|
1
|
Brown JJ, Ollier W, Arscott G, Ke X, Lamb
J, Day P and Bayat A: Genetic susceptibility to keloid scarring:
SMAD gene SNP frequencies in Afro-caribbeans. Exp Dermatol.
17:610–613. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
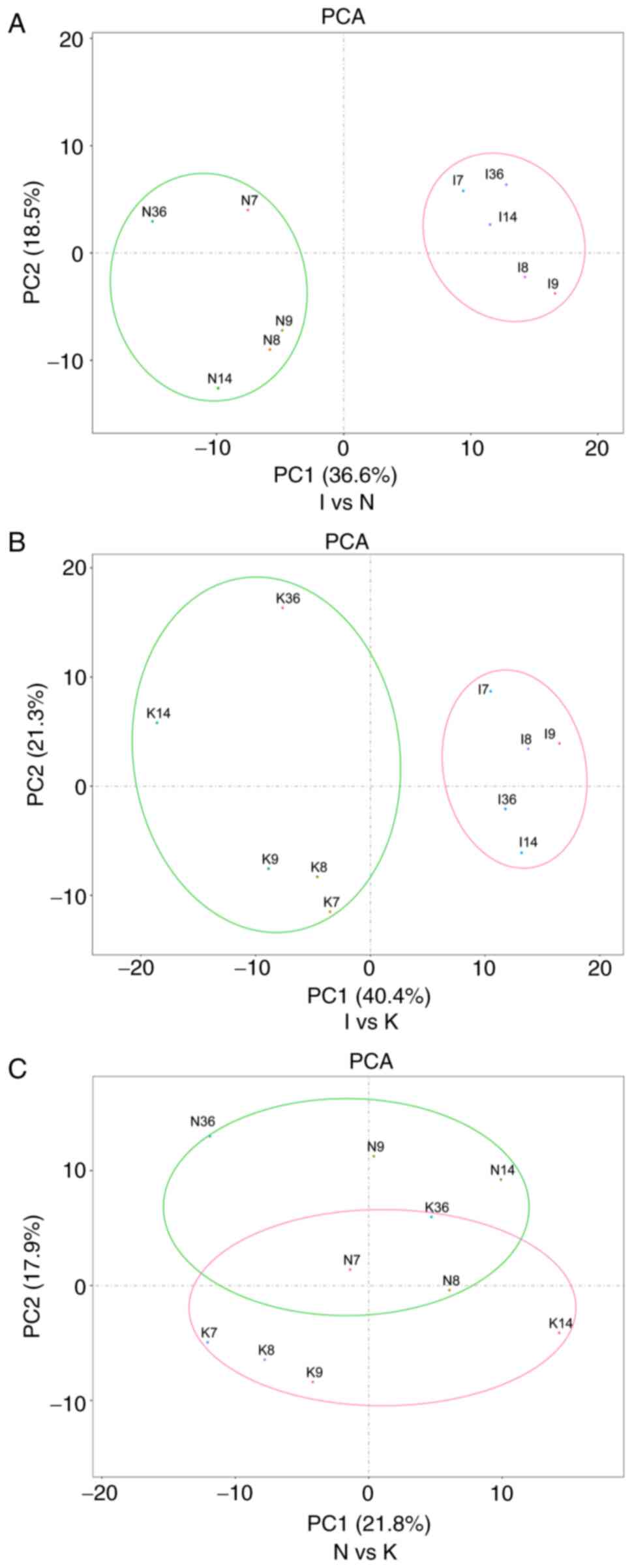
Chung S, Nakashima M, Zembutsu H and
Nakamura Y: Possible involvement of NEDD4 in keloid formation; its
critical role in fibroblast proliferation and collagen production.
Proc Jpn Acad Ser B Phys Biol Sci. 87:563–573. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Glass DA II: Current understanding of the
genetic causes of keloid formation. J Investig Dermatol Symp Proc.
18:S50–S53. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Tsai CH and Ogawa R: Keloid research:
Current status and future directions. Scars Burn Heal.
5:20595131198686592019.PubMed/NCBI
|
|
5
|
Song KX, Liu S, Zhang MZ, Liang WZ, Liu H,
Dong XH, Wang YB and Wang XJ: Hyperbaric oxygen therapy improves
the effect of keloid surgery and radiotherapy by reducing the
recurrence rate. J Zhejiang Univ Sci B. 19:853–862. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Ogawa R: Keloid and hypertrophic scars are
the result of chronic inflammation in the reticular dermis. Int J
Mol Sci. 18:6062017. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Diaz A, Tan K, He H, Xu H, Cueto I, Pavel
AB, Krueger JG and Guttman-Yassky E: Keloid lesions show increased
IL-4/IL-13 signaling and respond to Th2-targeting dupilumab
therapy. J Eur Acad Dermatol Venereol. 34:e161–e164. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Shi CK, Zhao YP, Ge P and Huang GB:
Therapeutic effect of interleukin-10 in keloid fibroblasts by
suppression of TGF-β/Smad pathway. Eur Rev Med Pharmacol Sci.
