Introduction
The F-box protein S-phase kinase associated protein
2 (Skp2), a component of the Skp1-Cullin 1-F-box E3
ubiquitin-ligase complex, promotes the degradation of the
cyclin-dependent kinase inhibitor p27, a key regulator of
G1 progression. An inverse correlation between Skp2 and
p27 protein level has been reported in tumors (1,2). Under
physiological conditions, Skp2 controls the initiation of mitosis
in that its expression peaks at the S and G2 phases, but
not at G0 and G1 phases (3,4). Skp2 is
key in the p27 ubiquitin degradation pathway and may inhibit the
proliferation of a variety of cell types through the
ubiquitin-proteasome pathway (4–6).
Skp2 overexpression has been observed in various
human cancers, including lymphomas (7), prostate cancer (8), melanoma (9), pancreatic cancer (10), gastric cancer (11), breast cancer (12), colorectal cancer (13) and lung cancer (14,15).
The involvement of Skp2 overexpression in metastasis
and aggressiveness has been reported in many tumors including
melanoma (9), pancreatic cancer
(10), breast (12), colorectal cancer (13) and lung cancer (1). Skp2 is also considered to be associated
with the occurrence, development and prognosis of malignant tumors
such as breast cancer (12), melanoma
(9), prostate cancer (8), colorectal cancer (14), and lung cancer (1).
Expression of Skp2 is significantly upregulated in
primary non-small cell lung cancer (NSCLC), compared with
non-tumorous lung tissues (16). In
patients with NSCLC, overexpression of the Skp2 protein is
associated with numerous clinicopathological parameters, including
histology, lymph node metastasis, smoking status, differentiation
and tumor stage (1,16). In addition, the positive expression of
Skp2 is correlated with a poor prognosis in patients with NSCLC
(1).
Concerning the two major pathological types of
NSCLC, Skp2 is notably overexpressed in lung squamous cell
carcinoma (LUSC), compared with lung adenocarcinoma (LUAD)
(1). Due to the limitation of sample
amounts, it remains unclear whether Skp2 expression is associated
with those clinicopathological features in LUAD and LUSC,
respectively; therefore, a study with a greater number of patients
and specimens is required to compare the clinicopathological and
prognostic implications with the Skp2 expression in these NSCLC
types.
In the present study, the correlation between
clinicopathological features and Skp2 protein expression was
investigated in 351 LUAD and 149 LUSC samples. Additionally, the
prognostic value of Skp2 protein expression was also investigated.
Bioinformatics analyses were performed to investigate Skp2
expression-associated mRNAs and lung cancer mutations.
Materials and methods
Cases and specimens
Patients were excluded due to chemotherapy or
radiotherapy prior to surgery. A total of 351 patients with LUAD
and 149 patients with LUSC who were diagnosed and underwent surgery
at Peking University People's Hospital (Beijing, China) between
January 2004 and December 2012 were enrolled in the present study.
The patients included 305 males and 195 females, with a median age
of 64 years (range, 24–86). All procedures performed in the current
study involving human participants were in accordance with the
ethical standards of the Ethics Committee of Peking University
People's Hospital. Informed consent was obtained from all
individual participants included in the study. All patients were
followed up following surgery at 3–6 month intervals, and the total
follow-up periods ranged from 4.8 to 7.6 years, with a median of
5.9 years. Complete clinical information from the clinical and
pathological records, including sex, age, tumor size, lymphatic
invasion, differentiation grade and pathological histology were
collected and summarized in Table
I.
 | Table I.Correlation between Skp2 protein
expression and clinicopathological parameters in lung cancers. |
Table I.
Correlation between Skp2 protein
expression and clinicopathological parameters in lung cancers.
|
| LUAD | LUSC |
|---|
|
|
|
|
|---|
|
|
|
| Skp2 expression |
|
|
| Skp2 expression |
|
|
|---|
|
|
|
|
|
|
|
|
|
|
|
|---|
| Histology
Clinicopathological parameters | Category | Cases | Low | High | Statistical
value | P-value | Cases | Low | High | Statistical
value | P-value |
|---|
| Total |
| 351 | 231 | 120 |
|
| 149 | 54 | 95 |
|
|
| Sex | Male | 172 | 102 | 70 |
6.325b | 0.012a | 133 | 48 | 85 | 0.012b | 0.912 |
|
| Female | 179 | 129 | 50 |
|
| 16 | 6 | 10 |
|
|
| Age | <65 | 186 | 129 | 57 |
2.207b | 0.137 | 81 | 34 | 47 | 2.525b | 0.112 |
|
| ≥65 | 165 | 102 | 63 |
|
| 68 | 20 | 48 |
|
|
|
Differentiation | Well | 107 | 91 | 16 | 36.308c |
<0.001a | 11 | 7 | 4 | 2.898c | 0.235 |
|
| Moderate | 122 | 78 | 44 |
|
| 68 | 25 | 43 |
|
|
|
| Poor | 122 | 62 | 60 |
|
| 70 | 22 | 48 |
|
|
| Smoking status
(cigarettes/year) | 0 | 232 | 163 | 69 | 16.198c |
<0.001a | 31 | 14 | 17 | 6.555c | 0.038a |
|
| 0<n≤400 | 62 | 41 | 21 |
|
| 30 | 7 | 23 |
|
|
|
| >400 | 57 | 27 | 30 |
|
| 88 | 33 | 55 |
|
|
| T | 1 | 157 | 110 | 47 |
4.410c | 0.221 | 30 | 9 | 21 | 0.653c | 0.884 |
|
| 2 | 170 | 110 | 60 |
|
| 86 | 35 | 51 |
