|
1
|
Siegel RL, Miller KD and Jemal A: Cancer
Statistics, 2017. CA Cancer J Clin. 67:7–30. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Goya T, Asamura H, Yoshimura H, Kato H,
Shimokata K, Tsuchiy R, Sohara Y, Miya T and Miyaoka E; Japanese
Joint Committee of Lung Cancer Registry, : Prognosis of 6644
resected non-small cell lung cancers in Japan: A Japanese lung
cancer registry study. Lung Cancer. 50:227–234. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Wu GX and Raz DJ: Lung cancer screening.
Cancer Treat Res. 170:1–23. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Witwer KW: Circulating microRNA biomarker
studies: Pitfalls and potential solutions. Clin Chem. 61:56–63.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Moretti F, D'Antona P, Finardi E, Barbetta
M, Dominioni L, Poli A, Gini E, Noonan DM, Imperatori A, Rotolo N,
et al: Systematic review and critique of circulating miRNAs as
biomarkers of stage I–II non-small cell lung cancer. Oncotarget.
8:94980–94996. 2017.PubMed/NCBI
|
|
6
|
He Y, Lin J, Kong D, Huang M, Xu C, Kim
TK, Etheridge A, Luo Y, Ding Y and Wang K: Current state of
circulating microRNAs as cancer biomarkers. Clin Chem.
61:1138–1155. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Tiberio P, Callari M, Angeloni V, Daidone
MG and Appierto V: Challenges in using circulating miRNAs as cancer
biomarkers. Biomed Res Int. 2015:7314792015. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Armand-Labit V and Pradines A: Circulating
cell-free microRNAs as clinical cancer biomarkers. Biomol Concepts.
8:61–81. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Larrea E, Sole C, Manterola L, Goicoechea
I, Armesto M, Arestin M, Caffarel MM, Araujo AM, Araiz M,
Fernandez-Mercado M and Lawrie CH: New concepts in cancer
biomarkers: Circulating miRNAs in liquid biopsies. Int J Mol Sci.
17:pii: E6272016. View Article : Google Scholar
|
|
10
|
Mestdagh P, Hartmann N, Baeriswyl L,
Andreasen D, Bernard N, Chen C, Cheo D, D'Andrade P, DeMayo M,
Dennis L, et al: Evaluation of quantitative miRNA expression
platforms in the microRNA quality control (miRQC) study. Nat
Methods. 11:809–815. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Schwarzenbach H, da Silva AM, Calin G and
Pantel K: Data normalization strategies for microRNA
quantification. Clin Chem. 61:1333–1342. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Wang K, Zhang S, Weber J, Baxter D and
Galas DJ: Export of microRNAs and microRNA-protective protein by
mammalian cells. Nucleic Acids Res. 38:7248–7259. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Arroyo JD, Chevillet JR, Kroh EM, Ruf IK,
Pritchard CC, Gibson DF, Mitchell PS, Bennett CF,
Pogosova-Agadjanyan EL, Stirewalt DL, et al: Argonaute2 complexes
carry a population of circulating microRNAs independent of vesicles
in human plasma. Proc Natl Acad Sci USA. 108:5003–5008. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Cortez MA, Bueso-Ramos C, Ferdin J,
Lopez-Berestein G, Sood AK and Calin GA: MicroRNAs in body
fluids-the mix of hormones and biomarkers. Nat Rev Clin Oncol.
8:467–477. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Vickers KC, Palmisano BT, Shoucri BM,
Shamburek RD and Remaley AT: MicroRNAs are transported in plasma
and delivered to recipient cells by high-density lipoproteins. Nat
Cell Biol. 13:423–433. 2011. View
Article : Google Scholar : PubMed/NCBI
|
|
16
|
Turchinovich A, Weiz L and Burwinkel B:
Extracellular miRNAs: The mystery of their origin and function.
Trends Biochem Sci. 37:460–465. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Bianchi F, Nicassio F, Marzi M, Belloni E,
Dall'olio V, Bernard L, Pelosi G, Maisonneuve P, Veronesi G and Di
Fiore PP: A serum circulating miRNA diagnostic test to identify
asymptomatic high-risk individuals with early stage lung cancer.