23:9085–9092. 2019.PubMed/NCBI
|
|
9
|
Gankande TU, Wood FM, Edgar DW, Duke JM,
DeJong HM, Henderson AE and Wallace HJ: A modified vancouver scar
scale linked with TBSA (mVSS-TBSA): Inter-rater reliability of an
innovative burn scar assessment method. Burns. 39:1142–1149. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Wang CH, Shan MJ, Liu H, Hao Y, Song KX,
Wu HW, Meng T, Feng C, Qi Z, Wang Z and Wang YB: Hyperbaric oxygen
treatment on keloid tumor immune gene expression. Chin Med J
(Engl). 134:2205–2213. 2021.PubMed/NCBI
|
|
11
|
Privé F, Luu K, Vilhjálmsson BJ and Blum
M: Performing highly efficient genome scans for local adaptation
with r package pcadapt version 4. Mol Biol Evol. 37:2153–2154.
2020. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Kanehisa M and Goto S: KEGG: Kyoto
encyclopedia of genes and genomes. Nucleic Acids Res. 28:27–30.
2000. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Huang DW, Sherman BT and Lempicki RA:
Systematic and integrative analysis of large gene lists using DAVID
bioinformatics resources. Nat Protoc. 4:44–57. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Szklarczyk D, Gable AL, Lyon D, Junge A,
Wyder S, Huerta-Cepas J, Simonovic M, Doncheva NT, Morris JH and
Bork P: STRING v11: Protein-protein association networks with
increased coverage, supporting functional discovery in genome-wide
experimental datasets. Nucleic Acids Res. 47:D607–D613. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Shannon P, Markiel A, Ozier O, Baliga NS,
Wang JT, Ramage D, Amin N, Schwikowski B and Ideker T: Cytoscape: A
software environment for integrated models of biomolecular
interaction networks. Genome Res. 13:2498–2504. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Ni M, Liu X, Wu J, Zhang D, Tian J, Wang
T, Liu S, Meng Z, Wang K, Duan X, et al: Identification of
candidate biomarkers correlated with the pathogenesis and prognosis
of non-small cell lung cancer via integrated bioinformatics
analysis. Front Genet. 9:4692018. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Yan J, Zuo G, Sherchan P, Huang L, Ocak U,
Xu W Travis Z, Wang W, Zhang J and Tang J: CCR1 Activation Promotes
Neuroinflammation Through CCR1/TPR1/ERK1/2 Signaling Pathway After
Intracerebral Hemorrhage in Mice. Neurotherapeutics. 17:1170–1183.
2020. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
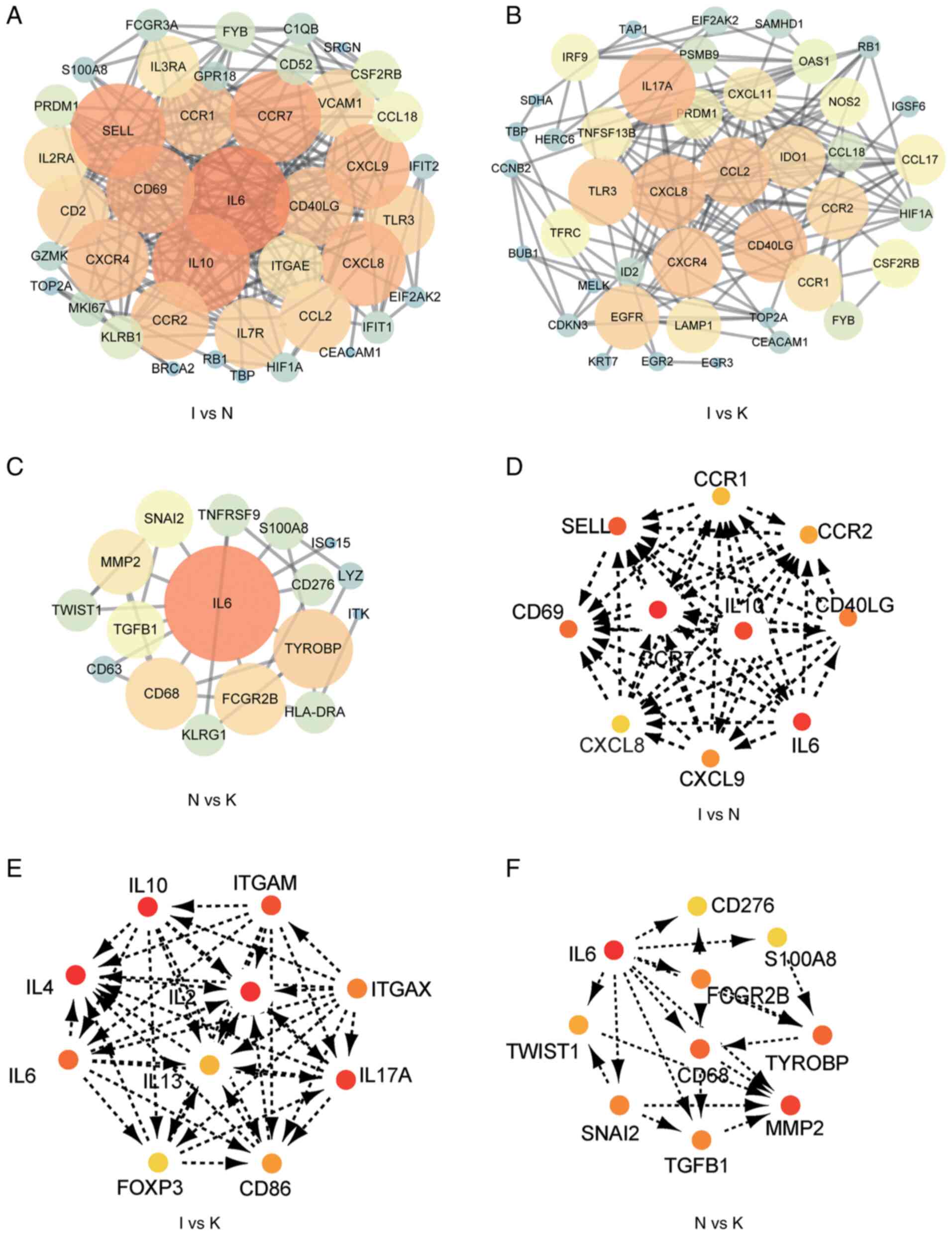
Chin CH, Chen SH, Wu HH, Ho CW, Ko MT and
Lin CY: cytoHubba: Identifying hub objects and sub-networks from
complex interactome. BMC Syst Biol. 8 (Suppl 4):S112014. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Song X, Du R, Gui H, Zhou M, Zhong W, Mao
C and Ma J: Identification of potential hub genes related to the
progression and prognosis of hepatocellular carcinoma through
integrated bioinformatics analysis. Oncol Rep. 43:133–146.
2020.PubMed/NCBI
|
|
20
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2-(Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Zhang Q, Yamaza T, Kelly AP, Shi S, Wang
