Introduction
Modifications to histone amino-terminals are
significant for the regulation of chromatin structure, interaction
with chromatin-associated proteins, transcription and DNA
replication (1–3). Histone lysine methylases (KMTs) are
responsible for modifications in amino-acid residues of the exposed
N-terminal domain of histones by methylation of lysines. As a
result, the gene is activated or repressed (2,3).
Methyltransferases of mixed lineage leukemia (MLL) and the SET and
MYND domain-containing protein (SMYD) family are involved in
trimethylation of lysine 4 at histone 3 (3). Expression of SMYD family members is
significantly altered in various human diseases (3,4). Their
participation has been studied in cancer, embryonic heart
development and inflammatory processes (4). The SMYD family consists of five members
with a different structure from the other KMTs: The SET domain is
split into two segments by an MYND domain, followed by a
cysteine-rich post SET domain (1,3,5,6). SMYD3
was the first member of the SMYD family for which catalytic
significance was also demonstrated for domains other than classical
SET (3,7). SMYD3 was recently shown also to
catalyze methylation of lysine 5 of histone 4 (3,8).
Specific SMYD3 binding elements in the target DNA (5′-CCCTCC-3′ or
5′-GGAGGG-3′) are present in gene promoter regions below
SMYD3, such as Nks2.8, WHT10B, and HTERT (1,9–11). SMYD3 activates transcription of
several oncogenes (e.g. C-Met, JUND and Wnt10B), cell
cycle regulating genes (e.g. CDK2 and β DNA topoisomerase)
and genes responsible for signal transduction (e.g. RAB40C
and GNRF2). However, SMYD3 inhibits the expression of some
tumor suppressor genes (e.g. RIZ1) by epigenetic regulation
(1,9,12,13).
Increased SMYD3 expression is significant for cell
viability, adhesion, migration and invasion (1,14). It
correlates with poor prognosis in various types of cancer. The
nucleus placed protein-coding gene SMYD3 is a selective
transcription enhancer of oncogenes and the process of cell
proliferation in liver and colon cancers. The mechanism is based on
interaction with RNA Pol II and H3K4me3 and, in the case of liver
tumors, is strongly associated with poor prognosis. Additionally,
SMYD3 has an impact on Ras/ERK signaling in lung and pancreatic
cancers by methylation of MAP3K2 kinase (3). A significant correlation has been found
between the genetic variant with the variable number tandem repeat
(VNTR) in the SMYD3 gene and the development of breast
cancer in Jordanian women (15).
Overexpression of SMYD3 also correlates with a more
aggressive phenotype of prostate cancer (16). The role of the SMYD3 gene is
also being studied in cholangiocarcinoma, esophageal squamous cell
carcinoma, cervical, ovarian, bladder, gastric cancers, chronic
lymphocytic leukemia and glioma (17–26). No
studies are being undertaken to evaluate SMYD3 expression in
papillary thyroid cancers, while one previous study confirmed its
overexpression in medullary thyroid cancers (24).
An enhancer of zeste homolog 2 (EZH2) is a histone
methyltransferase, the catalytic subunit of the Polycomb 2
repression complex responsible for the trimethylation of histone H3
lysine 27 (H3K27me3) (27,28). Research on embryonic EZH2-zero
(ESC) stem cells has shown residual H3K27me3, termed EZH1
methyltransferase. This may indicate at least partial compensation
of both enzymes (29–31). Genetic loss-of-function studies have
demonstrated a role for EZH2 in the establishment and physiology of
several cell types and tissues, such as the skin, heart and mammary
gland (29–33). Similar to SMYD3, EZH2 is highly
expressed in various types of cancer, which is often also
correlated with poor prognosis. The effect of EZH2 gene
expression on carcinogenesis is the promotion of survival,
proliferation, transformation of epithelium to mesenchyme, and the
invasion and drug resistance of cancers. However, tumor-suppressive
effects of EZH2 have also been identified. EZH2 has a significant
impact on immune cells (27). The
overexpression of EZH2 has been demonstrated in breast,
prostate, endometrial, bladder, liver, lung, ovarian cancer,
melanoma, glioblastoma and Natural killer/T-cell lymphoma.
Gain-of-function mutations are present in Non-Hodgkin's lymphoma
and melanoma. Through repression of the TIMP-3 metastatic
suppressor gene, EZH2 leads to progression and spread in prostate
and lung cancers and, through missense mutation in lymphomas, leads
to increased function of the mutated protein (28).
These genes are currently undergoing extensive
research in various types of cancers, including thyroid cancers.
These are seen as the goals for targeted therapeutic strategies in
oncology. Both genes have already been studied in medullary thyroid
cancer (MTC) (24). In addition,
research on the EZH2 gene has also been carried out on
poorly differentiated (PDTC) and anaplastic thyroid carcinoma (ATC)
(34), and also papillary thyroid
cancer (35). Our study aimed to
analyze EZH2 and SMYD3 gene expression in papillary
thyroid cancer (PTC), the most common form of malignancy in this
organ, and to correlate this with clinical outcome.
Material and methods
Tissue samples
Samples of resected thyroid tissue from consecutive
patients were collected: papillary thyroid cancers and thyroid
tissue from thyroids without cancer excised for nodular goiter. All
patients underwent primary thyroid surgery. We excluded patients
with mixed thyroid cancers. Tissue samples were stored in RNA
protective medium at −80°C until reverse transcription-quantitative
PCR (RT-qPCR) analysis. The Ethical Committee of Poznan University
of Medical Sciences approved the study (approval no. 228/14) and
each patient provided written informed consent.
Nucleic acid extraction and
validation
RNA was isolated from tissue specimens using the
Direct-zol RNA kit column system for high molecular weight RNA
isolation according to the manufacturer's protocol (Zymo Research).
In short, ~25 mg of tissue was homogenized in liquid nitrogen, and
suspended in TriReagent (GenoPlast). After chloroform addition, the
samples were centrifuged (12,000 × g, 15 min, 4°C) and the aqueous
phase was transferred in the column. The isolation, following the
protocol, finished with RNA recovery from silica matrix columns in
pre-warmed water. The quality, quantity and purity of RNA were
analyzed as described before (36)
with the use of a NanoPhotometer® NP-80 (IMPLEN). The
integrity was evaluated by electrophoretic separation in denaturing
conditions.
RT-qPCR
Complementary to RNA DNA (cDNA) was synthesized in a
three-step reaction conducted following the Transcriptor Reverse
Transcriptase manufacturer's protocol (Roche Diagnostics GmbH) in a
total volume of 20 µl. A mixture of 5 mM oligo(d)T10,
RNA (1 µg) and RNase-, DNase- and pyrogen-free water was incubated
for 10 min at 65°C. Subsequently, the samples were chilled.
Subsequently, 10 U/µl ribonuclease inhibitor (RNasin, Roche
Diagnostics GmbH), 10 U/µl of transcriptor reverse transcriptase
(Roche Diagnostics GmbH), 100 mM deoxyribonucleotide triphosphates
(Novayzm) and 1× reaction buffer (Roche Diagnostics GmbH) were
added. The subsequent steps of cDNA synthesis had been described
earlier (36).
