Introduction
Laryngeal squamous cell carcinoma (LSCC) was the
second most predominant type of head and neck squamous cell cancer
(HNSCC), and the sixth most common type of carcinoma worldwide in
2011 (1). Laryngeal cancer accounts
for 2–5% of all malignancies with ~200,000 deaths annually
(2); however, it has a major impact
on phonation, swallowing, respiration and quality of life (3). Histologically, >95% of laryngeal
carcinoma cases are identified as LSCC (4). For clinical and staging purposes,
laryngeal cancer is currently clustered in supraglottic, glottic
and subglottic regions, among which supraglottic LSCC is the second
most common type of laryngeal cancer (5) and presents different biological
properties, such as being more likely to recur and metastasize
(6). Notably, supraglottic LSCC is
more prevalent in some developing countries where alcohol and
smoking are common risk factors (7).
The occurrence and development of LSCC are
synergistically caused by extrinsic and inheritable
multi-etiological factors and their interactions. In addition to
environmental carcinogens, such as exposure to tobacco, alcohol and
radioactive rays, and infectious agents, including viral
infections, molecular and chromosomal alterations are the most
common inheritable etiological factors (8). Copy number variations (CNVs) are the
prevalent structural variations in the genome and have been
demonstrated to contribute to the genetic diversity of tumor
pathogenesis, being increasingly accepted as a major source of
inheritable variations (9). CNVs
have been demonstrated to result in loss-of-function variants of
tumor suppressor genes and/or gain-of-function variants of
oncogenes (10). Notably, high
levels of DNA CNVs in tumors are frequently limited to certain
chromosomal segments with renowned oncogenes that are overexpressed
or triggered (11).
Comparative genomic hybridization (CGH) has emerged
as a high-throughput genomic technology to facilitate the
aggregation of high-resolution data of cancer-associated genomic
imbalances. In the present study, high-resolution genomic
microarray CGH was applied to detect regions of gain or
amplification and losses in supraglottic LSCC. By integration of
CNVs with bioinformatic analysis, the possible candidate target
genes with strong clinical significance were identified, providing
novel insights into the molecular pathological mechanisms
underlying supraglottic LSCC. Additionally, associations between
genetic alternations and lymph node metastasis, tumor stage and
distant metastasis were statistically analyzed, and differentially
expressed genes (DEGs) between samples from patients with HNSCC and
normal controls were identified using microarray data from the Gene
Expression Omnibus (GEO) database to determine the optimal gene
combination with diagnostic value for HNSCC. The findings of the
present study may elucidate the mechanism underlying the
development of supraglottic LSCC and uncover potential diagnostic
and therapeutic biomarkers for supraglottic LSCC.
Materials and methods
Collection of tumor tissues
Tissues from 38 patients with primary supraglottic
LSCC were collected during excision surgery between November 2011
and October 2016 at the Department of Otorhinolaryngology, Head and
Neck Surgery, The First Hospital of Jilin University (Changchun,
China). All patients had histopathologically confirmed LSCC and TNM
stage was determined according to American Joint Committee on
Cancer staging system (12). The
present study included 31 male and 7 female patients aged between
52 and 75 years (median age, 62 years). The clinicopathological
features and risk factors of the patients with supraglottic LSCC
are summarized in Table I. All
patients had negative histories of exposure to either chemotherapy
or radiotherapy before surgery, and no other cancers were
detected.
 | Table I.Clinicopathological characteristics
of 38 patients with supraglottic laryngeal squamous cell carcinoma
for array-based comparative genomic hybridization analysis. |
Table I.
Clinicopathological characteristics
of 38 patients with supraglottic laryngeal squamous cell carcinoma
for array-based comparative genomic hybridization analysis.
| Features | Number | Percentage |
|---|
| Mean age at
diagnosis, | 62.87 |
|
| years (range) | (52–75) |
|
|
≤60 | 15 | 39.47 |
|
>60 | 23 | 60.53 |
| Sex |
|
|
|
Male | 31 | 81.58 |
|
Female | 7 | 18.42 |
| Histopathological
grading |
|
|
| Well
differentiated | 4 | 10.53 |
|
Moderately differentiated | 27 | 71.05 |
| Poorly
differentiated | 7 | 18.42 |
| AJCC TNM
classification |
|
|
| I | 2 | 5.26 |
| II | 11 | 28.95 |
|
III | 11 | 28.95 |
| IV | 14 | 36.84 |
| T
classification |
|
|
| T1 | 4 | 10.53 |
| T2 | 17 | 44.74 |
| T3 | 14 | 36.84 |
| T4 | 3 | 7.89 |
| N
classification |
|
|
| N0 | 20 | 52.63 |
| N1 | 6 | 15.79 |
| N2 | 10 | 26.32 |
| N3 | 2 | 5.26 |
| Metastasis |
|
|
| M0 | 38 | 100.00 |
| M1 | 0 | 0.00 |
| Smoking history,
years |
|
|
| 0 | 2 | 5.26 |
|
≤20 | 1 | 2.63 |
|
20–30 | 10 | 26.32 |
|
30–40 | 18 | 47.37 |
|
>40 | 7 | 18.42 |
| Alcoholic
history |
|
|
|
Alcoholic | 19 | 50.00 |
|
Non-alcoholic | 19 | 50.00 |
Written informed consent was provided by all
patients enrolled in the present study with the authorization of
the Ethics Committee of The First Hospital of Jilin University.
Tumor tissues and adjacent non-tumor tissues (>1 cm from tumor)
were surgically obtained at the Department of Otorhinolaryngology,
Head and Neck Surgery. The primary tumor tissues were obtained from
the center of the tumor lesion, confirmed by an independent
pathologist after surgical removal, immediately snap-frozen in
liquid nitrogen and stored at −80°C until further use. Samples were
first digested using proteinase K, followed by the
phenol-chloroform method for DNA isolation according to standard
protocols. Briefly, frozen tumor samples were grinded to small
pellets and treated with SE buffer (75 mM NaCl, 25 mM EDTA and 10N
NaOH pH 8.0), 20% SDS and proteinase K. After incubation overnight
at 50°C, DNA was extracted by phenol-chloroform extraction and
ethanol precipitation. The DNA was washed with 70% ethanol and air
dried. The DNA was dissolved in 50 µl TE buffer (10 mM Tris HCl pH
7.4 and 1 mM EDTA) and the DNA concentration was measured by
Nanodrop 2000 (Thermo Fisher Scientific, Inc.) and run on a 0.8%
agarose gel.
Array-based CGH (aCGH) assay
aCGH was conducted on a SurePrint 2×400 k oligomer
CGH chip (Agilent Technologies, Inc.) according to the
manufacturer's protocol. The reference DNA used was included in the
aforementioned labeling kit to identify CNVs (male reference DNA,
cat. no. 5190-3796; and female reference DNA, cat. no. 5190-3797;
Agilent Technologies, Inc.). The DNA from patients and the
reference DNA were labeled with different color fluorescent dyes
via PCR for 40 h at 48°C, namely with Cyanine 5 and Cyanine 3,
respectively, and then equal labeling products were mixed together
for competitive co-hybridization onto a chip by incubation in a
hybridization oven (Agilent Technologies, Inc.) for 40 h at 48°C.
The chip was washed using the washing buffer in the kit and scanned
with an MS200 laser scanner using the NimbleGen system (Roche
Applied Science).
Array data analysis
Images were analyzed using the Cytogenomics v4.0
software (Agilent Technologies, Inc.). The cut-off threshold of
positive CNVs was determined using a log2 ratio with
±0.15 in ≥10 neighboring probes coordinating gain or loss, due to
complicated DNA components from tumor tissues, such as tumor cell
diversity and mosaic variation. Amplifications were specified as
those with a smoothed log R ratio >0.5, while homozygous
deletions were specified as those with a smoothed log R ratio
<-0.5.
Affymetrix microarray data
The microarray gene expression profiling dataset
GSE6631 for HNSCC was downloaded from the GEO database (https://www.ncbi.nlm.nih.gov/sites/GDSbrowser?acc=GDS2520).
The HNSCC-associated dataset GSE6631 was from the GPL8300
Affymetrix Human Genome U95 Version 2 Array platform, and included
44 HNSCC tumors and 44 matching normal mucosa samples (13). GSE6631 series matrix .txt files and
GPL8300 platform .txt files were extracted for data processing.
Data processing and identification of
DEGs
The gene probe IDs in the matrix files were
converted to the gene symbols in the platform files using a Perl
script to generate a matrix file with the global standard gene
name. Each dataset was then standardized using the
normalizeBetweenArrays function of the limma R package v3.10.1
(http://www.bioconductor.org/packages/release/bioc/html/limma.html).
All gene expression data were transformed to a log2
scale. DEGs between HNSCC tumor tissues and control samples were
determined using the limma R package. P-values were adjusted using
the Benjamini-Hochberg false discovery rate method, and adjusted
P<0.05 and |log fold-change (FC)|>1 were used as the cut-off
criteria. Heatmaps and volcano maps were visualized using the
ggplot2 R package v3.0.0 (14).
