|
1
|
Weimers P and Munkholm P: The natural
history of IBD: Lessons learned. Curr Treat Options Gastroenterol.
16:101–111. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Hovde Ø, Høivik ML, Henriksen M, Solberg
IC, Småstuen MC and Moum BA: Malignancies in patients with
inflammatory bowel disease: Results from 20 Years of follow-up in
the IBSEN Study. J Crohns Colitis. 11:571–577. 2017.PubMed/NCBI
|
|
3
|
Pellino G, Sciaudone G, Patturelli M,
Candilio G, De Fatico GS, Landino I, Facchiano A, Vastarella A,
Canonico S, Riegler G and Selvaggi F: Relatives of Crohn's disease
patients and breast cancer: An overlooked condition. Int J Surg. 12
(Suppl 1):S156–S158. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Riegler G, Caserta L, Castiglione F,
Esposito I, Valpiani D, Annese V, Zoli G, Gionchetti P, Viscido A,
Sturniolo GC, et al: Increased risk of breast cancer in
first-degree relatives of Crohn's disease patients. An IG-IBD
study. Dig Liver Dis. 38:18–23. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Barrett T, Wilhite SE, Ledoux P,
Evangelista C, Kim IF, Tomashevsky M, Marshall KA, Phillippy KH,
Sherman PM, Holko M, et al: NCBI GEO: Archive for functional
genomics data sets-update. Nucleic Acids Res. 41((Database Issue)):
D991–D995. 2013.PubMed/NCBI
|
|
6
|
