Introduction
Liver cancer can be divided into two major
categories: primary and secondary liver cancer. Primary liver
cancer originates from the epithelial or mesenchymal tissue of the
liver, it is a malignant tumor with high incidence and great harm
in China (1,2). Hepatocellular carcinoma (HCC) is a type
of primary liver cancer, and it is one of the most common
malignancies worldwide (3). Most
patients with HCC are diagnosed with advanced disease, resulting in
low survival and poor prognosis due to lack of effective treatment
(4,5). Therefore, exploring the pathogenesis of
HCC and discovery of an effective treatment strategy is very
urgent.
Mounting evidence displays that the dysregulation of
miRNAs is involved in a variety of cancers and modulate tumor
development via targeting their mRNA, including HCC (6,7). For
example, miR-451a was shown to have inhibitory effect on HCC
tumorigenesis through targeting LPIN1 (8). Wu et al (9) reported that miR-3650 inhibited HCC
metastasis via targeting NFASC. However, miR-1307 enhanced HCC
metastasis and tumor growth by inhibiting DAB2 (10). Upregulation of miR-494 promoted the
development of HCC by targeting SIRT3 through TGF-β/SMAD signaling
pathway (11). Importantly, previous
studies have shown that miR-642a was downregulated in HCC and
associated with clinicopathological information (12). However, the role and precise
molecular mechanisms of miR-642a in HCC cell migration and invasion
has not been reported yet.
SEMA4C, a member of the semaphorin family, is
upregulated in various cancers and take part in tumor growth,
metastasis, and apoptosis through several signal transduction
pathways, which is closely related to the progression and
development of cancers. For example, Gurrapu et al (13) found that SEMA4C was overexpressed in
breast cancer and served as an oncogene in modulating cell growth.
Moreover, SEMA4C was significantly increased in tumor-associated
lymphatic endothelial cells and showed facilitating effects on
lymphatic metastasis (14).
Furthermore, SEMA4C promoted malignant glioma cell invasion and was
correlated with glioma poor survival (15). In addition, SEMA4C was proven to be
upregulated in HCC and regulated HCC cell invasion and migration
(16). However, whether SEMA4C
affected the role of miR-642a in HCC metastasis is not very
clear.
p38 MAPK signaling pathway is involved in cell
growth, differentiation, inflammatory response and other important
cellular physiological/pathological processes. Previous studies
have provided evidence that SEMA4C might be an activator for p38
MAPK pathway (17). Here, we
investigated whether p38 MAPK pathway participated in HCC
development regulated by miR-642a/SEMA4C axis.
In the present study, the role and potential
mechanism of miR-642a in HCC metastasis were investigated. The
findings show that miR-642a was decreased in HCC and re-expression
of miR-642a curbed HCC cell invasion and migration. On the
contrary, SEMA4C showed a facilitating effect on HCC metastasis.
SEMA4C was involved in HCC progression modulated by miR-642a, which
provides a potential target for treatment of HCC patients.
Patients and methods
HCC specimens
Sixty pairs of HCC and paracancerous tissues were
obtained from patients who underwent hepatectomy at The Second
People's Hospital of Lianyungang (Lianyungang, China) from August
2013 to September 2018. The patients received no radiotherapy or
chemotherapy before surgery. The study was approved by the Ethics
Committee of The Second People's Hospital of Lianyungang, and an
informed consent was signed by each patient. The fresh HCC tissues
were immediately frozen at −80°C for RT-PCR analysis.
Cell lines and cell culture
HCC cells (Huh7, HCCLM3) and normal liver epithelial
cells THLE-3 were cultivated in RPMI-1640 medium containing 20% FBS
and antibiotics at 37°C under the atmosphere of 5% CO2.
The cells were obtained from Shanghai Institute of Chinese Academy
of Sciences (Shanghai, China).
Cell transfection
Huh7 cells were added to 24-well plates containing
RPMI-1640 medium. The transfection was performed for 48 h using
Lipofectamine 3000 reagent (Invitrogen; Thermo Fisher Scientific,
Inc.) according to the manufacturer's instructions. The miR-642a
mimic, inhibitor and negative control used in this study were
purchased from GenePharma. The miR-642a mimic or inhibitor was
transfected into Huh7 cells to overexpress or silence miR-642a.
SEMA4C siRNA and con siRNA were synthesized by GenePharma, and they
were used to downregulate SEMA4C or utilize as a control. The
miR-642a mimics sequence was: 5′-AGGACAGGGGAGGATTGCAACG-3′. The
sequences of miR-642a inhibitor and scramble oligonucleotides were
as follows: 5′-CACAGACGGAGGCCAGGGGAGA-3′; and
5′-CCGAAACCUCGGUUGAUUGCGG-3′. Con-siRNA,
5′-UUCUCCGAACGUGUCACGUTT-3′; SEMA4C-siRNA,
5′-CCUAUGCCUUCCAGCCCAAdTdT-3′.
RNA extraction and RT-PCR
TRIzol reagent (Invitrogen; Thermo Fisher
Scientific, Inc.) was used to extract total RNA from HCC cells and
tissues. The mRNA expression was quantified by Platinum™ Taq DNA
polymerase. The sequences of the primers were as follows:
miR-642a-F: 5′-ATACAAAGCCTAAGATGAG-3′, miR-642a-R:
5′-GAGCAAGCTCCTATTCC-3′; SEMA4C-F: 5′-ACCTTGTGCCGCGTAAGACAG-3′,
SEMA4C-R: 5′-CGTCAGCGTCAGTGTCAGGAA-3′; U6-F:
5′-CTCGCTTCGGCAGCACATATACT-3′, U6-R: 5′-ACGCTTCACGAATTTGCGTGTC-3′;
GAPDH-F: 5′-GATCATTGCTCCTCCTGAGC-3′, GAPDH-R:
5′-ACTCCTGCTTGCTGATCCAC-3′. U6 and GAPDH were used as internal
controls. Relative expression of miR-642a and SEMA4C was measured
by the 2−ΔΔCT method.
Western blot analysis
Lysis buffer was used to extract proteins from HCC
tissues or cells. After centrifugation at 12,000 × g, 4°C for 30
min, the supernatant of tissues or cells was measured by BCA kit.
