Introduction
Hepatocellular carcinoma (HCC) is one of the most
common and frequently fatal malignances worldwide, with a life
expectancy of ~6 months from diagnosis (1–3).
Although treatment options for HCC have improved, at present,
resection and transplantation remain the only curative approaches
(4–6). Advances in curative treatment
strategies for HCC are partially being held back by a lack of
understanding of the molecular mechanisms surrounding the
pathogenesis of HCC. Therefore, elucidating the mechanisms
underlying HCC pathogenesis and progression is of significant
importance in improving the therapeutic options for HCC.
HCC is typically caused by chronic hepatitis B or C
virus infection and, to a lesser extent, through chronic exposure
to toxins or hereditary liver diseases (7,8). There
has been an increase in the identification of disease-associated
risk factors, such as histone deacetylases (HDACs) and vimentin, as
well as the signaling pathways involved, including activation of
the Wnt signaling pathway and inactivation of the cellular tumor
antigen p53 signaling pathway (9).
However, the complex nature of this disease has complicated
advances in understanding the pathological mechanisms of HCC.
HDACs and histone acetylases together regulate gene
expression by histone acetylation and de-acetylation. Aberrant
expression of these two enzymes may result in epigenetic alteration
and, consequently, dysregulated gene expression (10,11).
HDAC1 is the most extensively studied HDAC family member, and its
expression is increased in a range of different types of cancer,
including HCC, prostate cancer and gastric cancer (12–14).
Furthermore, upregulated expression of HDAC1 has been associated
with a decrease in tissue differentiation and decreased survival
rates (12,15–18).
Vimentin is an intermediate filament protein in the cytoplasm of
mesenchymal cells, and has frequently been used as a marker to
demonstrate epithelial-mesenchymal transition, which is associated
with the progression of cancer (19). Previous studies have demonstrated
that vimentin expression is associated with HCC progression and
that the serum vimentin level may serve as an additional marker of
HCC, suggesting a role of vimentin protein in HCC pathogenesis
(20–22).
Although HDAC1 and vimentin are both important
factors in HCC pathogenesis and progression, their roles in this
disease have usually been studied separately, and whether there are
interactions between HDAC1 and vimentin that affect HCC has not
been investigated (12,21). A previous study demonstrated an
association between HDAC1 dysregulation and vimentin expression in
HCC (23); however, the underlying
mechanisms have not been determined. Whether a causal relationship
between HDAC1 and vimentin expression exists was investigated in
the present study. The present data may provide valuable
information regarding potential therapeutic targets.
Materials and methods
Cell lines, plasmids and small
interfering (si)RNAs
The HCC cell line Hep3B and the normal human liver
cell line THLE-3 were purchased from The Type Culture Collection of
the Chinese Academy of Sciences, (Shanghai, China) and cultured in
Dulbecco's modified Eagle's medium (DMEM) supplemented with 10%
fetal bovine serum (both Gibco; Thermo Fisher Scientific, Inc.,
Waltham, MA, USA), penicillin (100 U/ml), and streptomycin (100
mg/ml) at 37°C in a humidified incubator containing 5%
CO2.
Human HDAC1, p65 and p50 gene sequences were
amplified from human cDNA library and subcloned into a pcDNA3.1(+)
vector (Thermo Fisher Scientific, Inc.). Gene sequences were
verified by Sanger sequencing. The control Renilla
luciferase reporter gene plasmid and the firefly luciferase
reporter gene vector pGL3-Basic were purchased from Promega
Corporation, (Madison, WI, USA). The vimentin promoter sequence was
amplified from the genome of THLE-3 cells, subcloned into
pGL3-Basic and termed ‘(−800/+72)Vimentin’. Vimentin promoter
truncations and mutations at the NF-κB and PEA3 binding sites were
performed on the full length (−800/+72)Vimentin and termed
‘(−725/+72)Vimentin’, ‘(−353/+72)Vimentin’, ‘(−261/+72)Vimentin’,
(−200/+72)Vimentin, NF-κB MUT and PEA3 MUT, respectively. The
sequences of the primers used for plasmid construction are listed
in Table I. HDAC1 siRNA (cat. no.
sc-29343) and control siRNA (cat. no. sc-37007) were purchased from
Santa Cruz Biotechnology, Inc. (Dallas, TX, USA).
 | Table I.Primers used for plasmid
construction. |
Table I.
Primers used for plasmid
construction.
| Primer name | Primer
sequence | Plasmid name |
|---|
| V-800-F |
5′-TAGGTACCGCCCGTTAGGTCCCTCGACA-3′ |
(−800/+72)Vimentin |
| V-725-F |
5′-TAGGTACCCGGGCCGGAGCAGCCCCCCT-3′ |
(−725/+72)Vimentin |
| V-353-F |
5′-TAGGTACCCCCAGCCCAGCGCTGAAGTA-3′ |
(−353/+72)Vimentin |
| V-261-F |
5′-TAGGTACCCCCCGCTTCTCGCTAGGTCC-3′ |
(−261/+72)Vimentin |
| V-200-F |
5′-GGGACCCTCTTTCCTAACGG-3′ |
(−200/+72)Vimentin |
| V-R |
5′-TCGGCCGGCTCGCGGTGCCC-3′ | Reverse primer for
all the vimentin plasmids |
| N-F |
5′-TCCCTATTGGATAATGCGCTCCGCGG-3′ | NF-κB MUT |
| N-R |
5′-CCTAGCGAGAAGCGGGGA-3′ |
|
| P-F |
5′-TTCCTAACGGAACCTTAAAAACAGCGCCCTCGG-3′ | PEA3 MUT |
| P-R |
5′-AGAGGGTCCCCTCCCACT-3′ |
|
| H-F |
5′-TAGAATTCATGGCGCAGACGCAGGGCAC-3′ | HDAC1 |
| H-R |
5′-TACTCGAGTCAGGCCAACTTGACCTCCT-3′ |
|
| p65-F |
5′-TAGGTACCATGGGACCCGAAAACGAGAG-3′ | p65 |
| p65-R |
5′-GAGAATTCTCAGGTCTGGGGACTTGCTT-3′ |
|
| p50-F |
5′-GCGGTACCATGGCAGAAGATGATCCATA-3′ | p50 |
| p50-R | 5′
GCGAATTCCTAAATTTTGCCTTCTAGAG-3′ |
|
Reverse transcription-quantitative
polymerase chain reaction (RT-qPCR)
RT-qPCR was performed as previously described, with
certain modifications (24). The
total RNA was extracted from cells using TRIzol® reagent
(Thermo Fisher Scientific, Inc.) and reverse-transcribed into cDNA
using Moloney murine leukemia virus reverse transcriptase (M-MLV
RT) (Promega Corporation). Total RNA (2 µg) was first mixed with
0.5 µg Oligo(dT)18 primer and heated to 70°C for 5 min.
