Introduction
Ovarian cancer (OC) is the most lethal gynecological
cancer (1). The main reason for the
high mortality rate of OC (70%) is it is asymptomatic in the early
stages of the disease, therefore, ~70% of OC cases are diagnosed in
advanced stages (2). The mortality
rate also depends on therapy resistance occurrence (3), distant metastasis in advanced stages
and epithelial-mesenchymal transition (EMT), which is the key
molecular process in metastasis (4). It has been reported that 90% of
primary ovarian tumors are of epithelial origin, described as the
surface epithelium of the ovary or the surface epithelial inclusion
cysts (5).
MicroRNAs (miRNAs/miR) are small [21-24 nucleotides
(nt) long] non-coding RNAs and can regulate the translation via
degradation of target mRNAs (6).
Each miRNA has multiple targets and a specific sequence of 6–8 nt
that binds to complementary sequences in the regulatory regions of
target genes (7). miRNAs may serve
a role as oncogenes (8) or tumor
suppressors (9) depending on the
target genes. Thus, they are marked regulators of complex processes
such as tumor metastasis and differentiation.
Epithelial cells are tightly interconnected by
various connecting structures such as tight junction complex,
cell-cell adhesion complex and desmosomes. E-cadherin (epithelial
cadherin) is responsible for cell-cell adhesion and binds to actin,
which acts as a cytoskeletal element, through membrane proteins
such as α- and β-catenin. The increased function of
E-cadherin/catenin affects the EMT process and metastasis (4).
Factors that cause EMT are considered to initiate
epithelial remodeling by impairing E-cadherin function or altering
its expression (10,11). Vimentin belongs to the intermediate
filament family that forms the cytoskeleton and is expressed in
mesenchymal cells. Altered expression level of vimentin is
associated with malignancy and causes poor prognosis in several
cancer types, such as prostate, lung and breast cancer, melanoma
and OC (12–15). Vimentin is a potential target in
cancer therapy due to its upregulation in patients with tumors and
the association with metastasis via its regulatory role in the EMT
process (13,16).
EMT-mediated cancer progression and metastasis are
regulated through several processes including miRNAs (17). miR-200 family can inhibit EMT and
induce the mesenchymal-to-epithelial transition (MET) process
(17). The EMT-inducing targets of
the miR-200 family are the zinc finger enhancer (E)-box binding
homeobox (ZEB) 1 and ZEB2 genes and elucidating their interaction
mechanism could be a notable step in the EMT-associated treatment
of cancer types such as breast, lung, colorectal, liver and ovarian
cancer (17). A list of up- and
downregulated-miRNAs in tumor tissues were previously identified
with epithelial OC (EOC) compared with simple ovarian cyst tissues
using a microarray, with miR-200c-3p reported to be markedly
upregulated in EOC samples (2). In
the present study, the function of miR-200c-3p was investigated
with one of its target genes, ZEB1, to elucidate the underlying
molecular mechanisms involved in the EMT process in high-grade
serous OC (HGSOC). Furthermore, E-cadherin and vimentin proteins
were evaluated immunohistochemically in terms of their roles in EMT
and MET in the development and progression of ovarian tumors. The
role of this miRNA and target genes in EMT will enhance current
understanding of how cancer metastasis occurs and targeting these
miRNAs will allow the potential development of novel ways to treat
cancer by using them as therapeutic agents in the future.
Patients and methods
Sample collection
A two-sample independent t-test was used to assess
statistical power to detect differences between tumor and healthy
groups. Cohen's D was used to determine effect sizes based on the
means and standard deviations of each group. Python (version
3.14.0; python.org) was used to perform power analysis, considering
a sample size of n=15 per group. Power test values for sample size
were visualized. A power level of 80% was used to assess whether
the available sample size was sufficiently large to detect
statistically significant differences.
The samples consist of healthy (n=15) and cancerous
ovarian tissues (n=15). The present study was approved by the
Istanbul Medical Faculty Clinical Research Ethics Committee
(approval no. 2019/1206; Istanbul, Turkey) and written informed
consent was obtained from each patient involved. All tissue samples
had been collected at the Department of Obstetrics and Gynecology,
Istanbul University Faculty of Medicine (Istanbul, Turkey) from
December 2019 to April 2020. Cancerous ovarian tissues were
resected from patients with HGSOC who did not undergo hormone
therapy or chemotherapy prior to surgery. Healthy ovarian tissues
were obtained from patients who underwent surgery for any reason
other than cancer, such as total abdominal hysterectomy or
uni/bilateral oophorectomy. All tissues were pathologically
diagnosed and graded by Department of Pathology, Istanbul
University Faculty of Medicine. All serous ovarian carcinoma tissue
samples were staged according to the International Federation of
Gynecology and Obstetrics staging system (18). After collection, tissue samples were
transferred to 10% neutral buffered formalin to prevent tissue
degradation for use in immunohistochemical studies and RNAlater
solution (Invitrogen; Thermo Fisher Scientific, Inc.) to protect
and stabilize RNAs for use in molecular genetic studies.
Subsequently, 500 mg was extracted from all tissue samples and
stored at −80°C until total RNA isolation.
Reverse transcription-quantitative PCR
(RT-qPCR)
Total RNA was isolated from tissues using the
PureLink RNA Mini Kit (Invitrogen; Thermo Fisher Scientific, Inc.)
according to the manufacturer's protocol. The quantity and quality
of the obtained total RNA were measured using NanoDrop 2000 (Thermo
Scientific, Inc.). All samples were kept at −80°C for further
steps.
To compare the expression of Homo sapiens
(hsa)-miR-200c-3p in healthy ovarian and tumor tissues, total RNA
samples were reverse transcribed into complementary DNA (cDNA)
using the TaqMan™ MicroRNA Reverse Transcription Kit (Thermo Fisher
Scientific, Inc.). U6 small nuclear RNA was used for normalization.
