Introduction
Pituitary adenomas represent one of the most
frequently occurring intracranial tumors globally, constituting 10
to 15% of all primary brain neoplasms (1). These tumors originate from the
adenohypophysis and can lead to severe clinical manifestations,
including systemic complications of the cardiovascular and
metabolic systems, as well as permanent visual deficits due to the
compression of surrounding structures, such as the optic chiasm, by
the tumor (mass effect). Although most pituitary adenomas are
histologically non-cancerous and show a slow-growing clinical
course, a certain group exhibits invasive characteristics, marked
by the penetration into adjacent tissues such as the cavernous
sinus, dura mater or sphenoid sinus (2). These invasive pituitary adenomas
(IPAs) are often associated with treatment resistance, increased
recurrence rates and worse clinical outcomes, despite lacking
malignant histopathological features (3–5).
Growing evidence suggests that metabolic reprogramming and
mitochondrial dysfunction serve pivotal roles in the acquisition of
aggressive phenotypes in various tumors. In the context of
pituitary adenomas, alterations in energy metabolism and
mitochondrial respiratory chains have been implicated in promoting
tumor cell survival and invasion. However, a systematic
understanding of the molecular landscape specifically associated
with these metabolic shifts remains limited.
Recent advances in omics technologies, including
proteomics and metabolomics, have provided powerful tools to
explore tumor biology at the systems level. Proteomic and
metabolomic approaches offer complementary insights into protein
expression dynamics and metabolic alterations, critical for
elucidating tumor biology. While proteomics provides a snapshot of
the functional effectors of the cell, metabolomics offers a
downstream functional readout of the cellular physiological state.
Integrated analysis of these datasets can reveal the molecular
mechanisms underlying tumor progression and metabolic reprogramming
(6,7). Such high-throughput screening is
essential for identifying key biomarkers that might be overlooked
by traditional single-molecule studies.
In the present study, quantitative profiling of
proteins and metabolites was performed in human pituitary adenoma
tissues, comparing invasive and non-invasive subtypes.
Specifically, the study focused on non-functioning pituitary
adenomas to minimize the confounding effects of excessive hormone
secretion on the metabolic profile. By identifying markedly
dysregulated molecular features and assessing their enrichment in
relevant biological pathways, particularly those involving
mitochondrial function and energy pathways, the present study aimed
to delineate the molecular and metabolic determinants of tumor
invasion. The findings indicate that coordinated molecular
perturbations contribute to the aggressive behavior of invasive
pituitary adenomas, providing novel insights into potential
therapeutic targets and supporting future mechanistic
investigations into the metabolic drivers of tumor progression.
Materials and methods
Patients
This study was designed as a prospective recruitment
of patients undergoing surgical resection for pituitary adenomas. A
comparative analysis of 2 patient cohorts was performed to
elucidate the proteomic and metabolic determinants associated with
varying degrees of pituitary tumor aggressiveness. Tumors assigned
a Knosp grade (8) of ≥2 were
classified as invasive, whereas those <2 were categorized as
non-invasive. The inclusion criteria for the pituitary adenoma
samples were as follows: i) Patients had not received any prior
medical, radiological, or surgical treatment before the study; ii)
a preoperative clinical diagnosis of pituitary adenoma; iii)
availability of tumor tissue samples resected during surgery; and
iv) postoperative pathological confirmation of the pituitary
adenoma diagnosis. The exclusion criteria included: i) Patients who
had undergone any prior medical, radiological or surgical
treatment; and ii) cases where the postoperative pathological
diagnosis excluded pituitary adenoma. Additionally, patients who
failed to complete the planned surgery or those from whom tumor
tissue samples could not be retained were excluded from the final
analysis. Patient tumor samples were collected between September
2023 and May 2024 at The First Affiliated Hospital of Xiamen
University (Xiamen, China). Statistical analyses were performed
using appropriate parametric or non-parametric tests depending on
data type and distribution. Continuous variables were compared
using unpaired Student's t-test for two-group comparisons or
one-way ANOVA followed by Tukey's post hoc test for multiple-group
comparisons. Categorical variables, including sex, were analyzed
using Fisher's exact test. A P-value <0.05 was considered to
indicate a statistically significant difference.
Proteomic analysis and protein
quantification
Freshly resected pituitary adenoma tissues were
taken from liquid nitrogen storage and quickly transferred onto
ice. Samples were reconstituted in a lysis buffer containing 8 M
urea and 1% SDS, supplemented with a 1% protease inhibitor cocktail
to prevent enzymatic degradation. Homogenization was performed
using a high-throughput tissue grinder for three consecutive cycles
of 180 sec. This procedure was followed by cryogenic, non-contact
sonication for 30 min. The resulting lysates were centrifuged at
16,000 × g for 30 min at 8°C. Protein concentrations in the
supernatants were quantified using the bicinchoninic acid (BCA)
assay, according to the manufacturer's protocol (BCA Protein Assay
Kit; Thermo Fisher Scientific, Inc.). Subsequently, SDS-PAGE was
employed to separate the extracted proteins using a 12% separation
gel, with 20 µg of protein loaded per lane.
Protein digestion
A total of 100 µg protein was dissolved in
triethylammonium bicarbonate (TEAB) buffer to a final concentration
of 100 mM. Reduction was performed using 10 mM
tris(2-carboxyethyl)phosphine at 37°C for 1 h to disrupt disulfide
bonds. Alkylation was then performed with 40 mM iodoacetamide at
room temperature for min in the dark to preserve the reactivity of
the alkylating agent. Following these treatments, the samples were
centrifuged at 10,000 × g for 20 min at 4°C to eliminate soluble
contaminants, yielding a protein pellet. The pellet was resuspended
in 100 µl TEAB buffer, maintaining the buffer concentration at 100
mM. Proteins were digested by adding trypsin at an
enzyme-to-substrate ratio of 1:50 and incubating overnight at 37°C
to produce peptides for subsequent analysis.
Peptide desalting and
quantification
Following trypsin-mediated proteolysis, peptides
were dried to completeness using a vacuum concentrator. The
resulting desiccated peptides were reconstituted in 0.1%
trifluoroacetic acid and purified via desalting with
hydrophilic-lipophilic balanced cartridges. After purification, the
samples were once again vacuum-dried. Peptide concentrations were
quantified based on ultraviolet absorbance using a NanoDrop One
spectrophotometer (Thermo Fisher Scientific, Inc.), following the
manufacturer's standard operating protocol.
Data-independent acquisition (DIA)
mass detection
Quantified peptide concentrations were subjected to
analytical processing using the Vanquish Neo liquid chromatography
(LC) system and the Compass HyStar interface developed by Bruker
Corporation. All analyses were performed by Shanghai Majorbio
Bio-Pharm Technology Co., Ltd. Mass spectrometry was conducted in
positive ionization mode. Chromatographic separation was performed
on a custom-packed analytical column (15 cm × 100 µm) filled with
1.7 µm particles designed to enhance peptide resolution. For the
electrospray source, the nitrogen gas temperature was set at 180°C,
the nebulizer pressure was maintained at 2.0 bar and the dry gas
flow rate was 8.0 l/min. The separation process employed a binary
solvent system consisting of solvent A (2% acetonitrile and 0.1%
formic acid in water) and solvent B (80% acetonitrile and 0.1%
formic acid in water), maintained over an 8-min linear gradient.
Subsequent peptide detection was performed using a timsTOF HT mass
spectrometer operated in DIA mode. Mass spectra were acquired
within the m/z ranges of 70-1,050 for MS1 and 150-2,000 for MS2,
enabling in-depth characterization of the peptide profiles. Mass
spectrometry (MS) acquisition was performed in dia-PASEF mode. Each
acquisition cycle consisted of multiple isolation windows, with
each window defined by a specific m/z range and its corresponding
ion mobility interval. The DIA isolation windows included the Mass
Begin and Mass End boundaries for each m/z segment, the associated
1/K0 mobility ranges (Begin/End) and the collision
energy (CE), which was dynamically ramped according to the ion
mobility interval. The total cycle time for one full dia-PASEF
acquisition cycle was 0.73 sec, and all remaining timsTOF
instrument parameters followed manufacturer-recommended settings
for quantitative proteomics.