After heating, the sample was immediately cooled on ice.
Subsequently, 5 µl 5X reaction buffer, 1.25 µl 10 mM dATP, 1.25 µl
10 mM dCTP, 1.25 µl 10 mM dGTP, 1.25 µl 10 mM dTTP, 25 units
recombinant RNasin ribonuclease inhibitor (Takara Biotechnology
Co., Inc.) and 200 units M-MLV RT (Promega Corporation) were added
and mixed, and incubated at 42°C for 60 min. Subsequently, the
vimentin RNA level was quantified using SYBR Green (Bio-Rad
Laboratories, Inc., Hercules, CA, USA), with GAPDH as an internal
control. The following thermal cycling protocol was used:
Polymerase activation and DNA denaturation (95°C, 1 min); 40 cycles
of denaturation (95°C, 10 sec) and annealing/extension (60°C, 30
sec), followed by melt-curve analysis (65–95°C with 0.5°C
increments, 2–5 sec/step). The following primers were used for
RT-qPCR. The primers for vimentin were: Forward,
5′-AGGAAATGGCTCGTCACCTTCGTGAATA-3′; reverse,
5′-GGAGTGTCGGTTGTTAAGAACTAGAGCT-3′. The primers for GAPDH were:
Forward, 5′-AGCCACATCGCTCAGACA-3′; reverse,
5′-TGGACTCCACGACGTACT-3′. The relative vimentin mRNA expression
level was calculated using the 2−ΔΔCq method (25).
Subcellular fractionation
Cell nuclear and cytoplasmic fractions were prepared
using NE-PER™ Nuclear and Cytoplasmic Extraction kit (Thermo Fisher
Scientific, Inc.), according to the manufacturer's protocol. Cells
were harvested and resuspended in DNA endonuclease I-Cre(I) by
vortexing for 15 sec. Subsequently, the cell suspension was mixed
with CRE II by vortexing for 10 sec and centrifuged at 16,000 × g
for 5 min at 4°C to obtain the cytoplasmic fraction from the
supernatant. The insoluble pellet, which contained the nuclear
fraction, was resuspended in the nuclear extraction reagent and
vortexed for 15 sec. After centrifugation at 16,000 × g for 10 min
at 4°C, the nuclear fraction was obtained from the supernatant.
Western blotting
For analysis using whole cell lysate, cells were
harvested and lysed using lysis buffer (Thermo Fisher Scientific,
Inc.) supplemented with protease inhibitors (Roche Applied Science,
Penzberg, Germany), and mixed with SDS-PAGE loading buffer. For
analysis using cytoplasmic and nuclear fractions, prepared
fractions were mixed with SDS-PAGE loading buffer directly. Protein
concentration was quantified using the Pierce BCA Protein Assay kit
(Thermo Fisher Scientific, Inc.) and 20 µg whole cell lysate or 10
µg cytoplasmic and nuclear fractions were separated using a 12%
SDS-PAGE gel and transferred onto a PVDF membrane. GAPDH was used
as a loading control. Non-specific binding sites on the membrane
were blocked with 5% skimmed milk for 1 h at room temperature. The
membrane was sequentially incubated with primary and corresponding
horseradish peroxidase (HRP)-conjugated secondary antibodies for 2
and 1 h at room temperature, respectively. After extensive washes,
immunobands were visualized using ECL substrate (Merck KGaA,
Darmstadt, Germany,) under a charge-couple device camera (Bio-Rad
Laboratories, Inc., Hercules, CA, USA). The following primary
antibodies were used in this study: Anti-HDAC1 antibody (cat. no.
ab19845; Abcam, Cambridge, UK), anti-p65 antibody (cat. no. sc-372;
Santa Cruz Biotechnology, Inc.), anti-vimentin antibody (cat. no.
ab92547; Abcam) and anti-GAPDH antibody (cat. no. g8795;
Sigma-Aldrich; Merck KGaA). HRP-conjugated secondary antibodies
[Goat Anti-Mouse IgG (H+L)-HRP; cat. no. BA1050 and Goat
Anti-Rabbit IgG (H+L)-HRP; cat. no. BA1056] were both obtained from
Wuhan Boster Biological Technology, Ltd., (Wuhan, China). All the
primary antibodies were used at a 1:1,000 dilution, and all the
secondary antibodies were used at a 1:100,000 dilution.
Transfection and luciferase reporter
gene activity assay
The luciferase reporter gene activity assay was
performed as previously described with modifications (26). Cells were first transfected with
firefly luciferase reporter gene plasmids with full length vimentin
promoter (−800/+72)Vimentin, vimentin truncations
(−725/+72)Vimentin, (−353/+72)Vimentin, (−261/+72)Vimentin,
(−200/+72)Vimentin, or the vimentin mutations NF-κB MUT or PEA3
MUT, in addition to an HDAC1 plasmid and control Renilla
luciferase reporter gene plasmid using Lipofectamine™ 2000,
according to the manufacturer's protocol (Thermo Fisher Scientific,
Inc.). Cells were incubated for 24 h post transfection, after which
they were lysed with lysis buffer and firefly and Renilla
luciferase activities were measured using Promega Dual Luciferase
Assay kit and the GloMax®-Multi Detection System,
according to the manufacturer's protocol (both from Promega
Corporation). The transfection efficiency was normalized to
Renilla luciferase activity. For the siRNA knockdown assay,
p65 siRNA (sc-29410; Santa Cruz Biotechnology, Inc.) or control
siRNA (sc-37007; Santa Cruz Biotechnology, Inc.) at a final
concentration of 100 nM was introduced into cells using X-tremeGENE
siRNA Transfection Reagent (Roche Applied Science) 24 h prior to
plasmid transfection. For the signaling pathway inhibition assay,
an NF-κB inhibitor (Celastrol; 300 nM) or Janus kinase (JNK)
inhibitor (SP600125; 10 µM; both InvivoGen. Inc., Toulouse, France)
was added to the cell culture 4–6 h post transfection. For p65 +
p50 co-transfection, plasmids encoding p65 and p50 at a ratio of
1:1 were mixed and transfected into cells using
Lipofectamine® 2000 according to the manufacturer's
protocol (Thermo Fisher Scientific. Inc.).
