|
1
|
Sung H, Ferlay J, Siegel RL, Laversanne M,
Soerjomataram I, Jemal A and Bray F: Global cancer statistics 2020:
GLOBOCAN estimates of incidence and mortality worldwide for 36
cancers in 185 countries. CA Cancer J Clin. 71:209–249. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Arnold M, Sierra MS, Laversanne M,
Soerjomataram I, Jemal A and Bray F: Global patterns and trends in
colorectal cancer incidence and mortality. Gut. 66:683–691. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Kocarnik JM, Shiovitz S and Phipps AI:
Molecular phenotypes of colorectal cancer and potential clinical
applications. Gastroenterol Rep (Oxf). 3:269–276. 2015.PubMed/NCBI
|
|
4
|
Guinney J, Dienstmann R, Wang X, de
Reyniès A, Schlicker A, Soneson C, Marisa L, Roepman P, Nyamundanda
G, Angelino P, et al: The consensus molecular subtypes of
colorectal cancer. Nat Med. 21:1350–1356. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Sawayama H, Miyamoto Y, Ogawa K, Yoshida N
and Baba H: Investigation of colorectal cancer in accordance with
consensus molecular subtype classification. Ann Gastroenterol Surg.
4:528–539. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Yuan ZX, Wang XY, Qin QY, Chen DF, Zhong
QH, Wang L and Wang JP: The prognostic role of BRAF mutation in
metastatic colorectal cancer receiving anti-EGFR monoclonal
antibodies: A meta-analysis. PLoS One. 8:e659952013. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Afrăsânie VA, Marinca MV, Alexa-Stratulat
T, Gafton B, Păduraru M, Adavidoaiei AM, Miron L and Rusu C: KRAS,
NRAS, BRAF, HER2 and microsatellite instability in metastatic
colorectal cancer-practical implications for the clinician. Radiol
Oncol. 53:265–274. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Gong J, Cho M, Sy M, Salgia R and Fakih M:
Molecular profiling of metastatic colorectal tumors using
next-generation sequencing: A single-institution experience.
Oncotarget. 8:42198–42213. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Ullah S, John P and Bhatti A: MicroRNAs
with a role in gene regulation and in human diseases. Mol Biol Rep.
41:225–232. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Bagga S, Bracht J, Hunter S, Massirer K,
Holtz J, Eachus R and Pasquinelli AE: Regulation by let-7 and lin-4
miRNAs results in target mRNA degradation. Cell. 122:553–563. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Bartel DP: Metazoan MicroRNAs. Cell.
173:20–51. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Peng Y and Croce CM: The role of MicroRNAs
in human cancer. Signal Transduct Target Ther. 1:150042016.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Zhang C, Sun C, Zhao Y, Wang Q, Guo J, Ye
B and Yu G: Overview of MicroRNAs as diagnostic and prognostic
biomarkers for high-incidence cancers in 2021. Int J Mol Sci.
23:113892022. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Garzon R, Fabbri M, Cimmino A, Calin GA
and Croce CM: MicroRNA expression and function in cancer. Trends
Mol Med. 12:580–587. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Backes C, Meese E and Keller A: Specific
miRNA disease biomarkers in blood, serum and plasma: Challenges and
prospects. Mol Diagn Ther. 20:509–518. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Jorgensen BG and Ro S: MicroRNAs and
‘sponging’ competitive endogenous RNAs dysregulated in colorectal
cancer: Potential as noninvasive biomarkers and therapeutic
targets. Int J Mol Sci. 23:21662022. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Mo WY and Cao SQ: MiR-29a-3p: A potential
biomarker and therapeutic target in colorectal cancer. Clin Transl
Oncol. 25:563–577. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Wang CZ, Deng F, Li H, Wang DD, Zhang W,
Ding L and Tang JH: MiR-101: A potential therapeutic target of
cancers. Am J Transl Res. 10:3310–3321. 2018.PubMed/NCBI
|
|
19
|
Yin H, Sun Y, Wang X, Park J, Zhang Y, Li
M, Yin J, Liu Q and Wei M: Progress on the relationship between
miR-125 family and tumorigenesis. Exp Cell Res. 339:252–260. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Wang D, Wang X, Song Y, Si M, Sun Y, Liu
X, Cui S, Qu X and Yu X: Exosomal miR-146a-5p and miR-155-5p
promote CXCL12/CXCR7-induced metastasis of colorectal cancer by
crosstalk with cancer-associated fibroblasts. Cell Death Dis.