The primer sequences of hsa-miR-200c-3p and U6 are shown in
Table I.
 | Table I.Primer sequences of ZEB1,
hsa-miR-200c-3p, U6 snRNA and GAPDH. |
Table I.
Primer sequences of ZEB1,
hsa-miR-200c-3p, U6 snRNA and GAPDH.
| Gene/miR | Primer sequence
(5′-3′) |
|---|
| ZEB1 | F:
AACTGCTGGGAGGATGACAC |
|
| R:
TCCTGCTTCATCTGCCTGA |
|
Hsa-miR-200c-3p | F:
GGCGTAATACTGCCGGGTA |
|
| R:
ATTGCGTGTCGTGGAGTCG |
| U6 snRNA | F:
CTCGCTTCGGCAGCACA |
|
| R:
AACGCTTCACGAATTTGCGT |
| GAPDH | F:
AGGGCTGCTTTTAACTCTGGT |
|
| R:
CCCCACTTGATTTTGGAGGGA |
mRNA targets of hsa-miR-200c-3p were identified
using the miRNA target prediction and functional annotation
database (miRDB; mirdb.org/). The expression of ZEB1 gene was also
analyzed using RT-qPCR. For cDNA synthesis, the iScript cDNA
Synthesis Kit (Bio-Rad Laboratories, Inc.) was used according to
the manufacturer's protocol. RT-qPCR was performed on the
Stratagene™ Mx3005p thermal cycler (Agilent Technologies) using the
iTaq Universal SYBR® Green Supermix kit (Bio-Rad
Laboratories, Inc.). GAPDH was used as the reference gene. Primer
sequences of ZEB1 and GAPDH genes are shown in Table I. TaqMan™ MicroRNA Master Mix
(Thermo Fisher Scientific, Inc.) was used for TaqMan-based RT-qPCR
assay.
Immunohistochemistry (IHC)
Tissue samples were fixed in 10% neutral buffered
formalin at room temperature for 24 h, dehydrated through a graded
ethanol series, cleared in xylene, and embedded in paraffin.
Sections of 3 µm thickness were cut from paraffin blocks and
mounted on positively charged slides. Immunohistochemical staining
was performed using the Ventana Benchmark Ultra fully automated
staining system (Roche Tissue Diagnostics) according to the
manufacturer's protocols. Antigen retrieval was performed using
Cell Conditioning 1 (CC1) solution (cat. no. 950-224; Roche Tissue
Diagnostics) at 95°C for 30 min. Permeabilization, blocking of
nonspecific binding, and quenching of endogenous peroxidase
activity were performed automatically by the Benchmark Ultra system
using proprietary reagents included in the detection kit.
Immunodetection was carried out using the UltraView Universal DAB
Detection Kit (cat. no. 760-500; Roche Tissue Diagnostics). Primary
and secondary antibody incubation was performed at 37°C, with
incubation times controlled automatically by the instrument
according to the manufacturer's recommendations. Slides were
counterstained with hematoxylin at room temperature using the
automated protocol and subsequently mounted. Immunohistochemical
staining and evaluation were performed using the Ventana BenchMark
Ultra system and Ventana System Software (version 12.1; Roche
Tissue Diagnostics). The primary antibodies are provided in
Table II.
 | Table II.Antibodies used in
immunohistochemical staining and their properties. |
Table II.
Antibodies used in
immunohistochemical staining and their properties.
| Antibody name | Source | Clone | Cat. no. | Dilution | Positive control
tissue |
|---|
| ZEB1 | Invitrogen; Thermo
Fisher Scientific, Inc. | Rabbit
polyclonal | PA5 82982 | 1:3,000 | Endometrium |
| E-cadherin | Roche Tissue
Diagnostics; Roche Diagnostics, Ltd. | EP700Y/Rabbit
monoclonal | 05973872001 | Ready to use | Breast
carcinoma |
| Vimentin | Roche Tissue
Diagnostics; Roche Diagnostics, Ltd. | V9/Mouse
monoclonal | 05278139001 | Ready to use | Skin |
Immunohistochemically stained sections were observed
and evaluated with a Nikon Eclipse E200 (Nikon Corporation) light
microscope. Microscopic images were obtained from the digital
imaging system Aperio ScanScope XD (Leica Biosystems Nussloch
GmbH). Two different criteria, which are the percentage of staining
area and staining intensity, were evaluated using a
semi-quantitative scoring system for ZEB1, E-cadherin and vimentin
(10,11,13).
The percentage of the staining area was obtained with the formula:
(Total number of positive cells/total number of tumor cells) ×100.
Data were scored as 0 (0%), 1 (1–10%), 2 (10–50%), 3 (<51%).
Staining intensity was determined according to the brown shade
formed in DAB chromogen+ cells. Scoring was evaluated as
0 (negative, -), 1 (weak, +), 2 (medium, ++) and 3 (strong, +++).
The sum of staining intensity and percent staining area scores was
used as the final staining score.
Statistical analysis
The RT-qPCR results of ZEB1 mRNA and hsa-miR-200c-3p
were analyzed according to Cq values. ΔΔCq values are calculated
using Microsoft Excel (Microsoft Corporation, Version 16.103.3) and
2−ΔΔCq demonstrated the fold-changes between the test
and control groups.
GraphPad Prism (version 9.3.0; Dotmatics) was used
for statistical analysis of RT-qPCR results. Data distribution was
evaluated for normality using the Shapiro-Wilk test RT-qPCR data
are expressed as mean ± SD and an independent sample t-test was
used for comparison. Since the IHC staining results were
categorical data, they are expressed as median (minimum-maximum)
and a Mann-Whitney U test was used for comparison. Correlation
analysis was performed using Spearman's rank correlation analysis
for all data. Receiver operating characteristic (ROC) curve
analysis was applied to continuous numerical data, and cut-off
values were determined by selecting the point on the ROC curve
closest to the upper-left corner (0,1). P<0.05 was considered to
indicate a statistically significant difference.