RNA expression pattern analysis was performed using
a LightCycler® 2.0 (Roche Diagnostics GmbH). Primer
sequences and TaqMan® hydrolysis probe positions for the
EZH2 and SMYD3 were determined using the Roche
Universal ProbeLibrary (UPL) Assay Design Center (http://qpcr.probefinder.com; last accessed on May 10,
2017). Ensembl (http://www.ensembl.org/), and protein-coding sequences
for EZH2 (ENST00000320356.7, ENST00000460911.5,
ENST00000476773.5, ENST00000478654.5, ENST00000350995.6,
ENST00000483967.5) were used to design the primers and hydrolysis
probes (Roche TaqMan Probe UPL #35; cat. no. 04687680001). The
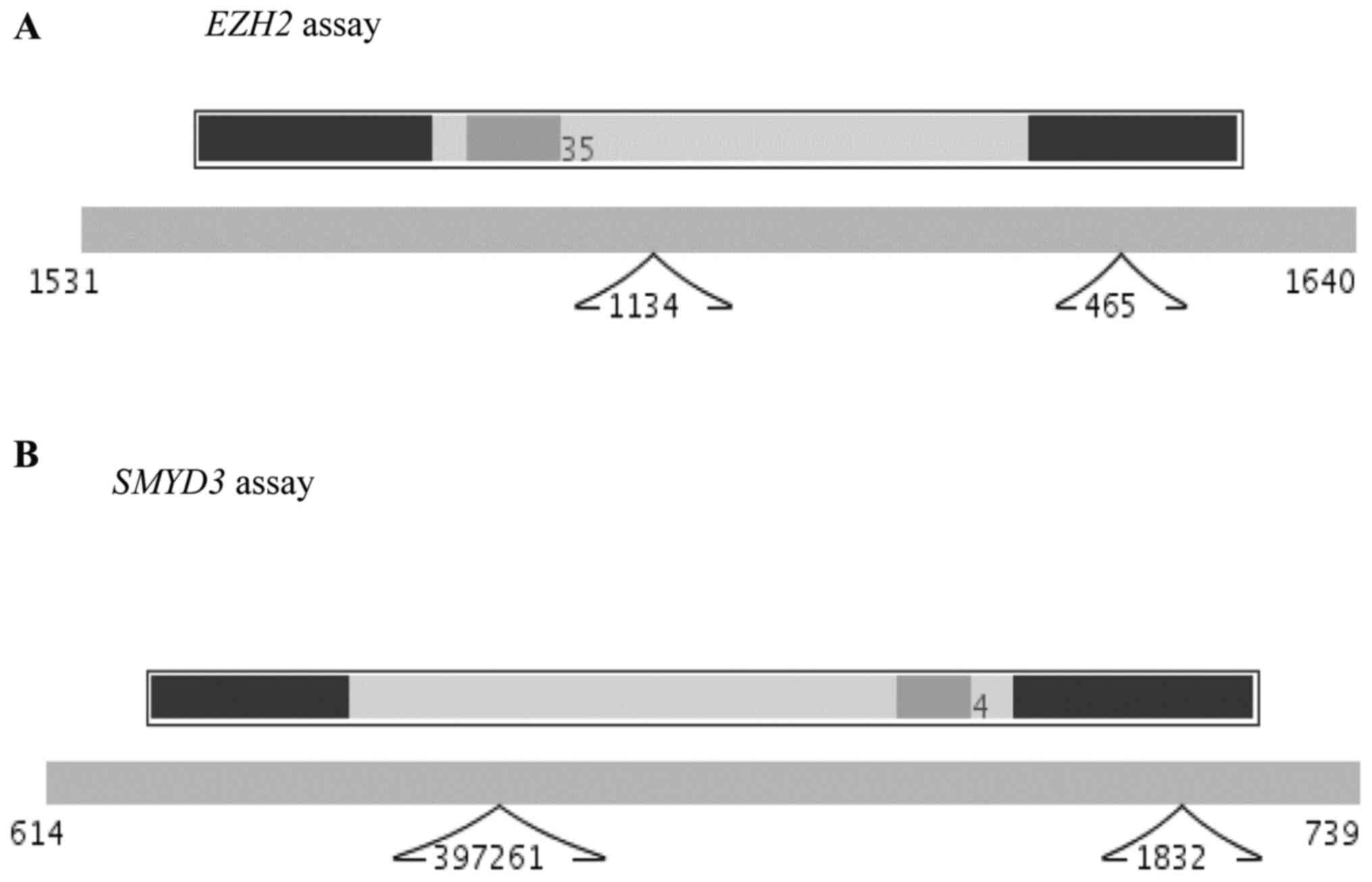
forward (5′-tgtggatactcctccaaggaa-3′) and reverse
(5′-gaggagccgtcctttttca-3′) primers flank the 90 nt amplicon
(Fig. 1A). The set of forward
(5′-cctgcctttgacctttttga-3′) and reverse
(5′-agatactgggatataggccaacac-3′) primers' for SMYD3 covered
the 106 nt amplicons with the TaqMan UPL #4 in between (Roche
Diagnostics GmbH; cat. no. 04685016001). The assay was designed for
both transcript variants (NCBI NM_001167740 and NM_022743.2)
(Fig. 1B). The hypoxanthine-guanine
phosphoribosyltransferase (HPRT) gene assay (Roche cat. no.
05532957001) served as an internal control.
The quantitative polymerase chain reactions had been
standardized earlier and conducted as described before (36) in a total volume of 20 µl with a 1×
LightCycler® FastStart TaqMan® Probe Master
mix. Each reaction was performed in duplicate on independently
synthesized cDNA, and the mean values were used for statistical
analyses. Reaction efficiencies were obtained from the genes'
standard curves (36). Raw data for
threshold values were analyzed by comparing them to appropriately
selected standard curves and the reference gene assay with the use
of LC 5.0.0.38 software, and presented as concentration ratios
(Cr).
Statistical analysis
Statistical analysis was performed with MedCalc
Statistical Software version 19.1.5 (MedCalc Software bv;
https://www.medcalc.org; 2020). The
D'Agostino-Pearson test analyzed normality. P<0.05 was
considered to indicate a statistically significant difference. The
Mann-Whitney test was used to compare non-normally distributed
parameters between the study and control groups, as well as between
analyzed subgroups. When data followed a normal distribution,
Student's t-test was used for comparison between groups. The
χ2 test was applied to compare descriptive parameters.
The Kruskal-Wallis test with Conover post-hoc test was used to
compare gene expression between thyroid cancer samples staged 1, 2,
3, and 4. The Spearman's correlation coefficient test was used to
find relationships between analyzed parameters.
Results
Patient characteristics
The study group consisted of 62 patients with
papillary thyroid cancers. There were 30 tissue samples in the
healthy control group. Clinical data are presented in Table I. The study, and the control groups
did not differ according to patients' age or sex.
 | Table I.Clinical data of the study and the
control groups. |
Table I.
Clinical data of the study and the
control groups.