13:3802022. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Yamada A, Horimatsu T, Okugawa Y, Nishida
N, Honjo H, Ida H, Kou T, Kusaka T, Sasaki Y, Yagi M, et al: Serum
miR-21, miR-29a, and miR-125b Are promising biomarkers for the
early detection of colorectal neoplasia. Clin Cancer Res.
21:4234–4242. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Peng X, Wang J, Zhang C, Liu K, Zhao L,
Chen X, Huang G and Lai Y: A three-miRNA panel in serum as a
noninvasive biomarker for colorectal cancer detection. Int J Biol
Markers. 35:74–82. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Rawlings-Goss RA, Campbell MC and Tishkoff
SA: Global population-specific variation in miRNA associated with
cancer risk and clinical biomarkers. BMC Med Genomics. 7:532014.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Panico A, Tumolo MR, Leo CG, Donno A,
Grassi T, Bagordo F, Serio F, Idolo A, Masi R, Mincarone P and
Sabina S: The influence of lifestyle factors on miRNA expression
and signal pathways: A review. Epigenomics. 13:145–164. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Rovčanin Dragović I, Popović N, Ždralević
M, Radulović L, Vuković T, Marzano F, Tullo A and Radunović M:
Inflammation-related microRNAs-146a and −155 are upregulated in
mild cognitive impairment subjects among older age population in
montenegro. J Alzheimers Dis. 90:625–638. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Andersen CL, Jensen JL and Ørntoft TF:
Normalization of real-time quantitative reverse transcription-PCR
data: A model-based variance estimation approach to identify genes
suited for normalization, applied to bladder and colon cancer data
sets. Cancer Res. 64:5245–5250. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Huang HY, Lin YCD, Cui S, Huang Y, Tang Y,
Xu J, Bao J, Li Y, Wen J, Zuo H, et al: miRTarBase update 2022: An
informative resource for experimentally validated miRNA-target
interactions. Nucleic Acids Res. 50(D1): D222–D230. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
29
|
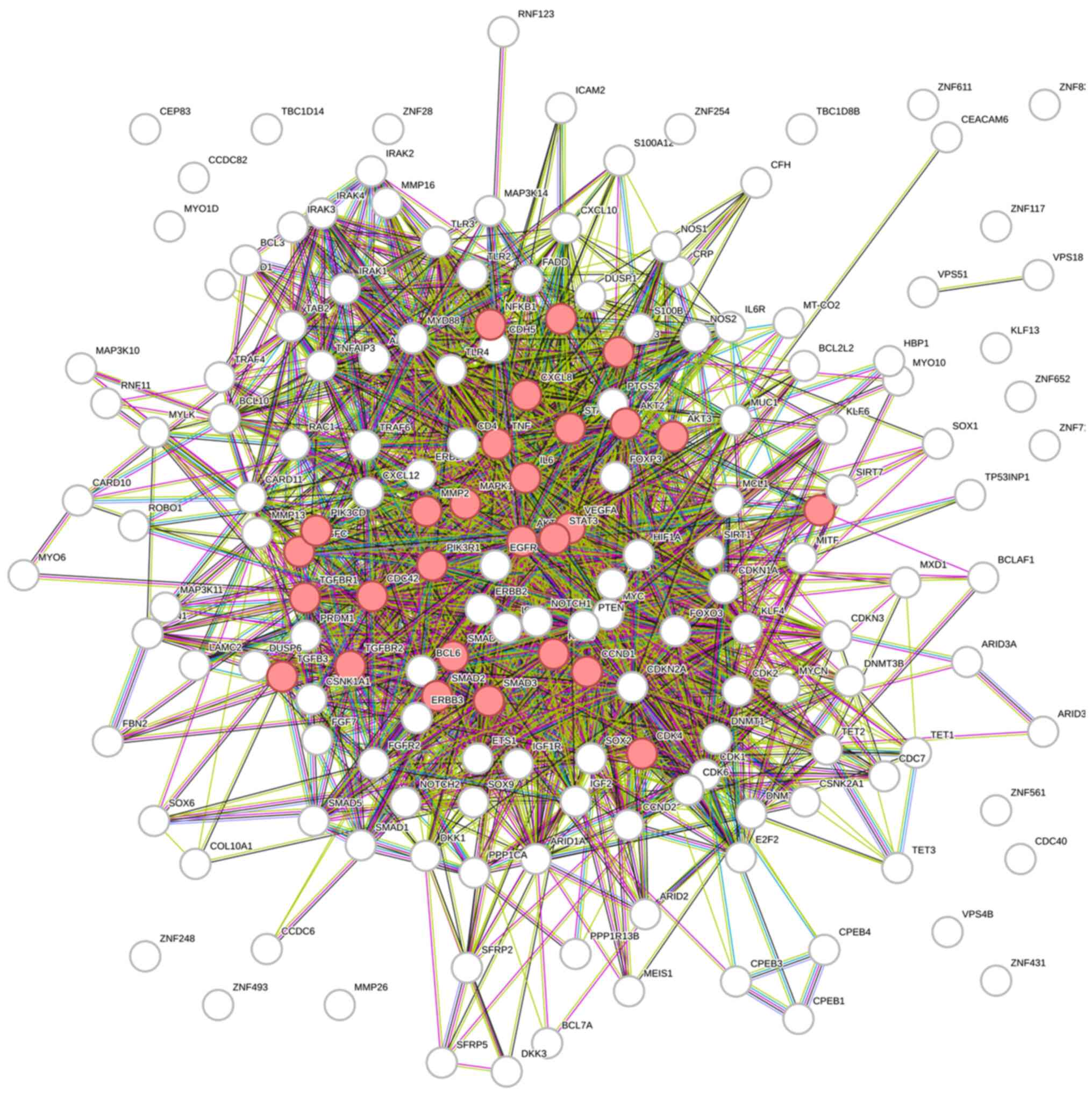
Szklarczyk D, Gable AL, Lyon D, Junge A,
Wyder S, Huerta-Cepas J, Simonovic M, Doncheva NT, Morris JH, Bork
P, et al: STRING v11: Protein-protein association networks with
increased coverage, supporting functional discovery in genome-wide
experimental datasets. Nucleic Acids Res. 47(D1): D607–D613. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Wang H, Li X, Li T, Wang L, Wu X, Liu J,
Xu Y and Wei W: Multiple roles of microRNA-146a in immune responses
and hepatocellular carcinoma (review). Oncol Lett. 18:5033–5042.
2019.PubMed/NCBI
|
|
31
|
Szklarczyk D, Gable AL, Nastou KC, Lyon D,