EMBO Mol Med. 3:495–503. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Chen X, Hu Z, Wang W, Ba Y, Ma L, Zhang C,
Wang C, Ren Z, Zhao Y, Wu S, et al: Identification of ten serum
microRNAs from a genome-wide serum microRNA expression profile as
novel noninvasive biomarkers for nonsmall cell lung cancer
diagnosis. Int J Cancer. 130:1620–1628. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Markou A, Sourvinou I, Vorkas PA, Yousef
GM and Lianidou E: Clinical evaluation of microRNA expression
profiling in non small cell lung cancer. Lung Cancer. 81:388–396.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
The World Health Organization histological
typing of lung tumours. Second edition. Am J Clin Pathol.
77:123–136. 1982. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Goldstraw P, Crowley J, Chansky K, Giroux
DJ, Groome PA, Rami-Porta R, Postmus PE, Rusch V and Sobin L;
International Association for the Study of Lung Cancer
International Staging Committee, ; Participating Institutions, :
The IASLC lung cancer staging project: proposals for the revision
of the TNM stage groupings in the forthcoming (seventh) edition of
the TNM Classification of malignant tumours. J Thorac Oncol.
2:706–714. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Kroh EM, Parkin RK, Mitchell PS and Tewari
M: Analysis of circulating microRNA biomarkers in plasma and serum
using quantitative reverse transcription-PCR (qRT-PCR). Methods.
50:298–301. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Wang P, Yang D, Zhang H, Wei X, Ma T,
Cheng Z, Hong Q, Hu J, Zhuo H, Song Y, et al: Early detection of
lung cancer in serum by a panel of MicroRNA biomarkers. Clin Lung
Cancer. 16:313–319.e1. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Wang RJ, Zheng YH, Wang P and Zhang JZ:
Serum miR-125a-5p, miR-145 and miR-146a as diagnostic biomarkers in
non-small cell lung cancer. Int J Clin Exp Pathol. 8:765–771.
2015.PubMed/NCBI
|
|
25
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Das AV and Pillai RM: Implications of miR
cluster 143/145 as universal anti-oncomiRs and their dysregulation
during tumorigenesis. Cancer Cell Int. 15:922015. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Danielson LS, Menendez S, Attolini CS,
Guijarro MV, Bisogna M, Wei J, Socci ND, Levine DA, Michor F and
Hernando E: A differentiation-based microRNA signature identifies
leiomyosarcoma as a mesenchymal stem cell-related malignancy. Am J
Pathol. 177:908–917. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
de Rie D, Abugessaisa I, Alam T, Arner E,
Arner P, Ashoor H, Åström G, Babina M, Bertin N, Burroughs AM, et
al: An integrated expression atlas of miRNAs and their promoters in
human and mouse. Nat Biotechnol. 35:872–878. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Fan L, Qi H, Teng J, Su B, Chen H, Wang C
and Xia Q: Identification of serum miRNAs by nano-quantum dots
microarray as diagnostic biomarkers for early detection of
non-small cell lung cancer. Tumour Biol. 37:7777–7784. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Arab A, Karimipoor M, Irani S, Kiani A,
Zeinali S, Tafsiri E and Sheikhy K: Potential circulating miRNA
signature for early detection of NSCLC. Cancer Genet.
216–217:1–158. 2017.
|
|
31
|
Taverna S, Giallombardo M, Gil-Bazo I,
Carreca AP, Castiglia M, Chacártegui J, Araujo A, Alessandro R,
Pauwels P, Peeters M and Rolfo C: Exosomes isolation and
characterization in serum is feasible in non-small cell lung cancer
patients: Critical analysis of evidence and potential role in
clinical practice. Oncotar. 7:28748–28760. 2016. View Article : Google Scholar
|
|
32
|
Lopatina T, Gai C, Deregibus MC, Kholia S
and Camussi G: Cross talk between cancer and mesenchymal stem cells
through extracellular vesicles carrying nucleic acids. Front Oncol.
