|
1
|
Siegel RL, Miller KD and Jemal A: Cancer
statistics, 2017. CA Cancer J Clin. 67:7–30. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Siegel RL, Miller KD and Jemal A: Cancer
statistics, 2016. CA Cancer J Clin. 66:7–30. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Chen W, Zheng R, Baade PD, Zhang S, Zeng
H, Bray F, Jemal A, Yu XQ and He J: Cancer statistics in China,
2015. CA Cancer J Clin. 66:115–132. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Antonarakis ES, Lu C, Wang H, Luber B,
Nakazawa M, Roeser JC, Chen Y, Mohammad TA, Chen Y, Fedor HL, et
al: AR-V7 and resistance to enzalutamide and abiraterone in
prostate cancer. N Engl J Med. 371:1028–1038. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Sahu B, Laakso M, Ovaska K, Mirtti T,
Lundin J, Rannikko A, Sankila A, Turunen JP, Lundin M, Konsti J, et
al: Dual role of FoxA1 in androgen receptor binding to chromatin,
androgen signalling and prostate cancer. EMBO J. 30:3962–3976.
2011. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Barbieri CE, Baca SC, Lawrence MS,
Demichelis F, Blattner M, Theurillat JP, White TA, Stojanov P, Van
Allen E, Stransky N, et al: Exome sequencing identifies recurrent
SPOP, FOXA1 and MED12 mutations in prostate cancer. Nat Genet.
44:685–689. 2012. View
Article : Google Scholar : PubMed/NCBI
|
|
7
|
Geng C, Rajapakshe K, Shah SS, Shou J,
Eedunuri VK, Foley C, Fiskus W, Rajendran M, Chew SA, Zimmermann M,
et al: Androgen receptor is the key transcriptional mediator of the
tumor suppressor SPOP in prostate cancer. Cancer Res. 74:5631–5643.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Kouzarides T: Histone methylation in
transcriptional control. Curr Opin Genet Dev. 12:198–209. 2002.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Berger SL: Histone modifications in
transcriptional regulation. Curr Opin Genet Dev. 12:142–148. 2002.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Kulis M, Queirós AC, Beekman R and
Martín-Subero JI: Intragenic DNA methylation in transcriptional
regulation, normal differentiation and cancer. Biochim Biophys
Acta. 1829:1161–1174. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Sharif J, Muto M, Takebayashi S, Suetake
I, Iwamatsu A, Endo TA, Shinga J, Mizutani-Koseki Y, Toyoda T,
Okamura K, et al: The SRA protein Np95 mediates epigenetic
inheritance by recruiting Dnmt1 to methylated DNA. Nature.
450:908–912. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Fukuda A and Hisatake K: Histone
methylation and transcriptional regulation. Seikagaku. 79:362–365.
2007.(In Japanese). PubMed/NCBI
|
|
13
|
An W: Histone acetylation and methylation:
Combinatorial players for transcriptional regulation. Subcell
Biochem. 41:351–369. 2007.PubMed/NCBI
|
|
14
|
Schlesinger Y, Straussman R, Keshet I,
Farkash S, Hecht M, Zimmerman J, Eden E, Yakhini Z, Ben-Shushan E,
Reubinoff BE, et al: Polycomb-mediated methylation on Lys27 of
histone H3 pre-marks genes for de novo methylation in cancer. Nat
Genet. 39:232–236. 2007. View
Article : Google Scholar : PubMed/NCBI
|
|
15
|
Kulis M and Esteller M: DNA methylation
and cancer. Adv Genet. 70:27–56. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
de Sousa E Melo F, Colak S, Buikhuisen J,
Koster J, Cameron K, de Jong JH, Tuynman JB, Prasetyanti PR,
Fessler E, van den Bergh SP, et al: Methylation of
cancer-stem-cell-associated Wnt target genes predicts poor
prognosis in colorectal cancer patients. Cell Stem Cell. 9:476–485.
2011. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Parrella P, Poeta ML, Gallo AP, Prencipe
M, Scintu M, Apicella A, Rossiello R, Liguoro G, Seripa D, Gravina
C, et al: Nonrandom distribution of aberrant promoter methylation
of cancer-related genes in sporadic breast tumors. Clin Cancer Res.
