|
1
|
Bray F, Ferlay J, Soerjomataram I, Siegel
RL, Torre LA and Jemal A: Global cancer statistics 2018: GLOBOCAN
estimates of incidence and mortality worldwide for 36 cancers in
185 countries. CA Cancer J Clin. 68:394–424. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Harbeck N and Gnant M: Breast cancer.
Lancet. 389:1134–1150. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Redig AJ and McAllister SS: Breast cancer
as a systemic disease: A view of metastasis. J Intern Med.
274:113–126. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Stratton MR, Campbell PJ and Futreal PA:
The cancer genome. Nature. 458:719–724. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Balmain A, Gray J and Ponder B: The
genetics and genomics of cancer. Nat Genet. 33 (Suppl):S238–S244.
2003. View
Article : Google Scholar
|
|
6
|
Nguyen DX and Massague J: Genetic
determinants of cancer metastasis. Nat Rev Genet. 8:341–352. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Irish JM, Kotecha N and Nolan GP: Mapping
normal and cancer cell signalling networks: Towards single-cell
proteomics. Nat Rev Cancer. 6:146–155. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Papaleo E, Gromova I and Gromov P: Gaining
insights into cancer biology through exploration of the cancer
secretome using proteomic and bioinformatic tools. Expert Rev
Proteomics. 14:1021–1035. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Aftab A, Shahzad S, Hussain HMJ, Khan R,
Irum S and Tabassum S: CDKN2A/P16INK4A variants association with
breast cancer and their in-silico analysis. Breast Cancer.
26:11–28. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Klahan S, Wong HS, Tu SH, Chou WH, Zhang
YF, Ho TF, Liu CY, Yih SY, Lu HF, Chen SC, et al: Identification of
genes and pathways related to lymphovascular invasion in breast
cancer patients: A bioinformatics analysis of gene expression
profiles. Tumour Biol. 39:10104283177055732017. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Tang J, Kong D, Cui Q, Wang K, Zhang D,
Gong Y and Wu G: Prognostic genes of breast cancer identified by
gene co-expression network analysis. Front Oncol. 8:3742018.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Cheng D, He H and Liang B: A
three-microRNA signature predicts clinical outcome in breast cancer
patients. Eur Rev Med Pharmacol Sci. 22:6386–6395. 2018.PubMed/NCBI
|
|
13
|
Dedeurwaerder S, Desmedt C, Calonne E,
Singhal SK, Haibe-Kains B, Defrance M, Michiels S, Volkmar M,
Deplus R, Luciani J, et al: DNA methylation profiling reveals a
predominant immune component in breast cancers. EMBO Mol Med.
3:726–741. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Lv C, Wu X, Wang X, Su J, Zeng H, Zhao J,
Lin S, Liu R, Li H, Li X and Zhang W: The gene expression profiles
in response to 102 traditional Chinese medicine (TCM) components: A
general template for research on TCMs. Sci Rep. 7:3522017.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Mounir M, Lucchetta M, Silva TC, Olsen C,
Bontempi G, Chen X, Noushmehr H, Colaprico A and Papaleo E: New
functionalities in the TCGAbiolinks package for the study and
integration of cancer data from GDC and GTEx. PLoS Comput Biol.
15:e1006701. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW,
Shi W and Smyth GK: limma powers differential expression analyses
for RNA-sequencing and microarray studies. Nucleic Acids Res.
