Introduction
Bladder cancer (BLCA) is the tenth most common
cancer type worldwide, with an estimated 549,000 new cases and
200,000 deaths each year according to the Global Cancer Statistics
2018 (1). BLCA is more common in men
than in women, with the morbidity and mortality rate in men being
four times higher than that in women worldwide (1). Transitional cell carcinoma is the most
common pathological subtype of BLCA, accounting for ~90% of all
BLCA cases (2). Squamous cell
carcinoma and adenocarcinoma are ranked second and third,
respectively (2). BLCA can be
divided into two types according to degree of invasiveness:
Non-muscle invasive BLCA (NMIBC) and muscle invasive BLCA (MIBC)
(3). NMIBC does not pose a major
threat to the lives of the patients; however, NMIBC has a high
recurrence rate, with a small number of patients progressing to
invasive BLCA (2,4). MIBC is invasive and has a high
mortality rate (4). Therefore, it is
necessary to identify novel molecular targets for the diagnosis and
treatment of BLCA.
Protein glycosylation is a common post-translational
modification regulated by glycosyltransferases and glycosidases
present in the endoplasmic reticulum-Golgi complex (5). The two most common types of
glycosylation are O-linked glycosylation and N-linked glycosylation
(6). Changes in the degree of
glycosylation are caused by the overexpression of glycoproteins
carrying certain carbohydrate chains, an increase or decrease in
the levels of nucleotide donors, and changes in the expression of
glycosyltransferases and glycosidases (7). Glycosylation serves a key role in a
number of important biological processes of tumors, including cell
adhesion, metastasis, cell signal transduction and cell metabolism
(8,9). Sialylation is one of the important
pathways of glycosylation modification and is a typical protein
terminal modification that mediates important biological functions
(10). It has been demonstrated that
abnormal sialylation is closely associated with malignant brain
tumors and multiple myeloma, and is usually accompanied by changes
in the expression levels of sialyl transferases (STs) (11,12).
The human ST family consists of 20 STs that catalyze
the binding of sialic acid of CMP-Neu5Ac to the terminal glycosyl
groups of the glycoconjugates (13–15).
According to the different types of glycosidic bonds formed,
salivary STs are further divided into 3 types: ST3 β-galactoside
α-2,3-ST (ST3GAL)1-6, ST6 β-galactoside α-2,6-ST1-2 and α-2,8-ST1
(ST8SIA1)-6 (16). ST8SIA1 is a
member of the ST family involved in the production of gangliosides
(17). The abnormal expression of
ST8SIA1 has been demonstrated to influence the metastasis of
triple-negative breast cancer (18);
however, to the best of our knowledge, the role of this enzyme in
BLCA remains unknown.
In the present study, differentially expressed
glycosyltransferases and their expression profile were evaluated
based on the data obtained from The Cancer Genome Atlas (TCGA) and
Gene Expression Profiling Interactive Analysis (GEPIA) databases.
The subsequent experiments performed in the present study aimed to
investigate the association between ST8SIA1 expression and the
clinicopathological characteristics of patients with BLCA, as well
as to reveal the effect and regulatory mechanism of ST8SIA1 on the
proliferation, migration and invasion of BLCA cells. The
downregulation and antitumor role of ST8SIA1 in BLCA may provide a
new strategy to diagnose and treat BLCA.
Materials and methods
Patient samples
A total of 136 BLCA tissue and adjacent normal
tissue samples (>5 cm away from the tumor edge) were collected
from randomly selected patients undergoing surgical resection
between May 2008 and July 2013 at The First Affiliated Hospital of
Dalian Medical University (Dalian, China). After surgical
resection, all samples were routinely processed and
paraffin-embedded. The inclusion criterion was primary and
metastatic BLCA tissues of patients. Tissue specimens from patients
who received radiotherapy or chemotherapy before surgical resection
were excluded. The age range of the patients was 34–82 years with a
mean age of 58.8 years. The ratio of male to female was 97:39.
Tumors were staged according to the 7th Edition of the American
Joint Committee on Cancer (19). Of
the 136 BLCA tissue, 39 were determined as early stage (I/II) and
97 as late stage (III/IV). The present study was approved by the
Institutional Research Ethics Committee of the First Affiliated
Hospital of Dalian Medical University (approval no. LCKY2015-08;
Dalian, China), and written informed consent was provided by the
patients with BLCA who underwent surgery.
Cell culture
SV-HUC-1 normal bladder epithelial cell, 5637 and
T24 BLCA cell lines were purchased from The Cell Bank of Type
Culture Collection of The Chinese Academy of Sciences. SV-HUC-1
normal bladder epithelial cells were cultured in F12/DMEM (Thermo
Fisher Scientific, Inc.) supplemented with 10% FBS (Thermo Fisher
Scientific, Inc.). The 5637 and T24 BLCA cell lines were cultured
inRPMI-1640 medium (Thermo Fisher Scientific, Inc.) supplemented
with 10% FBS (Thermo Fisher Scientific, Inc.). All cell cultures
were supplemented with 1% penicillin/streptomycin, and cells were
incubated with 5% CO2 at 37°C.
Identification of differentially
expressed glycosyltransferases
TCGA (https://cancergenome.nih.gov/; Project ID:TCGA-BLCA)
project provided mRNA expression profile data on 408 BLCA cases:
433 samples, including 19 normal and 414 BLCA samples. Some
patients provided both cancer tissue and adjacent normal tissue and
some patients provided >2 cancer tissues. Therefore, the number
of patient cases was less than the tissue samples obtained from the
TCGA database. The ‘edgeR’ package (http://www.bioconductor.org/help/search/index.html?q=edger/;
release number 3.13; Bioconductor) in R software (https://www.r-project.org/; release number 3.6.1) was
used to identify the differentially expressed mRNAs of
glycosyltransferases in BLCA tissues compared with normal bladder
tissues. The following thresholds were used to identify significant
differentially expressed genes: False discovery rate (FDR)<0.001
and |log2(fold change)|>2. After screening of
differentially expressed genes, the differentially expressed gene
ID were obtained. The gene ID were annotated using ENSEMBL
(https://www.ensembl.org/) to get the
differentially expressed gene symbol. Then, the expression levels
of ST8SIA1 in BLCA and normal tissue were assessed based on the
data obtained from GEPIA (http://gepia.cancer-pku.cn/).
Immunohistochemical staining
BLCA and adjacent normal tissue were fixed with 4%
paraformaldehyde at room temperature for 24 h. After paraffin
embedding, 4 µm-thick sections were cut and mounted onto slides.
Next, slides were deparaffinized with xylene, rehydrated by a
series of graded alcohol solutions as follows: 100, 100, 95, 85 and
75%. The antigen was retrieved with 10 mM sodium citrate buffer at
95°C for 20 mi and immersed in 3% hydrogen peroxide in methanol at
room temperature for 15 min to block the endogenous peroxidase
activity. The slides were washed with PBS twice, blocked with 5%
goat serum (Beijing Zhong Shan-Golden Bridge Biological Technology
Co., Ltd.) at room temperature for 20 min and incubated with a
primary anti-ST8SIA1 antibody (1:80; cat. no. 24918-1-AP;
ProteinTech Group, Inc.) at room temperature for 1 h. Following
washing with PBS, the PBS surrounding the tissue was wiped dry and
the samples were incubated with a secondary horseradish peroxidase
(HRP)-conjugated IgG antibody (1:100; cat. no. sc-2357; Santa Cruz
Biotechnology, Inc.) at room temperature for 45 min. The sections
were then treated with 3,3′-diaminobenzidine at room temperature
for 30 sec, counterstained with hematoxylin at room temperature for
5 min, dried, sealed with a neutral gum and observed under a light
microscope. The immunostaining intensity was then evaluated. The
immunostaining for ST8SIA1 expression was scored using ‘−’, ‘+’,
‘++’ and ‘+++’, to indicate a positive staining percentage of
<5, 5–25, 25–50 and >50%, respectively. All specimens were
examined by 2 pathologists of The First Affiliated Hospital of
Dalian Medical University (Dalian, China), who were blinded to the
clinical data.