Statistical analysis
Data are presented as the case number and all
experiments were performed once. Significant associations between
genomic aberrations and clinicopathological factors were determined
using the χ2 test in SPSS v20.0 (IBM Corp.). P<0.05
was considered to indicate a statistically significant
difference.
Results
Identification of recurrent CNVs in
supraglottic LSCC via aCGH
Genomic CNVs (gains, losses, amplifications and
homozygous deletions) were detected via aCGH in all 38 samples of
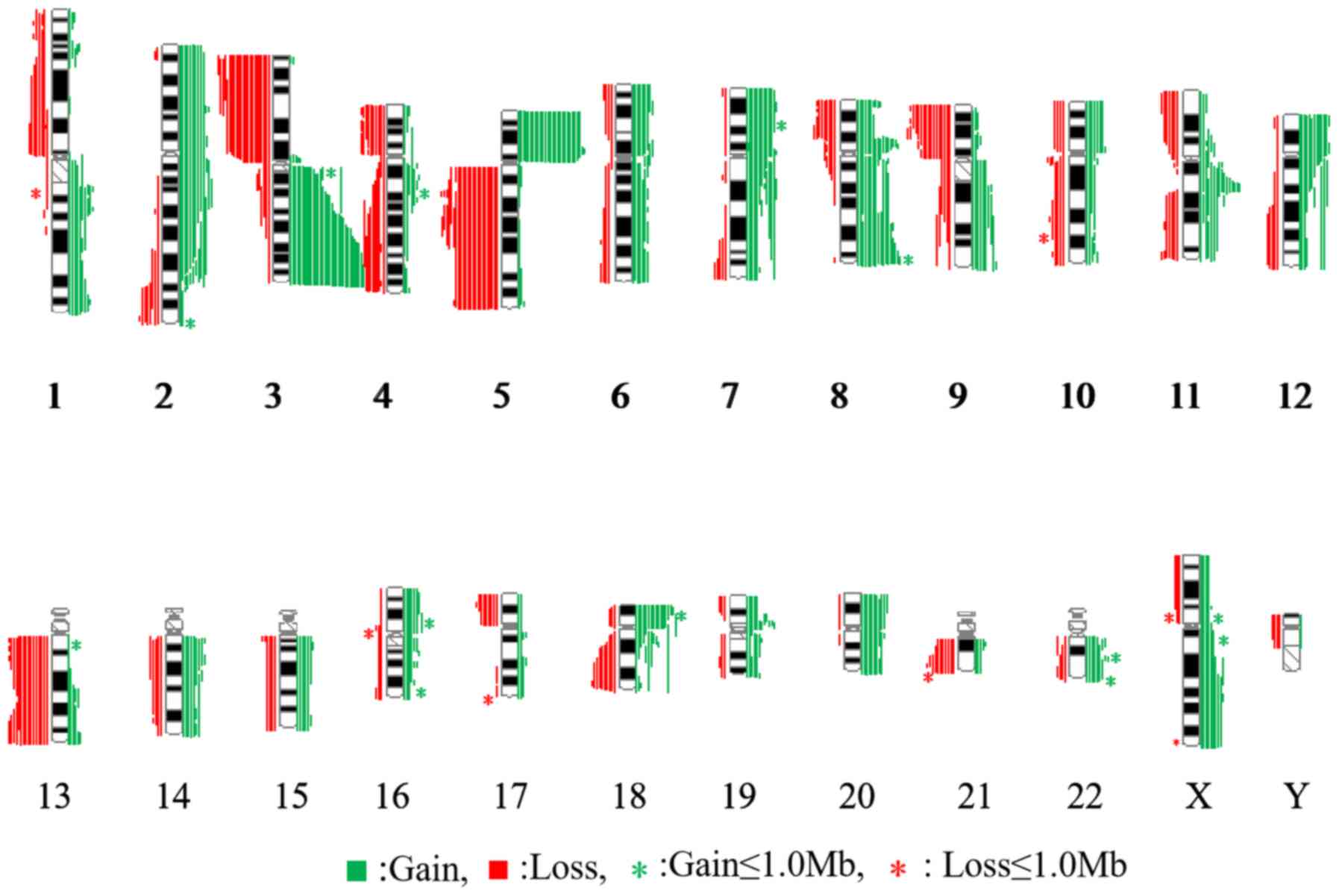
supraglottic LSCC. Genomic imbalance profiling is presented in
Fig. 1. Net gains (21 cases) were
more frequent compared with net losses (17 cases). As shown in
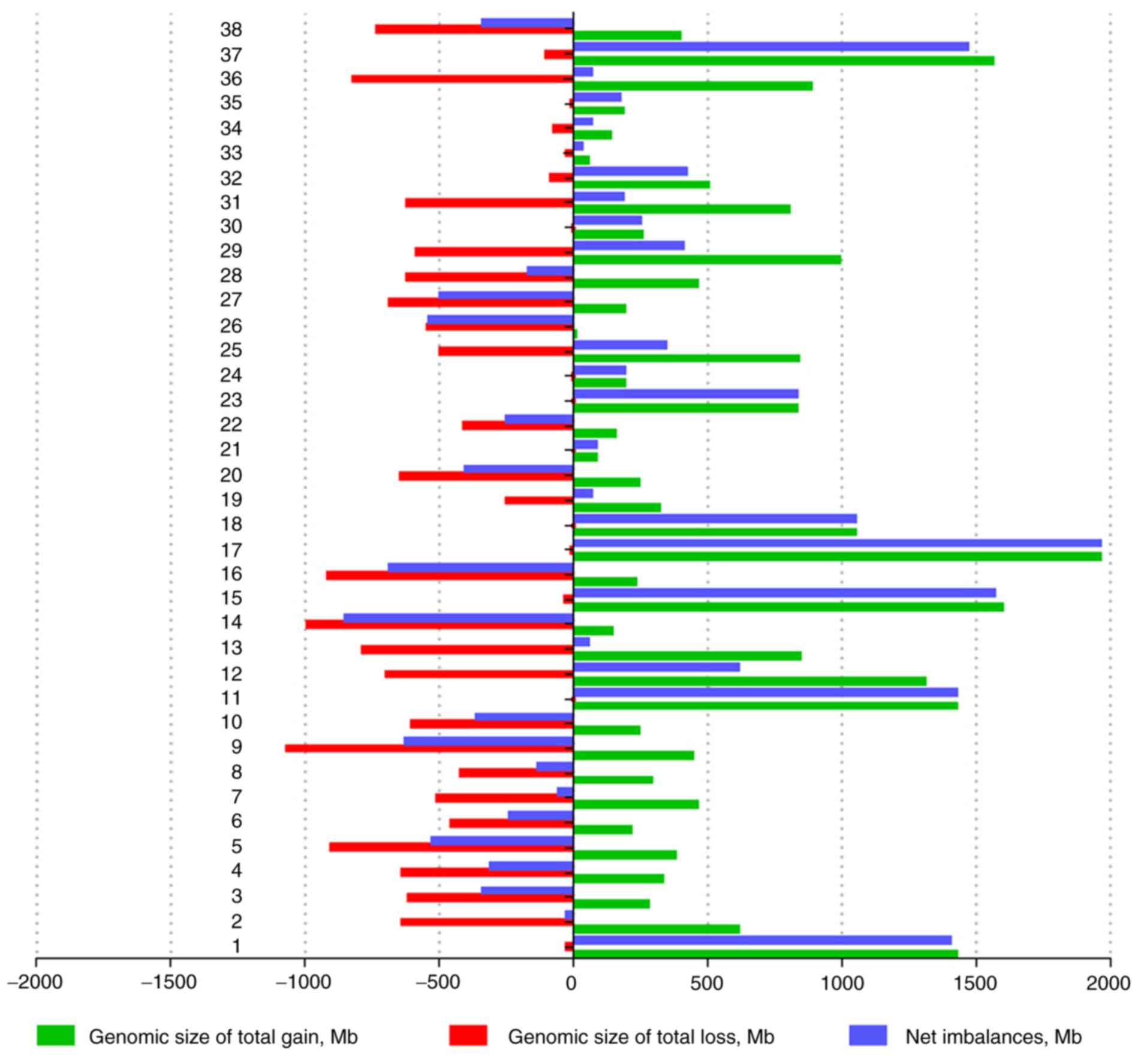
Fig. 2, the sizes of net genomic
imbalances per case ranged between a loss of 849.4 Mb (~29.9% of
the genome) and a gain of 1,958.6 Mb (~69.0% of genome). The
average number of gains per case was 15.7, ranging between 2 and
37, whereas the average number of losses per case was 9.2, ranging
between 0 and 24 (data not shown). The gain sizes ranged between
252 kb and 204 Mb, whereas the loss sizes ranged between 174 kb and
198 Mb. Overall, 41/946 (4.3%) of the total genomic imbalances were
<1 Mb, among which 31/41 (75.6%) were gains and 10/41 (24.4%)
were losses (data not shown). The most frequent genomic imbalances
were considered as those detected in ≥10/38 (26.3%) samples of
supraglottic LSCC and included 10 gains (2pter-q22.1, 3q26.1-qter,
5pter-p12, 7p22.3p14.1, 8p12p11.22, 8q24.13q24.3, 11q13.2q13.4,
12pter-p12.2, 18pter-p11.31 and 20p13p12.1), and 5 losses in
various chromosome regions (3pter-p21.32, 4q28.1-q35.2,
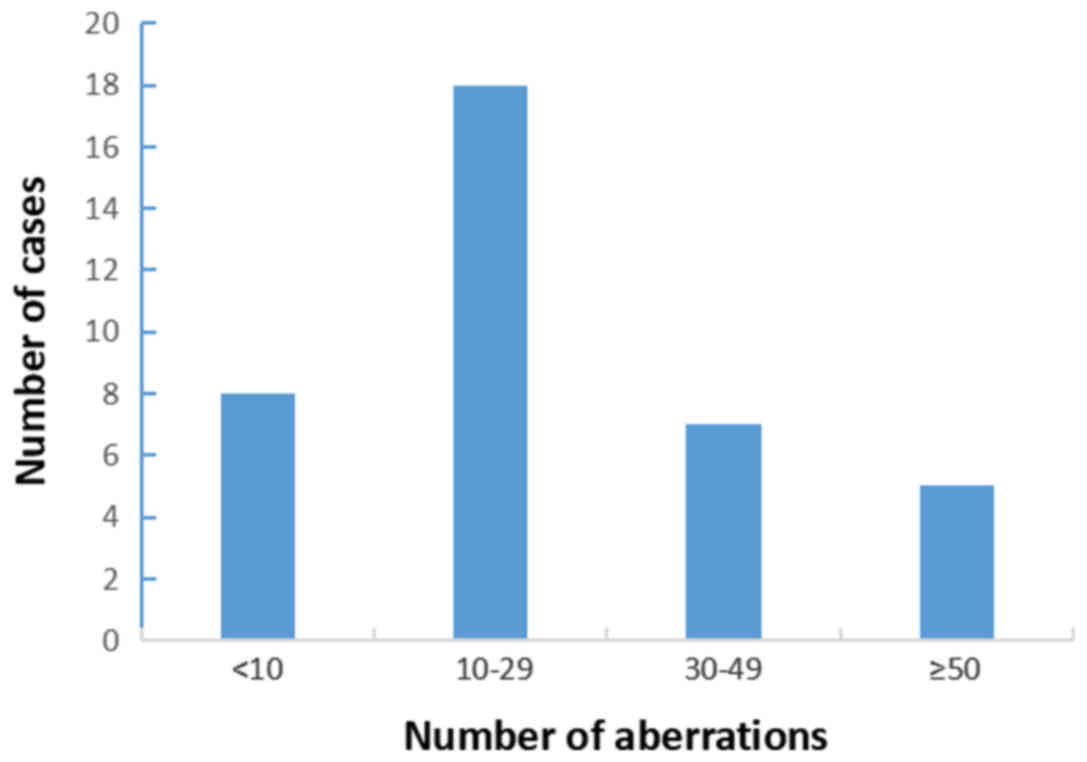
5q13.2-qter, 9pter-p21.3 and monosomy 13) (Table II). As presented in Fig. 3, nearly half of the cases had between
10 and 29 genetic alterations.
 | Table II.Frequently alternated loci and
interesting genes in supraglottic laryngeal squamous cell carcinoma
samples. |
Table II.
Frequently alternated loci and
interesting genes in supraglottic laryngeal squamous cell carcinoma
samples.
| Copy number
variation | Chromosome | Genomic
coordinates, bp | Frequency
(n=38) | Selected
interesting gene(s) |
|---|
| Gains | 2pter-q22.1 | 0-138532912 | 10 | EPCAM, MSH6, MSH2,
EHBP1, IL1RN, IL1B, GACAT3, BUB1, TET3, ODC1 |
|
| 3q26.1-qter |