43:e472015. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Wen Q, Yang Y, Chen XH, Pan XD, Han Q,
Wang D, Dang Y, Li XH, Yan J and Zhou JH: Competing endogenous RNA
screening based on long noncoding RNA-messenger RNA co-expression
profile in Hepatitis B virus-associated hepatocarcinogenesis. J
Tradit Chin Med. 37:510–521. 2017. View Article : Google Scholar
|
|
18
|
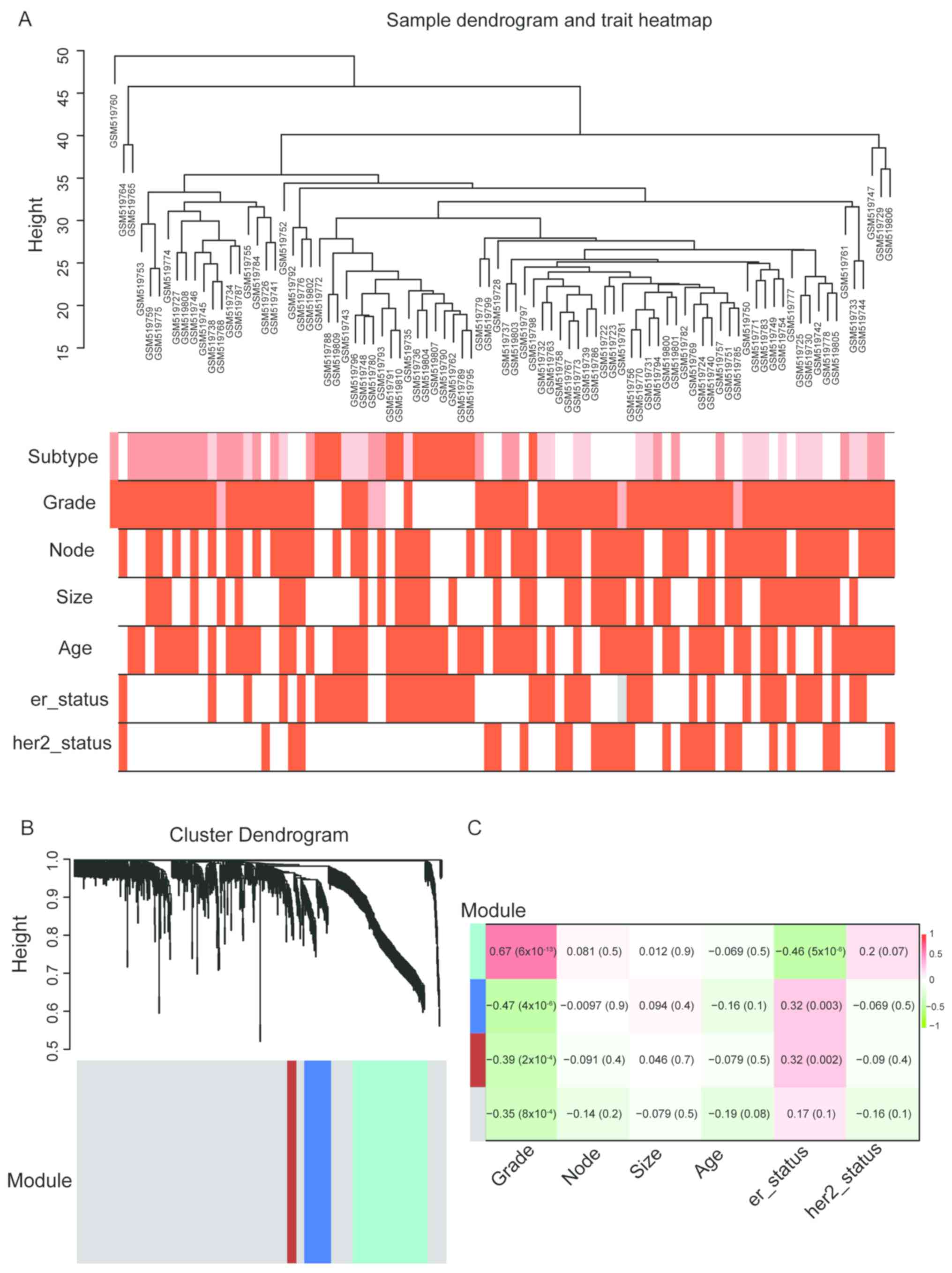
Langfelder P and Horvath S: WGCNA: An R
package for weighted correlation network analysis. BMC
Bioinformatics. 9:5592008. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Yuan L, Zeng G, Chen L, Wang G and Wang X,
Cao X, Lu M, Liu X, Qian G, Xiao Y and Wang X: Identification of
key genes and pathways in human clear cell renal cell carcinoma
(ccRCC) by co-expression analysis. Int J Biol Sci. 14:266–279.