10:5349–5354. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Hayami S, Yoshimatsu M, Veerakumarasivam
A, Unoki M, Iwai Y, Tsunoda T, Field HI, Kelly JD, Neal DE, Yamaue
H, et al: Overexpression of the JmjC histone demethylase KDM5B in
human carcinogenesis: Involvement in the proliferation of cancer
cells through the E2F/RB pathway. Mol Cancer. 9:592010. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Barrett A, Santangelo S, Tan K, Catchpole
S, Roberts K, Spencer-Dene B, Hall D, Scibetta A, Burchell J,
Verdin E, et al: Breast cancer associated transcriptional repressor
PLU-1/JARID1B interacts directly with histone deacetylases. Int J
Cancer. 121:265–275. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Wang D, Han S, Peng R, Jiao C, Wang X,
Yang X, Yang R and Li X: Depletion of histone demethylase KDM5B
inhibits cell proliferation of hepatocellular carcinoma by
regulation of cell cycle checkpoint proteins p15 and p27. J Exp
Clin Cancer Res. 35:372016. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Wang Z, Tang F, Qi G, Yuan S, Zhang G,
Tang B and He S: KDM5B is overexpressed in gastric cancer and is
required for gastric cancer cell proliferation and metastasis. Am J
Cancer Res. 5:87–100. 2015.PubMed/NCBI
|
|
22
|
Dai B, Hu Z, Huang H, Zhu G, Xiao Z, Wan
W, Zhang P, Jia W and Zhang L: Overexpressed KDM5B is associated
with the progression of glioma and promotes glioma cell growth via
downregulating p21. Biochem Biophys Res Commun. 454:221–227. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Bamodu OA, Huang WC, Lee WH, Wu A, Wang
LS, Hsiao M, Yeh CT and Chao TY: Aberrant KDM5B expression promotes
aggressive breast cancer through MALAT1 overexpression and
downregulation of hsa-miR-448. BMC Cancer. 16:1602016. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Catchpole S, Spencer-Dene B, Hall D,
Santangelo S, Rosewell I, Guenatri M, Beatson R, Scibetta AG,
Burchell JM and Taylor-Papadimitriou J: PLU-1/JARID1B/KDM5B is
required for embryonic survival and contributes to cell
proliferation in the mammary gland and in ER+ breast cancer cells.
Int J Oncol. 38:1267–1277. 2011.PubMed/NCBI
|
|
25
|
Xiang Y, Zhu Z, Han G, Ye X, Xu B, Peng Z,
Ma Y, Yu Y, Lin H, Chen AP and Chen CD: JARID1B is a histone H3
lysine 4 demethylase up-regulated in prostate cancer. Proc Natl
Acad Sci USA. 104:19226–19231. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
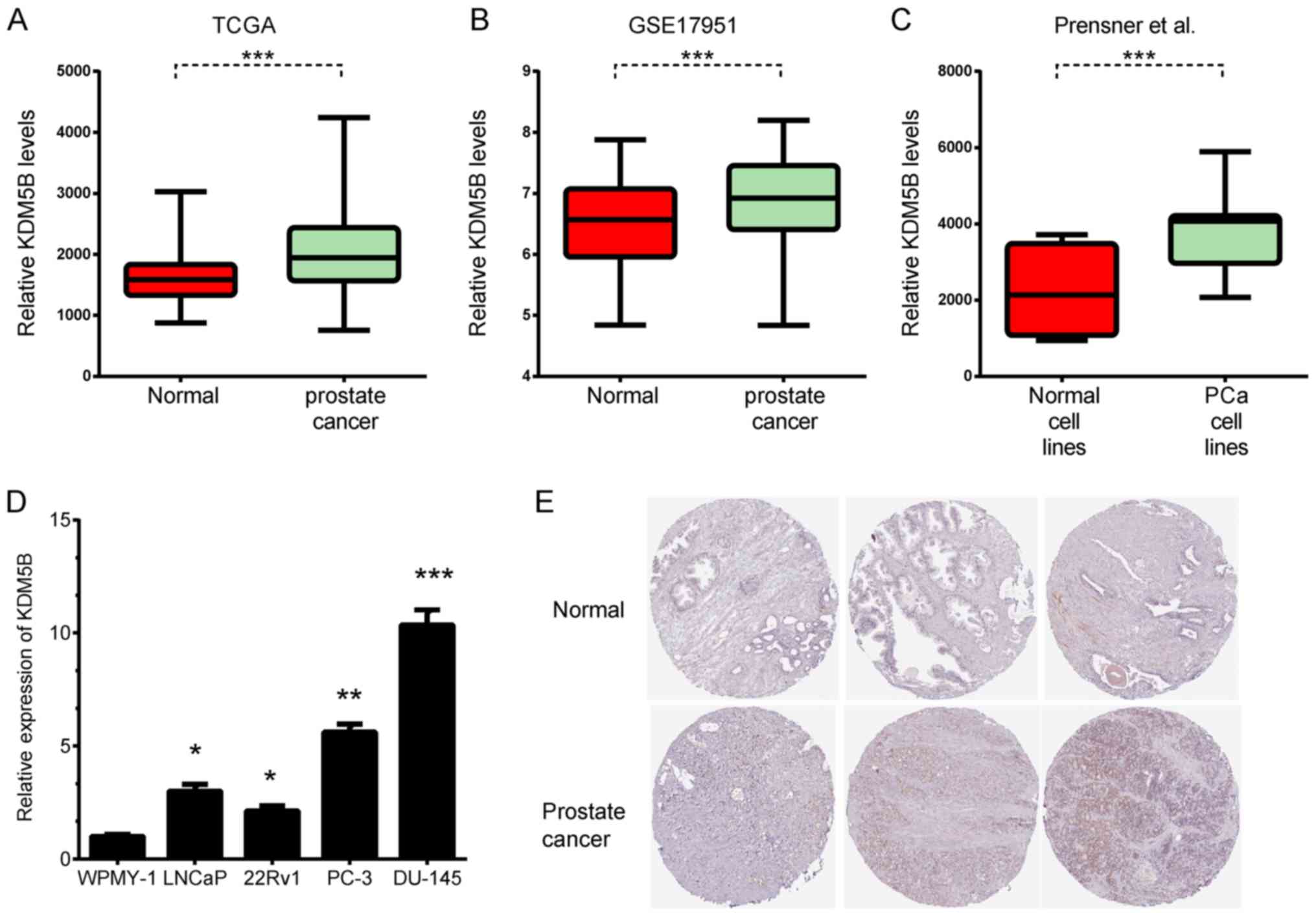
