Introduction
Bladder urothelial carcinoma (BLCA) occurs in the
bladder mucosa, is the most common malignancy of the urinary tract,
and its incidence increases with patient age (1). There were estimated to be >81,400
new cases and 17,000 BLCA-associated deaths in the United States in
2020 (2). Although patients with
BLCA undergo aggressive treatment, including surgery, chemotherapy,
radiotherapy, and immunotherapy, the 5-year overall survival (OS)
rate remains unsatisfactory and recurrence and progression rates
following BLCA treatment remain high, which places a considerable
financial burden on the healthcare system and impacts the quality
of life of patients (3,4). Patient prognosis is difficult to
predict as there are no clinical biomarkers or parameters that can
reliably determine disease progression (5–7).
Therefore, the identification of new biomarkers is crucial for the
early diagnosis, prognostic assessment, and treatment of bladder
cancer.
Ephrins are a class of cell surface ligands that
mediate the migration, rejection, and adhesion of neuronal,
vascular, and epithelial cells by binding to members of the Eph
tyrosine kinase receptor family (8). A previous study has shown that Eph
receptors and ephrins serve a key role in cancer cell
proliferation, invasion, metastasis, and angiogenesis as signaling
molecules involved in axon guidance (9). As a member of the Ephrin family,
Ephrin A3 (EFNA3) potentially plays a role in the pathology of
several cancer types (10). High
expression of EFNA3 in gastric cancer cells is correlated with
poorer patient prognosis and is an effective prognostic indicator
for the responsiveness to immunotherapy in gastric cancer (11). Meanwhile, the upregulation of EFNA3
is regulated by microRNA-210, which promotes the proliferation and
invasion of oral cancer cells (12).
Collectively, the results of the aforementioned
studies suggest an association between the EFNA3 gene and an
unfavorable cancer prognosis. However, the precise expression
levels and prognostic significance of the EFNA3 gene in the context
of bladder cancer remain unclear. Thus, the aim of the present
study was to investigate the expression of EFNA3 in BLCA, assess
its correlation with clinicopathological characteristics, and
evaluate its impact on patient prognosis. To achieve these
objectives, a publicly accessible database was initially utilized
to examine EFNA3 expression in BLCA. Subsequently,
immunohistochemistry (IHC) was conducted on tissues from 491
patients with BLCA to confirm the association of EFNA3 with this
disease. The results of the present study indicated that EFNA3 may
serve as a promising biomarker for determining both prognosis and
treatment of BLCA.
Materials and methods
UALCAN database and TCGA
UALCAN (http://ualcan.path.uab.edu) is a database that uses
The Cancer Genome Atlas (TCGA) database to collect clinical data
from 31 cancer types. EFNA3 gene expression information and basic
clinical features were obtained from TCGA (https://portal.gdc.cancer.gov) (13). UALCAN can analyze the relative
expression of a gene across tumor and normal samples, as well as
across various tumor subgroups that are based on the cancer stage,
tumor grade, race, body weight, or other clinicopathological
features. This resource serves as a platform for in silico
validation of target genes and for identifying tumor
subgroup-specific candidate biomarkers (14). Data in the UALCAN database was
examined and filtered for the EFNA3 gene in BLCA using the
following criteria: i) ‘gene symbol: EFNA3’ and ii) ‘TCGA dataset:
Bladder urothelial carcinoma’. Then the following conditions were
selected: ‘Expression’; ‘Survival’; ‘Correlation’; and ‘Pan-cancer
view’.
Gene Ontology (GO) and Kyoto
Encyclopedia of Genes and Genomes (KEGG) analyses
LinkedOmics (http://www.linkedomics.org/login.php) uses the data of
32 cancer types from TCGA. These data can be used to analyze the
relationship between mRNA gene expression and features such as
methylation and mutation sites. In the present study, LinkedOmics
was used to obtain information on EFNA3 expression in BLCA by
setting the following filter conditions: i) ‘Gene: EFNA3’; ii)
‘Analysis Type: Cancer vs. Normal analysis’; iii) ‘Data Type:
mRNA’; iv) ‘Cancer Type: Bladder urothelial carcinoma’; v) ‘Gene
Summary: P-value<0.05, fold change=all, gene rank=top 10%’; and
vi) ‘Statistical method: Pearson correlation test’. Following this,
‘Gene Set Enrichment Analysis (GSEA)’ tools were selected. Then the
following conditions were selected: ‘GO analysis (biological
process)’; ‘GO analysis (cellular component)’; ‘GO analysis
(molecular function)’; and ‘KEGG pathway’.
GSEA
GSEA is a computational method used to determine
whether a predefined gene set exhibits statistically significant
differences between two biological states. In the present study,
GSEA was used to identify potential pathways associated with EFNA3
expression and prognosis in BLCA. For this, data associated with
BLCA were downloaded from TCGA (https://portal.gdc.cancer.gov/), and GSEA software
(version 4.1.0; Broad Institute and the University of California)
was used for analysis. The gene expression profiles of patients
with BLCA were divided into high and low-expression groups based on
the median value of EFNA3 expression. Gene sets were considered
significantly enriched when the false discovery rate (FDR) <0.25
and P. adjust <0.05.
Protein-protein interaction networks
(PPIs)
The interacting proteins associated with EFNA3, and
the interaction network was analyzed using the STRING search tool
(http://string-db.org/). The necessary data were
obtained by searching for protein names, species and other
necessary information.
Patient samples
The present study was a retrospective study in which
bladder cancer pathological specimens were obtained. A
retrospective analysis of tissue sections stored in the
pathological database was performed. A total of 491 samples of BLCA
tissues were collected from patients undergoing surgical resection
at The Zhejiang Provincial People's Hospital (Hangzhou, China)
between January 1998 and December 2011. The patient cohort
consisted of 432 males and 59 females, aged 35–79 years old
(median, 63.2 years old). No patients had undergone radiotherapy or
chemotherapy prior to surgical resection. Moreover, 80 control
samples were collected from adjacent tissues located >5 cm from
the tumor edge, which is usually defined clinically as normal
tissue >5 cm from the tumor margin (samples were collected
between January 1998 and December 2011) (Table I). The samples were subsequently
used to prepare tissue microarrays (TMAs), which were constructed
by Shanghai Xinchao Biological Technology Co., Ltd. The study was a
retrospective analysis, approved by the Ethics Committee of
Zhejiang Provincial People's Hospital and an informed consent
waiver was obtained from the Ethics Committee of Zhejiang
Provincial People's Hospital (approval no. QT2022423). The study
was performed in accordance with the Declaration of Helsinki
(15) and all patient details were
removed to protect patient privacy throughout the process.
 | Table I.Expression of Ephrin A3 mRNA in BLCA
and non-carcinomatous bladder tissues. |
Table I.
Expression of Ephrin A3 mRNA in BLCA
and non-carcinomatous bladder tissues.
|
| Ephrin A3
expression |
|---|
|
|
|
|---|
| Sample | Number | Negative | Positive | P-value |
|---|
| BLCA | 491 | 209 | 282 | <0.01 |
| Non-carcinomatous
bladder tissues | 80 | 55 | 25 |
|
IHC and evaluation of EFNA3 protein
expression
The changes in EFNA3 protein expression were studied
using the TMA consisting of the 491 human BLCA samples and 80
non-cancerous human bladder tissue samples. Briefly, slides were
incubated at 68°C for 2 h, followed by deparaffinization,
dehydration in xylene, and rehydration. Subsequently, the sections
were immersed in antigen retrieval buffer and boiled at 120°C in a
pressure cooker for 3 min. The sections were then treated with 3%
H2O2 for 15 min to quench endogenous
peroxidase activity and 1% bovine serum albumin to prevent
non-specific binding. Then, the sections were incubated with rabbit
anti-EFNA3 antibody (1:500; cat. no. PA5-86397; Thermo Fisher
Scientific, Inc.) at 4°C overnight, washed three times with PBS,
incubated with biotin-labeled secondary antibody for 20 min at room
temperature and then horseradish peroxidase-conjugated antibody for
another 20 min. Finally, tissue sections were stained with
3,3-diaminobenzidine, counterstained with hematoxylin for 8 min at
room temperature, dehydrated, washed, and mounted.