Results
Patient characteristics
In the present study, 15 healthy and cancerous
ovarian tissues [13 were serous (86.6%), 1 was mucinous (6.6%) and
1 was heterologous carcinoma (6.6%)] were included. All serous
adenocarcinoma samples were in stage III–IV, except one stage III
sample. The age range of all patients in both groups was 43–74
years. The median age was 59 years (interquartile range, 52–67
years).
miRNA expression analysis
Concentrations of total RNAs isolated from healthy
and tumor ovarian tissues were measured with NanoDrop. The
concentration values were between 25.5–1,054.2 ng/µl. A260/280
purity values were between 1.93 and 2.13. According to the RT-qPCR
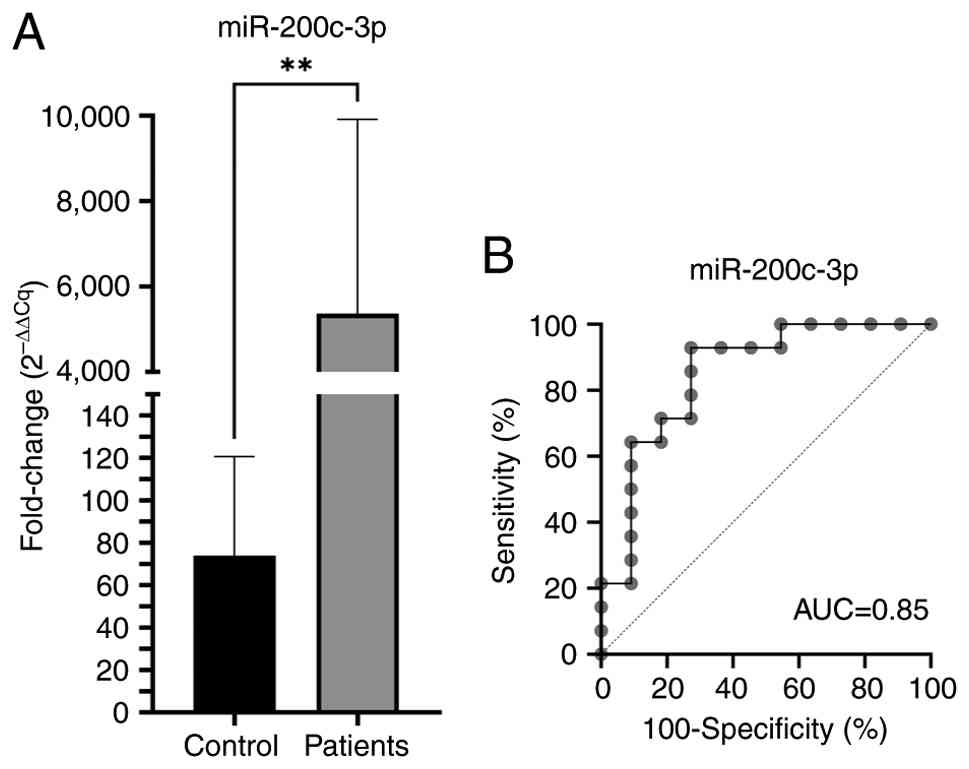
results, hsa-miR-200c-3p expression was significantly increased in
OC (P<0.01) compared with that in healthy tissues (Fig. 1A). ROC curve analysis results
revealed that hsa-miR-200c-3p has 90% sensitivity and 100%
specificity for OC (Fig. 1B).
Gene expression analysis
mRNA targets of hsa-miR-200c-3p were identified
using the miRNA target prediction and functional annotation
database (miRDB; mirdb.org/) and ZEB1 is identified as one of the
target genes (Table SI). RT-qPCR
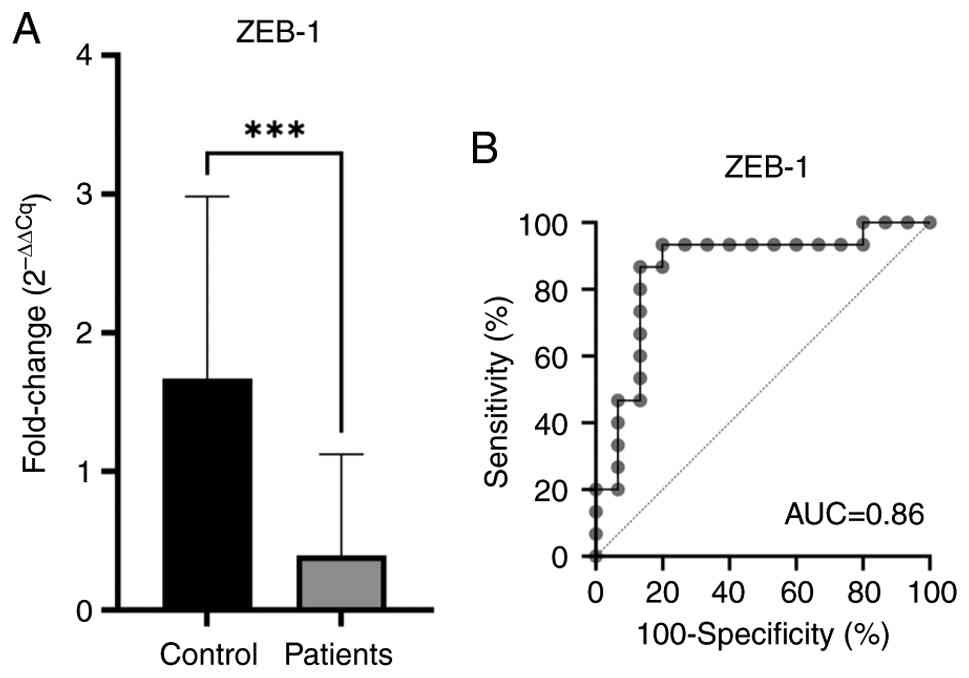
of healthy and tumor cDNA samples demonstrated that the expression
level of the ZEB1 was significantly decreased in OC tissues
compared with healthy tissues (P<0.001). The obtained relative
expression chart is displayed in Fig.
