Introduction
Primary liver cancer (including hepatocellular
carcinoma and intrahepatic cholangiocarcinoma) is the second cause
of cancer related death and one of the cancers with still
increasing incidence rate (1,2). Hepatocellular carcinoma, a major primary
liver cancer pathological type, shows very low 5-year survival rate
(<15%) owing to its late diagnosis and compromised underlying
liver function (3,4). Although surgical resection or
transplantation helps to improve the survival rate of patients,
there is still no effective treatment for advanced patients who are
ineligible for surgery, resulting in only a median overall survival
of 6.6 months for these patients (5,6).
Therefore, it is imperative to explore effective and safe
prognostic biomarkers for early diagnosis and therapeutic targets
to improve treatment strategies.
Autophagy is a dynamic degradation process for
delivering dysfunctional cellular components or foreign invaders to
lysosome to be digested by lysosomal hydrolase (7–9).
Autophagy-related gene (ATG) products play important roles in
autophagy. Autophagosome formation is mediated by two
ubiquitin-like conjuction systems composed of ATG proteins, which
culminate in conjugation of ATG12 to ATG5 and conversion of a
soluble form of microtubule-associated protein 1 light chain 3
(LC3-I) to phosphatidylethanolamine-conjugated membrane-bound form
(LC3-II) (10,11). Beclin1 is a component of the class III
phosphoinositide 3-kinase complex and also plays an essential role
in autophagy regulation. A considerable body of studies have shown
that autophagy plays a dual-side role in cancer pathogenesis. In
the initiation of tumor, autophagy may inhibit tumor formation by
degradation of damaged organelles or proteins (12). After the formation of tumor, however,
the tumor can utilize the autophagy as a survival mechanism to
counter metabolic stress (13).
Additionally, recent evidence has demonstrated that
autophagy is linked to increased tumor cell invasion because tumor
cell invasion requires the production of pro-migratory cytokines
mediated by autophagy (14,15). How autophagy activity influence cancer
metastasis remains to be determined.
It is known that heat shock protein 90 (HSP90)
chaperones are pivotal regulators of proteostasis in the
mitochondria of selective tumor cells. TNF receptor-associated
protein-1 (TRAP1), as a mitochondrial molecular chaperone of the
Hsp90 family, has attracted attention because of its high
expression in various tumors but low expression in normal tissues
(16,17). HSP90 chaperones can work as an
upstream connector between the control of protein folding and
adaptive mechanisms of autophagy and cancer metabolism. Some recent
studies have shown that the TRAP1 expression level and cellular
mitochondria autophagy are closely related (18–21).
However, the relationship between TRAP1 expression and autophagy in
liver cancer is still unknown.
In the present study, we aim to explore the effect
of rapamycin, invasive ability and hepatitis B (HBV) infection on
the level of autophagy and TRAP1 expression in four different liver
cancer cell lines and the relationship between autophagy and TRAP1
in liver cancer.
Materials and methods
Reagents
The following reagents were used: EMDM (Gibco;
Thermo Fisher Scientific, Inc., Waltham, MA, USA); FBS (Gemini Bio
Products, West Sacramento, CA, USA); TRIzol (Ambion; Thermo Fisher
Scientific, Inc.); MDC (Tiangen Biotech Co., Ltd., Beijing, China);
WIP organization cell lysis solution (Boaoseng, Beijing, China);
RNase Away (Ed lai, Beijing, China); BCA protein concentration
determination kit (Blue skies, China); SDS-PA (Nanjing KeyGen
Biotech. Co. Ltd., Nanjing, China); Reverse transcription kit
(Lisu, Shanghai, China); Primers (Hongxun, Suzhou, China); High
sensitive chemiluminescence detection kit (Qihaifutan, Shanghai,
China); TRAP1 Rabbit mAb (Cell Signaling Technology, Inc., Danvers,
MA, USA); Beclin1 Rabbit mAb (Cell Signaling Technology, Inc.); LC3
Rabbit mAb (Cell Signaling Technology, Inc.); Anti-rabbit IgG,
HRP-linked Antibody (Isotype: Goat; Cell Signaling Technology,
Inc.); Mouse anti β-actin mAb (Zhongshanjinqiao, China);
Anti-β-actin Mouse Monoclonal Antibody (Isotype: Goat;
Zhongshanjinqiao, China); and Protein Simple (San Jose, CA,
USA).
Cell lines and culture
Four liver cancer cell lines (Hep3B2.1–7, Sk-hep1,
HepG2 and HepG2.2.15 cell lines) were obtained from the Chinese
Academy of Sciences Library. All cell lines were cultured in EMDM
medium containing 10% FBS and were maintained at 37°C in 95% air
and 5% CO2.
RNA isolation and reverse
transcription
Total RNA was isolated from four liver cancer cell
lines with TRIzol according to manufacturer's instructions. After
the RNA concentration was measured, the integrity of the isolated
RNA was analyzed (NanoDrop 2000; Thermo Fisher Scientific, Inc.), 3
µg of RNA was reverse transcribed into cDNA according to the
manufacturer's protocol.