Cell transfection
T24 and 5637 cells were transfected with the
recombinant pcDNA3.1/ST8SIA1 or mock vectors. Non-transfected and
mock vector transfected cells were used as the control. The
pcDNA3.1/ST8SIA1 and mock vectors were synthesized by Public
Protein/Plasmid Library (http://www.geneppl.com/index.php; pcDNA3.1/ST8SIA1,
cat. no. BC046158; mock vector, cat. no. V790-20). Cells were
transfected with a mixture of 5 µg plasmids and 10 µl
Lipofectamine® 2000 transfection reagent (Thermo Fisher
Scientific, Inc.) at 37°C for 7 h. To select a stable population of
transfected cells, the culture medium was replaced with complete
medium containing 600 mg/ml G418 (Merck KGaA) after 48 h. After 2
months of screening with 300 mg/ml G418, two cell lines stably
expressing ST8SIA1 (ST8SIA1-1 and ST8SIA1-2) were established. The
modified expression was confirmed by reverse
transcription-quantitative PCR (RT-qPCR) and western blotting.
RT-qPCR
Total RNA was extracted from the tissue samples and
cultured cells using TRIzol® reagent (Thermo Fisher
Scientific, Inc.) according to the manufacturer's protocol. RNA was
subjected to reverse transcription to generate cDNA using a
PrimeScript RT reagent kit (Takara Bio, Inc.) for 2 min at 42°C, 15
min at 37°C and 5 sec at 85°C and stored at −20°C and amplified by
subsequent qPCR using a qPCR SYBR GreenSuperMix (TransGen Biotech
Co., Ltd.). The amplification conditions were as follows: 94°C for
30 sec, followed by 94°C for 5 sec and 60°C for 34 sec for 40
cycles. Relative changes in gene expression were analyzed using the
2−ΔΔCq method (20).
GAPDH was used as an internal control. Specific primers for GAPDH
and ST8SIA1 were purchased from Shanghai GenePharma Co., Ltd. The
primer sequences used were as follows: ST8SIA1 forward,
5′-CATTGAAGAAATGCGCGGTGG-3′ and reverse,
5′-GTTCTGAAACCTTTGCCGAATTATG-3′; GAPDH forward,
5′-TCCAAAATCAAGTGGGGCGA-3′ and reverse,
5′-AAATGAGCCCCAGCCTTCTC-3′.
Western blot analysis
Total protein was extracted from the cultured cells
and specimens using RIPA buffer (Beyotime Institute of
Biotechnology) and PMSF (dilution, 1:100; Beyotime Institute of
Biotechnology). A bicinchoninic acid assay kit (Beyotime Institute
of Biotechnology) was used for quantification. Equal amounts of
protein (30 µg/lane) were separated using 12% SDS-PAGE and
transferred onto PVDF membranes (Pall Life Sciences). The membranes
were blocked at room temperature for 2 h with 5% fat-free milk
prepared in 1XTris-Buffered Saline, 0.1% Tween® 20
Detergent (TBST). Western blot analysis was performed using
phosphorylated (p-)STAT3 (dilution, 1:500; cat. no. 39596;
ProteinTech Group, Inc.), STAT3 (dilution, 1:500; cat. no. AP0366;
Bioworld Technology, Inc.), ST8SIA1 (dilution, 1:500; cat. no.
24918-1-AP; ProteinTech Group, Inc.), Janus kinase 2 (JAK2;
dilution, 1:500; cat. no. 17670-1-AP; ProteinTech Group, Inc.),
p-JAK2 (dilution, 1:500; cat. no. AP0531; ABclonal Biotech Co.,
Ltd.), cyclin D1 (dilution, 1:500; cat. no. 26939-1-AP; ProteinTech
Group, Inc.), MMP2 (dilution, 1:500; cat. no. 10373-2-AP;
ProteinTech Group, Inc.), proliferating cell nuclear antigen (PCNA;
dilution, 1:500; cat. no. 10205-2-AP; ProteinTech Group, Inc.),
Bcl2 (dilution, 1:500; cat. no. 12789-1-AP; ProteinTech Group,
Inc.) and GAPDH (dilution, 1:500; cat. no. 10494-1-AP; ProteinTech
Group, Inc.) antibodies. The membranes were incubated with primary
antibodies at 4°C overnight. After three washes with TBST, the
membranes were incubated with a horseradish peroxidase-conjugated
goat anti-rabbit secondary antibody (dilution, 1:5,000; cat. no.
SA00001-2; ProteinTech Group, Inc.) at room temperature for 1 h.
The immunoreactivity was detected using an electrochemiluminescence
kit (Advansta, Inc.) and processed using Image Lab software v.5.2
(Bio-Rad Laboratories, Inc.).
Cell counting Kit-8 (CCK-8)
Cell proliferation was determined using aCCK-8 assay
(Dojindo Molecular Technologies, Inc.). BLCA cells were seeded into
a 96-well plate at a density of 5×103 cells per well in
triplicate. Cells were incubated at 37°C for 12, 24, 36 or 48 h,
and 10 µl CCK-8 reagent was added into each well. Cells were
incubated at 37°C with 5% CO2 for 2 h. A microplate
reader (Thermo Fisher Scientific, Inc.) was used to measure the
optical density values at 450 nm.
Cell colony formation assay
BLCA cells in the logarithmic growth phase were
digested into a single-cell suspension. Cells were seeded into
6-well culture plates (Corning, Inc.) at a density of 200 cells per
well and incubated at 37°C with 5% CO2 for 14 days.
Following three washes with PBS, cells were fixed with
paraformaldehyde (4%) at room temperature for 15 min and stained
with 0.1% crystal violet at room temperature for 30 min. The
colonies were imaged using a light optical microscope and the
number of colonies of >50 cells was counted using ImageJ
software v.4.0 (National Institutes of Health).
Flow cytometry
T24 and 5637 cells were collected as a single cell
suspension and fixed with 75% alcohol at 4°C overnight. The cells
were centrifuged for 5 min at 1,000 × g at room temperature and
then washed with ice-cold PBS. Cells were stained by incubation
with propidium iodide (50 µg/ml; Beyotime Institute of
Biotechnology) combined with RNase A (50 µg/ml; Beyotime Institute
of Biotechnology) for 30 min at 37°C. Cells were detected using a
flow cytometer (Accuri C6 Cytometer; BD Biosciences) and then
analyzed with FlowJo software v.10.2 (TreeStar, Inc.). Signals were
acquired from at least 1×106 cells per sample.
Wound healing assay
Cell migration was analyzed using a wound healing
assay. Non-transfected T24 or 5637, mock, ST8SIA1-1 and ST8SIA1-2
cells (2.5×106) were respectively transferred to a
six-well plate and allowed to grow to 100% confluence in complete
medium. A 200-µl sterile pipette tip was used to scratch a gap
through the monolayer. The cellular debris was dislodged by washing
with PBS thrice. The cells were then washed and imaged under a
light microscope at a magnification of ×10. Following cell culture
with the serum-free medium for 12 h, the same areas were imaged
again. Cell mobility was assessed by calculating the remaining open
area of the wound using ImageJ software v.4.0 (National Institutes
of Health) and analyzed with the following formula: (Width 0
h-Width 12 h)/Width 0 h ×100%.