162071193-197940241 | 31 | PIK3CA, LPP, CEP19,
CLDN1, SOX2, LIPH, BCL6 |
|
| 5pter-p12 | 0-46104594 | 26 | FGF10, LIFR, SDHA,
PRLR, TERT, ANKH, GDNF, IL7R, GHR |
|
| 7p22.3p14.1 | 0-42314328 | 13 | MAD1LA, PMS2,
CARD11, RALA, AHR, AQP1, MMD2 |
|
| 8p12-p11.22 |
33531006-38295039 | 17 | FGFR1, BRF2, DDHD2,
ERLIN2, ADRB3, STAR |
|
| 8q24.13-q24.3 |
124021960-142128937 | 19 | RNF139, MYC,
PRNCR1, NDUFB9, PVT1, SLA, LRRC6, NDRG1, KCNQ3 |
|
| 11q13.2q13.4 |
68442471-70576583 | 13 | CCND1, CTTN,
ORAOV1, MYEOV, FGF3, FGF4, FGF19, FADD, SHANK2 |
|
| 12pter-p12.2 | 0-20960518 | 14 | FGF6, CCND2, ATN1.
EPS8, ART4, EMG1, EGF23, GDF3, WNK1, GNB3 |
|
| 18pter-p11.31 | 0-3569480 | 20 | YES1, LPIN2,
SMCHD1, USP14, MYOM1 |
|
| 20p13p12.1 | 0-17519968 | 12 | SNAP25, PANK2,
MCM8, IDH3B, TGM6, MKKS, PDYN, PTPRA, HAO1, TASP1 |
| Losses | 3pter-p21.32 | 0-54006116 | 20 | BTD, MYL3, XPC,
RASSF1, DLEC1, BAP1, OFF1, GNAI2, MLH1, CTNNB1, TGFBR2 |
|
| 4q28.1-q35.2 |
135254483-191010484 | 10 | KLKB1, HHIP, EDNRA,
NR3C2, NEK1, LAT, FAT1, PALLD, TLR2 |
|
| 5q13.2-qter |
68974554-181354732 | 18 | FGFR4, RASA1, IRF1,
MSH3, HMMR, MCC, APC, NPM1, ARHG-AP26, TLX3, COX7C, TLX3, F12,
CCNH, APC, CD14, LOX, CHD1, CCNH, FER |
|
| 9pter-p21.3 | 0-21890680 | 16 | IFNA1, GLDC, DOCK8,
JAK2, MTAP, IL33, IFNA21 |
|
| Monosomy 13 | 0-96500000 | 13 | RB1, DLEU1, DELU2,
FLT3, BRCA2, RNF6, SUCLA2 |
The high-level copy number gains were analyzed in
the amplifications with a log2 ratio >0.5. A total of 41
amplified chromosome segmental regions were detected and are
summarized in Table III. Among
these regions, the 3q26.32q27.2 region was amplified in 8 cases and
gained in 23 cases, in which the size of the smallest region of
overlap (SRO) was ~9.48 Mb, including sex-determining region Y-box
2 (SOX2), eukaryotic translation initiation factor 4 gamma 1
(EIF4G1), fragile X-related gene 1 (FXR1), disheveled segment
polarity protein 3 (DVL3), defective n cullin neddylation 1 domain
containing 1 (DCUN1D1) and insulin like growth factor 2 mRNA
binding protein 2 (IGF2BP2) genes. The 8q24.21 region was amplified
in 7 cases and gained in 14 cases, with an SRO of ~672 kb in size,
including the CCDC26 long non-coding RNA (CCDC26) gene.
 | Table III.High copy number amplification/gain
segments and genes in supraglottic laryngeal squamous cell
carcinoma samples. |
Table III.