Reverse transcription-quantitative
polymerase chain reaction (RT-qPCR)
RT-qPCR was performed using SYBR-Green Supermix. The
reaction conditions were as follows: 95°C for 10 sec and 60°C for
32 sec, 40 cycles. Target genes were normalized to β-actin and were
quantified using the comparative 2−∆∆Cq method. TRAP1
expression levels were measured in triplicate, and the average was
calculated. β-actin was applied as an internal control. The primers
for β-actin (96 bp) were 5′-GATCAGCAGCAGAGTATG-3′ (F) and
5′-AGAGTGTACGCACTA-3′ (R). The primers for TRAP-1 (112 bp) were
5′-CAGGGTTCCACTTCCAAACA-3′ (F) and 5′-TGGAGATCAGCTCCCGTATAA-3′
(R).
Rapamycin-induced cell autophagy
A total of 3×105/ml cells in the
logarithmic growth phase were inoculated in a round-bottom 6-well
plate. The cells were exposed to rapamycin at 0, 5, 10, and 15
µmol/l for 10, 15, 20, 25 and 30 h. Next, the cells were washed
three times with PBS, dyed for 30 min (MDC) in the dark, and washed
three times with 1× wash buffer. Finally, infiltrating cells were
obtained with collection buffer. The cells were observed and
photographed by using a fluorescence microscope (Eclipse Ti-U;
Nikon Corporation, Tokyo, Japan). We found that activation of
autophagy was highest when the cells were induced by rapamycin for
20 h. Thus, we treated the cells with rapamycin for 20 h in the
present study.
Western blot analysis
Thirty micrograms of protein samples from each case
was separated with 10% sodium dodecyl sulfate-polyacrylamide gel
electrophoresis and were subsequently transferred to polyvinylidene
fluoride membranes. After transfer, the membranes were incubated
with blocking buffer of 5% non-fat dry milk in Tris-buffered saline
containing Tween-20 at room temperature for 1 h. The membranes were
incubated with primary antibodies (1:1,000 dilution) for 20 h at
4°C. After washing, secondary antibodies (1:5,000 dilution) were
added at room temperature for 1 h. Finally, ECL development and
fixing were considered.
Protein Simple western analysis
Protein levels were quantified by using WES™
(Protein Simple), an automated capillary-based size sorting system
(22). Protein samples and 5×
Fluorescent master were mixed in a micro-centrifuge tube (final
concentration, 0.2 mg/ml); then, the samples were denatured, and a
biotinylated ladder was used (95°C, 5 min). The primary antibody
was diluted 1:50 in Antibody Diluent II before use. The secondary
antibody was used without dilution (Protein Simple). Next, 150 µl
of Luminol-S and 150 µl of peroxide were combined in a
micro-centrifuge tube. The samples, wash buffer, primary
antibodies, secondary antibodies, blocking reagent, and
chemiluminescent substrate were dispensed into designated wells in