Transwell migration and invasion
assay
A total of ~2×104 cells in 150 µl
RPMI-1640 medium without FBS were added to the upper compartment of
a 24-well Transwell chamber (pore size, 8.0 µm; diameter, 6.5 mm;
Corning, Inc.), and 500 µl RPMI-1640 medium with 20% FBS was added
to the lower compartment. Following incubation at 37°C for 48 h in
a 5% CO2 incubator, the cells on the upper surface of
the filter were completely removed using a cotton swab. The 24-well
Transwell chambers were coated with 40 µl Matrigel® for
30 min at 37°C on the upward-facing side of the polycarbonate
membrane for the invasion assay. Subsequently, 4×104
cells in 150 µl serum-free medium were added to the upper chamber
and 500 µl medium containing 20% FBS was added into the lower
chambers. After incubation for 48 h at 37°C, the cells were fixed
with 4% paraformaldehyde at room temperature for 15 min and stained
with 0.1% crystal violet at room temperature for 10 min. The cells
were imaged using a light microscope and counted using ImageJ
software v.4.0 (National Institutes of Health).
Statistical analysis
All experiments were repeated at least three times.
Quantitative data are presented as the mean ± SD. Statistical
analysis was performed using GraphPad Prism 5.0 software (GraphPad
Software, Inc.). Differences between two groups were analyzed using
Student's unpaired t-test, whereas those among multiple groups were
analyzed using one-way ANOVA followed by the post hoc Tukey's test.
The χ2 test was performed for the comparisons of
categorical variables. P<0.05 was considered to indicate a
statistically significant difference.
Results
Identification of differentially
expressed STs in BLCA and normal tissues using TCGA and GEPIA
databases
To identify the STs associated with the malignant
progression of BLCA, TCGA (https://cancergenome.nih.gov/) was utilized to analyze
the differentially expressed glycosyltransferse genes between 414
BLCA and 19 normal tissue samples. Using FDR<0.001 and
|log2(fold change)|>2 as the selection criteria for
the standardized data, 13 glycosyltransferases, including only 1 ST
(ST8SIA1) with low expression levels in BLCA, were identified
(Fig. 1A and B). The expression
levels of ST8SIA1 were further assessed based on the data obtained
from GEPIA (http://gepia.cancer-pku.cn/). Decreased ST8SIA1
expression was identified in BLCA tissues compared with normal
tissues (Fig. 1C). These results
suggested that ST8SIA1 may serve an important role in the
development of BLCA.
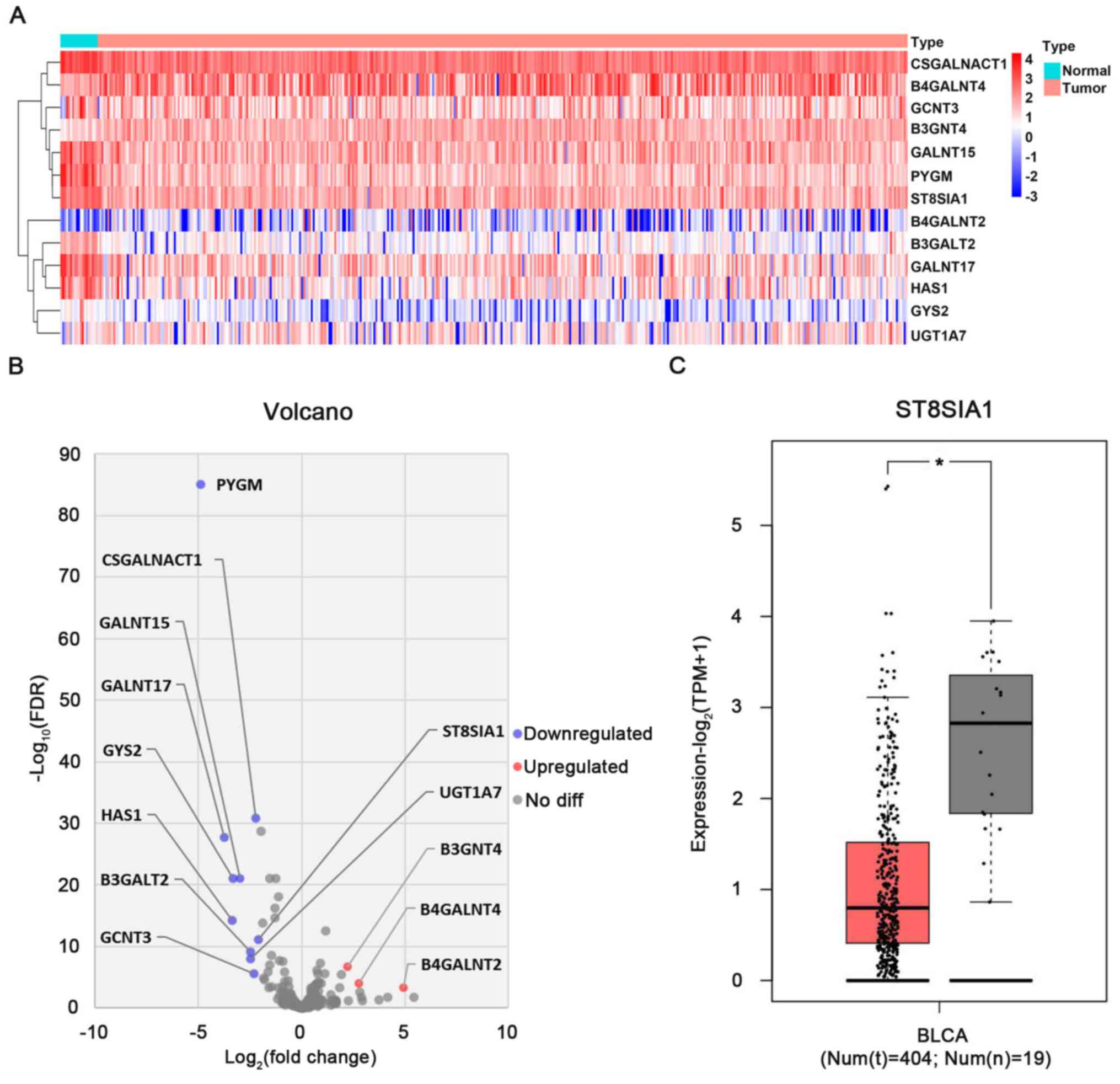 | Figure 1.Identification of differentially
expressed sialyl transferases in BLCA and normal tissues using TCGA
and GEPIA databases. (A) Heatmap analysis of differentially
expressed glycosyltransferase genes in TCGA between 19 normal and
414 BLCA tumor tissues. The red-to-blue scale is defined by
Log10(X+0.001) (X: The amount of gene expression for
counts data). (B) Volcano plot analysis of differentially expressed
glycosyltransferase genes in TCGA database between 19 normal and
414 BLCA tumor tissues. |log2(fold change)|>2 and