S, Brown J, Wang L, French SW, Shi S and Le AD: Tumor-like stem
cells derived from human keloid are governed by the inflammatory
niche driven by IL-17/IL-6 axis. PLoS One. 4:e77982009. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Limandjaja GC, Niessen FB, Scheper RJ and
Gibbs S: The keloid disorder: Heterogeneity, histopathology,
mechanisms and models. Front Cell Dev Biol. 8:3602020. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
van den Broek LJ, Limandjaja GC, Niessen
FB and Gibbs S: Human hypertrophic and keloid scar models:
Principles, limitations and future challenges from a tissue
engineering perspective. Exp Dermatol. 23:382–386. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Tan S, Khumalo N and Bayat A:
Understanding keloid pathobiology from a quasi-neoplastic
perspective: Less of a scar and more of a chronic inflammatory
disease with cancer-like tendencies. Front Immunol. 10:18102019.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Karnell JL, Rieder SA, Ettinger R and
Kolbeck R: Targeting the CD40-CD40L pathway in autoimmune diseases:
Humoral immunity and beyond. Adv Drug Deliv Rev. 141:92–103. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Tokunaga R, Zhang W, Naseem M, Puccini A,
Berger MD, Soni S, McSkane M, Baba H and Lenz HJ: CXCL9, CXCL10,
CXCL11/CXCR3 axis for immune activation-A target for novel cancer
therapy. Cancer Treat Rev. 63:40–47. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Zhong L, Bian L, Lyu J, Jin H, Liu Z, Lyu
L and Lu D: Identification and integrated analysis of microRNA
expression profiles in keloid. J Cosmet Dermatol. 17:917–924. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Ghazizadeh M: Essential role of IL-6
signaling pathway in keloid pathogenesis. J Nippon Med Sch.
74:11–22. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Jin Q, Gui L, Niu F, Yu B, Lauda N, Liu J,
Mao X and Chen Y: Macrophages in keloid are potent at promoting the
differentiation and function of regulatory T cells. Exp Cell Res.
362:472–476. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Suzuki M, Yokota M, Matsumoto T and Ozaki
S: Synergic effects of CD40 and CD86 silencing in dendritic cells
on the control of allergic diseases. Int Arch Allergy Immunol.
177:87–96. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Mburu YK, Wang J, Wood MA, Walker WH and
Ferris RL: CCR7 mediates inflammation-associated tumor progression.
Immunol Res. 36:61–72. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Huang H, Fu S and Liu D: Detection and
analysis of the hedgehog signaling pathway-related long non-coding
RNA (lncRNA) expression profiles in keloid. Med Sci Monit.
24:9032–9044. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Spangler JB, Moraga I, Mendoza JL and
Garcia KC: Insights into cytokine-receptor interactions from
cytokine engineering. Annu Rev Immunol. 33:139–167. 2015.
View Article : Google Scholar : PubMed/NCBI
|