the provided microplate. Following plate loading, separation and
immunodetection were performed automatically using default
settings. The incubation time of the primary antibody was 60 min
and that of the secondary antibodies was 30 min. The data were
analyzed using inbuilt Compass software (Protein Simple).
Cell growth curve
Overall, 1×104/ml cells in the
logarithmic growth phase were inoculated in a round-bottomed 6-well
plate. Starting on the second day, the cells with pancreatic enzyme
digestion and count three holes every day, averaged, and counted
for five consecutive days.
Statistical analysis
All P-values were two sided, and P<0.05 was
considered to indicate a statistically significant difference. The
experimental data were expressed as the mean ± standard deviation,
and analysis of variance with an LSD t-test was used to assess the
relationship between two groups. Statistical analyses were
performed using the SPSS v20.0 software (SPSS, Inc., Chicago, IL,
USA).
Results
Cell invasive ability, HBV infection
and rapamycin influenced cell growth
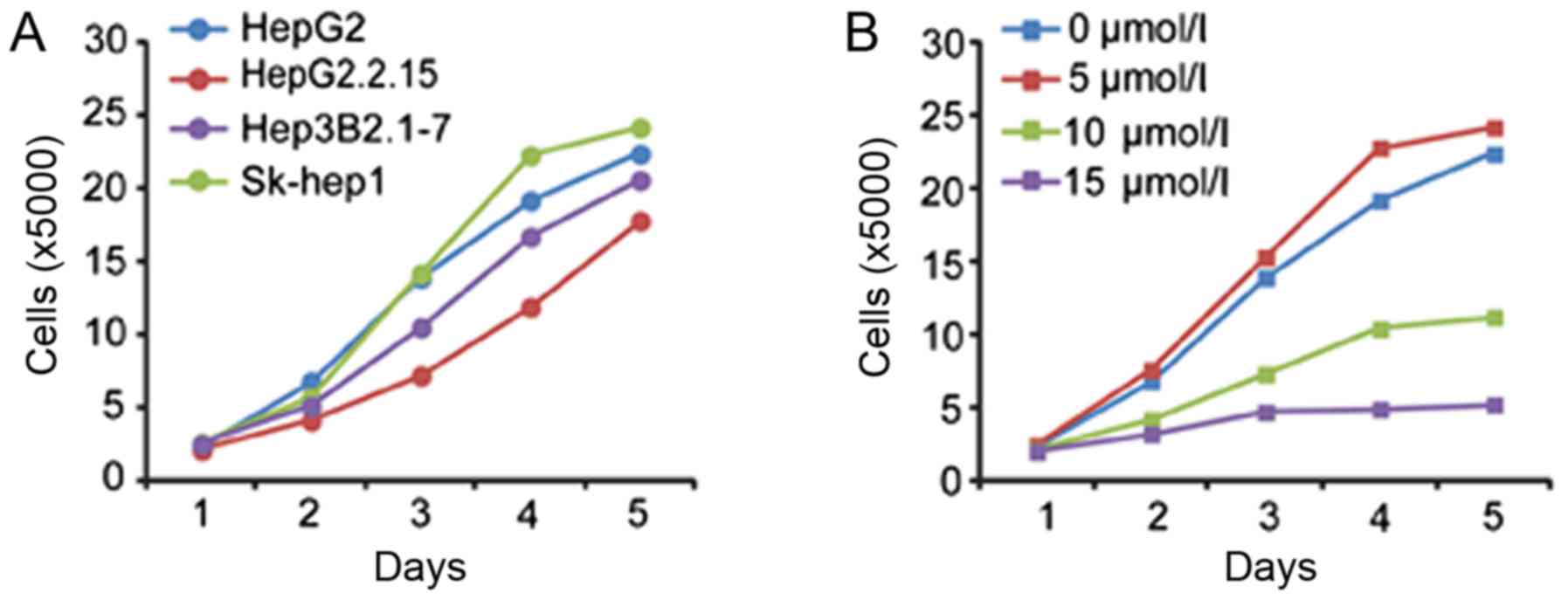
Herein, cell growth was examined in four liver
cancer cell lines. Hep3B2.1–7, HepG2 and Sk-hep1 cells, which
exhibit growing invasive ability, showed increasing cell
proliferation ability, which indicated that cell invasive ability
was positively related to cell growth. Additionally, the growth
rate of HepG2.2.15 cells was slower when compared with HepG2 cells
(P<0.05), suggesting that HBV infection inhibited the growth of
cells (Fig. 1A).
We also examined the effect of rapamycin on the
growth of HepG2 cells at 0, 5, 10 and 15 µmol/l for 20 h. As a
result, the growth rate of HepG2 cells was inhibited under the
treatment of 10 and 15 µmol/l rapamycin (P<0.05). The results
indicated that rapamycin exposure could inhibit the growth of HepG2
cells (Fig. 1B).
Basal level of autophagy was affected
by cell invasive ability and HBV infection
The basal level of autophagy was examined in four
liver cancer cell lines. Beclin1 and LC3-II/LC3-I are two specific
autophagy-related markers and their protein level were detected by
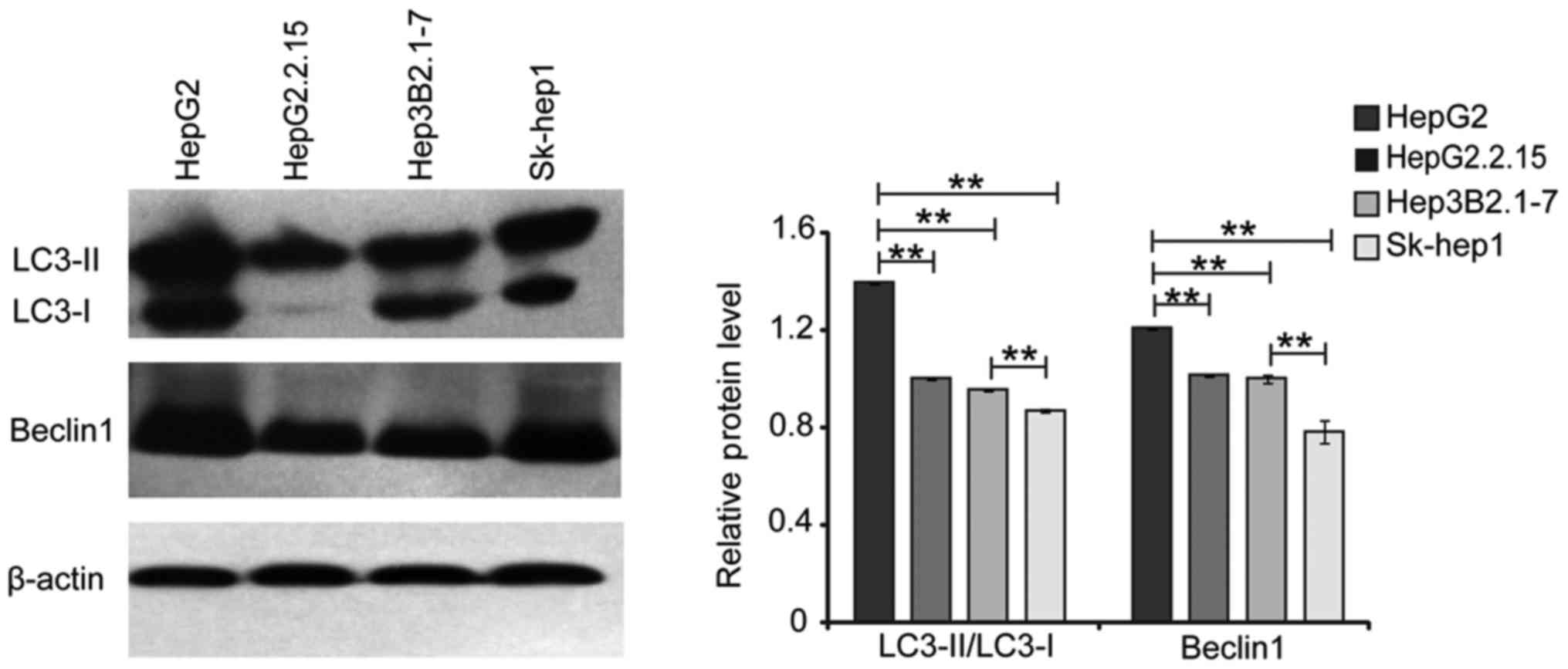
western blot. As shown in Fig. 2, the
expression level of Beclin1 and LC3-II/LC3-I was changed in the
same tendency among HepG2, Hep3B2.1–7, Sk-hep1 and HepG2.2.15
cells. HepG2 cell line, with medium invasive ability, showed the
highest expression level of Beclin1 and LC3-II/LC3-I. However, the
expression level of Beclin1 and LC3-II/LC3-I was lowest in the most
invasive Sk-hep1 cell line. These findings suggested that the basal
level of autophagy might be affected by cell invasive ability.