Chromatin immunoprecipitation (ChIP)
assay
ChIP analysis was performed using a Pierce™ Magnetic
ChIP kit (Thermo Fisher Scientific, Inc.) according to the
manufacturer's protocol. All reagents used were included in the kit
unless otherwise stated. Cells were first crosslinked with 1%
formaldehyde, lysed with membrane extraction buffer and digested
with MNase, and the chromatin smear was obtained by sonication
(3×20 sec pulses at 3 W power on ice with a 20-sec incubation on
ice between pulses). The obtained chromatin smear was then
sequentially incubated with 5 µg of either anti-p65 antibody (cat.
no. sc-372; Santa Cruz Biotechnology, Inc.) or normal rabbit IgG
overnight at 4°C and magnetic protein A/G beads for 2 h at 4°C.
Subsequent to washings, precipitated DNA was recovered from
magnetic beads with elution buffer and a PCR was performed with
Q5® High-Fidelity DNA Polymerase (New England BioLabs,
Inc., Ipswich, MA, USA) using vimentin promoter-specific primers:
Forward primer, 5′-GGGCTCCATGAGTCATATCC-3′ and reverse primer,
5′-ATCTGGCTCAAGACCTTTGC-3′. The following thermal cycling
conditions were used: Initial denaturation, 98°C, 30 sec; 35 of
cycles of denaturation (98°C, 10 sec), annealing (55°C, 10 sec) and
elongation (72°C, 30 sec); and final extension (72°C, 2 min).
Statistical analysis
All experiments were repeated three times. Data were
presented as mean ± standard deviation. A Student's t test or a
one-way analysis of variance with a Student-Newman-Keuls post hoc
were used to compare the differences between two groups or three or
more groups, respectively. P<0.05 was considered to indicate a
statistically significant difference. All statistical analysis was
performed with GraphPad PRISM 4.0.3 (GraphPad Software, Inc., La
Jolla, CA, USA).
Results
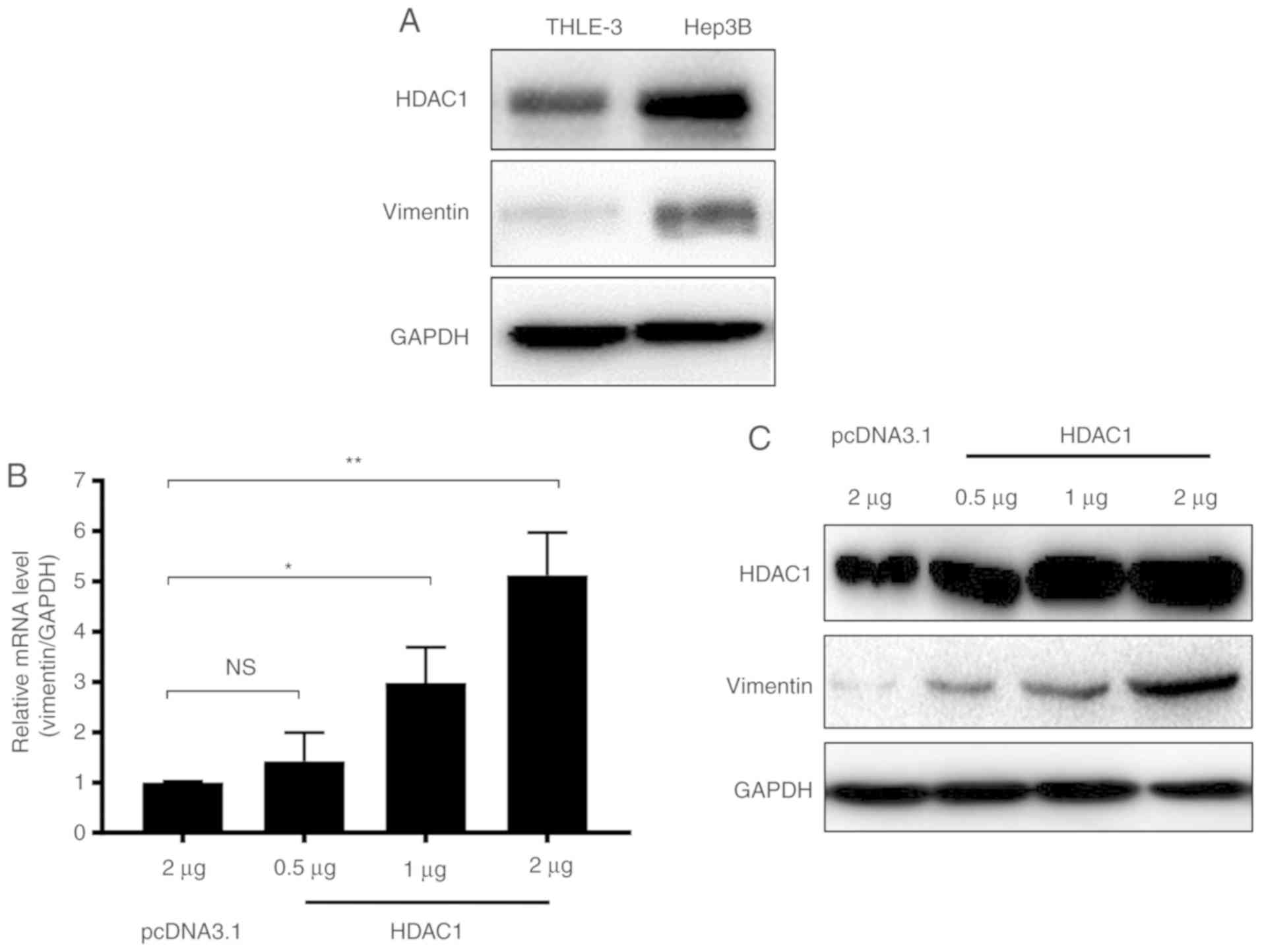
HDAC1 overexpression results in
increased vimentin expression
A previous study demonstrated that HDAC1
downregulation may result in a decrease in vimentin expression in
HCC (23). In the present study, to
confirm the association between HDAC1 and vimentin expression in
HCC, the expression of HDAC1 and vimentin in THLE-3 normal liver
cells and Hep3B HCC cells were determined. HDAC1 protein expression
levels were markedly increased in Hep3B cells compared with THLE-3
cells (Fig. 1A). Notably, similar to
HDAC1, the expression of vimentin also markedly increased in Hep3B
cells (Fig. 1A).
To further investigate whether vimentin expression
was associated with HDAC1 expression, THLE-3 cells were transfected
with increasing quantities of HDAC1, and the mRNA and protein
levels of vimentin were measured. Vimentin mRNA expression levels
were increased significantly when ≥1 µg of HDAC1 was transfected
(P<0.05; Fig. 1B). Western
blotting demonstrated that vimentin protein expression levels were
additionally increased in an HDAC1 dose dependent manner (Fig. 1C). Together, these data suggested
that HDAC1 and vimentin are increased in HCC cells, and HDAC1
expression may upregulate vimentin expression.