Manual IHC staining
quantification
Immunostaining was assessed and scored according to
the staining intensity by two independent observers blinded to the
clinical and pathological data. EFNA3-positive expression was
categorized into 4 classes based on staining intensity which were
assigned as follows: 0, no staining; 1, light yellow (weak
staining); 2, yellowish-brown (moderate staining); and 3, brown
(strong staining). The proportion of stained cells was scored as
follows: 0, <5% cells stained; 1, 6–25% cells stained; 2, 26–50%
cells stained; and 3, ≥50% cells stained. The staining index was
calculated by multiplying the intensity and proportion scores. A
staining index ≥4 was considered to reflect high EFNA3 expression
and an index of <4 was considered low EFNA3 expression.
Statistical analysis
SPSS (version 26.0; IBM Corp.) was used to perform
all statistical analyses. Categorical data were assessed for
statistical significance of differences using a χ2 test
or Fisher's exact test to compare the groups. Logistic regression
analyses were performed to determine the effects of the EFNA3 gene
and clinical factors on patient prognosis. Survival analysis was
performed using the Kaplan-Meier (KM) method combined with log-rank
test. KM analysis was the basis for plotting overall survival (OS)
curves. Patients were grouped according to EFNA3 expression and
assessed for survival status and OS time. KM survival curves were
plotted using GraphPad Prism version 10 (GraphPad Software, Inc.).
In addition, univariate and multivariate Cox regression analyses
were used to determine the relationship between EFNA3 gene
expression and clinicopathological characteristics of 491 patients
with BLCA. All tests were two-tailed statistical tests. P<0.05
was considered to indicate a statistically significant
difference.
Results
BLCA samples have significantly higher
EFNA3 expression levels than normal tissues
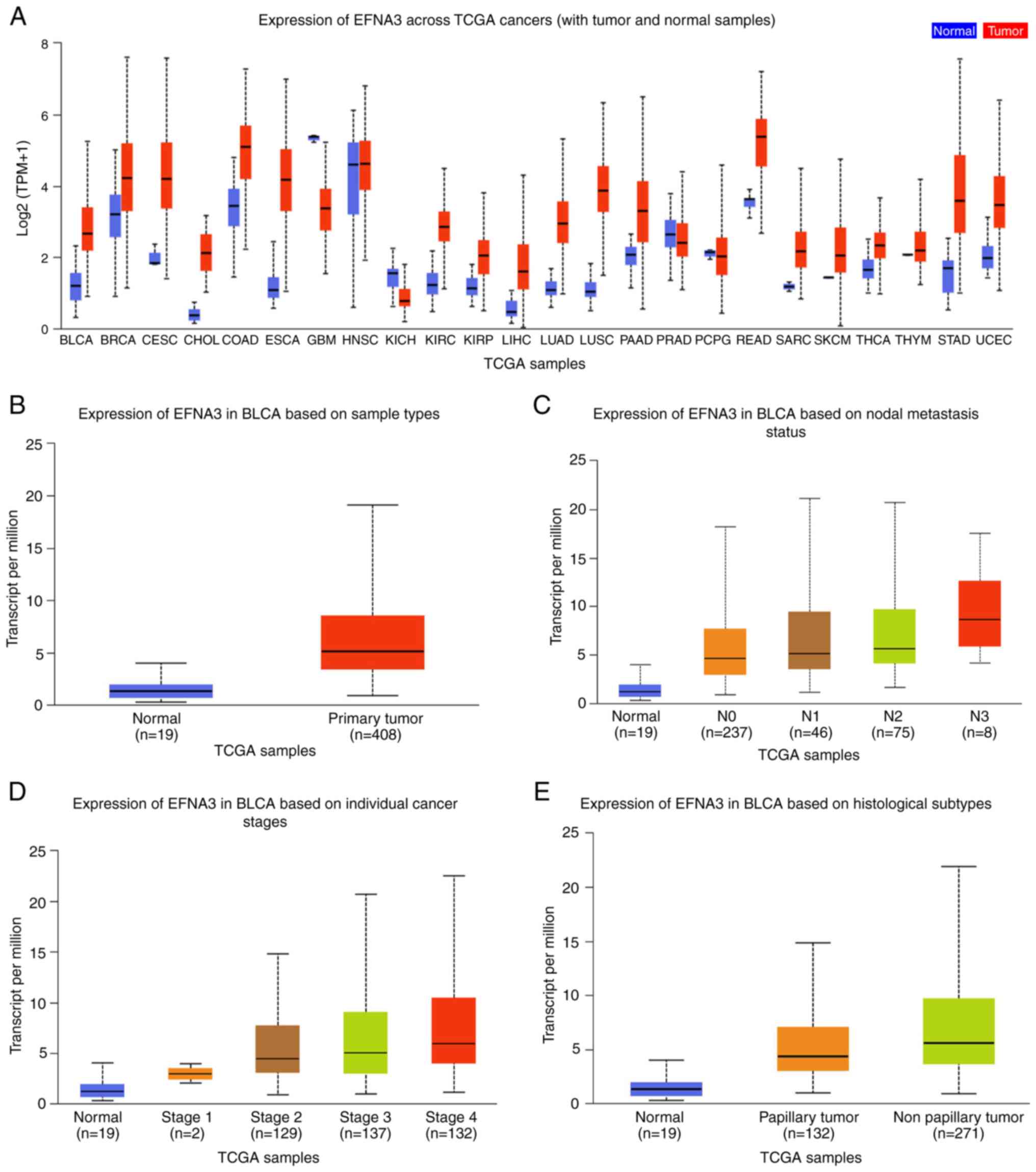
EFNA3 expression level differences in various tumor
and normal tissue samples were analyzed using TCGA data from
UALCAN. Upregulated EFNA3 expression was observed in a number of
cancer types, particularly in cervical squamous cell carcinoma and
esophageal cancer, while significant differences were observed
between the EFNA3 expression levels in bladder cancer and normal
tissues (Fig. 1A). After analyzing
19 normal bladder and 408 BLCA tissues in TCGA, the results
demonstrated that EFNA3 was expressed at high levels in patients
with advanced tumor-node-metastasis (TNM) stages and non-papillary
carcinomas (Fig. 1B-F).
