|
1
|
Kyle R and Rajkumar SV: Multiple myeloma.
Blood. 111:2962–2972. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Siegel RL, Miller KD and Jemal A: Cancer
statistics, 2018. CA. Cancer J Clin. 68:7–30. 2018. View Article : Google Scholar
|
|
3
|
Rastgoo N, Abdi J, Hou J and Chang H: Role
of epigenetics-microRNA axis in drug resistance of multiple
myeloma. J Hematol Oncol. 10:1212017. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Dupéré-Richer D and Licht JD: Epigenetic
regulatory mutations and epigenetic therapy for multiple myeloma.
Curr Opin Hematol. 24:336–344. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Dimopoulos K, Gimsing P and Grønbæk K: The
role of epigenetics in the biology of multiple myeloma. Blood
Cancer J. 4:e2072014. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
He L and Hannon GJ: Erratum: MicroRNAs:
Small RNAs with a big role in gene regulation. Nat Rev Genet.
5:522–531. 2004. View
Article : Google Scholar : PubMed/NCBI
|
|
7
|
Kloosterman WP and Plasterk RH: The
diverse functions of microRNAs in animal development and disease.
Dev Cell. 11:441–450. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Garzon R, Fabbri M, Cimmino A, Calin GA
and Croce CM: MicroRNA expression and function in cancer. Trends
Mol Med. 12:580–587. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Kuehbacher A, Urbich C and Dimmeler S:
Targeting microRNA expression to regulate angiogenesis. Trends
Pharmacol Sci. 29:12–15. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Pichiorri F, Suh SS, Ladetto M, Kuehl M,
Palumbo T, Drandi D, Taccioli C, Zanesi N, Alder H, et al:
MicroRNAs regulate critical genes associated with multiple myeloma
pathogenesis. Proc Natl Acad Sci USA. 105:12885–12890. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Abdi J, Jian H and Chang H: Role of
micro-RNAs in drug resistance of multiple myeloma. Oncotarget.
7:60723–60735. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
de Yébenes VG, Bartolomé-Izquierdo N and
Ramiro AR: Regulation of B cell development and function by