An NF-κB transcription factor-binding
site in the vimentin promoter is associated with HDAC1-induced
vimentin expression
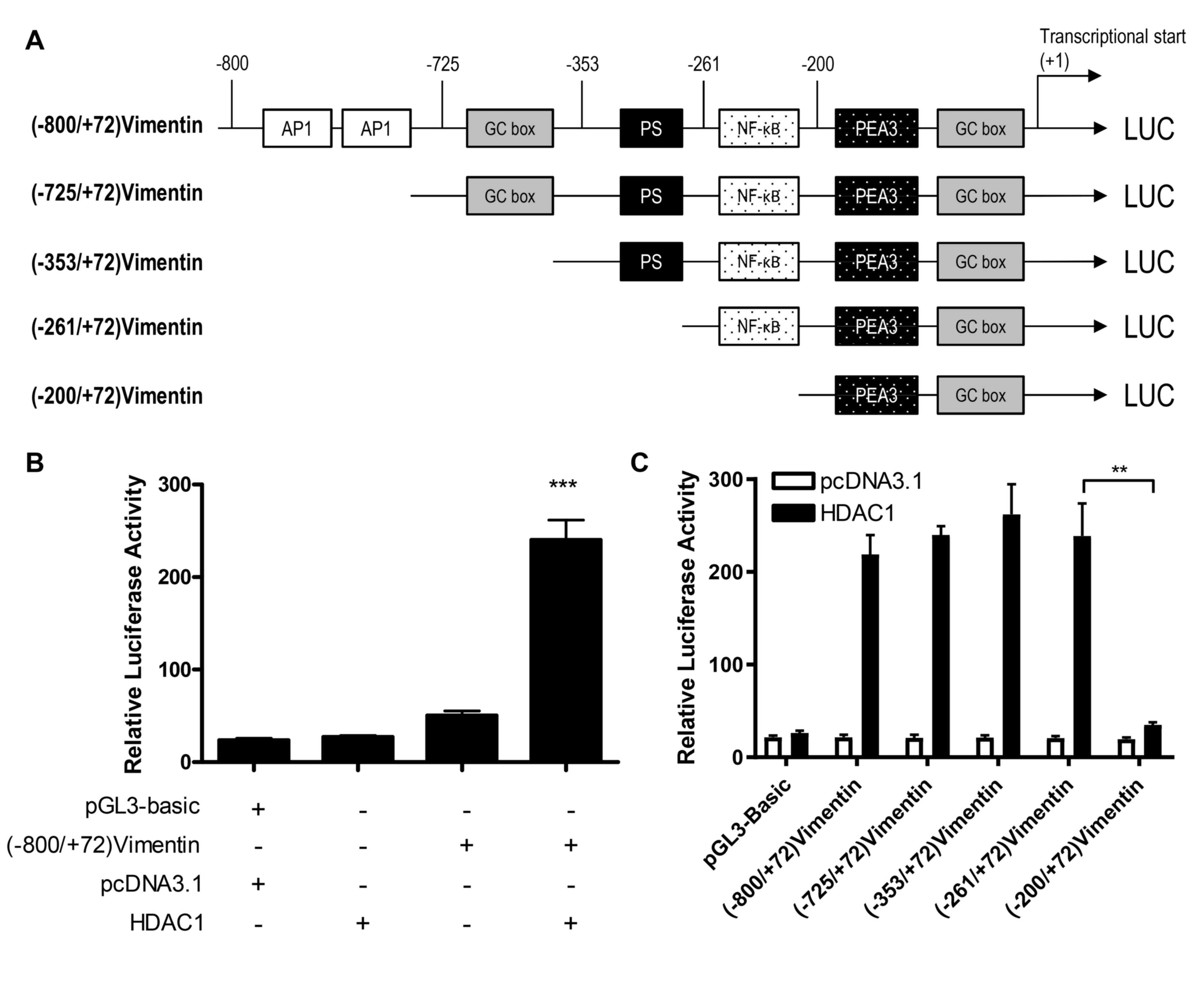
To determine whether HDAC1 upregulated vimentin
expression through the transactivation of a vimentin promoter,
luciferase reporter gene plasmids were constructed with gene
expression under the control of a full length or a 5-flanking
region of a truncated vimentin promoter sequence (Fig. 2A), and the activation of these
plasmids by HDAC1 was determined. HDAC1 overexpression
significantly activated the vimentin promoter (P<0.001 vs.
pcDNA3.1; Fig. 2B). Through the use
of truncation mutants, it was demonstrated that HDAC1-mediated
activation of the vimentin promoter was abrogated when the sequence
between −261 and −200 was removed; suggesting that the sequence in
this region was necessary for HDAC1 induced activation (Fig. 2C).
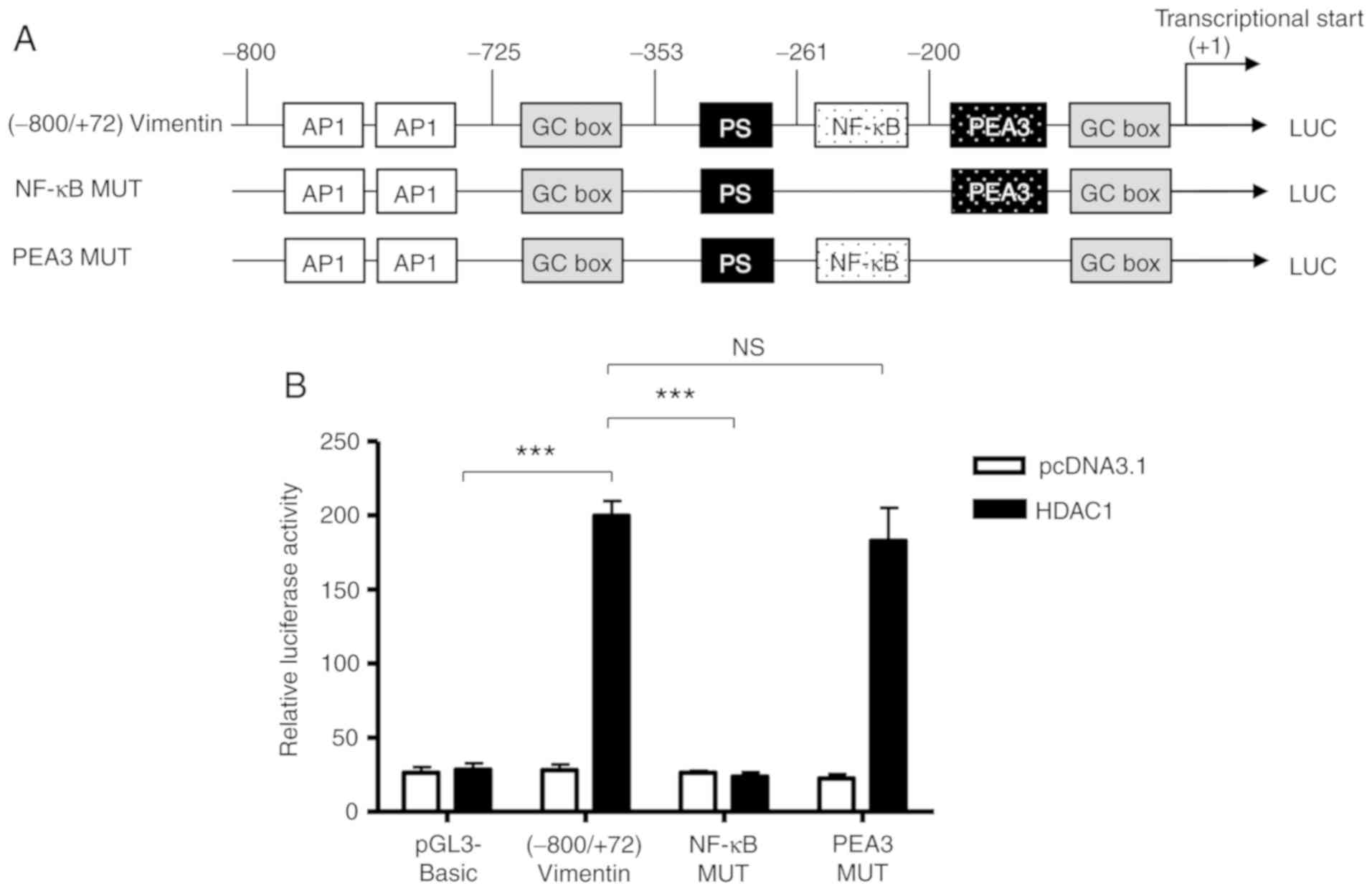
Based on the bioinformatics analysis performed
previously (26), there is a NF-κB
binding site between −261 and −200 (Fig.