2A.
ROC curve analysis results demonstrated that the
ZEB1 gene had 93% sensitivity and 100% specificity for OC (Fig. 2B). ROC curve analysis of mRNAs of
the ZEB1 gene in predicting OC is shown in Fig. 2B.
IHC analysis
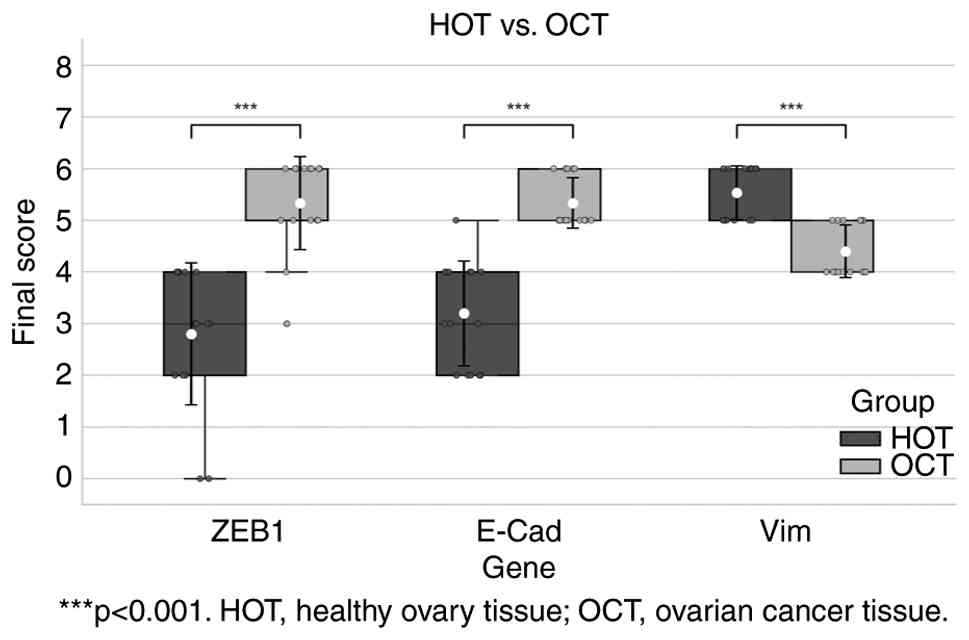
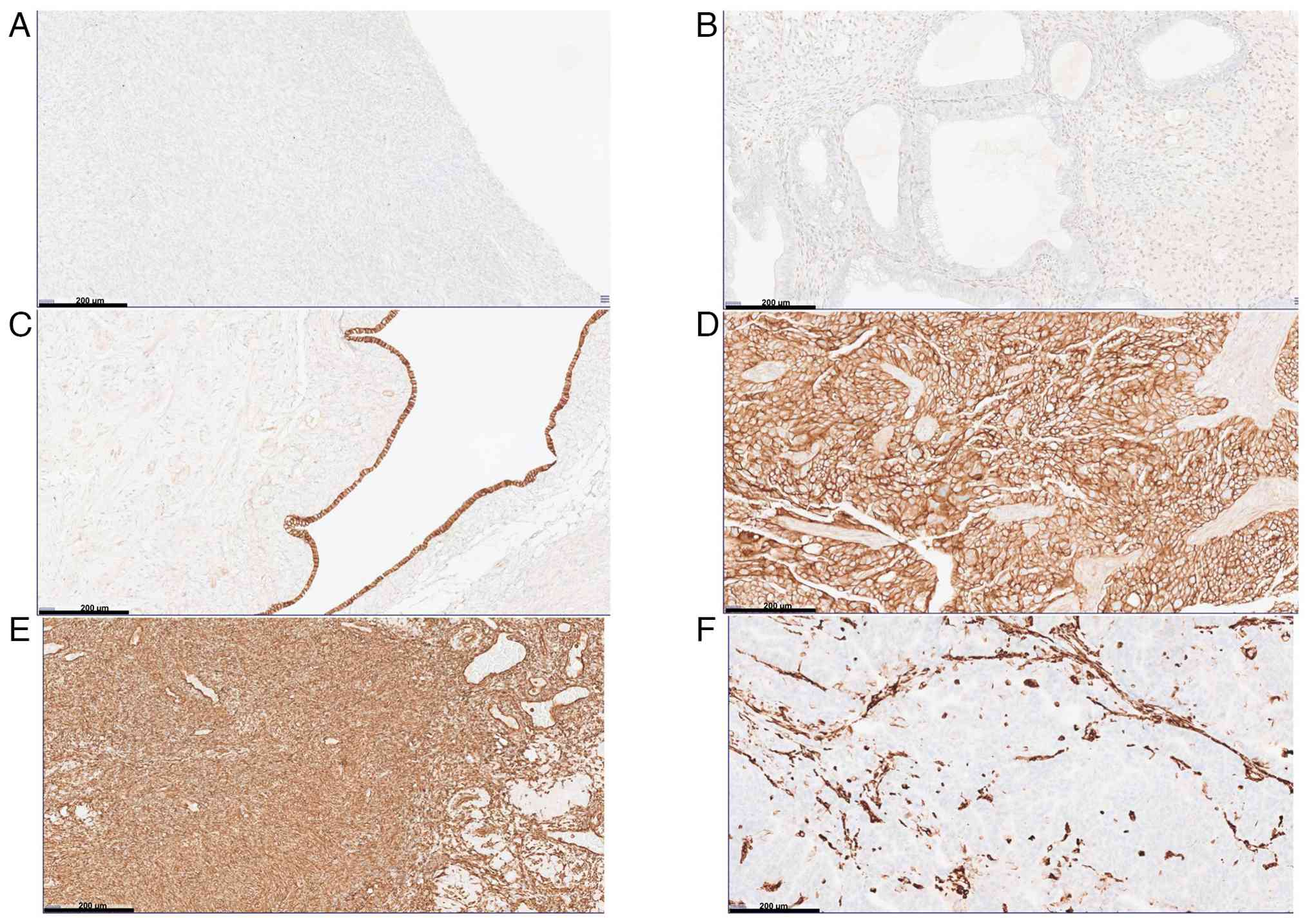
Semi-quantitative staining scores of antibodies are
displayed in Fig. 3 and Table III. Certain IHC staining images of
ZEB1, E-cadherin and vimentin proteins in healthy and cancerous
ovarian tissues are displayed in Fig.