Protein specimens (50 µg) were added onto SDS-PAGE and
electrophoresed at 60 V. Proteins were then transferred to
nitrocellulose filter membranes (NC). Subsequently, the membranes
were blocked with skim milk (5-10%) at 37°C for 1 h and then
incubated with the primary antibodies against SEMA4C (1:500;
sc-136445; Santa Cruz Biotechnology, Inc.), p38 (1:1000, ab170099;
Abcam), p-p38 (1:1000, ab47363; Abcam), MAPK (1:1000, ab185145;
Abcam), p-MAPK (1:2000, 4370; Cell Signaling Technology, Inc.,) at
4°C overnight. Then, the membranes were incubated with horseradish
peroxidase-labeled secondary antibody (1:5000; Santa Cruz
Biotechnology) for 1 h at 37°C. GAPDH (1:2000; ab181602; Abcam) was
used as the loading control. Finally, enhanced chemiluminescence
kit (ECL; EMD Millipore) was used to detect the signals.
Transwell assay
Transwell assays were used to measure cell migration
and invasion. For migration assay, the top and the lower chambers
were separated by Transwell chamber with 8-µm pore size
polycarbonic membrane (Costar) in 6-well plates. Huh7 cells
(5×105/well) with different transfection were added into
the top chambers and the lower chambers were fixed with DMEM
containing 10% FBS. Then, they were cultured for 24 h at 37°C. When
the cells migrated into the lower chambers, the cells were fixed
using 100% methanol, stained with 0.1% crystal violet, photographed
with an inverted microscope (Nikon 80i; Olympus) and counted with
image software. For invasion assay, except the top chamber with the
filter coated with Matrigel, it was the same as for the cell
migration assay.
Luciferase reporter assay
The pEZX-MT06 vector containing the full length of
SEMA4C 3′-UTR was purchased from GeneCopoeia Inc. Lipofectamine
2000 (11668027; Invitrogen; Thermo Fisher Scientific, Inc.) and
used to perform transfection with miR-642a mimic and luciferase
reporter vector containing wild-type (WT) or mutated (MuT) 3′UTR of
SEMA4C into Huh7 cells. The Dual Luciferase Reporter system (E1910;
Promega) was carried out to measure the luciferase activity of Huh7
cells treated with different transfection.
Statistical analysis
All the independent experiments were repeated at
least three times and results are shown as mean ± SD. Statistical
analysis and graph presentations were performed, respectively, by
SPSS v.19.0 software (SPSS, Inc.) and GraphPad Prism 6 software
(GraphPad Inc.). The differences between groups were compared using
Student's t-test or one-way analysis of variance (ANOVA) followed
by Tukey's post hoc test. Statistically significant difference was
considered as P<0.05.
Results
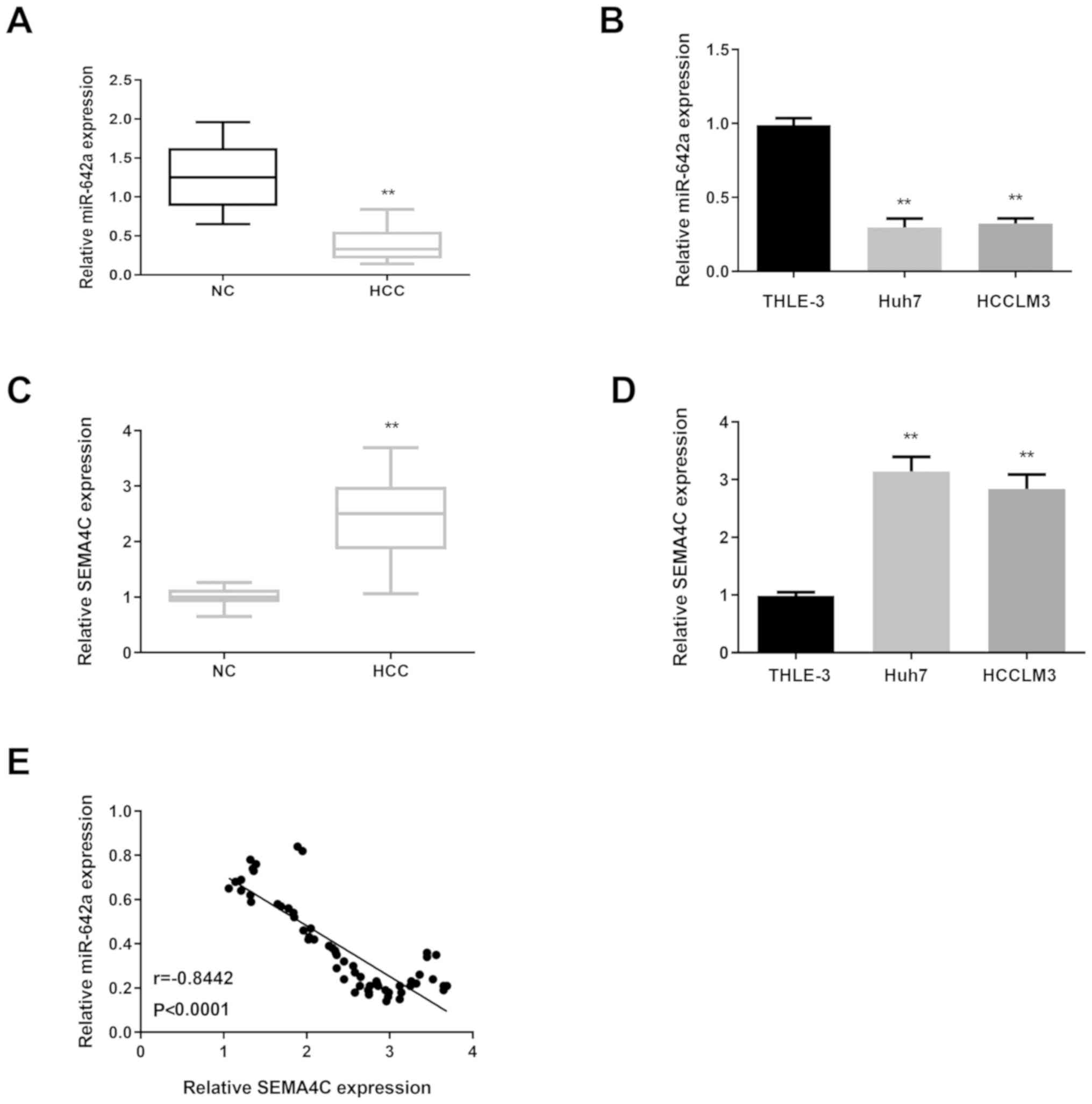
miR-642a expression is decreased and
SEMA4C is increased in HCC
To investigate the biological functions of miR-642a
and SEMA4C on HCC development, we first tested the expressional
level of miR-642a and SEMA4C in HCC tissues and cells. RT-PCR
results displayed that miR-642 expression was reduced in HCC
tissues and cells compared with normal controls (Fig. 1A and B). However, SEMA4C showed the
opposite expression in HCC tissues and cells (Fig. 1C and D). Then, the relationship
between miR-642a and SEMA4C was measured in HCC tissues and the
results revealed that they were negatively correlated (Fig. 1E).