2). To determine whether the NF-κB binding site was involved in
the HDAC-1 overexpression-induced vimentin promoter activation, the
ability of HDAC1 to activate a full-length vimentin promoter with a
mutated NF-κB binding site was determined (Fig. 3A). As shown in Figure. 3B, HDAC1 overexpression was unable
to activate the NF-κB mutated vimentin promoter; however, HDAC1
mediated activation of the vimentin promoter was still
significantly increased when the NF-κB neighboring domain, PEA3 was
mutated, with a potency similar to (−800/+72)Vimentin. The data
presented in Fig. 2C suggest that
the NF-κB binding site between-261 and −200 in vimentin promoter is
involved in the HDAC1-induced vimentin expression.
HDAC1 induces vimentin expression
through the NF-κB signaling pathway
To determine whether HDAC1 overexpression could
promote NF-κB binding with the vimentin promoter, a ChIP assay was
performed. The vimentin promoter sequence was not pulled down by an
anti-p65 antibody when HDAC1 was not overexpressed. However,
following HDAC1 overexpression, the vimentin promoter sequence was
detected in the anti-p65 antibody pull-down, indicating that HDAC1
overexpression could result in the binding of NF-κB with the
vimentin promoter (Fig. 4A). The
control immunoglobulin G antibody did not result in a pull-down of
the vimentin promoter sequence irrespective of HDAC1 expression
(Fig. 4A).
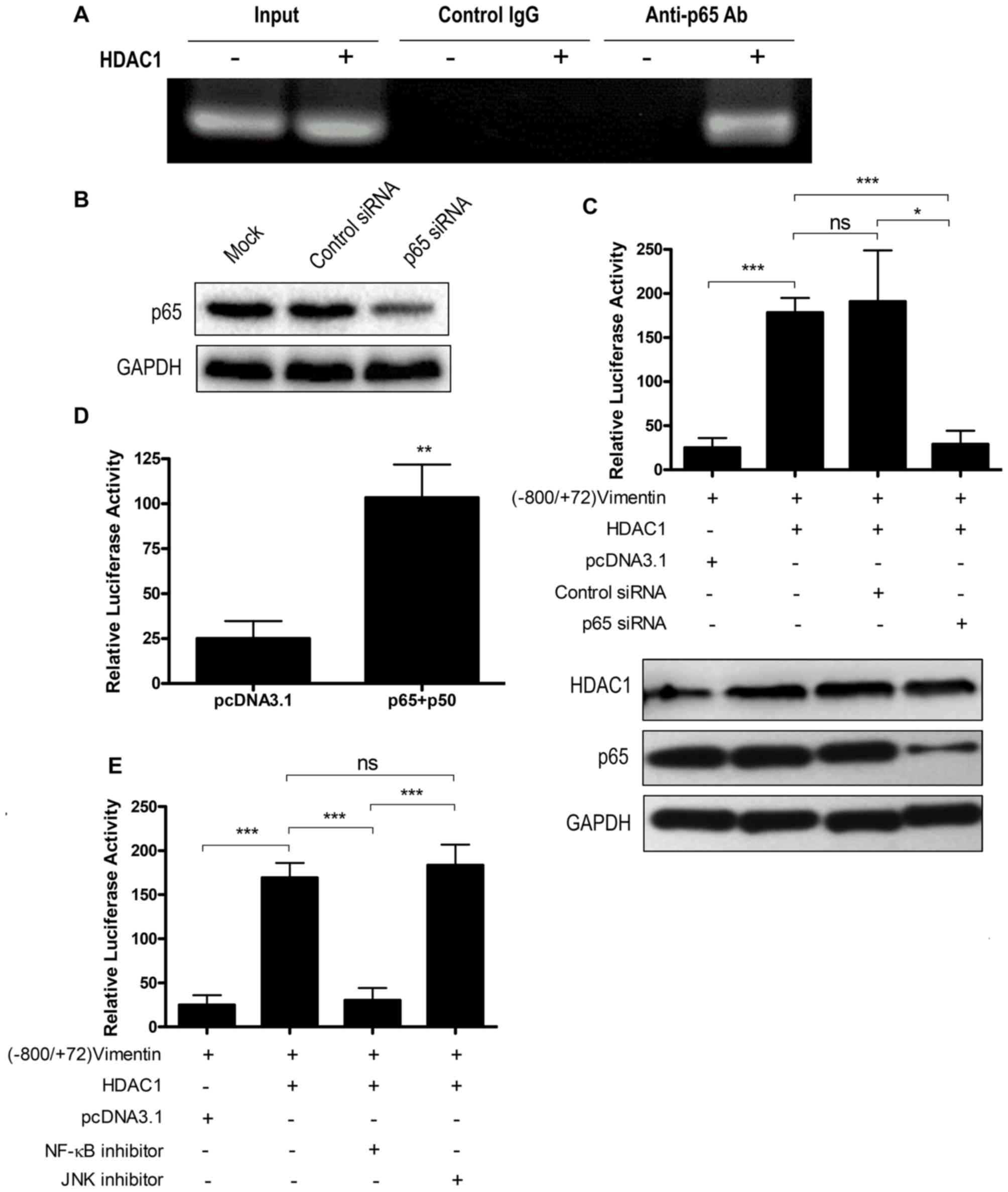 | Figure 4.HDAC1 overexpression upregulates
vimentin expression via the NF-κB signaling pathway. (A) Chromatin
immunoprecipitation assay using an anti-p65 antibody to detect
NF-κB binding to vimentin promoter. (B) THLE-3 cells were mock
transfected or transfected with control siRNA or p65 siRNA and p65
expression was determined by western blotting. (C) THLE-3 cells
were treated with p65 siRNA or control siRNA, and were further
transfected with (−800/+72) Vimentin together with or without HDAC1
and the luciferase activity was measured. ns, not significant;
*P<0.05, ***P<0.001. (D) THLE-3 cells were first transfected
with (−800/+72) Vimentin together with sham vector or a combination
of p65 and p50 and the luciferase activity was measured.