Kirsch R, Pyysalo S, Doncheva NT, Legeay M, Fang T, Bork P, et al:
The STRING database in 2021: Customizable protein-protein networks,
and functional characterization of user-uploaded gene/measurement
sets. Nucleic Acids Res. 49(D1): D605–D612. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
El-Far AH, Sroga G, Jaouni SKA and Mousa
SA: Role and mechanisms of RAGE-ligand complexes and
RAGE-inhibitors in cancer progression. Int J Mol Sci. 21:36132020.
View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Chandramouli A, Onyeagucha BC,
Mercado-Pimentel ME, Stankova L, Shahin NA, LaFleur BJ, Heimark RL,
Bhattacharyya AK and Nelson MA: MicroRNA-101 (miR-101)
post-transcriptionally regulates the expression of EP4 receptor in
colon cancers. Cancer Biol Ther. 13:175–183. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Cao H, Huang S, Liu A and Chen Z:
Up-regulated expression of miR-155 in human colonic cancer. J
Cancer Res Ther. 14:604–607. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Waghela BN, Vaidya FU, Ranjan K, Chhipa
AS, Tiwari BS and Pathak C: AGE-RAGE synergy influences programmed
cell death signaling to promote cancer. Mol Cell Biochem.
476:585–598. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Snelson M, Lucut E and Coughlan MT: The
role of AGE-RAGE signalling as a modulator of gut permeability in
diabetes. Int J Mol Sci. 23:17662022. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Li Y, Tan S, Shen Y and Guo L: miR-146a-5p
negatively regulates the IL-1β-stimulated inflammatory response via
downregulation of the IRAK1/TRAF6 signaling pathway in human
intestinal epithelial cells. Exp Ther Med. 24:6152022. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Garo LP, Ajay AK, Fujiwara M, Gabriely G,
Raheja R, Kuhn C, Kenyon B, Skillin N, Kadowaki-Saga R, Saxena S
and Murugaiyan G: MicroRNA-146a limits tumorigenic inflammation in
colorectal cancer. Nat Commun. 12:24192021. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Sathyanarayanan A, Chandrasekaran KS and
Karunagaran D: microRNA-146a inhibits proliferation, migration and
invasion of human cervical and colorectal cancer cells. Biochem
Biophys Res Commun. 480:528–533. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Lu D, Yao Q, Zhan C, Le-Meng Z, Liu H, Cai
Y, Tu C, Li X, Zou Y and Zhang S: MicroRNA-146a promote cell
migration and invasion in human colorectal cancer via
carboxypeptidase M/src-FAK pathway. Oncotarget. 8:22674–22684.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Chae YS, Kim JG, Lee SJ, Kang BW, Lee YJ,
Park JY, Jeon HS, Park JS and Choi GS: A miR-146a polymorphism
(rs2910164) predicts risk of and survival from colorectal cancer.