2018. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Wang T, Wu B, Zhang X, Zhang M, Zhang S,
Huang W, Liu T, Yu W, Li J and Yu X: Identification of gene
coexpression modules, hub genes, and pathways related to spinal
cord injury using integrated bioinformatics methods. J Cell
Biochem. Jan 17–2019.(Epub ahead of print). doi:
10.1002/jcb.27908.
|
|
21
|
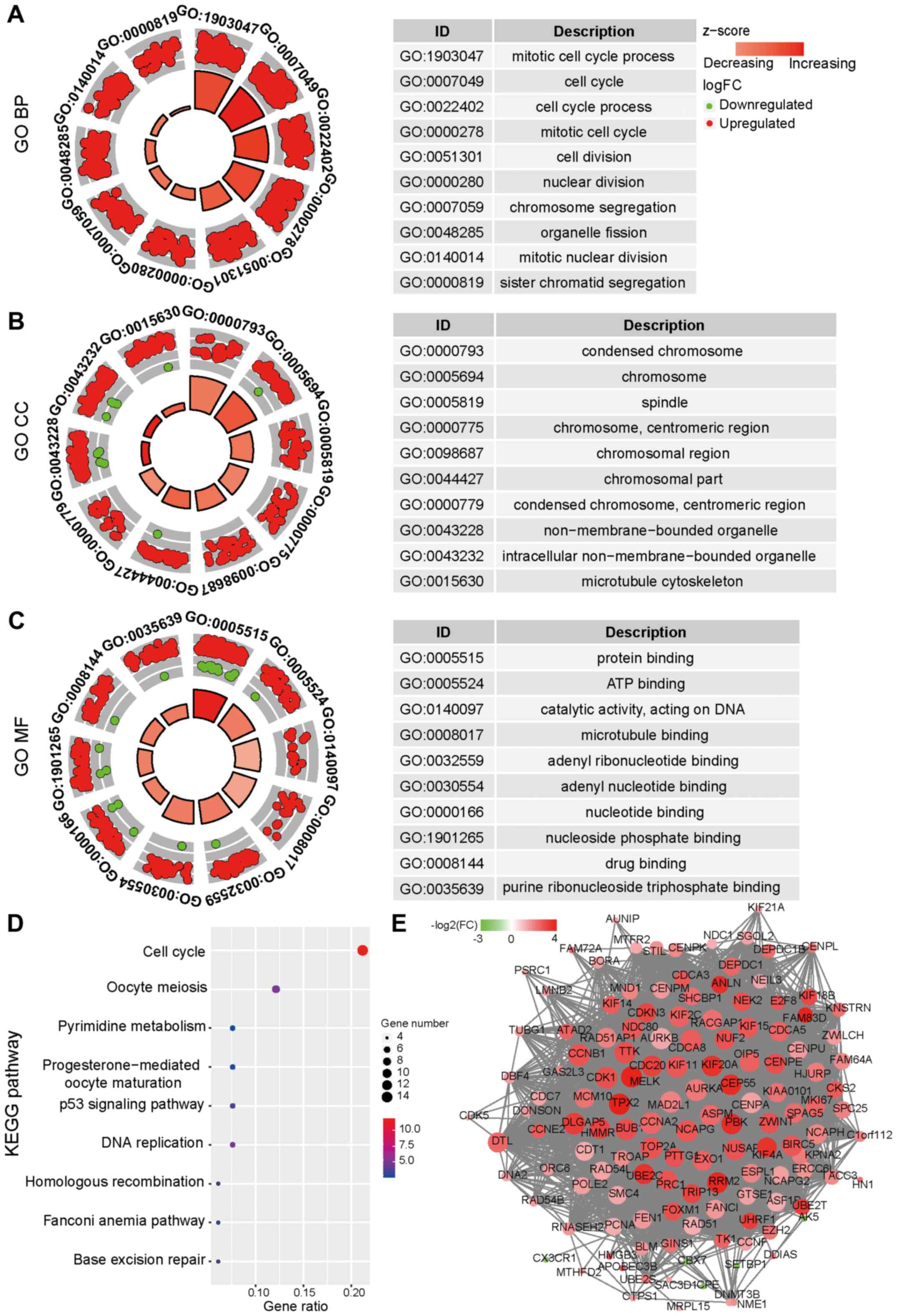
Yu G, Wang LG, Han Y and He QY:
ClusterProfiler: An R package for comparing biological themes among
gene clusters. OMICS. 16:284–287. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Walter W, Sánchez-Cabo F and Ricote M:
GOplot: An R package for visually combining expression data with
functional analysis. Bioinformatics. 31:2912–2914. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Mo XG, Liu W, Yang Y, Imani S, Lu S, Dan
G, Nie X, Yan J, Zhan R, Li X, et al: NCF2, MYO1F, S1PR4, and FCN1
as potential noninvasive diagnostic biomarkers in patients with
obstructive coronary artery: A weighted gene co-expression network
analysis. J Cell Biochem. Jan 17–2019.(Epub ahead of print).