Burczynski ME, Peterson RL, Twine NC,
Zuberek KA, Brodeur BJ, Casciotti L, Maganti V, Reddy PS, Strahs A,
Immermann F, et al: Molecular classification of Crohn's disease and
ulcerative colitis patients using transcriptional profiles in
peripheral blood mononuclear cells. J Mol Diagn. 8:51–61. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Labreche HG, Nevins JR and Huang E:
Integrating factor analysis and a transgenic mouse model to reveal
a peripheral blood predictor of breast tumors. BMC Med Genomics.
4:612011. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Lenz M, Müller FJ, Zenke M and Schuppert
A: Principal components analysis and the reported low intrinsic
dimensionality of gene expression microarray data. Sci Rep.
6:256962016. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Yu G, Wang LG, Han Y and He QY:
ClusterProfiler: An R package for comparing biological themes among
gene clusters. OMICS. 16:284–287. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Shannon P, Markiel A, Ozier O, Baliga NS,
Wang JT, Ramage D, Amin N, Schwikowski B and Ideker T: Cytoscape: A
software environment for integrated models of biomolecular
interaction networks. Genome Res. 13:2498–504. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
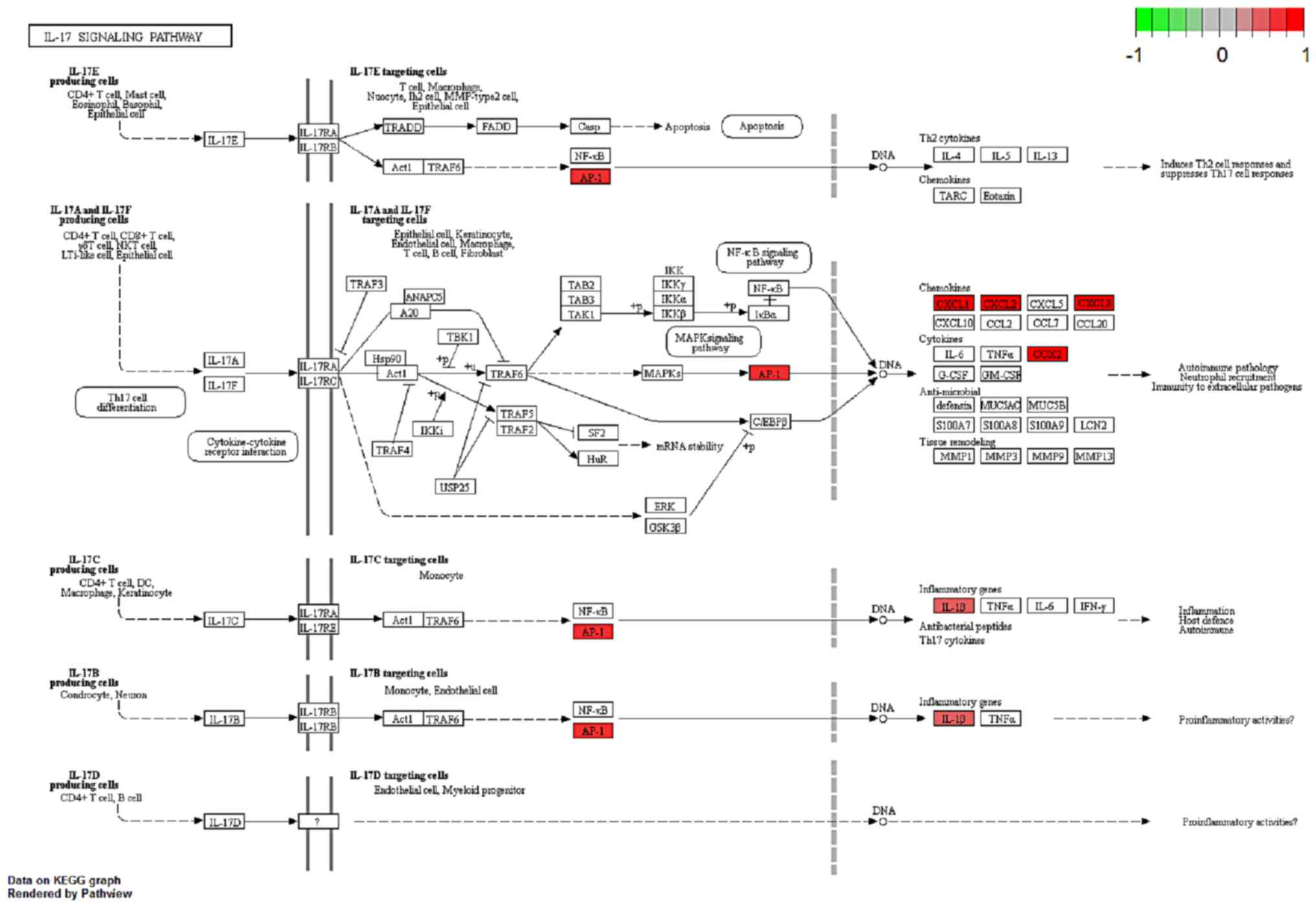
