|
1
|
Johnson BE: Divide and conquer to treat
lung cancer. N Engl J Med. 375:1892–1893. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Chen Z, Fillmore CM, Hammerman PS, Kim CF
and Wong KK: Non-small-cell lung cancers: A heterogeneous set of
diseases. Nat Rev Cancer. 14:535–546. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Siegel R, Naishadham D and Jemal A: Cancer
statistics, 2013. CA Cancer J Clin. 63:11–30. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Schiller JH, Harrington D, Belani CP,
Langer C, Sandler A, Krook J, Zhu J and Johnson DH; Eastern
Cooperative Oncology Group, : Comparison of four chemotherapy
regimens for advanced non-small-cell lung cancer. N Engl J Med.
346:92–98. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Petty RD, Nicolson MC, Kerr KM,
Collie-Duguid E and Murray GI: Gene expression profiling in
non-small cell lung cancer: From molecular mechanisms to clinical
application. Clin Cancer Res. 10:3237–3248. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Cheng N, Li X, Zhao C, Ren S, Chen X, Cai
W, Zhao M, Zhang Y, Li J, Wang Q and Zhou C: Microarray expression
profile of long non-coding RNAs in EGFR-TKIs resistance of human
non-small cell lung cancer. Oncol Rep. 33:833–839. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Wilusz JE, Sunwoo H and Spector DL: Long
noncoding RNAs: Functional surprises from the RNA world. Genes Dev.
23:1494–1504. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Zhou M, Guo M, He D, Wang X, Cui Y, Yang
H, Hao D and Sun J: A potential signature of eight long non-coding
RNAs predicts survival in patients with non-small cell lung cancer.
J Transl Med. 13:2312015. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Ghadimi K, Bahrami N, Fathi M, Farzanegan
B, Naji T, Emami M and Mohamadnia A: Diagnostic value of LunX mRNA
and CEA mRNA expression in pleural fluid of patients with non-small
cell lung cancer. Minerva Pneumol. 56:90–95. 2017.
|
|
10
|
Zhou HX, Yang MX, Wang Y, Cao WM, Lu KF,
Zong LY, Wu RQ and Zhang P: Plasma LUNX mRNA, a non-invasive
specific biomarker for diagnosis and prognostic prediction of
non-small cell lung cancer. Am J Cancer Res. 6:452–458.
2016.PubMed/NCBI
|
|
11
|
Xu C, Xu Y, Liao X, Liao R, Zhang L, Niu
K, Li T, Li D, Chen Z, Duan Y and Sun J: Plasma miRNAs in
predicting radiosensitivity in non-small cell lung cancer. Tumour
Biol. 37:11927–11936. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Wu C and Arora P: Long noncoding
RNA-microRNA-mRNA: A novel tripartite axis in the regulation of
cardiac hypertrophy. Circ Cardiovasc Genet. 7:729–731. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Ye S, Yang L, Zhao X, Song W, Wang W and
Zheng S: Bioinformatics method to predict two regulation mechanism:
TF-miRNA-mRNA and lncRNA-miRNA-mRNA in pancreatic cancer. Cell
Biochem Biophys. 70:1849–1858. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Yang ZH, Zheng R, Gao Y, Zhang Q and Zhang
H: Abnormal gene expression and gene fusion in lung adenocarcinoma
with high-throughput RNA sequencing. Cancer Gene Ther. 21:74–82.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Leidinger P, Brefort T, Backes C, Krapp M,
Galata V, Beier M, Kohlhaas J, Huwer H, Meese E and Keller A:
High-throughput qRT-PCR validation of blood microRNAs in non-small
cell lung cancer. Oncotarget. 7:4611–4623. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
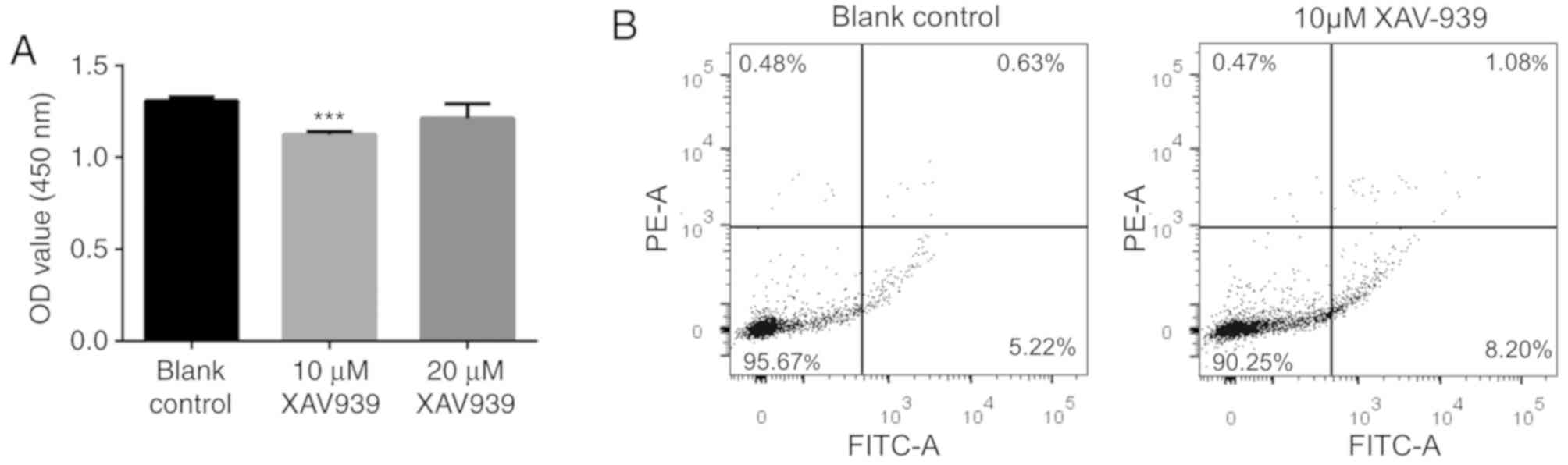
Tian X, Hou W, Bai S, Fan J, Tong H and
Bai Y: XAV939 promotes apoptosis in a neuroblastoma cell line via
telomere shortening. Oncol Rep. 32:1999–2006. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Guo W, Shen F, Xiao W, Chen J and Pan F:
Wnt inhibitor XAV939 suppresses the viability of small cell lung
cancer NCI-H446 cells and induces apoptosis. Oncol Lett.
14:6585–6591. 2017.PubMed/NCBI
|
|
18
|
Lin HH, Feng WC, Lu LC, Shao YY, Cheng AL
and Hsu CH: Abstract 2052: WNT/beta-catenin signaling inhibitors
improve the anti-proliferative effect of sorafenib against
hepatocellular carcinoma (HCC) cells. Cancer Res. 73:20522013.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Liu Y, Su D and Song T: Programmed cell
death 4 inhibits proliferation and differentiation and induces
apoptosis of human mesenchymal stem cells through suppressing the
Wnt/β-catenin pathway. RSC Adv. 7:26566–26573. 2017. View Article : Google Scholar
|
|
20
|
Ran M, Chen B and Li Z, Wu M, Liu X, He C,
Zhang S and Li Z: Systematic identification of long non-coding RNAs
in immature and mature porcine testes. Biol Reprod. 94:772016.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Bolger AM, Lohse M and Usadel B:
Trimmomatic: A flexible trimmer for Illumina sequence data.
Bioinformatics. 30:2114–2120. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Harrow J, Frankish A, Gonzalez JM,
Tapanari E, Diekhans M, Kokocinski F, Aken BL, Barrell D, Zadissa
A, Searle S, et al: GENCODE: The reference human genome annotation
for The ENCODE Project. Genome Res. 22:1760–1774. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Trapnell C, Pachter L and Salzberg SL:
TopHat: Discovering splice junctions with RNA-Seq. Bioinformatics.
25:1105–1111. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Lun AT, Chen Y and Smyth GK: It's
DE-licious: A recipe for differential expression analyses of
RNA-seq experiments using quasi-likelihood methods in edgeR.
Methods Mol Biol. 1418:391–416. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Gene Ontology Consortium, . The gene
ontology in 2010: Extensions and refinements. Nucleic Acids Res.