High copy number amplification/gain
segments and genes in supraglottic laryngeal squamous cell
carcinoma samples.
| No. | Chromosome | Amp | Gain | SRO | Size, Mb | Interesting
genes |
|---|
| 1 | 1p34.2p33 | 1 | 4 |
43184751-49838052 | 6.35 | MUTYH, NASP |
| 2 | 1p35.1p34.3 | 1 | 1 |
33752754-35149661 | 1.33 | CSMD2, DLGAP3 |
| 3 | 1q41q44 | 3 | 4 |
220602864-248250621 | 26.40 | ENAH, LEFTY, RAB4A,
CHML |
| 4 | 1q31.2q31.3 | 2 | 4 |
191220248-194858155 | 3.47 | UCHL5, RGS1,
CDC73 |
| 5 | 2p25.3p23.3 | 1 | 7 | 0-24153800 | 24.10 | TPO, DDX1, GREB1,
MYCN |
| 6 | 2q33.1 | 5 | 2 |
200862568-200901870 | 0.04 | CLK1, PPIL3 |
| 7 | 2q37.3 | 1 | 2 |
238403850-239073750 | 0.65 | TWIST2, HDAC4 |
| 8 | 3p12.1 | 1 | 3 |
84554500-85765787 | 1.21 | CADM2 |
| 9 | 3q26.32q27.2 | 8 | 23 |
176175382-185749890 | 9.48 | SOX2, EIF4G1, FXR1,
DVL3, DCUN1D1, IGF2BP2 |
| 10 | 5p15.33p13.3 | 2 | 24 | 0-31372588 | 31.30 | PDCD6, MTRR,
AHRR |
| 11 | 5q35.2q35.3 | 1 | 2 |
175916967-178314470 | 2.39 | FGFR4, NSD1,
GRK6 |
| 12 | 6p11.2 | 2 | 8 |
57317359-58726570 | 1.34 | PRIM2 |
| 13 | 6q21 | 1 | 6 |
108877605-110016036 | 1.13 | CD164, SESN1 |
| 14 | 7p11.2 | 3 | 14 |
54806239-56202945 | 1.39 | EGFR, VOPP1 |
| 15 | 7q22.1 | 3 | 6 |
98182043-100255629 | 2.07 | TRRAP, STAG3,
ARPC1B |
| 16 | 7q33q34 | 1 | 6 |
134991516-142413172 | 7.42 | AGK, HIPK2, SSBP1,
BRAF |
| 17 | 8p11.22 | 6 | 14 |
39419051-39849714 | 0.41 | ADAM2 |
| 18 | 8q24.21 | 7 | 14 |
128106781-128779314 | 0.66 | CCDC26 |
| 19 | 8q11.21 | 3 | 4 |
49155884-50221287 | 1.06 | SNTG1 |
| 20 | 9p24.1 | 3 | 2 |
7441659-8589761 | 1.10 | PTPRD |
| 21 | 10p11.21p11.1 | 1 | 9 |
37004391-39080681 | 2.07 | ZNF |
| 22 | 11p12 | 3 | 4 |
38695098-40869199 | 2.17 | LRRC4C |
| 23 | 11q13.1 | 2 | 8 |
64578176-65569224 | 0.97 | MEN1, CAPN1, EHD1,
BATF2 |
| 24 | 11q13.2q13.4 | 6 | 7 |
68442471-70556136 | 2.00 | CCND1, CTTN, FADD,
FGF4 |
| 25 | 12p12.1 | 3 | 11 |
23524152-26316175 | 2.79 | SOX5, BCAT1 |
| 26 | 12p13.33 | 3 | 13 | 141614-1969444 | 1.74 | WNT5B, RAD52 |
| 27 | 12q13.11q13.12 | 2 | 6 |
46221018-49404362 | 3.18 | VDR, HDAC7,
TROAP |
| 28 | 13q21.33q22.1 | 2 | 3 |
73340325-74328790 | 0.94 | KLF12 |
| 29 | 13q33.3q34 | 2 | 4 |
107817292-114754999 | 6.93 | LIG4, SOX1,
CDC16 |
| 30 | 14q11.2 | 3 | 7 |
20179223-20422416 | 0.23 | TEP1, PARP2 |
| 31 | 14q21.3q22.1 | 1 | 6 |
47205270-51017435 | 3.81 | CDKL1, POLE2 |
| 32 | 16q12.1 | 2 | 4 |
47280685-49712448 | 2.43 | ABCC11, PHKB |
| 33 | 18p11.32p11.31 | 3 | 18 | 0-3569480 | 3.56 | TYMS, CETN1,
USP14 |
| 34 | 18q11.1q22.3 | 1 | 4 |
18358334-69120712 | 50.70 | DSC1, LIPG, SMAD2,
MBD1 |
| 35 | 19q13.2 | 2 | 6 |
39598572-40098463 | 0.49 | CLC, FCGBP |
| 36 | 20p13p11.22 | 3 | 9 |
1193811-21924800 | 19.70 | PRNP, SNX5,
PLCB1 |
| 37 | 21q21.1 | 2 | 4 |
18548675-22735276 | 4.18 | NCAM2 |
| 38 | 22q11.21 | 4 | 7 |
18879043-21487181 | 2.60 | COMT, TBX1 |
| 39 | 22q13.31 | 2 | 7 |
45540029-46955726 | 1.41 | WNT7B |
| 40 | 22q13.33 | 2 | 7 |
49584858-49851099 | 0.26 | BRD1 |
| 41 | Xp11.23p11.22 | 2 | 4 |
48970719-49815538 | 0.82 | FOXP3, PLP2 |
Four interesting potential homozygous losses with a
log2 ratio >-0.4 were identified (Table IV), harboring putative tumor
suppressor genes, such as Nei-like DNA glycosylase 3 (NEIL3), CUB
and Sushi multiple domains 1 (CSMD1), cyclin dependent kinase
inhibitor 2A (CDKN2A) and protocadherin 20 (PCDH20).
 | Table IV.Potential homozygous segments and
target genes in supraglottic laryngeal squamous cell carcinoma
samples. |
Table IV.
Potential homozygous segments and
target genes in supraglottic laryngeal squamous cell carcinoma
samples.
| Chromosome
region | Log2
ratio | Genomic
coordinates, bp | Size, Mb | Target gene |
|---|
| 4q34.3 | 0.6 |
177027640-178012479 | 0.94 | NEIL3 |
| 8p23.3p23.2 | 0.8 | 0-4845472 | 4.84 | CSMD1 |
| 9p21.3 | 0.5 |
21952870-25578273 | 3.62 | CDKN2A |
| 13q21.2 | 0.6 |
61391312-614188052 | 1.10 | PCDH20 |
Genomic imbalances are associated with
lymph node metastasis, tumor stage, tumor differentiation and
smoking history
Patients with LSCC were divided into different
groups according to their clinicopathological characteristics as
follows: i) With or without lymph node metastasis; ii) tumor stages
I–II or III–IV; iii) well, moderate or poor differentiation; and
iv) with or without a smoking history. Differences in frequencies
(>10/38) of genomic aberrations among the different groups of
patients with supraglottic LSCC were analyzed using the
χ2 test, revealing a significant association between
3q26.1-qter gain and tumor stage, poor differentiation and smoking
history (Table V).
 | Table V.Genomic aberrations associated with
clinicopathological characteristics of the supraglottic laryngeal
squamous cell carcinoma samples. |
Table V.
Genomic aberrations associated with
clinicopathological characteristics of the supraglottic laryngeal
squamous cell carcinoma samples.