FDR<0.001 were set as the screening criteria. The blue and red
dots represent the downregulated and upregulated genes in BLCA
tumor tissues, and the grey dots represent genes that did not
satisfy the screening criteria. (C) Differential expression of the
ST8SIA1 gene in patients with BLCA was identified using data from
the GEPIA database. *P<0.05. BLCA, bladder cancer; FDR, false
discovery rate; GEPIA, Gene Expression Profiling Interactive
Analysis; Num(n), number of normal tissue samples; Num(t), number
of tumor tissue samples; ST8SIA1, α-2,8-sialyltransferase1; TCGA,
The Cancer Genome Atlas; TPM, transcripts per million. |
ST8SIA1 expression is markedly
downregulated in BLCA tissues and cells, and negatively associated
with the pathological grade and invasiveness in patients with
BLCA
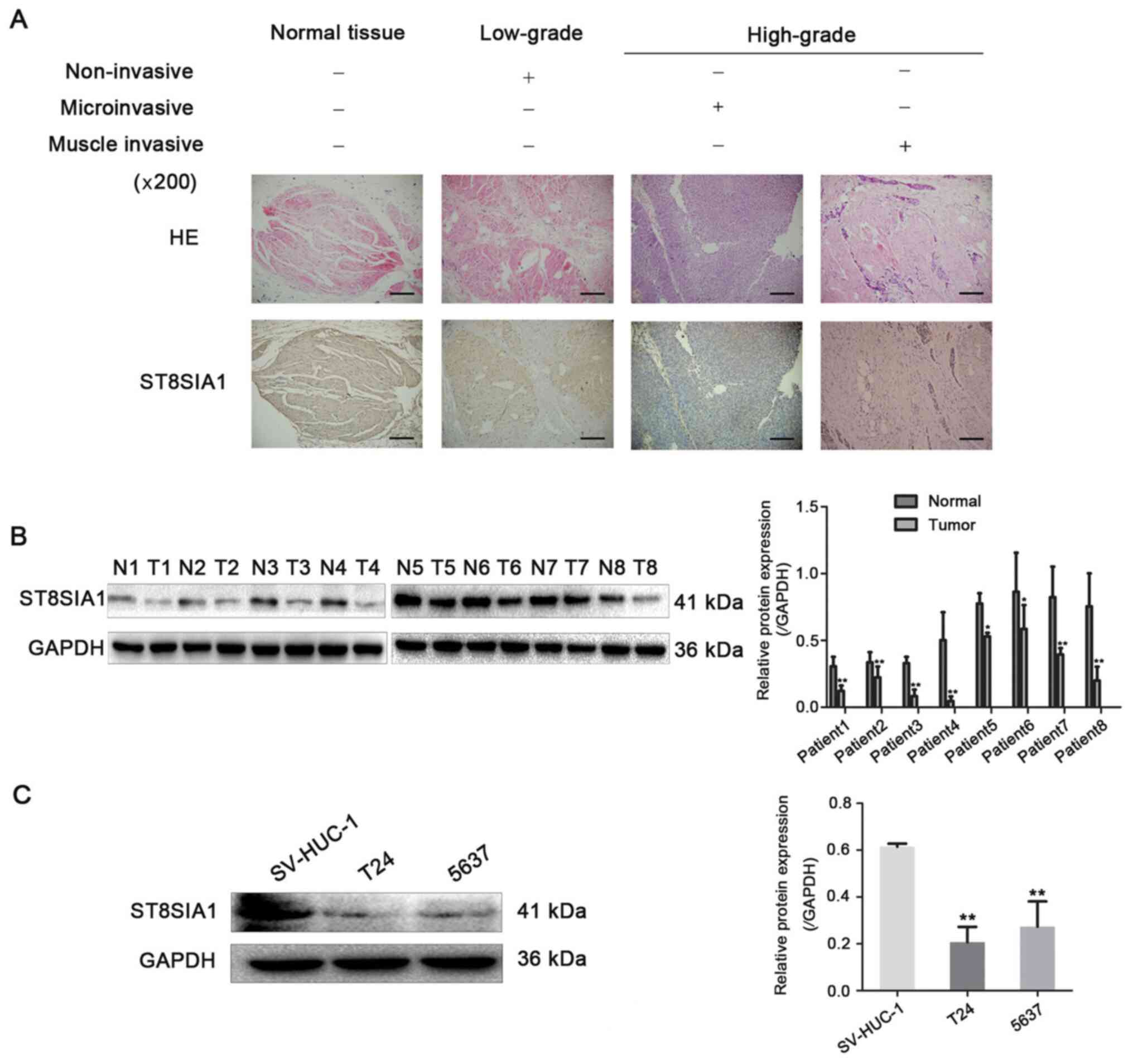
To validate the results obtained using TCGA,
immunohistochemical staining and western blot analysis were used to
detect the expression levels of ST8SIA1 in BLCA. The data indicated
that ST8SIA1 protein showed cytoplasm staining and the staining
intensity of ST8SIA1 in high-grade cancer tissues was lower
compared with that in low-grade cancer and adjacent normal tissues
(Fig. 2A). The expression levels of
ST8SIA1 in 8 pairs of BLCA and adjacent normal tissues were
detected by western blot analysis, and the results revealed that
the protein expression levels of ST8SIA1 were significantly reduced
in the tumor tissue samples (Fig.
2B). Furthermore, ST8SIA1 expression was detected in two BLCA
cell lines (5637 and T24) and a normal bladder epithelial cell line
(SV-HUC-1). The relative protein expression levels of ST8SIA1 were
significantly lower in the BLCA cell lines compared with the normal
bladder epithelial cell line (Fig.
2C). The association between ST8SIA1 protein expression and
major clinicopathological characteristics was further analyzed in
136 patients with BLCA. ST8SIA1 expression was negatively
associated with tumor grade and invasiveness (Table I). These data indicated that ST8SIA1
may act as a tumor suppressor in BLCA progression.
 | Table I.Association between ST8SIA1
expression and clinicopathological characteristics of patients with
bladder cancer. |
Table I.
Association between ST8SIA1
expression and clinicopathological characteristics of patients with
bladder cancer.
|
|
| ST8SIA1
expression |
|
|
|---|
|
|
|
|
|
|
|---|
| Variables | No. | −/+, n | ++/+++, n | χ2 | P-value |
|---|
| Age, years |
|
|
|
|
|
|
≤50 | 51 | 36 | 15 | 0.323 | 0.570 |
|
>50 | 85 | 56 | 29 |
|
|
| Sex |
|
|
|
|
|
|
Female | 39 | 25 | 14 | 0.314 | 0.575 |
|
Male | 97 | 67 | 30 |
|
|
| Grade |
|
|
| 61.710 | <0.01 |
| Low
(I/II) | 39 | 7 | 32 |
|
|
| High
(III/IV) | 97 | 85 | 12 |
|
|
| Invasion |
|
|
| 49.041 | <0.01 |
|
Non-invasive | 38 | 9 | 29 |
|
|
|
Microinvasive | 25 | 18 | 7 |
|
|
| Muscle
invasive | 73 | 65 | 8 |
|
|
ST8SIA1 inhibits the proliferation of
5637 and T24 cells
To explore the roles of ST8SIA1 in the malignant
progression of BLCA cells, T24 and 5637 cell lines stably
overexpressing ST8SIA1 (ST8SIA1-1 and ST8SIA1-2) were established.