Moreover, when compared to HepG2 cells, the protein level of
Beclin1 and LC3-II/LC3-I were significantly decreased in
HBV-infected HepG2.2.15 cells, indicating that the HBV infection
might inhibit basal level of cell autophagy.
Rapamycin triggered autophagy in four
liver cancer cell lines
Rapamycin is a mTOR-independent autophagy inducer.
Four liver cancer cell lines were further treated with indicated
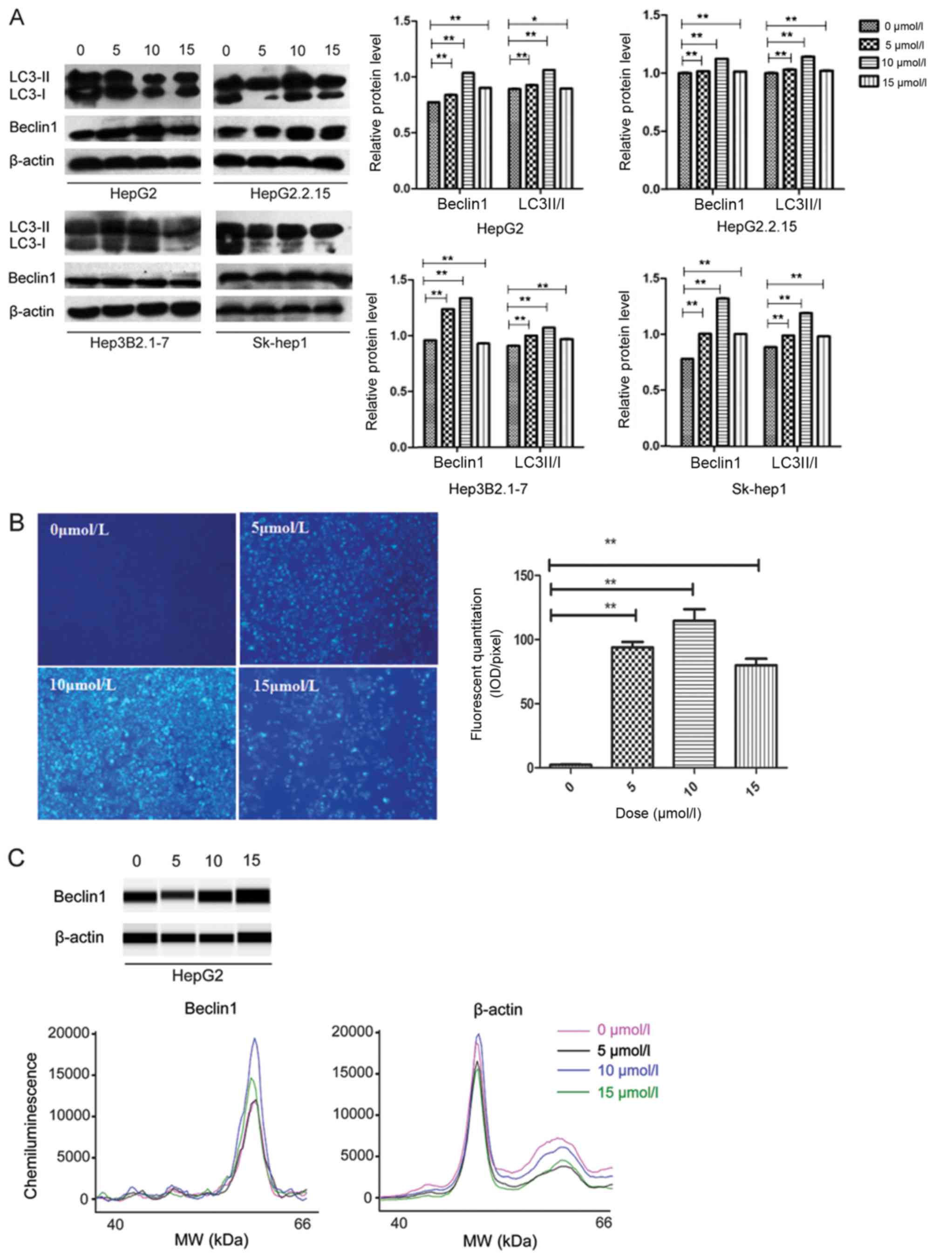
concentration of rapamycin for 20 h. As shown in Fig. 3A, compared with control group, the
expression levels of Beclin1 and LC3-II/LC3-I were significantly
increased in rapamycin-treated cells (P<0.05). In addition, the
highest expression levels were shown in four liver cancer cell
lines under the treatment of 10 µmol/l rapamycin. These results
were further confirmed in HepG2 cells by using MDC method and
Protein Simple Western. The MDC-positive cells and the protein
level of Beclin1 were obviously increased in HepG2 cells after
treated with rapamycin and peaked at 10 µmol/l (Fig. 3B and C). Taken together, these
observations reveal that the autophagy could be triggered by
rapamycin in four liver cancer cells.