|
|
|
| 3 | 13 | 6 | 7 |
|
| 26 | 8 | 18 |
|
|
|
| 4 | 11 | 5 | 6 |
|
| 7 | 2 | 5 |
|
|
| N | 0 | 233 | 174 | 59 | 24.081c |
<0.001a | 72 | 23 | 49 | 2.564c | 0.464 |
|
| 1 | 35 | 17 | 18 |
|
| 33 | 13 | 20 |
|
|
|
| 2 | 79 | 37 | 42 |
|
| 43 | 17 | 26 |
|
|
|
| x | 4 | 3 | 1 |
|
| 1 | 1 | 0 |
|
|
| M | 0 | 334 | 219 | 115 |
0.181b | 0.670 | 146 | 52 | 94 | 1.227b | 0.268 |
|
| 1 | 17 | 12 | 5 |
|
| 3 | 2 | 1 |
|
|
| TNM stage | I | 215 | 165 | 50 | 33.202c |
<0.001a | 54 | 19 | 35 | 1.127c | 0.771 |
|
| II | 44 | 19 | 25 |
|
| 41 | 14 | 27 |
|
|
|
| III | 75 | 35 | 40 |
|
| 51 | 19 | 32 |
|
|
|
| IV | 17 | 12 | 5 |
|
| 3 | 2 | 1 |
|
|
| Diameter | ≤3 cm | 270 | 195 | 75 | 21.368b |
<0.001a | 47 | 15 | 32 | 0.556b | 0.456 |
|
| >3 cm | 81 | 36 | 45 |
|
| 102 | 39 | 63 |
|
|
Normal tissue was collected from at >2 cm away
from the primary cancer site and histopathologically identified as
normal by a pathologist at the Peking University People's Hospital.
Fresh tissue samples from each patient were formalin-fixed (10%) at
room temperature overnight, paraffin-embedded and constructed into
tissue microarrays (TMA).
TMA and immunohistochemistry
(IHC)
TMA were prepared using carefully identified
representative core-tissue specimens (2 mm in diameter) as
previously described (17). Sections
4 µm thick from the recipient blocks were cut. Antigen retrieval
was performed in a pressure cooker, followed by treatment with 3%
hydrogen peroxide for 15 min to block endogenous peroxidase
activity. IHC analysis was performed using an antibody against Skp2
at room temperature for 2 h (1:200 dilution; cat. no. 2652; Cell
Signaling Technology, Beverly, MA, USA). The sections were rinsed
and incubated for 30 min with biotinylated second antibody in room
temperature. Expression of Skp2 was detected using a
diaminobenzidine reaction for 30 min and hematoxylin
counterstaining for 30 sec at room temperature. There were two
independent pathologists, who were blind to the patients' clinical
data, who scored the staining intensity level as follows: Level-0,
negative; level-1, weak positive; level-2, moderate positive;
level-3, strong positive, and the percentage of positive cells.
Skp2 protein expression assessment was based on the method of a
semi-quantitatively scoring system (18). The final score was obtained from
multiplying the staining intensity level and the percentage of
positively stained cells (A for level-1, B for level-2, C for
level-3). The calculation formula was as follows: Score = 1×A% +
2×B% + 3×C%. The semi-quantitative method provided further
information regarding the expression of Skp2 and reduced bias,
whilst analyzing the correlation with clinicopathological
characters.
Skp2 co-expression analysis of LUAD
and LUSC differential mRNAs
The Cancer Genome Atlas (TCGA) mRNA-expression data
for LUSC and LUAD were obtained from Firehose (http://gdac.broadinstitute.org/) using the
Bioconductor package RTCGAToolbox (version 2.4.0; http://bioconductor.org/packages/RTCGAToolbox/). For
LUAD, a provisional dataset of 162 total tumors was used, and for
LUSC, 240 samples were used.
Differential mRNAs were screened using the
Bioconductor package edgeR (version 3.16.5; http://bioconductor.org/packages/edgeR/) (19). The FDR threshold was set to 0.05, in
order to identify significantly deviating genes between LUAD and
LUSC. The LogFC threshold was set to 1.5 to select mRNAs for the
following co-expression analysis. The R statistical language
(version3.4.1; http://www.r-project.org/) was used to conduct these
analyses.
The spearman rank correlation method was used to
assess the co-expression of the differential mRNAs. The top 50
Skp2-associated mRNAs were cluster analyzed and plotted by R
package heatmap (version1.0.8; http://CRAN.R-project.org/package=pheatmap). The
highest associated mRNAs were processed in The Database for
Annotation, Visualization and Integrated Discovery functional
annotation bioinformatics analysis (20,21).
TCGA gene expression and mutation data for LUAD and
LUSC were obtained from the cBioPortal for Cancer Genomics
(www.cbioportal.org) (22,23). For
LUAD, the provisional dataset of 522 total tumors was used.
Mutation data were obtained for tumor protein P53 (TP53), KRAS,
Kelch-like ECH-associated protein 1 (KEAP1), serine/threonine
kinase 11, epidermal growth factor receptor, neurofibromin 1 (NF1),
SET domain containing 2, RNA binding motif protein 10, MGA, MET,
AT-rich interaction domain 1A,
phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit α
(PIK3CA), SWI/SNF-associated matrix-associated actin dependent
regulator of chromatin subfamily A member 4, RB transcriptional
corepressor 1 (RB1), cyclin dependent kinase inhibitor 2A (CDKN2A),
U2 small nuclear RNA auxiliary factor 1, Ras-like without CAAX 1,
Erb-B2 receptor tyrosine kinase 2 and ALK. For LUSC, data were
obtained from the provisional dataset of 504 samples. Mutation data
were obtained for TP53, CDKN2A, phosphatase and tensin homolog
(PTEN), PIK3CA, KEAP1, major histocompatibility complex class I A,
nuclear factor erythroid 2 like 2 (NFE2L2), NOTCH1 and RB1.