38((Database Issue)): D331–D335. 2010.PubMed/NCBI
|
|
26
|
Huang DW, Sherman BT, Tan Q, Kir J, Liu D,
Bryant D, Guo Y, Stephens R, Baseler MW, Lane HC and Lempicki RA:
DAVID bioinformatics resources: Expanded annotation database and
novel algorithms to better extract biology from large gene lists.
Nucleic Acids Res. 35((Web Server Issue)): W169–W175. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Szklarczyk D, Franceschini A, Wyder S,
Forslund K, Heller D, Huerta-Cepas J, Simonovic M, Roth A, Santos
A, Tsafou KP, et al: STRING v10: Protein-protein interaction
networks, integrated over the tree of life. Nucleic Acids Res.
43((Database Issue)): D447–D452. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Shannon P, Markiel A, Ozier O, Baliga NS,
Wang JT, Ramage D, Amin N, Schwikowski B and Ideker T: Cytoscape: A
software environment for integrated models of biomolecular
interaction networks. Genome Res. 13:2498–2504. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Opsahl T, Agneessens F and Skvoretz J:
Node centrality in weighted networks: Generalizing degree and
shortest paths. Social Networks. 32:245–251. 2010. View Article : Google Scholar
|
|
30
|
Pearson K: Note on regression and
inheritance in the case of two parents. Proc R Soc Lond.
58:240–242. 1895. View Article : Google Scholar
|
|
31
|
Benjamini Y and Hochberg Y: Controlling
the false discovery rate: A practical and powerful approach to
multiple testing. J R Statist Soc. 57:289–300. 1995.
|
|
32
|
Yu G, Wang LG, Han Y and He QY:
clusterProfiler: An R package for comparing biological themes among
gene clusters. OMICS. 16:284–287. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Resnik P: Semantic similarity in a
taxonomy: An information-based measure and its application to
problems of ambiguity in natural language. J Artif Intell Res.
11:95–130. 1999. View Article : Google Scholar
|
|
34
|
Wang JZ, Du Z, Payattakool R, Yu PS and
Chen CF: A new method to measure the semantic similarity of GO
terms. Bioinformatics. 23:1274–1281. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Yu G, Li F, Qin Y, Bo X, Wu Y and Wang S:
GOSemSim: An R package for measuring semantic similarity among GO
terms and gene products. Bioinformatics. 26:976–978. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Kuleshov MV, Jones MR, Rouillard AD,
Fernandez NF, Duan Q, Wang Z, Koplev S, Jenkins SL, Jagodnik KM,
Lachmann A, et al: Enrichr: A comprehensive gene set enrichment
analysis web server 2016 update. Nucleic Acids Res. 44((W1)):
W90–W97. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Zhan Y, Zang H, Feng J, Lu J, Chen L and
Fan S: Long non-coding RNAs associated with non-small cell lung
cancer. Oncotarget. 8:69174–69184. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Zheng K, Wu G, Qin X, Wang Y, Xia S and
Meng X: Effects of XAV939 on proliferation and glycolysis of
hepatocellular carcinoma cells. J Mod Oncol. 13:2023–2026.
2016.
|
|
39
|
Teng H, Wang P, Xue Y, Liu X, Ma J, Cai H,
Xi Z, Li Z and Liu Y: Role of HCP5-miR-139-RUNX1 feedback loop in
regulating malignant behavior of glioma cells. Mol Ther.
24:1806–1822. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Muys BR, Lorenzi JC, Zanette DL, Lima e
Bueno Rde B, de Araújo LF, Dinarte-Santos AR, Alves CP, Ramão A, de
Molfetta GA, Vidal DO and Silva WA Jr: Placenta-Enriched LincRNAs
MIR503HG and linc00629 decrease migration and invasion potential of
JEG-3 cell line. PLoS One. 11:e01515602016. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Pachera E, Assassi S, Cintora GS,
Frank-Bertoncelj M, Haunerdinger V, Dobrota R, Brock M, Vettori S,
Hellerbrand C, Feghali-Bostwick CA, et al: OP0284 long noncoding
RNA MIR503HG is a novel factor in the pathogenesis of systemic
sclerosis. Ann Rheum Dis. 74 (Suppl 2):S180.1–180. 2015. View Article : Google Scholar
|
|
42
|
Grimminger PP, Maus MK, Schneider PM,
Metzger R, Hölscher AH, Sugita H, Danenberg PV, Alakus H and
Brabender J: Glutathione S-transferase PI (GST-PI) mRNA expression
and DNA methylation is involved in the pathogenesis and prognosis
of NSCLC. Lung Cancer. 78:87–91. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Scharer CD, Mccabe CD, Aliseyed M, Berger
MF, Bulyk ML and Moreno CS: Genome-wide promoter analysis of the
SOX4 transcriptional network in prostate cancer cells. Cancer Res.