View Article : Google Scholar
|
|
24
|
Yang Y, Lu Q, Shao X, Mo B, Nie X, Liu W,
Chen X, Tang Y, Deng Y and Yan J: Development of a three-gene
prognostic signature for hepatitis B virus associated
hepatocellular carcinoma based on integrated transcriptomic
analysis. J Cancer. 9:1989–2002. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Wang N, Guo H, Dong Z, Chen Q, Zhang X,
Shen W, Bao Y and Wang X: Establishment and validation of a
7-microRNA prognostic signature for non-small cell lung cancer.
Cancer Manag Res. 10:3463–3471. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Colwill K; Renewable Protein Binder
Working Group, ; Gräslund S: A roadmap to generate renewable
protein binders to the human proteome. Nat Methods. 8:551–558.
2011. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Xu H, Zhang Y, Qi L, Ding L, Jiang H and
Yu H: NFIX Circular RNA promotes glioma progression by regulating
miR-34a-5p via notch signaling pathway. Front Mol Neurosci.
11:2252018. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Subramanian A, Tamayo P, Mootha VK,
Mukherjee S, Ebert BL, Gillette MA, Paulovich A, Pomeroy SL, Golub
TR, Lander ES and Mesirov JP: Gene set enrichment analysis: A
knowledge-based approach for interpreting genome-wide expression
profiles. Proc Natl Acad Sci USA. 102:15545–15550. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Hu Z, Qin J, Zhang H, Wang D, Hua Y, Ding
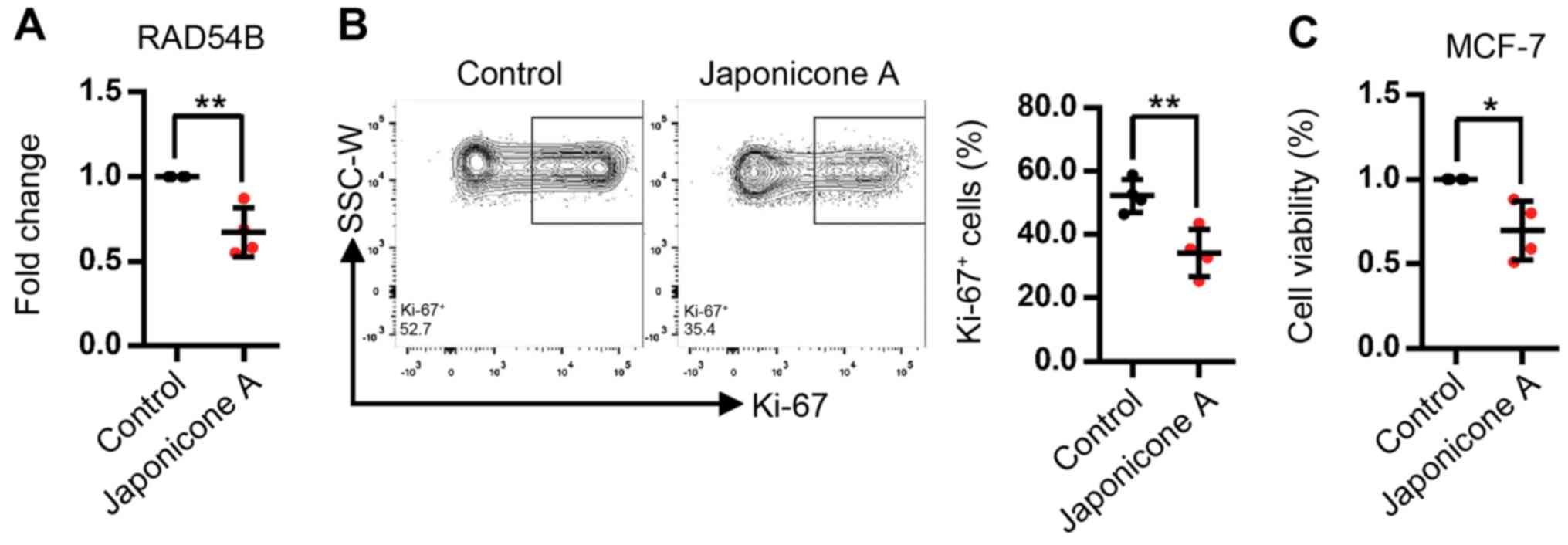
J, Shan L, Jin H, Zhang J and Zhang W: Japonicone A antagonizes the
activity of TNF-α by directly targeting this cytokine and
selectively disrupting its interaction with TNF receptor-1. Biochem
Pharmacol. 84:1482–1491. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Qin JJ, Jin HZ, Fu JJ, Hu XJ, Wang Y, Yan
SK and Zhang WD: Japonicones A-D, bioactive dimeric sesquiterpenes
from Inula japonica Thunb. Bioorg Med Chem Lett. 19:710–713.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Li X, Yang X, Liu Y, Gong N, Yao W, Chen
P, Qin J, Jin H, Li J, Chu R, et al: Japonicone A suppresses growth
of Burkitt lymphoma cells through its effect on NF-κB. Clin Cancer
Res. 19:2917–2928. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Du Y, Gong J, Tian X, Yan X, Guo T, Huang
M, Zhang B, Hu X, Liu H, Wang Y, et al: Japonicone A inhibits the
growth of non-small cell lung cancer cells via
mitochondria-mediated pathways. Tumour Biol. 36:7473–7482. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Patrick J: survivalROC: Time-dependent ROC
curve estimation from censored survival data. https://cran.r-project.org/web/packages/survivalROC/index.htmlMay.
