|
1
|
Allemani C, Matsuda T, Di Carlo V,
Harewood R, Matz M, Nikšić M, Bonaventure A, Valkov M, Johnson CJ,
Estève J, et al: Global surveillance of trends in cancer survival
2000–14 (CONCORD-3): Analysis of individual records for 37 513 025
patients diagnosed with one of 18 cancers from 322 population-based
registries in 71 countries. Lancet. 391:1023–1075. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Venook AP, Papandreou C, Furuse J and de
Guevara LL: The incidence and epidemiology of hepatocellular
carcinoma: A global and regional perspective. Oncologist. 15 (Suppl
4):S5–S13. 2010. View Article : Google Scholar
|
|
3
|
Stepien M, Fedirko V, Duarte-Salles T,
Ferrari P, Freisling H, Trepo E, Trichopoulou A, Bamia C,
Weiderpass E, Olsen A, et al: Prospective association of liver
function biomarkers with development of hepatobiliary cancers.
Cancer Epidemiol. 40:179–187. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Bhangui P, Adam R, Vibert E, Azoulay D,
Samuel D and Castaing D: 41 resection or transplantation for early
hepatocellular carcinoma in a cirrhotic liver-size does matter. J
Clin Exp Hepatol. 1:1512011. View Article : Google Scholar
|
|
5
|
Simoneau E, Hassanain M, Madkhali A,
Salman A, Nudo CG, Chaudhury P and Metrakos P:
(18)F-Fluorodeoxyglucose positron-emission tomography could have a
prognostic role in patients with advanced hepatocellular carcinoma.
Curr Oncol. 21:e551–e556. 2014. View Article : Google Scholar : PubMed/NCBIPubMed/NCBI
|
|
6
|
Trevisani F, Cantarini MC, Wands JR and
Bernardi M: Recent advances in the natural history of
hepatocellular carcinoma. Carcinogenesis. 29:1299–1305. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Zheng H, Zou AE, Saad MA, Wang XQ, Kwok
JG, Korrapati A, Li P, Kisseleva T, Wang-Rodriguez J and Ongkeko
WM: Alcohol-dysregulated microRNAs in hepatitis B virus-related
hepatocellular carcinoma. PLoS One. 12:e01785472017. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Wang XJ, Zhang DL, Xu ZG, Ma ML, Wang WB,
Li LL, Han XL, Huo Y, Yu X and Sun JP: Understanding cadherin EGF
LAG seven-pass G-type receptors. J Neurochem. 131:699–711. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Usui T, Shima Y, Shimada Y, Hirano S,
Burgess RW, Schwarz TL, Takeichi M and Uemura T: Flamingo, a
seven-pass transmembrane cadherin, regulates planar cell polarity
under the control of Frizzled. Cell. 98:585–595. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Langenhan T, Aust G and Hamann J: Sticky
signaling-adhesion class G protein-coupled receptors take the
stage. Sci Signal. 6:re32013. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Jeong P, Ha YS, Cho IC, Yun SJ, Yoo ES,
Kim IY, Choi YH, Moon SK and Kim WJ: Three-gene signature predicts
disease progression of non-muscle invasive bladder cancer. Oncol
Lett. 2:679–684. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Anwar SL and Lehmann U: DNA methylation,
microRNAs, and their crosstalk as potential biomarkers in
hepatocellular carcinoma. World J Gastroenterol. 20:7894–7913.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Ricketts CJ, Morris MR, Gentle D, Brown M,
Wake N, Woodward ER, Clarke N, Latif F and Maher ER: Genome-wide
CpG island methylation analysis implicates novel genes in the
pathogenesis of renal cell carcinoma. Epigenetics. 7:278–290. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Erkan M, Weis N, Pan Z, Schwager C,
Samkharadze T, Jiang X, Wirkner U, Giese NA, Ansorge W, Debus J, et
al: Organ-, inflammation- and cancer specific transcriptional
fingerprints of pancreatic and hepatic stellate cells. Mol Cancer.