microRNAs. Immunol Rev. 253:25–39. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
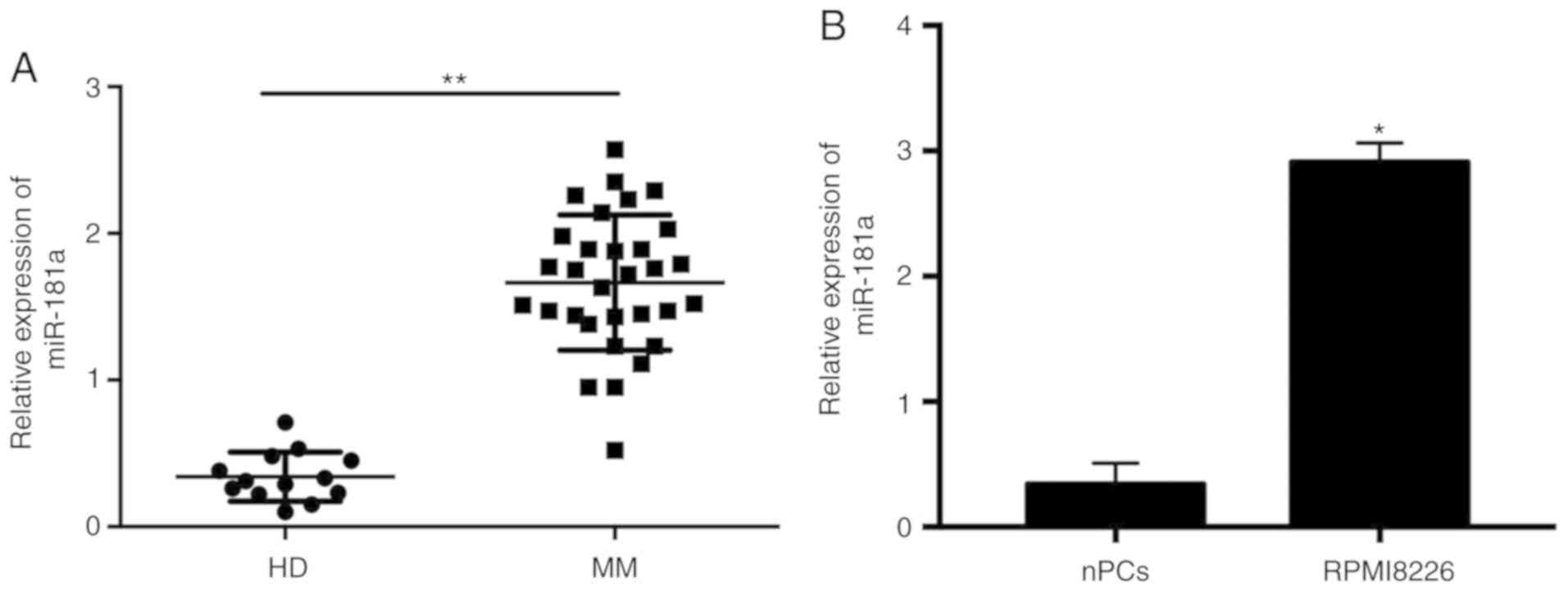
Peng J, Thakur A, Zhang S, Dong Y, Wang X,
Yuan R, Zhang K and Guo X: Expressions of miR-181a and miR-20a in
RPMI8226 cell line and their potential as biomarkers for multiple
myeloma. Tumour Biol. 36:8545–8552. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
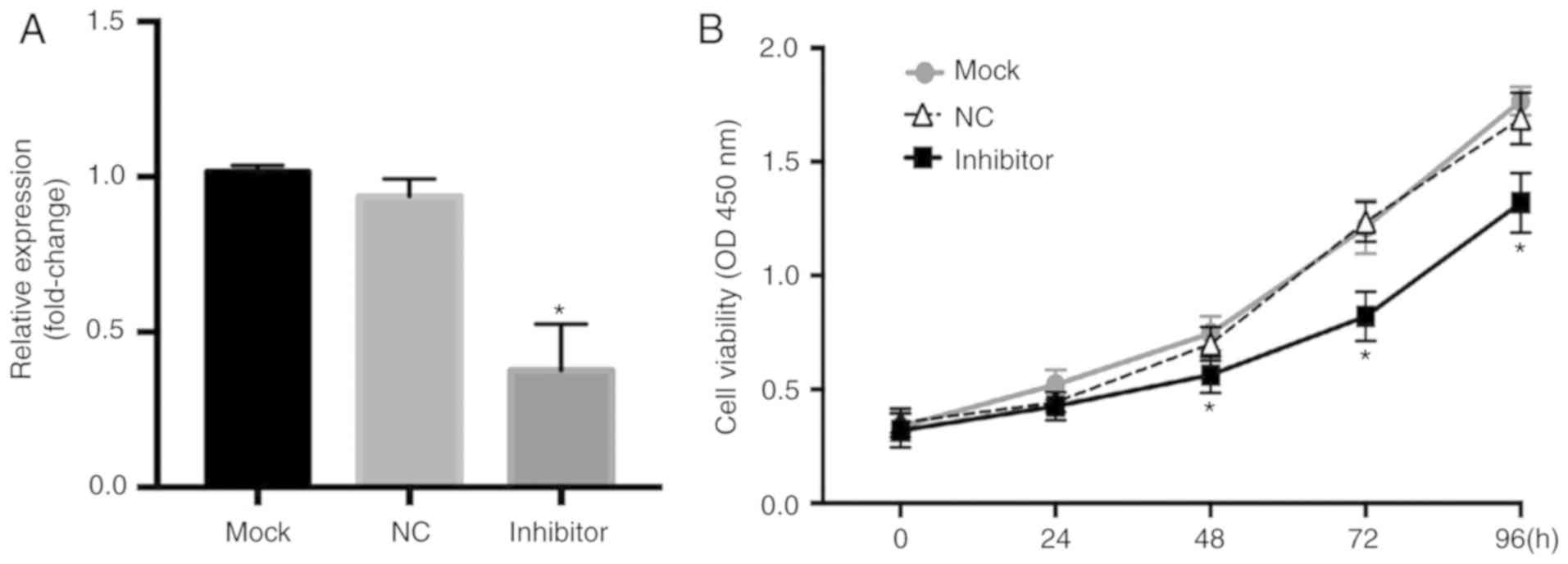
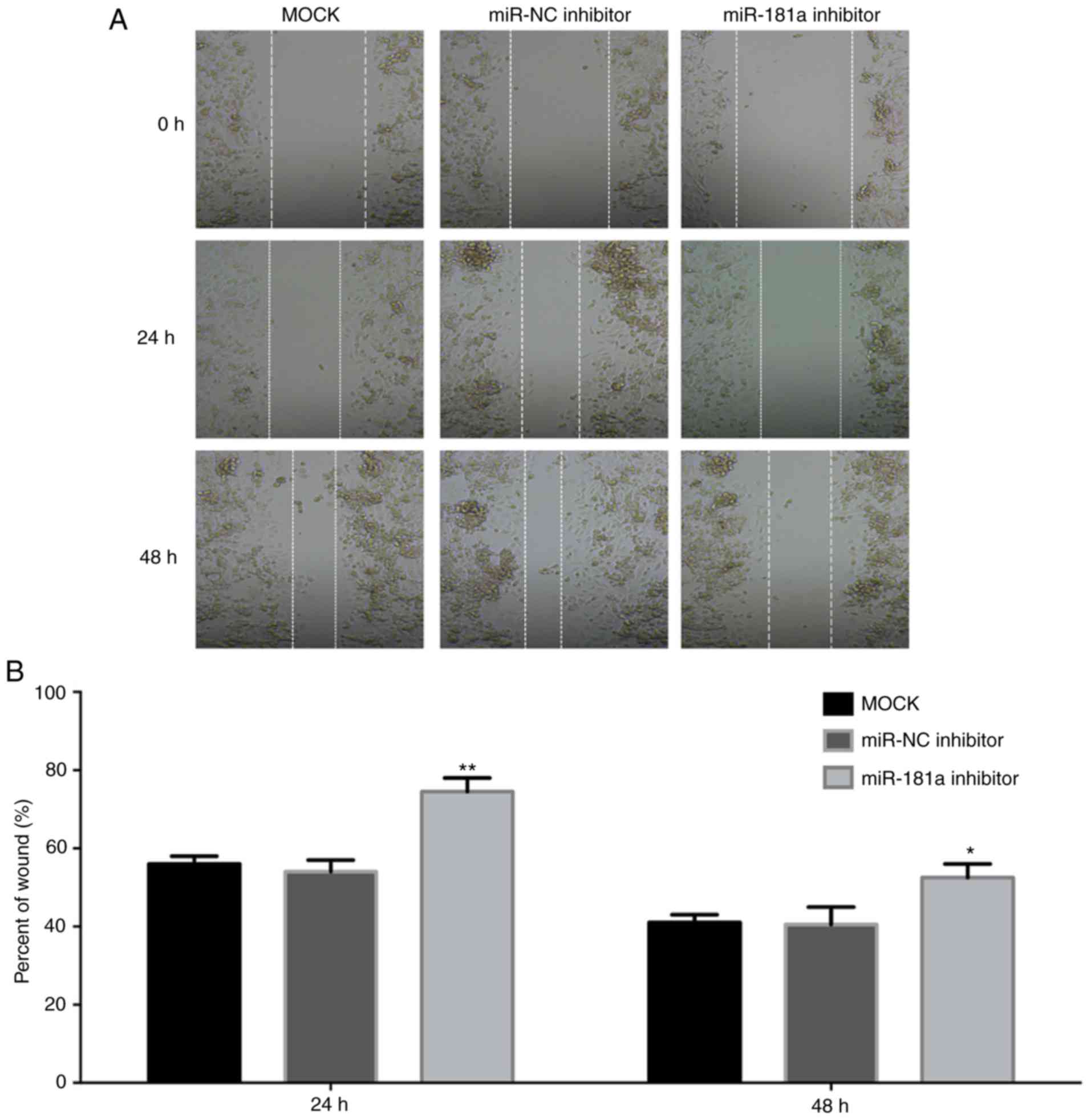
Liu N, Yang J, Yuan R, Peng J, Liu L and
Guo X: Effects of miR-181a on the biological function of multiple
myeloma. Oncol Rep. 42:291–300. 2019.PubMed/NCBI
|
|
15
|
Cao Y, Zhao D, Li P, Wang L, Qiao B, Qin
X, Li L and Wang Y: MicroRNA-181a-5p impedes IL-17-induced
non-small cell lung cancer proliferation and migration through
targeting VCAM-1. Cell Physiol Biochem. 42:346–356. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Greipp PR, San Miguel J, Durie BG, Crowley
JJ, Barlogie B, Bladé J, Boccadoro M, Child JA, Avet-Loiseau H,
Kyle RA, et al: International staging system for multiple myeloma.