|
Characteristics | Papillary thyroid
cancer group (n=62) | Control group
(n=30) | P-value |
|---|
| Mean ± SD age,
years | 51±16 | 46±16 | 0.17 |
| Sex |
|
| 0.15 |
|
Female | 43 | 25 |
|
|
Male | 19 | 5 |
|
| Histological
variant |
|
| – |
|
Conventional | 54 | – |
|
|
Follicular | 4 | – |
|
|
Oncocytic | 3 | – |
|
| Tall
cell | 1 | – |
|
| Staging at
diagnosis |
|
| – |
| I | 35 | – |
|
| II | 12 | – |
|
|
III | 13 | – |
|
| IV | 12 | – |
|
| Metastases to the
lymph nodes (%) | 14 (22.6) | – | – |
| Multifocality
(%) | 27 (43.5) | – | – |
| Distant metastases
(%) | 1 (1.6) | – | – |
EZH2 expression
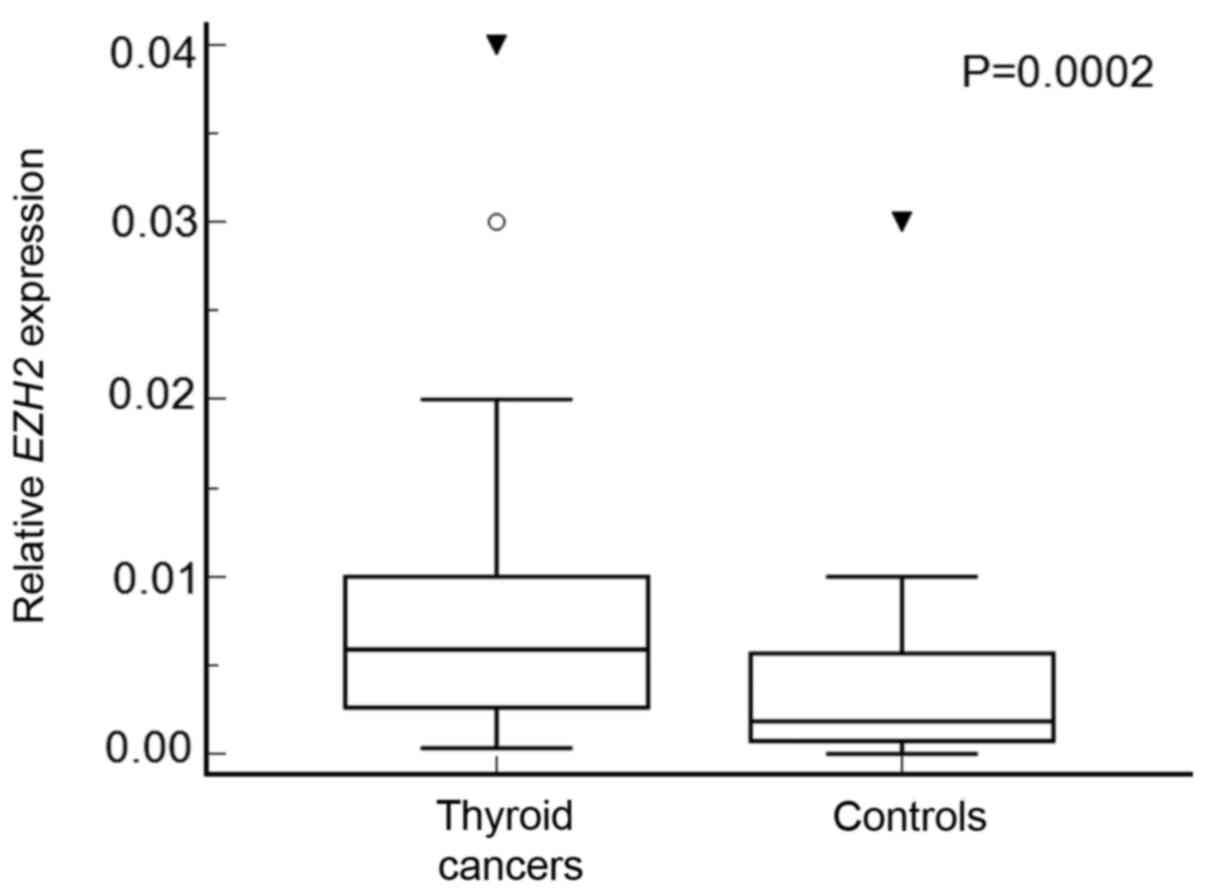
EZH2 expression was found in all thyroid
cancer samples and 25 out of 30 samples of benign lesions. We found