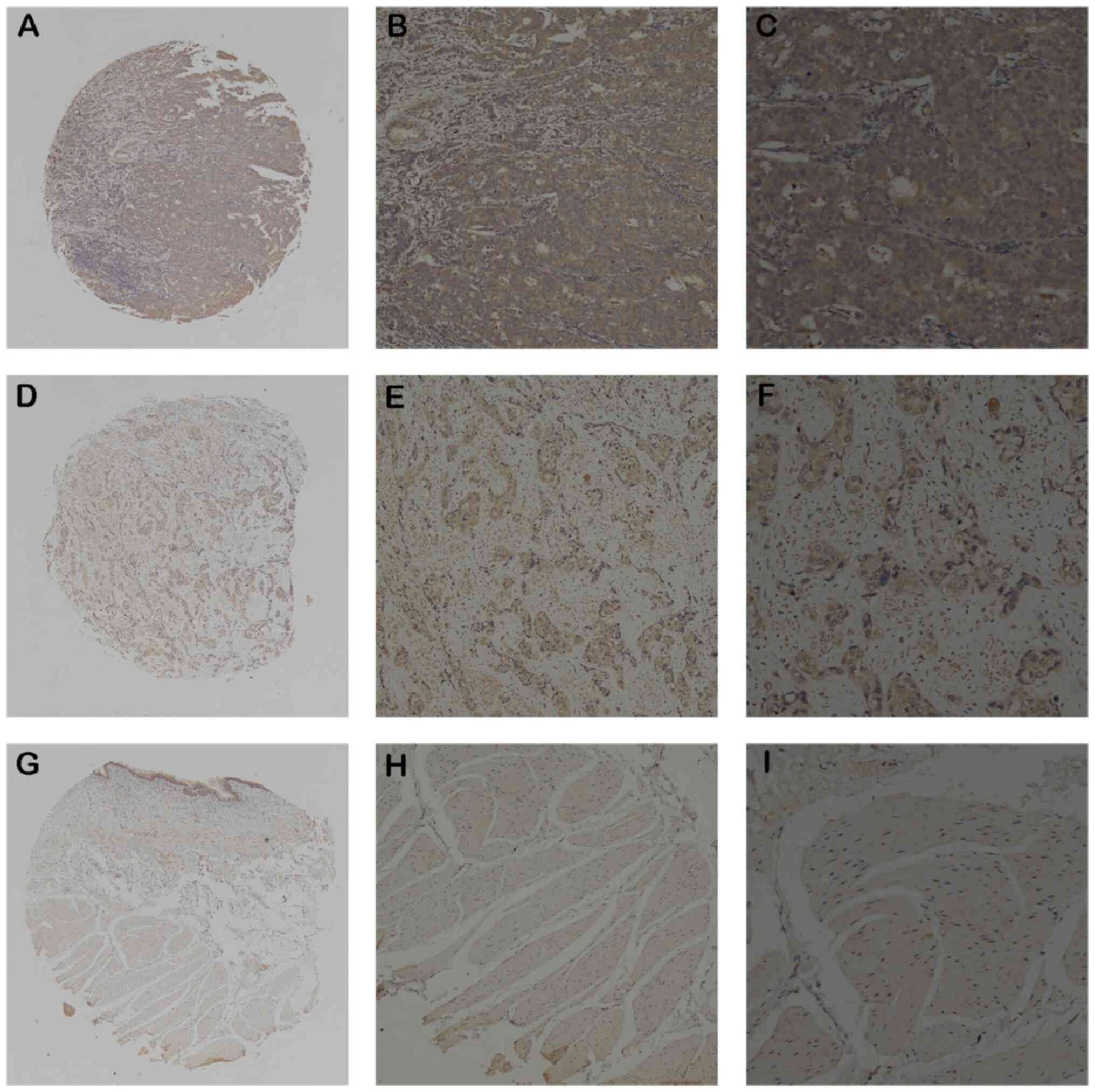
IHC was then used to assess the EFNA3 protein
expression levels in BLCA and normal bladder tissues from patients.
The results demonstrated that non-tumor tissues had little EFNA3
expression and BLCA tissues had markedly high expression (Fig. 2A-I). EFNA3 protein expression was
detected in 282 out of 491 (57.4%) BLCA samples and 25 out of 80
(31.3%) normal bladder tissues, indicating a significant
upregulation of expression in BLCA tissues. In addition, a positive
association between high EFNA3 expression and tumor size, invasion
depth, lymph node metastasis, distant metastasis, vascular
invasion, and histological grade was found, while there was no
significant association with age and sex (Table II).
 | Table II.Relationship between Ephrin A3
expression and the pathological parameters of bladder urothelial
carcinoma. |
Table II.
Relationship between Ephrin A3
expression and the pathological parameters of bladder urothelial
carcinoma.
|
| EFNA3
expression |
|---|
|
|
|
|---|
| Clinical
parameters | Low | High | χ2 | P-value |
|---|
| Sex |
|
| 0.280 | 0.597 |
|
Male | 182 | 250 |
|
|
|
Female | 27 | 32 |
|
|
| Age, years |
|
| 1.935 | 0.164 |
|
<55 | 87 | 100 |
|
|
|
≥55 | 122 | 182 |
|
|
| Size of tumor |
|
| 37.221 |
<0.0001c |
| <3
cm | 136 | 105 |
|
|
| ≥3
cm | 73 | 177 |
|
|
| Invasion depth |
|
| 26.182 |
<0.0001c |
|
Ta-T1 | 60 | 30 |
|
|
|
T2-T4 | 149 | 252 |
|
|
| Lymph node
metastasis |
|
| 8.658 | 0.003a |
| No | 171 | 198 |
|
|
|
Yes | 38 | 84 |
|
|
| Distant
metastasis |
|
| 10.264 | 0.001b |
| No | 198 | 242 |
|
|
|
Yes | 11 | 40 |
|
|
| Vascular
invasion |
|
| 19.687 |
<0.0001c |
|
Negative | 181 | 196 |
|
|
|
Positive | 28 | 86 |
|
|
| Histological
grade |
|
| 11.436 | 0.01a |
| Low
grade (I–II) | 104 | 132 |
|
|
| High
grade (III–IV) | 105 | 150 |
|
|
Clinical significance of EFNA3
expression in BLCA prognosis
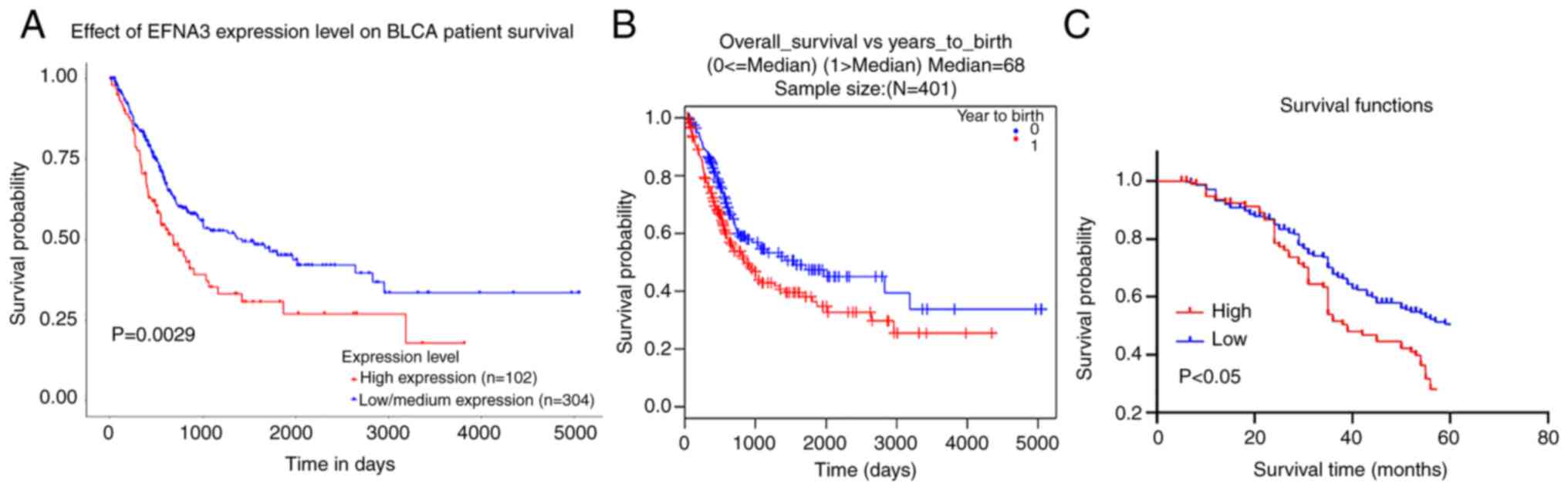
KM survival data demonstrated that the overall
survival time was shorter in the high EFNA3 expression group
compared with the low expression group (Fig. 3A and B). An analysis of the clinical
sample data demonstrated that the mean survival time of patients
with high EFNA3 expression was 38±1.03 months, which was
significantly shorter than that of the low expression group
(45±1.21 months) (Fig. 3C). ln
addition, a univariate analysis of factors affecting survival was
conducted and demonstrated that survival time was associated with
tumor size (P=0.026), lymph node metastasis (P<0.01), vascular
invasion (P<0.01), depth of invasion (P<0.01), distant
metastasis (P=0.009), histological grade (P<0.01), and EFNA3
expression (P<0.01) (Table
III). After entering these factors into a Cox proportional risk
regression model, the results demonstrated that vascular invasion,
histological grade, and depth of invasion were independent factors
affecting the prognosis of patients with BLCA (Table III). Although EFNA3 expression was
not an independent factor, it contributed to the development of
BLCA, which may be one of the adverse factors affecting patient
prognosis. The EFNA3 gene was also subjected to binary logistic
regression analysis and the results revealed that EFNA3 (P=0.036)
had some therapeutic value in the early diagnosis of bladder
cancer.
 | Table III.Univariate and multivariate analysis
of the correlation between clinicopathological parameters and
survival time of patients with bladder urothelial carcinoma. |
Table III.