Protein identification
The raw files from DIA were evaluated utilizing
Spectronaut software (version 19; Biognosys AG). In the course of
conducting the database search, parameters were set to restrict
peptide lengths within the range of 7 to 52 amino acids. For
enzymatic cleavage, trypsin/P was designated, permitting up to two
missed cleavage sites. Fixed modifications included
carbamidomethylation of cysteine residues, while methionine
oxidation and N-terminal acetylation of proteins were treated as
variable modifications. Rigorous identification criteria were
implemented, ensuring that both protein and peptide-level false
discovery rates (FDR) remained <1%, with peptide confidence
established at 99%. The extracted ion chromatogram width was
limited to a maximum of 75 ppm. Protein quantification was
conducted using the MaxLFQ algorithm (9).
Proteomics statistical analyses
The proteomic data analysis was performed using the
Majorbio Cloud Platform (version 3.0; http://cloud.majorbio.com), accessible at
https://cloud.majorbio.com. To evaluate the differences in protein
expression between the two experimental groups, the R software
(version 4.3.1; R Foundation for Statistical Computing) and the
‘statmod’ package (version 1.5.0) (10). Protein fold changes (FC) and
corresponding P-values were computed to identify differentially
expressed proteins (DEPs). DEPs were defined by stringent criteria:
FC >1.2 or <0.83 combined with a P-value <0.05, ensuring
the reliable selection of proteins with statistically significant
expression differences. To evaluate the consistency of biological
replicates and the degree of proteomic similarity between samples,
Pearson's correlation analysis was performed. The entire proteomic
dataset was further subjected to functional annotation using
authoritative databases, including Gene Ontology (GO; http://geneontology.org/) and the Kyoto Encyclopedia
of Genes and Genomes (KEGG; http://www.genome.jp/kegg/). This annotation
facilitated the interpretation of biological roles and pathways
associated with the DEPs. Enrichment analyses of GO terms and KEGG
pathways were subsequently performed to reveal the biological
relevance of these proteins. Additionally, protein-protein
interaction (PPI) networks were constructed employing the STRING
database (version 11.5; https://string-db.org/; STRING Consortium)
(11) to visualize the interactions
among DEPs, thereby enhancing comprehension of their functional
interplay within the examined biological system.
Metabolite extraction
A total of ~100 mg of solid tissue was transferred
into a 2 ml microcentrifuge tube, followed by the addition of a 6
mm stainless steel grinding bead to facilitate homogenization.
Metabolite extraction commenced upon adding 800 µl methanol-water
solution (4:1, v/v) to each sample, supplemented with four internal
standards, including L-2-chlorophenylalanine at a concentration of
0.02 mg/ml. The homogenization procedure was carried out utilizing
a frozen tissue grinder (model Wonbio-96c; Shanghai Wonbio
Biotechnology Co., Ltd.), with the temperature maintained at −10°C
and the operating frequency set at 50 Hz for a duration of 6 min.
Following the initial homogenization, the resulting homogenates
underwent ultrasonic-assisted extraction, which was performed at a
controlled temperature of 5°C and a frequency of 40 kHz for a
period of 30 min. Once the extraction process was completed, the
samples were left to incubate at −20°C for 30 min. This was
succeeded by centrifugation at a high speed of 13,000 × g for 15
min at 4°C, ensuring the proper separation of the components within
the samples for subsequent analysis. The supernatants obtained were
gathered and placed into autosampler vials for further analysis
using LC-MS/MS.
Quality control (QC) sample
To guarantee stability within the system and uphold
the quality of analytical control, a pooled QC sample was created
by amalgamating equal portions from each individual sample. The
preparation and analysis processes for these QC samples were
consistent with those applied to the experimental samples. Acting
as a representative for the complete dataset, the pooled QC was
injected at regular intervals-namely, following every six
analytical runs-to evaluate and oversee instrument performance and
signal stability during the LC-MS/MS analysis.
Ultra performance LC-tandem MS
(UPLC-MS/MS) analysis
The samples were analyzed utilizing LC-MS/MS on a
cutting-edge UHPLC-Orbitrap Exploris 240 system (Thermo Fisher
Scientific, Inc.). This advanced system was equipped with an
ACQUITY HSS T3 column (100 mm in length and has an internal
diameter of 2.1 mm with a particle size of 1.8 µm; Waters
Corporation), which measures 100 mm in length and has an internal
diameter of 2.1 mm with a particle size of 1.8 µm, and was provided
by Waters Corporation. The experiments were performed at Shanghai
Majorbio Bio-Pharm Technology Co. Ltd., ensuring access to
high-level analytical resources. To facilitate the chromatographic
process, a mobile phase system was utilized, consisting of two
primary solvents. Solvent A was prepared by dissolving 0.1% formic
acid in a water-acetonitrile mixture at a volumetric ratio 2:98,
whereas Solvent B consisted of pure acetonitrile containing 0.1%
formic acid. Chromatographic separation was conducted under tightly
controlled conditions, maintaining a flow rate of 0.40 ml/min and a
column temperature of 40°C. For each analysis, 5 µl sample was
injected to ensure precision and reproducibility.
The MS analysis employed a UHPLC system coupled to a
UHPLC-Orbitrap Exploris 240 mass spectrometer equipped with an
electrospray ionization source, capable of operating in both
positive and negative ion modes. Instrument parameters were
carefully optimized, with the source temperature set at 400°C,
sheath gas flow at 40 arbitrary units and auxiliary gas flow at 10
arbitrary units. The ion spray voltage floating was adjusted to
−2,800 V in negative mode and 3,500 V in positive mode to promote
efficient ionization of analytes. Fragmentation was facilitated
using MS/MS with normalized collision energies of 20, 40 and 60 V
applied via a rolling collision energy approach to enhance
fragmentation efficiency and specificity. Data acquisition was
performed using a data-dependent acquisition strategy, collecting
mass spectra across an m/z range of 70 to 1,050. This comprehensive
acquisition method ensured enhanced data quality and coverage,
supporting thorough identification and characterization of
compounds.
Metabolic data analysis
The initial processing of raw data derived from
LC/MS was performed using Progenesis QI software (version 3.0;
Waters Corporation). This sophisticated platform enabled the
construction of a three-dimensional data matrix saved in CSV
format, incorporating essential elements such as sample
identifiers, metabolite names and corresponding mass spectral
intensity values, all critical for subsequent analyses. Internal
standard peaks and known contaminants including noise, column bleed
and derivatization reagent signals were carefully removed to ensure
data integrity. Following this refinement, data redundancy was
minimized by pooling similar peaks. Metabolite identification was
performed through comprehensive cross-referencing against multiple
databases, primarily the Human Metabolome Database (HMDB,
http://www.hmdb.ca/), the METLIN database (https://metlin.scripps.edu/) and the proprietary
Majorbio Database (MIDB) maintained by Shanghai Majorbio
Biotechnology Co., Ltd. This rigorous approach facilitated accurate
and reliable metabolite profiling.
To ensure reliable compound annotation, metabolite
identification followed explicit criteria based on the
database-matching workflow. Specifically, precursor ions were
required to meet a mass accuracy threshold of <10 ppm, and
metabolite assignment further required a MS/MS spectral match score
above the identification cutoff used in Progenesis QI. Precursor
and fragment ions were matched against entries in HMDB, METLIN and
the Majorbio in-house MIDB database to support putative metabolite
identities.
The data matrix, which was annotated based on
database searches, was subsequently uploaded to the Majorbio cloud
platform (https://cloud.majorbio.com) for
additional analysis. The preprocessing of this data matrix involved
retaining metabolic features that were present in ≥80% of the
samples from any experimental group. For the treatment of missing
values, the minimum value observed within the dataset was used for
substitution, and each metabolic feature underwent normalization
relative to the overall signal sum. To address variability that
could stem from inconsistencies in sample preparation and
instrument fluctuations, mass spectrometric peak intensities were
subjected to sum normalization, resulting in a normalized data
matrix. Variables linked to QC samples showing a relative standard
deviation >30% were removed, and the data was log10 transformed
to stabilize the variance, culminating in the final matrix ready
for subsequent analyses. Variance analysis was then performed on
this refined dataset.
For the purpose of conducting multivariate
statistical analyses, the R package ‘ropls’ (version 1.6.2;
http://bioconductor.org/packages/ropls/) was employed
to execute principal component analysis (PCA) alongside orthogonal
partial least squares discriminant analysis (OPLS-DA). The model's
robustness was assessed via a seven-cycle interactive
cross-validation method. Metabolites were deemed statistically
significant if they exhibited a variable importance in projection
(VIP) score >1 within the OPLS-DA framework and a P-value
<0.05, evaluated using an unpaired Student's t-test.