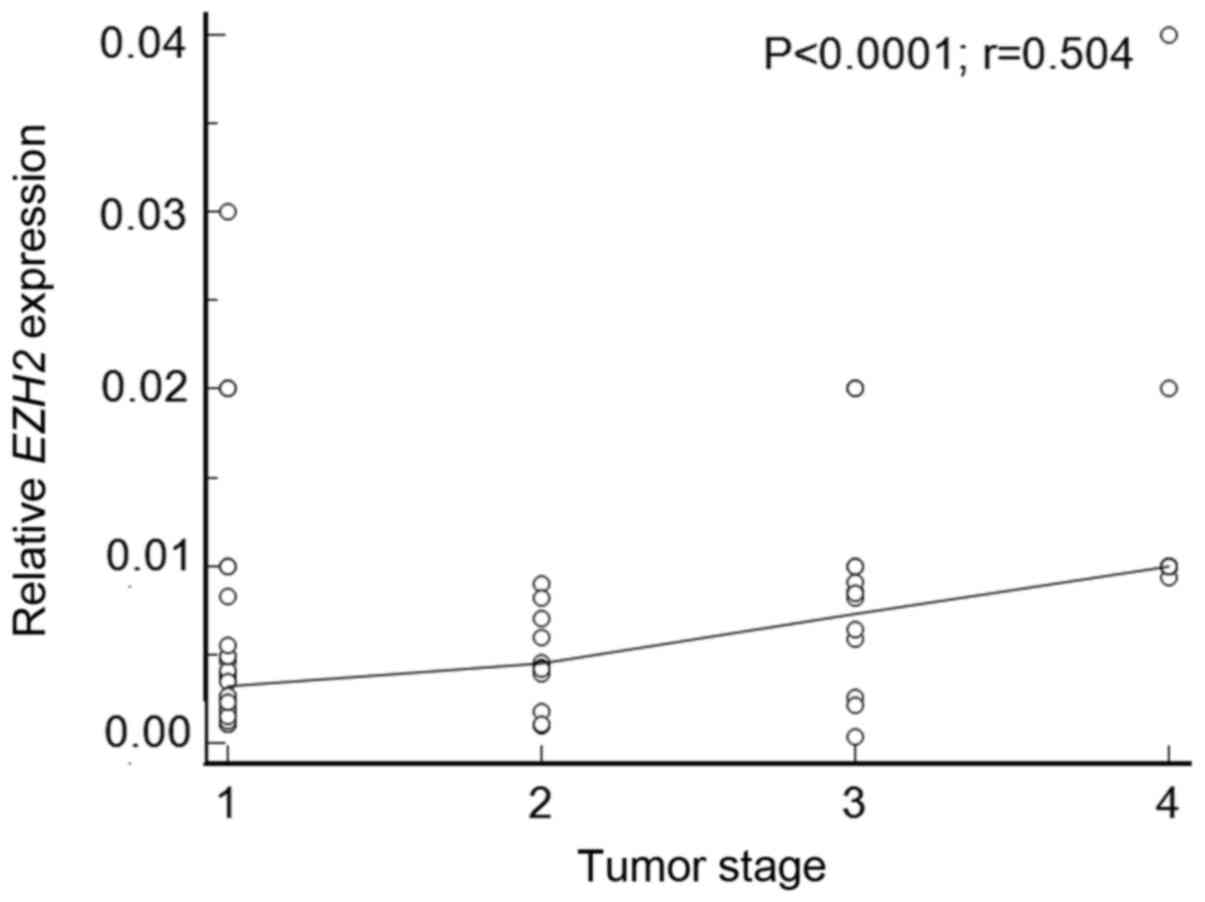
EZH2 overexpression in thyroid cancers (P=0.0002) (Fig. 2). EZH2 expression positively
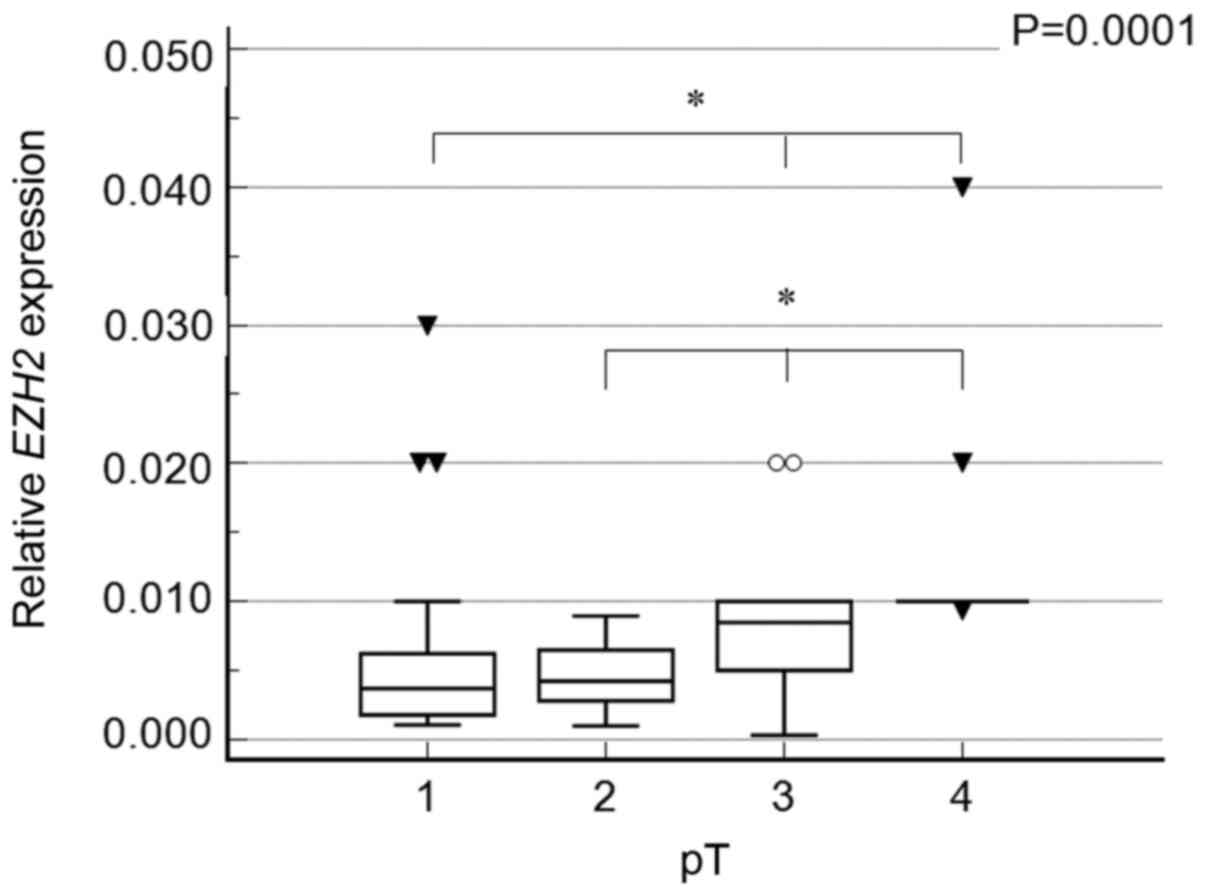
correlated with tumor stage (P<0.0001; r=0.504; Fig. 3), and multiple comparison analysis
revealed the highest expression in samples staged pT4 (P=0.0001)
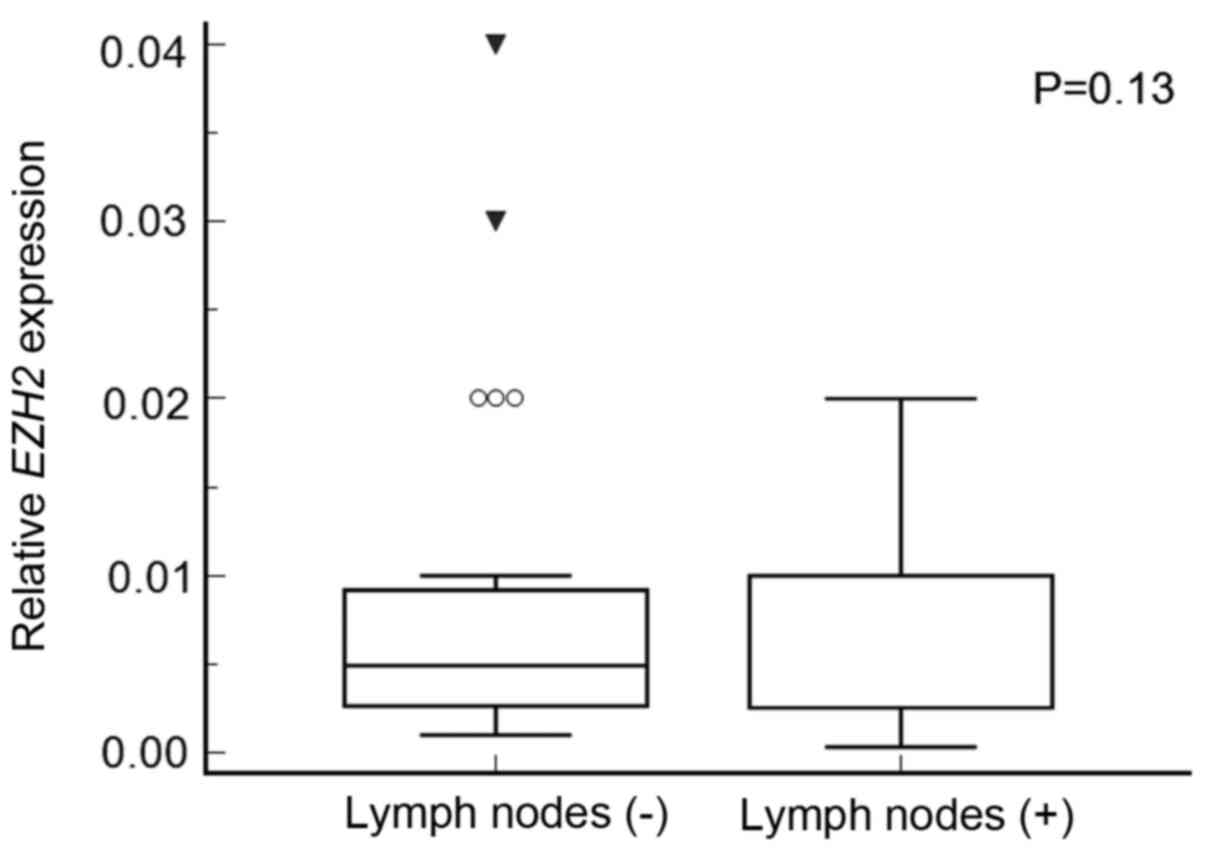
(Fig. 4). We did not observe
EZH2 overexpression in patients with lymph node involvement
(Fig. 5), and there was no
association between EZH2 expression and multifocality
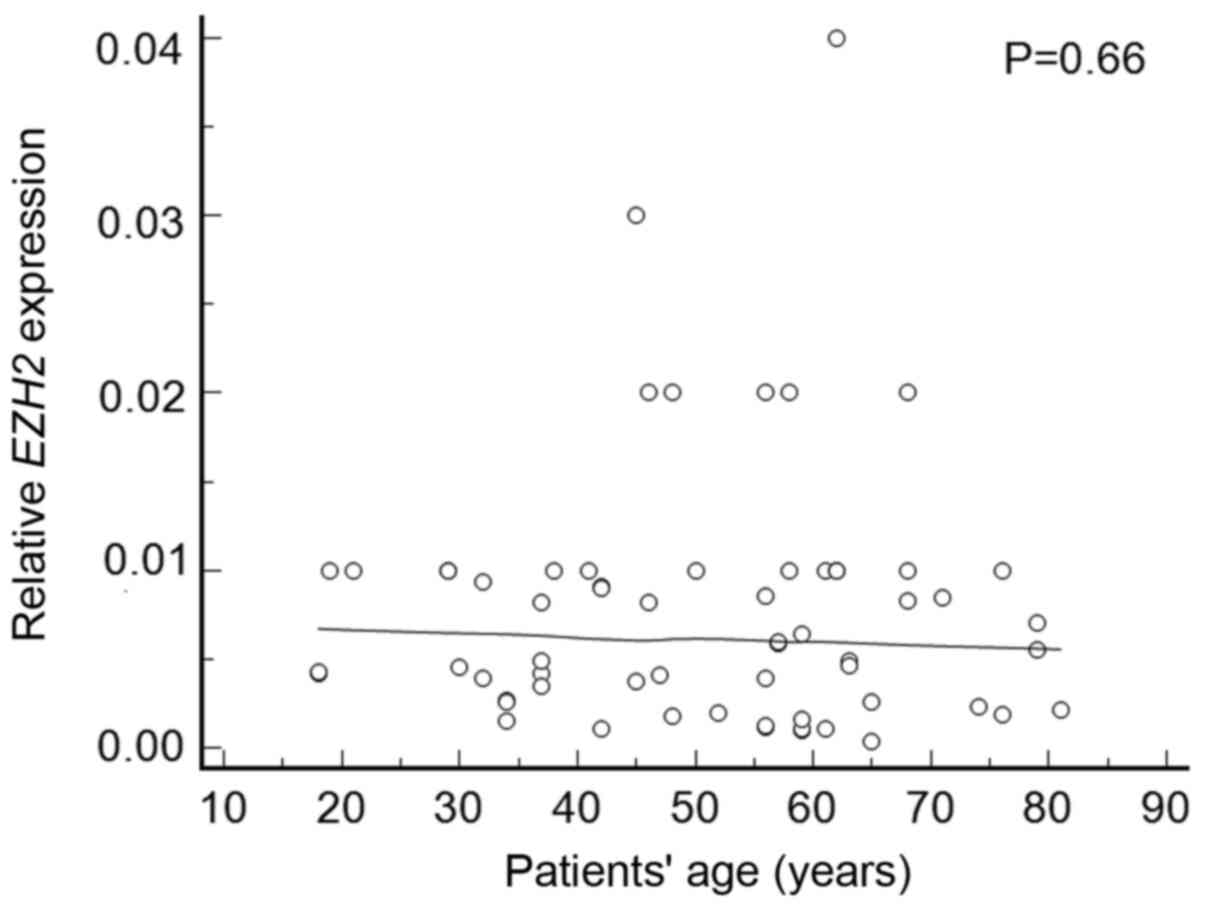
(P=0.13 and P=0.49, respectively). Also, patients' age did not
correlate with EZH2 expression levels (P=0.66) (Fig. 6).
