Introduction
Immunotherapy has become an important therapeutic
strategy for the management of several types of cancers. Antibodies
against T cell suppressor receptor cytotoxic T lymphocyte antigen 4
(CTLA4) and programmed cell death protein 1 (PD-1) or its ligand,
programmed death-ligand 1 (PD-L1), have been widely used in cancer
immunotherapy (1). The success of
these immune checkpoint inhibitors has demonstrated the potential
of tumor-specific CD8+ T cells in the prevention and
treatment of cancer. Unlike the effects usually observed with
oncogene-targeting drugs or standard chemotherapy, reversing the
inhibition of CD8+ T cells can lead to tumor regression
or elimination, as well as lasting clinical remission (2). However, for many indications (lung
cancer and colon cancer), only 15% of patients exhibit a clinical
response to checkpoint inhibitors, such as anti-CTLA4 and anti-PD-1
antibodies, when administered alone (3). Several factors can limit the use of
CD8+ T cell immunotherapy, including the difficulty in
recognizing tumor-specific peptides presented by MHC class I
molecules and the ability of tumor cells to evade being killed by
CD8+ T cells (4,5). It
is therefore important to elucidate how the tumor and tumor
microenvironment affects the function of CD8+ T cells,
as well as the effect of cancer immunotherapy.
Zinc finger DHHC-type palmitoyltransferase 9
(ZDHHC9) is an integral membrane protein and a member of the zinc
finger DHHC domain-containing protein family (6). The encoded protein mediates the
palmitoylation of proteins as a palmitoyltransferase.
Palmitoylation is a reversible posttranslational lipid modification
that can attach long-chain fatty acids, such as palmitic acid, to
cysteine residues and contribute to the vesicular transport and
subcellular localization of modified proteins (7). It has been previously reported that
ZDHHC9 is associated with neurological and neurodevelopmental
disorders (8). Loss-of-function
mutations of ZDHHC9 can lead to X-linked intellectual disability
and an increase in the incidence of epilepsy (9,10).
ZDHHC9 inactivation mitigates the leukemogenic potential of
oncogenic Nras (11), enhancer of
zeste 2 polycomb repressive complex 2 subunit (12), and TEA domain transcription factor
4 (13). Of note, ZDHHC9 also
promotes the growth of breast cancer by palmitoylating PD-L1,
affecting the stability of PD-L1 and protecting tumor cells from
T-cell immune surveillance (14).
ZDHHC9 expression is upregulated in colon cancer, especially in
microsatellite stable (MSS) tumors (15); however, how it regulates the
development of colon cancer remains unclear. ZDHHC9 regulates PD-L1
expression in breast cancer, but whether it also regulates the
immune microenvironment of colon cancer is still unclear. Thus, the
role of ZDHHC9 in colon cancer, especially the role of ZDHHC9 in
the antitumor immunity of colon cancer patients, requires further
investigation.
In the present study, it was shown that ZDHHC9
inhibited the proliferation of colon cancer cells in vitro
but promoted the growth of colon cancer in vivo by
inhibiting the T cell-mediated tumor immune response. It was also
demonstrated that ZDHHC9 positively regulated the gene expression
of PD-L1 by affecting the activation of the IFN-γ-induced JAK/STAT1
signaling pathway in colon cancer.
Materials and methods
Cell culture
MC38 mouse colon cancer cells and human DLD-1 colon
cancer cells were purchased from National Infrastructure of Cell
Line Resource and cultured in DMEM (HyClone; Cytiva) supplemented
with 10% FBS (Gibco; Thermo Fisher Scientific, Inc.). Cells were
incubated with 5% CO2 at 37°C. ZDHHC9-knockdown
(ZDHHC9-KD) cells in MC38 were prepared using the CRISPR system
with the pLV hU6-sgRNA hUbC-dCas9-KRAB-T2a-Puro plasmid (cat. no.
71236; Addgene, Inc.) (16). The
following sgRNA sequences were used: ZDHHC9-sgRNA top,
5′-caccGTAGCGACTTCTCCCTGACA-3′; ZDHHC9-sgRNA bottom,
5′-aaacTGTCAGGGAGAAGTCGCTAC-3′.
sgRNA was phosphorylated, annealed, and ligated with
the plasmid and transformed into E. coli competent DH5α
cells. The clones were selected and then sequenced by Sangon
Biotech, Co., Ltd. Next, the plasmid was prepared and transfected
into MC38 cells, and then the ZDHHC9-KD cell line was screened
using puromycin. Control cells were transfected with empty control
plasmid. ZDHHC9-KD DLD-1 cells were constructed using RNAi. The
following siRNA sequences (Shanghai GenePharma Co., Ltd.) were
used: 5′-GGGACUGACUGGAUUUCAUTT-3′ (sense) and
5′-AUGAAAUCCAGUCAGUCCCTT-3′ (anti-sense). DLD-1 cells were
transfected with ZDHHC9 siRNA for 72 h using RNAiMAX (Invitrogen;
Thermo Fisher Scientific, Inc.). The ZDHHC9-OE cell line was
constructed by transfecting the MIGR1-ZDHHC9 plasmid into MC38
cells, control cells were transfected with empty MIGR1-vector
plasmid (cat. no. 27490; Addgene, Inc.). The sequences of the
primers for ZDHHC9, used to construct ZDHHC9-OE cell line, were as
follows: Sense, 5′-AATTAGATCTCTCGAATGTCTGTGATGGTGGTAAGA-3′ and
antisense, 5′-GGGGGGGGCGGAATTCTTCTCAGCTTCGGATGCCTCCT-3′.
Data collection and processing
ZDHHC9 gene expression in 270 colon cancer tissues
and 41 adjacent tissues in TCGA was assessed using GEPIA
(http://gepia.cancer-pku.cn/). The
patients were divided into a low and a high expression group based
on median ZDHHC9 expression. Overall survival analysis was then
performed.
ZDHHC9-related immune infiltration
analysis
Tumor Immune Estimation Resource (TIMER) (https://cistrome.shinyapps.io/timer/) is
a web server for the comprehensive analysis of tumor-infiltrating
immune cells. Based on this database, the correlation between
ZDHHC9 expression and CD8+ T cells, neutrophils, and
macrophages was analyzed. In addition, the relationship between
ZDHHC9 and IFN-γ expression was analyzed.