Anticancer Res. 33:3233–3239. 2013.PubMed/NCBI
|
|
42
|
Li MW, Gao L, Dang YW, Li P, Li ZY, Chen G
and Luo DZ: Protective potential of miR-146a-5p and its underlying
molecular mechanism in diverse cancers: A comprehensive
meta-analysis and bioinformatics analysis. Cancer Cell Int.
19:1672019. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Fellizar A, Refuerzo V, Ramos JD and
Albano PM: Expression of specific microRNAs in tissue and plasma in
colorectal cancer. J Pathol Transl Med. May 3–2022.(Epub ahead of
print). View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Wang A, Deng S, Chen X, Yu C, Du Q, Wu Y,
Chen G, Hu L, Hu C and Li Y: miR-29a-5p/STAT3 positive feedback
loop regulates TETs in colitis-associated colorectal cancer.
Inflamm Bowel Dis. 26:524–533. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Wang J, Chen X, Xie C, Sun M, Hu C, Zhang
Z, Luan L, Zhou J, Zhou J, Zhu X, et al: MicroRNA miR-29a inhibits
colon cancer progression by downregulating B7-H3 expression:
Potential molecular targets for colon cancer therapy. Mol
Biotechnol. 63:849–861. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
He D, Yue Z, Li G, Chen L, Feng H and Sun
J: Low serum levels of miR-101 are associated with poor prognosis
of colorectal cancer patients after curative resection. Med Sci
Monit. 24:7475–7481. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Chen LG, Xia YJ and Cui Y: Upregulation of
miR-101 enhances the cytotoxic effect of anticancer drugs through
inhibition of colon cancer cell proliferation. Oncol Rep.
38:100–108. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Gong J, Chu Y, Xu M, Huo J and Lv L:
Esophageal squamous cell carcinoma cell proliferation induced by
exposure to low concentration of cigarette smoke extract is
mediated via targeting miR-101-3p/COX-2 pathway. Oncol Rep.
35:463–471. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Zhang X, Li T, Han YN, Ge M, Wang P, Sun
L, Liu H, Cao T, Nie Y, Fan D, et al: miR-125b promotes colorectal
cancer migration and invasion by dual-targeting CFTR and CGN.
Cancers (Basel). 13:57102021. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Yu X, Shi W, Zhang Y, Wang X, Sun S, Song
Z, Liu M, Zeng Q, Cui S and Qu X: CXCL12/CXCR4 axis induced
miR-125b promotes invasion and confers 5-fluorouracil resistance
through enhancing autophagy in colorectal cancer. Sci Rep.
7:422262017. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Cappuzzo F, Sacconi A, Landi L, Ludovini
V, Biagioni F, D'Incecco A, Capodanno A, Salvini J, Corgna E,
Cupini S, et al: MicroRNA signature in metastatic colorectal cancer
patients treated with anti-EGFR monoclonal antibodies. Clin
Colorectal Cancer. 13:37–45.e4. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Zhou XG, Huang XL, Liang SY, Tang SM, Wu
SK, Huang TT, Mo ZN and Wang QY: Identifying miRNA and gene modules
of colon cancer associated with pathological stage by weighted gene
co-expression network analysis. Onco Targets Ther. 11:2815–2830.
2018. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Liu HN, Liu TT, Wu H, Chen YJ, Tseng YJ,
Yao C, Weng SQ, Dong L and Shen XZ: Serum microRNA signatures and
metabolomics have high diagnostic value in colorectal cancer using
two novel methods. Cancer Sci. 109:1185–1194. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Yagi T, Iinuma H, Hayama T, Matsuda K,
Nozawa K, Tsukamoto M, Shimada R, Akahane T, Tsuchiya T, Ozawa T
and Hashiguchi Y: Plasma exosomal microRNA-125b as a monitoring
biomarker of resistance to mFOLFOX6-based chemotherapy in advanced
and recurrent colorectal cancer patients. Mol Clin Oncol.
11:416–424. 2019.PubMed/NCBI
|
|
55
|
Nishida N, Yokobori T, Mimori K, Sudo T,
Tanaka F, Shibata K, Ishii H, Doki Y, Kuwano H and Mori M: MicroRNA
miR-125b is a prognostic marker in human colorectal cancer. Int J
Oncol. 38:1437–1443. 2011.PubMed/NCBI
|
|
56
|
Ak S, Tunca B, Tezcan G, Cecener G, Egeli
U, Yilmazlar T, Ozturk E and Yerci O: MicroRNA expression patterns
of tumors in early-onset colorectal cancer patients. J Surg Res.