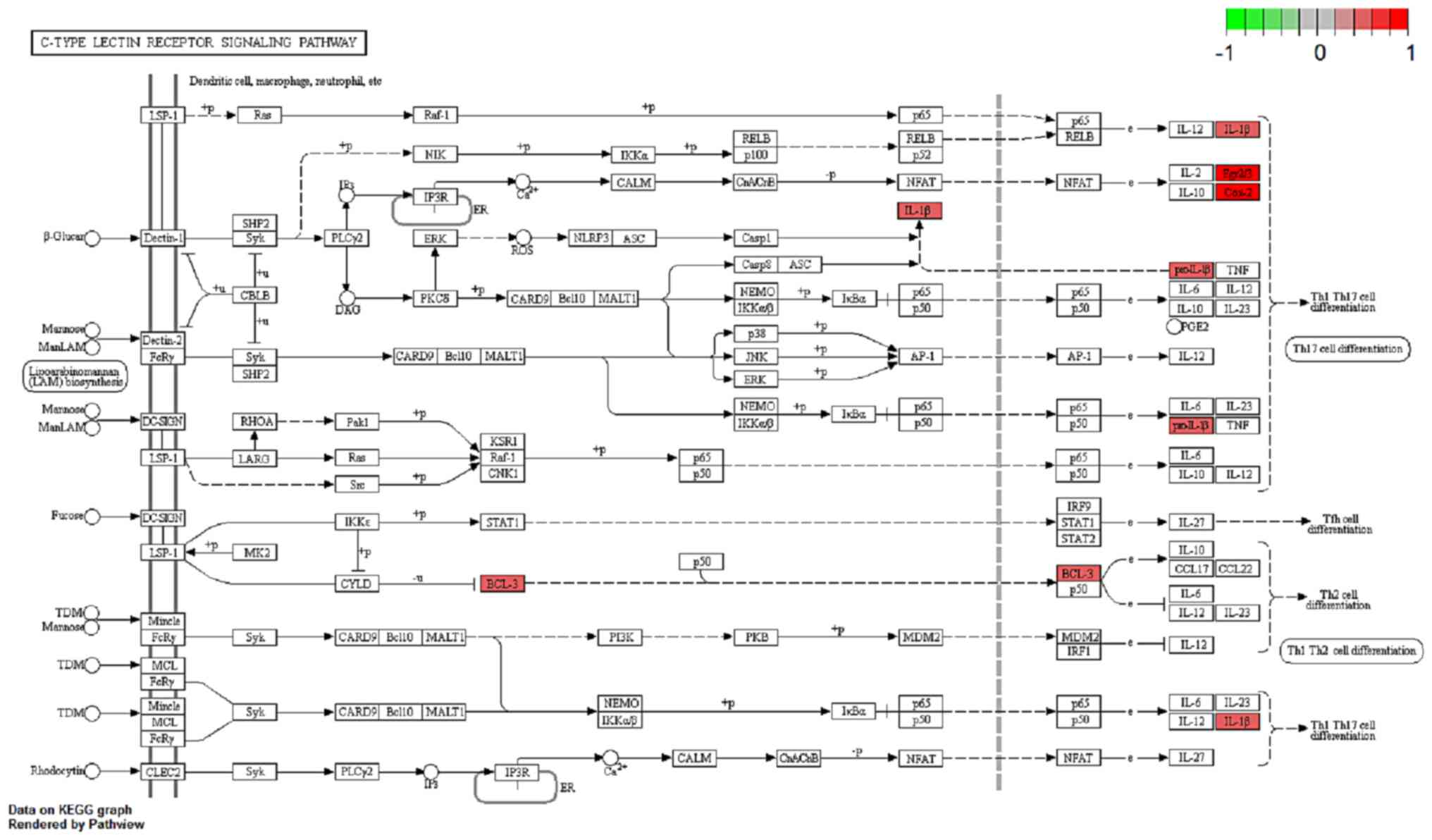
Luo W and Brouwer C: Pathview: An
R/Bioconductor package for pathway-based data integration and
visualization. Bioinformatics. 29:1830–1831. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Welte T and Zhang XHF: Interleukin-17
could promote breast cancer progression at several stages of the
disease. Mediators Inflamm. 2015:8043472015. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Wolfkamp SC, Verstege MI, Vogels EW,
Meisner S, Verseijden C, Stokkers PC and te Velde AA: Single
nucleotide polymorphisms in C-type lectin genes, clustered in the
IBD2 and IBD6 susceptibility loci, may play a role in the
pathogenesis of inflammatory bowel diseases. Eur J Gastroenterol
Hepatol. 24:965–970. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Hohenberger M, Cardwell LA, Oussedik E and
Feldman SR: Interleukin-17 inhibition: Role in psoriasis and
inflammatory bowel disease. J Dermatolog Treat. 29:13–18. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Tambuwala MM: Natural nuclear factor kappa
beta inhibitors: Safe therapeutic options for inflammatory bowel
disease. Inflamm Bowel Dis. 22:719–23. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Wang W, Nag SA and Zhang R: Targeting the
NFκB signaling pathways for breast cancer prevention and therapy.