SMYD3 expression
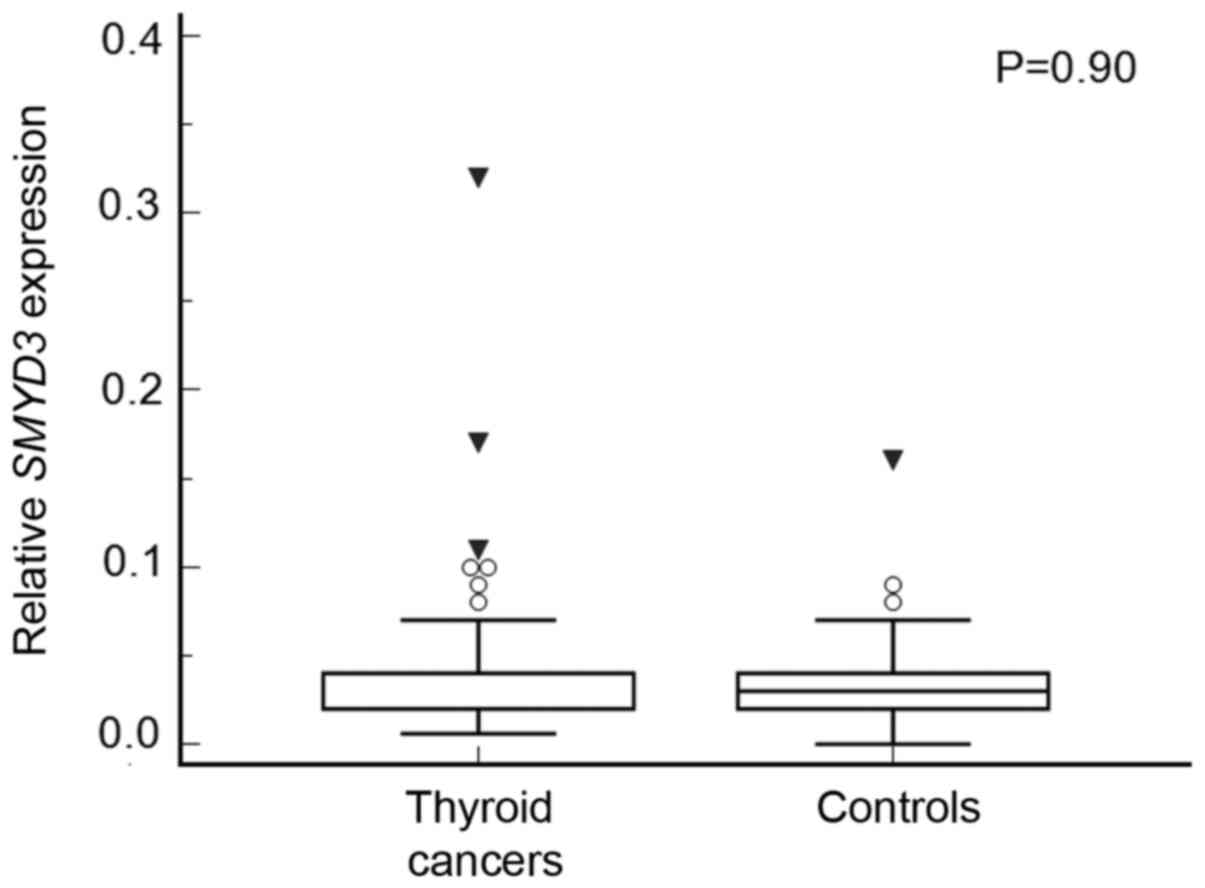
SMYD3 expression was found in all thyroid
cancer samples and 29 out of 30 healthy tissues, and the expression
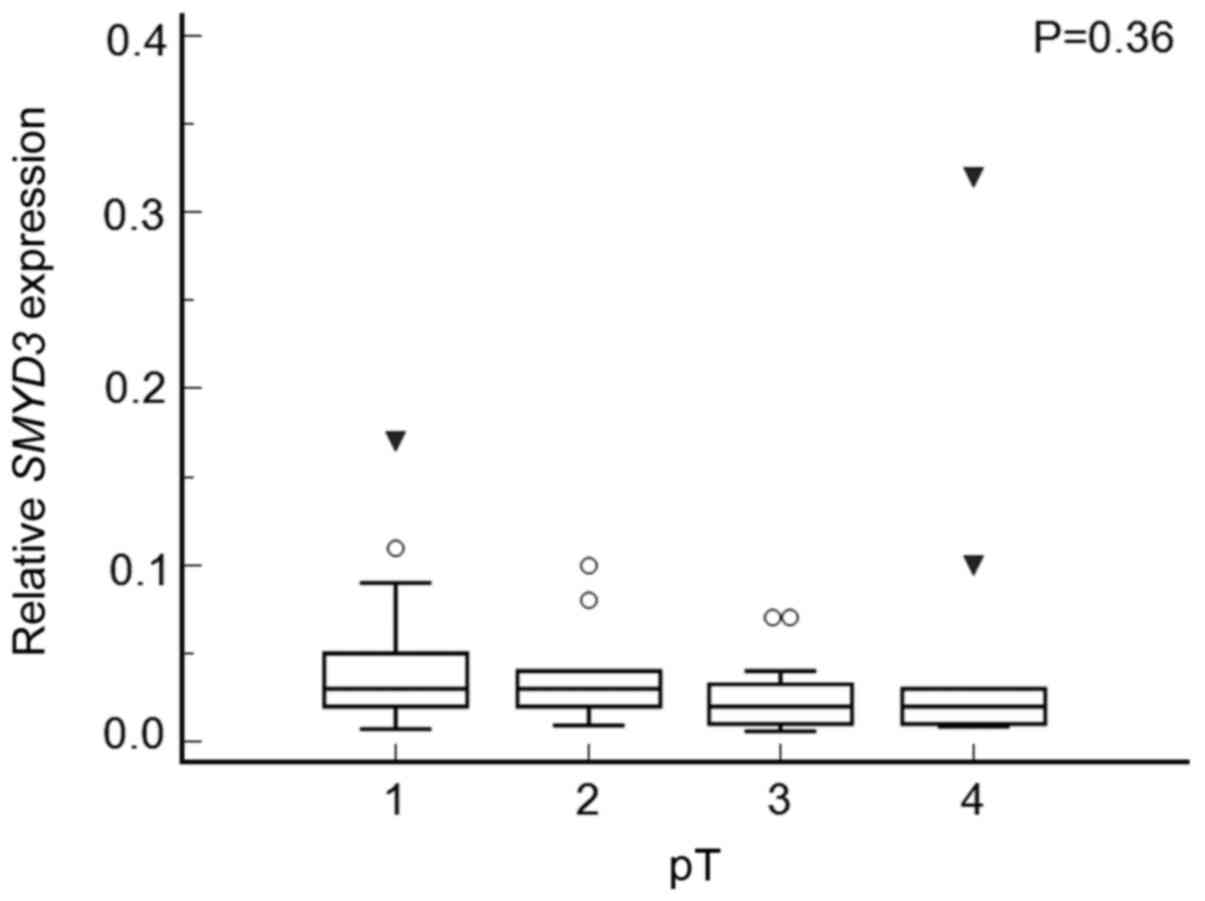
levels were similar in both groups (P=0.90) (Fig. 7). Also, there were no differences in
SMYD3 expression between tumors staged pT1, pT2, pT3 or pT4
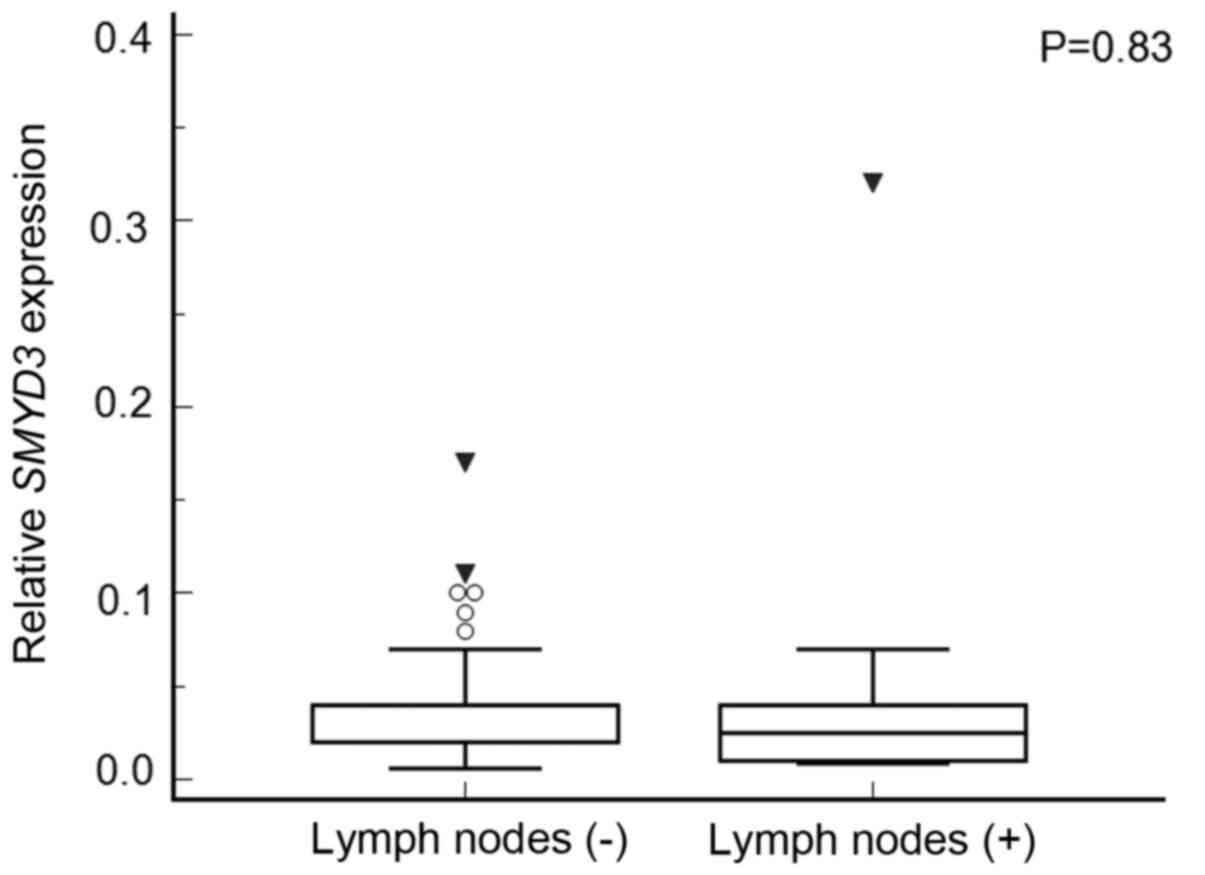
(P=0.37) (Fig. 8). Patients with
metastases to the lymph nodes did not have higher SMYD3
expression than those without (P=0.83) (Fig. 9). We did not observe any correlation
between SMYD3 expression and multifocality (P=0.45).
Discussion
We found histone methyltransferase EZH2
overexpression in papillary thyroid cancer (PTC), while
SMYD3 expression was not elevated. EZH2 gene
expression was found in all papillary thyroid cancer samples, but
also in most, as many as five sixths of healthy thyroid tissue
samples. These were significantly higher expression rates, in both
the study and control groups, than those obtained by Xue et
al (35). However, they examined
the expression of the EZH2 gene both by real-time PCR, as in
our study, and immunohistochemistry (IHC), and they presented the
expression percentages for IHC (35). However, in papillary thyroid
carcinomas, statistically significant overexpression of the
EZH2 gene was found in our study. Therefore, the EZH2
gene may be associated with the development of papillary thyroid
cancer. Xue et al (35)
obtained similar results. This is the case in papillary thyroid
cancer, as well as in other thyroid cancers, as shown in Table II (24,34,35).
 | Table II.Studies on EZH2 gene in
thyroid cancers. |
Table II.
Studies on EZH2 gene in
thyroid cancers.
| First author,
year | Number of patients,
type of thyroid cancer | Number of patients
in the control group | Analytical
technique | Results | Refs. |
|---|
| Masudo, 2018 | 67 cases of PDTC
and 48 cases of ATC | 30 adjacent healthy
and differentiated thyroid carcinoma tissue | IHC | EZH2 was expressed
in PDTC and ATC, but not in the normal thyroid gland or DTC;
EZH-positivity increased in the order of DTC, PDTC, and ATC
(P<0.01); higher EZH2 expression correlated with more reduced
survival in PDTC (P=0.004) and ATC (P=0.166) | (34) |
| Sponziello,
2014 | 54 MTCs; 13
familial MTCs and 41 sporadic forms; 33 hosted an RET
mutation and 13 an RAS somatic mutation | – | qPCR | A significant
increase in EZH2 and SMYD3 gene expression in more
aggressive diseases (i.e. occurrence of metastases; persistent
disease; disease-related death); the increase in EZH2 and
SMYD3 did not correlate with the mutational status of
RET or RAS genes | (24) |
| Xue, 2019 | 65 PTCs | 30 adjacent healthy
thyroid tissues | qPCR and IHC | Higher EZH2
expression in PTC tissues than in healthy thyroid tissues;
EZH2 expression is associated with lymph node metastasis and
is recurrent; inhibition of EZH2 in PTC cell lines
downregulates cellular proliferation and migration. PTC is a
disease with a high incidence in females and E2-ERα upregulates
EZH2 expression | (35) |
The expression of the EZH2 gene positively
correlated with the tumor stage, in the case of tumorous staged
