|
1
|
Khodavirdipour A, Zarean R and
Safaralizadeh R: Evaluation of the anti-cancer effect of Syzygium
cumini ethanolic extract on HT-29 colorectal cell line. J
Gastrointest Cancer. Jun 6–2020.(Epub ahead of print). View Article : Google Scholar
|
|
2
|
Zoratto F, Rossi L, Verrico M, Papa A,
Basso E, Zullo A, Tomao L, Romiti A, Lo Russo G and Tomao S: Focus
on genetic and epigenetic events of colorectal cancer pathogenesis:
Implications for molecular diagnosis. Tumour Biol. 35:6195–6206.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Palii SS and Robertson KD: Epigenetic
control of tumor suppression. Crit Rev Eukaryot Gene Expr.
17:295–316. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Baylin SB and Ohm JE: Epigenetic gene
silencing in cancer-a mechanism for early oncogenic pathway
addiction? Nat Rev Cancer. 6:107–116. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Krishna SS, Majumdar I and Grishin NV:
Structural classification of zinc fingers: Survey and summary.
Nucleic Acids Res. 31:532–550. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Laity JH, Lee BM and Wright PE: Zinc
finger proteins: New insights into structural and functional
diversity. Curr Opin Struct Biol. 11:39–46. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Yang L, Hamilton SR, Sood A, Kuwai T,
Ellis L, Sanguino A, Lopez-Berestein G and Boyd DD: The previously
undescribed ZKSCAN3 (ZNF306) is a novel ‘driver’ of colorectal
cancer progression. Cancer Res. 68:4321–4330. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Li J, Wang Y, Fan X, Mo X, Wang Z, Li Y,
Yin Z, Deng Y, Luo N, Zhu C, et al: ZNF307, a novel zinc finger
gene suppresses p53 and p21 pathway. Biochem Biophys Res Commun.
363:895–900. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Huang C, Jia Y, Yang S, Chen B, Sun H,
Shen F and Wang Y: Characterization of ZNF23, a KRAB-containing
protein that is downregulated in human cancers and inhibits cell
cycle progression. Exp Cell Res. 313:254–263. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Hu R, Peng G, Dai H, Breuer EK,
Stemke-Hale K, Li K, Gonzalez-Angulo AM, Mills GB and Lin SY:
ZNF668 functions as a tumor suppressor by regulating p53 stability
and function in breast cancer. Cancer Res. 71:6524–6534. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Nagase T, Ishikawa K, Suyama M, Kikuno R,
Hirosawa M, Miyajima N, Tanaka A, Kotani H, Nomura N and Ohara O:
Prediction of the coding sequences of unidentified human genes.
XII. The complete sequences of 100 new cDNA clones from brain which
code for large proteins in vitro. DNA Res. 5:355–364. 1998.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Gianfrancesco F, Esposito T, Ombra MN,
Forabosco P, Maninchedda G, Fattorini M, Casula S, Vaccargiu S,
Casu G, Cardia F, Deiana I, et al: Identification of a novel gene
and a common variant associated with uric acid nephrolithiasis in a
Sardinian genetic isolate. Am J Hum Genet. 72:1479–1491. 2003.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Medina-Escobedo M, González-Herrera L,
Villanueva-Jorge S and Martín-Soberanis G: Metabolic abnormalities
and polymorphisms of the vitamin D receptor (VDR) and ZNF365 genes
in children with urolithiasis. Urolithiasis. 42:395–400. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Haritunians T, Jones MR, McGovern DP, Shih
DQ, Barrett RJ, Derkowski C, Dubinsky MC, Dutridge D, Fleshner PR,
Ippoliti A, et al: Variants in ZNF365 isoform D are associated with
Crohn's disease. Gut. 60:1060–1067. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Cao S, Chee SP, Yu HG, Sukavatcharin S, Wu
L, Kijlstra A, Hou S and Yang P: Investigation of the association
of Vogt-Koyanagi-Harada syndrome with IL23R-C1orf141 in Han Chinese
Singaporean and ADO-ZNF365-EGR2 in Thai. Br J Ophthalmol.