Curr Med Chem. 22:264–89. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Abdulamir AS, Hafidh RR and Bakar FA:
Molecular detection, quantification, and isolation of Streptococcus
gallolyticus bacteria colonizing colorectal tumors:
Inflammation-driven potential of carcinogenesis via IL-1, COX-2,
and IL-8. Mol Cancer. 9:2492010. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Lee YA, Choi HM, Lee SH, Yang HI, Yoo MC,
Hong SJ and Kim KS: Synergy between adiponectin and interleukin-1β
on the expression of interleukin-6, interleukin-8, and
cyclooxygenase-2 in fibroblast-like synoviocytes. Exp Mol Med.
44:440–447. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Kameyama H, Nagahashi M, Shimada Y, Tajima
Y, Ichikawa H, Nakano M, Sakata J, Kobayashi T, Narayanan S, Takabe
K and Wakai T: Genomic characterization of colitis-associated
colorectal cancer. World J Surg Oncol. 16:1212018. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Han YM, Koh J, Kim JW, Lee C, Koh SJ, Kim
B, Lee KL, Im JP and Kim JS: NF-kappa B activation correlates with
disease phenotype in Crohn's disease. PLoS One. 12:e01820712017.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Kmieć Z, Cyman M and Ślebioda TJ: Cells of
the innate and adaptive immunity and their interactions in
inflammatory bowel disease. Adv Med Sci. 62:1–16. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Schreiber S, Nikolaus S and Hampe J:
Activation of nuclear factor κB in inflammatory bowel disease. Gut.
42:477–484. 1998. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Gálvez J: Role of Th17 cells in the
pathogenesis of human IBD. ISRN Inflamm. 2014:9284612014.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Bassolas-Molina H, Raymond E, Labadia M,
Labadia M, Wahle J, Ferrer-Picón E, Panzenbeck M, Zheng J, Harcken
C, Hughes R, et al: An RORγ oral inhibitor modulates IL-17
responses in peripheral blood and intestinal Mucosa of Crohn's
disease patients. Front Immunol. 9:23072018. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Schwarzmaier D, Foell D, Weinhage T, Varga
G and Dabritz J: Peripheral monocyte functions and activation in
patients with quiescent Crohn's disease. PLoS One. 8:e627612013.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Brand S: Crohn's disease: Th1, Th17 or
Both? The change of a paradigm: New immunological and genetic
insights implicate Th17 cells in the pathogenesis of Crohn's
disease. Gut. 58:1152–1167. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Kim H, Kim Y, Bae S, Kong JM, Choi J, Jang
M, Choi J, Hong JM, Hwang YI, Kang JS and Lee WJ: Direct
interaction of CD40 on tumor cells with CD40L on T cells increases
the proliferation of tumor cells by enhancing TGF-β production and
Th17 differentiation. PLoS One. 10:e01257422015. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Wang J, Cai D, Ma B, Wu G and Wu J:
Skewing the balance of regulatory T-cells and T-helper 17 cells in
breast cancer patients. J Int Med Res. 39:691–701. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Ma K, Yang L, Shen R, Kong B, Chen W,
Liang J, Tang G and Zhang B: Th17 cells regulate the production of
CXCL1 in breast cancer. Int Immunopharmacol. 56:320–329. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Alinejad V, Dolati S, Motallebnezhad M and
Yousefi M: The role of IL17B-IL17RB signaling pathway in breast
cancer. Biomed Pharmacother. 88:795–803. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Lu H, Clauser KR, Tam WL, Fröse J, Ye X,
Eaton EN, Reinhardt F, Donnenberg VS, Bhargava R, Carr SA and
Weinberg RA: A breast cancer stem cell niche supported by
juxtacrine signalling from monocytes and macrophages. Nat Cell
Biol. 16:1105–1117. 2014. View
Article : Google Scholar : PubMed/NCBI
|
|
32
|
Li B, Lu Y, Yu L, Han X, Wang H, Mao J,
Shen J, Wang B, Tang J, Li C and Song B: miR-221/222 promote cancer
stem-like cell properties and tumor growth of breast cancer via
targeting PTEN and sustained Akt/NF-κB/COX-2 activation. Chem Biol
Interact. 277:33–42. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Nomura A, Gupta VK, Dauer P, Sharma NS,
Dudeja V, Merchant N, Saluja AK and Banerjee S: NFκB-mediated
invasiveness in CD133+ pancreatic TICs is regulated by
autocrine and paracrine activation of IL1 signaling. Mol Cancer
Res. 16:162–172. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Ha H, Debnath B and Neamati N: Role of the
CXCL8-CXCR1/2 axis in cancer and inflammatory diseases.