pT4, it was the highest. Xue et al (35) did not observe statistically
significant differences between tumors <=1 cm and tumors >1
cm, or between those that extended beyond the thyroid tissue and
those that did not. However, we did not observe EZH2
overexpression in patients with lymph node involvement, as had been
obtained by Xue et al (35).
Also, we did not find a relationship between EZH2 expression
and multifocality. In both studies, age did not significantly
correlate with EZH2 gene expression. Correlation with
aggressiveness in thyroid cancers was described by Sponziello et
al (24) based on their study on
medullary thyroid carcinomas (MTC). Sponziello et al
(24) examined the expression of
epigenetic regulators in medullary thyroid carcinomas (MTCs) and
correlated this with clinical outcome and RET or RAS
mutational status. In the case of a more aggressive disease, they
noted a significant increase in EZH2 and SMYD3 gene
expression (more than 3 and 2-fold, respectively). They determined
the aggressiveness of the disease, according to the current
guidelines (37), based on the
occurrence of lymph nodes and distant metastases, persistence after
primary treatment and disease-related death. Noticeably, they did
not observe a significant correlation between the overexpression of
EZH2 and SMYD3 and the mutational status of
RET or RAS genes. Therefore, the researchers
suggested that EZH2 and SMYD3 mRNA expression may be
useful prognostic biomarkers, and further studies are needed to
investigate their possibility of use in therapy of MTC patients
(24). Also, Masudo et al
(34) claim that EZH2
overexpression may be a useful prognostic marker for more
aggressive thyroid cancers. This is justified by their
statistically significant increase in EZH-positivity in
order from differentiated (DTC), then poorly differentiated (PDTC)
to anaplastic forms of thyroid cancers (ATC). Also, higher
EZH2 expression correlated with more reduced survival in the
case of less differentiated cancers (34). Similarly, the prognostic significance
of the EZH2 gene has already been observed in cancers of
other organs, including the prostate, lung or lymphomas (28). Currently, increasing numbers of
studies are being developed that expand the range of thyroid
cancers tested, as well as molecular mechanisms associated with the
impact of the EZH2 gene on carcinogenesis (38). EZH2 is important in medullary thyroid
cancer by affecting ERK and AKT signaling pathways. It also
controls genes of the Wnt/β-catenin (24). It has been observed that increased
expression of EZH2 in PTC cell lines upregulates cellular
proliferation and migration by affecting the E2-ERα signaling
pathway (35). Researchers have
observed the role of long noncoding RNA PVT1 in the development of
thyroid cancer through its involvement in the modulation of cell
proliferation by recruiting EZH2 and regulating the
thyroid-stimulating hormone receptor (TSHR) (39). In their search for differences
between thyroid follicular cancer and thyroid follicular adenoma
scientists have used network-based genetic profiling, which
includes the EZH2 gene (40).