Statistical analysis
Median score and 5–95 percentile were plotted. Skp2
expression among different groups according to pathological
grading, lymph node metastasis, smoking status and differentiation
grade were evaluated by Kruskal-Wallis H-test which is a
non-parametric method for testing whether samples originate from
the same distribution. Wilcoxon matched-pairs signed rank test were
used to compare Skp2 expression in LUAD and LUSC Pearson's
χ2 was used to compare Skp2 expression in other clinical
features, including histological, age, sex, tumor size and distant
metastasis. Disease-free survival (DFS) was defined as the period
between surgery and the date of recurrence, and overall survival
(OS) was considered as the interval from surgery to the date of
mortality or final follow-up. Kaplan-Meier method with the log-rank
test was performed to determine the difference in survival between
groups. Cox proportional hazards regression analysis was performed
to estimate the hazard ratios for positive risk factors with the
backward conditional elimination method (24). P<0.05 was considered to indicate a
statistically significant difference (two sided). SPSS 22.0
statistic software (IBM Corp., Armonk, NY, USA) and GraphPad Prism
6 software (GraphPad Software, Inc., La Jolla, CA, USA) were used
to perform the computation for all statistical analyses.
Results
Correlation between Skp2 protein
expression assessed by TMA-IHC and clinicopathological parameters
in LUSC and LUAD
Skp2 protein expression was assessed by TMA-IHC
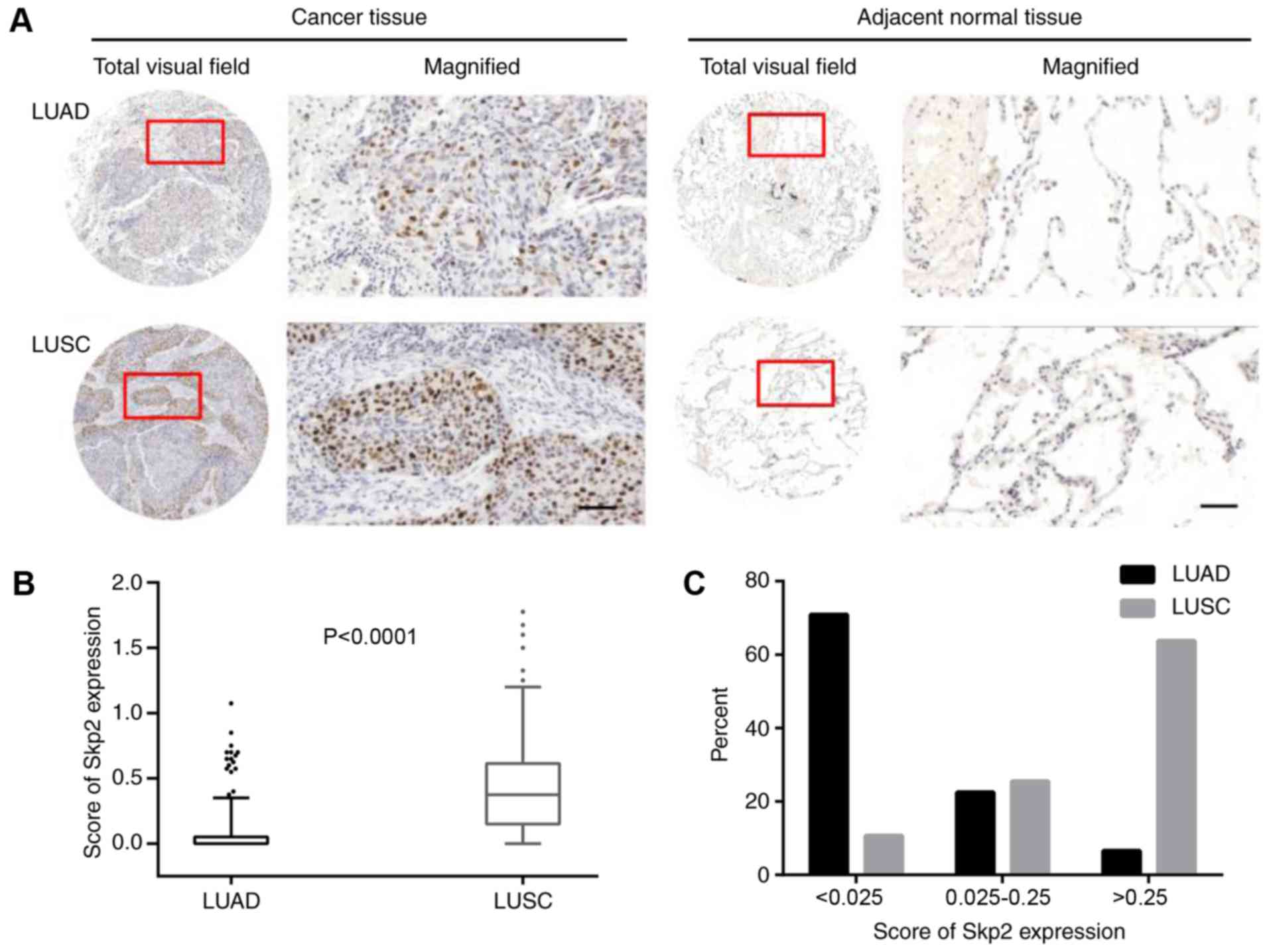
analysis in the primary lung cancer tissues. Skp2 staining was
primarily localized in the nuclear area, which is consistent with a
previous study (2) (Fig. 1). Another previous study indicated
that Skp2 expression was significantly higher in LUSC, compared
with LUAD, which was verified in the present study (P<0.0001;
Fig. 1A and B). The median score of
Skp2 in LUAD and LUSC were 0.00 and 0.375, respectively. In NSCLC
(LUAD and LUSC), the 25th, 50th, 75th quartiles of Skp2 expression
final score were 0.000, 0.025 and 0.250. Distribution of the Skp2
expression score in the three intervals were notably different. The
majority of Skp2 expression scores were <0.025 in the LUAD
group, while scores >0.250 were observed more frequently in LUSC
(Fig. 1C). Based on these data,
subgroups analysis was proceeded in LUAD and LUSC, separately. A
score of 0.025 was used to define high expression and low
expression in LUAD, and a score of 0.250 in LUSC was used
identically.