Univariate and multivariate analysis
of the correlation between clinicopathological parameters and
survival time of patients with bladder urothelial carcinoma.
| A, Univariate |
|---|
|
|---|
| Covariate | Coefficient | Standard error | HR | 95% CI | P-value |
|---|
| Age range, years:
>55 vs. ≤55 | 0.267 | 0.138 | 1.31 | 0.996–1.712 | 0.053 |
| Tumor size: ≥3 cm
vs. <3 cm | 0.297 | 0.133 | 1.35 | 1.037–1.748 | 0.026a |
| Sex: male vs.
female | 0.129 | 0.196 | 1.14 | 0.775–1.670 | 0.511 |
| Lymph node
metastasis: positive vs. negative | 0.524 | 0.15 | 1.69 | 1.258–2.267 |
<0.0001c |
| Vascular invasion:
positive vs. negative | 1.082 | 0.145 | 2.95 | 2.219–3.922 |
<0.0001c |
| Distant metastasis:
positive vs. negative | 0.533 | 0.205 | 1.7 | 1.140–2.549 | 0.009b |
| Ephrin A3
expression: high vs. low | 0.516 | 0.137 | 1.68 | 1.280–2.192 |
<0.0001c |
| Depth of invasion:
Ta-T1 vs. T2-T4 | 0.954 | 0.206 | 2.6 | 1.736–3.885 |
<0.0001c |
| Histological
grade | 0.563 | 0.069 | 1.76 | 1.533–2.011 |
<0.0001c |
|
| B,
Multivariate |
|
|
Covariate |
Coefficient | Standard
error | HR | 95% CI | P-value |
|
| Tumor size: ≥3 cm
vs. <3 cm | −0.036 | 0.139 | 0.97 | 0.734–1.268 | 0.797 |
| Lymph node
metastasis: positive vs. negative | 0.135 | 0.16 | 1.15 | 0.836–1.568 | 0.399 |
| Vascular invasion:
positive vs. negative | 0.547 | 0.161 | 1.73 | 1.259–2.371 | 0.001b |
| Distant metastasis:
positive vs. negative | 0.386 | 0.218 | 1.47 | 0.960–2.254 | 0.076 |
| Ephrin A3
expression: high vs. low | 0.275 | 0.15 | 1.32 | 0.982–1.765 | 0.066 |
| Depth of invasion:
Ta-T1 vs. T2-T4 | 0.745 | 0.213 | 2.11 | 1.387–3.197 |
<0.0001c |
| Histological
grade | 0.459 | 0.079 | 1.58 | 1.355–1.848 |
<0.0001c |
Genes co-expressed with EFNA3 in BLCA
(GO and KEGG pathway analyses)
Using the LinkedOmics web tool, co-expression of the
EFNA3 gene in 408 BLCA samples from TCGA database was investigated.
Genes positively correlated with EFNA3 (red spots; FDR=0.05) and
genes negatively correlated with EFNA3 (green spots; FDR=0.05) are
shown in the volcano plot in Fig.
4A. The top 50 genes positively and negatively correlated with
EFNA3 are shown in the heat map in Fig.
4B and C. The top three positively associated genes were
ephrinA4 (EFNA4), ADAM metalloprotease domain 15 (ADAM15) and
serine protease 27 (PRSS27) (Fig.
4D-F), and the top three negatively associated genes were
centrosomal protein 120, DENN domain-containing 4A and KIAA1109
(Fig. 4G-I). The results of the GO
enrichment analysis of the LinkedOmics data demonstrated that genes
co-expressed with EFNA3 were located mostly in a mitochondrial
protein complex, respiratory chain, anchored component of membrane,
Sm-like protein family complex or the cytosol (Fig. 5A). The genes were involved in
mitochondrial gene expression, NADH dehydrogenase complex assembly,
the nucleoside monophosphate metabolic process, protein
localization in the endoplasmic reticulum and the ephrin receptor
signaling pathway (Fig. 5B). This
co-expression of genes plays an important role in ribosomal RNA
binding, ephrin receptor binding, unfolded protein binding,
threonine-type peptidase activity, and oxidoreductase activity,
acting on a heme group of donors (Fig.
5C). The KEGG pathway analysis revealed that the genes
co-expressed with EFNA3 were primarily enriched in the ribosome,
Parkinson's disease, the proteasome, carbon metabolism, fructose
and mannose metabolism, the spliceosome and systemic lupus
erythematosus (Fig. 5D). These
findings indicated that EFNA3 had a broad transcriptome impact.
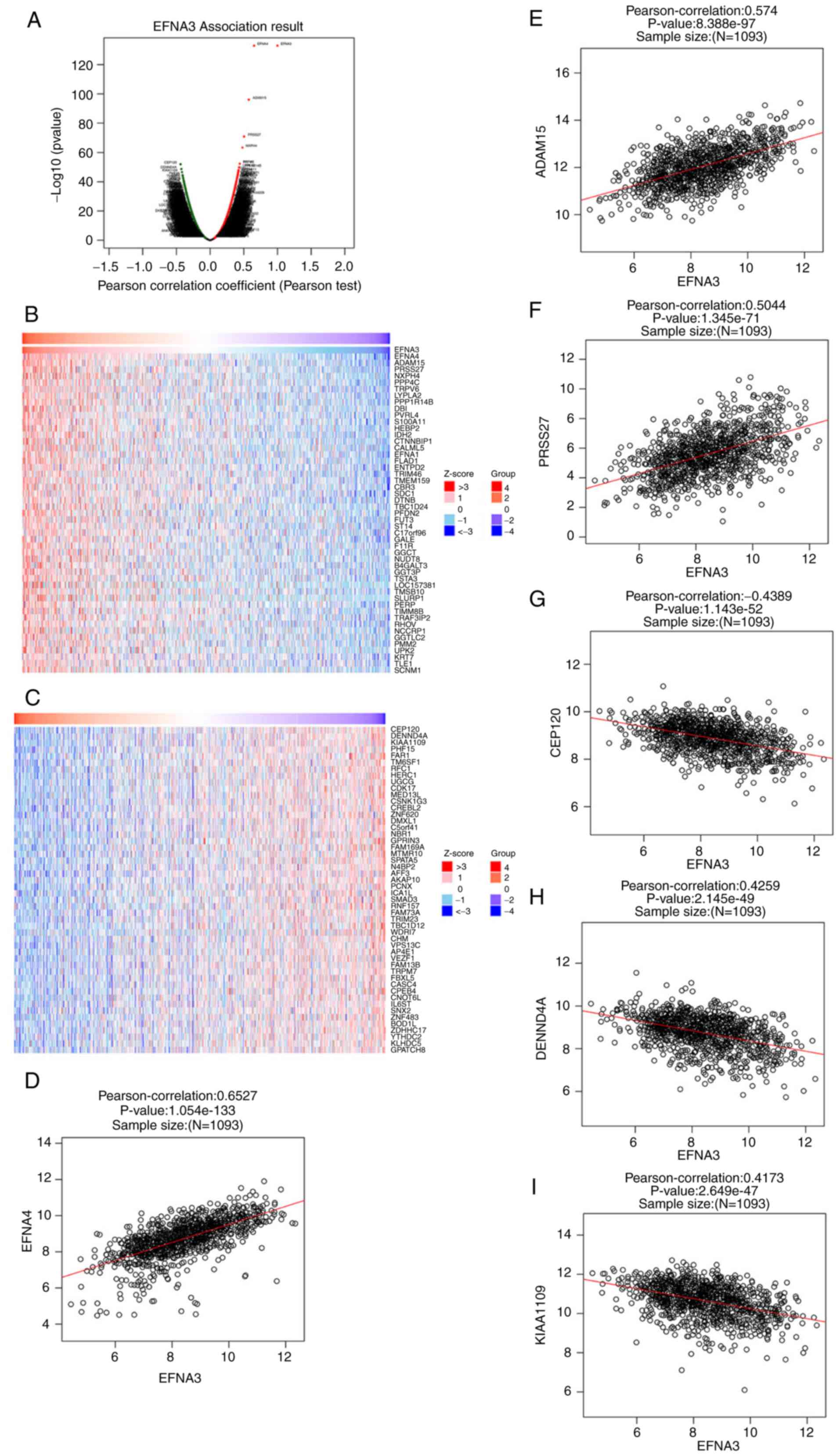 | Figure 4.Genes co-expressed with EFNA3 in BLCA
(using LinkedOmics). (A) Pearson's test was used to analyze
correlations between EFNA3, and genes differentially expressed in
BLCA. (B and C) Heat maps showing the top 50 genes positively and
negatively correlated with EFNA3 in BLCA. Red indicates positively
correlated genes and green indicates negatively correlated genes.
