ as an output. A clinical application
designed to screen a target group not in need of invasive testing,
such as biopsy and conization could be desirable as well.
Therefore, although the clinical significance of the classification
into three groups (severe dysplasia, CIS and IC) is currently
limited, the present study demonstrated that deep learning, by
inputting only images, could be used to classify colposcopy
images.
The final validation accuracy was ~50%, which is
better than a random result (33%). To the best of our knowledge, no
report using ‘deep learning’ for classification of images from
colposcopy exists to date. Although previous studies have used
automation diagnosis, deep learning was not employed and the
patient cohort was different than the current study (CIN3 and
invasive cancer) (14–16). As such, the present study cannot be
directly compared to the previous literature, in order to evaluate
whether the 50% accuracy result was good or poor. However, although
this result may not be satisfactory in terms of deep-learning
tasks, it may be sufficient in the clinic considering the
difficulty of distinguishing among severe dysplasia, CIS and IC
even among experts.
The present study aimed, not to stress the accuracy
rate itself, but rather to demonstrate that gynecologists, who are
not specialists in artificial intelligence or machine learning, can
utilize deep learning in clinical practice. The barriers to using
artificial intelligence and deep learning will likely be decreased
in the near future. Thus, as much relevant clinical information as
possible should be stored appropriately for future use. For
instance, the images used in this study contained as few as 150×150
pixels and 3 RGB channels. Images of this size could be obtained by
most users, even those using smart phones. Our facility is a cancer
center, and the patient population could have been biased in terms
of disease conditions because those who require operations or
intensive observation tend to be referred to our hospital.
Consequently, the collected images could also be biased.
Furthermore, as mentioned above, there were many more images taken
of lesions considered to be more severe. Therefore, the same
proportion of images is likely stored at many other facilities,
regardless of the diagnosis and the apparent severity of the
lesion.
The current study investigated a method for applying
deep learning to colposcopy image classification. The accuracy on
the final validation dataset reached ~50%. Although this result is
preliminary, it suggests that clinicians and researchers, who are
not specialists in artificial intelligence or machine learning, can
utilize deep learning. Furthermore, these findings suggest that as
much relevant clinical practice information as possible, including
colposcopy data and images, should be stored for future use.
References
|
1
|
Schmidhuber J: Deep learning in neural
networks: An overview. Neural Netw. 61:85–117. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Esteva A, Kuprel B, Novoa RA, Ko J,
Swetter SM, Blau HM and Thrun S: Dermatologist-level classification
of skin cancer with deep neural networks. Nature. 542:115–118.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Janowczyk A and Madabhushi A: Deep
learning for digital pathology image analysis: A comprehensive
tutorial with selected use cases. J Pathol Inform. 7:292016.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Rajkomar A, Lingam S, Taylor AG, Blum M
and Mongan J: High-throughput classification of radiographs using
deep convolutional neural networks. J Digit Imaging. 30:95–101.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Yu L, Chen H, Dou Q, Qin J and Heng PA:
Automated melanoma recognition in dermoscopy images via very deep
residual networks. Ieee Trans Med Imaging. 36:994–1004. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Zhang YC and Kagen AC: Machine learning
interface for medical image analysis. J Digit Imaging. 30:615–621.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Rampasek L and Goldenberg A: Tensorflow:
Biology's gateway to deep learning? Cell Syst. 2:12–14. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Ginsburg O, Bray F, Coleman MP, Vanderpuye
V, Eniu A, Kotha SR, Sarker M, Huong TT, Allemani C, Dvaladze A, et
al: The global burden of women's cancers: A grand challenge in
global health. Lancet. 389:847–860. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Cook DA, Smith LW, Law J, Mei W, van
Niekerk DJ, Ceballos K, Gondara L, Franco EL, Coldman AJ, Ogilvie
GS, et al: Aptima HPV assay versus hybrid capture® 2 HPV
test for primary cervical cancer screening in the HPV FOCAL trial.
J Clin Virol. 87:23–29. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Coste J, Cochand-Priollet B, de Cremoux P,
Le Galès C, Cartier I, Molinié V, Labbé S, Vacher-Lavenu MC and
Vielh P; French Society of Clinical Cytology Study Group, : Cross
sectional study of conventional cervical smear, monolayer cytology
and human papillomavirus DNA testing for cervical cancer screening.
BMJ. 326:7332003. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
den Boon JA, Pyeon D, Wang SS, Horswill M,
Schiffman M, Sherman M, Zuna RE, Wang Z, Hewitt SM and Pearson R:
Molecular transitions from papillomavirus infection to cervical
precancer and cancer: Role of stromal oestrogen receptor
signalling. Proc Natl Acad Sci USA. 112:pp. E3255–E3264. 2015;
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Garcia-Arteaga JD, Kybic J and Li W:
Automatic colposcopy video tissue classification using higher order
entropy-based image registration. Comput Biology Med. 41:960–970.
2011. View Article : Google Scholar
|
|
13
|
Balas C: A novel optical imaging method
for the early detection, quantitative grading and mapping of
cancerous and precancerous lesions of cervix. IEEE Trans Biomed
Eng. 48:96–104. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Acosta-Mesa HG, Cruz-Ramirez N and
Hernández-Jiménez R: Aceto-white temporal pattern classification
using k-NN to identify precancerous cervical lesion in colposcopic
images. Comput Biol Med. 39:778–784. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Acosta-Mesa HG, Rechy-Ramirez F,
Mezura-Montes E, Cruz-Ramirez N and Hernández Jiménez R:
Application of time series discretization using evolutionary
programming for classification of precancerous cervical lesions. J
Biomed Inform. 49:73–83. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Simoes PW, Izumi NB, Casagrande RS, Venson
R, Veronezi CD, Moretti GP, da Rocha EL, Cechinel C, Ceretta LB,
Comunello E, et al: Classification of images acquired with
colposcopy using artificial neural networks. Cancer Infor.
13:119–124. 2014. View Article : Google Scholar
|
|
17
|
Medved D, Nugues P and Nilsson J:
Predicting the outcome for patients in a heart transplantation
queue using deep learning. Conf Proc IEEE Eng Med Biol Soc.
2017:pp. 74–77. 2017; PubMed/NCBI
|