191:113–122. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Zeng ZL, Lu JH, Wang Y, Sheng H, Wang YN,
Chen ZH, Wu QN, Zheng JB, Chen YX, Yang DD, et al: The lncRNA
XIST/miR-125b-2-3p axis modulates cell proliferation and
chemotherapeutic sensitivity via targeting Wee1 in colorectal
cancer. Cancer Med. 10:2423–2441. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Yu M, Yi Z, Chen S, Chen X and Xie X:
MIR4435-2HG, miR-125b-5p, and Sema4D axis affects the
aggressiveness of colorectal cancer cells. Folia Histochem
Cytobiol. 60:191–202. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
59
|
Cenariu D, Zimta AA, Munteanu R, Onaciu A,
Moldovan CS, Jurj A, Raduly L, Moldovan A, Florea A, Budisan L, et
al: Hsa-miR-125b therapeutic role in colon cancer is dependent on
the mutation status of the TP53 gene. Pharmaceutics. 13:6642021.
View Article : Google Scholar : PubMed/NCBI
|
|
60
|
Volinia S, Calin GA, Liu CG, Ambs S,
Cimmino A, Petrocca F, Visone R, Iorio M, Roldo C, Ferracin M, et
al: A microRNA expression signature of human solid tumors defines
cancer gene targets. Proc Natl Acad Sci USA. 103:2257–2261. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
61
|
Qu YL, Wang HF, Sun ZQ, Tang Y, Han XN, Yu
XB and Liu K: Up-regulated miR-155-5p promotes cell proliferation,
invasion and metastasis in colorectal carcinoma. Int J Clin Exp
Pathol. 8:6988–6994. 2015.PubMed/NCBI
|
|
62
|
Rodriguez A, Vigorito E, Clare S, Warren
MV, Couttet P, Soond DR, van Dongen S, Grocock RJ, Das PP, Miska
EA, et al: Requirement of bic/microRNA-155 for normal immune
function. Science. 316:608–611. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
63
|
Valeri N, Gasparini P, Fabbri M, Braconi
C, Veronese A, Lovat F, Adair B, Vannini I, Fanini F, Bottoni A, et
al: Modulation of mismatch repair and genomic stability by miR-155.
Proc Natl Acad Sci USA. 107:6982–6987. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
64
|
Xu Y, Han S, Lei K, Chang X, Wang K, Li Z
and Liu J: Anti-Warburg effect of rosmarinic acid via miR-155 in
colorectal carcinoma cells. Eur J Cancer Prev. 25:481–489. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
65
|
Liu J, Chen Z, Xiang J and Gu X:
MicroRNA-155 acts as a tumor suppressor in colorectal cancer by
targeting CTHRC1 in vitro. Oncol Lett. 15:5561–5568.
2018.PubMed/NCBI
|
|
66
|
Wang XJ, Zeng B, Lin S, Chen M and Chi P:
An integrated miRNA-lncRNA signature predicts the survival of stage
II colon cancer. Ann Clin Lab Sci. 49:730–739. 2019.PubMed/NCBI
|
|
67
|
Ulivi P, Canale M, Passardi A, Marisi G,
Valgiusti M, Frassineti GL, Calistri D, Amadori D and Scarpi E:
Circulating plasma levels of miR-20b, miR-29b and miR-155 as
predictors of bevacizumab efficacy in patients with metastatic
colorectal cancer. Int J Mol Sci. 19:3072018. View Article : Google Scholar : PubMed/NCBI
|
|
68
|
Chen J, Wang W, Zhang Y, Chen Y and Hu T:
Predicting distant metastasis and chemoresistance using plasma
miRNAs. Med Oncol. 31:7992014. View Article : Google Scholar : PubMed/NCBI
|
|
69
|
Hongliang C, Shaojun H, Aihua L and Hua J:
Correlation between expression of miR-155 in colon cancer and serum
carcinoembryonic antigen level and its contribution to recurrence
and metastasis forecast. Saudi Med J. 35:547–553. 2014.PubMed/NCBI
|
|
70
|
Lv ZC, Fan YS, Chen HB and Zhao DW:
Investigation of microRNA-155 as a serum diagnostic and prognostic
biomarker for colorectal cancer. Tumor Biol. 36:1619–1625. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
71
|
Zhang GJ, Xiao HX, Tian HP, Liu ZL, Xia SS
and Zhou T: Upregulation of microRNA-155 promotes the migration and
invasion of colorectal cancer cells through the regulation of
claudin-1 expression. Int J Mol Med. 31:1375–1380. 2013. View Article : Google Scholar : PubMed/NCBI
|