2019
|
|
35
|
Vickers AJ and Elkin EB: Decision curve
analysis: A novel method for evaluating prediction models. Med
Decis Making. 26:565–574. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Marshall B: rmda: Risk model decision
analysis. https://cran.r-project.org/web/packages/rmda/index.htmlMarch
20–2018
|
|
37
|
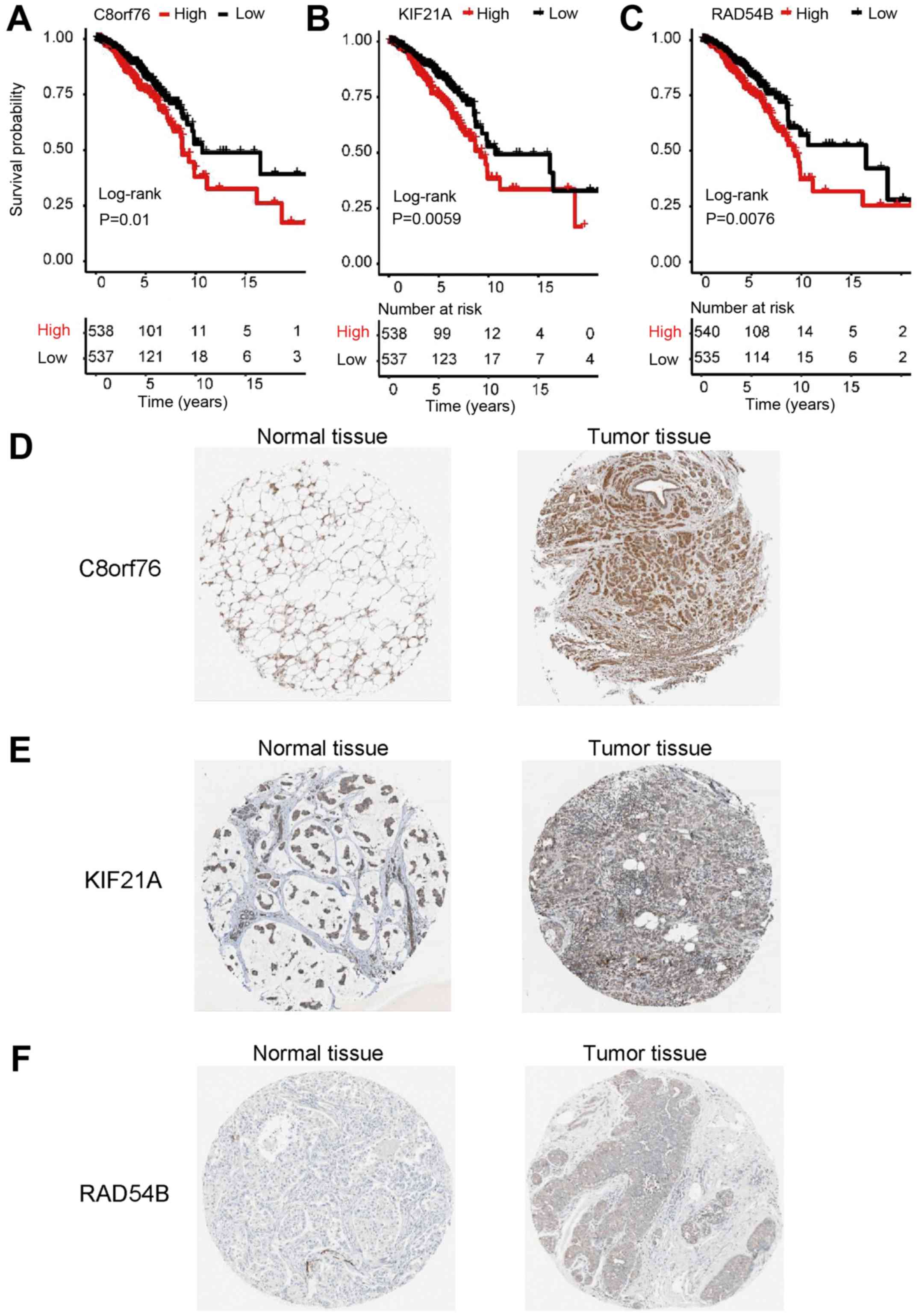
Uhlen M, Zhang C, Lee S, Sjöstedt E,
Fagerberg L, Bidkhori G, Benfeitas R, Arif M, Liu Z, Edfors F, et
al: A pathology atlas of the human cancer transcriptome. Science.
357(pii): eaan25072017. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Pontén F, Jirström K and Uhlen M: The
human protein atlas-a tool for pathology. J Pathol. 216:387–393.
2008. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Zhou J, Li G, Zheng Y, Shen HM, Hu X, Ming
QL, Huang C, Li P and Gao N: A novel autophagy/mitophagy inhibitor
liensinine sensitizes breast cancer cells to chemotherapy through
DNM1L-mediated mitochondrial fission. Autophagy. 11:1259–1279.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Nagini S: Breast cancer: Current molecular
therapeutic targets and new players. Anticancer Agents Med Chem.
17:152–163. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Hiramoto T, Nakanishi T, Sumiyoshi T,
Fukuda T, Matsuura S, Tauchi H, Komatsu K, Shibasaki Y, Inui H,
Watatani M, et al: Mutations of a novel human RAD54 homologue,
RAD54B, in primary cancer. Oncogene. 18:3422–3426. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Miyagawa K, Tsuruga T, Kinomura A, Usui K,
Katsura M, Tashiro S, Mishima H and Tanaka K: A role for RAD54B in
homologous recombination in human cells. EMBO J. 21:175–180. 2002.