The metabolites demonstrating considerable
differences between the two groups were comprehensively organized
into different biochemical pathways by employing metabolic
enrichment and pathway analysis, utilizing the information
available from the KEGG database. This process involved
categorizing the metabolites based on their corresponding
biological functions or pathways. To assess whether certain
functional groups of metabolites were statistically
overrepresented, an enrichment analysis was performed. This
approach allowed for a broader interpretation of the data, moving
beyond the identification of single metabolites to a more holistic
group-level understanding. The enrichment analysis was made
possible through the utilization of the Python package
‘scipy.stats’ (https://docs.scipy.org/doc/scipy/), which served a
crucial role in pinpointing the biological pathways that were most
pertinent to the specific experimental conditions under study.
Statistical correction for multiple
comparisons
For all metabolomic and proteomic comparisons,
P-values were adjusted using the Benjamini Hochberg FDR procedure,
as implemented on the Shanghai Majorbio Cloud Platform.
Differential metabolites and proteins were considered significant
only after FDR correction in addition to predefined fold-change or
VIP thresholds.
Results
Each of the two groups, invasive and non-invasive,
included 8 patients. The clinical and radiological characteristics
of these cohorts are detailed in Table
I. Patients in the invasive group were significantly older than
those in the non-invasive group (59.3±11.1 vs. 55.4±8.1 years;
P=0.0006). Regarding tumor morphology, the invasive group
demonstrated a substantially larger median preoperative tumor
volume [7,777.5 (5,398.9, 9,202.6) cm3] compared with
the non-invasive group [3,117.7 (1,570.7, 3,605.9) cm3;
P=0.0121]. Additionally, all 8 patients in the invasive cohort were
classified as Knosp grade ≥2, whereas none of the patients in the
non-invasive cohort met this threshold (P>0.9999). No
significant difference was observed in the sex distribution between
the two groups (P=0.6199). These results confirm significant
baseline differences in tumor aggressiveness and volume between the
two study groups (Table I).
 | Table I.Baseline characteristics of the
patients with pituitary adenoma included in the study. |
Table I.
Baseline characteristics of the
patients with pituitary adenoma included in the study.
| Characteristic | Invasive group
(n=8) | Non-invasive group
(n=8) |
P-valuea |
|---|
| Age, years | 59.3 (11.1) | 55.4 (8.1) | 0.0006 |
| Sexb, n (%) |
|
| 0.6199 |
|
Male | 5 (62.5) | 3 (37.5) |
|
|
Female | 3 (37.5) | 5 (62.5) |
|
| Tumor
volumec,
cm3 | 7,777.5 (5,398.9,
9,202.6) | 3,117.7 (1,570.7,
3,605.9) | 0.0121 |
| Invasive (Knosp
grade ≥2), n (%) | 8 (100) | 0 (0) | >0.9999 |
Identification of DEPs
Quantitative proteomics analysis revealed 614 DEPs
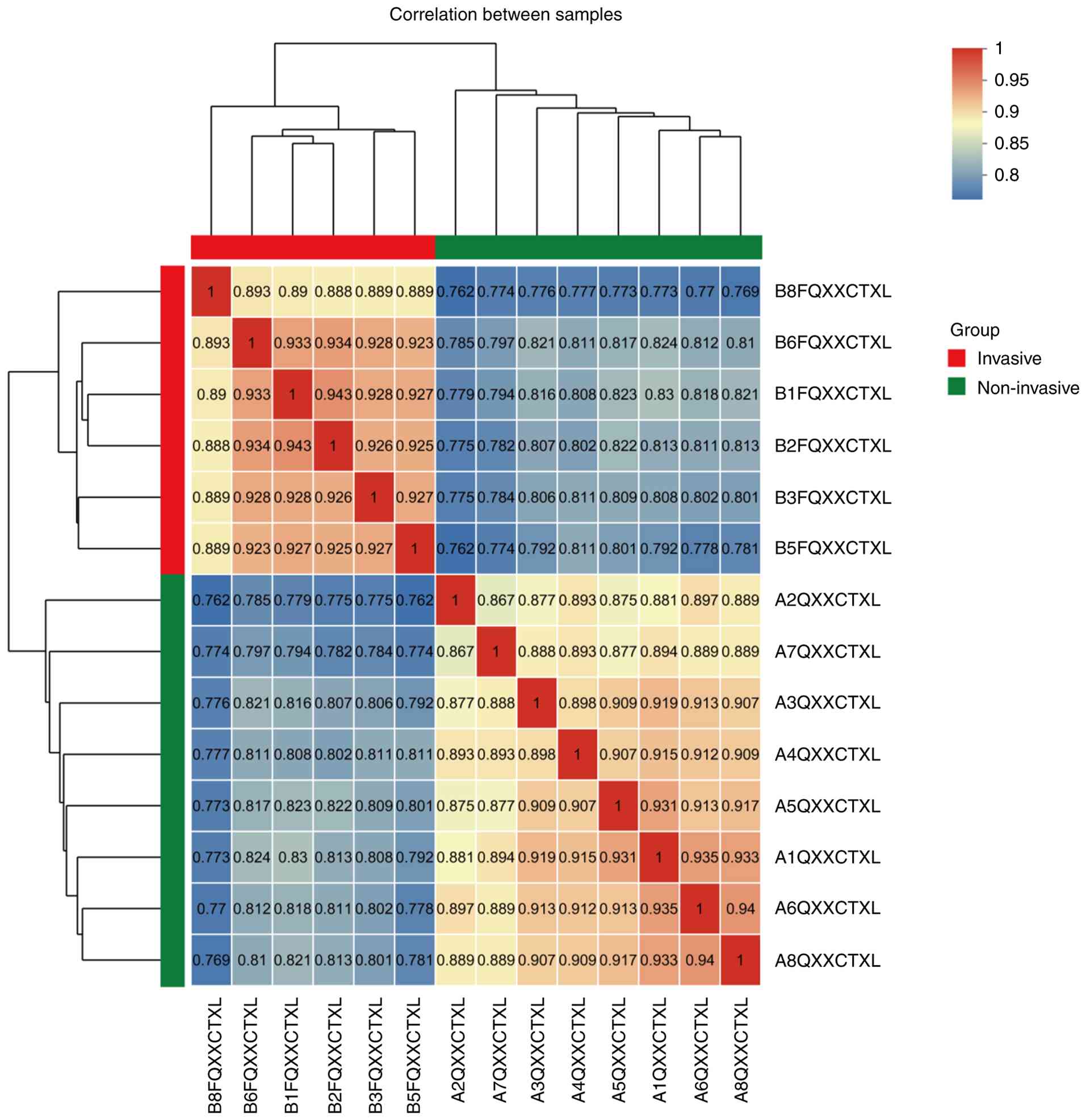
between the invasive and non-invasive groups. To gain insight into
group-specific functional differences, unsupervised hierarchical
clustering was performed on these proteins (Fig. 1). The results indicated low
Pearson's correlation coefficients of protein expression among
samples within groups, but high Pearson's correlation coefficients
between groups, suggesting clear proteomic differences between the
groups, whereas the proteomic profiles within each group were
relatively stable.