Overexpression of SMYD3 was not
characteristic of papillary thyroid cancer in our study. Expression
of this gene was observed in every test sample and in almost every
control sample. Moreover, expression levels in study and control
samples were similar. There was also no correlation between
SMYD3 gene expression and the markers of greater disease
aggression. No studies on the expression of the SMYD3 gene
in papillary thyroid cancer have previously been performed.
However, our study was justified due to the overexpression of both
the EZH2 and SMYD3 genes observed by Sponziello et
al, as well as the correlation of both genes with greater
aggressiveness of medullary thyroid cancers (MTCs) (24). A similar correlation between the
expression level of these genes and tumor aggression has been
observed in cancers of other organs, e.g. liver or prostate
(3,16). Chemical probes are being developed to
target SMYD3 selectively (41).
In both the study and the control groups, the
majority of patients were women. It has been known for many years
that PTC is a disease with a high prevalence in women, which is
also confirmed by the current research. This tendency is emphasized
by Xue et al (35) who
recently published their research on the EZH2 gene in PTC on
similarly numerous research (n=65) and control (n=30) groups.
Moreover, the latest trends show that among women, the highest
increase in incidence was observed in 2014: 22.2 new cases were
diagnosed per 100,000 people (42).
Also, researchers have noted that the disproportion between women
and men is particularly intensified during the reproductive period
(35,43). It has been found that estrogen can
increase PTC growth, progression and metastasis and that E2
treatment can significantly increase EZH2 levels (35,44,45). The
effectiveness of the specific EZH2 inhibitor GSK126 also confirms
the above observations (35). Most,
as many as 25 thyroid cancers were in stage I. This means that
often the tumor was not larger than 2 cm and did not grow outside
the thyroid gland (37). Samples in
stages II-IV were similarly numerous. In thyroid cancers,
metastases to the lymph nodes were observed in 23% of patients.
Although PTC is often localized, the lymphatic tract is the most
common for metastasis, and the site of metastasis is usually local
lymph nodes (46). Literature data
show the influence of both genes on the fate of individual cells,
so it would be reasonable to compare cells from the same thyroid
that are neoplastic to those unchanged. On the other hand, thyroid
cancers might be multifocal, and molecular alterations proceed
cancer. So it could also potentially affect achieved results.
Our results indicate that overexpression of the
EZH2 gene may be associated with the development of
papillary thyroid cancer. Therefore, the EZH2 gene may also
be a potential therapeutic target in papillary thyroid cancer.
Acknowledgements
Not applicable.
Funding
The National Science Center partially supported this
work in Poland (grant no. DEC 2012/07/N/NZ5/01736).
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
NSG designed the study, was involved in data
collection, analyzed data, and wrote and revised the manuscript. SS
and PZ conducted the manuscript preparation and data
analysis/interpretation. MA carried out the experimental studies
and data analysis. AC was involved in data collection and data
analysis. PG conceived the study and was involved in data analysis.
MR made substantial contribution to acquisition of samples and
clinical data, and revised the paper. NSG and MA confirm the
authenticity of all the raw data. All authors read and approved the
final manuscript.
Ethics approval and consent to
participate
The Ethical Committee of Poznan University of
Medical Sciences approved the present study (approval no. 228/14).
Written informed consent was obtained from all patients.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Zou JN, Wang SZ, Yang JS, Luo XG, Xie JH
and Xi T: Knockdown of SMYD3 by RNA interference down-regulates
c-Met expression and inhibits cells migration and invasion induced
by HGF. Cancer Lett. 280:78–85. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Kouzarides T: Chromatin modifications and
their function. Cell. 128:693–705. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Giakountis A, Moulos P, Sarris ME, Hatzis
P and Talianidis I: Smyd3-associated regulatory pathways in cancer.
Semin Cancer Biol. 42:70–80. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Varier RA and Timmers HT: Histone lysine
methylation and demethylation pathways in cancer. Biochim Biophys
Acta. 1815:75–89. 2011.PubMed/NCBI
|
|
5
|
Spellmon N, Holcomb J, Trescott L,
Sirinupong N and Yang Z: Structure and function of SET and MYND
domain-containing proteins. Int J Mol Sci. 16:1406–1428. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Foreman KW, Brown M, Park F, Emtage S,
Harriss J, Das C, Zhu L, Crew A, Arnold L, Shaaban S and Tucker P:
Structural and functional profiling of the human histone
methyltransferase SMYD3. PLoS One. 6:e222902011. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Xu S, Wu J, Sun B, Zhong C and Ding J:
Structural and biochemical studies of human lysine
methyltransferase Smyd3 reveal the important functional roles of
its post-SET and TPR domains and the regulation of its activity by
DNA binding. Nucleic acids Res. 39:4438–4449. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Van Aller GS, Reynoird N, Barbash O,
Huddleston M, Liu S, Zmoos AF, McDevitt P, Sinnamon R, Le B, Mas G,
et al: Smyd3 regulates cancer cell phenotypes and catalyzes histone
H4 lysine 5 methylation. Epigenetics. 7:340–343. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Hamamoto R, Furukawa Y, Morita M, Iimura
Y, Silva FP, Li M, Yagyu R and Nakamura Y: SMYD3 encodes a histone
methyltransferase involved in the proliferation of cancer cells.
Nat Cell Biol. 6:731–740. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Hamamoto R, Silva FP, Tsuge M, Nishidate
T, Katagiri T, Nakamura Y and Furukawa Y: Enhanced SMYD3 expression
is essential for the growth of breast cancer cells. Cancer Sci.
97:113–118. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Liu C, Fang X, Ge Z, Jalink M, Kyo S,
Björkholm M, Gruber A, Sjöberg J and Xu D: The telomerase reverse
transcriptase (hTERT) gene is a direct target of the histone
methyltransferase SMYD3. Cancer Res. 67:2626–2631. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Guo N, Chen R, Li Z, Liu Y, Cheng D, Zhou
Q, Zhou J and Lin Q: Hepatitis C virus core upregulates the
methylation status of the RASSF1A promoter through regulation of
SMYD3 in hilar cholangiocarcinoma cells. Acta Biochim Biophys Sin.
43:354–361. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Tsuge M, Hamamoto R, Silva FP, Ohnishi Y,
Chayama K, Kamatani N, Furukawa Y and Nakamura Y: A variable number
of tandem repeats polymorphism in an E2F-1 binding element in the
5′ flanking region of SMYD3 is a risk factor for human cancers. Nat
Genet. 37:1104–1107. 2005. View
Article : Google Scholar : PubMed/NCBI
|
|
14
|
Luo XG, Ding Y, Zhou QF, Ye L, Wang SZ and
Xi T: SET and MYND domain-containing protein 3 decreases
sensitivity to dexamethasone and stimulates cell adhesion and
migration in NIH3T3 cells. J Biosci Bioeng. 103:444–450. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
AL-Eitan LN and Rababa'h DM: Correlation
between a variable number tandem repeat (VNTR) polymorphism in
SMYD3 gene and breast cancer: A genotype-phenotype study. Gene.