TRAP1 expression level was associated
with cell autophagy
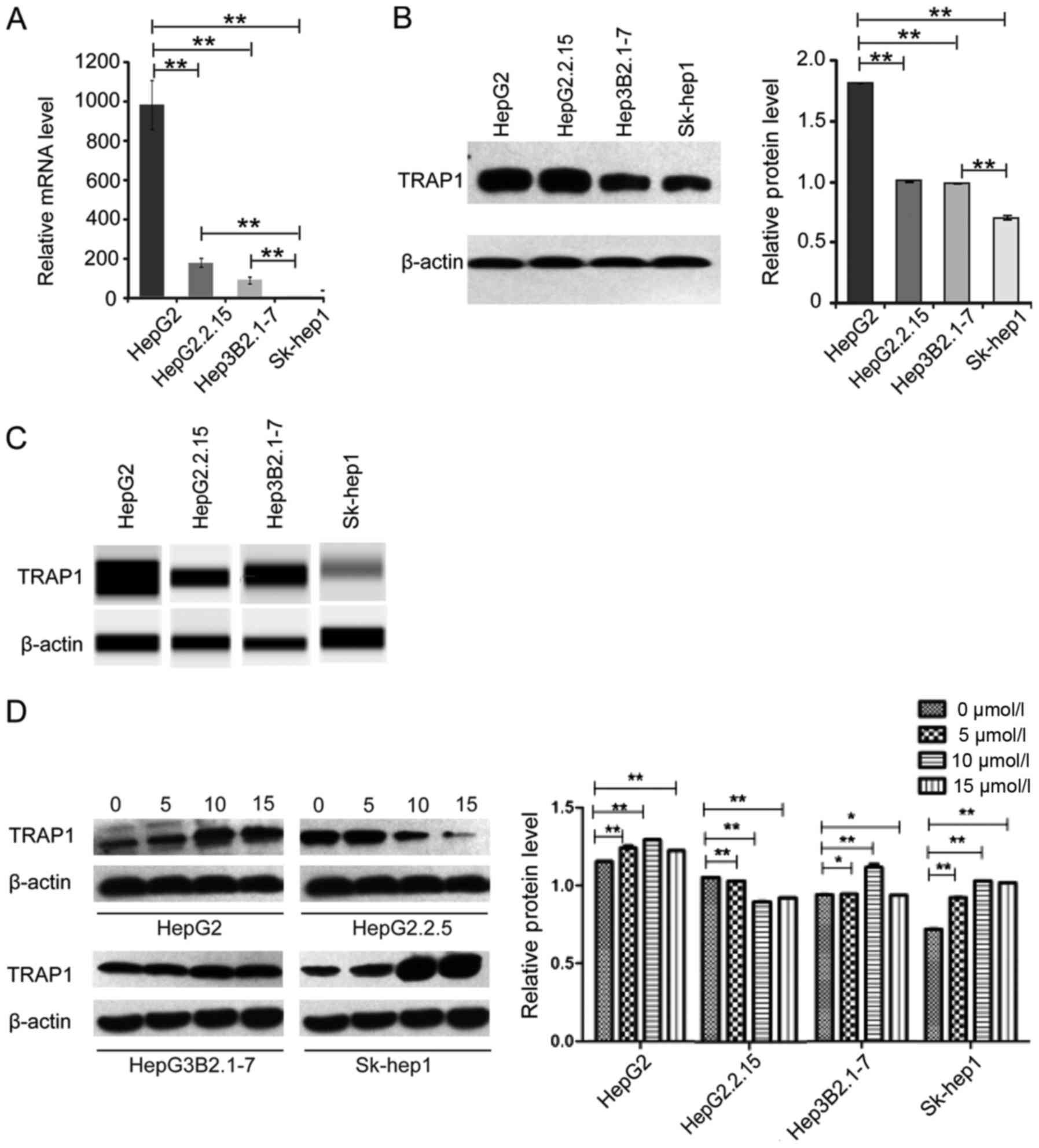
We further examined the mRNA and protein level of
TRAP1 in four liver cancer cell lines. The expression level of
TRAP1 was measured by RT-qPCR, western blot and Protein Simple
Western. Intriguingly, a similar change of TRAP1 expression and
autophagy activity was observed in four liver cancer cell lines
with or without rapamycin induction. In detail, the expression
level of TRAP1 was quite different in HepG2, Hep3b2.1–7 and Sk-hep1
cells, which representing different invasive ability. And the
highest expression level of TRAP1 was observed in HepG2 cells which
presented highest basal autophagy activity. Moreover, similar to
autophagy activity, the TRAP1 expression was significantly
inhibited in HepG2.2.15 cells (Fig.