|
|
| Stage, n (%) |
|
| pN status, n
(%) |
|
| Differentiation, n
(%) |
|
| Smoking history, n
(%) |
|
|
|---|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|---|
| Chromosome | Gain/loss | Positive | Negative | c2 | P-value | Stage I–II | Stage III–IV | c2 | P-value | Well | Moderate | Poor | c2 | P-value | Yes | No | c2 | P-value |
|---|
| 3q26.1-qter | Yes | 15 | 16 | 0.070 | 0.791 | 8 | 23 | 5.281 | 0.022 | 5 | 25 | 1 | 9.243 | 0.004 | 31 | 0 | 14.424 | <0.001 |
| gain | (31) | (48.39) | (51.61) |
|
| (25.81) | (74.19) |
|
| (16.13) | (80.65) | (3.23) |
|
| (100.00) | (0.00) |
|
|
|
| No | 3 | 4 |
|
| 5 | 2 |
|
| 2 | 2 | 3 |
|
| 4 | 3 |
|
|
|
| (7) | (42.86) | (57.14) |
|
| (71.43) | (28.57) |
|
| (28.57) | (28.57) | (42.86) |
|
| (57.14) | (42.86) |
|
|
| 5pter-p12 | Yes | 12 | 14 | 0.049 | 0.825 | 8 | 18 | 0.433 | 0.510 | 4 | 18 | 4 | 2.005 | 0.342 | 25 | 1 | 1.856 | 0.173 |
| gain | (26) | (46.15) | (53.85) |
|
| (30.77) | (69.23) |
|
| (15.38) | (69.23) | (15.38) |
|
| (96.15) | (3.85) |
|
|
|
| No | 6 | 6 |
|
| 5 | 7 |
|
| 3 | 9 | 0 |
|
| 10 | 2 |
|
|
|
| (12) | (50.00) | (50.00) |
|
| (41.67) | (58.33) |
|
| (25.00) | (75.00) | (0.00) |
|
| (83.33) | (16.67) |
|
|
| 18pter-p11.31 | Yes | 7 | 13 | 2.591 | 0.107 | 7 | 13 | 0.012 | 0.914 | 3 | 15 | 2 | 0.372 | 0.830 | 18 | 2 | 0.257 | 0.618 |
| gain | (20) | (35.00) | (65.00) |
|
| (35.00) | (65.00) |
|
| (15.00) | (75.00) | (10.00) |
|
| (90.00) | (10.00) |
|
|
|
| No | 11 | 7 |
|
| 6 | 12 |
|
| 4 | 12 | 2 |
|
| 17 | 1 |
|
|
|
| (18) | (61.11) | (38.89) |
|
| (33.33) | (66.67) |
|
| (22.22) | (66.67) | (11.11) |
|
| (94.44) | (5.56) |
|
|
| 8q24.13-q23.3 | Yes | 10 | 9 | 0.422 | 0.516 | 5 | 14 | 1.052 | 0.305 | 2 | 14 | 3 | 2.193 | 0.341 | 17 | 2 | 0.362 | 0.547 |
| gain | (19) | (52.63) | (47.37) |
|
| (26.32) | (73.68) |
|
| (10.53) | (73.68) | (15.79) |
|
| (89.47) | (10.53) |
|
|
|
| No | 8 | 11 |
|
| 8 | 11 |
|
| 5 | 13 | 1 |
|
| 18 | 1 |
|
|
|
| (19) | (42.11) | (57.89) |
|
| (42.11) | (57.89) |
|
| (26.32) | (68.42) | (5.26) |
|
| (94.74) | (5.26) |
|
|
| 3pter-p21.31 | Yes | 9 | 11 | 0.095 | 0.758 | 8 | 12 | 0.629 | 0.428 | 5 | 14 | 1 | 2.096 | 0.396 | 18 | 2 | 0.257 | 0.618 |
| loss | (20) | (45.00) | (55.00) |
|
| (40.00) | (60.00) |
|
| (25.00) | (70.00) | (5.00) |
|
| (90.00) | (10.00) |
|
|
|
| No | 9 | 9 |
|
| 5 | 13 |
|
| 2 | 13 | 3 |
|
| 17 | 1 |
|
|
|
| (18) | (50.00) | (50.00) |
|
| (27.78) | (72.22) |
|
| (11.11) | (72.22) | (16.67) |
|
| (94.44) | (5.56) |
|
|
| 5q13.2-qter | Yes | 10 | 8 | 0.920 | 0.338 | 8 | 10 | 1.591 | 0.207 | 6 | 10 | 2 | 5.215 | 0.053 | 18 | 0 | 2.931 | 0.232 |
| loss | (18) | (55.56) | (44.44) |
|
| (44.44) | (55.56) |
|
| (33.33) | (55.56) | (11.11) |
|
| (100.00) | (0.00) |
|
|
|
| No | 8 | 12 |
|
| 5 | 15 |
|
| 1 | 17 | 2 |
|
| 17 | 3 |
|
|
|
| (20) | (40.00) | (60.00) |
|
| (25.00) | (75.00) |
|
| (5.00) | (85.00) | (10.00) |
|
| (85.00) | (15.00) |
|
|
Identification of DEGs between
patients with HNSCC and normal controls
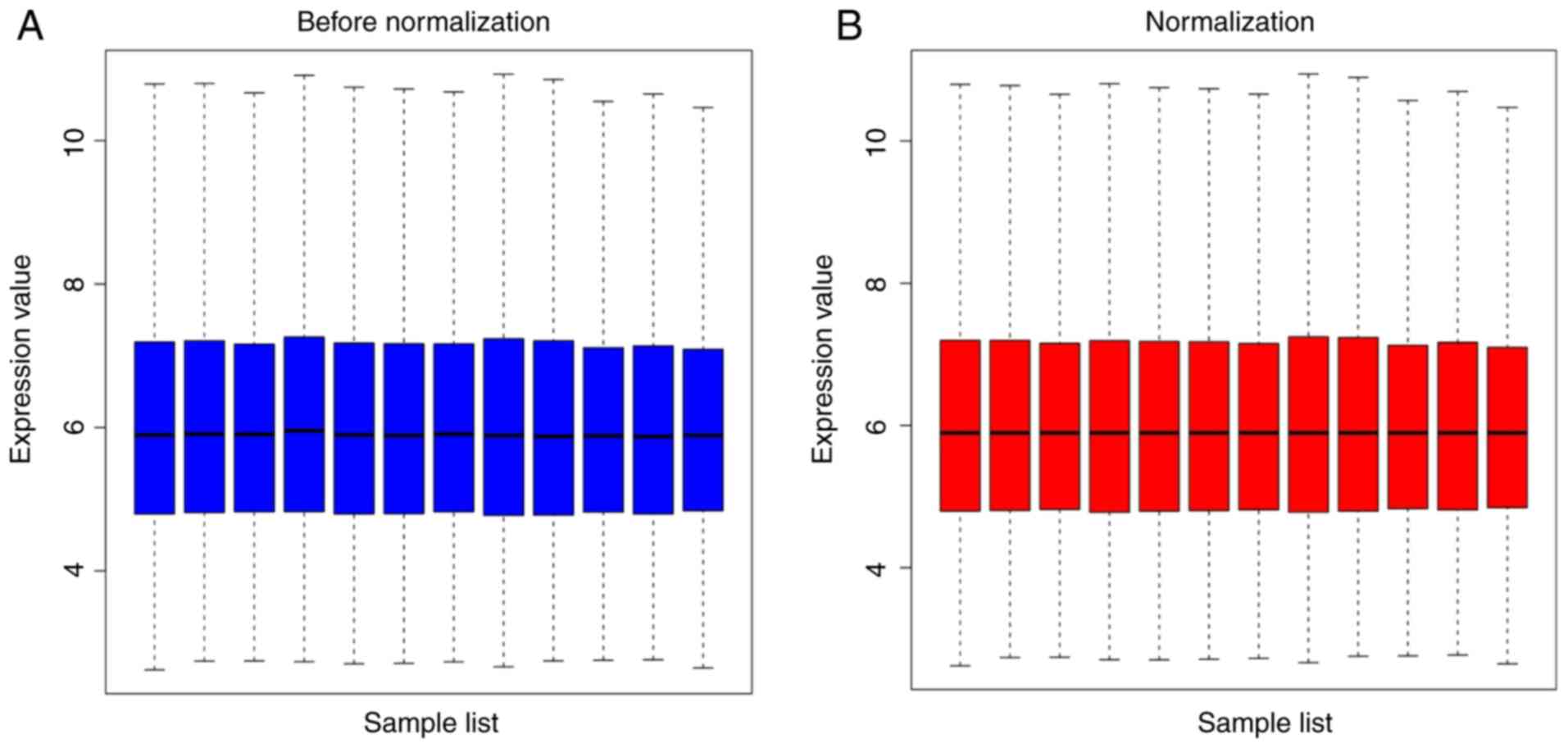
The expression levels in the HNSCC chip dataset
GSE6631 were normalized (Fig. 4). A
total of 140 DEGs were detected between 44 HNSCC tumors and their
paired normal mucosa samples using adjusted P<0.05 and |log
FC|>1 as cut-off criteria, among which 89 DEGs were
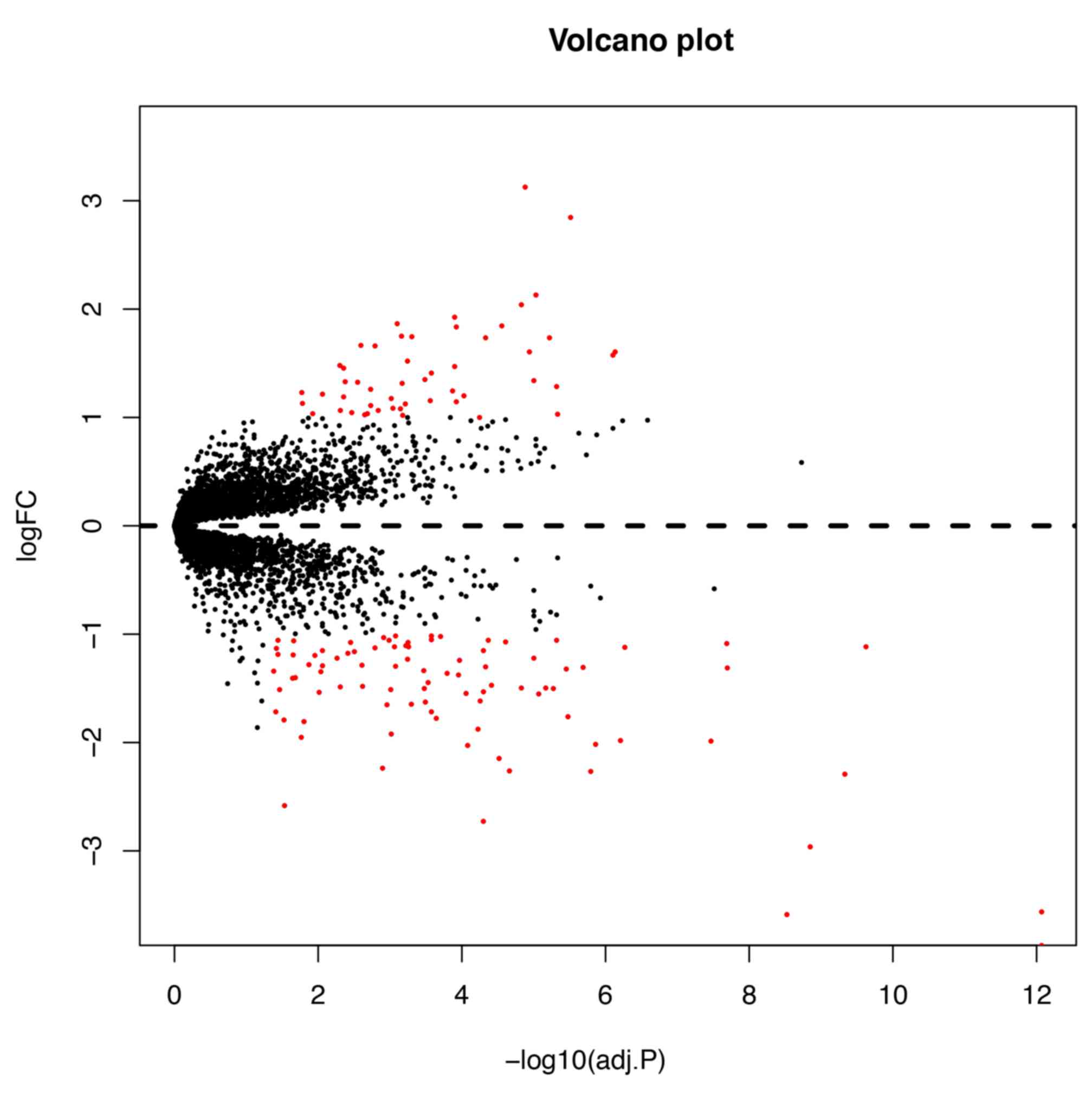
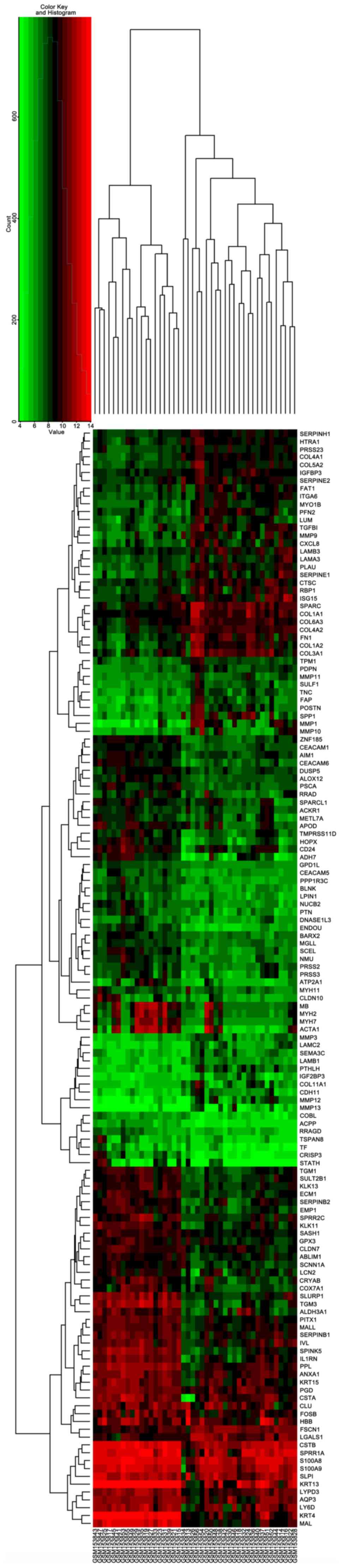
downregulated and 51 were upregulated. The volcano plot and heatmap
of DEGs are shown in Figs. 5 and
6, respectively. The chromosome
regions of upregulated and downregulated DEGs are listed in
Table VI.
 | Table VI.Differentially expressed genes with
|logFC|>2 and their chromosome regions. |
Table VI.
Differentially expressed genes with
|logFC|>2 and their chromosome regions.
| Gene symbol | logFC | Adj.P | Chromosome
region |
|---|
| MMP1 | 3.127560595 |
1.31×10−5 | 11q22.2 |
| SPP1 | 2.84711336 |
3.07×10−6 | 4q22.1 |
| FN1 | 2.129994375 |
9.38×10−6 | 2q35 |
| COL1A2 | 2.038465913 |
1.49×10−5 | 7q21.3 |
| KRT4 | −3.871821922 |
8.53×10−13 | 12q13.13 |
| TGM3 | −3.587447271 |
2.97×10−9 | 20p13 |
| MAL | −3.560291761 |
8.53×10−13 | 2q11.1 |
| SPINK5 | −2.962023367 |
1.41×10−9 | 5q32 |
| KRT13 | −2.725726018 |
5.06×10−5 | 17q21.2 |
| ENDOU | −2.289484712 |
4.67×10−10 | 12q13.1 |