RT-qPCR and western blot analysis demonstrated that T24 (Fig. 3A and B, top panel) and 5637 (Fig. 3A and B, bottom panel) cell lines with
a stable overexpression of ST8SIA1 were successfully constructed
using molecular cloning and liposome transfection technology. To
investigate the effects of ST8SIA1 upregulation on the
proliferation and colony formation of BLCA cells, CCK-8, colony
formation and flow cytometry assays were performed. The CCK-8 assay
revealed that the proliferation of T24 (Fig. 3C, left panel) and 5637 (Fig. 3C, right panel) cells stably
overexpressing ST8SIA1 (ST8SIA1-1 and ST8SIA1-2) was significantly
reduced compared with that of mock and non-transfected cells. To
further confirm this result, a colony formation assay was performed
and demonstrated that ST8SIA1 overexpression decreased the number
of T24 (Fig. 3D, top panel) and 5637
(Fig. 3D, bottom panel) cell
colonies. The flow cytometry results revealed that ST8SIA1
overexpression markedly arrested T24 (Fig. 3E, top panel) and 5637 (Fig. 3E, bottom panel) cells in the S phase
of the cell cycle. These results suggested that ST8SIA1
downregulated the proliferation ability of 5637 and T24 cells.
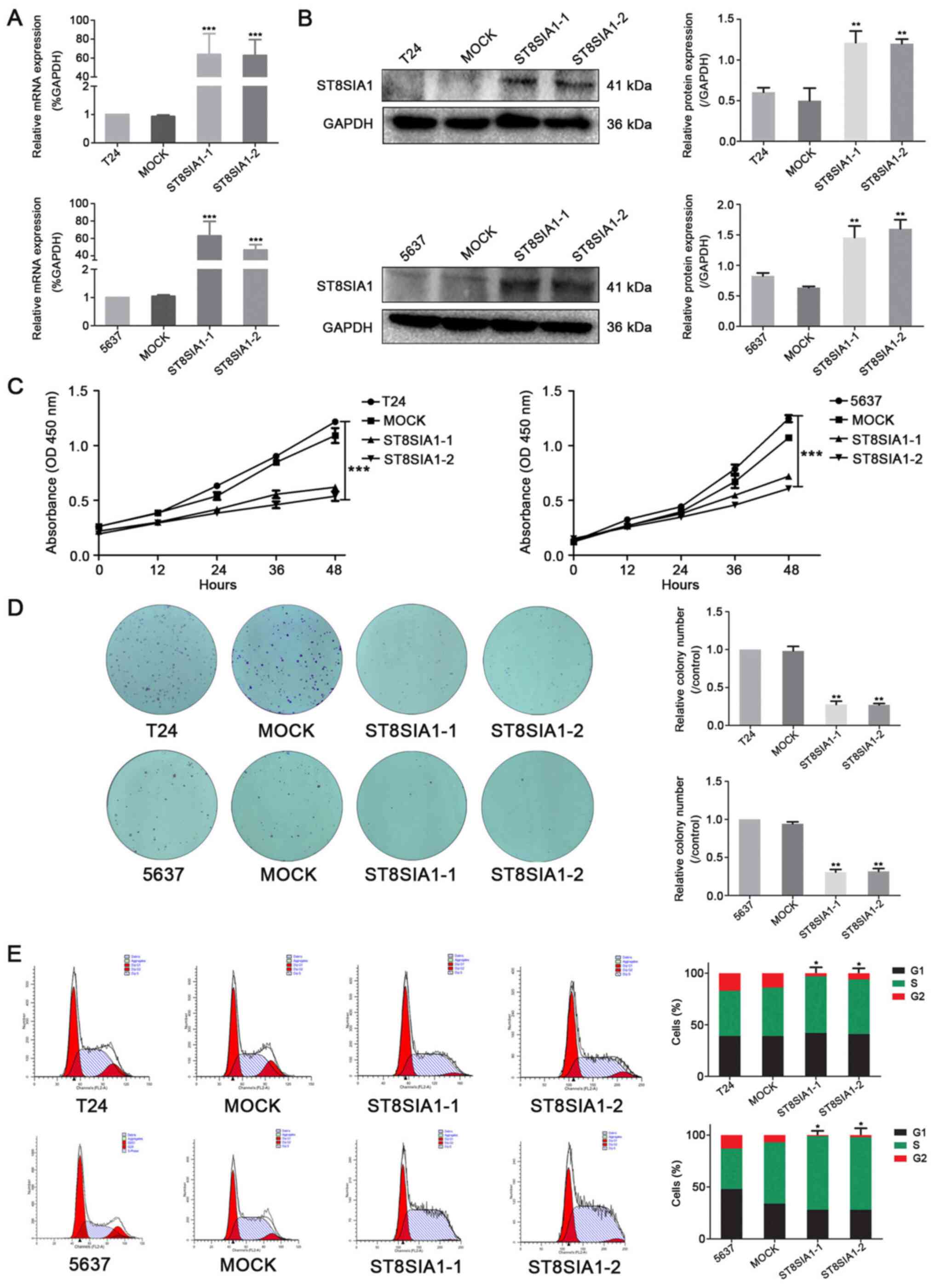 | Figure 3.ST8SIA1 inhibits the proliferation of
5637 and T24 cells. (A) Reverse transcription-quantitative PCR
analysis ofST8SIA1 mRNA expression in non-transfected T24 or 5637
cells, mock and stably transfected cells (ST8SIA1-1 and ST8SIA1-2).
Relative changes in gene expression were normalized to GAPDH and
analyzed using the 2−ΔΔCq method. ***P<0.001 vs.
mock. (B) Western blot analysis of ST8SIA1 protein expression in
non-transfected T24 or 5637, mock, ST8SIA1-1 and ST8SIA1-2 cells.