Cell proliferation assay
The cells were incubated at 37°C for 2 h with 10 µl
CCK-8 solution (cat. no. 40203ES60; Dojindo Molecular Technologies,
Inc.), and absorbance was measured at 450 nm using a microplate
reader. Proliferation rates were determined after 24, 48, 72, or 96
h.
In vitro killing assay
To prepare effector cells, OT-1 mice were
subcutaneously immunized using OVA. All experiments involving mice
were conducted in accordance with the National Institute of Health
Guide for the Care and Use of Laboratory Animals and were approved
by the Scientific Investigation Board of the Naval Medical
University (Shanghai, China). The mice were kept in a special
pathogen-free facility with free access to drinking water and a
pellet-based diet. After 7 days, mice were euthanized by injection
of 150 mg/kg pentobarbital, and the spleen and mesenteric lymph
nodes of OT-1 mice were collected. CD8+ T cells were
separated with magnetic beads that had been activated with
anti-CD3/anti-CD28 antibodies. ZDHHC9-KD and control cells were
incubated with 2 µmol/l OVA (257–264) polypeptide. The supernatant
was then washed out, and the cell count was adjusted to
1×105/ml. Effector cells were added to 100 µl target
cell suspension according to different effector-target (E:T) ratios
of 1:2, 1:1, and 2:1. The cells were cultured at 37°C and 5%
CO2 for 6 h, FVS780 fluorescent antibody (1:1,000; cat.
no. 565388; BD Biosciences) was added, and the cells were
centrifuged at 3,000 × g at 4°C for 30 min and resuspended for flow
cytometry (Attune NxT; Thermo Fisher Scientific, Inc.). Flowjo_v10
(FlowJo LLC) was used to analyze the data. The specific killing
ability was equivalent to the proportion of FVS780-positive cells.
The above animal experiments were approved by the Ethics Committee
of the Navy Medical University.
Mouse model and tumor studies
For the in vivo studies, 8-week-old male
C57BL/6 mice were randomly divided into groups (n=5 per group). The
average starting weight of mice was ~21 g. Stable ZDHHC9-KD and
control cells were digested and resuspended at a concentration of
1.5×106 cells/200 µl in PBS. A 200-µl cell suspension
was injected into the right hip of each mouse subcutaneously. A
total of 5 mice per group were raised under the same conditions,
and the size of the tumors was measured every 3 days; the tumor
volume was calculated as follows: Volume=length × width2
×0.5. On the 21st day, the mice were sacrificed; mice were
euthanized by injection of 150 mg/kg pentobarbital.
Tumor-infiltrating CD8+
T-cell responses
The tumor was cut into pieces and digested using 200
µg/ml collagenase (cat. no. LS004188; Worthington Biochemical
Corporation) and 20 µg/ml DNase (cat. no. LS002138; Worthington
Biochemical Corporation) at 37°C for 30 min. The digested tumor
tissue suspension was filtered through a 0.45-µm filter, and the
filtered single-cell suspension was collected and treated with
Golgiplug Protein Transport Inhibitor (cat. no. 555029; BD
Biosciences), according to the manufacturer's instructions. The
cells were then labeled with CD8 (1:100; cat. no. 11-0081-82;
Ebioscience; Thermo Fisher Scientific, Inc.) and an intracellular
IFN-γ antibody (1:100; cat. no. 554413; BD Biosciences, Inc.) at
4°C for 15 min, and assessed using a flow cytometer (Attune NxT;
Thermo Fisher Scientific, Inc.).
RNA isolation and reverse
transcription-quantitative PCR (RT-qPCR)
RNA was extracted from cells and reverse-transcribed
into cDNA using an M-MLV RT kit (cat. no. 2641A; Takara Bio, Inc.)
according to the manufacturer's protocol. The reverse transcription
temperature protocol was: 95°C for 10 sec, 58°C for 15 sec and 72°C
for 35 sec.
The sequences of the primers used for amplification
were: β-actin forward, 5′-AGTGTGACGTGACATCCGT-3′ and reverse,
5′-GCAGCTCAGTAACAGTCCGC-3′ reverse; ZDHHC9 forward,
5′-AAGGTGACACGGAAATGGGAG-3′ and reverse,
5′-CGACACTCGAGAGCAAGAAGAA-3′; and PD-L1 forward,
5′-GGTGCCGACTACAAGCGAAT-3′ and reverse,
5′-AGCCCTCAGCCTGACATGTC-3′.
Western blotting
Total protein was extracted from the cells using
M-per (cat. no. 78505; Thermo Fisher Scientific, Inc.), and then
the concentration of the protein was measured by BCA assay (cat.
no. BCA1; Merck KGaA). The final protein amount was 15 µg per well.
The proteins were separated on 10% gels using SDS-PAGE, transferred
to PVDF membranes (EMD Millipore), and incubated with primary
antibodies overnight at 4°C, followed by incubation with a
horseradish peroxidase (HRP)-conjugated (1:2,000; cat. no.
7076/7074; Cell Signaling Technology, Inc.) secondary antibody for
2 h at 37°C. Signals were visualized using Western Bright ECL HRP
(EMD Millipore). The following primary antibodies were used for
western blotting: ZDHHC9 (1:1,000; cat. no. ab74504; Abcam, Inc.),
STAT1 (1:1,000; cat. no. 14994; Cell Signaling Technology, Inc.),
p-STAT1 (1:1,000; cat. no. 14994; Cell Signaling Technology, Inc.),
JAK1 (1:1,000; cat. no. ab133666; Abcam, Inc.), p-JAK1 (1:1,000;
cat. no. ab133666; Abcam, Inc.) and GAPDH (1:5,000; cat. no.
amab91153; MilliporeSigma).
Statistical analysis
Data are presented as the mean ± SD. A Student's
t-test was used for comparisons between the two groups. A one-way
ANOVA followed by post hoc Bonferroni's correction was used for
comparison between multiple groups. P<0.05 was considered to
indicate a statistically significant difference. All statistical
analyses were performed using GraphPad Prism version 8.0 (GraphPad
Software, Inc.).