| SLURP1 | −2.269044972 |
1.61×10−6 | 8q24.3 |
| HOPX | −2.264251872 |
2.17×10−5 | 4q12 |
| CRISP3 | −2.145114443 |
3.06×10−5 | 6p12.3 |
| IL1RN | −2.024761541 |
8.36×10−5 | 2q14.1 |
| PPL | −2.019045519 |
1.37×10−6 | 16p13.3 |
Candidate target genes of gains and
losses in supraglottic LSCC
The Affymetrix microarray expression profiling data
of HNSCC and the gene expression profiling of supraglottic LSCC
samples detected by aCGH were integratively analyzed to identify
the potential target genes of downregulated serine peptidase
inhibitor Kazal type 5 (SPINK5), which is closely associated with
squamous cell carcinoma. Analysis of candidate target genes of
gains in 3q26.32q27.2 and 8q24.21 revealed upregulation of SOX2,
EIF4G1, FXR1, DVL3, DCUN1D1, IGF2BP2 and CCDC26 (Table III), and downregulation of CDKN2A,
SPINK5, PCDH20, CSMD1 and NEIL3 in supraglottic LSCC (Table IV).
Discussion
The present study evaluated genetic alternations in
LSCC using aCGH and bioinformatics analysis of microarray
expression profiling. Primary LSCC accounts for 20% of all HNSCC
and is associated with a high risk of developing metastatic
squamous cell carcinoma (15).
Supraglottic LSCC is emerging as the second most common type of
laryngeal malignancy (5). In the
present study, 38 primary supraglottic LSCC cases and paired
controls were screened via aCGH and the high-density Affymetrix
Genome-wide Human Genome U95 Version 2 Array from the GEO database
to analyze CNVs (gains, losses, amplifications and homozygous
deletions) and their association with clinicopathological
characteristics. Important roles of certain chromosomal regions and
gene loci were identified for the biology and prognosis of
supraglottic LSCC.
Chromosomal instability is a numerical and
structural chromosomal abnormality commonly resulting in DNA CNVs
and genetic heterogeneity that may trigger cancer development
processes, including LSCC tumorigenesis (16). Compared with the detection of
expression profiles using conventional karyotyping methods, aCGH
has been demonstrated to be more effective in the determination of
molecular profiles with a 5–10 times higher resolution (17,18). In
the present study, a total of 598 gains and amplifications, and 348
losses were identified using aGCH. The frequency of gains and
amplifications was 1.72 times higher than that of losses. A total
of 21/38 cases with genomic imbalances had net genomic gains
(range, 54.86–1,958.63 Mb) and 17/38 cases had net genomic losses
(range, 22.02–849.44 Mb), demonstrating a wider range in the net
genomic gains than in the losses. The most frequent genomic
imbalances detected in the present study included gains such as
3q26.1-qter (31/38), 5pter-p12 (26/38) and 18pter-p11.31 (20/38),
and losses such as 3pter-p21.32 (20/38), 5q13.2-qter (18/38) and
9pter-p21.3 (16/38). Previous studies have reported that the gain
of genomic material of 3q has a high prevalence in squamous cell
carcinomas originating from the head and neck region (19,20), and
that it is associated with disease-specific survival time (21), consistent with previous studies in
LSCC (22–24). The chromosomal region of 3q,
particularly 3q26-29, was demonstrated to be a defining feature of
squamous cell carcinomas, with an ~75% predicted occurrence in
HNSCC (25). Furthermore, gains of
11q13 and loss of 18q23 were positively associated with a poor
outcome in patients with LSCC (23,26,27).
In the present study, amplifications were detected
in 41 segmental regions, among which 3q26.32q27.2 and 8q24.21 were
the most repeatedly involved interesting regions. Amplification of
3q26.32q27.2 was one of the most relevant findings of the present
study. In 31/38 cases with copy number gains of 3q26.1-qter, 23
cases exhibited gains and 8 cases amplifications. Amplification of
the 3q26q27 region is one of the most recurrent genetic alterations
in HNSCC and other carcinomas, and has been demonstrated to be
associated with tumor progression and a poor prognosis (20,28). In
the present study, the SRO located in locus 3q26q27 harbored
several oncogenes, including SOX2, EIF4G1, FXR1, DVL3, DCUN1D1 and
IGF2BP2. SOX2 has been demonstrated to be a candidate tumor driver
gene in squamous cell carcinoma of the lung and esophagus (29–31).