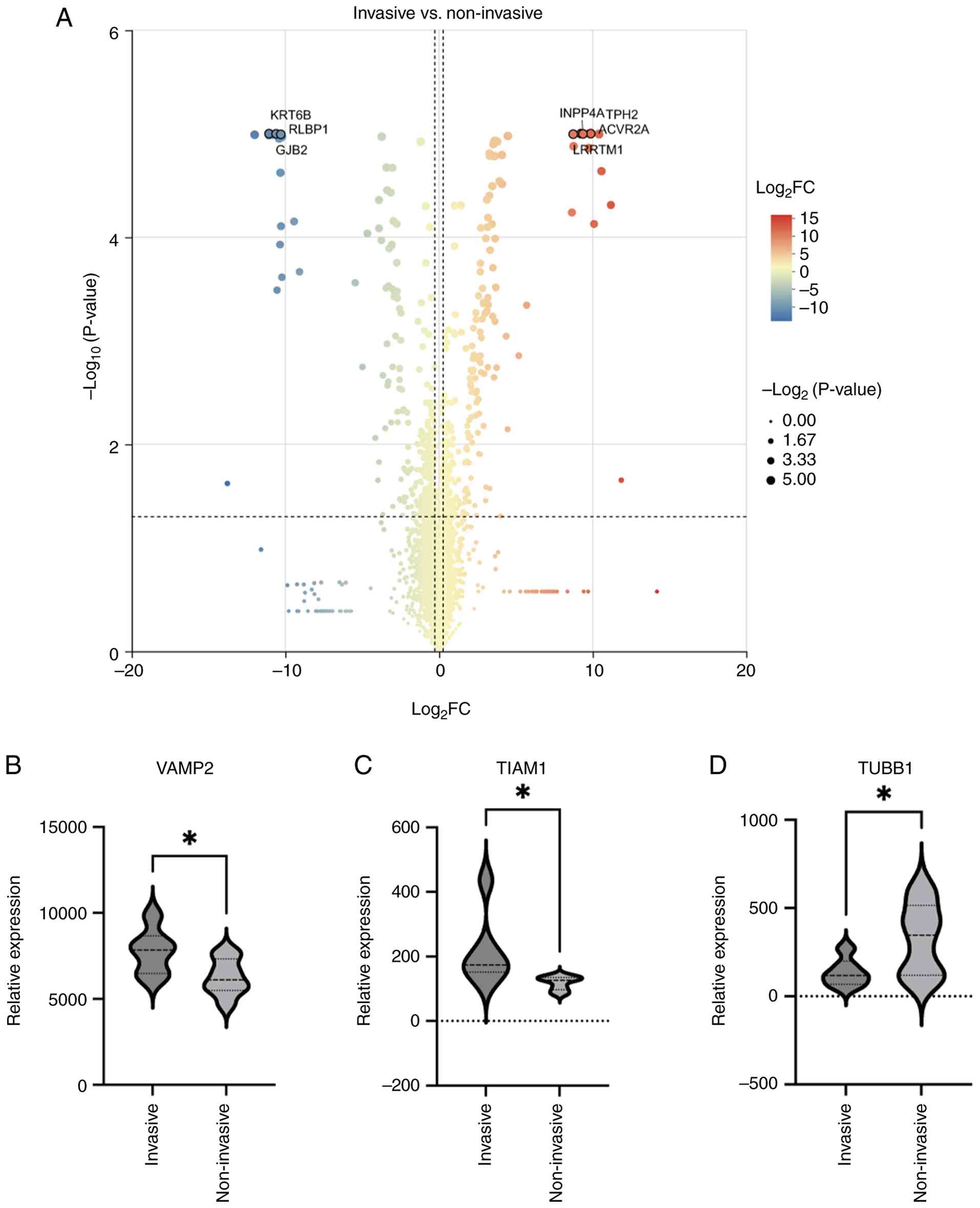
Subsequent comparative analyses were conducted to
identify and visualize DEPs between the invasive and non-invasive
groups using volcano plots. In total, 286 proteins exhibited
upregulation, while 328 proteins showed downregulation in the
invasive group when compared with the non-invasive cohort (Fig. 2A). The most upregulated proteins in
the invasive group included vesicle-associated membrane protein 2
(VAMP2) and T-cell lymphoma invasion and metastasis-inducing
protein 1, while tubulin b1 class VI (TUBB1) was among the most
significantly downregulated proteins (Fig. 2B-D).
Pathway enrichment analysis of
DEPs
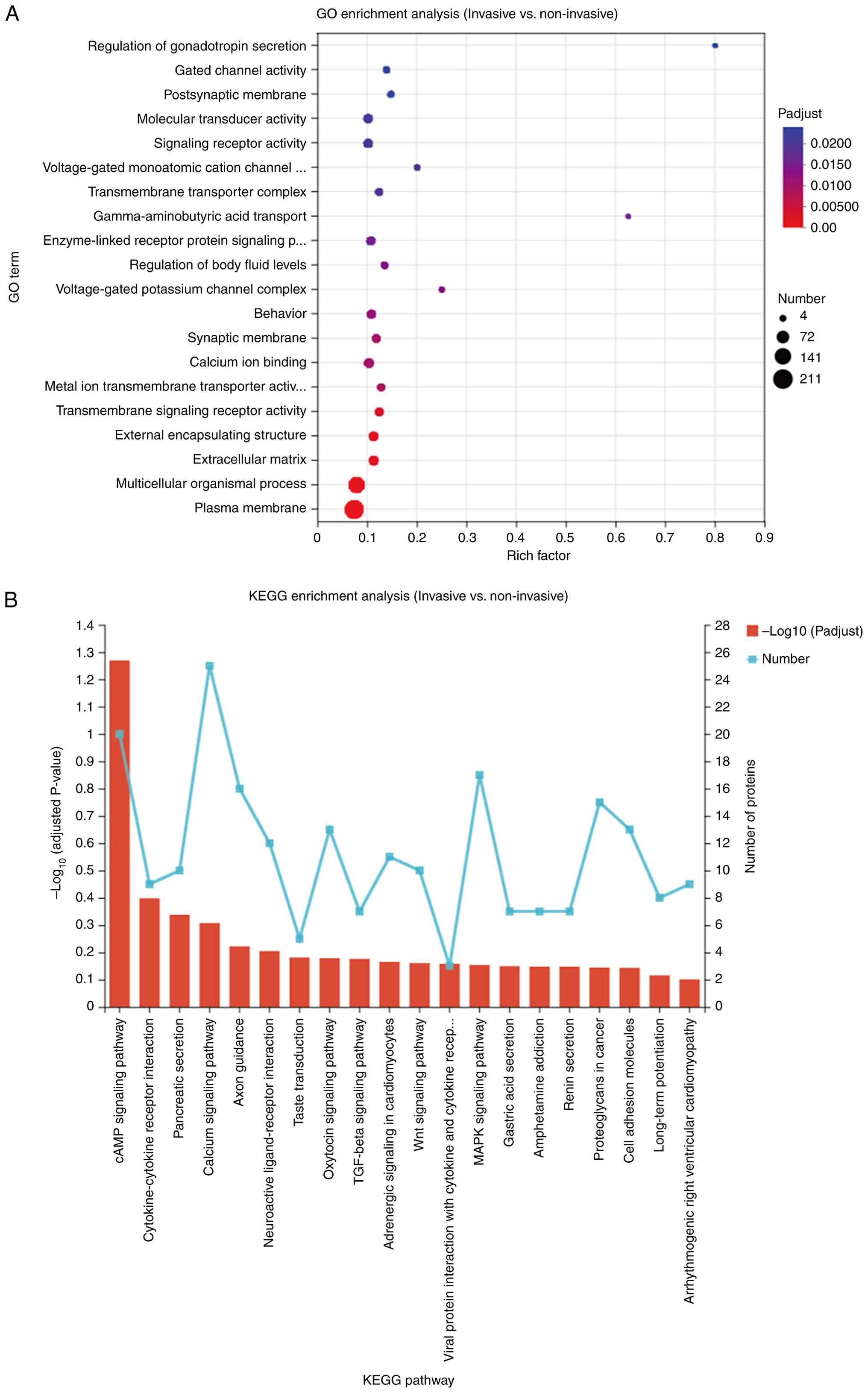
An analysis of enrichment was performed on the DEPs
to assess their functional significance. The GO analysis indicated
a notable enrichment of DEPs in various biological processes and
components, including the ‘regulation of gonadotropin secretion’,
‘gated channel activity’ and their localization at the
‘postsynaptic membrane’ (Fig. 3A).
Furthermore, a pathway analysis utilizing the KEGG suggested that
these proteins were primarily associated with the ‘cyclic adenosine
monophosphate (cAMP) signaling pathway’, ‘cytokine-cytokine
receptor interaction’, as well as ‘pancreatic secretion’ (Fig. 3B).
