|
1
|
Bray F, Ferlay J, Soerjomataram I, Siegel
RL, Torre LA and Jemal A: Global cancer statistics 2018: GLOBOCAN
estimates of incidence and mortality worldwide for 36 cancers in
185 countries. CA Cancer J Clin. 68:394–424. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Global Burden of Disease Cancer
Collaboration, ; Fitzmaurice C, Dicker D, Pain A, Hamavid H,
Moradi-Lakeh M, MacIntyre MF, Allen C, Hansen G, Woodbrook R, et
al: The global burden of cancer 2013. JAMA Oncol. 1:505–527. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
International Agency for Research on
Cancer (IARC), . Cancer Fact Sheets, Digestive organs. IARC; Lyon:
2020, https://gco.iarc.fr/today/fact-sheets-cancerDecember.
2020
|
|
4
|
National Cancer Institute (NIH), . Cancer
Stat Facts: Stomach cancer. NIH; Bethesda, MD: 2021, https://seer.cancer.gov/statfacts/html/stomach.html
|
|
5
|
Bevan S and Houlston RS: Genetic
predisposition to gastric cancer. QJM. 92:5–10. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Ushijima T and Sasako M: Focus on gastric
cancer. Cancer Cell. 5:121–125. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Wang P, Wang Y, Hang B, Zou X and Mao JH:
A novel gene expression-based prognostic scoring system to predict
survival in gastric cancer. Oncotarget. 7:55343–55351. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Cui J, Li F, Wang G, Fang X, Puett JD and
Xu Y: Gene-expression signatures can distinguish gastric cancer
grades and stages. PLoS One. 6:e178192011. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Mattioni M, Soddu S, Porrello A,
D'Alessandro R, Spila A and Guadagni F: Serum anti-p53 antibodies
as a useful marker for prognosis of gastric carcinoma. Int J Biol
Markers. 22:302–306. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Takeno A, Takemasa I, Doki Y, Yamasaki M,
Miyata H, Takiguchi S, Fujiwara Y, Matsubara K and Monden M:
Integrative approach for differentially overexpressed genes in
gastric cancer by combining large-scale gene expression profiling
and network analysis. Br J Cancer. 99:1307–1315. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Heuser VD, Kiviniemi A, Lehtinen L, Munthe
S, Kristensen BW, Posti JP, Sipilä JOT, Vuorinen V, Carpén O and
Gardberg M: Multiple formin proteins participate in glioblastoma
migration. BMC Cancer. 20:7102020. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Zhao S, Cai J, Zhang X, Cui J and Jiu Y:
Different formins restrict localization of distinct tropomyosins on
dorsal stress fibers in osteosarcoma cells. Cytoskeleton (Hoboken).
77:16–24. 2020. View
Article : Google Scholar : PubMed/NCBI
|
|
13
|
Alvarez DE and Agaisse H: A role for the
small GTPase Rac1 in vaccinia actin-based motility. Small GTPases.
6:119–122. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Schulte A, Stolp B, Schönichen A,
Pylypenko O, Rak A, Fackler OT and Geyer M: The human formin FHOD1
contains a bipartite structure of FH3 and GTPase-binding domains
required for activation. Structure. 16:1313–1323. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Heuser VD, Mansuri N, Mogg J, Kurki S,
Repo H, Kronqvist P, Carpén O and Gardberg M: Formin proteins FHOD1
and INF2 in triple-negative breast cancer: Association with basal
markers and functional activities. Breast Cancer (Auckl).
12:11782234187922472018.PubMed/NCBI
|
|
16
|
Sun BO, Fang Y, Li Z, Chen Z and Xiang J:
Role of cellular cytoskeleton in epithelial-mesenchymal transition
process during cancer progression. Biomed Rep. 3:603–610. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Li DJ, Feng ZC, Li XR and Hu G:
Involvement of methylation-associated silencing of formin 2 in
colorectal carcinogenesis. World J Gastroenterol. 24:5013–5024.
2018. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Zhu L, Wang H, Jiang C, Li W, Zhai S, Cai
X, Wang X, Liao L, Tao F, Jin D, et al: Clinically applicable
53-Gene prognostic assay predicts chemotherapy benefit in gastric
cancer: A multicenter study. EBioMedicine. 61:1030232020.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
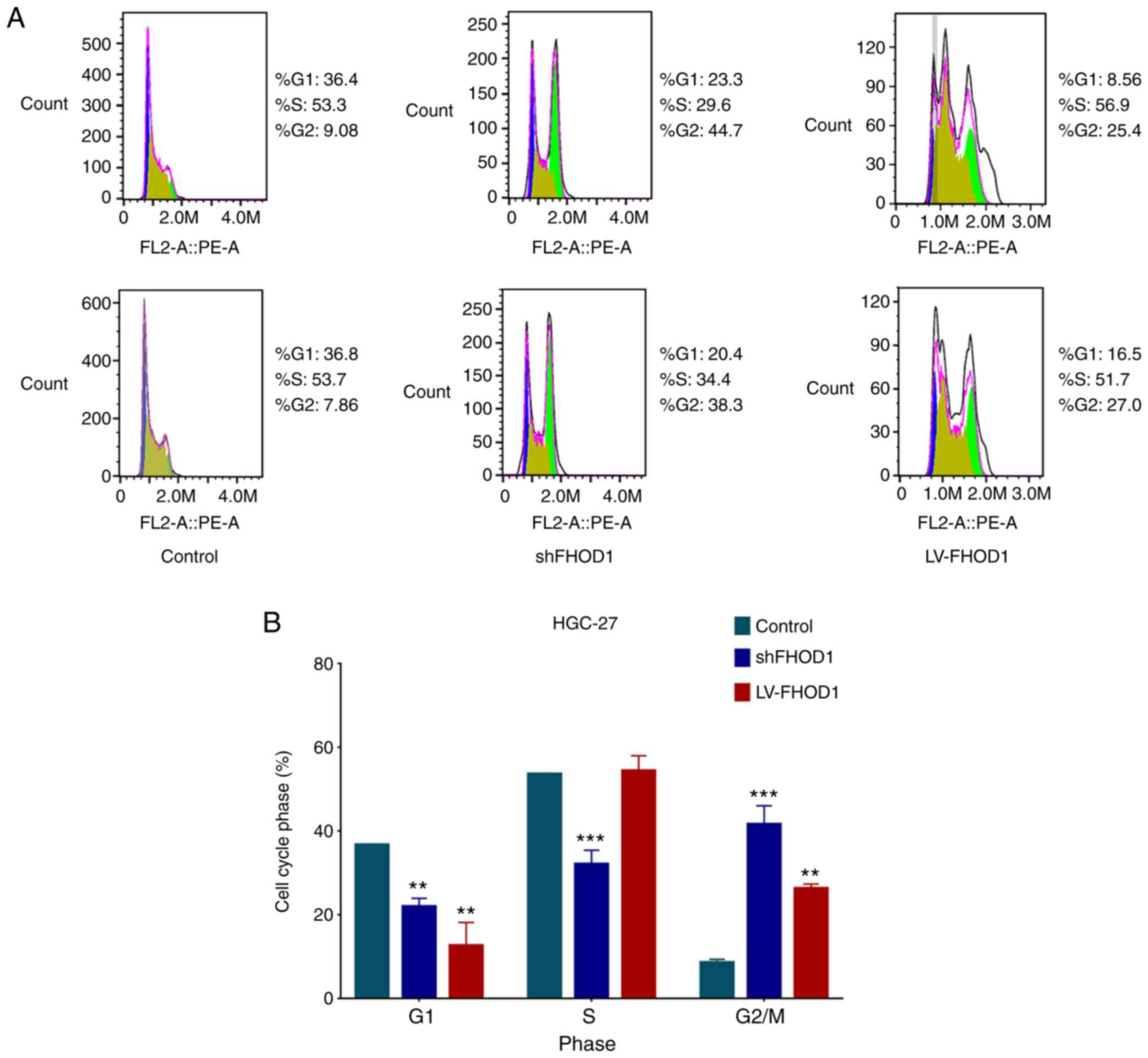
20
|
Bertoli C, Skotheim JM and de Bruin RA:
Control of cell cycle transcription during G1 and S phases. Nat Rev
Mol Cell Biol. 14:518–528. 2013. View
Article : Google Scholar : PubMed/NCBI
|
|
21
|
Zhang H, Wang Y, Wang Y, Wu D, Lin E and
Xia Q: Intratumoral and intertumoral heterogeneity of HER2
immunohistochemical expression in gastric cancer. Pathol Res Pract.
216:1532292020. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Dai J, Peng T and Yu X: NK6 homeobox 2
regulated gastrokin-2 suppresses gastric cancer cell proliferation
and invasion via Akt signaling pathway. Cell Biochem Biophys.
79:123–131. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Han X, Zhang HB, Li XD and Wang ZA: Long
non-coding RNA X-inactive-specific transcript contributes to
cisplatin resistance in gastric cancer by sponging miR-let-7b.
Anticancer Drugs. 31:1018–1025. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Gao N, Yang F, Chen S, Wan H, Zhao X and
Dong H: The role of TRPV1 ion channels in the suppression of
gastric cancer development. J Exp Clin Cancer Res. 39:2062020.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Li Z, Gao H, Liu Y, Wu H, Li W, Xing Y,
Zhang Z and Zhang X: Genetic variants in the regulation region of
TLR4 reduce the gastric cancer susceptibility. Gene.
767:1451812021. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Jurmeister S, Baumann M, Balwierz A,
Keklikoglou I, Ward A, Uhlmann S, Zhang JD, Wiemann S and Sahin Ö:
MicroRNA-200c represses migration and invasion of breast cancer
cells by targeting actin-regulatory proteins FHOD1 and PPM1F. Mol
Cell Biol. 32:633–651. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Gasteier JE, Madrid R, Krautkrämer E,
Schröder S, Muranyi W, Benichou S and Fackler OT: Activation of the
Rac-binding partner FHOD1 induces actin stress fibers via a
ROCK-dependent mechanism. J Biol Chem. 278:38902–38912. 2003.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Gardberg M, Kaipio K, Lehtinen L, Mikkonen
P, Heuser VD, Talvinen K, Iljin K, Kampf C, Uhlen M, Grénman R, et
al: FHOD1, a formin upregulated in epithelial-mesenchymal
transition, participates in cancer cell migration and invasion.
PLoS One. 8:e749232013. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Haraguchi T, Kondo M, Uchikawa R,
Kobayashi K, Hiramatsu H, Kobayashi K, Chit UW, Shimizu T and Iba
H: Dynamics and plasticity of the epithelial to mesenchymal
transition induced by miR-200 family inhibition. Sci Rep.
6:211172016. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Perdigão-Henriques R, Petrocca F,
Altschuler G, Thomas MP, Le MT, Tan SM, Hide W and Lieberman J:
miR-200 promotes the mesenchymal to epithelial transition by
suppressing multiple members of the Zeb2 and Snail1 transcriptional
repressor complexes. Oncogene. 35:158–172. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Peippo M, Gardberg M, Lamminen T, Kaipio
K, Carpén O and Heuser VD: FHOD1 formin is upregulated in melanomas
and modifies proliferation and tumor growth. Exp Cell Res.
350:267–278. 2017. View Article : Google Scholar : PubMed/NCBI
|