**P<0.01 vs. pcDNA3.1. (E) THLE-3 cells were transfected with
(−800/+72) Vimentin with or without HDAC1 for 4–6 h, then treated
with NF-κB or JNK signaling pathway inhibitors and the luciferase
activity was measured. ns, not significant; ***P<0.001. HDAC1,
histone deacetylase 1; NF-κB, nuclear factor κ-light-chain-enhancer
of activated B cells; p50, NF-κB p105 subunit; p65, transcription
factor p65; siRNA, small interfering RNA. |
To confirm the participation of NF-κB in HDAC1
overexpression-induced vimentin expression, a p65 knockdown by
siRNA interference was performed. p65 siRNA knockdown efficiency
was first confirmed by western blotting (Fig. 4B). p65 knockdown abrogated the
activation of the vimentin promoter by HDAC1 overexpression
(P<0.05), whereas control siRNA interference had no effect
(P>0.05; Fig. 4C). NF-κB
overexpression may increase the protein expression levels of all
the NF-κB forms, including the unphosphorylated form in the
cytoplasm and the phosphorylated form in the nucleus (27), and the increase of phosphorylated
NF-κB may enhance vimentin expression. Therefore, the role of NF-κB
overexpression on the transactivation of vimentin was investigated.
The results demonstrated that NF-κB (p65+p50) overexpression could
activate the vimentin promoter (P<0.01; Fig. 4D).
To investigate the involvement of the NF-κB
signaling pathway in HDAC1 overexpression-induced vimentin
expression, a signaling pathway inhibition assay was performed.
Vimentin promoter activation was significantly decreased when NF-κB
inhibitor (P<0.001), but not JNK inhibitor, was added (Fig. 4E). Taken together, these data
indicated that HDAC1 overexpression-induced vimentin upregulation
via the NF-κB signaling pathway.
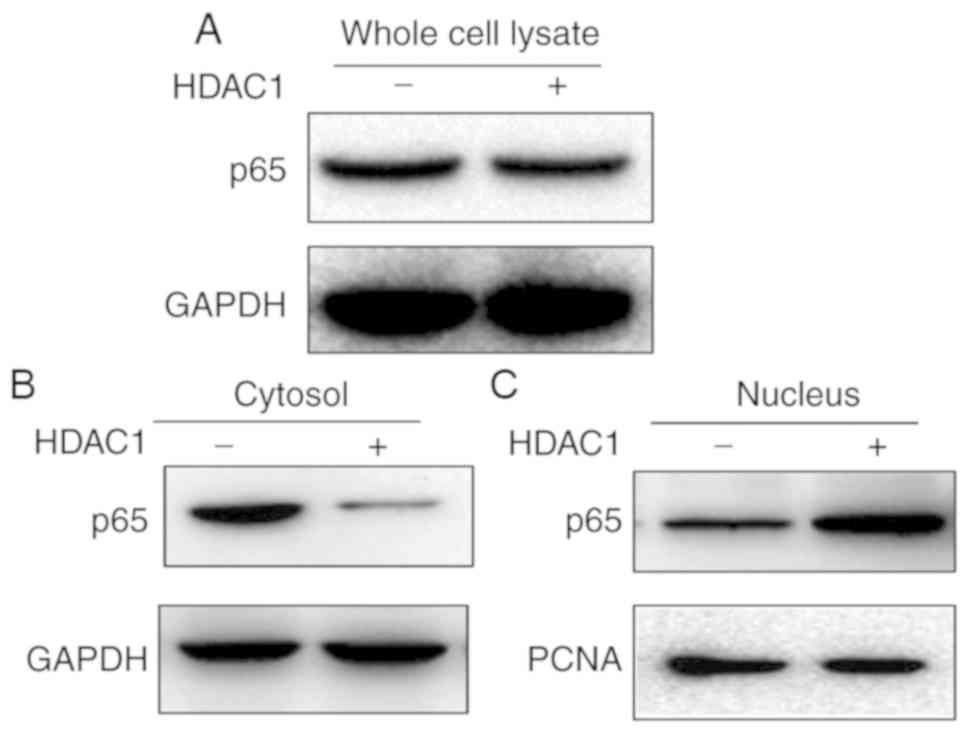
HDAC-1 induces NF-κB translocation
from the cytoplasm into the nucleus
As the NF-κB signaling pathway was modulated in
HDAC1-induced vimentin expression, the impact of HDAC1
overexpression on NF-κB expression and translocation was
subsequently investigated. First, the whole cell lysate of THLE-3
cells with or without HDAC1 overexpression were tested for p65
expression. As presented in Fig. 5A,
p65 expression did not demonstrate any apparent alterations in
expression before or after HDAC1 overexpression. The cytoplasmic
and nucleic fractions were further separated and the expression
levels of p65 in both fractions were determined. p65 was present in
both fractions when HDAC1 was not overexpressed (Fig. 5B). However, when HDAC1 was
overexpressed, a near complete translocation of p65 from the
cytoplasm into the nucleus was observed, indicating that HDAC1
overexpression modulates the NF-κB signaling pathway by promoting
NF-κB translocation from the cytoplasm into the nucleus.
Together, these data demonstrated that HDAC1 is
overexpressed in HCC and this HDAC1 overexpression could upregulate
vimentin expression through the NF-κB signaling pathway. The
present study demonstrated a causal relationship between HDAC1 and
vimentin in HCC, providing valuable information for understanding
the pathogenesis of HCC, as well as serving as potential targets
for the treatment of HCC.