SEMA4C serves as a target of
miR-642a
Based on the above results, we needed to clarify
whether SEMA4C acted as a candidate target of miR-642a in HCC. For
verifying this hypothesis, PITA database was first applied to
determine the binding sites of SEMA4C with miR-642a. As presented
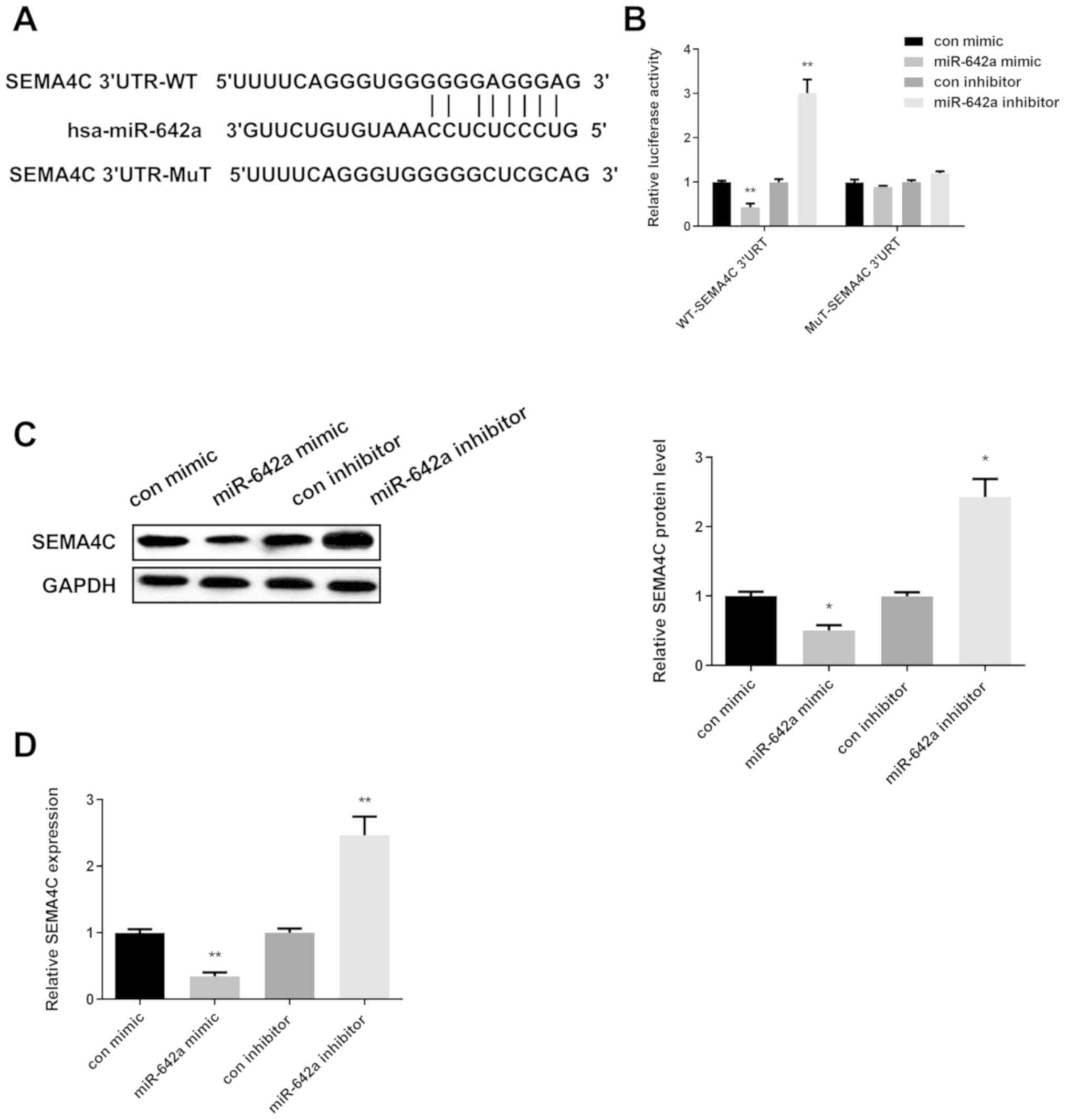
in Fig. 2A, they have the binding
sites. Then, dual-luciferase reporter assay was applied to further
confirm whether SEMA4C was the direct target of miR-642a in HCC
cells. The findings revealed that the luciferase activity in Huh7
cells co-transfected with miR-642a mimic and SEMA4C-3′-UTR-WT was
significantly decreased, while increased in Huh7 cells
co-transfected with miR-642a inhibitor and SEMA4C-3′-UTR-WT.