9:882010. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
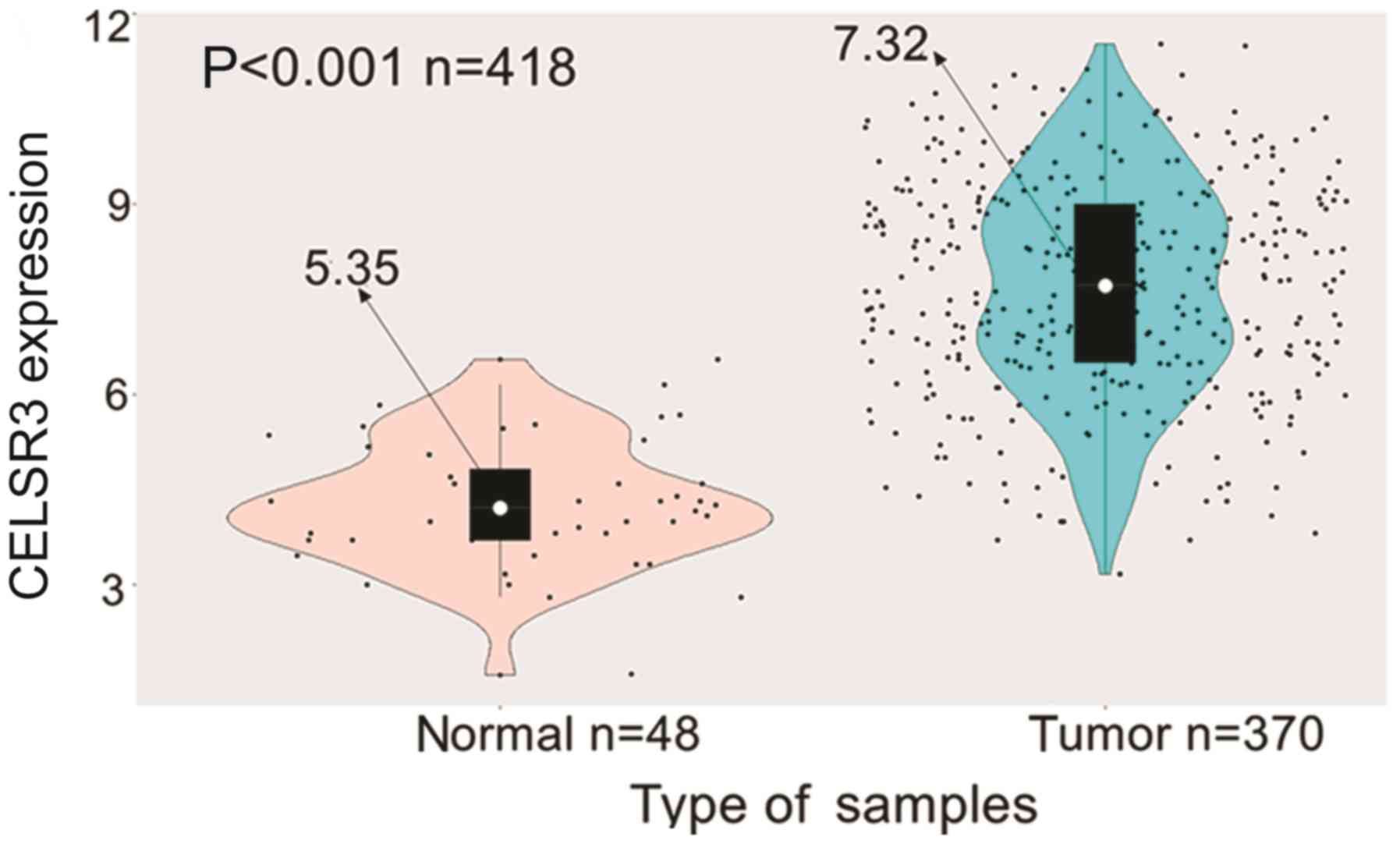
Gu X, Li H, Sha L, Mao Y, Shi C and Zhao
W: CELSR3 mRNA expression is increased in hepatocellular carcinoma
and indicates poor prognosis. PeerJ. 7:e78162019. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Khor GH, Froemming GR, Zain RB, Abraham TM
and Lin TK: Involvement of CELSR3 hypermethylation in primary oral
squamous cell carcinoma. Asian Pac J Cancer Prev. 17:219–223. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Xie Z, Dang Y, Wu H, He R, Ma J, Peng Z,
Rong M, Li Z, Yang J, Jiang Y, et al: Effect of CELSR3 on the cell
cycle and apoptosis of hepatocellular carcinoma cells. J Cancer.
11:2830–2844. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Hayes CN and Chayama K: MicroRNAs as
biomarkers for liver disease and hepatocellular carcinoma. Int J
Mol Sci. 17:2802016.
|
|
19
|
Tat TT, Maroney PA, Chamnongpol S, Coller
J and Nilsen TW: Cotranslational microRNA mediated messenger RNA
destabilization. Elife. 5:e128802016. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Zhang X, Xu X, Ge G, Zang X, Shao M, Zou
S, Zhang Y, Mao Z, Zhang J, Mao F, et al: miR498 inhibits the
growth and metastasis of liver cancer by targeting ZEB2. Oncol Rep.