GAPDH served as a control. **P<0.01 vs. mock. (C) Proliferation
of non-transfected T24 or 5637 cells, mock, ST8SIA1-1 and ST8SIA1-2
cells was detected using a Cell Counting Kit-8 assay at 0, 12, 24,
36 and 48 h. ***P<0.001 vs. mock. (D) The clonogenic ability of
non-transfected T24 or 5637, mock, ST8SIA1-1 and ST8SIA1-2 cells
was analyzed using a cell colony formation assay (magnification,
×100). **P<0.01 vs. mock. (E) Flow cytometry was used to analyze
the cell cycle of non-transfected T24 or 5637, mock, ST8SIA1-1 and
ST8SIA1-2 cells. *P<0.05 vs. mock. OD, optical density; ST8SIA1,
α-2,8-sialyltransferase1. |
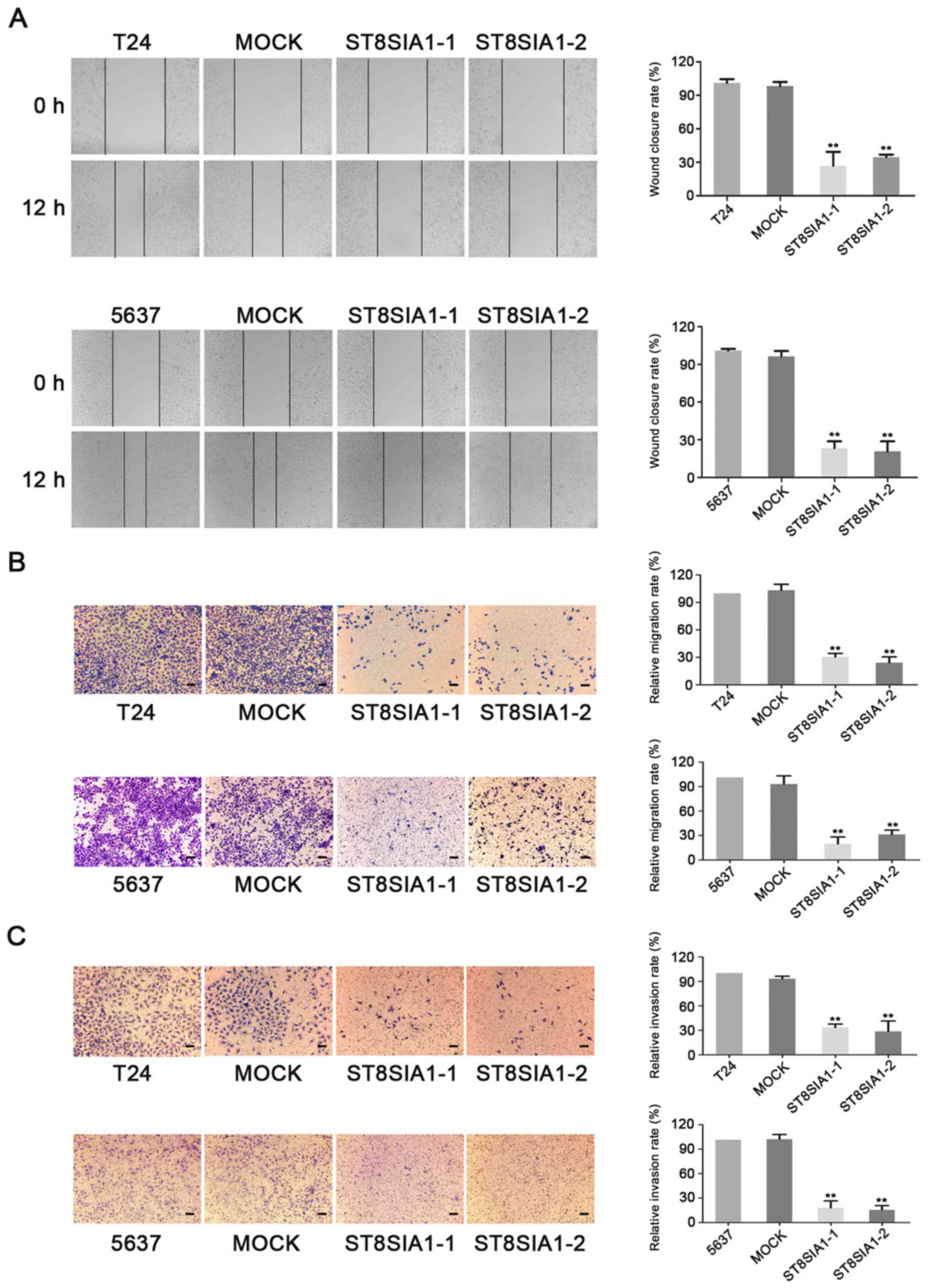
ST8SIA1 inhibits the migration and
invasion of 5637 and T24 cells
To further verify the effects of ST8SIA1 on the
malignant progression of BLCA cells, wound healing and Transwell
assays were used to analyze migration and invasion following
ST8SIA1 overexpression in 5637 and T24 cells. The results of the
wound healing assay demonstrated that the motility of T24 (Fig. 4A, top panel) and 5637 (Fig. 4A, bottom panel) cells stably
overexpressing ST8SIA1 (ST8SIA1-1 and ST8SIA1-2) was markedly
reduced compared with that of mock and non-transfected cells within
12 h. Furthermore, the frequencies of T24 (Fig. 4B and C, top panel) and 5637 (Fig. 4B and C, bottom panel) cells that
migrated and invaded into the lower chamber were significantly
decreased following ST8SIA1 overexpression. These data indicated
that ST8SIA1 attenuated the migration and invasion ability of 5637
and T24 cells.
ST8SIA1 suppresses the JAK2/STAT3
signaling pathway in 5637 and T24 cells
To explore the molecular mechanisms of the antitumor
effect of ST8SIA1 in BLCA cells, the activation and expression
levels of JAK/STAT, as well as the involved signaling molecules
were detected by western blot analysis in 5637 and T24 cells
following ST8SIA1 overexpression. The results demonstrated that
ST8SIA1 overexpression in T24 (Fig.
5A) and 5637 (Fig. 5B) cells
markedly reduced the ratio of the phosphorylated JAK2/total JAK2
and the phosphorylated STAT3/total STAT3, as well as the expression
levels of the downstream targets of JAK/STAT signaling pathway,
such as MMP2, PCNA, cyclin D1 and Bcl2 compared with mock and
non-transfected cells. These results suggested that ST8SIA1
inhibited the activation of JAK2/STAT3 signaling pathway.
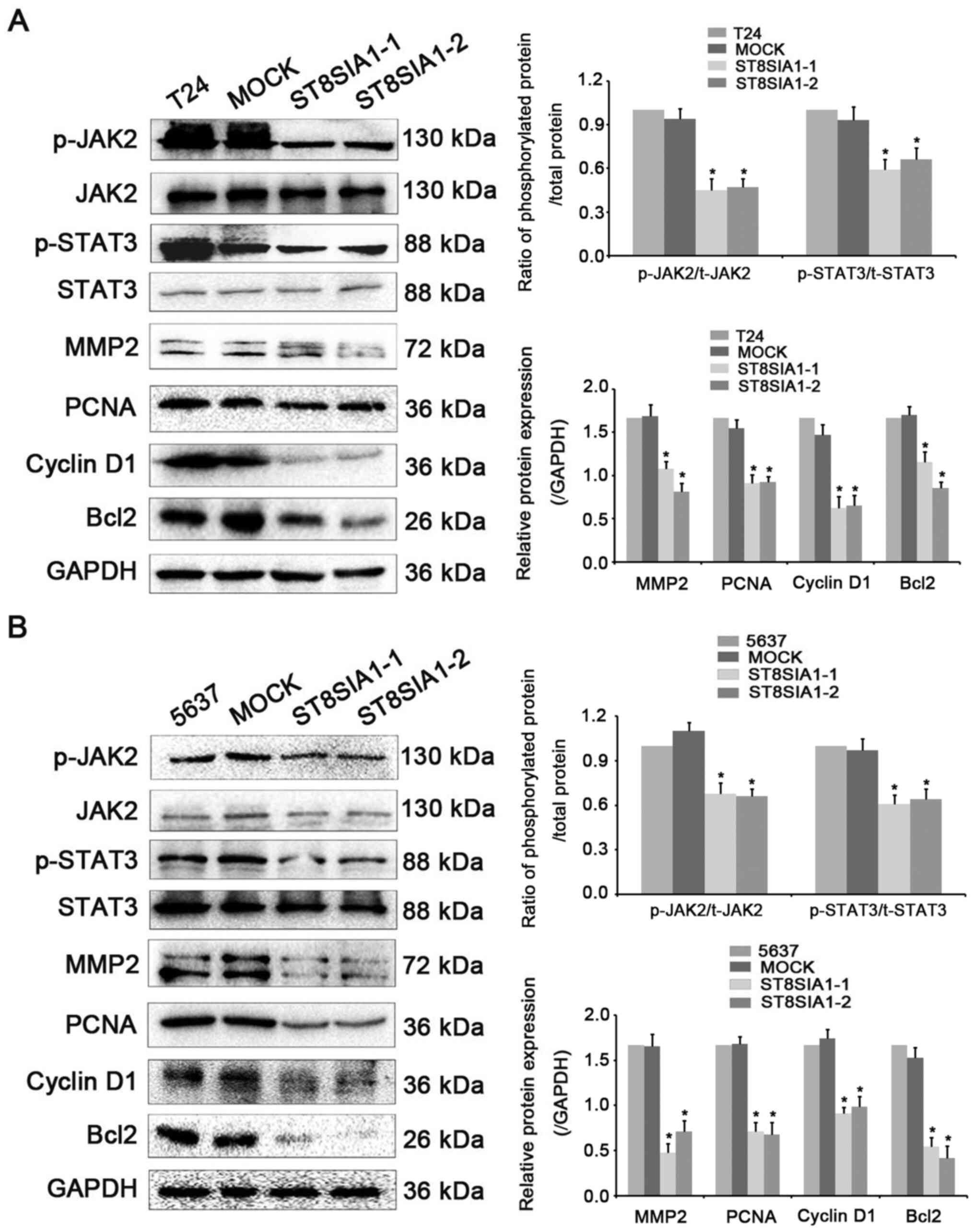 | Figure 5.ST8SIA1 suppresses the JAK2/STAT3
signaling pathway in 5637 and T24 cells. (A) Western blot analysis
of the phosphorylation levels of JAK2 and STAT3, and protein
expression levels of JAK2, STAT3, MMP2, PCNA, cyclin D1 and Bcl2 in
non-transfected T24, mock, ST8SIA1-1 and ST8SIA1-2 cells. GAPDH
served as a control. *P<0.05 vs. mock. (B) Western blot analysis
of the protein expression levels of phosphorylated JAK2,
phosphorylated STAT3, JAK2, STAT3, MMP2, PCNA, cyclin D1 and Bcl2
in non-transfected 5637, mock, ST8SIA1-1 and ST8SIA1-2 cells. GAPDH
served as a control. *P<0.05 vs. mock. JAK2, Janus kinase 2; p-,
phosphorylated; PCNA, proliferating cell nuclear antigen; ST8SIA1,
α-2,8-sialyltransferase 1. |
Discussion
Sialylation is the process of adding sialic acid to
the ends of oligosaccharides and glycoproteins by STs and
glycosidase. Sialylation is involved in embryonic development,
reprogramming, tumorigenesis and immune response (21). Elevated mRNA expression levels of
ST3GAL2 and ST3GAL3 have been reported to serve an important role
in the progression and metastasis of oral cancer (22). Glycoproteins carrying ST3GAL4 have
been identified in gastric cancer cells overexpressing sialyl Lewis
x, and the potential of these proteins as biomarkers of gastric
cancer was evaluated (23). Zhang
et al (24) reported that
high expression levels of ST3GAL3 are associated with an increased
resistance of ovarian cancer cells to paclitaxel, and that the
downregulation of ST3GAL3 could lead to paclitaxel-induced
apoptosis. A previous study had demonstrated that caveolin-1 could
upregulate α2,6-sialylation on integrin α5β1, and promote integrin
α5β1-dependent hepatocarcinoma cell adhesion (25). These results indicated that it is
necessary to investigate the roles and mechanisms of various
members of the ST family in various types of cancer.
Abnormal expression of STsis involved in the
regulation of the malignant progression of cancer (26). Accumulating evidence has demonstrated
that ST8SIA1 can influence the growth and metastasis of a number of