However, there was no significant difference in SEMA4C-3′-UTR-MUT
(Fig. 2B). Next, we measured whether
miR-642a regulated SEMA4C expression in Huh7 cells. Results of
Western blot and RT-PCR analyses showed that re-expression of
miR-642 was reduced, while silence of miR-642 elevated SEMA4C
expression (Fig. 2C and D). The
conclusion drawn from the above results suggested that miR-642a
regulated SEMA4C expression by binding to its 3′UTR in HCC.
miR-642a represses HCC malignancy
The biological functions of miR-642a on HCC
malignancy was measured. The miR-642a mimic or inhibitor was
transfected into Huh7 cells to measure the effect of miR-642a on
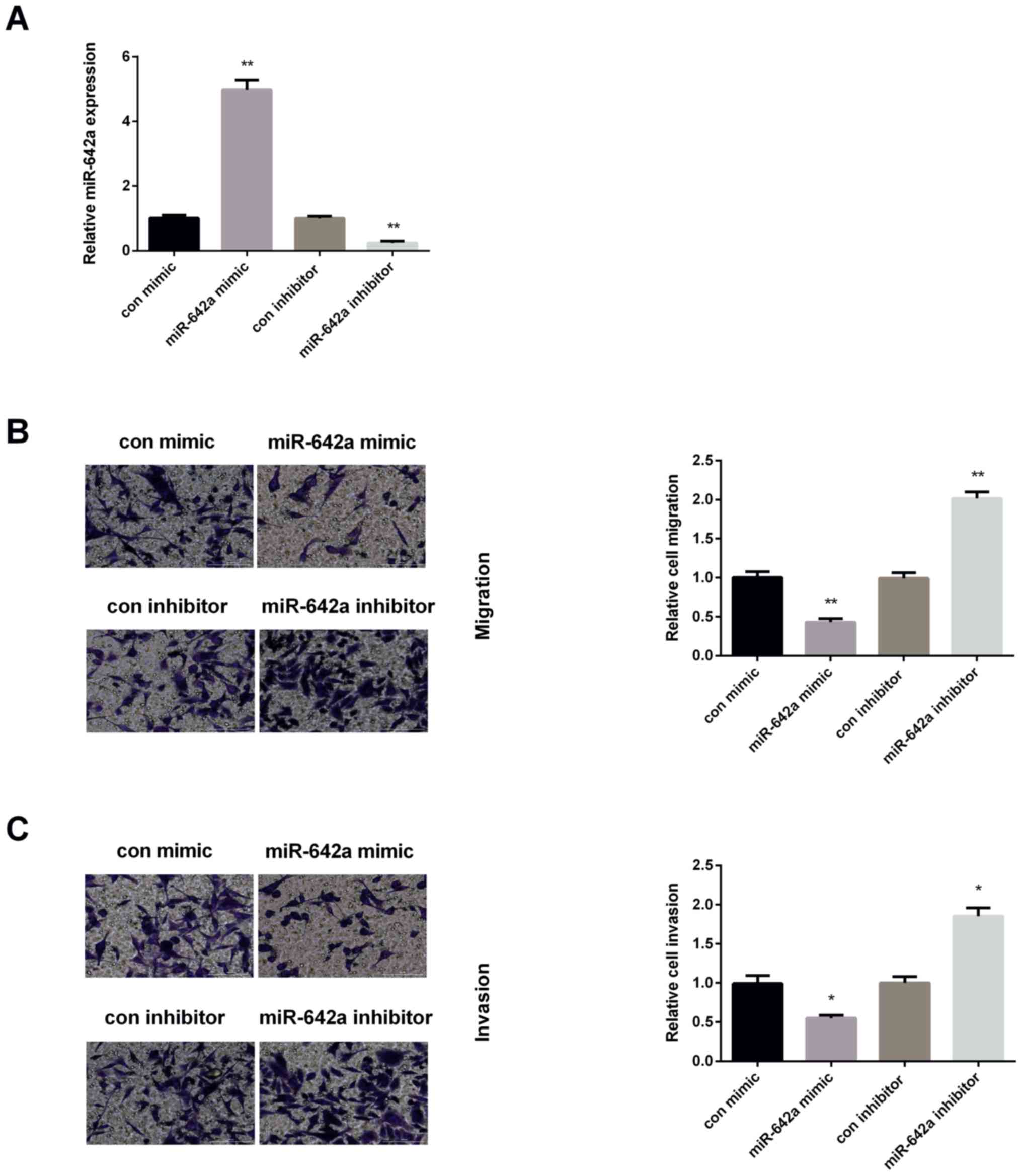
HCC cell migration and invasion. As shown in Fig. 3A, the transfection efficiency of
miR-642a was successful. The expression of miR-642a was increased
significantly in Huh7 cells after overexpression of miR-642a, while
decreased after knockdown of miR-642a. Then, Transwell assays were
applied to examine HCC cell migration and invasion. As presented in
Fig. 3B, the number of migrated
cells declined significantly in miR-642a mimic group, whereas were
elevated in miR-642a inhibitor group. Moreover, the invasive cells
of Huh7 were decreased after re-expression of miR-642a, while
increased after inhibiting miR-642a (Fig. 3C).
Silencing SEMA4C suppresses HCC
malignancy
Next, the effects of SEMA4C on HCC cell migration
and invasion were detected. Due to the higher expression of SEMA4C