4.
 | Table III.Semi-quantitative scoring of
immunohistochemical staining of ZEB1, E-cadherin and vimentin
antibodies. |
Table III.
Semi-quantitative scoring of
immunohistochemical staining of ZEB1, E-cadherin and vimentin
antibodies.
|
| ZEB1 | E-cadherin | Vimentin |
|---|
|
|
|
|
|
|---|
|
| HOT | OCT | HOT | OCT | HOT | OCT |
|---|
|
|
|
|
|
|
|
|
|---|
| Sample no. | PSA | SI | FS | PSA | SI | FS | PSA | SI | FS | PSA | SI | FS | PSA | SI | FS | PSA | SI | FS |
|---|
| 1 | 2 | 1 | 3 | 2 | 1 | 3 | 1 | 1 | 2 | 3 | 3 | 6 | 3 | 3 | 6 | 2 | 3 | 5 |
| 2 | 2 | 1 | 3 | 3 | 3 | 6 | 3 | 2 | 5 | 3 | 3 | 6 | 3 | 3 | 6 | 2 | 2 | 4 |
| 3 | 0 | 0 | 0 | 3 | 2 | 5 | 3 | 1 | 4 | 3 | 3 | 6 | 3 | 3 | 6 | 2 | 3 | 5 |
| 4 | 1 | 1 | 2 | 3 | 3 | 6 | 2 | 2 | 4 | 3 | 3 | 6 | 3 | 3 | 6 | 2 | 2 | 4 |
| 5 | 0 | 0 | 0 | 3 | 3 | 6 | 1 | 1 | 2 | 2 | 3 | 5 | 3 | 3 | 6 | 2 | 2 | 4 |
| 6 | 2 | 2 | 4 | 3 | 3 | 6 | 1 | 1 | 2 | 2 | 3 | 5 | 3 | 3 | 6 | 2 | 3 | 5 |
| 7 | 1 | 1 | 2 | 3 | 3 | 6 | 1 | 1 | 2 | 3 | 3 | 6 | 2 | 3 | 5 | 3 | 2 | 5 |
| 8 | 1 | 1 | 2 | 3 | 3 | 6 | 2 | 2 | 4 | 2 | 3 | 5 | 3 | 3 | 6 | 2 | 2 | 4 |
| 9 | 2 | 2 | 4 | 2 | 2 | 4 | 1 | 1 | 2 | 2 | 3 | 5 | 2 | 3 | 5 | 2 | 2 | 4 |
| 10 | 2 | 2 | 4 | 3 | 2 | 5 | 2 | 1 | 3 | 2 | 3 | 5 | 2 | 3 | 5 | 3 | 2 | 5 |
| 11 | 2 | 2 | 4 | 3 | 3 | 6 | 2 | 2 | 4 | 2 | 3 | 5 | 2 | 3 | 5 | 2 | 2 | 4 |
| 12 | 2 | 1 | 3 | 3 | 2 | 5 | 2 | 2 | 4 | 2 | 3 | 5 | 3 | 3 | 6 | 2 | 2 | 4 |
| 13 | 2 | 1 | 3 | 3 | 2 | 5 | 1 | 2 | 3 | 2 | 3 | 5 | 2 | 3 | 5 | 2 | 2 | 4 |
| 14 | 2 | 2 | 4 | 3 | 3 | 6 | 2 | 2 | 4 | 2 | 3 | 5 | 2 | 3 | 5 | 2 | 2 | 4 |
| 15 | 2 | 2 | 4 | 2 | 3 | 5 | 1 | 2 | 3 | 2 | 3 | 5 | 2 | 3 | 5 | 3 | 2 | 5 |
IHC statistical analysis
Immunohistochemical staining results in healthy and
cancerous ovarian tissues (Fig. 3)
were compared using a Mann-Whitney U test. Expression of ZEB1
protein in patients with OC was significantly increased compared
with the in healthy individuals (U=11; P<0.001). The results
demonstrated that the levels of E-cadherin protein in patients with
OC were significantly increased compared with healthy individuals
(U=5; P<0.001). In addition, the levels of vimentin protein in
patients with OC were demonstrated to be significantly reduced
compared with healthy individuals (U=21; P<0.001). The
correlation between five different parameters (hsa-miR-200c-3p,
ZEB1 mRNA, ZEB1, E-cadherin and vimentin proteins) in OC was
evaluated and correlation coefficients were calculated (Fig. S1). In the correlation analysis, the
relationship with the correlation coefficient was accepted as a
medium linear relationship ±0.3–0.7 and the relationship with the
correlation coefficient was considered as a strong linear
relationship ±0.7–1. Strong negative correlations were detected
between hsa-miR-200c-3p and ZEB1 mRNA (P=−0.75) and between ZEB1
and vimentin proteins (P=−0.78; both P<0.05). Correlations
between other parameters were moderately correlated. Parameters
exhibiting moderate positive correlations were ZEB1 mRNA with
vimentin protein (Ρ=0.47), hsa-miR-200c-3p with E-cadherin protein
(Ρ=0.58), hsa-miR-200c-3p with ZEB1 protein (Ρ=0.53) and E-cadherin
with ZEB1 protein (Ρ=0.66; all P<0.05). Moderately negatively
correlated parameters were ZEB1 mRNA and E-cadherin protein
(P=−0.66, P=0.00007), ZEB1 mRNA and ZEB1 protein (P=−0.51,
P=0.004), hsa-miR-200c-3p with vimentin (P=−0.53, P=0.01), and
vimentin and E-cadherin proteins (P=−0.56, P=0.001).