100:436–442. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Lindström S, Vachon CM, Li J, Varghese J,
Thompson D, Warren R, Brown J, Leyland J, Audley T, Wareham NJ, et
al: Common variants in ZNF365 are associated with both mammographic
density and breast cancer risk. Nat Genet. 43:185–187. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Couch FJ, Gaudet MM, Antoniou AC, Ramus
SJ, Kuchenbaecker KB, Soucy P, Beesley J, Chen X, Wang X, Kirchhoff
T, et al: Common variants at the 19p13.1 and ZNF365 loci are
associated with ER subtypes of breast cancer and ovarian cancer
risk in BRCA1 and BRCA2 mutation carriers. Cancer Epidemiol
Biomarkers Prev. 21:645–657. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Kleihues P and Sobin LH: World Health
Organization classification of tumors. Cancer. 88:28872000.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Edge SB and Compton CC: The American Joint
Committee on Cancer: The 7th edition of the AJCC cancer staging
manual and the future of TNM. Ann Surg Oncol. 17:1471–1474. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Tao Q, Huang H, Geiman TM, Lim CY, Fu L,
Qiu GH and Robertson KD: Defective de novo methylation of viral and
cellular DNA sequences in ICF syndrome cells. Hum Mol Genet.
11:2091–2102. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
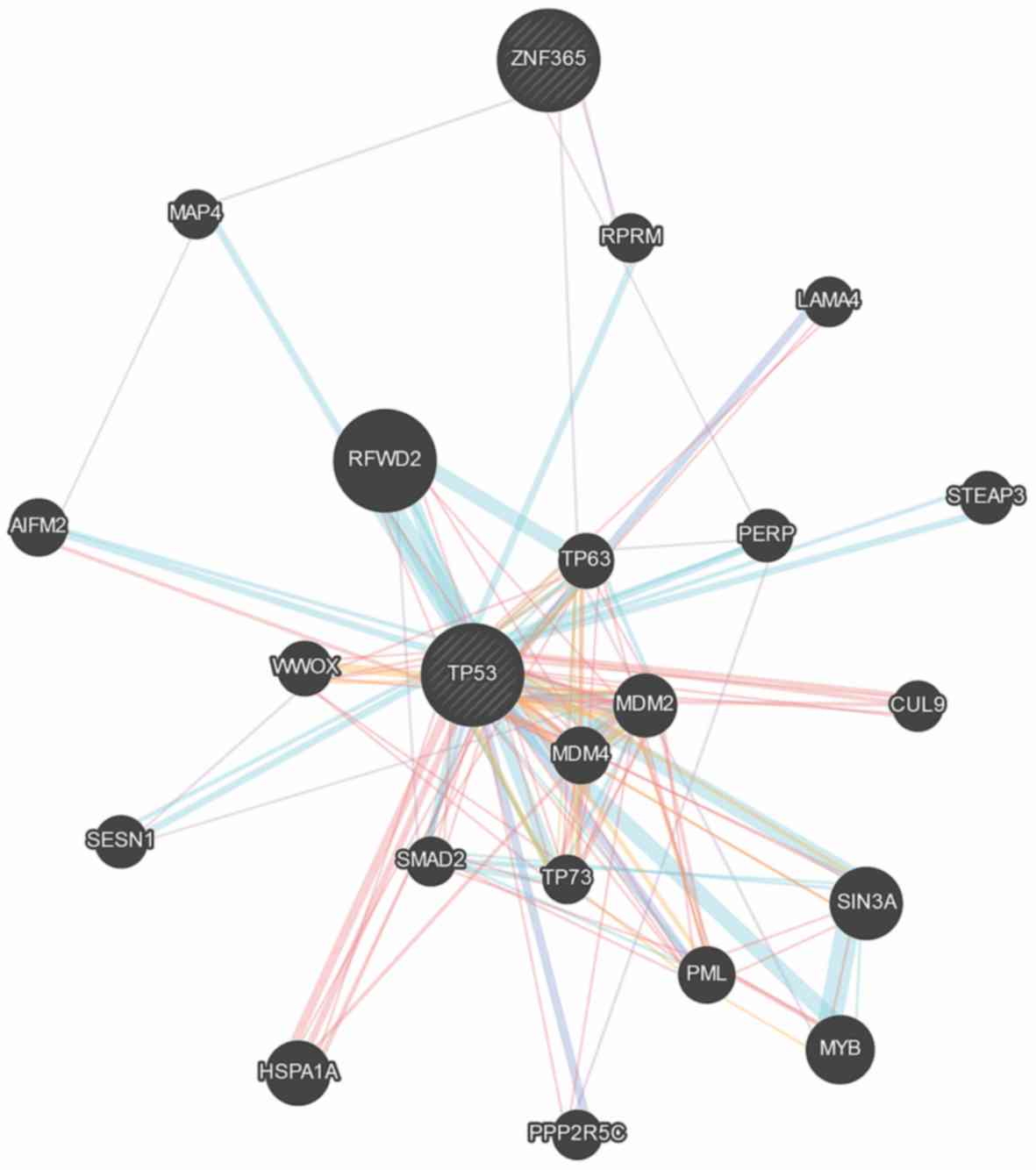
Warde-Farley D, Donaldson SL, Comes O,
Zuberi K, Badrawi R, Chao P, Franz M, Grouios C, Kazi F, Lopes CT,
et al: The GeneMANIA prediction server: Biological network
integration for gene prioritization and predicting gene function.