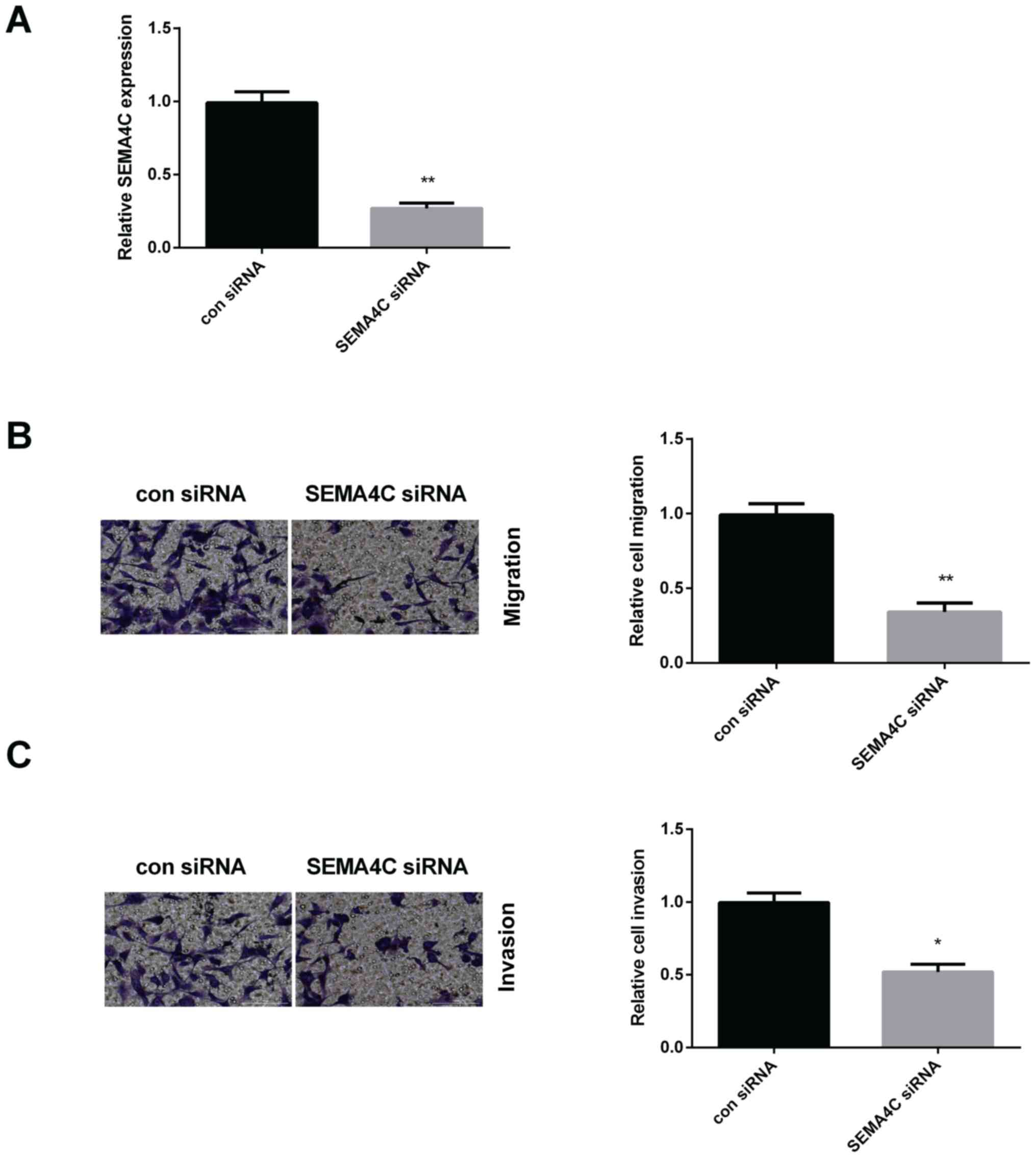
in HCC, it was decreased by SEMA4C siRNA. As expected, SEMA4C
expression was declined remarkably after silencing SEMA4C in Huh7
cells (Fig. 4A). Transwell assay was
then applied to measure the migration and invasion of Huh7 cells
affected by SEMA4C siRNA. As presented in Fig. 4B, SEMA4C siRNA suppressed Huh7 cell
migration. Moreover, silencing SEMA4C repressed Huh7 cell invasion
(Fig. 4C).
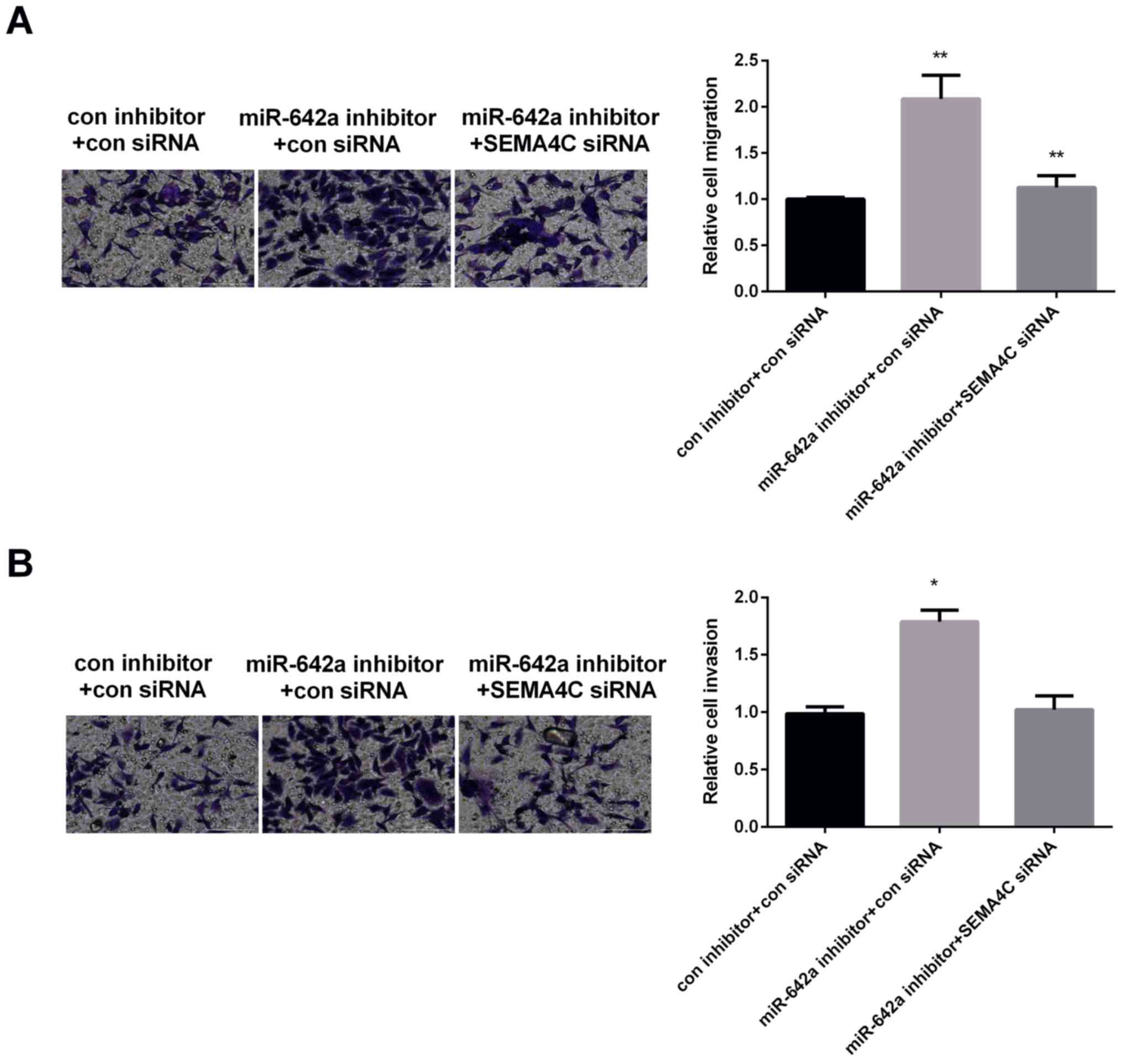
The effect of SEMA4C on miR-642 in the
modulation of HCC malignancy
The effect of SEMA4C on miR-642a in regulating HCC
malignancy was explored due to their opposite effect on HCC
invasion and migration. Huh7 cells were treated with different
transfections: con inhibitor+con siRNA, miR-642a inhibitor+con
siRNA, miR-642a inhibitor+SEMA4C siRNA. Transwell migration assay
results displayed that the migration of Huh7 cells was increased
obviously by silencing miR-642a, while decreased by silencing
miR-642 and SEMA4C, suggesting that SEMA4C could rescue miR-642a
suppression effect on cell migration (Fig. 5A). Moreover, the results of Transwell
invasion assay demonstrated that downregulation of miR-642a
increased HCC cell invasion, whereas was reduced by downregulation
of miR-642a and SEMA4C, indicating that SEMA4C could overturn the
inhibition effect of miR-642a on HCC cell invasion (Fig. 5B). Correctively, miR-642a curbed the
invasion and migration of HCC cells by modulating SEMA4C.