41:1638–1648. 2019.PubMed/NCBI
|
|
21
|
Liu H: MicroRNAs in breast cancer
initiation and progression. Cell Mol Life Sci. 69:3587–3599. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Chu R, Mo G, Duan Z, Huang M, Chang J, Li
X and Liu P: miRNAs affect the development of hepatocellular
carcinoma via dysregulation of their biogenesis and expression.
Cell Commun Signal. 12:452014. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Fang T, Lv H, Lv G, Li T, Wang C, Han Q,
Yu L, Su B, Guo L, Huang S, et al: Tumor-derived exosomal
miR-1247-3p induces cancer-associated fibroblast activation to
foster lung metastasis of liver cancer. Nat Commun. 9:1912018.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Liu J, Lichtenberg T, Hoadley KA, Poisson
LM, Lazar AJ, Cherniack AD, Kovatich AJ, Benz CC, Levine DA, Lee
AV, et al: An integrated TCGA pan-cancer clinical data resource to
drive high-quality survival outcome analytics. Cell.
173:400–416.e11. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Robinson MD, McCarthy DJ and Smyth GK:
edgeR: A Bioconductor package for differential expression analysis
of digital gene expression data. Bioinformatics. 26:139–140. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Lu TX and Rothenberg ME: MicroRNA. J
Allergy Clin Immunol. 141:1202–1207. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Kertesz M, Iovino N, Unnerstall U, Gaul U
and Segal E: The role of site accessibility in microRNA target
recognition. Nat Genet. 39:1278–1284. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Rehmsmeier M, Steffen P, Hochsmann M and
Giegerich R: Fast and effective prediction of microRNA/target
duplexes. RNA. 10:1507–1517. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Chung TK, Lau TS, Cheung TH, Yim SF, Lo
KW, Siu NS, Chan LK, Yu MY, Kwong J, Doran G, et al: Dysregulation
of microRNA-204 mediates migration and invasion of endometrial
cancer by regulating FOXC1. Int J Cancer. 130:1036–1045. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Shannon P, Markiel A, Ozier O, Baliga NS,
Wang JT, Ramage D, Amin N, Schwikowski B and Ideker T: Cytoscape: A
software environment for integrated models of biomolecular
interaction networks. Genome Res. 13:2498–2504. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Xie C, Mao X, Huang J, Ding Y, Wu J, Dong
S, Kong L, Gao G, Li CY and Wei L: KOBAS 2.0: A web server for
annotation and identification of enriched pathways and diseases.
Nucleic Acids Res. 39:W316–W322. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Lee JW, Son MH, Cho HW, Ma YE, Yoo KH,
Sung KW and Koo HH: Clinical significance of MYCN amplification in
patients with high-risk neuroblastoma. Pediatr Blood Cancer.
65:e272572018. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Laisné M, Gupta N, Kirsh O, Pradhan S and
Defossez PA: Mechanisms of DNA methyltransferase recruitment in
mammals. Genes (Basel). 9:6172018. View Article : Google Scholar
|
|
35
|
Roos L, van Dongen J, Bell CG, Burri A,
Deloukas P, Boomsma DI, Spector TD and Bell J: Integrative DNA
methylome analysis of pan-cancer biomarkers in cancer discordant
monozygotic twin-pairs. Clin Epigenetics. 8:72016. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Li H, Zhao X, Li C, Sheng C and Bai Z:
Integrated analysis of lncRNA-associated ceRNA network reveals
potential biomarkers for the prognosis of hepatitis B virus-related
hepatocellular carcinoma. Cancer Manag Res. 11:877–897. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Zhao Y, Xue F, Sun J, Guo S, Zhang H, Qiu
B, Geng J, Gu J, Zhou X, Wang W, et al: Genome-wide methylation
profiling of the different stages of hepatitis B virus-related
hepatocellular carcinoma development in plasma cell-free DNA
reveals potential biomarkers for early detection and high-risk
monitoring of hepatocellular carcinoma. Clin Epigenetics. 6:302014.