Results
ZDHHC9 expression is upregulated in
colon cancer and is positively associated with a poorer
prognosis
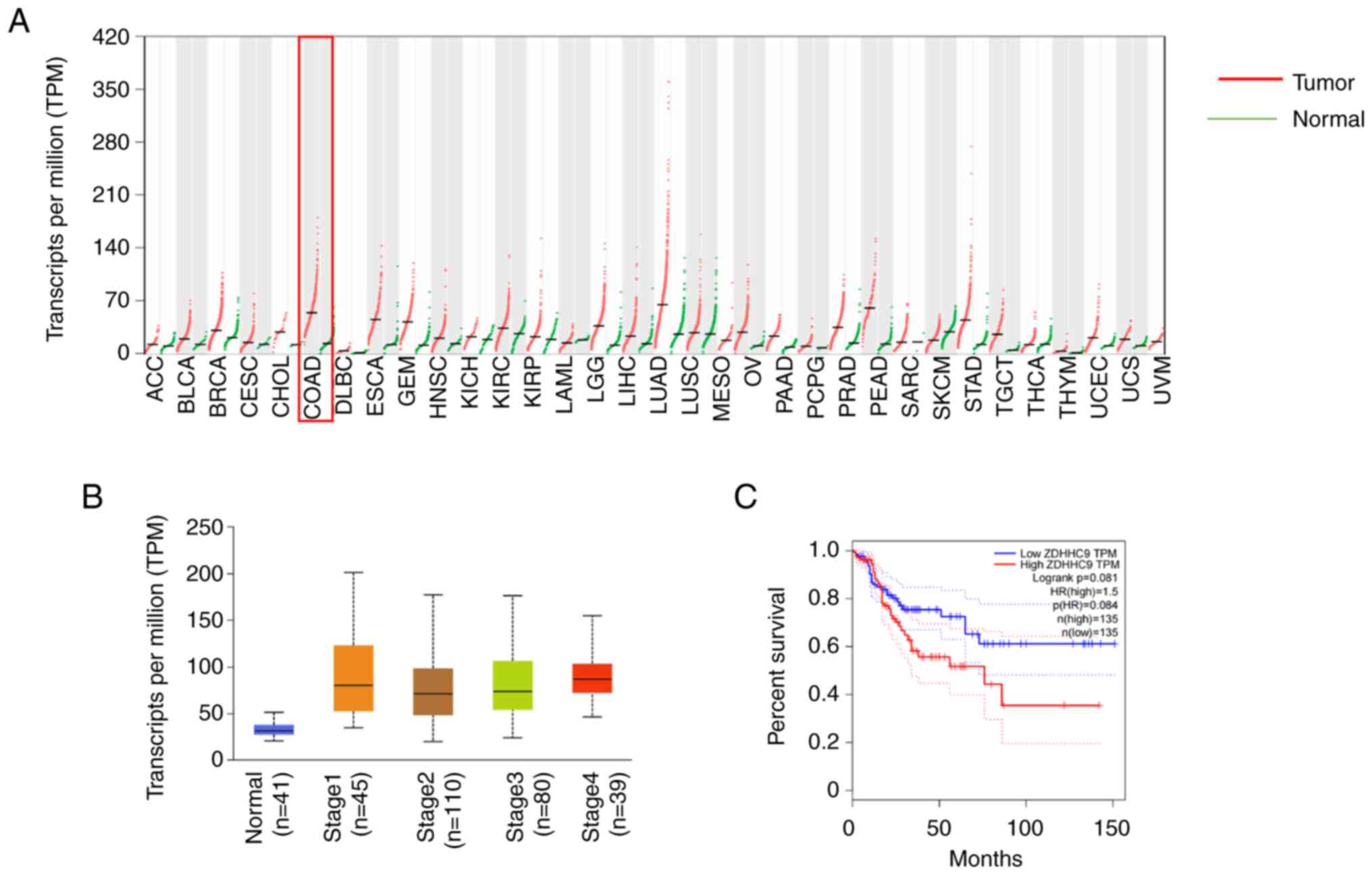
The expression profiles of ZDHHC9 in several cancers
based on data from TCGA were analyzed using GEPIA (http://gepia.cancer-pku.cn/), and the results
indicated that ZDHHC9 mRNA was upregulated in numerous cancers,
including colon, lung, esophageal, and other gastrointestinal
cancers. In particular, the expression of ZDHHC9 in colon cancer
was higher than that in normal tissues (Fig. 1A). ZDHHC9 expression in tissues of
different TNM stages was analyzed, and the results showed that the
expression of ZDHHC9 in stages 1, 2, 3, and 4 of colon cancer was
significantly higher than that in the normal tissues (Fig. 1B). In addition, data on 270
patients with colon cancer from TCGA were also analyzed, and it was
found that patients with higher ZDHHC9 expression had markedly
lower survival times (Fig.
1C).