J Clin Oncol. 23:3412–3420. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Di Leva G, Garofalo M and Croce CM:
MicroRNAs in cancer. Annu Rev Pathol. 9:287–314. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Bi C and Chng WJ: MicroRNA: Important
player in the pathobiology of multiple myeloma. Biomed Res Int.
2014:5215862014. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Huang S, Wu S, Ding J, Lin J, Wei L, Gu J
and He X: MicroRNA-181a modulates gene expression of zinc finger
family members by directly targeting their coding regions. Nucleic
Acids Res. 38:7211–7218. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Li L, Xu QH, Dong YH, Li GX, Yang L, Wang
LW and Li HY: MiR-181a upregulation is associated with
epithelial-to-mesenchymal transition (EMT) and multidrug resistance
(MDR) of ovarian cancer cells. Eur Rev Med Pharmacol Sci.
20:2004–2010. 2016.PubMed/NCBI
|
|
22
|
Ghorbani S, Talebi F, Chan WF, Masoumi F,
Vojgani M, Power C and Noorbakhsh F: MicroRNA-181 variants regulate
T cell phenotype in the context of autoimmune neuroinflammation.
Front. Immunol. 8:7582017.
|
|
23
|
Verduci L, Azzalin G, Gioiosa S, Carissimi
C, Laudadio I, Fulci V and Macino G: microRNA-181a enhances cell
proliferation in acute lymphoblastic leukemia by targeting EGR1.
Leuk Res. 39:479–485. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Yyusnita, Norsiah, Zakiah I, Chang KM,
Purushotaman VS, Zubaidah Z and Jamal R: MicroRNA (miRNA)
expression profiling of peripheral blood samples in multiple
myeloma patients using microarray. Malaysian J Pathol. 34:133–143.
2012.
|
|
25
|
Li Y, Li D, Yan Z, Qi K, Chen L, Zjang Z,
Fan G, Li H, Xu K and Li Z: Potential relationship and clinical
significance of miRNAs and Th17 cytokines in patients with multiple
myeloma. Leuk Res. 38:1130–1135. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Lionetti M, Musto P, Di Martino MT, Fabris
S, Agnelli L, Todoerti K, Tuana G, Mosca L, Gallo Cantafio ME,
Grieco V, et al: Biological and clinical relevance of miRNA
expression signatures in primary plasma cell leukemia. Clin Cancer
Res. 19:3130–3142. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Baldin V, Lukas J, Marcote MJ, Pagano M
and Draetta G: Cyclin D1 is a nuclear protein required for cell
cycle progression in G1. Genes Dev. 7:812–821. 1993. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Casimiro MC, Velasco-Velázquez M,
Aguirre-Alvarado C and Pestell RG: Overview of cyclins D1 function
in cancer and the CDK inhibitor landscape: Past and present. Expert
Opin Investig Drugs. 23:295–304. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Lesage D, Troussard X and Sola B: The
enigmatic role of cyclin D1 in multiple myeloma. Int J Cancer.
115:171–176. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Hafner M, Landthaler M, Burger L, Khorshid
M, Hausser J, Berninger P, Rothballer A, Ascano M Jr, Jungkamp AC,
Munschauer M, et al: Transcriptome-wide identification of
RNA-binding protein and microRNA target sites by PAR-CLIP. Cell.
141:129–141. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Balakrishnan I, Yang X, Brown J,
Ramakrishnan A, Torok-Storb B, Kabos P, Hesselberth JR and Pillai
MM: Genome-wide analysis of miRNA-mRNA interactions in marrow
stromal cells. Stem Cells. 32:662–673. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Farazi TA, Ten Hoeve JJ, Brown M,
Mihailovic A, Horlings HM, van de Vijver MJ, Tuschl T and Wessels
LF: Identification of distinct miRNA target regulation between
breast cancer molecular subtypes using AGO2-PAR-CLIP and patient
datasets. Genome Biol. 15:R92014. View Article : Google Scholar : PubMed/NCBI
|