EIF4G1 is an RNA-binding protein that serves an important role in
the occurrence and development of breast cancer, squamous cell lung
carcinoma and other types of cancer (32,33). Jin
et al (34) reported an
upregulation of FXR1 expression in colorectal cancer, in which it
acted as an oncogene. DVL3 is a family member of the Wnt receptor
complex and the most critical regulator of the Wnt signaling
pathway, and is involved in cervical, breast, liver, prostate, lung
and esophageal cancer (35). DCUN1D1
expression is upregulated in human laryngeal carcinoma tissues
(36). IGF2BP2, also known as IMP2,
has been reported to be dysregulated in several types of cancer,
including colorectal cancer (37),
colon cancer (38), esophageal
adenocarcinoma (39) and breast
cancer (40).
Notably, the reciprocal loss of 3p and gain of 3q
were detected in 20/38 cases in the present study, together with
18/38 cases of reciprocal gain of 5p and loss of 5q. Frequent
occurrence of the reciprocal loss and gain of chromosomes 3 and 5
were observed in various epithelial tumors. In particular, the
isochromosome 3q has been detected in lung cancer and HNSCC
(41,42). Therefore, formation of isochromosome
3q may be an intermediate mechanism of somatic chromosomal
aberrations, leading to a reciprocal change during epithelial cell
carcinogenesis.
In addition, 3 cases of amplification and 14 of gain
in the 7p11.2 chromosomal fragment harboring the oncogene epidermal
growth factor receptor (EGFR) were detected in the present study.
As a tyrosine kinase receptor, EGFR is widely expressed in human
epithelial cell membranes. Amplification and upregulation of EGFR
have been observed in patients with LSCC and esophageal squamous
cell carcinoma (ESCC) with a poor prognosis, serving a crucial role
in ESCC progression (43). The
association analysis between genomic aberrations and
clinicopathological features in the present study revealed that
gains of 3q26.1-qter were associated with tumor stage, poor
differentiation and smoking history.
Four homozygous segments possibly containing
putative tumor suppressor genes were discovered in the present
study, including NEIL3, CSMD1, CDKN2A and PCDH20. CDKN2A is
considered to be the most commonly affected gene after TP53 in
terms of homozygous deletion, promoter hypermethylation, loss of
heterozygosity and point mutations in various types of cancer,
including LSCC (44,45). CSMD1 has been demonstrated to act as
a tumor suppressor in human breast, colorectal and head and neck
cancer (46–48), whereas PCDH20 has been proposed as a
tumor suppressor gene in hepatic carcinoma and lung cancer
(49,50). However, the role of NEIL3 as a tumor
suppressor gene remains unclear. Wang et al (51) proposed SPINK5 as a novel tumor
suppressor targeting the Wnt/β-catenin signaling pathway in
esophageal cancer.
In conclusion, using integration analysis of CNVs
from the GEO database and the gene expression profiling of
supraglottic LSCC samples detected via aCGH, potential candidate
target genes with strong clinical significance were identified,
such as the upregulation of SOX2, EIF4G1, FXR1, DVL3, DCUN1D1,
IGF2BP2 and CCDC26 expression, and downregulation of CDKN2A,
SPINK5, PCDH20, CSMD1 and NEIL3 expression, providing novel
insights into the molecular pathological mechanisms underlying
supraglottic LSCC. However, the findings of the present study were
limited by the relatively small sample size. Future studies should
involve the cooperation of researchers from different countries and
regions to further verify the findings of the present study, and
should investigate the mechanisms underlying the development of
supraglottic LSCC, potentially leading to the identification of
novel diagnostic and therapeutic biomarkers for supraglottic
LSCC.
Acknowledgements
Not applicable.
Funding
No funding was received.
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request. The GSE6631 dataset is available from the GEO database
(https://www.ncbi.nlm.nih.gov/sites/GDSbrowser?acc=GDS2520).
Authors' contributions
WZ conceived and designed the study. WY, PW and XiaW
acquired the data. DL and SLu analyzed and interpreted the data. DL
drafted the manuscript. SLi and XinW critically revised the
manuscript for important intellectual content and analyzed the
data. WZ and SLi approved the final manuscript. All authors read
and approved the final manuscript.
Ethics approval and consent to
participate
Written informed consent was provided by all
patients enrolled in the present study with the authorization of
the Ethics Committee of The First Hospital of Jilin University
(approval no. 2017-415; Changchun, China).
Patient consent for publication
Written informed consent for publication was
provided by all patients.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Jemal A, Bray F, Center MM, Ferlay J, Ward
E and Forman D: Global cancer statistics. CA Cancer J Clin.
61:69–90. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Siegel RL, Miller KD and Jemal A: Cancer
statistics, 2016. CA Cancer J Clin. 66:7–30. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Yamazaki H, Suzuki G, Nakamura S, Hirano
S, Yoshida K, Konish K, Teshima T and Ogawa K: Radiotherapy for
locally advanced resectable T3-T4 laryngeal cancer-does laryngeal
preservation strategy compromise survival? J Radiat Res. 59:77–90.
2018. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Marioni G, Marchese-Ragona R, Cartei G,
Marchese F and Staffieri A: Current opinion in diagnosis and
treatment of laryngeal carcinoma. Cancer Treat Rev. 32:504–515.
2006. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Yilmaz T, Suslu N, Atay G, Gunaydin RO,
Bajin MD and Ozer S: The effect of midline crossing of lateral
supraglottic cancer on contralateral cervical lymph node
metastasis. Acta Otolaryngol. 135:484–488. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Elegbede AI, Rybicki LA, Adelstein DJ,
Kaltenbach JA, Lorenz RR, Scharpf J and Burkey BB: Oncologic and
functional outcomes of surgical and nonsurgical treatment of
advanced squamous cell carcinoma of the supraglottic larynx. JAMA
Otolaryngol Head Neck Surg. 141:1111–1117. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Koirala K: Epidemiological study of
laryngeal carcinoma in Western Nepal. Asian Pac J Cancer Prev.
16:6541–6544. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Fan CY: Genetic alterations in head and
neck cancer: Interactions among environmental carcinogens, cell
cycle control, and host DNA repair. Curr Oncol Rep. 3:66–71. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Conrad DF, Pinto D, Redon R, Feuk L,
Gokcumen O, Zhang Y, Aerts J, Andrews TD, Barnes C, Campbell P, et
al: Origins and functional impact of copy number variation in the
human genome. Nature. 464:704–712. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Hu X, Moon JW, Li S, Xu W, Wang X, Liu Y
and Lee JY: Amplification and overexpression of CTTN and CCND1 at
chromosome 11q13 in Esophagus squamous cell carcinoma (ESCC) of
North Eastern Chinese population. Int J Med Sci. 13:868–874. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Beroukhim R, Mermel CH, Porter D, Wei G,
Raychaudhuri S, Donovan J, Barretina J, Boehm JS, Dobson J,
Urashima M, et al: The landscape of somatic copy-number alteration
across human cancers. Nature. 463:899–905. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Edge SB and Compton CC: The American joint
committee on cancer: The 7th edition of the AJCC cancer staging
manual and the future of TNM. Ann Surg Oncol. 17:1471–1474. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Kuriakose MA, Chen WT, He ZM, Sikora AG,
Zhang P, Zhang ZY, Qiu WL, Hsu DF, McMunn-Coffran C, Brown SM, et
al: Selection and validation of differentially expressed genes in
head and neck cancer. Cell Mol Life Sci. 61:1372–1383. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Wickham H: ggplot2: Elegant graphics for
data analysis. Springer-Verlag; New York: ISBN 978-3-319-24277-4.
2016
|
|
15
|
Karatzanis AD, Psychogios G, Waldfahrer F,
Kapsreiter M, Zenk J, Velegrakis GA and Iro H: Management of
locally advanced laryngeal cancer. J Otolaryngol Head Neck Surg.
43:42014. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Myllykangas S, Tikka J, Bohling T,
Knuutila S and Hollmen J: Classification of human cancers based on
DNA copy number amplification modeling. BMC Med Genomics. 1:152008.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Vissers LE, de Vries BB, Osoegawa K,
Janssen IM, Feuth T, Choy CO, Straatman H, van der Vliet W, Huys
EH, van Rijk A, et al: Array-based comparative genomic
hybridization for the genomewide detection of submicroscopic
chromosomal abnormalities. Am J Hum Genet. 73:1261–1270. 2003.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Shaw-Smith C, Redon R, Rickman L, Rio M,
Willatt L, Fiegler H, Firth H, Sanlaville D, Winter R, Colleaux L,
et al: Microarray based comparative genomic hybridisation
(array-CGH) detects submicroscopic chromosomal deletions and
duplications in patients with learning disability/mental
retardation and dysmorphic features. J Med Genet. 41:241–248. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Riazimand SH, Welkoborsky HJ, Bernauer HS,
Jacob R and Mann WJ: Investigations for fine mapping of
amplifications in chromosome 3q26.3–28 frequently occurring in
squamous cell carcinomas of the head and neck. Oncology.