Discussion
OC is the most lethal gynecological malignancies
worldwide with an estimated global incidence of 6–7 cases/100,000
women per year (5). The incidence
of OC increases with age particularly after the mid-40s and because
of the lack of specific symptoms, the early diagnosis of OC remains
difficult in patients with OC (19). The absence of a highly sensitive
biomarker for early diagnosis of disease has led to research into
to the identification of gene-based biomarkers to be used in the
clinic. miRNAs are non-coding small RNAs (~22 nt) which regulate
gene expression by suppressing the translation of targeted mRNAs or
triggering the degradation of mRNA (6). They function as both oncogene and
tumor suppressors in the pathogenesis of cancer. miRNAs serve a
role in cell pathways such as cell invasion, carcinogenesis and
metastasis (20). The use of
miRNA-based biomarkers in the early diagnosis and prognosis of OC
provides more specific and sensitive results compared with other
biomarkers such as members of the miR-200 family, miR-21, and
miR-125b (20).
In our previous study (2), hsa-miR-200c-3p was significantly
increased (P=0.0000965; log FC=7.873392) in tumor tissues with EOC
compared with simple ovarian cyst tissues by microarray and was
validated using qPCR (2). The
present study aimed to determine differences in hsa-miR-200c-3p
expression in OC tissues and healthy ovarian tissues and to analyze
the target gene of candidate mRNAs. The present study demonstrated
a marked upregulation of hsa-miR-200c-3p in OC tissues, supporting
its potential role as a diagnostic biomarker consistent with
previous reports (21–23). Therefore, these findings suggest a
potential association with ovarian cancer biology, indicating that
hsa-miR-200c-3p may have value as a candidate biomarker, although
its role in tumor progression and early diagnosis requires further
validation.
Using bioinformatic tool, miRDB, the present study
determined that hsa-miR-200c-3p serves a role in the EMT pathway
and one of its target genes in this pathway was ZEB1. Investigation
of the EMT pathway may help elucidate the metastasis mechanism of
OC. Thus, the expression level of the ZEB1 gene was assessed and
demonstrated to be downregulated in OC tissues.
EMT is a key mechanism facilitating tumor invasion
and metastasis (4,10,11).
Among the transcription factors regulating EMT, ZEB1 has been
identified as a direct target of hsa-miR-200c-3p (24). It regulates EMT in cancer
development and progression by suppressing the adhesion molecule
E-cadherin (24), expressed by the
cadherin-1 (CDH1) gene, which controls the activity of genes
involved in chemical signal transduction, cell maturation and
regulating cellular movement in the cell. ZEB1 binds to E-boxes
within the E-cadherin promoter and stops expression of E-cadherin
(25). These results suggest that
decreased suppression of CDH1 expression, potentially driven by
hsa-miR-200c-3p-mediated regulation of its target ZEB1, may
contribute to the enhancement of EMT. According to the staining
results, the levels of ZEB1 protein were increased in OC tissues
compared with healthy tissues and were demonstrated to be
negatively correlated with hsa-miR-200c-3p expression. The observed
inverse correlation between ZEB1 mRNA and protein levels suggests
the involvement of post-transcriptional regulatory mechanisms
(26).
IHC results demonstrated increased levels of
E-cadherin in patients with OC compared with that in healthy women.
This result is contrary to the previously reported loss of
E-cadherin in the EMT process in OC (27). This may suggest that there is a
transition process from mesenchymal to epithelium in ovarian
carcinogenesis. In addition, the ovarian surface epithelium may
have trigger abnormal epithelial differentiation. In epithelial
tumors, E-cadherin expression decreases as cancer progresses so
that cells can become more mobile. Loss of E-cadherin in cells is a
key marker of EMT (28). It is
hypothesized that this process is different in OC, indicating that
E-cadherin may be a biomarker that can potentially identify OC at
an early stage and be a key target in the treatment of the disease
in the future.
Vimentin is a type III filament protein that is
highly expressed in mesenchymal cells, such as fibroblasts and
smooth muscle cells (29), and is a
mesenchymal biomarker of EMT. The present study showed that
vimentin protein had a weaker staining pattern in tumorous ovarian
tissues compared with healthy subjects. Huo et al (30) reported that vimentin expression is
decreased in drug-resistant ovarian cancer cells compared with
control cells. The authors stated that the downregulation of
vimentin provides stem cell properties to the cells and causes them
to develop drug resistance by cell cycle arrest in the
G2 phase (30). Vimentin
may be a potential therapeutic target used to treat drug-resistant
OC in the future.