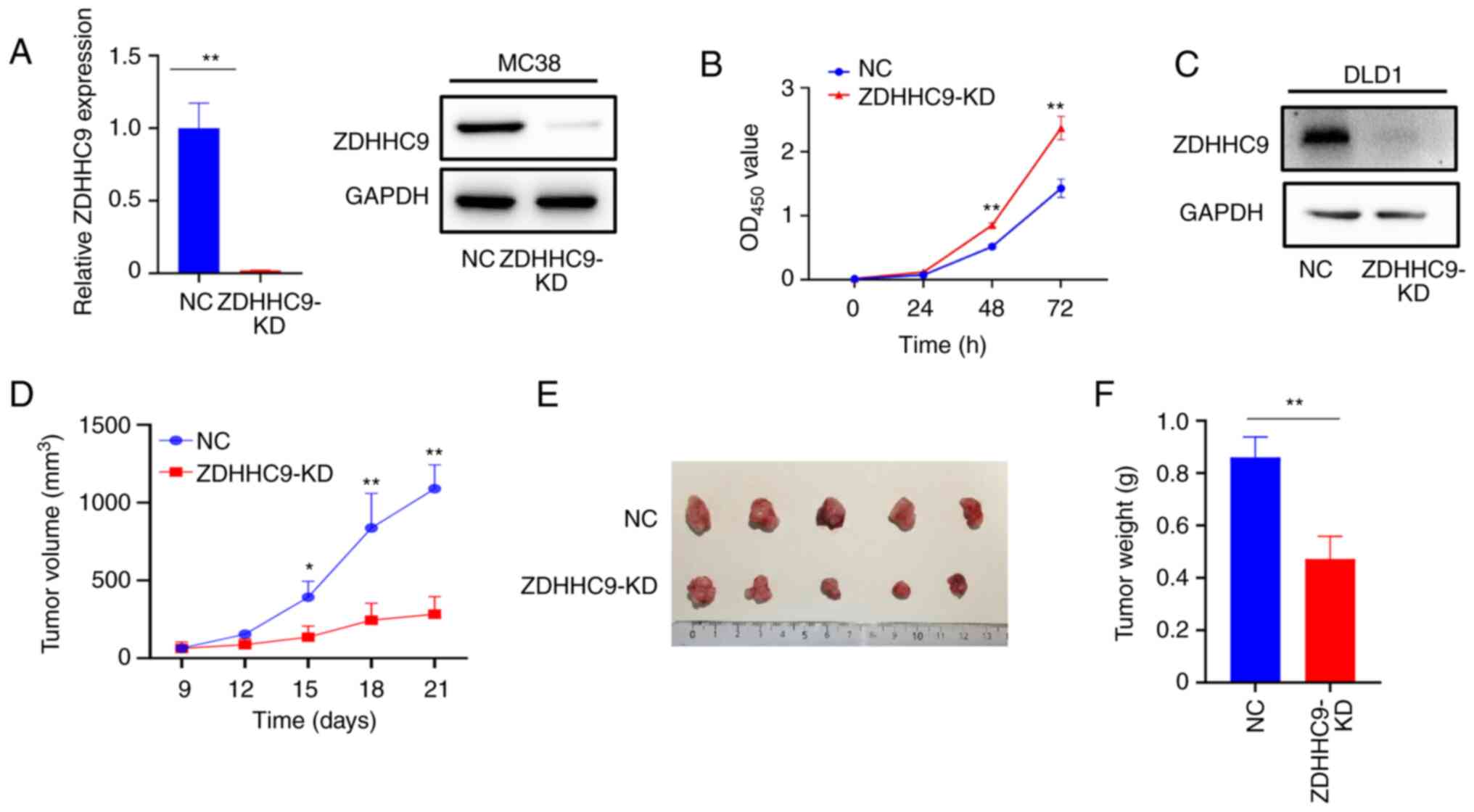
To determine whether ZDHHC9 affected colon cancer
growth, CRISPR-mediated knockdown was performed to knock down the
expression of ZDHHC9 in MC38 mouse colon cancer cells. As shown in
Fig. 2A, the mRNA and protein
expression levels of ZDHHC9 in MC38 cells were markedly lower
following knock down. A cell proliferation assay was performed to
observe the effect of ZDHHC9 expression on the in vitro
proliferation of MC38 cells. As shown in Fig. 2B, knock down of ZDHHC9 expression
increased the proliferation of MC38 cells. A transplantation tumor
model was then established in C57BL/6 mice by subcutaneously
injecting ZDHHC9-KD or control MC38 cells. The mice were sacrificed
21 days after injection. ZDHHC9 was then knocked down in DLD1 cells
(Fig. 2C). As shown in Fig. 2D-F, the growth of ZDHHC9-KD MC38
cells in vivo was significantly decreased compared with that
of mice injected with control MC38 cells. These results suggest
that ZDHHC9 plays an important role in the occurrence and
development of colon cancer.
Bioinformatics analysis of the
relationship between ZDHHC9 and the immune system
The data in Fig. 2
shows that ZDHHC9 differentially affected the growth of MC38 cells
in vitro and in vivo. ZDHHC9 has been reported to
protect breast cancer cells from the immune surveillance of T cells
(14). It was hypothesized that
ZDHHC9 promoted the in vivo growth of MC38 colon cancer
cells by affecting the immune response against MC38 cells. To
investigate the role of ZDHHC9 in immunity against colon cancer,
TIMER was used to analyze the relationship between ZDHHC9
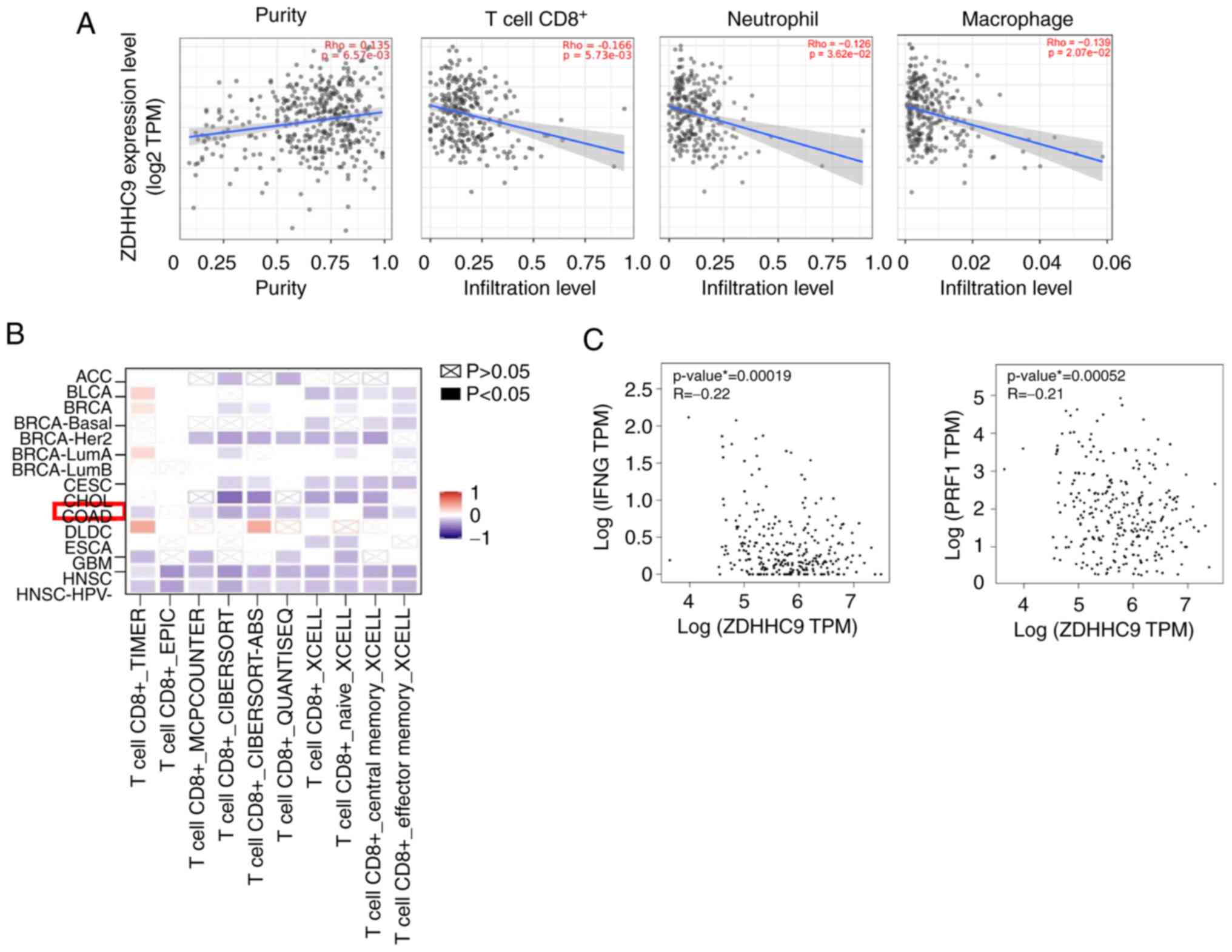
expression and immune infiltration. As shown in Fig. 3A, ZDHHC9 was significantly
positively correlated with tumor purity in colon adenocarcinoma
(COAD; cor.=0.135, P=6.57×10−3), indicating that ZDHHC9
was primarily expressed in tumor cells. By contrast, a higher
ZDHHC9 expression in COAD was significantly negatively correlated
with the infiltration of immune cells, particularly CD8+
T cells (cor.=−0.166, P=2.36×10−16), neutrophils
(cor.=−0.126, P=3.62×10−15), and macrophages
(cor.=−0.139, P=2.07×10−15). Consistently, multiple
databases showed that ZDHHC9 expression was significantly
negatively correlated with CD8+ T cells in COAD
(Fig. 3B). Furthermore, ZDHHC9
expression was markedly negatively correlated with IFN-γ and
perforin-1 (PRF1) expression in COAD tissues (Fig. 3C). These results indicated that
CD8+ T cell infiltration and activation were negatively