63:385–392. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Singh B, Stoffel A, Gogineni S, Poluri A,
Pfister DG, Shaha AR, Pathak A, Bosl G, Cordon-Cardo C, Shah JP and
Rao PH: Amplification of the 3q26.3 locus is associated with
progression to invasive cancer and is a negative prognostic factor
in head and neck squamous cell carcinomas. Am J Pathol.
161:365–371. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Ashman JN, Patmore HS, Condon LT, Cawkwell
L, Stafford ND and Greenman J: Prognostic value of genomic
alterations in head and neck squamous cell carcinoma detected by
comparative genomic hybridisation. Br J Cancer. 89:864–869. 2003.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Huang Q, Yu GP, McCormick SA, Mo J, Datta
B, Mahimkar M, Lazarus P, Schaffer AA, Desper R and Schantz SP:
Genetic differences detected by comparative genomic hybridization
in head and neck squamous cell carcinomas from different tumor
sites: Construction of oncogenetic trees for tumor progression.
Genes Chromosomes Cancer. 34:224–233. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Jarmuz-Szymczak M, Pelinska K,
Kostrzewska-Poczekaj M, Bembnista E, Giefing M, Brauze D,
Szaumkessel M, Marszalek A, Janiszewska J, Kiwerska K, et al:
Heterogeneity of 11q13 region rearrangements in laryngeal squamous
cell carcinoma analyzed by microarray platforms and fluorescence in
situ hybridization. Mol Biol Rep. 40:4161–4171. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Patmore HS, Ashman JN, Stafford ND,
Berrieman HK, MacDonald A, Greenman J and Cawkwell L: Genetic
analysis of head and neck squamous cell carcinoma using comparative
genomic hybridisation identifies specific aberrations associated
with laryngeal origin. Cancer Lett. 258:55–62. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Fields AP, Justilien V and Murray NR: The
chromosome 3q26 OncCassette: A multigenic driver of human cancer.
Adv Biol Regul. 60:47–63. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Ambrosio EP, Silveira CG, Drigo SA,
Sacomano Vde S, Molck MC, Rocha RM, Domingues MA, Soares FA,
Kowalski LP and Rogatto SR: Chromosomal imbalances exclusively
detected in invasive front area are associated with poor outcome in
laryngeal carcinomas from different anatomical sites. Tumour Biol.
34:3015–3026. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Jarvinen AK, Autio R, Haapa-Paananen S,
Wolf M, Saarela M, Grenman R, Leivo I, Kallioniemi O, Makitie AA
and Monni O: Identification of target genes in laryngeal squamous
cell carcinoma by high-resolution copy number and gene expression
microarray analyses. Oncogene. 25:6997–7008. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Singh B, Gogineni SK, Sacks PG, Shaha AR,
Shah JP, Stoffel A and Rao PH: Molecular cytogenetic
characterization of head and neck squamous cell carcinoma and
refinement of 3q amplification. Cancer Res. 61:4506–4513.
2001.PubMed/NCBI
|
|
29
|
Gen Y, Yasui K, Zen Y, Zen K, Dohi O, Endo
M, Tsuji K, Wakabayashi N, Itoh Y, Naito Y, et al: SOX2 identified
as a target gene for the amplification at 3q26 that is frequently
detected in esophageal squamous cell carcinoma. Cancer Genet
Cytogenet. 202:82–93. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Hussenet T, Dali S, Exinger J, Monga B,
Jost B, Dembele D, Martinet N, Thibault C, Huelsken J, Brambilla E
and du Manoir S: SOX2 is an oncogene activated by recurrent 3q26.3
amplifications in human lung squamous cell carcinomas. PLoS One.
5:e89602010. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Weina K and Utikal J: SOX2 and cancer:
Current research and its implications in the clinic. Clin Transl
Med. 3:192014. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Badura M, Braunstein S, Zavadil J and
Schneider RJ: DNA damage and eIF4G1 in breast cancer cells
reprogram translation for survival and DNA repair mRNAs. Proc Natl
Acad Sci USA. 109:18767–18772. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Bauer C, Brass N, Diesinger I, Kayser K,
Grasser FA and Meese E: Overexpression of the eukaryotic
translation initiation factor 4G (eIF4G-1) in squamous cell lung
carcinoma. Int J Cancer. 98:181–185. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Jin X, Zhai B, Fang T, Guo X and Xu L:
FXR1 is elevated in colorectal cancer and acts as an oncogene.
Tumour Biol. 37:2683–2690. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Chen XQ, Jiang J, Wang XT, Zhang CL, Ji AY
and Chen XJ: Role and mechanism of Dvl3 in the esophageal squamous
cell carcinoma. Eur Rev Med Pharmacol Sci. 22:7716–7725.
2018.PubMed/NCBI
|
|
36
|
Shuang Y, Li C, Zhou X, Huang Y and Zhang
L: MicroRNA-195 inhibits growth and invasion of laryngeal carcinoma
cells by directly targeting DCUN1D1. Oncol Rep. 38:2155–2165. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Ye S, Song W, Xu X, Zhao X and Yang L:
IGF2BP2 promotes colorectal cancer cell proliferation and survival
through interfering with RAF-1 degradation by miR-195. FEBS Lett.
590:1641–1650. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Liu W, Li Z, Xu W, Wang Q and Yang S:
Humoral autoimmune response to IGF2 mRNA-binding protein (IMP2/p62)
and its tissue-specific expression in colon cancer. Scand J
Immunol. 77:255–260. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Barghash A, Golob-Schwarzl N, Helms V,
Haybaeck J and Kessler SM: Elevated expression of the IGF2 mRNA
binding protein 2 (IGF2BP2/IMP2) is linked to short survival and
metastasis in esophageal adenocarcinoma. Oncotarget. 7:49743–49750.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Liu W, Li Y, Wang B, Dai L, Qian W and
Zhang JY: Autoimmune response to IGF2 mRNA-Binding Protein 2
(IMP2/p62) in breast cancer. Scand J Immunol. 81:502–507. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Manor E, Tetro S and Bodner L:
Translocation (12;14) and other chromosome abnormalities in
squamous cell carcinoma of the tongue. Eur Arch Otorhinolaryngol.
267:1273–1276. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Tai AL, Mak W, Ng PK, Chua DT, Ng MY, Fu
L, Chu KK, Fang Y, Qiang Song Y, Chen M, et al: High-throughput
loss-of-heterozygosity study of chromosome 3p in lung cancer using
single-nucleotide polymorphism markers. Cancer Res. 66:4133–4138.
2006. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Jiang D, Li X, Wang H, Shi Y, Xu C, Lu S,
Huang J, Xu Y, Zeng H, Su J, et al: The prognostic value of EGFR
overexpression and amplification in Esophageal squamous cell
carcinoma. BMC Cancer. 15:3772015. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Hu N, Wang C, Su H, Li WJ, Emmert-Buck MR,
Li G, Roth MJ, Tang ZZ, Lu N, Giffen C, et al: High frequency of
CDKN2A alterations in esophageal squamous cell carcinoma from a
high-risk Chinese population. Genes Chromosomes Cancer. 39:205–216.
2004. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Willem P, Brown J and Schouten J: A novel
approach to simultaneously scan genes at fragile sites. BMC Cancer.
6:2052006. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Escudero-Esparza A, Bartoschek M, Gialeli
C, Okroj M, Owen S, Jirstrom K, Orimo A, Jiang WG, Pietras K and
Blom AM: Complement inhibitor CSMD1 acts as tumor suppressor in
human breast cancer. Oncotarget. 7:76920–76933. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Jung AR, Eun YG, Lee YC, Noh JK and Kwon
KH: Clinical significance of CUB and Sushi multiple domains 1
inactivation in head and neck squamous cell carcinoma. Int J Mol
Sci. 19(pii): E39962018. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Zhang R and Song C: Loss of CSMD1 or 2 may
contribute to the poor prognosis of colorectal cancer patients.
Tumour Biol. 35:4419–4423. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Lv J, Zhu P, Yang Z, Li M, Zhang X, Cheng
J, Chen X and Lu F: PCDH20 functions as a tumour-suppressor gene
through antagonizing the Wnt/β-catenin signalling pathway in
hepatocellular carcinoma. J Viral Hepat. 22:201–211. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Imoto I, Izumi H, Yokoi S, Hosoda H,
Shibata T, Hosoda F, Ohki M, Hirohashi S and Inazaw J: Frequent
silencing of the candidate tumor suppressor PCDH20 by epigenetic
mechanism in non-small-cell lung cancers. Cancer Res. 66:4617–4626.
2006. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Wang Q, Lv Q, Bian H, Yang L, Guo KL, Ye
SS, Dong XF and Tao LL: A novel tumor suppressor SPINK5 targets
Wnt/β-catenin signaling pathway in esophageal cancer. Cancer Med.
8:2360–2371. 2019. View Article : Google Scholar : PubMed/NCBI
|