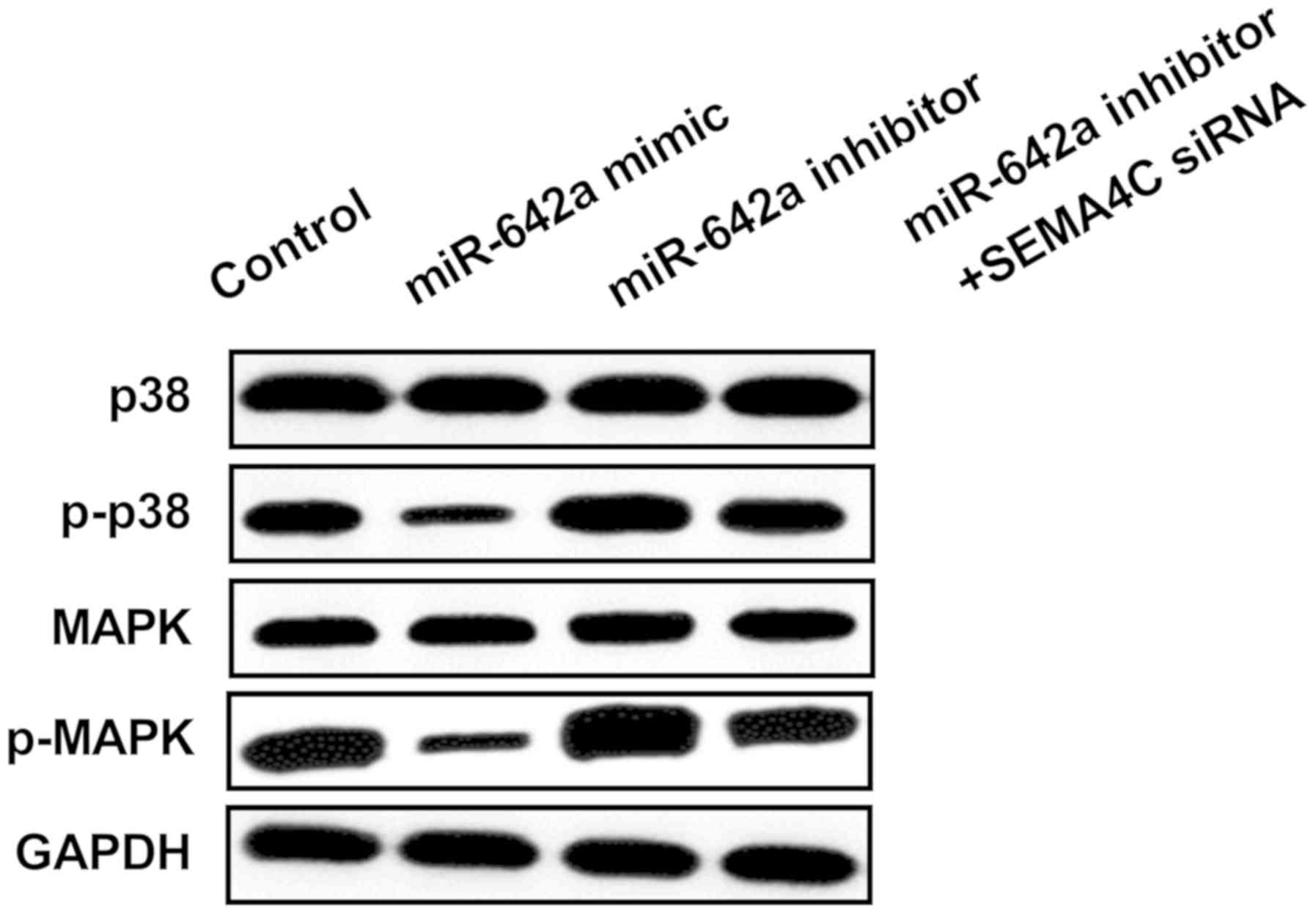
p38 MAPK signaling pathway is involved
in HCC development regulated by miR-642a/SEMA4C axis
Finally, we investigated whether p38 MAPK signaling
pathway was modulated by miR-642a/SEMA4C axis to explore the
precise molecular mechanism of miR-642a in the development of HCC.
Results of Western blot analysis displayed that the downstream
genes of p38 MAPK pathway were inhibited by miR-642a upregulation,
while enhanced by miR-642a inhibitor. Moreover, silenced SEMA4C was
able to repress the activation of phosphorylation of p38 and MAPK
induced by miR-642a inhibitor (Fig.
6). These results demonstrated that miR-642a impeded the
activation of p38 MAPK signaling pathway by suppressing SEMA4C in
HCC.
Discussion
Our findings displayed that miR-642a expression was
declined while SEMA4C was raised in HCC tissues and cells.
Moreover, miR-642a showed inhibitory effect on HCC malignancy and
SEMA4C showed the opposite effect. Importantly, SEMA4C was proved
to be the target of miR-642a in regulation of HCC development.
Moreover, SEMA4C could rescue the inhibitory effect of miR-642a on
HCC malignancy. In addition, p38 MAPK signaling pathway was
modulated by miR-642a/SEMA4C axis. Taken together, the research
demonstrated that miR-642a inhibited HCC development by repressing
SEMA4C through p38 MAPK signaling pathway, providing a potential
target for treatment of HCC patients.