View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Wesoly J, Agarwal S, Sigurdsson S, Bussen
W, Van Komen S, Qin J, van Steeg H, van Benthem J, Wassenaar E,
Baarends WM, et al: Differential contributions of mammalian Rad54
paralogs to recombination, DNA damage repair, and meiosis. Mol Cell
Biol. 26:976–989. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Yasuhara T, Suzuki T, Katsura M and
Miyagawa K: Rad54B serves as a scaffold in the DNA damage response
that limits checkpoint strength. Nat Commun. 5:54262014. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Wang R, Li Y, Chen Y and Wang L, Wu Q, Guo
Y, Li Y, Liu J and Wang L: Inhibition of RAD54B suppresses
proliferation and promotes apoptosis in hepatoma cells. Oncol Rep.
40:1233–1242. 2018.PubMed/NCBI
|
|
46
|
Mathur S and Hoskins C: Drug development:
Lessons from nature. Biomed Rep. 6:612–614. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Clardy J and Walsh C: Lessons from natural
molecules. Nature. 432:829–837. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Ertl P and Schuffenhauer A:
Cheminformatics analysis of natural products: Lessons from nature
inspiring the design of new drugs. Prog Drug Res. 66:217, 219–235.
2008.
|
|
49
|
Qin JJ, Jin HZ, Zhu JX, Fu JJ, Hu XJ, Liu
XH, Zhu Y, Yan SK and Zhang WD: Japonicones E-L, dimeric
sesquiterpene lactones from Inula japonica Thunb. Planta
Med. 76:278–283. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
West AP, Khoury-Hanold W, Staron M, Tal
MC, Pineda CM, Lang SM, Bestwick M, Duguay BA, Raimundo N, MacDuff
DA, et al: Mitochondrial DNA stress primes the antiviral innate
immune response. Nature. 520:553–557. 2015. View Article : Google Scholar : PubMed/NCBI
|