(D-F) The positive correlation between EFNA3 and the top three
genes (EFNA4, ADAM15, and PRSS27). (G-I) The negative correlation
between EFNA3 and the top three genes (CEP120, DENND4A, and
KIAA1109). ADAM15, A distintergase and metalloprotease domain 15;
BLCA, bladder urothelial carcinoma; CEP120, centrosomal protein
120; DENND4A, DENN domain-containing 4A; EFNA3, ephrin A3; EFNA4,
ephrinA4; PRSS27, serine protease 27. |
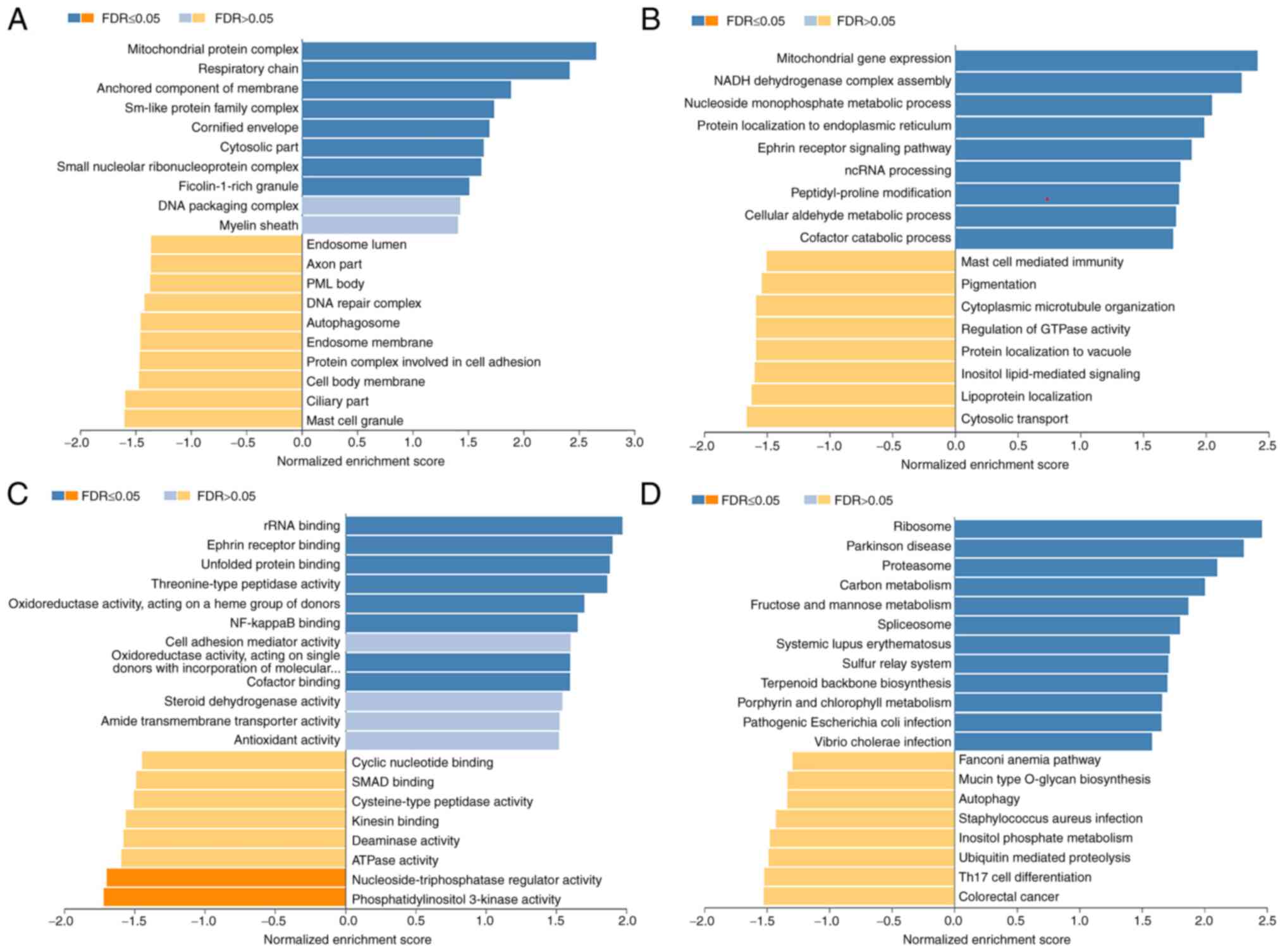 | Figure 5.Significantly enriched GO annotations
and KEGG pathways of EFNA3 in BLCA (using LinkedOmics). The
significantly enriched GO annotations and KEGG pathways of genes
co-expressed with EFNA3 in BLCA were analyzed using gene set
enrichment analysis. The GO annotations were divided into (A)
cellular components, (B) biological processes, and (C) molecular
functions. (D) KEGG pathway analysis. The x-axis represents the
NES, and the y-axis represents the GO term. BLCA, bladder
urothelial carcinoma; EFNA3, ephrin A3; GO, Gene Ontology; KEGG,
Kyoto Encyclopedia of Genes and Genomes; NES, normalized enrichment
score; FDR, false discovery rate; p.adj, adjusted P-value. |
GSEA
To investigate the potential functional processes of
EFNA3 in BLCA, TCGA data were used to perform a GSEA to locate KEGG
pathways that were enriched in samples with high EFNA3 expression.
Significantly enriched pathways were chosen based on their
normalized enrichment score. The results demonstrated that
‘REPRODUCTION’, ‘M_PHASE’, ‘NEURONAL_SYSTEM’,
‘EPIGENETIC_REGULATION_OF_GENE_EXPRESSION’, ‘HCMV_EARLY_EVENTS’ and
‘CHROMOSOME_MAINTENANCE’ were mostly enriched in samples with high
EFNA3 expression (Table IV and
Fig. 6A-F). These data implied that
EFNA3 may aid in the advancement of BLCA by participating in a
number of cancer-related signaling pathways.