Theranostics. 7:1543–1588. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Liu Q, Li A, Tian Y, Wu JD, Liu Y, Li T,
Chen Y, Han X and Wu K: The CXCL8-CXCR1/2 pathways in cancer.
Cytokine Growth Factor Rev. 31:61–71. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Dinarello CA: An interleukin-1 signature
in breast cancer treated with interleukin-1 receptor blockade:
Implications for treating cytokine release syndrome of checkpoint
inhibitors. Cancer Res. 78:5200–5202. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Perez-Yepez EA, Ayala-Sumuano JT, Lezama R
and Meza I: A novel β-catenin signaling pathway activated by IL-1β
leads to the onset of epithelial-mesenchymal transition in breast
cancer cells. Cancer Lett. 354:164–171. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Naldini A, Filippi I, Miglietta D,
Moschetta M, Giavazzi R and Carraro F: Interleukin-1β regulates the
migratory potential of MDAMB231 breast cancer cells through the
hypoxia-inducible factor-1α. Eur J Cancer. 46:3400–3408. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Filippi I, Carraro F and Naldini A:
Interleukin-1β affects MDAMB231 breast cancer cell migration under
hypoxia: Role of HIF-1α and NFκB transcription factors. Mediators
Inflamm. 2015:7894142015. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
O'Connor PM, Lapointe TK, Beck PL and
Buret AG: Mechanisms by which inflammation may increase intestinal
cancer risk in inflammatory bowel disease. Inflamm Bowel Dis.
16:1411–1420. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Reader J, Holt D and Fulton A:
Prostaglandin E2 EP receptors as therapeutic targets in breast
cancer. Cancer Metastasis Rev. 30:449–463. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Wang X, Reyes ME, Zhang D, Funakoshi Y,
Trape AP, Gong Y, Kogawa T, Eckhardt BL, Masuda H, Pirman DA Jr, et
al: EGFR signaling promotes inflammation and cancer stem-like
activity in inflammatory breast cancer. Oncotarget. 8:67904–67917.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Maroni P, Bendinelli P, Matteucci E and
Desiderio MA: The therapeutic effect of miR-125b is enhanced by the
prostaglandin endoperoxide synthase 2/cyclooxygenase 2 blockade and
hampers ETS1 in the context of the microenvironment of bone
metastasis. Cell Death Dis. 9:4722018. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Põld M, Zhu LX, Sharma S, Burdick MD, Lin
Y, Lee PP, Põld A, Luo J, Krysan K, Dohadwala M, et al:
Cyclooxygenase-2-dependent expression of angiogenic CXC chemokines
ENA-78/CXC Ligand (CXCL) 5 and interleukin-8/CXCL8 in human
non-small cell lung cancer. Cancer Res. 64:1853–1860. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Gross V, Andus T, Leser HG, Roth M and
Schölmerich J: Inflammatory mediators in chronic inflammatory bowel
diseases. Klin Wochenschr. 69:981–987. 1991. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Deshmukh SK, Srivastava SK, Poosarla T,
Dyess DL, Holliday NP, Singh AP and Singh S: Inflammation,
immunosuppressive microenvironment and breast cancer: Opportunities
for cancer prevention and therapy. Ann Transl Med. 7:5932019.
View Article : Google Scholar : PubMed/NCBI
|