correlated with ZDHHC9 expression in colon cancer tissues. We
analyzed the function of ZDHHC9 in the immune system using
bioinformatics tools and found that it affected CD8+ T
cells, neutrophils, and macrophages, but the effect on
CD8+ T cells was the most notable, thus, CD8+
T cells were chosen for further analysis.
ZDHHC9 promotes tumor growth by
inhibiting the response of T cells
To investigate whether ZDHHC9 affected the function
of T cells, T-cell killing experiments were performed.
CD8+ T cells were isolated from OT-I mice and
co-cultured with MC38 cells treated with OVA (257–264 aa) peptide.
As shown in Fig. 4A, the mortality
rate of MC38 cells was significantly increased when co-cultured
with OT-I T cells at E:T ratios of 2:1, 1:1, and 1:2 for 6 h. The
method of gating cells is demonstrated in Fig. S1. The inhibition of ZDHHC9
expression significantly increased the mortality of MC38 cells.
These data suggested that the expression of ZDHHC9 in tumor cells
could protect the cells from an attack by CD8+ T cells,
thus playing an important role in suppressing tumor immunity.
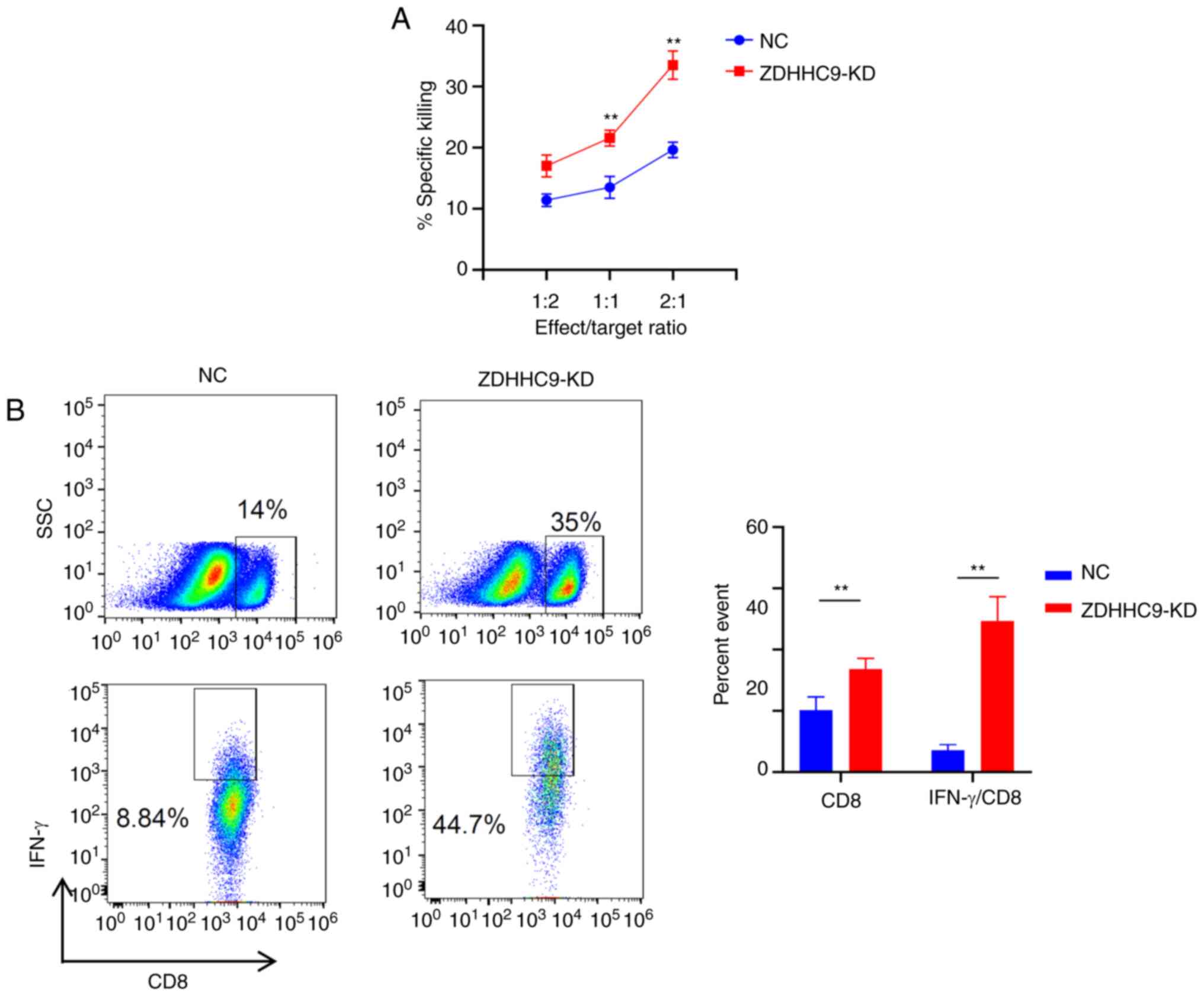 | Figure 4.ZDHHC9 promotes tumor growth by
inhibiting the T cell response. (A) T cell killing experiments
in vitro. ZDHHC9-deficient and NC cells were co-cultured
with OT-1 T cells at an E:T ratio of 1:2, 1:1, and 2:1 for 6 h, and
an FVS780 fluorescent antibody was used to detect the mortality
rate of tumor cells. (B) Detection of CD8 and IFN-γ secreted by CD8
in tumor-infiltrating lymphocytes using flow cytometry. On the 27th
day, the tumor tissue was excised, digested, and flow cytometry was
used to detect CD8 and IFN-γ expression in the cells (n=3).
**P<0.01. ZDHHC9, zßinc finger DHHC-type palmitoyltransferase 9;
NC, negative control; KD, knockdown; NC, negative control; SSC,
side scatter; E:T, Effect: Target. |
FACS analysis was then used to examine the changes
that occurred in intratumoral immune cell infiltration. The
distribution of different immune cell subpopulations was evaluated
in the tumor microenvironment. In tumors harvested from mice
injected with ZDHHC9-KD cells, the percentage of CD8+
cytotoxic T cells was significantly higher than that in the control
group (NC) (Fig. 4B). Flow
cytometry was further used to analyze the effect of ZDHHC9 on the
ability of CD8+ T cells to secrete cytokines. It was
found that ZDHHC9 knockdown in MC38 cells markedly increased the
secretion of IFN-γ by CD8+ T cells (Fig. 4B). These results showed that ZDHHC9