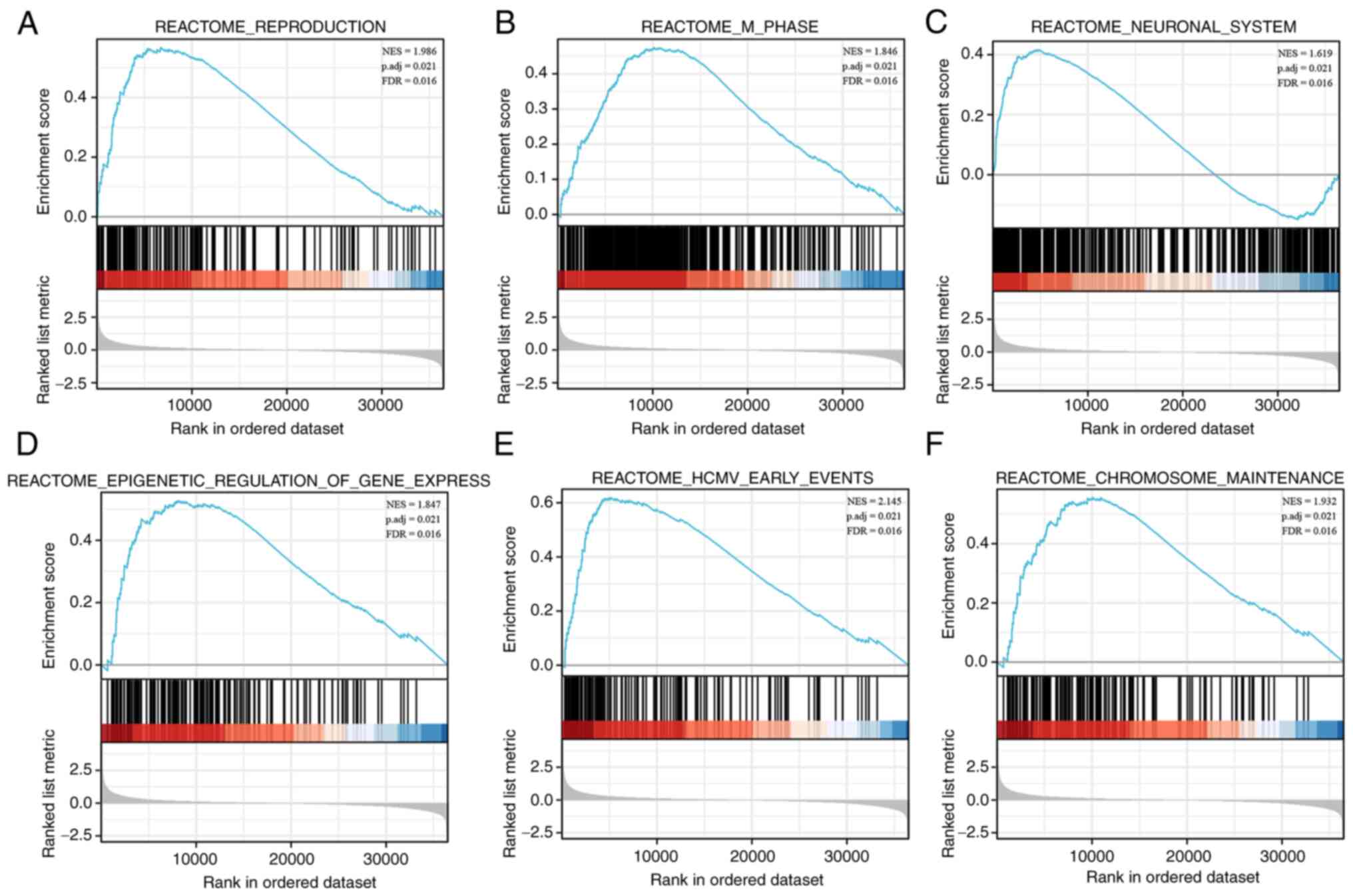 | Figure 6.GSEA reveals potential signaling
pathways of EFNA3. The analysis showed that (A) reproduction, (B)
M-phase, (C) neuronal system, (D) epigenetic regulation of gene
expression, (E) HCMV-early event, and (F) chromosome maintenance
were enriched in the EFNA3 high expression group. The top panels
indicate the enrichment scores for each gene, while the bottom
panels show the ranking metrics of each gene. y-axis, ranking
metric values; x-axis, ranks for all genes. BLCA, bladder
urothelial carcinoma; EFNA3, ephrin A3; GSEA, Gene Set Enrichment
Analysis; HCMV, human cytomegalovirus. |
 | Table IV.Gene sets enriched in the high ephrin
A3 expression phenotype of bladder urothelial carcinoma. |
Table IV.
Gene sets enriched in the high ephrin
A3 expression phenotype of bladder urothelial carcinoma.
| Gene set name | Normalized
enrichment score | Nominal
P-value | False discovery
rate q-value |
|---|
|
REACTOME_REPRODUCTIONa | 1.98614599 | 0.00175131 | 0.01645493 |
|
REACTOME_M_PHASEa | 1.84556691 | 0.00175439 | 0.01645493 |
|
REACTOME_NEURONAL_SYSTEMa | 1.61850743 | 0.00176056 | 0.01645493 |
|
REACTOME_EPIGENETIC_REGULATION_OF_GENE_EXPRESSIONa | 1.84724447 | 0.00176367 | 0.01645493 |
|
REACTOME_HCMV_EARLY_EVENTSa | 2.14477408 | 0.0017762 | 0.01645493 |
|
REACTOME_CHROMOSOME_MAINTENANCEa | 1.93203135 | 0.00177936 | 0.01645493 |
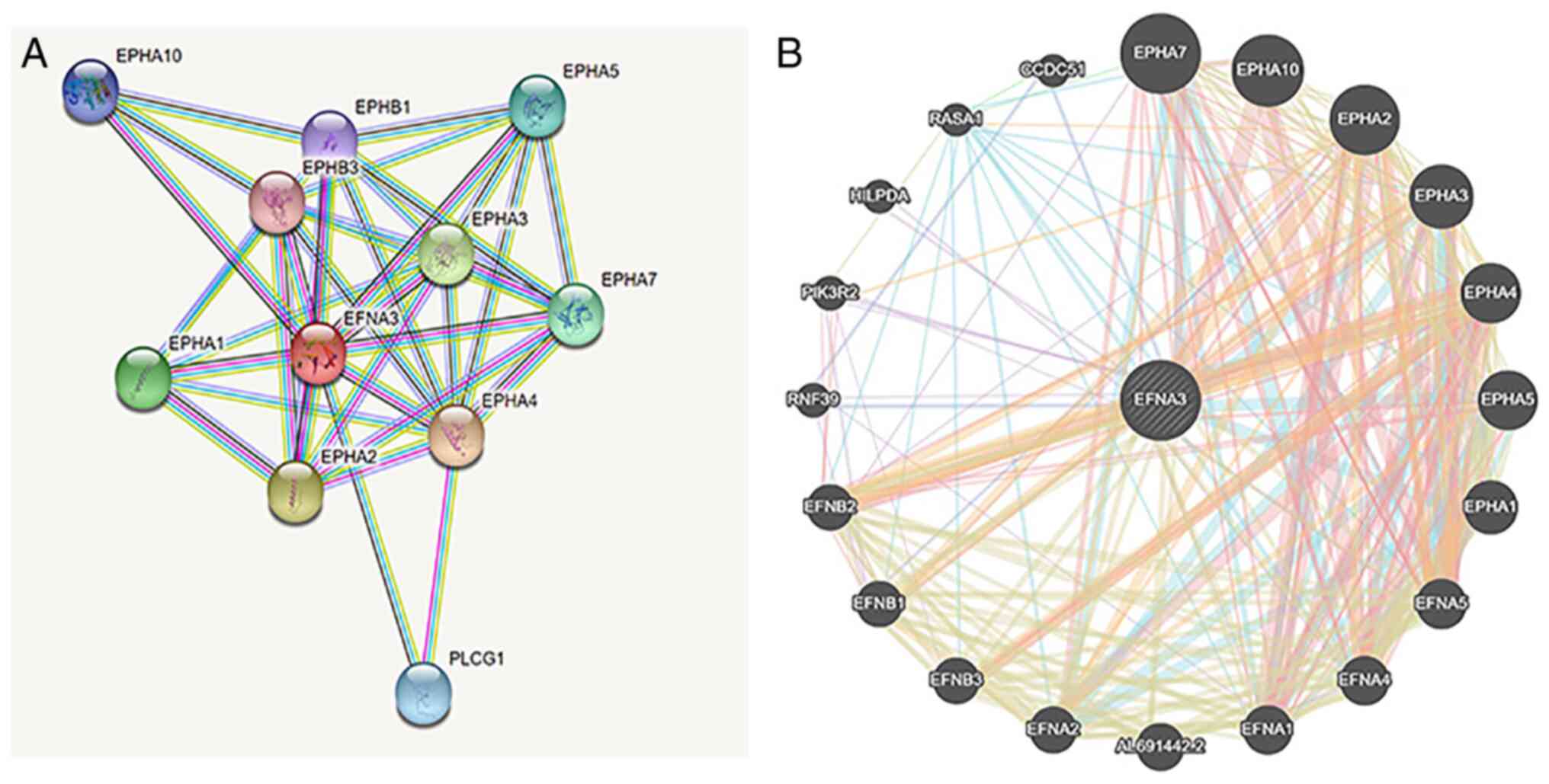
PPI networks
The STRING database was used to generate PPI
networks of EFNA3 in BLCA. The results demonstrated that EFNA3
interacted with Eph receptor A1 (16), EPHA2, EPHA3, EPHA4, EPHA5, EPHA7,
EPHA10, EPHB1, EPHB3, and phospholipase C γ1 (Fig. 7A). In addition, the protein
interactions were analyzed using the GeneMANIA tool and the data
demonstrated that EPHA1, EPHA3, EPHA5, EFNA4, Ras P21 protein
activator 1, EPHA2, EPHA10, EPHA4, EFNA5, EFNB3, EFNB2, EFNB1, and
PIK3R2 interacted with EFNA3 (Fig.