Identification of differentially
expressed metabolites (DEMs)
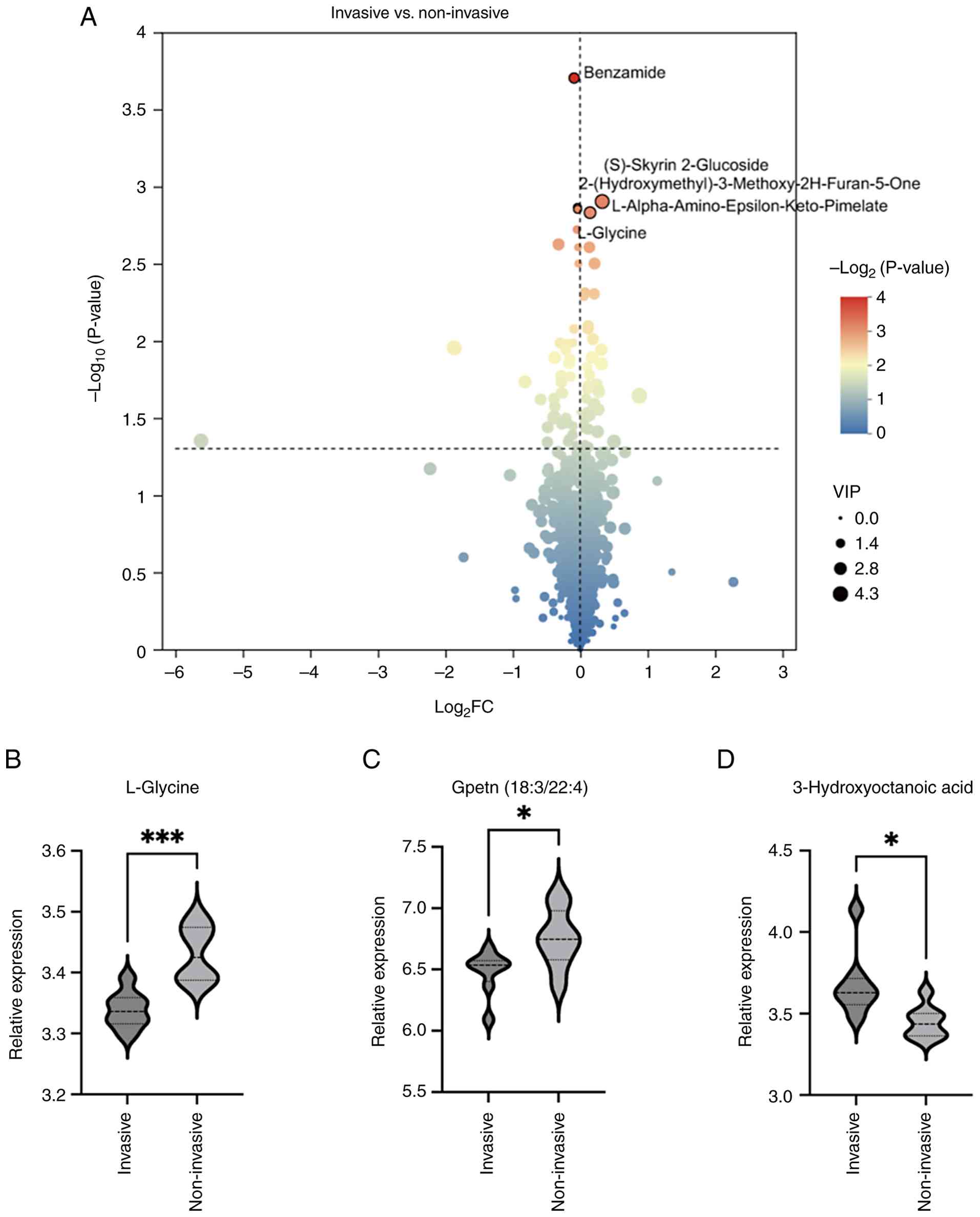
A comparative study was performed to identify and
illustrate the DEMs between the groups classified as invasive and
non-invasive, utilizing a volcano plot for visualization. In the
invasive group, 42 metabolites showed increased expression, while
32 metabolites were found to be decreased when compared with the
non-invasive group (Fig. 4A).
Compared with the non-invasive group, the metabolites that showed a
significant increase in the invasive group included
3-hydroxyoctanoic acid, whereas L-glycine and Gpetn(18:3/22:4)
exhibited a notable decrease (Fig.
4B-D).
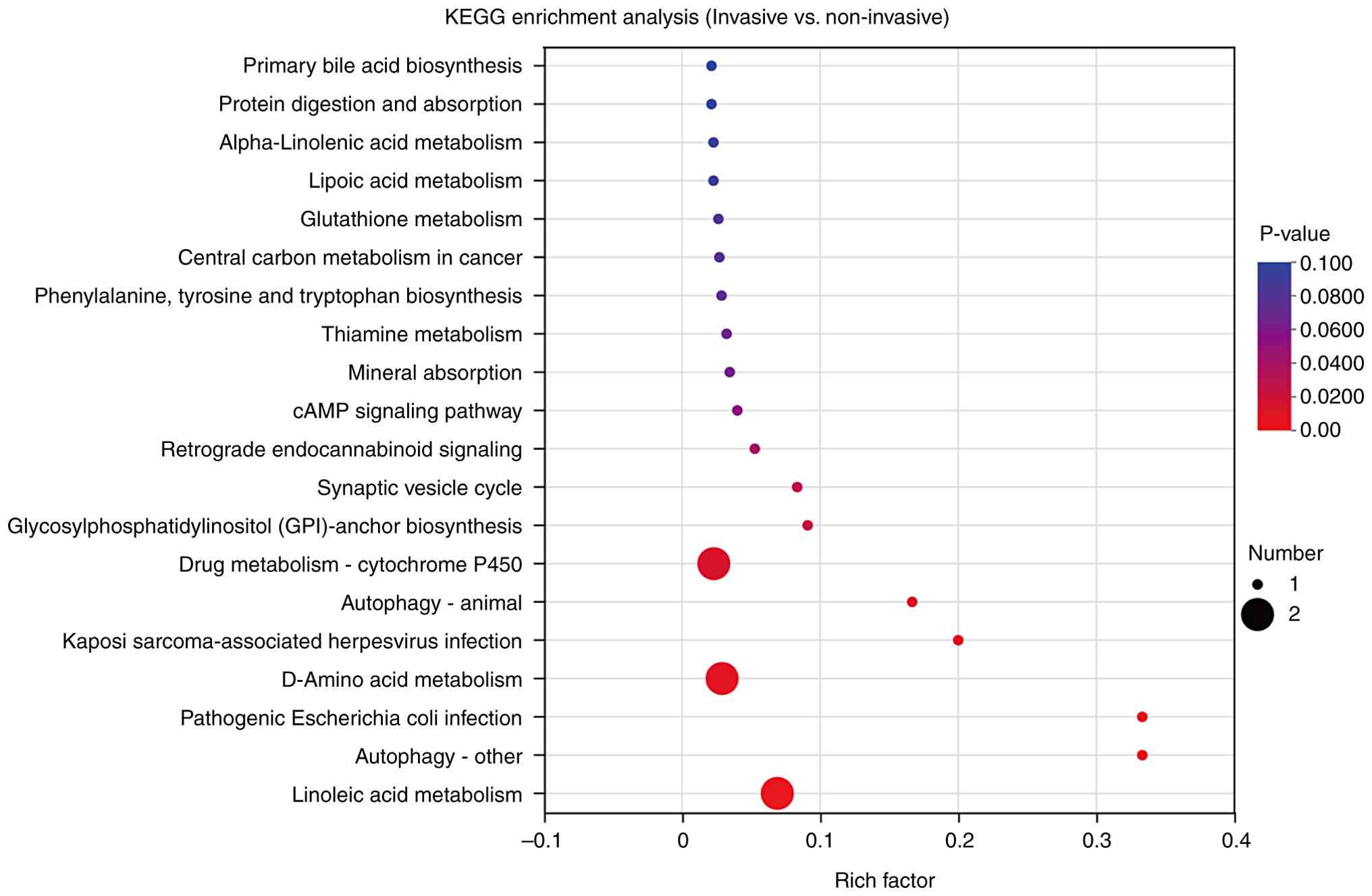
Metabolomics enrichment analysis
The examination of KEGG pathway enrichment for the
DEMs within the invasive and non-invasive groups revealed
significant enrichment across multiple biological pathways. These
pathways included the ‘primary bile acid biosynthesis’, ‘protein
digestion and absorption’, ‘alpha-linolenic acid metabolism’, the
‘cAMP signaling pathway’, the ‘synaptic vesicle cycle’ and pathways
associated with b ‘pathogenic Escherichia coli (E.
coli) infection’ (Fig. 5).