Increasing evidence has been reported that miRNAs
modulated various molecular pathways in tumor development via
targeting their specific mRNAs as either tumor suppressors or
oncogenes (18–20). Previous studies stated that miRNA
expression profiles varied from cancer to cancer. For example,
miR-642a was involved in pancreatic neuroendocrine neoplasms and
correlated with Ki67 score (21).
Moreover, Nordentoft et al (22) revealed that increasing miR-642
generally increased cisplatin sensitivity of bladder cancer and
might form a novel target for treatment of patients. However, Epis
et al (23) showed an
under-expression of miR-642 in prostate cancer and overexpression
of miR-642 suppressed cell proliferation. In the present study, it
was found that miR-642a was under-expressed in HCC tissues and
cells which is consistent with the report that miR-642a expression
was downregulated in HCC and took part in HCC proliferation and
metastasis regulated by Linc00974 and KRT19 (12). In this study, we also found that
re-expression of miR-642a repressed HCC invasion and migration.
SEMA4C was reported highly expressed in numerous
cancer tissues, such as in gastric cancer, esophageal cancer and
rectal cancer (24). SEMA4C usually
take part in tumor development as a target of miRNA. For instance,
it was a target of miR-25 in regulating the epithelial-mesenchymal
transition of cervical cancer (25).
Also, SEMA4C was regulated by miR-125b in regulating
epithelial-mesenchymal transition of lung cancer (26). Li et al (27) stated that miR-138 suppressed
non-small cell lung cancer cell proliferation through targeting
GIT1 and SEMA4C. In our study, the SEMA4C expression was raised in
HCC and silencing SEMA4C was able to curb HCC malignancy. These
results are in line with a previous study that SEMA4C was
overexpressed in HCC and involved in HCC tumor growth and invasion
(16). Furthermore, we first found
that SEMA4C was a specific target of miR-642a in the regulation of
HCC malignancy and SEMA4C could rescue miR-642a inhibitory effect
on HCC malignancy.
In the tumor development and metastasis, p38 MAPK
signaling pathway is activated as a frequent event (28,29). p38
MAPK pathway was dysregulated in HCC tissues compared with the
corresponding normal tissues (30).
Increasing number of studies have demonstrated that miRNAs
interacted with signaling pathways in a variety of cancers. In the
present study, we found that p38 MAPK signaling pathway was
inhibited by miR-642a mimic, whereas activated by miR-642a
inhibitor. Furthermore, SEMA4C siRNA attenuated the activation of
p38 MAPK signaling pathway induced by miR-642a inhibitor.
There are limitations in the present study. The role
of the miR-642a/SEMA4C axis should be verified in vivo. The
effect of SEMA4C overexpression or knockdown on p38 MAPK signaling
pathway remain to be done, which will guide the investigation into
the inmost mechanism. The association of miR-642a with clinical
characteristics of patients should be done in future studies.
In conclusion, our findings revealed that miR-642a
suppressed, while SEMA4C promoted HCC malignancy. This is the first
time we state that miR-642a directly targeted SEMA4C to modulate
HCC development and SEMA4C could overturn miR-642a inhibitory
effect on HCC. Taken together, by targeting SEMA4C, miR-642a
impeded HCC malignancy through suppressing p38 MAPK signaling
pathway.
Acknowledgements
Not applicable.
Funding
The study was supported by the Fujian Natural
Science Foundation (Fujian, China) (grant no. 2019J01188).
Availability of data and materials
The datasets used and analyzed during the present
study are available from the corresponding author on reasonable
request.
Authors' contributions
QK contributed to the conception and design of the
study. HL and JH analyzed and interpreted the patient general data.
DJ, JF, YS, LZ, XY and NX performed RT-PCR, Transwell assay,
western blot analysis and luciferase reporter assay. ZY and YD were
also involved in the conception of the study. QK gave final
approval of the version to be published. All authors read and
approved the final version of the manuscript.
Ethics approval and consent to
participate
The study was approved by the Ethics Committee of
The Second People's Hospital of Lianyungang (Lianyungang, China)
and an informed consent was signed by each patient.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Siegel RL, Miller KD and Jemal A: Cancer
Statistics, 2017. CA Cancer J Clin. 67:7–30. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Fornari F, Ferracin M, Trerè D, Milazzo M,
Marinelli S, Galassi M, Venerandi L, Pollutri D, Patrizi C, Borghi
A, et al: Circulating microRNAs, miR-939, miR-595, miR-519d and
miR-494, identify cirrhotic patients with HCC. PLoS One.
10:e01414482015. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Bray F, Ferlay J, Soerjomataram I, Siegel
RL, Torre LA and Jemal A: Global cancer statistics 2018: GLOBOCAN
estimates of incidence and mortality worldwide for 36 cancers in
185 countries. CA Cancer J Clin. 68:394–424. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Kassahun WT, Fangmann J, Harms J, Hauss J
and Bartels M: Liver resection and transplantation in the
management of hepatocellular carcinoma: A review. Exp Clin
Transplant. 4:549–558. 2006.PubMed/NCBI
|
|
5
|
Dilou N, Patouillard B and Audigier JC:
Staging systems in hepatocellular carcinoma. Gastroenterol Clin
Biol. 28:359–366. 2004.(In French). View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Ono K, Kuwabara Y and Han J: MicroRNAs and
cardiovascular diseases. FEBS J. 278:1619–1633. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Lim LP, Lau NC, Garrett-Engele P, Grimson
A, Schelter JM, Castle J, Bartel DP, Linsley PS and Johnson JM:
Microarray analysis shows that some microRNAs downregulate large
numbers of target mRNAs. Nature. 433:769–773. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Zhao S, Li J, Zhang G, Wang Q, Wu C, Zhang
Q, Wang H, Sun P, Xiang R and Yang S: Exosomal miR-451a functions
as a tumor suppressor in hepatocellular carcinoma by targeting
LPIN1. Cell Physiol Biochem. 53:19–35. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Wu J, Huang WJ, Xi HL, Liu LY, Wang ST,
Fan WZ and Peng BG: Tumor-suppressive miR-3650 inhibits tumor
metastasis by directly targeting NFASC in hepatocellular carcinoma.