7B).
Discussion
Bladder cancer is the second most common malignancy
of the urinary tract and ranks tenth among the most prevalent types
of cancer worldwide (4). Although
patients are now able to receive effective treatment with advances
in surgical techniques and the advent of neoadjuvant chemotherapy,
bladder cancer remains highly susceptible to recurrence after
treatment and places a great burden on families of patients
(17). Immunotherapy, such as
immune checkpoint blockade, now offers a novel avenue of treatment
for BLCA, improving the survival of patients with advanced bladder
cancer; however, the results remain unsatisfactory (18,19).
Therefore, there is an urgent need for new biomarkers both for
early screening and for the assessment of treatment effectiveness
in bladder cancer.
Ephrins are cell surface ligands that bind to Eph
receptors on adjacent cells. Ephrins act as cell signaling
molecules in pathways that repel and attract each other, mediating
the migration and adhesion of cancer cells (20). The present study focused on EFNA3,
which the present study showed to be significantly differentially
expressed in bladder cancer cells and has a high prognostic value.
A previous study has identified EFNA3 as a key driver of
hepatocellular carcinogenesis and progression in the hypoxic
microenvironment (21). Notably,
several studies have found that EFNA3 is involved in tumor
angiogenesis (22,23). However, to the best of our
knowledge, the impact of EFNA3 in BLCA is unknown.
In the present study, it was found that EFNA3
expression in BLCA tissues was significantly higher than that in
normal bladder tissues, through the analyses of several online
databases. It was also demonstrated that EFNA3 was positively
associated with pathological grade and tumor stage. In addition,
EFNA3 protein expression was detected by IHC, and it was found that
the EFNA3 expression levels were higher in BLCA tissues than in
normal tissues, a result consistent with the mRNA results obtained
from bioinformatics analyses. In summary, high expression of EFNA3
may have a facilitating effect on the proliferation and invasion of
tumor cells.
Next, the clinicopathological features of patients
with BLCA were correlated with the EFNA3 immunohistochemical
scoring in the present study. The results demonstrated that EFNA3
was not only highly expressed in BLCA but was also associated with
tumor size (P<0.05), depth of invasion (P<0.05), distant
metastasis (P=0.001), lymph node metastasis (P=0.003), vascular
invasion (P<0.05), and pathological grade (P=0.01). However,
EFNA3 expression was not associated with patient age or sex. In the
present study, univariate survival analysis was conducted to assess
the prognostic value of EFNA3 in BLCA and it was found that EFNA3
expression levels, tumor size, lymph node metastasis, distant
metastasis, vascular invasion, TNM stage, and pathological grade
may affect the prognosis of BLCA. However, multivariate survival
analysis demonstrated that only vascular invasion, pathological
grade, and TNM stage were independent prognostic factors of BLCA.
This suggested that the mechanism of BLCA development is complex
and that non-independent prognostic factors affecting its prognosis
may be associated with other factors, such as tumor size and
distant metastases. Moreover, survival analysis of 491 samples was
assessed and it was found that patients with BLCA with high EFNA3
expression had a worse prognosis than those with low expression.
This result was consistent with the results of the KM Plotter
database analysis. Therefore, it can be concluded that high EFNA3
expression promotes tumor cell growth and can be used to predict
tumor metastasis and progression in BLCA.
To further understand the role of EFNA3 in BLCA, GO
and KEGG analyses on genes co-expressed with EFNA3 were conducted
in the present study. The results demonstrated that EFNA3 played an
important role in the structure and function of the mitochondria,
ribosomes, DNA, and proteins. The most notable site of enrichment
was the ribosome. Increased synthesis of ribosomes leads to a
corresponding increase in protein synthesis, which ultimately
affects the development of tumor cells and plays an important role
in the development of cancer (24,25).
The enrichment of EFNA3 in the ribosome suggests that its
expression levels are closely related to ribosome biosynthesis in
bladder cancer cells. However, this hypothesis needs to be verified
by relevant experiments in subsequent studies. Nonetheless, the
present study provides new insights into the mechanism of BLCA
development.
In the present study, the results of the GSEA
demonstrated that EFNA3 was involved in biological processes, such
as the cell cycle, transcription factors, and epigenetics. Previous
studies have shown that epigenetic changes are closely associated
with tumor development and prognosis in bladder cancer and that
alterations in DNA hypermethylation and histone acetylation affect
the aberrant expression of a large number of genes (26,27).
Based on these findings, the following possible regulatory network
for EFNA3 is suggested: EFNA3 may influence cell proliferation and
apoptosis by regulating the cell cycle and DNA replication and by
regulating transcription factor activity at the transcriptional
level.
The top three genes positively associated with EFNA3
in the present study were EFNA4, ADAM15, and PRSS27. It has been
demonstrated that EFNA4 was expressed in 82.9% of osteosarcoma
cases and its high expression was associated with poor prognosis
(28). Moreover, knock down of
ADAM15 mRNA expression significantly reduced the invasive ability
of bladder cancer cells through vascular endothelial monolayer,
suggesting that ADAM15 may be involved in the proliferation and
migration of bladder cancer cells (29). In addition, PRSS27 mRNA expression
was higher in resected esophageal squamous cell carcinoma (ESCC)
tissue samples than in normal esophageal mucosal tissues,
suggesting that high PRSS27 expression is an indicator of poor
prognosis in patients with ESCC (30). In summary, EFNA3 co-expression of
related genes may play an important role in tumor development. It
is therefore hypothesized that EFNA3 may promote the development of
BLCA.
Unfortunately, the present study inevitably has some
limitations, the first of which is the small sample size. In this
retrospective analysis, the use of different surgical methods and
resection ranges in patients with different pathological stages has
led to challenges in obtaining sufficient paired samples of
non-cancerous bladder tissue are certainly limitations. For
example, the surgical margin distance for non-muscle invasive
bladder cancer is often <5 cm, which leads to the fact that
noncancerous bladder tissue may not be taken in every bladder
cancer specimen. At the same time, it is worth stating that RNA
sequencing data in public databases may not be representative of
the entire BLCA population. Secondly, TMAs were used instead of
whole tissue sections, which may not reflect the full heterogeneity
of primary BLCA. The third point is that tools such as Image J for
automated semiquantitative analysis of EFNA3 staining were lacking.
In the present study, gene expression levels in tissue sections
were assessed based on the number of positively stained cells and
the intensity of staining, rather than using histochemistry score
(H-score), Allred-score, and immunoreactive scoring systems. This
allows for possible visual bias and lack of accuracy in IHC
scoring. Finally, IHC-based EFNA3 protein expression was assessed,
which remains a semi-quantitative method. Therefore, more
quantitative examinations are required.
In conclusion, the understanding of the mechanisms
of the molecular oncogenic action in BLCA of EFNA3 has been
enriched by merging oncogene expression data from public databases
with IHC results from clinical samples. The present study
demonstrated that EFNA3 plays a significant role in BLCA, and since
upregulation of EFNA3 expression in BLCA was associated with a
shorter survival time and reduced survival, it may bring novel
approaches to BLCA prevention and therapy. Further understanding
EFNA3 expression in BLCA will aid in the identification of patients
with a high metastatic potential. As a result, EFNA3 expression
levels could be used to predict bladder cancer invasion and
prognosis in the future.
Acknowledegments
Not applicable.
Funding
This study was supported by the Zhejiang Provincial Science and
Technology Department Public Welfare Technology Research Program
(grant no. LGF19H160025), Zhejiang Provincial Science and
Technology Department Public Welfare Technology Research Program
(grant no. LBY21H050001), Zhejiang Provincial Health Department
Project (grant no. 2023KY049).
Availability of data and materials
The datasets generated and analyzed during the
current study are available in the TCGA repository, LinkedOmics,
STRING (http://string-db.org/), UALCAN, GEPIA,
and GeneMANIA. The datasets used and/or analyzed during the present
study are available from the corresponding author on reasonable
request.