4A-C). Furthermore, the TRAP1 expression level significantly
enhanced in four rapamycin-treated liver cancer cell lines except
HepG2.2.15 cells. And the TRAP1 expression peaked under the
treatment of 10 µmol/l in HepG2, Hep3B2.1–7 and Sk-hep1 cells.
Taken together, rapamycin might induce autophagy and TRAP1
expression in liver cancer cells, and the cell autophagy and TRAP1
expression might be positively correlated.
Discussion
In this study, we provided evidence that cell
invasive ability, rapamycin and HBV infection were factors that
affected the level of autophagy and TRAP1 expression in four liver
cancer cell lines. Additionally, TRAP1 expression was positively
correlated with the cell autophagy level, suggesting that TRAP1 and
cell autophagy might be closely related.
Primary liver cancer patients have a high recurrence
and mortality; however, they remain the main factors of poor
prognosis (23). Liver cancer
development is a multi-stage and multi-factor process, involving in
multiple gene interactions and cell signaling pathways. Currently,
specific clinical biomarkers are still not available, which has
leaded to low early diagnosis. Increasing studies have identified
the crucial role of autophagy in liver diseases (24). The dysregulation of autophagy is
associated with viral hepatitis, non-alcoholic fatty liver disease,
alcoholic liver disease, fibrosis, cirrhosis, and hepatocellular
carcinoma (HCC) (25–28).
In the present study, we detected the cell growth of
liver cancer cell lines with different invasive abilities, HBV
infection and rapamycin exposure. The results showed that highly
invasive Sk-hep1 cells grew faster than HepG2 cells and Hep3B2.1–7
cells. HepG2.2.15 cells, which were infected with HBV, grew slower
than HepG2 cells. These results indicated that cell invasive
ability might positively correlated with cell growth, and HBV
infection might inhibit the growth of liver cancer cells. However,
the potential mechanism remains to be determined. Rapamycin, a
small-molecule material, is currently used to induce autophagy. We
found that the growth rate of HepG2 cells was inhibited when
exposed to 10 and 15 µmol/l rapamycin. The results demonstrated
that high cell autophagy activity may inhibit cell growth. Some
studies have shown that autophagy in cancer cells such as gastric
cancer and liver cancer cells serves to ‘maintain survival’ and
‘induce death’, two opposing functions so autophagy could play a
bidirectional role in those processes.
HBV infection is the most important cause of liver
cancer (29). Some studies have found
that HBV has a close relationship with cell autophagy (30–32).
Moreover, TRAP1 expression is reported to associate with cell
autophagy (33). To understand more
about the pathogenesis of liver cancer, the expression level of
Beclin1, LC3-II/LC3-Iand TRAP1 in four liver cancer cell lines was
investigated by using RT-qPCR, western blot and Protein Simple
Western. The results showed that the expression level of TRAP1,
Beclin1 and LC3-II/LC3-I was highest in HepG2 cells and lowest in
Sk-hep1 cells, suggesting that higher invasive ability and HBV
infection could inhibit autophagy in liver cancer cell lines.
Additionally, the expression level of TRAP1 showed a positive
correlation with the autophagy level. The expression levels of
Beclin1, LC3-II/LC3-I and TRAP1 was concentration-dependently
increased in HepG2, Sk-hep1 and Hep3B2.1–7 cells after treated with
0, 5 and 10 µmol/l rapamycin. However, the expression levels of
Beclin1, LC3-II/LC3-I and TRAP1 were reduced when these cells were
exposed to 15 µmol/l rapamycin. The expression level of TRAP1 was
positively correlated with autophagy activity in HepG2, Sk-hep1 and
Hep3B2.1–7 cells. Interestingly, the TRAP1 level in HepG2.2.15
cells exposed with 0, 5 and 10 µmol/l rapamycin was negatively
correlated with the rapamycin concentration, suggesting that the
HBV infection can inhibit the rapamycin-induced TRAP1 expression.
We found that the cell invasive ability, HBV infection and
autophagy level can influence TRAP1 expression. However, the
specific mechanism needs further study.