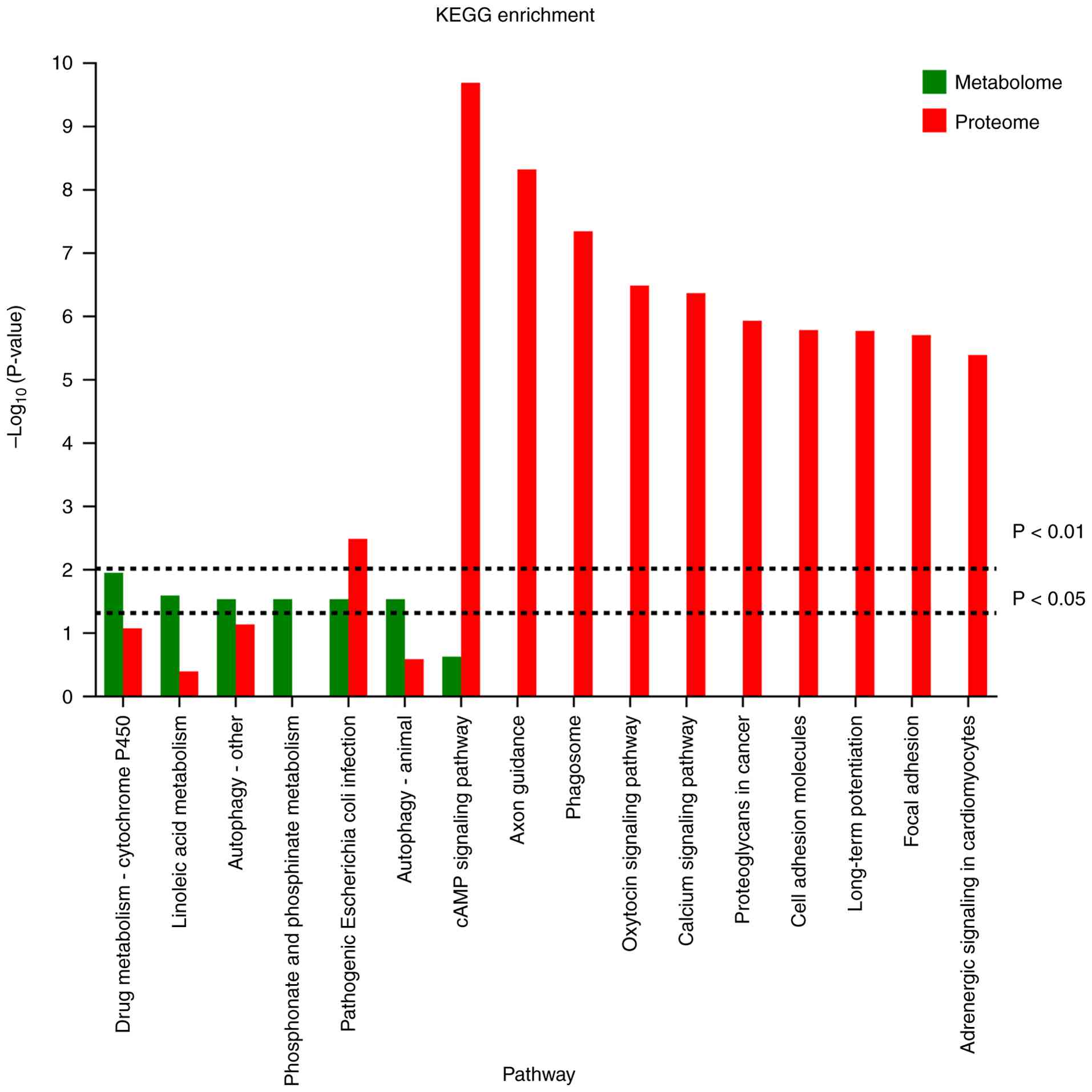
Combined proteomics and metabolomics
analyses
The results indicated that the proteins and
metabolites that were expressed differently in the invasive and
non-invasive groups were significantly enriched in pathways related
to pathogenic E. coli infection, the synaptic vesicle cycle
and cAMP signaling. This convergence suggests that these biological
pathways may be critically involved in the mechanisms underlying
pituitary tumor invasiveness (Fig.
6).
Discussion
The present study employed an integrative proteomic
and metabolomic approach to elucidate the molecular mechanisms
underlying the invasive phenotype of pituitary adenomas. By
comparing IPAs with non-invasive adenomas, distinct molecular
profiles were characterized at both protein and metabolite levels,
identifying pathways potentially contributing to tumor invasiveness
(12). The findings revealed unique
proteomic and metabolic changes in invasive tumors, marked by
differential expression of key proteins and metabolites enriched in
three critical pathways: The cAMP signaling cascade, pathogenic
E. coli infection and the synaptic vesicle cycle. These
results offer new insights into the molecular mechanisms driving
the aggressive behavior observed in a subset of pituitary
adenomas.
cAMP, originally identified as the first second
messenger, is critical in intracellular signaling pathways and
regulates a broad spectrum of physiological and pathological
processes (13). Its regulatory
actions are chiefly mediated through activation of protein kinase A
(PKA), which initiates downstream signaling events, including
modulation of gene transcription via the cAMP response
element-binding protein (14).
Furthermore, PKA phosphorylates various kinases, such as Raf and
glycogen synthase kinase 3 (15,16).
Altered signaling of cAMP-PKA has been linked to various human
tumors. Previous research has indicated that the cAMP signaling
pathway serves a role in several types of cancer, with its
tumor-promoting effects observed in hepatocellular carcinoma, lung
cancer, ovarian cancer, colorectal cancer and melanoma (17–21).
Moreover, as a classical signaling cascade, the cAMP pathway has
been demonstrated to promote the progression of pituitary adenomas
(22–24). The present data provide further
evidence supporting the involvement of the cAMP pathway in the
invasiveness of pituitary adenomas.
The KEGG pathway annotated as ‘pathogenic
Escherichia coli infection’ emerged as enriched in IPAs;
however, this does not imply the physical presence of bacteria
within the tumor tissue. Instead, this KEGG category mainly
reflects a set of host cellular processes that pathogenic E.
coli manipulate during infection, particularly actin
cytoskeleton remodeling, focal adhesion reorganization, membrane
trafficking and junctional disruption (25,26).
These same host-driven processes are well-recognized hallmarks of
cancer cell invasion (27).
Pathogenic E. coli effectors such as Tir, EspF and Map
activate MAPK, Rho GTPases and NF-κB pathways to induce
cytoskeletal rearrangement and promote pedestal formation in
epithelial cells; however, these pathways are fundamentally host
signaling modules that can be engaged in numerous non-infectious
contexts, including tumor progression (28). Therefore, the enrichment of this
pathway in invasive adenomas likely reflects activation of
actin-modifying, adhesion-modulating and endocytosis-related host
programs that overlap mechanistically with those annotated under
bacterial infection (27). In this
sense, the ‘E. coli infection’ label represents a functional
signature of cytoskeletal and membrane-dynamics pathways relevant
to cellular motility and invasion, rather than evidence for
microbial involvement.
However, beyond host signaling, the emerging role of
the tumor microbiome must also considered. Although the pituitary
gland has traditionally been viewed as a sterile environment,
landmark studies have previously revealed the presence of
intratumoral bacteria across various tumor types, suggesting a
potential role in carcinogenesis and therapeutic response (29–31).
In the specific context of pituitary adenomas, recent
investigations have identified distinct alterations in the gut
microbiota of patients with invasive tumors compared with those
with non-invasive subtypes, implying a functional ‘gut-pituitary
axis’ (32). Furthermore,
theoretical frameworks suggest that microbial metabolites could
influence pituitary tumor behavior and hormonal secretion through
systemic circulation (33). While
the present study focuses on host proteomic changes, the detection
of infection-related pathways highlights the need to consider these
potential microbial-host crosstalk mechanisms in future
comprehensive studies.
The synaptic vesicle cycle refers to the tightly
regulated process by which neurons release neurotransmitters at
synapses and then recycle the vesicle components for repeated use
(34). This cycle involves several
key steps: Vesicle docking, priming, Ca2+-dependent
exocytosis, endocytosis and vesicle recycling (35). Essential proteins in this pathway
include SNARE complexes, synaptotagmins, Rab GTPases and
clathrin-mediated endocytosis components. These proteins coordinate
membrane fusion and retrieval, maintaining synaptic transmission
fidelity and plasticity (36).
Although traditionally studied in the nervous system, previous
evidence suggests that components of the synaptic vesicle cycle are
aberrantly expressed or repurposed in various cancers. For
instance, proteins such as synaptotagmin have been found to be
dysregulated in gliomas (37). In
particular, previous studies have revealed that functional synaptic
connections can form between neurons and malignant brain tumors,
such as high-grade gliomas (38–40).
By contrast, direct evidence for such synaptic-like interactions in
pituitary adenomas is currently lacking. Therefore, while the
upregulation of synaptic vesicle cycle components in IPAs is
intriguing, any similarities to neuron-glioma interactions should
be regarded strictly as hypothesis-generating rather than
indicative of established tumor-neuron synaptic communication
(41). In the present study, the
significant enrichment of synaptic vesicle recycling pathways in
IPAs suggests that elements of vesicle trafficking and membrane
dynamics may be exploited to promote tumor invasiveness.