In LUAD, Skp2 expression varied in different groups
according to sex, differentiation, smoking history,
Tumor-Node-Metastasis (TNM) stage (25), lymph node metastasis and tumor
diameter (P<0.05; Table I). In
LUSC, Skp2 protein expression correlated with smoking status
(P<0.05; Table I), whilst it
demonstrated limited correlation with other clinical features.
Prognostic significance of Skp2
protein expression in LUAD
There were three patients who succumbed during the
follow-up, shortly following leaving the hospital. Survival
analyses were performed in 348 patients with LUAD, of which 120
patients had a high expression of Skp2 and 228 patients had a low
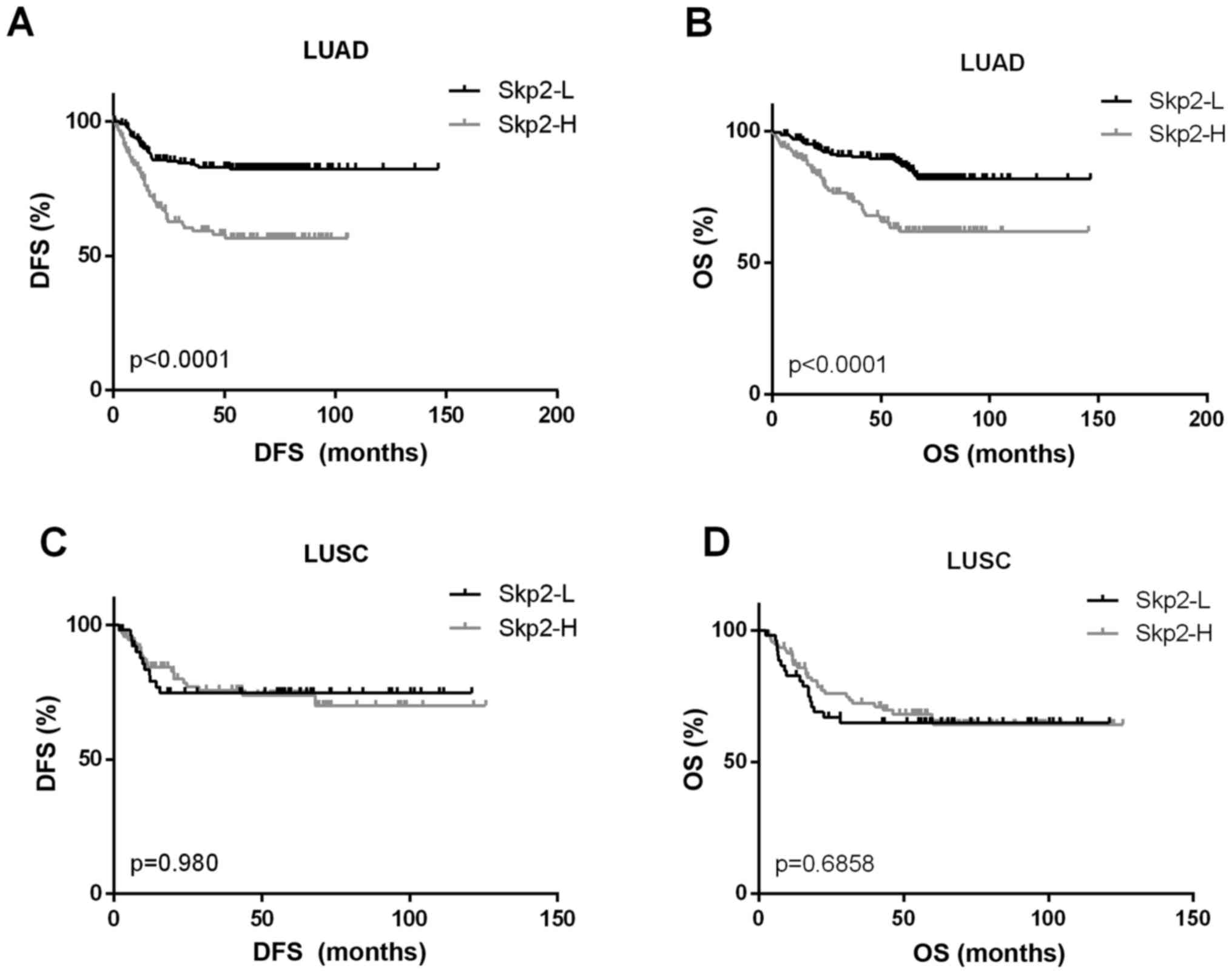
expression. Kaplan-Meier survival analysis with log-rank test
indicated that a high Skp2 protein expression in patients with LUAD
was significantly associated with a reduced DFS and OS (P<0.001;
Fig. 2A and B). The 5-year OS in the
Skp2-H group was 61.9%, compared with a rate of 86.9% in the Skp2-L
group. For patients with LUSC, no association was observed between
Skp2 expression and the outcome of patients (Fig. 2C and D).