molecules can modulate the tumor microenvironment by repressing the
activation and function of CD8+ T cells.
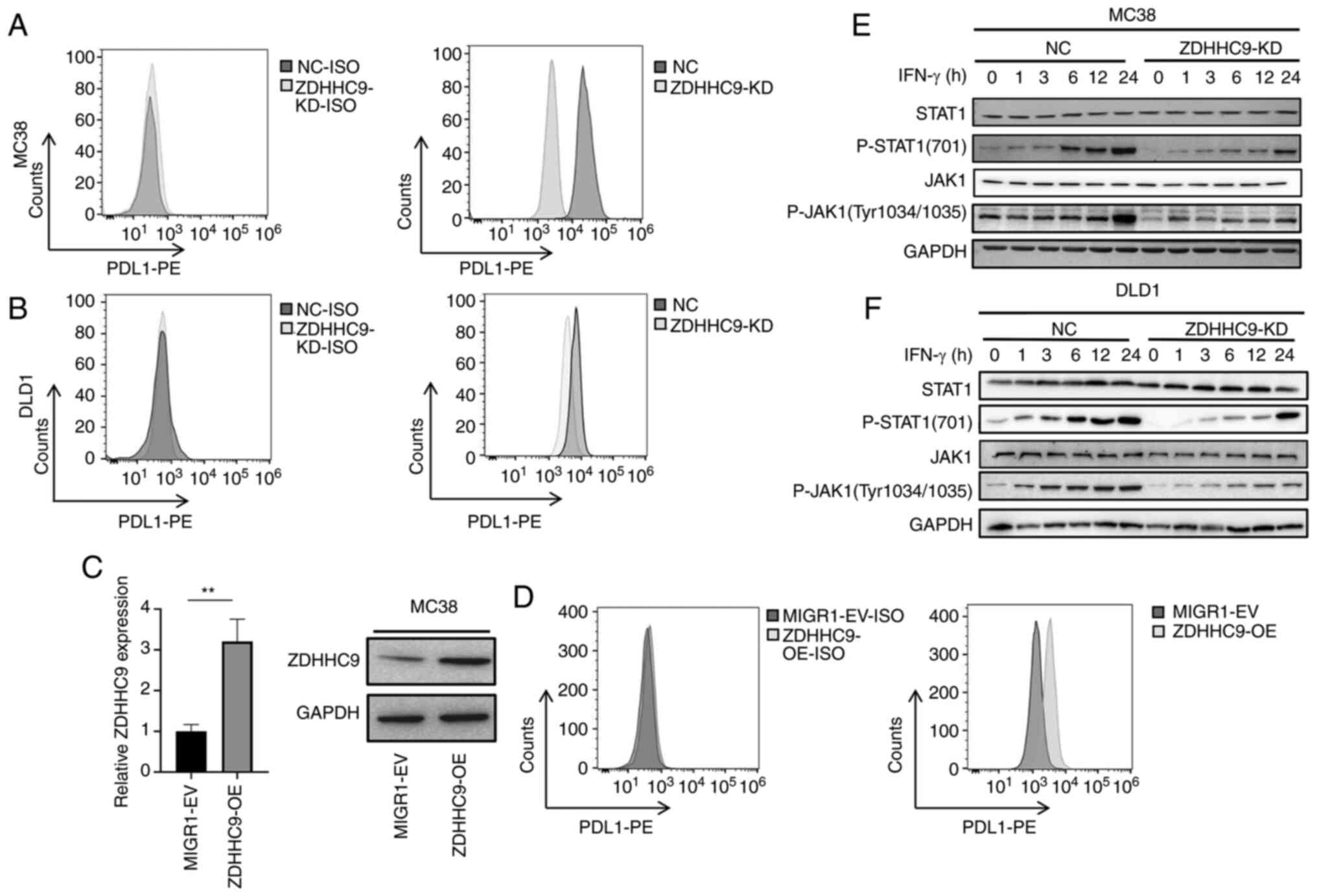
ZDHHC9 promotes PD-L1 expression
The above experimental results showed that ZDHHC9
inhibited the activation of CD8+ T cells in the tumor
microenvironment in colon cancer, thus promoting tumor growth in
vivo. The mechanism through which ZDHHC9 in tumor cells
promoted T cells to induce immune tolerance was next assessed. The
expression levels of PD-L1 protein were detected in ZDHHC9-KD MC38
cells. The results showed that the expression levels of PD-L1 in
colon cancer cells with ZDHHC9 deletion were lower than that in
control cells (Fig. 5A). ZDHHC9
expression was knocked down in the DLD1 cells (Fig. 2D). The expression levels of PD-L1
in DLD-1 cells were reduced following ZDHHC9 knockdown compared
with the control cells (Fig. 5B).
A MIGR1-ZDHHC9-MC38 cell line overexpressing ZDHHC9 was next
constructed (Fig. 5C). In MC38
cells overexpressing ZDHHC9, the expression of PD-L1 was also
increased (Fig. 5D), which further
confirmed the positive regulation of ZDHHC9 on the expression of
PD-L1.
To explore the mechanism by which ZDHHC9 upregulated
PD-L1 expression, the effect of ZDHHC9 on the expression of PD-L1
and activation of the JAK/STAT1 signaling pathway was measured. As
ZDHHC9 increased PD-L1 expression, and as PD-L1 gene expression is
regulated by the transcription factor STAT1 (17), whether ZDHHC9 affected the
activation of STAT1 was assessed. As anticipated, ZDHHC9 knockdown
markedly decreased the phosphorylation levels of STAT1 and JAK1 in
MC38 and DLD-1 cells (Fig. 5E and
F).
Discussion
In the present study, data from TCGA was used to
analyze the expression of ZDHHC9 in colon cancer. It was found that
ZDHHC9 expression was significantly increased in colon cancer and
was negatively correlated with the survival time of patients with
colon cancer. The knockdown of ZDHHC9 expression increased the
proliferation of MC38 cells in vitro; however, the knockdown
of ZDHHC9 expression decreased the growth of MC38 cells in
vivo.
The relationship between ZDHHC9 expression and the
activation of the immune system in colon cancer tissues was
analyzed, and it was found that ZDHHC9 expression in colon cancer
was negatively correlated with CD8+ T cell infiltration
and the expression of T cell effector molecules, such as IFN-γ.
Next, the effect of ZDHHC9 on tumor growth was
assessed in vivo using a subcutaneous transplantation model