In the present study, IPAs displayed significant
enrichment of synaptic vesicle cycling components. This observation
raises an important mechanistic question: How might the cAMP-PKA
pathway intersect with vesicle-trafficking regulation in tumor
cells? In neurons, cAMP-activated PKA is a central modulator of
presynaptic release. PKA phosphorylates synapsin I, promoting the
mobilization of synaptic vesicles from the reserve pool to the
readily releasable pool, thereby increasing vesicle availability
for exocytosis (42). PKA also
phosphorylates key SNARE-associated regulators such as tomosyn,
reducing its inhibitory interaction with syntaxin-1 and
facilitating SNARE-complex assembly (43) and snapin, whose phosphorylation
strengthens the SNAP-25 interaction and enhances vesicle priming
(44).
Although VAMP2 itself is not a direct PKA substrate,
PKA-driven modulation of upstream regulators creates a biochemical
environment that favors efficient assembly of VAMP2 with syntaxin-1
and SNAP-25, ultimately increasing vesicle fusion probability
(45). This mechanistic framework
is highly relevant to the proteomic findings. VAMP2, one of the
most significantly upregulated proteins in invasive adenomas, is a
v-SNARE essential for membrane fusion and has been implicated in
promoting integrin trafficking and cancer cell migration. In
non-neuronal cancer systems, VAMP2 mediates α5β1-integrin delivery
to the plasma membrane, and its depletion reduces cell adhesion,
motility and viability (46,47).
Therefore, in a tumor context with heightened cAMP-PKA activity,
increased phosphorylation of synapsin, tomosyn and snapin may
enhance vesicle priming, accelerate SNARE-complex assembly and
potentiate trafficking of pro-invasive cargos (such as integrins,
metalloproteinases and growth factors) to the plasma membrane. Such
coordinated enhancement of vesicle dynamics provides a biologically
plausible rationale for the concurrent enrichment of cAMP signaling
and the synaptic vesicle cycle in IPAs. Together, these data
support a model in which invasive tumors co-opt neuro-modulatory
exocytosis programs to facilitate membrane remodeling,
microenvironmental interactions and ultimately invasion.
In addition to pathway-level alterations, several of
the top-ranked DEPs and DEMs provide further insight into the
invasive phenotype. Although VAMP2 has already been discussed in
the context of vesicle trafficking, its upregulation in IPAs is
consistent with prior evidence showing that VAMP2 facilitates
integrin delivery, membrane fusion and motility in cancer cells
(47), supporting its relevance to
invasion. Conversely, the marked downregulation of TUBB1, a
β-tubulin isoform involved in microtubule stability, aligns with
previous studies demonstrating that altered tubulin composition can
enhance cytoskeletal plasticity, chromosomal instability, and
migratory capacity in malignant cells (48). On the metabolic side, the elevated
level of 3-hydroxyoctanoic acid, a hydroxy fatty acid and
endogenous agonist of HCA3, is notable because activation of this
pathway has been linked to inflammatory signaling, metabolic
rewiring and lipid utilization in tumor settings (49). Activation of the HCA3 pathway has
been increasingly linked to inflammatory signaling, metabolic
rewiring and lipid utilization within various tumor
microenvironments. According to recent findings, 3-hydroxyoctanoic
acid acts as a potent agonist that induces natural biased signaling
at the HCA3 receptor (49). Unlike
other structurally similar agonists, 3-hydroxyoctanoic acid leads
to the notable recruitment of β-arrestin-2, a process that has been
shown to be relevant for regulating cell-cell adhesion and altering
spheroid density in 3D cell culture models. Specifically, the
activation of HCA3 by 3-hydroxyoctanoic acid can result in less
dense and larger cellular structures, suggesting that this
metabolic signaling may weaken intercellular adhesion within the
tumor mass. Furthermore, the HCA3-mediated signal involves Gβγ
subunits and downstream effectors such as PI3K, Rac1 and Ras/Rho,
which are crucial components for cellular motility and chemotaxis.
In the context of IPAs, the accumulation of 3-hydroxyoctanoic acid
may therefore prime tumor cells for a more infiltrative phenotype
by facilitating microtubule remodeling and vesicle-mediated
secretion (49). While
traditionally viewed as an anti-inflammatory mediator, the HCA3
axis in pituitary adenomas likely coordinates metabolic adaptation
to support the energetic demands of tumor invasion into adjacent
structures such as the cavernous sinus. Together, these molecular
signatures suggest that IPAs may coordinate vesicle-mediated
secretion, microtubule remodeling and fatty acid-driven metabolic
adaptation to support a more motile and infiltrative phenotype.
From a clinical perspective, these molecular
alterations offer potential biomarkers for the prediction of tumor
invasiveness and may inform therapeutic strategies. For example,
the cAMP analogue Tacladesine can inhibit the growth of lung
cancer, colon cancer, breast cancer, fibrosarcoma and leukemia
in vitro and in vivo (50). Moreover, the integration of
proteomic and metabolomic data enhances the ability to map
functional networks, offering a more comprehensive view of tumor
biology than either approach alone.
However, the present study has several limitations.
A limitation is the relatively small sample size, which might
reduce the statistical power required to detect minor differences
and restrict the applicability of the results. Additionally,
although the omics-focused analyses reveal connections between
molecular alterations and invasiveness, they do not confirm a
causal relationship. Functional validation of candidate proteins
and metabolites is needed to confirm their mechanistic roles in
tumor invasion. Finally, the heterogeneity of pituitary adenoma
subtypes (such as hormonally active vs. inactive) was not
stratified in the current analysis, which may confound some of the
observed molecular patterns.
Future studies should expand the cohort size and
include longitudinal data to assess whether these molecular markers
are associated with treatment response or recurrence. Moreover,
both in vivo and in vitro studies are necessary to
assess the importance of crucial pathways, including cAMP signaling
and synaptic vesicle cycling, in facilitating tumor invasiveness.
Single-cell multi-omics approaches may further uncover
cell-type-specific drivers of invasion and help to delineate
tumor-microenvironment interactions. Finally, a specific limitation
of the present study is the lack of metagenomic or 16S rRNA
sequencing analysis. Since the present workflow was designed for
host proteomics and metabolomics, we could not directly assess the
microbiome composition. Therefore, although the enrichment of
infection-related pathways was interpreted primarily as host
cytoskeletal mimicry, the potential influence of the intratumoral
or gut microbiome cannot be definitively ruled out. Future research
should incorporate microbiome-specific approaches to explore
whether direct bacterial-tumor interactions contribute to the
invasive phenotype of pituitary adenomas.
In summary, the present integrative analysis
highlights several signaling and metabolic pathways that may
contribute to the invasive phenotype of pituitary adenomas. These
findings advance the understanding of pituitary tumor biology and
open new avenues for prognostic biomarker development and
therapeutic intervention
Acknowledgements
Not applicable.
Funding
The present work was supported by the Natural Science Foundation
of Fujian Province (grant nos. 2022J05315, 2023J011625, 2023J011622
and 2024J08322), the Medical Project of Xiamen Municipal Bureau of
Science and Technology (grant no. 3502Z2024ZD1007), Xiamen Health
High Quality Development Technology Plan Project (grant nos.
2024GZL-GG95 and 2024GZL-CX38), Medical and Health Guidance Program
of Xiamen City, China (grant no. 3502Z20224ZD1012) and the
Scientific Research Initiation Fund for Introduced Talents of the
First Affiliated Hospital of Xiamen University (grant no.
XYJ2023001).
Availability of data and materials
The proteomics data generated in the present study
may be found in the iProX Consortium via the ProteomeXchange
partner repository under the accession number PXD071754 or at the
following URL:
https://proteomecentral.proteomexchange.org/cgi/GetDataset?ID=PXD071754.
The metabolomics data generated in the present study may be found
in the MetaboLights repository under the accession number
MTBLS13478 or at the following URL:
https://www.omicsdi.org/dataset/metabolights_dataset/MTBLS13478.
Authors' contributions
JM, XC and JS designed the study and oversaw the
implementation of the project. WZ and CW were responsible for the
acquisition of clinical samples. BL, SC and GT carried out the data
processing and interpreted the proteomic and metabolomic findings.
JM, XC, WZ and BL wrote the original draft. BL, CW and JS reviewed
and edited the manuscript. XC, PZ and JS were involved with
conceptualization of the study. ZL and YL were responsible for the
management of the proteomic and metabolomic databases and performed
the formal statistical analysis. SC and GT performed project
administration. JM, XC, WZ, ZL, CW and JL contributed to funding
acquisition. JM and JS confirm the authenticity of all the raw
data. All authors read and approved the final version of the
manuscript.