Cox proportional hazards model was used to determine
the association of six factors (sex, age, differentiation,
pathological stage, smoking status and Skp2 protein expression)
with the OS of patients with LUAD. As a result, Skp2 protein highly
expression was demonstrated to be an independent prognostic factor
of OS, with a relative risk of 1.845 (P=0.030). Poor
differentiation (P=0.016) and high stage (P<0.001) were
associated with a reduced OS also (Table
II).
 | Table II.Multivariate analysis for prognostic
factors in patients with lung adenocarcinoma. |
Table II.
Multivariate analysis for prognostic
factors in patients with lung adenocarcinoma.
|
| OS |
|---|
|
|
|
|---|
| Variable | Regression
coefficient | S.E. | Relative risk | 95%CI | P-value |
|---|
| Sex (male vs.
female) | −0.221 | 0.273 | 0.801 | 0.469–1.368 | 0.417 |
| Age (<65 vs.
≥65) | 0.536 | 0.277 | 1.709 | 0.993–2.941 | 0.053 |
| Differentiation
(well/moderate vs. poor) | 1.282 | 0.531 | 3.604 | 1.272–10.214 | 0.016a |
| TNM stage (stage I
vs. stages II–IV) | 1.484 | 0.324 | 4.413 | 2.340–8.320 |
<0.001a |
| Smoking status
(non-smokerb vs.
smoker) | −0.322 | 0.354 | 0.752 | 0.362–1.452 | 0.364 |
| Skp2 expression
(low vs. high) | 0.613 | 0.283 | 1.845 | 1.060–3.212 | 0.030a |
Histological differential mRNAs and
frequent mutations association with Skp2
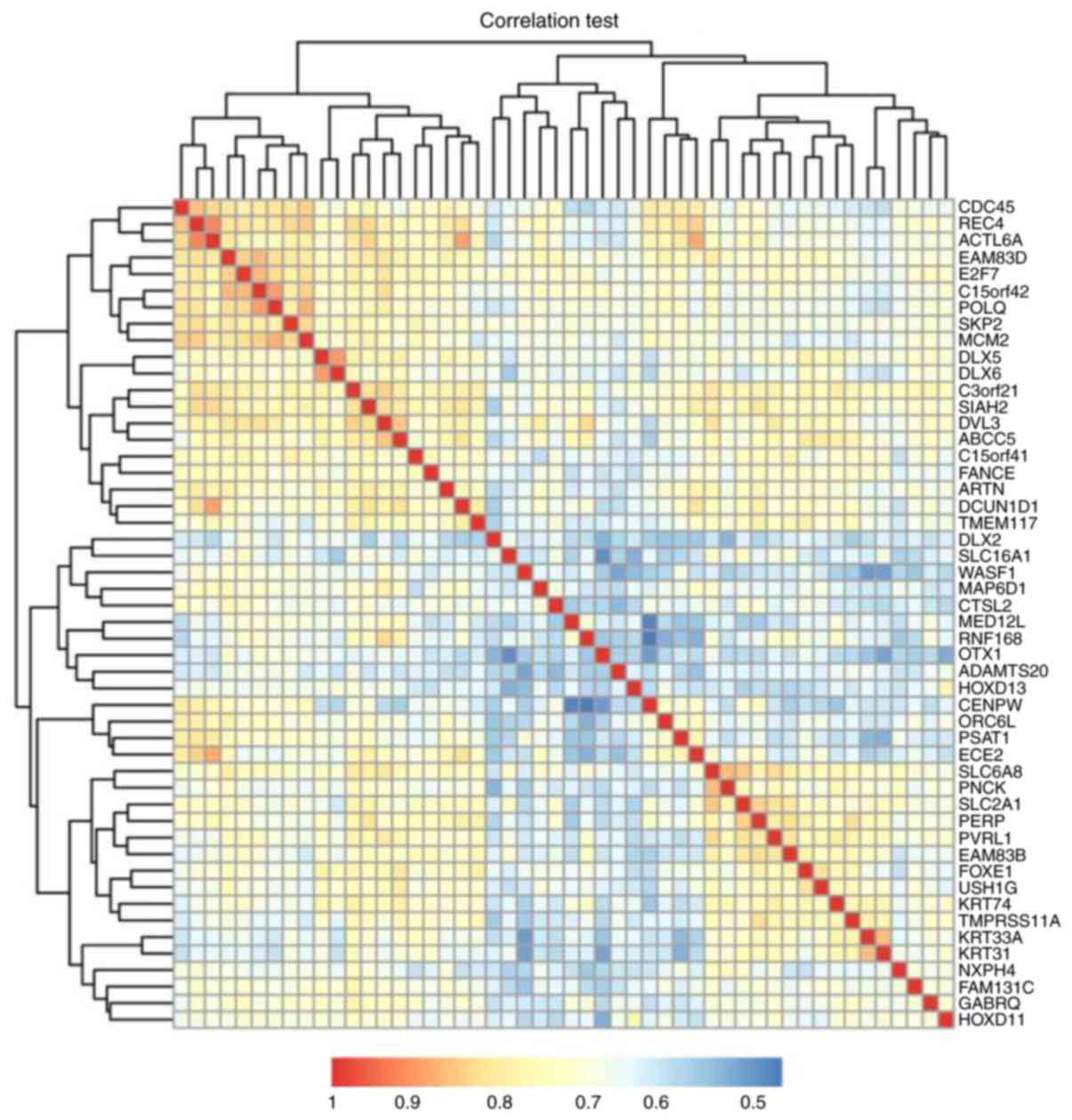
Association analysis and cluster analysis screened
nine Skp2 co-expression mRNAs [cell division cycle 45 (CDC45),
replication factor C subunit 4 (RFC4), actin like 6A (ACTL6A),
family with sequence similarity 83 member D, E2F transcription
factor 7, chromosome 15 open reading frame 42 (C15orf42), DNA
polymerase θ (POLQ), Skp2 and minichromosome maintenance complex
component 2 (MCM2)], which are also differential between LUAD and
LUSC. Skp2, CDC45 and MCM2 are involved in the G1/S
transition of the mitotic cell cycle. CDC45, MCM2, RFC4, POLQ and
ACTL6A participate in DNA replication and repair (Fig. 3).