Nucleic Acids Res. 38((Web Server Issue)): W214–W220. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Ying J, Li H, Seng TJ, Langford C,
Srivastava G, Tsao SW, Putti T, Murray P, Chan AT and Tao Q:
Functional epigenetics identifies a protocadherin PCDH10 as a
candidate tumor suppressor for nasopharyngeal, esophageal and
multiple other carcinomas with frequent methylation. Oncogene.
25:1070–1080. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Naccarati A, Polakova V, Pardini B,
Vodickova L, Hemminki K, Kumar R and Vodicka P: Mutations and
polymorphisms in TP53 gene-an overview on the role in colorectal
cancer. Mutagenesis. 27:211–218. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Zhang Y, Shin SJ, Liu D, Ivanova E,
Foerster F, Ying H, Zheng H, Xiao Y, Chen Z, Protopopov A, et al:
ZNF365 promotes stability of fragile sites and telomeres. Cancer
Discov. 3:798–811. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Amre DK, Mack DR, Morgan K, Israel D,
Deslandres C, Seidman EG, Lambrette P, Costea I, Krupoves A, Fegury
H, et al: Susceptibility loci reported in genome-wide association
studies are associated with Crohn's disease in Canadian children.
Aliment Pharmacol Ther. 31:1186–1191. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Hirota T, Takahashi A, Kubo M, Tsunoda T,
Tomita K, Sakashita M, Yamada T, Fujieda S, Tanaka S, Doi S, et al:
Genome-wide association study identifies eight new susceptibility
loci for atopic dermatitis in the Japanese population. Nat Genet.
44:1222–1226. 2012. View
Article : Google Scholar : PubMed/NCBI
|
|
27
|
Zhang Y, Park E, Kim CS and Paik JH:
ZNF365 promotes stalled replication forks recovery to maintain
genome stability. Cell Cycle. 12:2817–2828. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Negrini S, Gorgoulis VG and Halazonetis
TD: Genomic instability-an evolving hallmark of cancer. Nat Rev Mol
Cell Biol. 11:220–228. 2010. View
Article : Google Scholar : PubMed/NCBI
|
|
29
|
Li H, Zhang J, Tong JHM, Chan AWH, Yu J,
Kang W and To KF: Targeting the oncogenic p53 mutants in colorectal
cancer and other solid tumors. Int J Mol Sci. 20:59992019.
View Article : Google Scholar
|
|
30
|
Lane DP: Cancer. p53, guardian of the
genome. Nature. 358:15–16. 1992. View
Article : Google Scholar : PubMed/NCBI
|
|
31
|
Harris SL and Levine AJ: The p53 pathway:
Positive and negative feedback loops. Oncogene. 24:2899–2908. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Shieh SY, Ikeda M, Taya Y and Prives C:
DNA damage-induced phosphorylation of p53 alleviates inhibition by
MDM2. Cell. 91:325–334. 1997. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Soussi T and Beroud C: Assessing TP53
status in human tumours to evaluate clinical outcome. Nat Rev
Cancer. 1:233–240. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Sigal A and Rotter V: Oncogenic mutations
of the p53 tumor suppressor: The demons of the guardian of the
genome. Cancer Res. 60:6788–6793. 2000.PubMed/NCBI
|
|
35
|
Liu YY, Patwardhan GA, Bhinge K, Gupta V,
Gu X and Jazwinski SM: Suppression of glucosylceramide synthase
restores p53-dependent apoptosis in mutant p53 cancer cells. Cancer
Res. 71:2276–2285. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Vedeld HM, Andresen K, Eilertsen IA,
Nesbakken A, Seruca R, Gladhaug IP, Thiis-Evensen E, Rognum TO,
Boberg KM and Lind GE: The novel colorectal cancer biomarkers CDO1,
ZSCAN18 and ZNF331 are frequently methylated across
gastrointestinal cancers. Int J Cancer. 136:844–853. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Jones PA: DNA methylation and cancer.
Cancer Res. 46:461–466. 1986.PubMed/NCBI
|
|
38
|
Riggs AD and Jones PA: 5-methylcytosine,
gene regulation, and cancer. Adv Cancer Res. 40:1–30. 1983.
View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Baylin SB: DNA methylation and gene
silencing in cancer. Nat Clin Pract Oncol. 2 (Suppl 1):S4–S11.
2005. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Moore LD, Le T and Fan G: DNA methylation
and its basic function. Neuropsychopharmacology. 38:23–38. 2013.
View Article : Google Scholar : PubMed/NCBI
|