Aging (Albany NY). 11:3432–3434. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Chen S, Wang L, Yao B, Liu Q and Guo C:
miR-1307-3p promotes tumor growth and metastasis of hepatocellular
carcinoma by repressing DAB2 interacting protein. Biomed
Pharmacother. 117:1090552019. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Zhang J, Zhu Y, Hu L, Yan F and Chen J:
miR-494 induces EndMT and promotes the development of HCC
(hepatocellular carcinoma) by targeting SIRT3/TGF-β/SMAD signaling
pathway. Sci Rep. 9:72132019. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Tang J, Zhuo H, Zhang X, Jiang R, Ji J,
Deng L, Qian X, Zhang F and Sun B: A novel biomarker Linc00974
interacting with KRT19 promotes proliferation and metastasis in
hepatocellular carcinoma. Cell Death Dis. 5:e15492014. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Gurrapu S, Pupo E, Franzolin G, Lanzetti L
and Tamagnone L: Sema4C/PlexinB2 signaling controls breast cancer
cell growth, hormonal dependence and tumorigenic potential. Cell
Death Differ. 25:1259–1275. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Wei JC, Yang J, Liu D, Wu MF, Qiao L, Wang
JN, Ma QF, Zeng Z, Ye SM, Guo ES, et al: Tumor-associated lymphatic
endothelial cells promote lymphatic metastasis by highly expressing
and secreting SEMA4C. Clin Cancer Res. 23:214–224. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Le AP, Huang Y, Pingle SC, Kesari S, Wang
H, Yong RL, Zou H and Friedel RH: Plexin-B2 promotes invasive
growth of malignant glioma. Oncotarget. 6:7293–7304. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Lu J, Lin Y, Li F, Ye H, Zhou R, Jin Y, Li
B, Xiong X and Cheng N: miR-205 suppresses tumor growth, invasion,
and epithelial-mesenchymal transition by targeting SEMA4C in
hepatocellular carcinoma. FASEB J. 32:fj201800113R2018. View Article : Google Scholar
|
|
17
|
Wu H, Wang X, Liu S, Wu Y, Zhao T, Chen X,
Zhu L, Wu Y, Ding X, Peng X, et al: Sema4C participates in myogenic
differentiation in vivo and in vitro through the p38 MAPK pathway.
Eur J Cell Biol. 86:331–344. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Wang ZF, Liao F, Wu H and Dai J: Glioma
stem cell-derived exosomal miR-26a promotes angiogenesis of
microvessel endothelial cells in glioma. J Exp Clin Cancer Res.
38:2012019. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Zhang J, Hou L, Liang R, Chen X, Zhang R,
Chen W and Zhu J: CircDLST promotes the tumorigenesis and
metastasis of gastric cancer by sponging miR-502-5p and activating
the NRAS/MEK1/ERK1/2 signaling. Mol Cancer. 18:802019. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Ma M, Dai J, Tang H, Xu T, Yu S, Si L, Cui
C, Sheng X, Chi Z, Mao L, et al: MicroRNA-23a-3p inhibits mucosal
melanoma growth and progression through targeting adenylate cyclase
1 and attenuating cAMP and MAPK pathways. Theranostics. 9:945–960.
2019. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Thorns C, Schurmann C, Gebauer N,
Wallaschofski H, Kümpers C, Bernard V, Feller AC, Keck T, Habermann
JK, Begum N, et al: Global microRNA profiling of pancreatic
neuroendocrine neoplasias. Anticancer Res. 34:2249–2254.
2014.PubMed/NCBI
|
|
22
|
Nordentoft I, Birkenkamp-Demtroder K,
Agerbæk M, Theodorescu D, Ostenfeld MS, Hartmann A, Borre M,
Ørntoft TF and Dyrskjøt L: miRNAs associated with chemo-sensitivity
in cell lines and in advanced bladder cancer. BMC Med Genomics.
5:402012. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Epis MR, Giles KM, Kalinowski FC, Barker
A, Cohen RJ and Leedman PJ: Regulation of expression of
deoxyhypusine hydroxylase (DOHH), the enzyme that catalyzes the
activation of eIF5A, by miR-331-3p and miR-642-5p in prostate
cancer cells. J Biol Chem. 287:35251–35259. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Ye SM, Han M, Kan CY, Yang LL, Yang J, Ma
QF and Wang SX: Expression and clinical significance of Sema4C in
esophageal cancer, gastric cancer and rectal cancer. Zhonghua Yi
Xue Za Zhi. 92:1954–1958. 2012.(In Chinese). PubMed/NCBI
|
|
25
|
Song J and Li Y: miR-25-3p reverses
epithelial-mesenchymal transition via targeting Sema4C in
cisplatin-resistance cervical cancer cells. Cancer Sci. 108:23–31.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Zhang Y and Huang S: Up-regulation of
miR-125b reverses epithelial-mesenchymal transition in
paclitaxel-resistant lung cancer cells. Biol Chem. Aug
20–2015.(Epub ahead of print). doi: 10.1515/hsz-2015-0153.
View Article : Google Scholar
|
|
27
|
Li J, Wang Q, Wen R, Liang J, Zhong X,
Yang W, Su D and Tang J: miR-138 inhibits cell proliferation and
reverses epithelial-mesenchymal transition in non-small cell lung
cancer cells by targeting GIT1 and SEMA4C. J Cell Mol Med.
19:2793–2805. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Chen J, Ji T, Wu D, Jiang S, Zhao J, Lin H
and Cai X: Human mesenchymal stem cells promote tumor growth via
MAPK pathway and metastasis by epithelial mesenchymal transition
and integrin α5 in hepatocellular carcinoma. Cell Death Dis.
10:4252019. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Chatterjee S, Patra D, Chakraborti U,
Sengupta D, Ghosh P, Basu A, Sadhukhan GC and Chowdhury KD:
Association of p38 MAPK-p53-Fas aggregation in S-allyl cysteine
mediated regulation of hepatocarcinoma. Environ Toxicol.
34:928–940. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Zhang GP, Yue X and Li SQ: Cathepsin C
interacts with TNF-alpha/p38 MAPK signaling pathway to promote
proliferation and metastasis in hepatocellular carcinoma. Cancer
Res Treat. 52:10–23. 2020. View Article : Google Scholar : PubMed/NCBI
|