In the TCGA cohort of patients with LUAD and LUSC,
Skp2 expression was highly expressed in the TP53, NF1 and RB1
mutation groups, compared with the unaltered group. The KRAS
mutation group had a lower Skp2 expression in LUAD. Whilst in LUSC,
it was determined that there was a significant deviation of Skp2
expression in the four other commonly mutated genes, PTEN, PIK3CA,
KEAP1 and NFE2L2 (Table III).
 | Table III.Deviation of S-phase kinase
associated protein 2 expression with lung cancer mutations. |
Table III.
Deviation of S-phase kinase
associated protein 2 expression with lung cancer mutations.
|
|
| Mean | SD |
|
|
|---|
|
|
|
|
|
|
|
|---|
| Histology | Gene (n/%) | Altered group | Unaltered
group | Altered group | Unaltered
group | P-value | Q-value |
|---|
| LUAD | TP53
(111/21.3%) | 7.44 | 7.05 | 1.08 | 0.97 |
9.270×10−4 |
7.036×10−3 |
|
| KRAS
(103/19.7%) | 6.90 | 7.20 | 0.79 | 1.04 |
1.639×10−3 | 0.0324 |
|
| NF1 (29/13%) | 7.61 | 7.10 | 1.02 | 1.00 | 0.0102 | 0.193 |
|
| RB1 (17/7.4%) | 7.98 | 7.10 | 1.00 | 0.99 |
3.696×10−4 | 0.0178 |
| LUSC | TP53
(150/29.8%) | 9.22 | 8.79 | 0.83 | 0.96 |
6.32×10−7 |
6.708×10−6 |
|
| PTEN
(74/14.7%) | 9.24 | 8.87 | 0.86 | 0.95 |
1.091×10−3 | 0.119 |
|
| PIK3CA
(251/49.8) | 9.06 | 8.77 | 0.83 | 1.03 |
5.430×10−4 |
2.352×10−3 |
|
| KEAP1
(40/7.9%) | 9.28 | 8.89 | 0.90 | 0.94 | 0.0104 | 0.0869 |
|
| NFE2L2
(58/11.5%) | 9.16 | 8.88 | 0.88 | 0.95 |
7.959×10−3 | 0.0348 |
|
| RB1 (33/6.5%) | 9.39 | 8.88 | 0.90 | 0.94 |
1.199×10−3 | 0.0394 |
Discussion
LUAD and LUSC are the NSCLC types with the highest
prevalence, accounting for 80–85% of lung cancer types. Large-scale
sequencing studies have revealed the genomic differences between
LUAD and LUSC (26). The molecular
mechanisms underlying LUAD and LUSC are considerably different
(27). A previous study demonstrated
that the Skp2 relative gene copy number aberrations were detected
in NSCLC, and these changes are reflected at the mRNA and protein
expression levels (16).
In the present study, Skp2 protein has different
expression patterns in LUSC and LUAD; therefore the association
between Skp2 expression and clinicopathological parameters in
patients with LUAD and LUSC were evaluated. The data indicated that
clinicopathological and prognostic implications based on Skp2
expression in LUAD and LUSC should be considered different.
The data indicated that high expression of Skp2 in
LUSC would result in abnormal activation of DNA replication and
G1/S transition pathways. This indicated that
alterations of lung cancer associated oncogenes modulating these
pathways may contribute to the aberration of Skp2 expression. The
association between lung cancer mutations and Skp2 expression were
then investigated. Skp2 was expressed higher in the lung
cancer-associated genes altered group, except in the KRAS mutation
group. KRAS has a mutation rate of 19.7% in LUAD, which is more
frequent than LUSC; however, PTEN, PIK3CA, KEAP1 and NFE2L2, which
induce the proliferation of cancer cells, are present more
frequently in LUSC. Different driver gene mutation frequencies may
be one reason for the distinct expression patterns of Skp2 in LUSC
and LUAD; however, this would not fully explain the difference of
Skp2 expression between LUAD and LUSC. Further study focusing on
the regulation of Skp2 expression are required, including the
regulated relationship with deviated miRNAs and lncRNAs.
Skp2 has a function in promoting cell growth
(28). A previous study primarily
focused on the T stage (29). In the
present study, Skp2 overexpression was associated with the tumor
diameter more significantly than T stage (P<0.001 vs. P=0.221;
Table I). Ina ddition to the T stage,
continued studies must also consider pleural invasion and the
distance to the carina of the trachea, but these features are not
affected by tumor proliferation ability.