Discussion
HDACs are enzymes involved in the transcriptional
regulation of genes; the expression of HDACs is increased in a
number of different types of tumors, including HCC, prostate cancer
and gastric cancer (12–14) and HDAC inhibitors are effective in
suppressing tumor growth at an early stage (28). In HCC, the best studied HDAC family
member is HDAC1, which is expressed at aberrantly high levels in
HCC cells (29,30). Vimentin, one of the most widely
expressed type III intermediate filament proteins, is significantly
increased in a number of different types of solid tumors and is
regularly used as a marker for epithelial-mesenchymal transition, a
process closely associated with tumor progression (31). In HCC, the association of elevated
vimentin expression with tumor development, especially metastasis,
has been widely studied and the serum vimentin expression level is
used as a marker for HCC (21,22,32,33).
Although HDAC1 and vimentin have been widely studied in HCC and
other tumors, the association between them has not been
investigated, to the best of our knowledge. A previous study
demonstrated that downregulation of HDAC1 resulted in a decrease of
vimentin expression, indicating a potential link between these two
proteins. In the present study a causal relationship was confirmed
between HDAC1 and vimentin. Furthermore, the signaling pathway
responsible for the HDAC1 overexpression-induced vimentin
expression was identified. The findings of our current study would
not only provide information for the understanding of the
pathogenesis of HCC, but also provide potential targets for this
disease or possibly other solid tumors.
There are 18 known HDACs in mammals and the majority
of them have been associated with tumor progression, and HDACs have
been demonstrated to function distinctively in cancer cells
(34,35). For example, HDAC2, but not HDAC1, has
a suppressive effect on cell growth in breast cells (34). Additionally, knockdown of HDAC1 and
HDAC2 may exert differing effects on cell survival (35). In HCC, HDAC1 and HDAC2 are both
associated with tumor growth (28).
However, HDAC1 and HDAC2 possess a high degree of resemblance in
their genomic sequence (36).
Therefore, it may be interesting to investigate, albeit beyond the
scope of the present study, whether HDAC2 has an effect on vimentin
expression and whether the effect is different from that of HDAC1,
as well as the potential mechanisms.
Regulation of vimentin expression has been
investigated by other studies and different signaling pathways are
involved under different circumstances. In IL-6- promoted head and
neck tumor metastasis, vimentin expression is activated through the
JAK/STAT3/SNAIL signaling pathway (37), while in tubular epithelial cells,
hypoxia-induced vimentin expression is mediated through the Notch
signaling pathway (38). TGF-β1
regulates vimentin expression in skeletal myogenic cells through a
signaling pathway involving Smad/AP1/SP1 (24). In the present study, HDAC1
overexpression-induced vimentin expression in HCC cells was through
the NF-κB signaling pathway. Given the complex regulation network
surrounding vimentin expression (39), whether the NF-κB signaling pathway is
also responsible for vimentin expression in other tumors remains to
be determined.
Together, the present study demonstrated that HDAC1
is overexpressed in HCC and that HDAC1 overexpression could
upregulate vimentin expression through the NF-κB signaling pathway
in HCC cells. Furthermore a causal relationship between HDAC1 and
vimentin in HCC was identified, which may improve our understanding
of HCC pathogenesis, as well as identifying potential targets for
HCC treatment.
Acknowledgements
Not applicable.
Funding
The present study was supported by Natural Science
Foundation of Guangdong Province, Initiation Project for PhD (grant
no. S2012040007235).
Availability of data and materials
The datasets used and/or analyzed during the present
study are available from the corresponding author on reasonable
request.
Authors' contributions
HZ and JX designed the experiment. HZ and CZ
performed the assays. HZ and YW analyzed the data. HZ wrote the
manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
World-Health-Organization, . Cancer (fact
sheet) 2018. https://www.who.int/en/news-room/fact-sheets/detail/cancerSeptember
12–2018
|
|
2
|
Ito Y, Takeda T, Higashiyama S, Sakon M,
Wakasa KI, Tsujimoto M, Monden M and Matsuura N: Expression of
heparin binding epidermal growth factor-like growth factor in
hepatocellular carcinoma: An immunohistochemical study. Oncol Rep.
8:903–907. 2001.PubMed/NCBI
|
|
3
|
Forner A, Llovet JM and Bruix J:
Hepatocellular carcinoma. Lancet. 379:1245–1255. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Bosch FX, Ribes J, Diaz M and Cleries R:
Primary liver cancer: Worldwide incidence and trends.
Gastroenterology. 127:S5–S16. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Waly Raphael S, Yangde Z and YuXiang C:
Hepatocellular carcinoma: Focus on different aspects of management.
ISRN Oncol. 2012:4216732012.PubMed/NCBI
|
|
6
|
Llovet JM, Burroughs A and Bruix J:
Hepatocellular carcinoma. Lancet. 362:1907–1917. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Morgan RL, Baack B, Smith BD, Yartel A,
Pitasi M and Falck-Ytter Y: Eradication of Hepatitis C virus
infection and the development of hepatocellular carcinomaa: A
meta-analysis of observational studies. Ann Intern Med.
158:329–337. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Gordon SC, Lamerato LE, Rupp LB, Li J,
Holmberg SD, Moorman AC, Spradling PR, Teshale EH, Vijayadeva V,
Boscarino JA, et al: Antiviral therapy for chronic hepatitis B
virus infection and development of hepatocellular carcinoma in a US
population. Clin Gastroenterol Hepatol. 12:885–893. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Kirstein MM and Vogel A: The pathogenesis
of hepatocellular carcinoma. Dig Dis. 32:545–553. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Kaliman P, Álvarez-López MJ, Cosín-Tomás
M, Rosenkranz MA, Lutz A and Davidson RJ: Rapid changes in histone
deacetylases and inflammatory gene expression in expert meditators.
Psychoneuroendocrinology. 40:96–107. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Shankar S and Srivastava RK: Histone
deacetylase inhibitors: mechanisms and clinical significance in
cancer: HDAC inhibitor-induced apoptosis. Adv Exp Med Biol.
615:261–298. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Rikimaru T, Taketomi A, Yamashita Y,
Shirabe K, Hamatsu T, Shimada M and Maehara Y: Clinical
significance of histone deacetylase 1 expression in patients with
hepatocellular carcinoma. Oncology. 72:69–74. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Halkidou K, Gaughan L, Cook S, Leung HY,
Neal DE and Robson CN: Upregulation and nuclear recruitment of
HDAC1 in hormone refractory prostate cancer. Prostate. 59:177–189.