Cell autophagy is a biological process in eukaryotes
with lysosome degradation. According to the cell environment, level
of autophagy, and different roles of autophagy, it can be divided
into basal autophagy and induced autophagy. Basal autophagy is a
low-level cell autophagy activity that plays an important role in
updating intracellular substances and maintaining the cell
steady-state (34). Compared with
basal autophagy, the reaction level of induced autophagy is higher,
and the level of autophagy quickly climbs in a short time in the
absence of external nutrients, an energy supply or other
stimulation from the extracellular environment. An appropriate
window of induced autophagy can maintain cell survival, but
excessive induced autophagy can cause cell death (35). However, how autophagy switches from
‘maintain survival’ to ‘induce death’ is not fully clear. The
present study also found that inducing autophagy was positively
correlated with TRAP1 expression. Thus, TRAP1 may be one of the
triggers for the transformation. Additionally, it is thought that
TRAP1 is an important factor related to cancer progression and
prognosis. It can repair cell death with the properties of the
matrix protein, Cyclophilin-D. Several preclinical studies,
including in vivo studies in both normal and gene knock-out
animal models, had confirmed that TRAP1 was a novel and efficient
treatment target in various tumors. Although TRAP1 can maintain the
mitochondrial integrity and function by inhibiting the fold of
damaged proteins and promoting the refold process of denatured
proteins, TRAP1 has dual effects on the regulation of mitochondrial
apoptosis (36). Taken together,
these results demonstrated that TRAP1 might play different roles in
cell autophagy via the mitochondrial pathway. Thus, it is necessary
to explore the factors that can affect TRAP1 expression and cell
autophagy in liver cancer cell lines, and investigate the mechanism
underlying the ‘maintain survival’ and ‘induce death’, which
provide clues to improve the prognosis of liver cancer.
Acknowledgements
The authors would like to thank Miss Shan Wang
(Department of Toxicology, Guangzhou Key Laboratory of
Environmental Pollution and Health Risk Assessment, School of
Public Health, Sun Yat-sen University, Guangzhou, P.R. China) for
their assistance in revising the article.
Funding
The present study was supported financially by a
grant from the National Natural Science Foundation of China (grant
no. 81460446).
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
YS, HZ and YL were involved in the conception and
design of the study, the acquisition and interpretation of data,
drafting the article, and giving final approval of the version to
be published. LY, MZ and XS contributed towards the design of the
study. YY and WC analyzed and interpreted the data. YZ and JM
acquired the data, and revised the article. All authors read and
approved the final manuscript.
Ethics approval and consent to
participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Kassebaum NJ, Bertozzi-Villa A, Coggeshall
MS, Shackelford KA, Steiner C, Heuton KR, Gonzalez-Medina D, Barber
R, Huynh C, Dicker D, et al: Global, regional, and national levels
and causes of maternal mortality during 1990–2013: A systematic
analysis for the Global Burden of Disease Study 2013. Lancet.
384:980–1004. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Bosch FX and Ribes J: The epidemiology of
primary liver cancer: Global epidemiology. Perspect Med Virol.
6:1–16. 2002. View Article : Google Scholar
|
|
3
|
Altekruse SF, McGlynn KA and Reichman ME:
Hepatocellular carcinoma incidence, mortality, and survival trends
in the United States From 1975 to 2005. J Clin Oncol. 27:1485–1491.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Sapisochin G, Goldaracena N, Laurence JM,
Dib M, Barbas A, Ghanekar A, Cleary SP, Lilly L, Cattral MS,
Marquez M, et al: The extended Toronto criteria for liver
transplantation in patients with hepatocellular carcinoma: A
prospective validation study. Hepatology. 64:2077–2088. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Cheng AL, Thongprasert S, Lim HY,
Sukeepaisarnjaroen W, Yang TS, Wu CC, Chao Y, Chan SL, Kudo M,
Ikeda M, et al: Randomized, open-label phase 2 study comparing
frontline dovitinib versus sorafenib in patients with advanced
hepatocellular carcinoma. Hepatology. 64:774–784. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Zhu AX, Rosmorduc O, Evans TR, Ross PJ,
Santoro A, Carrilho FJ, Bruix J, Qin S, Thuluvath PJ, Llovet JM, et
al: SEARCH: A phase III, randomized, double-blind,
placebo-controlled trial of sorafenib plus erlotinib in patients
with advanced hepatocellular carcinoma. J Clin Oncol. 33:559–566.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Jing K and Lim K: Why is autophagy
important in human diseases? Exp Mol Med. 44:69–72. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Wen X, Wu J, Wang F, Liu B, Huang C and
Wei Y: Deconvoluting the role of reactive oxygen species and
autophagy in human diseases. Free Radic Biol Med. 65:402–410. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Yang Z and Klionsky DJ: Eaten alive: A
history of macroautophagy. Nat Cell Biol. 12:814–822. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Mizushima N: Autophagy: Process and
function. Genes Dev. 21:2861–2873. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Klionsky DJ, Cregg JM, Dunn WA Jr, Emr SD,
Sakai Y, Sandoval IV, Sibirny A, Subramani S, Thumm M, Veenhuis M
and Ohsumi Y: A unified nomenclature for yeast utophagy-related
genes. Dev Cell. 5:539–545. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Mathew R and White E: Why sick cells
produce tumors: The protective role of autophagy. Autophagy.
3:502–505. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
White E: Deconvoluting the
context-dependent role for autophagy in cancer. Nat Rev Cancer.