The present study demonstrated an increased level of
epithelial proteins and miR-200c, but a decreased level of
mesenchymal proteins. These results support the MET process instead
of EMT in ovarian carcinogenesis (31). The histological structure of the
ovarian surface contains cells having both epithelial and
mesenchymal phenotypes. The ovarian surface cells contain
epithelial markers such as mucin and E-cadherin and also
mesenchymal markers such as N-cadherin and vimentin (32). The hypothesis that OC originates
from the ovarian surface epithelium may be supported by the
transition of surface epithelial cells from mesenchymal to
epithelial structure. Tumor cells can become invasive or metastatic
without losing their epithelial properties or molecular markers and
without triggering the expression levels of genes of mesenchymal
cells. Therefore, it may not always be appropriate to describe the
terms EMT or MET with metastasis (33). Collectively, the present study
results indicated a MET-like phenotype in OC, rather than a
classical EMT pattern. This finding supports the hypothesis that OC
may originate from the ovarian surface epithelium, which exhibits
both epithelial and mesenchymal characteristics. However, the
limited sample size and the fact that 87% of the patients had
advanced-stage OC may have limited the ability to detect
differences in gene expression between disease stages. In the
future, analysis of a large number of patient samples including all
stages of OC may help to elucidate the ovarian carcinogenesis
process.
The present study has some limitations that should
be acknowledged. First, the relatively small sample size may limit
the generalizability of the findings. Moreover, the analysis was
conducted on a single ovarian cancer cohort, and no subgroup
analyses were performed according to disease stage, histological
subtype, or tissue of origin, which may have obscured potential
stage- or origin-specific differences. Finally, the lack of
validation in independent cohorts limits the strength of the
conclusions.
In conclusion, the present study results support the
hypothesis that hsa-miR-200c-3p negatively regulates ZEB1
expression by binding to the promoter region of the ZEB1 gene. Burk
U et al showed that miR-200c and its families bind to at
least two highly conserved sites in their putative promoter to the
ZEB1. Our hypothesis is based on this finding (34). The ZEB1 protein, whose levels are
then decreased in the cell, cannot sufficiently bind to the
promoter of the CDH1 gene expressing E-cadherin. Therefore, the
levels of E-cadherin protein increase. The findings suggest that
hsa-miR-200c-3p promotes epithelial characteristics in OC by
repressing ZEB1, leading to increased E-cadherin expression. This
regulatory axis may represent a potential diagnostic biomarker and
therapeutic target for OC in the future.
Supplementary Material
Supporting Data
Supporting Data
Acknowledgements
The abstract was presented at the 2nd International
Multidisciplinary Cancer Research Congress Jul 21-Jul 24 2022 in
Giresun, Turkey and published as abstract no. 19 in MOKAD
(Molecular Cancer Research Association of Turkey) abstract book.
The authors would like to thank Mr Mehmet Ulas Bilir (Faculty of
Science, Department of Molecular Biology and Genetics, Istanbul
University, Istanbul, Turkey) for technical support in statistical
analysis step.
Funding
The present study was supported by the Istanbul University
Scientific Research Projects Department (grant no. 35391).
Availability of data and materials
The RT-qPCR data generated in the present study may
be found in Mendeley database at the following URL: https://data.mendeley.com/datasets/9zbnzv8dh8/1; doi:
10.17632/9zbnzv8dh8.1. The remaining data generated in the present
study may be requested from the corresponding author.
Authors' contributions
MS and TG designed and supervised the present study.
EA, MS and ST contributed to sample and data collection, devised
the methodology, validation of data and statistical analysis. EA,
MS and TG contributed to the formal analysis, writing and editing
of the manuscript. ST performed sample handling and medical
assistance. TG and ST confirm the authenticity of all the raw data.
All authors read and approved the final version of the
manuscript.
Ethics approval and consent to
participate
The present study was performed in line with the
principles of the Declaration of Helsinki. The present study was
approved by the Istanbul Medical Faculty Clinical Research Ethics
Committee (approval no. 2019/1206; Istanbul, Turkey). All patients
and volunteers provided written informed consent.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
Authors' information
ORCID: MS, 0000-0002-1941-7135; EA,
0000-0003-3807-0330; ST, 0000-0002-9069-0185; TG,
0000-0003-3514-5210.
References
|
1
|
Jayson GC, Kohn EC, Kitchener HC and
Ledermann JA: Ovarian cancer. Lancet. 384:1376–1388. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Gumusoglu-Acar E, Gunel T, Hosseini MK,
Dogan B, Tekarslan EE, Gurdamar B, Cevik N, Sezerman U, Topuz S and
Aydinli K: Metabolic pathways of potential miRNA biomarkers derived
from liquid biopsy in epithelial ovarian cancer. Oncol Lett.
25:1422023. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Fantone S, Marzioni D and Tossetta G:
NRF2/KEAP1 signaling inhibitors in gynecologic cancers. Expert Rev
Anticancer Ther. 24:1191–1194. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Thiery JP, Acloque H, Huang RY and Nieto
MA: Epithelial-mesenchymal transitions in development and disease.
Cell. 139:871–890. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Cho KR and Shih Ie M: Ovarian cancer. Annu
Rev Pathol. 4:287–313. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Lee RC and Ambros V: An extensive class of
small RNAs in Caenorhabditis elegans. Science. 294:862–864. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Bartel DP: MicroRNAs: Target recognition
and regulatory functions. Cell. 136:215–233. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Mattiske S, Suetani RJ, Neilsen PM and
Callen DF: The oncogenic role of miR-155 in breast cancer. Cancer
Epidemiol Biomarkers Prev. 21:1236–1243. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Bonci D, Coppola V, Musumeci M, Addario A,
Giuffrida R, Memeo L, D'Urso L, Pagliuca A, Biffoni M, Labbaye C,
et al: The miR-15a-miR-16-1 cluster controls prostate cancer by
targeting multiple oncogenic activities. Nat Med. 14:1271–1277.