69:709–717. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Chen QL, Zheng WL, Yao WJ, Nie LW, Cheng
SH and Ma WL: Analysis of SOX4 gene mutation in non-small cell lung
cancer tissues. Zhonghua Yi Xue Yi Chuan Xue Za Zhi. 24:505–509.
2007.PubMed/NCBI
|
|
45
|
Tang T, Huan L, Zhang S, Zhou H, Gu L,
Chen X and Zhang L: MicroRNA-212 functions as a tumor-suppressor in
human non-small cell lung cancer by targeting SOX4. Oncol Rep.
38:2243–2250. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Li Y, Chen P, Zu L, Liu B, Wang M and Zhou
Q: MicroRNA-338-3p suppresses metastasis of lung cancer cells by
targeting the EMT regulator Sox4. Am J Cancer Res. 6:127–140.
2016.PubMed/NCBI
|
|
47
|
Foss KM, Sima C, Ugolini D, Neri M, Allen
KE and Weiss GJ: miR-1254 and miR-574-5p: Serum-based microRNA
biomarkers for early-stage non-small cell lung cancer. J Thorac
Oncol. 6:482–488. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Lee CW, Kang MR, Yun J, Oh SJ and Kang JS:
Abstract 4387: Up-regulation of VHL by miR-1273C inhibits renal
cell carcinoma. Cancer Res. 74:43872014. View Article : Google Scholar
|
|
49
|
Li C, Zheng X, Han Y, Lv Y, Lan F and Zhao
J: XAV939 inhibits the proliferation and migration of lung
adenocarcinoma A549 cells through the WNT pathway. Oncol Lett.
15:8973–8982. 2018.PubMed/NCBI
|
|
50
|
Lachaier E, Louandre C, Ezzoukhry Z, Godin
C, Mazière JC, Chauffert B and Galmiche A: Ferroptosis, a new form
of cell death relevant to the medical treatment of cancer. Med Sci
(Paris). 30:779–783. 2014.(In French). View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Dixon SJ, Lemberg KM, Lamprecht MR, Skouta
R, Zaitsev EM, Gleason CE, Patel DN, Bauer AJ, Cantley AM, Yang WS,
et al: Ferroptosis: An iron-dependent form of nonapoptotic cell
death. Cell. 149:1060–1072. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Jiang Y, He Y, Liu S and Tao Y: Chromatin
remodeling factor lymphoid-specific helicase inhibits ferroptosis
through lipid metabolic genes in lung cancer progression. Chin J
Cancer. 36:822017. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Alvarez SW, Sviderskiy VO, Terzi EM,
Papagiannakopoulos T, Moreira AL, Adams S, Sabatini DM, Birsoy K
and Possemato R: NFS1 undergoes positive selection in lung tumours
and protects cells from ferroptosis. Nature. 551:639–643. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
54
|
He L, Vasiliou K and Nebert DW: Analysis
and update of the human solute carrier (SLC) gene superfamily. Hum
Genomics. 3:195–206. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Sehm T, Rauh M, Wiendieck K, Buchfelder M,
Eyüpoglu IY and Savaskan NE: Temozolomide toxicity operates in a
xCT/SLC7a11 dependent manner and is fostered by ferroptosis.
Oncotarget. 7:74630–74647. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Jiang L: p53 promotes ferroptosis during
ROS stress to suppress tumorigenesis. Cancer Dis. 5:4652015.
View Article : Google Scholar
|
|
57
|
Ji XJ, Qian J, Rahman J, Harris B,
Hoeksema MD, Chen H, Eisenberg R and Young J: Abstract A10: SLC7A11
contributes to the pathogenesis of lung cancer. Mol Cancer Res.
14:A102016. View Article : Google Scholar
|