The prognostic value of Skp2 expression in NSCLC has
been controversial. Osoegawa et al (1) indicated that a high Skp2 protein
expression was an independent poor prognostic marker in NSCLC;
however, Zhu et al (30)
reported that Skp2 overexpression was not prognostically
significant alone, but along with a RAS mutation was a significant
independent poor prognostic marker in NSCLC. In the present study,
it was demonstrated that a high expression of Skp2 may indicate a
poor prognosis in LUAD, but it was not observed in LUSC.
Additionally, in KRAS-mutated LUAD, Skp2 had a lower expression
level, compared with the unaltered group. This may indicate that
Skp2 may have clinicopathological and prognostic implications when
it is expressed at a relatively lower level.
Detection of Skp2 overexpression may be a beneficial
marker for prognosis. Skp2 may provide sputum-based markers that
have the potential to improve the early detection of LUSC (31). Skp2 can also be detected in the
peripheral blood of patients with NSCLC, and has demonstrated high
diagnostic value (32). These studies
provide evidence that detection of Skp2 expression level should be
conducted prior to surgery, assisting with the decision for surgery
procedure; however, the molecular mechanism underlying regulation
of Skp2 in lung cancer remains unclear and further in vitro
and in vivo investigations are required.
To conclude, Skp2 protein expression has different
patterns between LUAD and LUSC. The clinicopathological and
prognostic implications based on Skp2 expression in LUAD and LUSC
should be considered different. Differential expression of Skp2
protein expression was associated with an unfavorable outcome in
patients with LUAD, but not in LUSC. Skp2-H LUSC may have robust
proliferation ability.
Acknowledgements
Not applicable.
Funding
The present study was funded by the National High
Technology Research and Development Program of China (863 Program)
(grant nos. ss2014AA020602 and ss2014AA020604).
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
KZ analyzed and interpreted the data regarding the
Skp2 expression and clinicopathological parameters, and also was a
major contributor in writing the manuscript. QH and JC performed
the tissue microarray and immunohistochemical analyze. FY and JW
made substantial contributions to conception and design. All
authors read and approved the final manuscript.
Ethics approval and consent to
participate
All procedures performed in the current study
involving human participants were in accordance with the ethical
standards of the Ethics Committee of Peking University People's
Hospital and with the 1964 Helsinki declaration and its later
amendments or comparable ethical standards. Informed consent was
obtained from all individual participants included in the
study.
Consent for publication
Written informed consent for the publication of any
associated data and accompanying images was obtained from all the
patient, or parent, guardian or next of kin (in case of deceased
patients).
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Osoegawa A, Yoshino I, Tanaka S, Sugio K,
Kameyama T, Yamaguchi M and Maehara Y: Regulation of p27 by S-phase
kinase-associated protein 2 is associated with aggressiveness in
non-small-cell lung cancer. J Clin Oncol. 22:4165–4173. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Gstaiger M, Jordan R, Lim M, Catzavelos C,
Mestan J, Slingerland J and Krek W: Skp2 is oncogenic and
overexpressed in human cancers. Proc Natl Acad Sci USA.
98:5043–5048. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Nakayama KI, Hatakeyama S and Nakayama K:
Regulation of the cell cycle at the G1-S transition by proteolysis
of cyclin E and p27Kip1. Biochem Biophys Res Commun. 282:853–860.
2001. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Nakayama K, Nagahama H, Minamishima YA,
Matsumoto M, Nakamichi I, Kitagawa K, Shirane M, Tsunematsu R,
Tsukiyama T, Ishida N, et al: Targeted disruption of Skp2 results
in accumulation of cyclin E and p27(Kip1), polyploidy and
centrosome overduplication. EMBO J. 19:2069–2081. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Sicari BM, Troxell R, Salim F, Tanwir M,
Takane KK and Fiaschi-Taesch N: c-myc and skp2 coordinate p27
degradation, vascular smooth muscle proliferation, and neointima
formation induced by the parathyroid hormone-related protein.
Endocrinology. 153:861–872. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Suzuki S, Fukasawa H, Misaki T, Togawa A,
Ohashi N, Kitagawa K, Kotake Y, Liu N, Niida H, Nakayama K, et al:
The amelioration of renal damage in Skp2-deficient mice canceled by
p27 Kip1 deficiency in Skp2−/− p27−/− mice.
PLoS One. 7:e362492012. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Lim MS, Adamson A, Lin Z, Perez-Ordonez B,
Jordan RC, Tripp S, Perkins SL and Elenitoba-Johnson KS: Expression
of Skp2, a p27(Kip1) ubiquitin ligase, in malignant lymphoma:
Correlation with p27(Kip1) and proliferation index. Blood.
100:2950–2956. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Wang Z, Gao D, Fukushima H, Inuzuka H, Liu
P, Wan L, Sarkar FH and Wei W: Skp2: A novel potential therapeutic
target for prostate cancer. Biochim Biophys Acta. 1825:11–17.
2012.PubMed/NCBI
|
|
9
|
Rose AE, Wang G, Hanniford D, Monni S, Tu
T, Shapiro RL, Berman RS, Pavlick AC, Pagano M, Darvishian F, et
al: Clinical relevance of SKP2 alterations in metastatic melanoma.
Pigment Cell Melanoma Res. 24:197–206. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Schüler S, Diersch S, Hamacher R, Schmid
RM, Saur D and Schneider G: SKP2 confers resistance of pancreatic
cancer cells towards TRAIL-induced apoptosis. Int J Oncol.
38:219–225. 2011.PubMed/NCBI
|
|
11
|
Osoegawa A, Yoshino I, Tanaka S, Sugio K,
Kameyama T, Yamaguchi M and Maehara Y: Regulation of p27 by S-phase
kinase-associated protein 2 is associated with aggressiveness in
non-small-cell lung cancer. J Cinical Oncol. 22:4165–4173. 2004.