Authors' contributions
QZ and HW designed the study. HW, YFW, KL and CFZ
collected the data. YFW, YNH and ZFS analyzed and interpreted the
data. YNH and KL confirm the authenticity of all the data. YFW and
HW wrote the manuscript. All authors edited and reviewed the
manuscript.
Ethics approval and consent to
participate
Ethical approval was received for the study from the
Clinical Research Ethics Committee of Zhejiang Provincial People's
Hospital (approval no. QT2022423; Hangzhou, China). Informed
consent was waived by the Clinical Research Ethics Committee of
Zhejiang Provincial People's Hospital. All methods were performed
in accordance with relevant guidelines and regulations.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Amara CS, Vantaku V, Lotan Y and Putluri
N: Recent advances in the metabolomic study of bladder cancer.
Expert Rev Proteomics. 16:315–324. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Siegel RL, Miller KD and Jemal A: Cancer
statistics, 2020. CA Cancer J Clin. 70:7–30. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Lenis AT, Lec PM, Chamie K and Mshs MD:
Bladder cancer: A review. JAMA. 324:1980–1991. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Rouprêt M, Babjuk M, Burger M, Capoun O,
Cohen D, Compérat EM, Cowan NC, Dominguez-Escrig JL, Gontero P,
Hugh Mostafid A, et al: European association of urology guidelines
on upper urinary tract urothelial carcinoma: 2020 update. Eur Urol.
79:62–79. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
He A, He S, Peng D, Zhan Y, Li Y, Chen Z,
Gong Y, Li X and Zhou L: Prognostic value of long non-coding RNA
signatures in bladder cancer. Aging (Albany NY). 11:6237–6251.
2019. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Dal Moro F, Valotto C, Guttilla A and
Zattoni FJU: Urinary markers in the everyday diagnosis of bladder
cancer. Urologia. 80:265–275. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Bellmunt J, Valderrama BP, Puente J,
Grande E, Bolós MV, Lainez N, Vázquez S, Maroto P, Climent MÁ,
Garcia Del Muro X, et al: Recent therapeutic advances in urothelial
carcinoma: A paradigm shift in disease management. Crit Rev Oncol
Hematol. 174:1036832022. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Husain A, Chiu YT, Sze KM, Ho DW, Tsui YM,
Suarez EMS, Zhang VX, Chan LK, Lee E, Lee JM, et al:
Ephrin-A3/EphA2 axis regulates cellular metabolic plasticity to
enhance cancer stemness in hypoxic hepatocellular carcinoma. J
Hepatol. 77:383–396. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Chen J: Regulation of tumor initiation and
metastatic progression by Eph receptor tyrosine kinases. Adv Cancer
Res,. 114:1–20. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Deng M, Tong R, Zhang Z, Wang T, Liang C,
Zhou X and Hou G: EFNA3 as a predictor of clinical prognosis and
immune checkpoint therapy efficacy in patients with lung
adenocarcinoma. Cancer Cell Int. 21:5352021. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Zheng P, Liu X, Li H, Gao L, Yu Y, Wang N
and Chen H: EFNA3 is a prognostic biomarker correlated with immune
cell infiltration and immune checkpoints in gastric cancer. Front
Genet,. 12:7965922022. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Wang L, Song Y, Wang H, Liu K, Shao Z and
Shang Z: MiR-210-3p-EphrinA3-PI3K/AKT axis regulates the
progression of oral cancer. J Cell Mol Med. 24:4011–4022. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Blum A, Wang P and Zenklusen JC: SnapShot:
TCGA-Analyzed Tumors. Cell. 173:5302018. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Chandrashekar DS, Karthikeyan SK, Korla
PK, Patel H, Shovon AR, Athar M, Netto GJ, Qin ZS, Kumar S, Manne
U, et al: UALCAN: An update to the integrated cancer data analysis
platform. Neoplasia. 25:18–27. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
World Medical Association, . World Medical
Association Declaration of Helsinki: Ethical principles for medical
research involving human subjects. JAMA. 310:2191–2194. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Stephenson AJ, Wood DP, Kattan MW, Klein
EA, Scardino PT, Eastham JA and Carver BS: Location, extent and
number of positive surgical margins do not improve accuracy of
predicting prostate cancer recurrence after radical prostatectomy.
J Urol. 182:1357–1363. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Expert Panel on Urological Imaging, .
Allen BC, Oto A, Akin O, Alexander LF, Froemming AT, Fulgham PF,
Halpern DJ, Gettle LM, Maranchie JK, et al: ACR Appropriateness
Criteria® Post-Treatment Surveillance of Bladder Cancer: 2021
Update. J Am Coll Radiol. 18((5S)): S126–S138. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Bhuvaneswari R, Gan YY, Lucky SS, Chin WW,
Ali SM, Soo KC and Olivo M: Molecular profiling of angiogenesis in
hypericin mediated photodynamic therapy. Mol Cancer. 7:562008.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Wu J, Xie RY, Cao CZ, Shang BQ, Shi HZ and
Shou JZ: Disease management of clinical complete responders to
neoadjuvant chemotherapy of muscle-invasive bladder cancer: A
review of literature. Front Oncol. 12:8164442022. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Nievergall E, Lackmann M and Janes PW:
Eph-dependent cell-cell adhesion and segregation in development and
cancer. Cell Mol Life Sci. 69:1813–1842. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Gomez-Maldonado L, Tiana M, Roche O,
Prado-Cabrero A, Jensen L, Fernandez-Barral A, Guijarro-Muñoz I,
Favaro E, Moreno-Bueno G, Sanz L, et al: EFNA3 long noncoding RNAs
induced by hypoxia promote metastatic dissemination. Oncogene.
34:2609–2620. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Wang H, Wang L, Zhou X, Luo X, Liu K,
Jiang E, Chen Y, Shao Z and Shang Z: OSCC exosomes regulate
miR-210-3p Targeting EFNA3 to promote oral cancer angiogenesis
through the PI3K/AKT Pathway. Biomed Res Int.
2020:21256562020.PubMed/NCBI
|
|
23
|
Besnier M, Gasparino S, Vono R, Sangalli
E, Facoetti A, Bollati V, Cantone L, Zaccagnini G, Maimone B,
Fuschi P, et al: miR-210 enhances the therapeutic potential of
bone-marrow-derived circulating proangiogenic cells in the setting
of limb ischemia. Mol Ther. 26:1694–1705. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Catez F, Dalla Venezia N, Marcel V, Zorbas
C, Lafontaine DLJ and Diaz JJ: Ribosome biogenesis: An emerging
druggable pathway for cancer therapeutics. Biochem Pharmacol.
159:74–81. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Pelletier J, Thomas G and Volarević S:
Ribosome biogenesis in cancer: New players and therapeutic avenues.
Nat Rev Cancer. 18:51–63. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Enokida H and Nakagawa M: Epigenetics in
bladder cancer. Int J Clin Oncol. 13:298–307. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Awadalla A, Abol-Enein H, Hamam ET, Ahmed
AE, Khirallah SM, El-Assmy A, Mostafa SA, Babalghith AO, Ali M,
Abdel-Rahim M, et al: Identification of epigenetic interactions
between miRNA and gene expression as potential prognostic markers
in bladder cancer. Genes (Basel). 13:16292022. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Abdou AG, Abd el-Wahed MM, Asaad NY,
Samaka RM and Abdallaha R: Ephrin A4 expression in osteosarcoma,
impact on prognosis, and patient outcome. Indian J Cancer.
47:46–52. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Lorenzatti Hiles G, Bucheit A, Rubin JR,
Hayward A, Cates AL, Day KC, El-Sawy L, Kunju LP, Daignault S, Lee
CT, et al: ADAM15 is functionally associated with the metastatic
progression of human bladder cancer. PLoS One. 11:e01501382016.
View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Kataoka A, Yamada K, Hagiwara T, Terayama
M, Sugimoto T, Nohara K, Igari T, Yokoi C and Kawamura YI:
Expression status of serine protease 27: A prognostic marker for
esophageal squamous cell carcinoma treated with preoperative
chemotherapy/chemoradiotherapy. Ann Surg Oncol. 28:5373–5381. 2021.
View Article : Google Scholar : PubMed/NCBI
|