Wang Y, Xia XQ, Jia Z, Sawyers A, Yao H,
Wang-Rodriquez J, Mercola D and McClelland M: In silico estimates
of tissue components in surgical samples based on expression
profiling data. Cancer Res. 70:6448–6455. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Prensner JR, Iyer MK, Balbin OA,
Dhanasekaran SM, Cao Q, Brenner JC, Laxman B, Asangani IA, Grasso
CS, Kominsky HD, et al: Transcriptome sequencing across a prostate
cancer cohort identifies PCAT-1, an unannotated lincRNA implicated
in disease progression. Nat Biotechnol. 29:742–749. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Budczies J, Klauschen F, Sinn BV, Győrffy
B, Schmitt WD, Darb-Esfahani S and Denkert C: Cutoff Finder: A
comprehensive and straightforward Web application enabling rapid
biomarker cutoff optimization. PLoS One. 7:e518622012. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Taylor BS, Schultz N, Hieronymus H,
Gopalan A, Xiao Y, Carver BS, Arora VK, Kaushik P, Cerami E Reva B,
et al: Integrative genomic profiling of human prostate cancer.
Cancer Cell. 18:11–22. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Ferro M, Lucarelli G, Bruzzese D, Di
Lorenzo G, Perdonà S, Autorino R, Cantiello F, La Rocca R, Busetto
GM, Cimmino A, et al: Low serum total testosterone level as a
predictor of upstaging and upgrading in low-risk prostate cancer
patients meeting the inclusion criteria for active surveillance.
Oncotarget. 8:18424–18434. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
de Cobelli O, Terracciano D, Tagliabue E,
Raimondi S, Galasso G, Cioffi A, Cordima G, Musi G, Damiano R,
Cantiello F, et al: Body mass index was associated with upstaging
and upgrading in patients with low-risk prostate cancer who met the
inclusion criteria for active surveillance. Urol Oncol.
33:201.e1–e8. 2015. View Article : Google Scholar
|
|
33
|
de Cobelli O, Buonerba C, Terracciano D,
Bottero D, Lucarelli G, Bove P, Altieri V, Coman I, Perdonà S,
Facchini G, et al: Urotensin II receptor on preoperative biopsy is
associated with upstaging and upgrading in prostate cancer. Future
Oncol. 11:3091–3098. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Sreekumar A, Poisson LM, Rajendiran TM,
Khan AP, Cao Q, Yu J, Laxman B, Mehra R, Lonigro RJ, Li Y, et al:
Metabolomic profiles delineate potential role for sarcosine in
prostate cancer progression. Nature. 457:910–914. 2009. View Article : Google Scholar : PubMed/NCBI
|