View Article : Google Scholar
|
|
12
|
Wang Z, Fukushima H, Inuzuka H, Wan L, Liu
P, Gao D, Sarkar FH and Wei W: Skp2 is a promising therapeutic
target in breast cancer. Front Oncol. 1:187022012. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Tian YF, Chen TJ, Lin CY, Chen LT, Lin LC,
Hsing CH, Lee SW, Sheu MJ, Lee HH, Shiue YL, et al: SKP2
overexpression is associated with a poor prognosis of rectal cancer
treated with chemoradiotherapy and represents a therapeutic target
with high potential. Tumor Biol. 34:1107–1117. 2013. View Article : Google Scholar
|
|
14
|
Hershko DD: Oncogenic properties and
prognostic implications of the ubiquitin ligase Skp2 in cancer.
Cancer. 112:1415–1424. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Timmerbeul I, Garrett-Engele CM, Kossatz
U, Chen X, Firpo E, Grünwald V, Kamino K, Wilkens L, Lehmann U,
Buer J, et al: Testing the importance of p27 degradation by the
SCFskp2 pathway in murine models of lung and colon cancer. Proc
Natl Acad Sci USA. 103:14009–14014. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Yokoi S, Yasui K, Mori M, Iizasa T,
Fujisawa T and Inazawa J: Amplification and overexpression of SKP2
are associated with metastasis of non-small-cell lung cancers to
lymph nodes. Am J Pathol. 165:175–180. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Hewitt SM: Tissue microarrays as a tool in
the discovery and validation of predictive biomarkers. Methods Mol
Biol. 823:201–214. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Remmele W and Stegner HE: Recommendation
for uniform definition of an immunoreactive score (IRS) for
immunohistochemical estrogen receptor detection (ER-ICA) in breast
cancer tissue. Pathologe. 8:138–140. 1987.(In German). PubMed/NCBI
|
|
19
|
Robinson MD and Oshlack A: A scaling
normalization method for differential expression analysis of
RNA-seq data. Genome Biol. 11:R252010. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
da Huang W, Sherman BT and Lempicki RA:
Systematic and integrative analysis of large gene lists using DAVID
bioinformatics resources. Nat Protoc. 4:44–57. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
da Huang W, Sherman BT and Lempicki RA:
Bioinformatics enrichment tools: Paths toward the comprehensive
functional analysis of large gene lists. Nucleic Acids Res.
37:1–13. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Gao J, Aksoy BA, Dogrusoz U, Dresdner G,
Gross B, Sumer SO, Sun Y, Jacobsen A, Sinha R, Larsson E, et al:
Integrative analysis of complex cancer genomics and clinical
profiles using the cBioPortal. Sci Signal. 6:pl12013. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Cerami E, Gao J, Dogrusoz U, Gross BE,
Sumer SO, Aksoy BA, Jacobsen A, Byrne CJ, Heuer ML, Larsson E, et
al: The cBio cancer genomics portal: An open platform for exploring
multidimensional cancer genomics data. Cancer Discov. 2:401–404.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Costacou T, Lopes-Virella MF, Zgibor JC,
Virella G, Otvos J, Walsh M and Orchard TJ: Markers of endothelial
dysfunction in the prediction of coronary artery disease in type 1
diabetes. The Pittsburgh epidemiology of diabetes complications
study. J Diabetes Complications. 19:183–193. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Edge SB and Compton CC: The American Joint
Committee on Cancer: The 7th edition of the AJCC cancer staging
manual and the future of TNM. Ann Surg Oncol. 17:1471–1474. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Campbell JD, Alexandrov A, Kim J, Wala J,
Berger AH, Pedamallu CS, Shukla SA, Guo G, Brooks AN, Murray BA, et
al: Distinct patterns of somatic genome alterations in lung
adenocarcinomas and squamous cell carcinomas. Nat Genet.
48:607–616. 2016. View
Article : Google Scholar : PubMed/NCBI
|
|
27
|
Liu J, Yang XY and Shi WJ: Identifying
differentially expressed genes and pathways in two types of
non-small cell lung cancer: Adenocarcinoma and squamous cell
carcinoma. Genet Mol Res. 13:95–102. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Su L, Han D, Wu J and Huo X: Skp2
regulates non-small cell lung cancer cell growth by Meg3 and
miR-3163. Tumour Biol. 37:3925–3931. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Takanami I: The prognostic value of
overexpression of Skp2 mRNA in non-small cell lung cancer. Oncol
Rep. 13:727–731. 2005.PubMed/NCBI
|
|
30
|
Zhu CQ, Blackhall FH, Pintilie M, Iyengar
P, Liu N, Ho J, Chomiak T, Lau D, Winton T, Shepherd FA and Tsao
MS: Skp2 gene copy number aberrations are common in non-small cell
lung carcinoma, and its overexpression in tumors with ras mutation
is a poor prognostic marker. Clin Cancer Res. 10:1984–1991. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Jiang F, Todd NW, Li R, Zhang H, Fang H
and Stass SA: A panel of sputum-based genomic marker for early
detection of lung cancer. Cancer Prev Res (Phila). 3:1571–1578.
2010. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Chiang IT, Wang WS, Liu HC, Yang ST, Tang
NY and Chung JG: Curcumin alters gene expression-associated DNA
damage, cell cycle, cell survival and cell migration and invasion
in NCI-H460 human lung cancer cells in vitro. Oncology Rep.
34:1853–1874. 2015. View Article : Google Scholar
|