of colon cancer in mice, and it was found that ZDHHC9 promoted
tumor growth in mice and significantly increased the percentage of
IFN-γ+/CD8+ T cells in the tumor
microenvironment. Through a specific T cell killing model, it was
found that ZDHHC9-deficient tumor cells were vulnerable to
cytotoxic T cells in vitro. These in vivo and in
vitro experimental results are consistent with the results of
TCGA correlation analysis, suggesting that ZDHHC9 promotes the
growth of colon cancer by inhibiting the T cell-mediated tumor
immune response.
It has been reported that ZDHHC9 increases the
protein expression of PD-L1 by palmitoylating and stabilizing PD-L1
in breast cancer (14). The
expression of PD-L1 in colon cancer cells was detected, and it was
found that ZDHHC9 increased the protein expression levels of PD-L1.
Further experiments showed that ZDHHC9 increased the activation of
the JAK1/STAT1 pathway, which has been demonstrated to increase
PD-L1 expression. It was therefore concluded that ZDHHC9 may
increase PD-L1 expression and affect the function of
CD8+ T cells.
In the present study, it was determined that ZDHHC9
can play a role in tumor immunity and inhibit the killing effect of
CD8+ T cells to promote tumor growth in colon cancer.
Only MC38 cells were used to evaluate the effect of ZDHHC9 on cell
proliferation and the association with the immune system. Thus, the
lack of evaluation of ZDHHC9 in DLD1 cells can be considered a
limitation and an area for further study. However, the tumor immune
microenvironment is a complex environment (16,18),
with B cells, tumor-associated macrophages, myeloid-derived
suppressor cells, regulatory T cells, neutrophils, and natural
killer cells (19). In addition to
PD-L1, tumor cells express numerous molecules that affect the tumor
immune microenvironment. Whether ZDHHC9 regulates other molecules
in addition to PD-L1 to affect T-cell tumor immunity needs further
investigation.
In conclusion, the results of the present study
suggested that ZDHHC9 can inhibit CD8+ T cells to
promote colon cancer and that it can increase PD-L1 expression
through the JAK1/STAT1 pathway.
Supplementary Material
Supporting Data
Acknowledgements
Not applicable.
Funding
This study was supported by grants from the National Key
Research and Development Program of China (grant nos.
2016YFA0502201 and 2018YFC1002801), National Natural Science
Foundation of China (grant nos. 81571550 and 81771698) and Shanghai
Key Laboratory of Cell Engineering (grant no. 14DZ2272300).