2004. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Choi JH, Kwon HJ, Yoon BI, Kim JH, Han SU,
Joo HJ and Kim DY: Expression profile of histone deacetylase 1 in
gastric cancer tissues. Jpn J Cancer Res. 92:1300–1304. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Quint K, Agaimy A, Di Fazio P, Montalbano
R, Steindorf C, Jung R, Hellerbrand C, Hartmann A, Sitter H,
Neureiter D and Ocker M: Clinical significance of histone
deacetylases 1, 2, 3, and 7: HDAC2 is an independent predictor of
survival in HCC. Virchows Arch. 459:129–139. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Seo J, Min SK, Park HR, Kim DH, Kwon MJ,
Kim LS and Ju YS: Expression of Histone Deacetylases HDAC1, HDAC2,
HDAC3, and HDAC6 in invasive ductal carcinomas of the breast. J
Breast Cancer. 17:323–331. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Ropero S and Esteller M: The role of
histone deacetylases (HDACs) in human cancer. Mol Oncol. 1:19–25.
2007. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Peng L and Seto E: Deacetylation of
nonhistone proteins by HDACs and the implications in cancer. Handb
Exp Pharmacol. 206:39–56. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Thiery JP: Epithelial-mesenchymal
transitions in tumour progression. Nature Rev Cancer. 2:442–454.
2002. View
Article : Google Scholar
|
|
20
|
Hu L, Lau SH, Tzang CH, Wen JM, Wang W,
Xie D, Huang M, Wang Y, Wu MC, Huang JF, et al: Association of
Vimentin overexpression and hepatocellular carcinoma metastasis.
Oncogene. 23:298–302. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Mitra A, Satelli A, Xia X, Cutrera J,
Mishra L and Li S: Cell-surface Vimentin: A mislocalized protein
for isolating csVimentin(+) CD133(−) novel stem-like hepatocellular
carcinoma cells expressing EMT markers. Int J Cancer. 137:491–496.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Sun S, Poon RT, Lee NP, Yeung C, Chan KL,
Ng IO, Day PJ and Luk JM: Proteomics of hepatocellular carcinoma:
Serum vimentin as a surrogate marker for small tumors (≤2 cm). J
Proteome Res. 9:1923–1930. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Zhou H, Wang J, Peng G, Song Y and Zhang
C: A novel treatment strategy in hepatocellular carcinoma by
down-regulation of histone deacetylase 1 expression using a shRNA
lentiviral system. Int J Clin Exp Med. 8:17721–17729.
2015.PubMed/NCBI
|
|
24
|
Wu Y, Zhang X, Salmon M, Lin X and Zehner
ZE: TGFbeta1 regulation of vimentin gene expression during
differentiation of the C2C12 skeletal myogenic cell line requires
Smads, AP-1 and Sp1 family members. Biochim Biophys Acta.
1773:427–439. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
livak kj and schmittgen td: analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Huang W, Hu K, Luo S, Zhang M, Li C, Jin
W, Liu Y, Griffin GE, Shattock RJ and Hu Q: Herpes simplex virus
type 2 infection of human epithelial cells induces CXCL9 expression
and CD4+ T cell migration via activation of
p38-CCAAT/enhancer-binding protein-β pathway. J Immunol.
188:6247–6257. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Lizzul PF, Aphale A, Malaviya R, Sun Y,
Masud S, Dombrovskiy V and Gottlieb AB: Differential expression of
phosphorylated NF-kappaB/RelA in normal and psoriatic epidermis and
downregulation of NF-kappaB in response to treatment with
etanercept. J Invest Dermatol. 124:1275–1283. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Ler SY, Leung CH, Khin LW, Lu GD,
Salto-Tellez M, Hartman M, Iau PT, Yap CT and Hooi SC: HDAC1 and
HDAC2 independently predict mortality in hepatocellular carcinoma
by a competing risk regression model in a Southeast Asian
population. Oncol Rep. 34:2238–2250. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Sun TY, Xie HJ, Li Z, Kong LF and Ding YZ:
Analysis of miRNAs related to abnormal HDAC1 expression in
hepatocellular carcinoma. Int J Clin Exp Med. 9:2016.
|
|
30
|
Yoo YG, Na TY, Seo HW, Seong JK, Park CK,
Shin YK and Lee MO: Hepatitis B virus X protein induces the
expression of MTA1 and HDAC1, which enhances hypoxia signaling in
hepatocellular carcinoma cells. Oncogene. 27:3405–3413. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Mani SA, Guo W, Liao MJ, Eaton EN, Ayyanan
A, Zhou AY, Brooks M, Reinhard F, Zhang CC, Shipitsin M, et al: The
epithelial-mesenchymal transition generates cells with properties
of stem cells. Cell. 133:704–715. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Salvi A, Bongarzone I, Ferrari L, Abeni E,
Arici B, De Bortoli M, Scuri S, Bonini D, Grossi I, Benetti A, et
al: Molecular characterization of LASP-1 expression reveals
vimentin as its new partner in human hepatocellular carcinoma
cells. Int J Oncol. 46:1901–1912. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Dong Q, Zhu X, Dai C, Zhang X, Gao X, Wei
J, Sheng Y, Zheng Y, Yu J, Xie L, et al: Osteopontin promotes
epithelial-mesenchymal transition of hepatocellular carcinoma
through regulating vimentin. Oncotarget. 7:12997–13012. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Harms KL and Chen X: Histone deacetylase 2
modulates p53 transcriptional activities through regulation of
p53-DNA binding activity. Cancer Res. 67:3145–3152. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Lei WW, Zhang KH, Pan XC, Wang DM, Hu Y,
Yang YN and Song JG: Histone deacetylase 1 and 2 differentially
regulate apoptosis by opposing effects on extracellular
signal-regulated kinase 1/2. Cell Death Dis. 1:e442010. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
De Ruijter AJ, Van Gennip AH, Caron HN,
Stephan K and Van Kuilenburg AB: Histone deacetylases (HDACs):
Characterization of the classical HDAC family. Biochem J.
370:737–749. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Yadav A, Kumar B, Datta J, Teknos TN and
Kumar P: IL-6 promotes head and neck tumor metastasis by inducing
epithelial-mesenchymal transition via the JAK-STAT3-SNAIL signaling
pathway. Mol Cancer Res. 9:1658–1667. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Du R, Sun W, Xia L, Zhao A, Yu Y, Zhao L,
Wang H, Huang C and Sun S: Hypoxia-induced down-regulation of
microRNA-34a promotes EMT by targeting the Notch signaling pathway
in tubular epithelial cells. PLoS One. 7:e307712012. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Sarria AJ, Nordeen SK and Evans RM:
Regulated expression of vimentin cDNA in cells in the presence and
absence of a preexisting vimentin filament network. J Cell Biol.
111:553–565. 1990. View Article : Google Scholar : PubMed/NCBI
|