12:401–410. 2012. View
Article : Google Scholar : PubMed/NCBI
|
|
14
|
Kenerson HL, Yeh MM, Kazami M, Jiang X,
Riehle KJ, McIntyre RL, Park JO, Kwon S, Campbell JS and Yeung RS:
Akt and mTORC1 have different roles during liver tumorigenesis in
mice. Gastroenterology. 144:1055–1065. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Zeng H, Yang K, Cloer C, Neale G, Vogel P
and Chi H: mTORC1 couples immune signals and metabolic programming
to establish T(reg)-cell function. Nature. 499:485–490. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Costantino E, Maddalena F, Calise S,
Piscazzi A, Tirino V, Fersini A, Ambrosi A, Neri V, Esposito F and
Landriscina M: TRAP1, a novel mitochondrial chaperone responsible
for multi-drug resistance and protection from apoptotis in human
colorectal carcinoma cells. Cancer Lett. 279:39–46. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Choi H, Merceron C, Mangiavini L, Seifert
EL, Schipani E, Shapiro IM and Risbud MV: Hypoxia promotes
noncanonical autophagy in nucleus pulposus cells independent of
MTOR and HIF1A signaling. Autophagy. 12:1631–1646. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Caino MC, Chae YC, Vaira V, Ferrero S,
Nosotti M, Martin NM, Weeraratna A, O'Connell M, Jernigan D,
Fatatis A, et al: Metabolic stress regulates cytoskeletal dynamics
and metastasis of cancer cells. J Clin Invest. 123:2907–2920. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Siegelin MD: Inhibition of the
mitochondrial Hsp90 chaperone network: A novel, efficient treatment
strategy for cancer? Cancer Lett. 333:133–146. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Costa AC, Loh SH and Martins LM:
Drosophila Trap1 protects against mitochondrial dysfunction in a
PINK1/parkin model of Parkinson's disease. Cell Death Dis.
4:e4672013. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Chen JF, Wu QS, Xie YX, Si BL, Yang PP,
Wang WY, Hua Q and He Q: TRAP1 ameliorates renal tubulointerstitial
fibrosis in mice with unilateral ureteral obstruction by protecting
renal tubular epithelial cell mitochondria. FASEB J. 31:4503–4514.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Harris VM: Protein detection by Simple
Western™analysis. Methods Mol Biol. 1312:465–468. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Xiao S, Chang RM, Yang MY, Lei X, Liu X,
Gao WB, Xiao JL and Yang LY: Actin-like 6A predicts poor prognosis
of hepatocellular carcinoma and promotes metastasis and
epithelial-mesenchymal transition. Hepatology. 63:1256–1271. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Huang Q, Zhan L, Cao H, Li J, Lyu Y, Guo
X, Zhang J, Ji L, Ren T, An J, et al: Increased mitochondrial
fission promotes autophagy and hepatocellular carcinoma cell
survival through the ROS-modulated coordinated regulation of the
NFKB and TP53 pathways. Autophagy. 12:999–1014. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Wang L, Liu X, Nie J, Zhang J, Kimball SR,
Zhang H, Zhang WJ, Jefferson LS, Cheng Z, Ji Q and Shi Y: ALCAT1
controls mitochondrial etiology of fatty liver diseases, linking
defective mitophagy to steatosis. Hepatology. 61:486–496. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Ding WX, Li M and Yin XM: Selective taste
of ethanol-induced autophagy for mitochondria and lipid droplets.
Autophagy. 7:248–249. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Lodder J, Denaës T, Chobert MN, Wan J,
El-Benna J, Pawlotsky JM, Lotersztajn S and Teixeira-Clerc F:
Macrophage autophagy protects against liver fibrosis in mice.
Autophagy. 11:1280–1292. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Ueno T and Komatsu M: Autophagy in the
liver: Functions in health and disease. Nat Rev Gastroenterol
Hepatol. 14:170–184. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Samal J, Kandpal M and Vivekanandan P:
Molecular mechanisms underlying occult hepatitis B virus infection.
Clin Microbiol Rev. 25:142–163. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Shin GC, Kang HS, Lee AR and Kim KH:
Hepatitis B virus-triggered autophagy targets TNFRSF10B/death
receptor 5 for degradation to limit TNFSF10/TRAIL response.
Autophagy. 12:2451–2466. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Lan SH, Wu SY, Zuchini R, Lin XZ, Su IJ,
Tsai TF, Lin YJ, Wu CT and Liu HS: Autophagy suppresses
tumorigenesis of hepatitis B virus-associated hepatocellular
carcinoma through degradation of microRNA-224. Hepatology.
59:505–517. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Sir D, Ann DK and Ou JH: Autophagy by
hepatitis B virus and for hepatitis B virus. Autophagy. 6:548–549.
2010. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Altieri DC: Mitochondrial HSP90s and tumor
cell metabolism. Autophagy. 9:244–245. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Guo XL, Hu F, Zhang SS, Zhao QD, Zong C,
Ye F, Guo SW, Zhang JW, Li R, Wu MC and Wei LX: Inhibition of p53
increases chemosensitivity to 5-FU in nutrient-deprived
hepatocarcinoma cells by suppressing autophagy. Cancer Lett.
346:278–284. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Amaravadi R, Kimmelman AC and White E:
Recent insights into the function of autophagy in cancer. Genes
Dev. 30:1913–1930. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Pridgeon JW, Olzmann JA, Chin LS and Li L:
PINK1 protects against oxidative stress by phosphorylating
mitochondrial chaperone TRAP1. PLoS Biol. 5:e1722007. View Article : Google Scholar : PubMed/NCBI
|