6:1252016. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Le HB, Zhu WY, Chen DD, He JY, Huang YY,
Liu XG and Zhang YK: Evaluation of dynamic change of serum miR-21
and miR-24 in pre- and post-operative lung carcinoma patients. Med
Oncol. 29:3190–3197. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Aushev VN, Zborovskaya IB, Laktionov KK,
Girard N, Cros MP, Herceg Z and Krutovskikh V: Comparisons of
microRNA patterns in plasma before and after tumor removal reveal
new biomarkers of lung squamous cell carcinoma. PLoS One.
8:e786492013. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Li J, Liu Y, Wang C, Deng T, Liang H, Wang
Y, Huang D, Fan Q, Wang X, Ning T, et al: Serum miRNA expression
profile as a prognostic biomarker of stage II/III colorectal
adenocarcinoma. Sci Rep. 5:129212015. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Leidinger P, Keller A, Backes C, Huwer H
and Meese E: MicroRNA expression changes after lung cancer
resection: A follow-up study. RNA Biol. 9:900–910. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Sanfiorenzo C, Ilie MI, Belaid A, Barlési
F, Mouroux J, Marquette CH, Brest P and Hofman P: Two panels of
plasma microRNAs as non-invasive biomarkers for prediction of
recurrence in resectable NSCLC. PLoS One. 8:e545962013. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Tang D, Shen Y, Wang M, Yang R, Wang Z,
Sui A, Jiao W and Wang Y: Identification of plasma microRNAs as
novel noninvasive biomarkers for early detection of lung cancer.
Eur J Cancer Prev. 22:540–548. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Geng Q, Fan T, Zhang B, Wang W, Xu Y and
Hu H: Five microRNAs in plasma as novel biomarkers for screening of
early-stage non-small cell lung cancer. Respir Res. 15:1492014.
View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Yu H, Jiang L, Sun C, Li Guo L, Lin M,
Huang J and Zhu L: Decreased circulating miR-375: A potential
biomarker for patients with non-small-cell lung cancer. Gene.
534:60–65. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Yang JS, Li BJ, Lu HW, Chen Y, Lu C, Zhu