728:1442812020. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Vieira FQ, Costa-Pinheiro P, Almeida-Rios
D, Graça I, Monteiro-Reis S, Simões-Sousa S, Carneiro I, Sousa EJ,
Godinho MI, Baltazar F, et al: SMYD3 contributes to a more
aggressive phenotype of prostate cancer and targets Cyclin D2
through H4K20me3. Oncotarget. 6:13644–13657. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Zeng B, Li Z, Chen R, Guo N, Zhou J, Zhou
Q, Lin Q, Cheng D, Liao Q, Zheng L and Gong Y: Epigenetic
regulation of miR-124 by hepatitis C virus core protein promotes
migration and invasion of intrahepatic cholangiocarcinoma cells by
targeting SMYD3. FEBS Lett. 586:3271–3278. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Zhu Y, Zhu MX, Zhang XD, Xu XE, Wu ZY,
Liao LD, Li LY, Xie YM, Wu JY, Zou HY, et al: SMYD3 stimulates EZR
and LOXL2 transcription to enhance proliferation, migration, and
invasion in esophageal squamous cell carcinoma. Hum Pathol.
52:153–163. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Wang SZ, Luo XG, Shen J, Zou JN, Lu YH and
Xi T: Knockdown of SMYD3 by RNA interference inhibits cervical
carcinoma cell growth and invasion in vitro. BMB Rep. 41:294–299.
2008. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Jiang Y, Lyu T, Che X, Jia N, Li Q and
Feng W: Overexpression of SMYD3 in ovarian cancer is associated
with ovarian cancer proliferation and apoptosis via methylating
H3K4 and H4K20. J Cancer. 10:4072–4084. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Wu X, Xu Q, Chen P, Yu C, Ye L, Huang C
and Li T: Effect of SMYD3 on biological behavior and H3K4
methylation in bladder cancer. Cancer Manag Res. 11:8125–8133.
2019. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Wang L, Wang QT, Liu YP, Dong QQ, Hu HJ,
Miao Z, Li S, Liu Y, Zhou H, Zhang TC, et al: ATM signaling pathway
is implicated in the SMYD3-mediated proliferation and migration of
gastric cancer cells. J Gastric Cancer. 17:295–305. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Khatami F and Tavangar SM: Genetic and
epigenetic of medullary thyroid cancer. Iran Biomed J. 22:142–150.
2018.PubMed/NCBI
|
|
24
|
Sponziello M, Durante C, Boichard A, Dima
M, Puppin C, Verrienti A, Tamburrano G, Di Rocco G, Redler A,
Lacroix L, et al: Epigenetic-related gene expression profile in
medullary thyroid cancer revealed the overexpression of the histone
methyltransferases EZH2 and SMYD3 in aggressive tumours. Mol Cell
Endocrinol. 392:8–13. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Lin F, Wu D, Fang D, Chen Y, Zhou H and Ou
C: STAT3-induced SMYD3 transcription enhances chronic lymphocytic
leukemia cell growth in vitro and in vivo. Inflamm Res. 68:739–749.
2019. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Dai B, Wan W, Zhang P, Zhang Y, Pan C,
Meng G, Xiao X, Wu Z, Jia W, Zhang J and Zhang L: SET and MYND
domain-containing protein 3 is overexpressed in human glioma and
contributes to tumorigenicity. Oncol Rep. 34:2722–2730. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Gan L, Yang Y, Li Q, Feng Y, Liu T and Guo
W: Epigenetic regulation of cancer progression by EZH2: From
biological insights to therapeutic potential. Biomark Res.
6:102018. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Kim KH and Roberts CW: Targeting EZH2 in
cancer. Nat Med. 22:128–134. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Bae WK and Hennighausen L: Canonical and
non-canonical roles of the histone methyltransferase EZH2 in
mammary development and cancer. Mol Cell Endocrinol. 382:593–597.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Shen X, Liu Y, Hsu YJ, Fujiwara Y, Kim J,
Mao X, Yuan GC and Orkin SH: EZH1 mediates methylation on histone
H3 lysine 27 and complements EZH2 in maintaining stem cell identity
and executing pluripotency. Mol Cell. 32:491–502. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Ezhkova E, Lien WH, Stokes N, Pasolli HA,
Silva JM and Fuchs E: EZH1 and EZH2 cogovern histone H3K27
trimethylation and are essential for hair follicle homeostasis and
wound repair. Genes Dev. 25:485–498. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Pal B, Bouras T, Shi W, Vaillant F,
Sheridan JM, Fu N, Breslin K, Jiang K, Ritchie ME, Young M, et al:
Global changes in the mammary epigenome are induced by hormonal
cues and coordinated by Ezh2. Cell Rep. 3:411–426. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Laible G, Wolf A, Dorn R, Reuter G, Nislow
C, Lebersorger A, Popkin D, Pillus L and Jenuwein T: Mammalian
homologues of the Polycomb-group gene Enhancer of zeste mediate
gene silencing in Drosophila heterochromatin and at S. cerevisiae
telomeres. EMBO J. 16:3219–3232. 1997. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Masudo K, Suganuma N, Nakayama H, Oshima
T, Rino Y, Iwasaki H, Matsuzu K, Sugino K, Ito K, Kondo T, et al:
EZH2 overexpression as a useful prognostic marker for aggressive
Behaviour in thyroid cancer. In vivo. 32:25–31. 2018.PubMed/NCBI
|
|
35
|
Xue L, Yan H, Chen Y, Zhang Q, Xie X, Ding
X, Wang X, Qian Z, Xiao F, Song Z, et al: EZH2 upregulation by ERα
induces proliferation and migration of papillary thyroid carcinoma.
BMC cancer. 19:10942019. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Andrusiewicz M, Slowikowski B, Skibinska
I, Wolun-Cholewa M and Dera-Szymanowska A: Selection of reliable
reference genes in eutopic and ectopic endometrium for quantitative
expression studies. Biomed Pharmacother. 78:66–73. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Haddad RI, Nasr C, Bischoff L, Busaidy NL,
Byrd D, Callender G, Dickson P, Duh QY, Ehya H, Goldner W, et al:
NCCN guidelines insights: Thyroid carcinoma, version 2.2018. J Natl
Compr Canc Netw. 16:1429–1440. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Wang LJ and Cai HQ: miR-1258: A novel
microRNA that controls TMPRSS4 expression is associated with
malignant progression of papillary thyroid carcinoma. Endokrynol
Pol. 71:146–152. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Zhou Q, Chen J, Feng J and Wang J: Long
noncoding RNA PVT1 modulates thyroid cancer cell proliferation by
recruiting EZH2 and regulating thyroid-stimulating hormone receptor
(TSHR). Tumour Biol. 37:3105–3113. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Hossain MA, Asa TA, Rahman MM, Uddin S,
Moustafa AA, Quinn JMW and Moni MA: Network-based genetic profiling
reveals cellular pathway differences between follicular thyroid
carcinoma and follicular thyroid adenoma. Int J Environ Res Public
Health. 17:13732020. View Article : Google Scholar
|
|
41
|
Fabini E, Manoni E, Ferroni C, Rio AD and
Bartolini M: Small-molecule inhibitors of lysine methyltransferases
SMYD2 and SMYD3: Current trends. Future Med Chem. 11:901–921. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Roman BR, Morris LG and Davies L: The
thyroid cancer epidemic, 2017 perspective. Curr Opin Endocrinol
Diabetes Obes. 24:332–336. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Al-Zahrani AS and Ravichandran K:
Epidemiology of thyroid cancer: A review with special reference to
Gulf cooperation council (GCC) states. Gulf J Oncolog. 17–28.
2007.PubMed/NCBI
|
|
44
|
Kinoshita Y, Takasu K, Yuri T, Yoshizawa
K, Emoto Y, Tsubura A and Shikata N: Estrogen receptor-and
progesterone receptor-positive diffuse sclerosing variant of
papillary thyroid carcinoma: A case report. Case Rep Oncol.
6:216–223. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Huang Y, Dong W, Li J, Zhang H, Shan Z and
Teng W: Differential expression patterns and clinical significance
of estrogen receptor-alpha and beta in papillary thyroid carcinoma.
BMC cancer. 14:3832014. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Lu J, Gao J, Zhang J, Sun J, Wu H, Shi X,
Teng L and Liang Z: Association between BRAF V600E mutation and
regional lymph node metastasis in papillary thyroid carcinoma. Int
J Clin Exp Pathol. 8:793–799. 2015.PubMed/NCBI
|