2008. View
Article : Google Scholar : PubMed/NCBI
|
|
10
|
Huber O, Bierkamp C and Kemler R:
Cadherins and catenins in development. Curr Opin Cell Biol.
8:685–691. 1996. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Jeanes A, Gottardi CJ and Yap AS:
Cadherins and cancer: How does cadherin dysfunction promote tumor
progression? Oncogene. 27:6920–6929. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Fletcher DA and Mullins RD: Cell mechanics
and the cytoskeleton. Nature. 463:485–492. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Satelli A and Li S: Vimentin in cancer and
its potential as a molecular target for cancer therapy. Cell Mol
Life Sci. 68:3033–3046. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Leung D, Price ZK, Lokman NA, Wang W,
Goonetilleke L, Kadife E, Oehler MK, Ricciardelli C, Kannourakis G
and Ahmed N: Platinum-resistance in epithelial ovarian cancer: An
interplay of epithelial-mesenchymal transition interlinked with
reprogrammed metabolism. J Transl Med. 20:5562022. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Goldie KN, Wedig T, Mitra AK, Aebi U,
Herrmann H and Hoenger A: Dissecting the 3-D structure of vimentin
intermediate filaments by cryo-electron tomography. J Struct Biol.
158:378–385. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Mendez MG, Kojima S and Goldman RD:
Vimentin induces changes in cell shape, motility, and adhesion
during the epithelial to mesenchymal transition. FASEB J.
24:1838–1851. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Park SM, Gaur AB, Lengyel E and Peter ME:
The miR-200 family determines the epithelial phenotype of cancer
cells by targeting the E-cadherin repressors ZEB1 and ZEB2. Genes
Dev. 22:894–907. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Berek JS, Kehoe ST, Kumar L and
Friedlander M: Cancer of the ovary, fallopian tube, and peritoneum:
2021 FIGO staging system. Int J Gynecol Obstet. 155 (Suppl
1):S61–S85. 2021. View Article : Google Scholar
|
|
19
|
Paul A and Paul S: The breast cancer
susceptibility genes (BRCA) in breast and ovarian cancers. Front
Biosci (Landmark Ed). 19:605–618. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Pal MK, Jaiswar SP, Dwivedi VN, Tripathi
AK, Dwivedi A and Sankhwar P: MicroRNA: A new and promising
potential biomarker for diagnosis and prognosis of ovarian cancer.
Cancer Biol Med. 12:328–341. 2015.PubMed/NCBI
|
|
21
|
Zuberi M, Mir R, Das J, Ahmad I, Javid J,
Yadav P, Masroor M, Ahmad S, Ray PC and Saxena A: Expression of
serum miR-200a, miR-200b, and miR-200c as candidate biomarkers in
epithelial ovarian cancer and their association with
clinicopathological features. Clin Transl Oncol. 17:779–787. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Wang W, Wu LR, Li C, Zhou X, Liu P, Jia X,
Chen Y and Zhu W: Five serum microRNAs for detection and predicting
of ovarian cancer. Eur J Obstet Gynecol Reprod Biol X.
3:1000172019.PubMed/NCBI
|
|
23
|
Meng X, Muller V, Milde-Langosch K,
Trillsch F, Pantel K and Schwarzenbach H: Diagnostic and prognostic
relevance of circulating exosomal miR-373, miR-200a, miR-200b and
miR-200c in patients with epithelial ovarian cancer. Oncotarget.
7:16923–16935. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Panda H, Pelakh L, Chuang TD, Luo X,
Bukulmez O and Chegini N: Endometrial miR-200c is altered during
transformation into cancerous states and targets the expression of
ZEBs, VEGFA, FLT1, IKKbeta, KLF9, and FBLN5. Reprod Sci.
19:786–796. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Humphries B and Yang C: The microRNA-200
family: Small molecules with novel roles in cancer development,
progression and therapy. Oncotarget. 6:6472–6498. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Baldi P and Long AD: A Bayesian framework
for the analysis of microarray expression data: Regularized t-test
and statistical inferences of gene changes. Bioinformatics.
17:509–519. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Sawada K, Mitra AK, Radjabi AR, Bhaskar V,
Kistner EO, Tretiakova M, Jagadeeswaran S, Montag A, Becker A,
Kenny HA, et al: Loss of E-cadherin promotes ovarian cancer
metastasis via alpha 5-integrin, which is a therapeutic target.
Cancer Res. 68:2329–2339. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Savagner P: Leaving the neighborhood:
Molecular mechanisms involved during epithelial-mesenchymal
transition. Bioessays. 23:912–923. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Steinert PM and Roop DR: Molecular and
cellular biology of intermediate filaments. Annu Rev Biochem.
57:593–625. 1988. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Huo Y, Zheng Z, Chen Y, Wang Q, Zhang Z
and Deng H: Downregulation of vimentin expression increased drug
resistance in ovarian cancer cells. Oncotarget. 7:45876–45888.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Yao D, Dai C and Peng S: Mechanism of the
mesenchymal-epithelial transition and its relationship with
metastatic tumor formation. Mol Cancer Res. 9:1608–1620. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Shah AS, Scheele WH, Glant MD and Fugere
P: Raloxifene HCl is not stimulatory in the endometrium as assessed
by the blaustein criteria and an estrogenicity scoring system. Prim
Care Update Ob Gyns. 5:1671998. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Christiansen JJ and Rajasekaran AK:
Reassessing epithelial to mesenchymal transition as a prerequisite
for carcinoma invasion and metastasis. Cancer Res. 66:8319–8326.
2006. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Burk U, Schubert J, Wellner U, Schmalhofer
O, Vincan E, Spaderna S and Brabletz T: A reciprocal repression
between ZEB1 and members of the miR-200 family promotes EMT and
invasion in cancer cells. EMBO Rep. 9:582–589. 2008. View Article : Google Scholar : PubMed/NCBI
|