Availability of data and materials
The datasets used and/or analyzed during the present
study are available from the corresponding author on reasonable
request.
Authors' contributions
HC and HA designed the experiments. XC, LZ, DY and
SC obtained the data and contributed to the data analysis. XC
drafted the manuscript. HC and HA performed critical revision of
the manuscript. GW, QY, XM, JX performed the statistical analysis.
Administrative and technical support was provided by GW, QY, XM,
JX, HC and HA were responsible for the study concept and design. XC
and SC confirmed the authenticity of all the raw data. All authors
read and approved the final manuscript.
Ethics approval and consent to
participate
This study was approved by the Committee on Ethics
of Medicine, Naval Medical University, PLA. All experiments
involving mice were conducted in accordance with the National
Institute of Health Guide for the Care and Use of Laboratory
Animals and were approved by the Scientific Investigation Board of
the Naval Medical University (Shanghai, China).
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Chen SJ, Wang SC and Chen YC: The
immunotherapy for colorectal cancer, lung cancer and pancreatic
cancer. Int J Mol Sci. 22:128362021. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Fiorica F, Belluomini L, Giuliani J,
Urbini B, Milella M, Frassoldati A, Pilotto S and Giorgi C:
Abscopal effect and resistance reversion in nivolumab-treated
non-small-cell lung cancer undergoing palliative radiotherapy: A
case report. Immunotherapy. 13:971–976. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Morad G, Helmink BA, Sharma P and Wargo
JA: Hallmarks of response, resistance, and toxicity to immune
checkpoint blockade. Cell. 185:5762022. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Dersh D, Holly J and Yewdell JW: A few
good peptides: MHC class I-based cancer immunosurveillance and
immunoevasion. Nat Rev Immunol. 21:116–128. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Watson RA, Tong O, Cooper R, Taylor CA,
Sharma PK, de Los Aires AV, Mahé EA, Ruffieux H, Nassiri I,
Middleton MR and Fairfax BP: Immune checkpoint blockade sensitivity
and progression-free survival associates with baseline CD8(+) T
cell clone size and cytotoxicity. Sci Immunol. 6:eabj88252021.
View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Kouskou M, Thomson DM, Brett RR, Wheeler
L, Tate RJ, Pratt JA and Chamberlain LH: Disruption of the Zdhhc9
intellectual disability gene leads to behavioural abnormalities in
a mouse model. Exp Neurol. 308:35–46. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Masurel-Paulet A, KalscheuerV M, Lebrun N,
Hu H, Levy F, Thauvin-Robinet C, Darmency-Stamboul V, El Chehadeh
S, Thevenon J, Chancenotte S, et al: Expanding the clinical
phenotype of patients with a ZDHHC9 mutation. Am J Med Genet A.
64:789–795. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Hawkins E, Akarca D, Zhang M, Brkić D,
Woolrich M, Baker K and Astle D: Functional network dynamics in a
neurodevelopmental disorder of known genetic origin. Hum Brain
Mapp. 41:530–544. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Shimell JJ, Shah BS, Cain SM, Thouta S,
Kuhlmann N, Tatarnikov I, Jovellar DB, Brigidi GS, Kass J,
Milnerwood AJ, et al: The X-linked intellectual disability gene
Zdhhc9 is essential for dendrite outgrowth and inhibitory synapse
formation. Cell Rep. 29:2422–2437. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Schirwani S, Wakeling E, Smith K, Study
DDD and Balasubramanian M: Expanding the molecular basis and
phenotypic spectrum of ZDHHC9-associated X-linked intellectual
disability. Am J Med Genet A. 176:1238–1244. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Liu P, Jiao B, Zhang R, Zhao H, Zhang C,
Wu M, Li D, Zhao X, Qiu Q, Li J and Ren R: Palmitoylacyltransferase
Zdhhc9 inactivation mitigates leukemogenic potential of oncogenic
Nras. Leukemia. 30:1225–1228. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Chen X, Ma H, Wang Z, Zhang S, Yang H and
Fang Z: EZH2 palmitoylation mediated by ZDHHC5 in p53-mutant glioma
drives malignant development and progression. Cancer Res.
77:4998–5010. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Noland CL, Gierke S, Schnier PD, Murray J,
Sandoval WN, Sagolla M, Dey A, Hannoush RN, Fairbrother WJ and
Cunningham CN: Palmitoylation of TEAD transcription factors is
required for their stability and function in hippo pathway
signaling. Structure. 24:179–186. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Yang Y, Hsu JM, Sun L, Chan LC, Li CW, Hsu
JL, Wei Y, Xia W, Hou J, Qiu Y and Hung MC: Palmitoylation
stabilizes PD-L1 to promote breast tumor growth. Cell Res.
29:83–86. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Mansilla F, Birkenkamp-Demtroder K,
Kruhøffer M, Sørensen FB, Andersen CL, Laiho P, Aaltonen LA,
Verspaget HW and Orntoft TF: Differential expression of DHHC9 in
microsatellite stable and instable human colorectal cancer
subgroups. Br J Cancer. 96:1896–1903. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Sharma P and Allison JP: Dissecting the
mechanisms of immune checkpoint therapy. Nat Rev Immunol. 20:75–76.
2020. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Cerezo M, Guemiri R, Druillennec S,
Girault I, Malka-Mahieu H, Shen S, Allard D, Martineau S, Welsch C,
Agoussi S, et al: Translational control of tumor immune escape via
the eIF4F-STAT1-PD-L1 axis in melanoma. Nat Med. 24:1877–1886.
2018. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Lv B, Wang Y, Ma D, Cheng W, Liu J, Yong
T, Chen H and Wang C: Immunotherapy: Reshape the tumor immune
microenvironment. Front Immunol. 13:8441422022. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Yenyuwadee S, Aliazis K, Wang Q,
Christofides A, Shah R, Patsoukis N and Boussiotis VA: Immune
cellular components and signaling pathways in the tumor
microenvironment. Semin Cancer Biol. 86:187–201. 2022. View Article : Google Scholar : PubMed/NCBI
|