View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Kazi JU and Rönnstrand L: FMS-like
Tyrosine kinase 3/FLT3: From basic science to clinical
implications. Physiol Rev. 99:1433–1466. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Lebron MB, Brennan L, Damoci CB, Prewett
MC, O'Mahony M, Duignan IJ, Credille KM, DeLigio JT, Starodubtseva
M, Amatulli M, et al: A human monoclonal antibody targeting the
stem cell factor receptor (c-Kit) blocks tumor cell signaling and
inhibits tumor growth. Cancer Biol Ther. 15:1208–1218. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
40
|
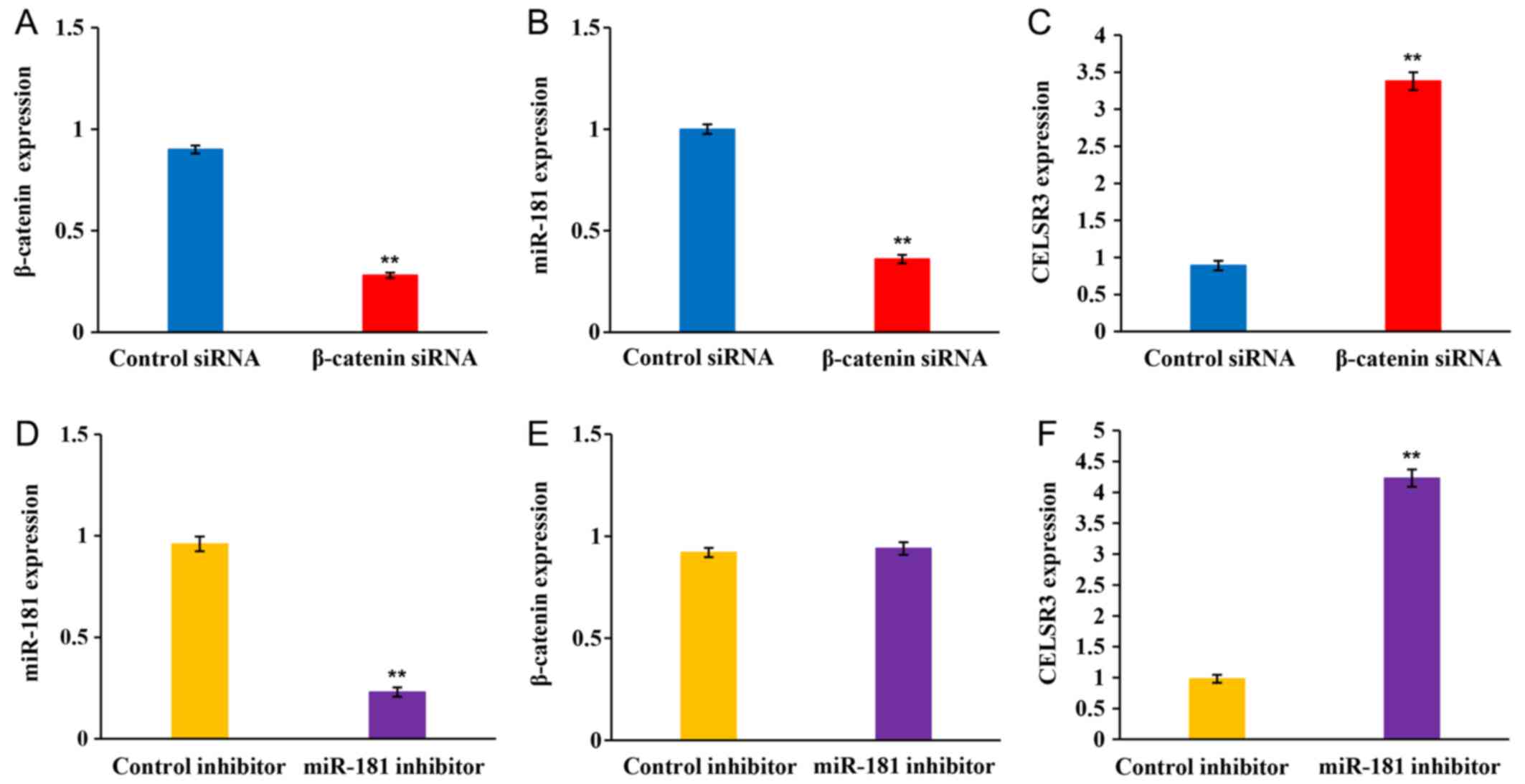
Ji J, Yamashita T and Wang XW:
Wnt/beta-catenin signaling activates microRNA-181 expression in
hepatocellular carcinoma. Cell Biosci. 1:42011. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Lu Y, Wang L, Liu P, Yang P and You M:
Gene-expression signature predicts postoperative recurrence in
stage I non-small cell lung cancer patients. PLoS One.
7:e308802012. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Zhu CQ, Ding K, Strumpf D, Weir BA,
Meyerson M, Pennell N, Thomas RK, Naoki K, Ladd-Acosta C, Liu N, et
al: Prognostic and predictive gene signature for adjuvant
chemotherapy in resected non-small-cell lung cancer. J Clin Oncol.
28:4417–4424. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Subramanian J and Simon R: Gene
expression-based prognostic signatures in lung cancer: Ready for
clinical use? J Natl Cancer Inst. 102:464–474. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Feng J, Xu Y, Wang M, Ruan Y, So KF,
Tissir F, Goffinet A and Zhou L: A role for atypical cadherin
Celsr3 in hippocampal maturation and connectivity. J Neurosci.
32:13729–13743. 2012. View Article : Google Scholar : PubMed/NCBI
|