RX, Liu SH, Yi QT, Li J and Song CH: Serum miR-152, miR-148a,
miR-148b, and miR-21 as novel biomarkers in non-small cell lung
cancer screening. Tumour Biol. 36:3035–3042. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Lv S, Xue J, Wu C, Wang L, Wu J, Xu S,
Liang X and Lou J: Identification of a panel of serum microRNAs as
biomarkers for early detection of lung adenocarcinoma. J Cancer.
8:48–56. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
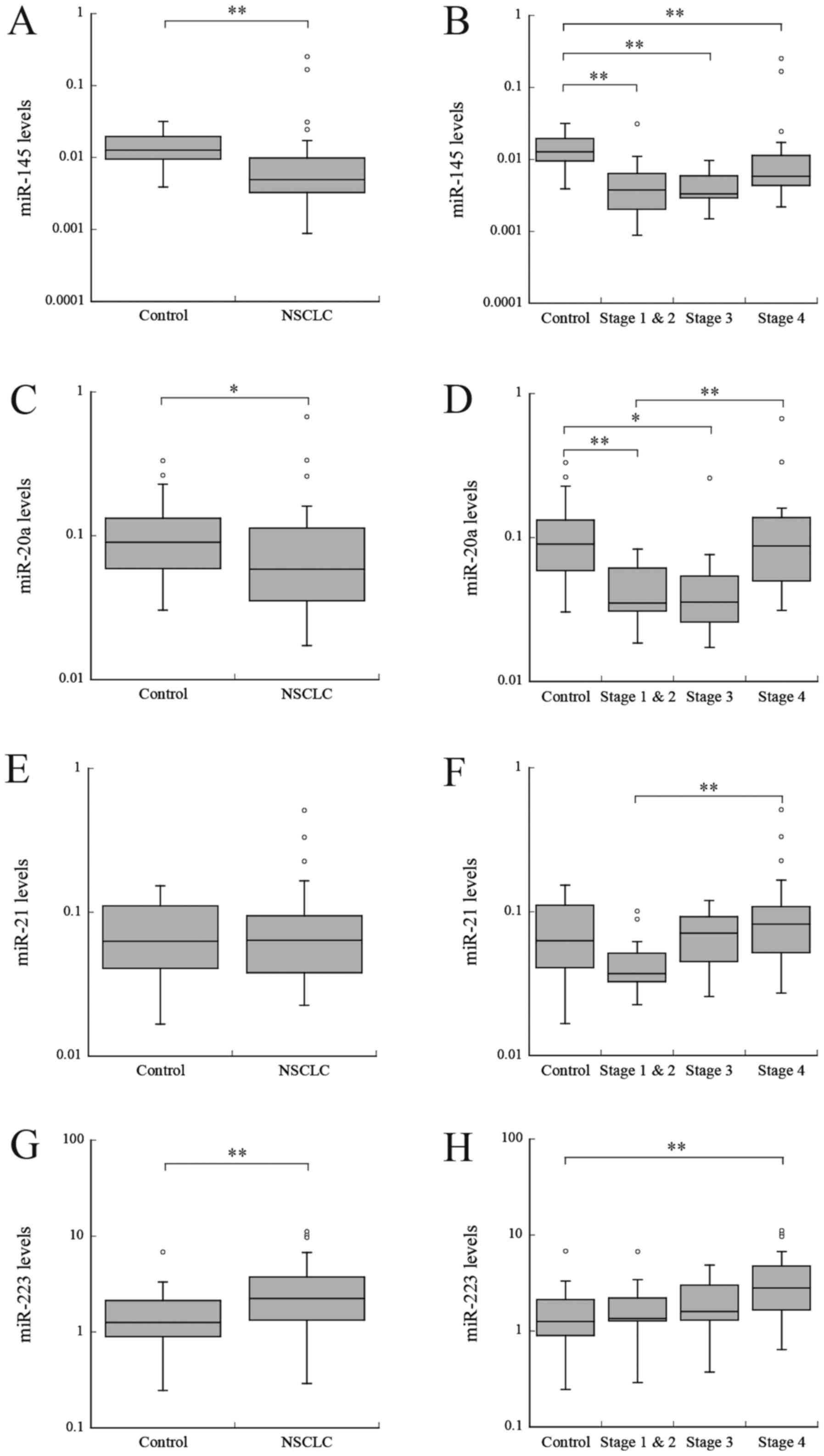
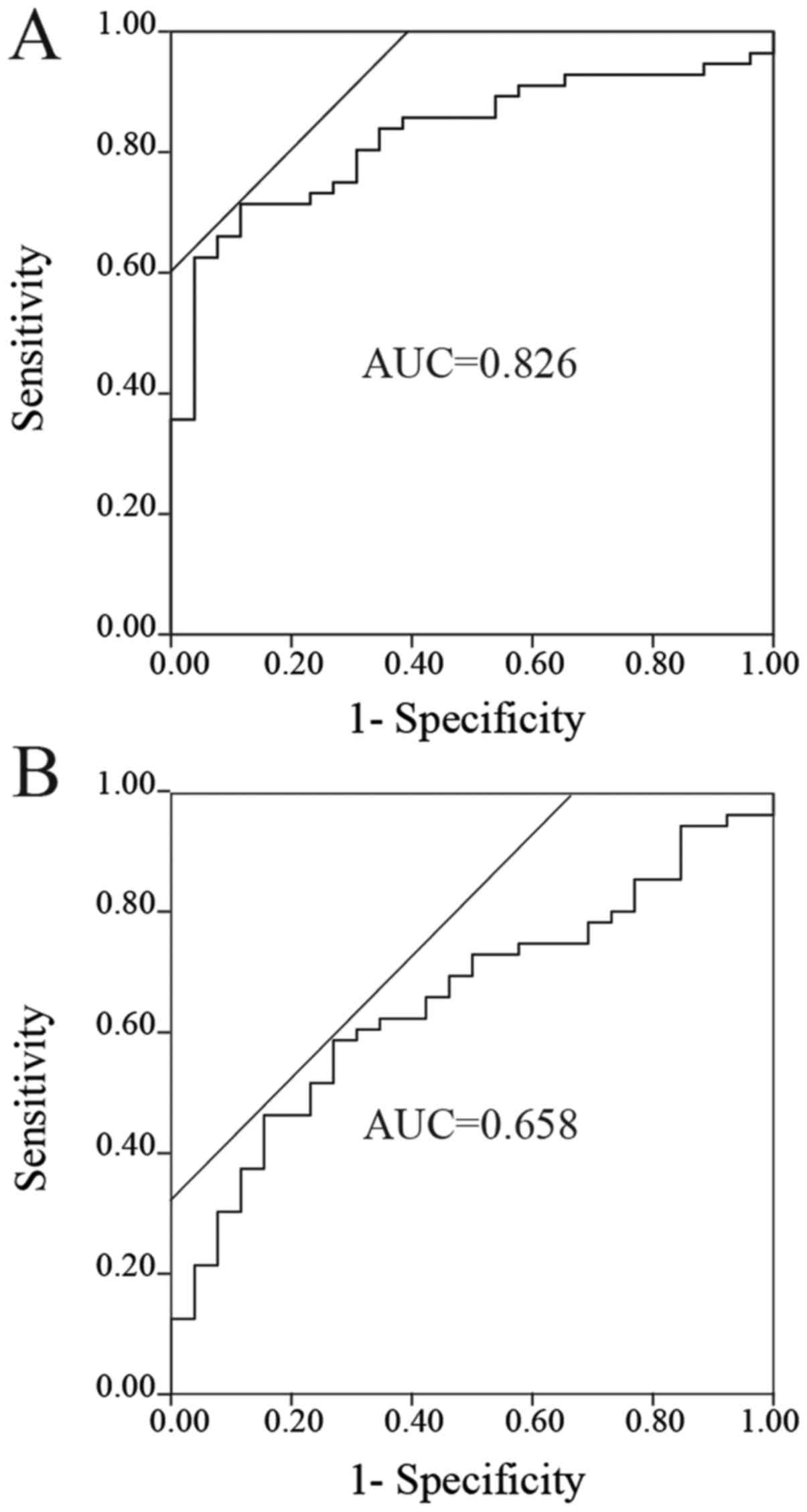
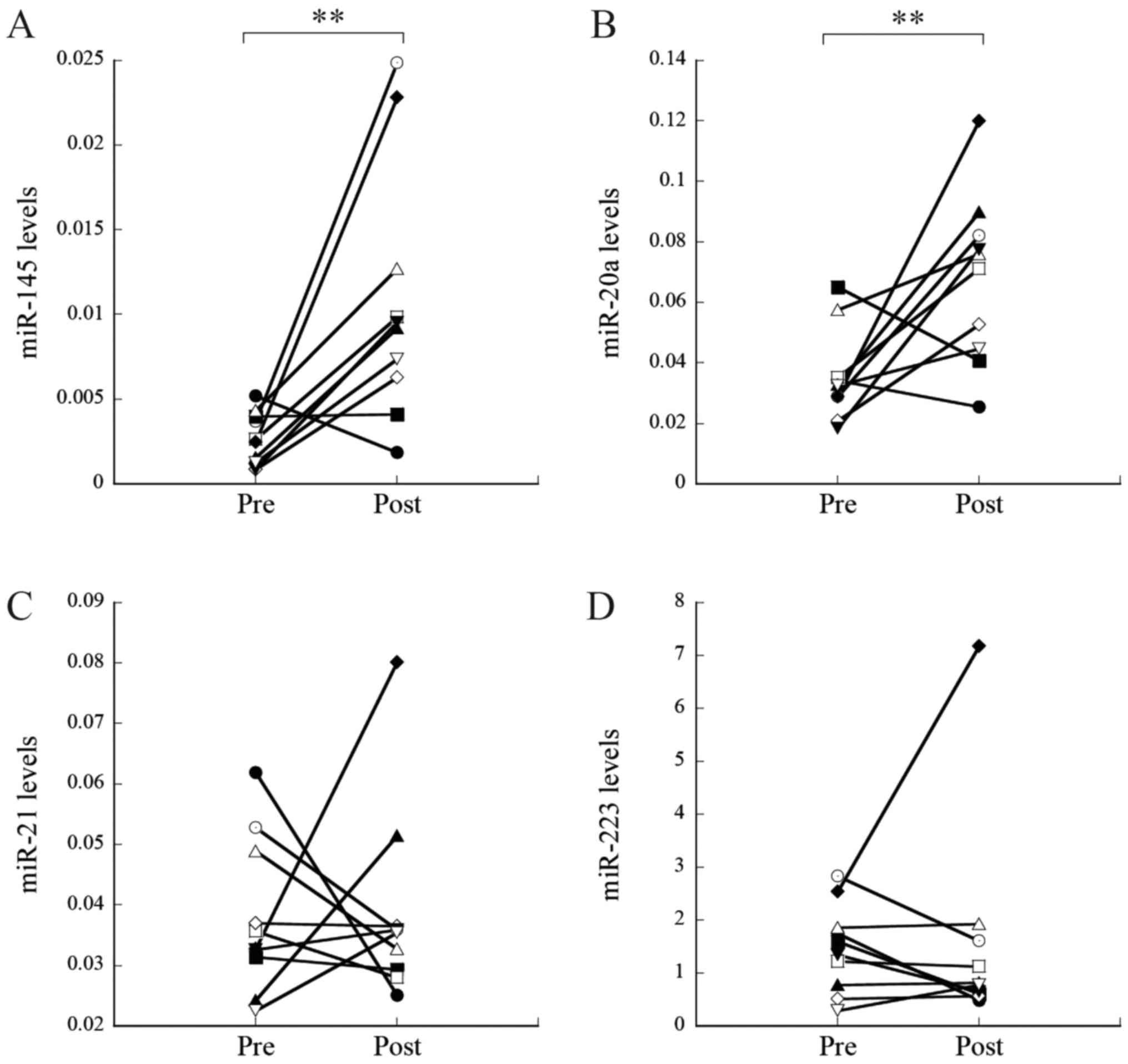
Zhang H, Mao F, Shen T, Luo Q, Ding Z,
Qian L and Huang J: Plasma miR-145, miR-20a, miR-21 and miR-223 as
novel biomarkers for screening early-stage non-small cell lung
cancer. Oncol Lett. 13:669–676. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Zhou X, Wen W, Shan X, Zhu W, Xu J, Guo R,
Cheng W, Wang F, Qi LW, Chen Y, et al: A six-microRNA panel in
plasma was identified as a potential biomarker for lung
adenocarcinoma diagnosis. Oncotarget. 8:6513–6525. 2017.PubMed/NCBI
|
|
45
|
O'Driscoll L: Extracellular nucleic acids
and their potential as diagnostic, prognostic and predictive
biomarkers. Anticancer Res. 27:1257–1265. 2007.PubMed/NCBI
|
|
46
|
Zampetaki A and Mayr M: Analytical
challenges and technical limitations in assessing circulating
miRNAs. Thromb Haemost. 108:592–598. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Xiang M, Zeng Y, Yang R, Xu H, Chen Z,
Zhong J, Xie H, Xu Y and Zeng X: U6 is not a suitable endogenous
control for the quantification of circulating microRNAs. Biochem
Biophys Res Commun. 454:210–214. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Sromek M, Glogowski M, Chechlinska M,
Kulinczak M, Szafron L, Zakrzewska K, Owczarek J, Wisniewski P,
Wlodarczyk R, Talarek L, et al: Changes in plasma miR-9, miR-16,
miR-205 and miR-486 levels after non-small cell lung cancer
resection. Cell Oncol (Dordr). 40:529–536. 2017. View Article : Google Scholar : PubMed/NCBI
|