cancer types, including triple-negative breast cancer (18), colorectal cancer (27) and cervical cancer (28). In the present study, 13
differentially expressed glycosyltransferase genes were screened
using TCGA; however, only 1 ST, ST8SIA1, was identified to exhibit
low expression levels in BLCA. The data obtained from the GEPIA
database indicated that the mRNA expression levels of ST8SIA1 were
decreased in BLCA tissues compared with in normal bladder tissues.
The downregulation of ST8SIA1 was also validated in BLCA tissues
and cells by immunohistochemistry and western blotting,
respectively. However, to the best of our knowledge, the clinical
significance of ST8SIA1 expression and its association with
clinicopathological characteristics in patients with BLCA has not
yet been determined. In the present study, clinical data from 136
patients with BLCA revealed that ST8SIA1 was negatively associated
with the pathological grade and degree of invasion of BLCA. These
results suggested that ST8SIA1 may act as a tumor suppressor in
BLCA.
ST8SIA1 is a key enzyme that converts
monosialoganglioside-3 (GM3) to disialoganglioside-3 (GD3)
(29). ST8SIA1 can protect nerves
from GD3-mediated apoptosis (30).
Mandal et al (31) have
reported that ST8SIA1 overexpression can inhibit the growth of
neovascularization cells and block cells at the S phase of the cell
cycle to induce apoptosis in pancreatic cancer cells. In the
present study, to investigate the roles of ST8SIA1 in the malignant
progression of BLCA, two BCLA cell lines (T24 and 5637) stably
overexpressing ST8SIA1 were established. Functional assays
indicated that the overexpression of ST8SIA1 inhibited the
proliferation, migration and invasion of BLCA cells, suggesting
that ST8SIA1 may have an inhibitory effect on the malignant
behavior of BLCA cells. It was suggested that this phenomenon may
be due to the methylation of the ST8SIA1 gene in the development of
BLCA, with the subsequent inhibition of ST8SIA1 expression and
activity in tumor cells (32).
The abnormal activation of JAK/STAT signaling is a
key factor contributing to tumor growth and metastasis (33). To explore the molecular mechanism of
the effect of ST8SIA1 on the occurrence and development of BLCA,
the association between ST8SIA1 and the JAK/STAT signaling pathway
was detected. The results demonstrated that ST8SIA1 overexpression
inhibited the phosphorylation of JAK2 and STAT3 but did not affect
their protein expression. The protein expression levels of JAK/STAT
pathway-targeting signaling molecules, such as MMP2, PCNA, cyclin
D1 and Bcl2, were markedly reduced following ST8SIA1 upregulation.
These findings suggested that ST8SIA1 may attenuate the
proliferation and metastasis of BLCA by suppressing the JAK-STAT
signaling pathway. Wang et al (34) reported that GM3 inhibited the
adhesion, proliferation and EGFR phosphorylation of YTS-1, T24,
5637 and KK47 human BLCA cell lines. Therefore, it was suggested
that ST8SIA1 may decrease the phosphorylation of EGFR, JAK2 and
STAT3 by increasing the sialylation of GM3. However, the detailed
molecular mechanism requires additional studies and
verification.
In conclusion, to the best of our knowledge, the
present study was the first to report that ST8SIA1 expression was
reduced in human BLCA tissue and cell lines. The expression levels
of ST8SIA1 in human BLCA tissue were negatively associated with the
pathological grade and invasiveness of BLCA. Furthermore, the
present study provided evidence that ST8SIA1 could decrease the
proliferation, migration and invasion of BLCA cells by blocking the
activation of JAK/STAT signaling and downregulating the expression
levels of JAK/STAT pathway-targeting signaling molecules. The
expression pattern and antitumor role of ST8SIA1 in BLCA suggested
that ST8SIA1 should be further evaluated as a potential candidate
target for the diagnosis and treatment of BLCA.
Acknowledgements
Not applicable.
Funding
The present study was supported by the Project of
Education Department in Liaoning Province (grant no. LNSJYT201917)
and Natural Science Foundation of Liaoning Province (grant no.
2019-ZD-0640).
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
SY, SW and XS analyzed and interpreted the data. SY,
SW, XS, YW, JZ and JL performed the experiments and analyzed the
data. SY and SW wrote the manuscript. DY and YJ edited and revised
the manuscript. DY and YJ proposed the conception of the study and
designed the project. SY, XS and YJ confirmed the authenticity of
all the raw data. All authors read and approved the final
manuscript.
Ethics approval and consent to
participate
The present study was approved by the Institutional
Research Ethics Committee of the First Affiliated Hospital of
Dalian Medical University (approval no. LCKY2015-08; Dalian,
China), and written informed consent was provided by the patients
with bladder cancer who underwent surgery.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Bray F, Ferlay J, Soerjomataram I, Siegel
RL, Torre LA and Jemal A: Global cancer statistics 2018: GLOBOCAN
estimates of incidence and mortality worldwide for 36 cancers in
185 countries. CA Cancer J Clin. 68:394–424. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Ohadian Moghadam S and Nowroozi MR:
Toll-like receptors: The role in bladder cancer development,
progression and immunotherapy. Scand J Immunol. 90:e128182019.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Tan TZ, Mathieu R, Tan KT, Huang YJ and
Jean-Paul T: Molecular subtypes of urothelial bladder cancer:
Results from a meta-cohort analysis of 2411 Tumors. Eur Urol.
75:423–432. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
McConkey DJ and Lerner SP: SIU-ICUD
consultation on bladder cancer: Basic science. World J Urol.
37:15–29. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Moremen KW, Tiemeyer M and Nairn AV:
Vertebrate protein glycosylation: Diversity, synthesis and
function. Nat Rev Mol Cell Biol. 13:448–462. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Tang L and Chen X, Zhang X, Guo Y, Su J,
Zhang J, Peng C and Chen X: N-Glycosylation in progression of skin
cancer. Med Oncol. 36:502019. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Silsirivanit A: Glycosylation markers in
cancer. Adv Clin Chem. 89:189–213. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Munkley J: Glycosylation is a global
target for androgen control in prostate cancer cells. Endocr Relat
Cancer. 24:R49–R64. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Hoja-Lukowicz D, Przybylo M, Duda M,
Pochec E and Bubka M: On the trail of the glycan codes stored in
cancer-related cell adhesion proteins. Biochim Biophys Acta Gen
Subj. 1861:3237–3257. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Vajaria BN and Patel PS: Glycosylation: A
hallmark of cancer? Glycoconj J. 34:147–156. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Veillon L, Fakih C, Abou-El-Hassan H,
Kobeissy F and Mechref Y: Glycosylation changes in brain cancer.
ACS Chem Neurosci. 9:51–72. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Natoni A, Bohara R, Pandit A and O'Dwyer
M: Targeted approaches to inhibit sialylation of multiple myeloma
in the bone marrow microenvironment. Front Bioeng Biotechnol.
7:2522019. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Natoni A, Macauley MS and O'Dwyer ME:
Targeting selectins and their ligands in cancer. Front Oncol.
6:932016. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Weijers CA, Franssen MC and Visser GM:
Glycosyltransferase-catalyzed synthesis of bioactive
oligosaccharides. Biotechnol Adv. 26:436–456. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Li Y and Chen X: Sialic acid metabolism
and sialyltransferases: Natural functions and applications. Appl
Microbiol Biotechnol. 94:887–905. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Bhide GP and Colley KJ: Sialylation of
N-glycans: Mechanism, cellular compartmentalization and function.
Histochem Cell Biol. 147:149–174. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Park HJ, Kim JH, Yoon JS, Choi YJ, Choi
YH, Kook KH and Choi JH: Identification and functional
characterization of ST3GAL5 and ST8SIA1 variants in patients with
thyroid-associated ophthalmopathy. Yonsei Med J. 58:1160–1169.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Nguyen K, Yan Y, Yuan B, Dasgupta A, Sun
J, Mu H, Do KA, Ueno NT, Andreeff M and Battula VL: ST8SIA1
regulates tumor growth and metastasis in TNBC by activating the
FAK-AKT-mTOR signaling pathway. Mol Cancer Ther. 17:2689–2701.
2018. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Charlton ME, Adamo MP, Sun L and Deorah S:
Bladder cancer collaborative stage variables and their data
quality, usage, and clinical implications: A review of SEER data,
2004–2010. Cancer. 120 (Suppl 23):3815–3825. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Li F and Ding J: Sialylation is involved
in cell fate decision during development, reprogramming and cancer
progression. Protein Cell. 10:550–565. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Mehta KA, Patel KA, Pandya SJ and Patel
PS: ‘Aberrant sialylation plays a significant role in oral squamous
cell carcinoma progression’. J Oral Pathol Med. 49:253–259. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Gomes C, Almeida A, Barreira A, Calheiros
J, Pinto F, Abrantes R, Costa A, Polonia A, Polonia A, Osório H, et
al: Carcinoembryonic antigen carrying SLeX as a new
biomarker of more aggressive gastric carcinomas. Theranostics.
9:7431–7446. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Zhang X, Yang X, Chen M, Zheng S, Li J,
Lin S and Wang X: ST3Gal3 confers paclitaxel-mediated
chemoresistance in ovarian cancer cells by attenuating caspase-8/3
signaling. Mol Med Rep. 20:4499–4506. 2019.PubMed/NCBI
|
|
25
|
Yu S, Fan J, Liu L, Zhang L, Wang S and
Zhang J: Caveolin-1 up-regulates integrin α2,6-sialylation to
promote integrin α5β1-dependent hepatocarcinoma cell adhesion. FEBS
Lett. 587:782–787. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Ghosh S: Sialylation and sialyltransferase
in insects. Glycoconj J. 35:433–441. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Shan Y, Liu Y, Zhao L, Liu B, Li Y and Jia
L: MicroRNA-33a and let-7e inhibit human colorectal cancer
progression by targeting ST8SIA1. Int J Biochem Cell Biol.
90:48–58. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Danolic D, Heffer M, Wagner J, Skrlec I,
Alvir I, Mamic I, Susnjar L, Banovic M, Danolić D and Puljiz M:
Role of ganglioside biosynthesis genetic polymorphism in cervical
cancer development. J Obstet Gynaecol. 40:1127–1132. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Yeh SC, Wang PY, Lou YW, Khoo KH, Hsiao M,
Hsu TL and Wong CH: Glycolipid GD3 and GD3 synthase are key drivers
for glioblastoma stem cells and tumorigenicity. Proc Natl Acad Sci
USA. 113:5592–5597. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Bernardo A, Harrison FE, McCord M, Zhao J,
Bruchey A, Davies SS, Jackson Roberts L II, Mathews PM, Matsuoka Y,
Ariga T, et al: Elimination of GD3 synthase improves memory and
reduces amyloid-beta plaque load in transgenic mice. Neurobiol
Aging. 30:1777–1791. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Mandal C, Sarkar S, Chatterjee U,
Schwartz-Albiez R and Mandal C: Disialoganglioside GD3-synthase
over expression inhibits survival and angiogenesis of pancreatic
cancer cells through cell cycle arrest at S-phase and disruption of
integrin-β1-mediated anchorage. Int J Biochem Cell Biol.
53:162–173. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Weisenberger DJ: Characterizing DNA
methylation alterations from the cancer genome atlas. J Clin
Invest. 124:17–23. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Arumuggam N, Bhowmick NA and Rupasinghe
HP: A Review: Phytochemicals targeting JAK/STAT signaling and IDO
expression in cancer. Phytother Res. 29:805–817. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Wang H, Isaji T, Satoh M, Li D, Arai Y and
Gu J: Antitumor effects of exogenous ganglioside GM3 on bladder
cancer in an orthotopic cancer model. Urology. 81:210.e11–5. 2013.
View Article : Google Scholar : PubMed/NCBI
|