Ethics approval and consent to
participate
Approval of the research protocol by an
Institutional Reviewer Board. The present study was reviewed and
approved by the Ethics Committee of The First Affiliated Hospital
of Xiamen University [approval no. 2023KYEC (070)]. The present
study was performed in accordance with the principles of the
Declaration of Helsinki and written informed consent about the
participation in the research work was obtained from all patients
before surgery.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
Glossary
Abbreviations
Abbreviations:
|
DEP
|
differentially expressed protein
|
|
DEM
|
differentially expressed
metabolite
|
|
LC-MS/MS
|
liquid chromatography-tandem mass
spectrometry
|
References
|
1
|
Molitch ME: Diagnosis and treatment of
pituitary adenomas: A review. JAMA. 317:516–524. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Lefevre E, Chasseloup F, Hage M, Chanson
P, Buchfelder M and Kamenický P: Clinical and therapeutic
implications of cavernous sinus invasion in pituitary adenomas.
Endocrine. 85:1058–1065. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Lu L, Wan X, Xu Y, Chen J, Shu K and Lei
T: Prognostic factors for recurrence in pituitary adenomas: Recent
progress and future directions. Diagnostics (Basel). 12:9772022.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Chen L, White WL, Spetzler RF and Xu B: A
prospective study of nonfunctioning pituitary adenomas:
Presentation, management, and clinical outcome. J Neurooncol.
102:129–138. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Raverot G, Ilie MD, Lasolle H, Amodru V,
Trouillas J, Castinetti F and Brue T: Aggressive pituitary tumours
and pituitary carcinomas. Nat Rev Endocrinol. 17:671–684. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
6
|
James EL and Parkinson EK: Serum
metabolomics in animal models and human disease. Curr Opin Clin
Nutr Metabolic Care. 18:478–483. 2015.PubMed/NCBI
|
|
7
|
Kwon YW, Jo HS, Bae S, Seo Y, Song P, Song
M and Yoon JH: Application of proteomics in cancer: Recent trends
and approaches for biomarkers discovery. Front Med. 8:7473332021.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Knosp E, Steiner E, Kitz K and Matula C:
Pituitary adenomas with invasion of the cavernous sinus space: A
magnetic resonance imaging classification compared with surgical
findings. Neurosurgery. 33:610–617. 1993. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Cox J, Hein MY, Luber CA, Paron I, Nagaraj
N and Mann M: Accurate proteome-wide label-free quantification by
delayed normalization and maximal peptide ratio extraction, termed
MaxLFQ. Mol Cell Proteomics. 13:2513–2526. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Smyth GK: Linear models and empirical
bayes methods for assessing differential expression in microarray
experiments. Stat Appl Genet Mol Biol. 3:1–25. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Szklarczyk D, Kirsch R, Koutrouli M,
Nastou K, Mehryary F, Hachilif R, Gable AL, Fang T, Doncheva NT,
Pyysalo S, et al: The STRING database in 2023: Protein-protein
association networks and functional enrichment analyses for any
sequenced genome of interest. Nucleic Acids Res. 51:D638–D646.
2023. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Casado-Vela J, Cebrián A, del Pulgar MT
and Lacal JC: Approaches for the study of cancer: Towards the
integration of genomics, proteomics and metabolomics. Clin Transl
Oncol. 13:617–628. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Sassone-Corsi P: The cyclic AMP pathway.
Cold Spring Harb Perspect Biol. 4:a0111482012. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Steven A, Friedrich M, Jank P, Heimer N,
Budczies J, Denkert C and Seliger B: What turns CREB on? And off?
And why does it matter? Cell Mol Life Sci. 77:4049–4067. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Häfner S, Adler HS, Mischak H, Janosch P,
Heidecker G, Wolfman A, Pippig S, Lohse M, Ueffing M and Kolch W:
Mechanism of inhibition of raf-1 by protein kinase a. Mo Cell Biol.
14:6696–6703. 1994. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Jensen J, Brennesvik EO, Lai YC and
Shepherd PR: GSK-3beta regulation in skeletal muscles by adrenaline
and insulin: Evidence that PKA and PKB regulate different pools of
GSK-3. Cell Signal. 19:204–210. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Wang M, Li Y, Wang R, Wang Z, Chen K, Zhou
B, Zhou Z and Sun X: PKA RIα/a-kinase anchoring proteins 10
signaling pathway and the prognosis of colorectal cancer. J
Gastroenterol Hepatol. 30:496–503. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Finger EC, Castellini L, Rankin EB,
Vilalta M, Krieg AJ, Jiang D, Banh A, Zundel W, Powell MB and
Giaccia AJ: Hypoxic induction of AKAP12 variant 2 shifts
PKA-mediated protein phosphorylation to enhance migration and
metastasis of melanoma cells. Proc Natl Acad Sci USA.
112:4441–4446. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Shaikh D, Zhou Q, Chen T, Ibe JCF, Raj JU
and Zhou G: cAMP-dependent protein kinase is essential for
hypoxia-mediated epithelial-mesenchymal transition, migration, and
invasion in lung cancer cells. Cell Signal. 24:2396–2406. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Tonucci FM, Almada E, Borini-Etichetti C,
Pariani A, Hidalgo F, Rico MJ, Girardini J, Favre C, Goldenring JR,
Menacho-Marquez M and Larocca MC: Identification of a CIP4 PKA
phosphorylation site involved in the regulation of cancer cell
invasiveness and metastasis. Cancer Lett. 461:65–77. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
McKenzie AJ, Campbell SL and Howe AK:
Protein kinase A activity and anchoring are required for ovarian
cancer cell migration and invasion. PLoS One. 6:e265522011.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Bizzi MF, Bolger GB, Korbonits M and
Ribeiro-Oliveira Jr: Phosphodiesterases and cAMP pathway in
pituitary diseases. Front Endocrinol (Lausanne). 10:1412019.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Lania A, Mantovani G and Spada A: cAMP
pathway and pituitary tumorigenesis. Ann Endocrinol (Paris).
73:73–75. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Pertuit M, Barlier A, Enjalbert A and
Gérard C: Signalling pathway alterations in pituitary adenomas:
Involvement of gsα, cAMP and mitogen-activated protein kinases. J
Neuroendocrinol. 21:869–877. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Singh AP, Sharma S, Pagarware K, Siraji
RA, Ansari I, Mandal A, Walling P and Aijaz S: Enteropathogenic E.
coli effectors EspF and Map independently disrupt tight junctions
through distinct mechanisms involving transcriptional and
post-transcriptional regulation. Sci Rep. 8:37192018. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Croxen MA and Finlay BB: Molecular
mechanisms of escherichia coli pathogenicity. Nat Rev Microbiol.
8:26–38. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Li X and Wang J: Mechanical tumor
microenvironment and transduction: Cytoskeleton mediates cancer
cell invasion and metastasis. Int J Biol Sci. 16:2014–2028. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Aseervatham J: Cytoskeletal remodeling in
cancer. Biology (Basel). 9:3852020.PubMed/NCBI
|
|
29
|
Nejman D, Livyatan I, Fuks G, Gavert N,
Zwang Y, Geller LT, Rotter-Maskowitz A, Weiser R, Mallel G, Gigi E,
et al: The human tumor microbiome is composed of tumor
type-specific intracellular bacteria. Science. 368:973–980. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Niño JL, Wu H, LaCourse KD, Kempchinsky
AG, Baryiames A, Barber B, Futran N, Houlton J, Sather C, Sicinska
E, et al: Effect of the intratumoral microbiota on spatial and
cellular heterogeneity in cancer. Nature. 611:810–817. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Kalaora S, Nagler A, Nejman D, Alon M,
Barbolin C, Barnea E, Ketelaars SLC, Cheng K, Vervier K, Shental N,
et al: Identification of bacteria-derived HLA-bound peptides in
melanoma. Nature. 592:138–143. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Hu J, Yang J, Chen L, Meng X, Zhang X, Li
W, Li Z and Huang G: Alterations of the gut microbiome in patients
with pituitary adenoma. Pathol Oncol Res. 28:16104022022.
View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Nie D, Li C and Zhang Y: PitNETs and the
gut microbiota: Potential connections, future directions. Front
Endocrinol (Lausanne). 14:12559112023. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Südhof TC: The synaptic vesicle cycle. Ann
Rev Neurosci. 27:509–547. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Chanaday NL, Cousin MA, Milosevic I,
Watanabe S and Morgan JR: The synaptic vesicle cycle revisited: New
insights into the modes and mechanisms. J Neurosci. 39:8209–8216.
2019. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Rizo J and Südhof TC: The membrane fusion
enigma: SNAREs, Sec1/Munc18 proteins, and their accomplices-guilty
as charged? Annu Rev Cell Dev Biol. 28:279–308. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Sheng B, Jiang Y, Wu D, Lai N, Ye Z, Zhang
B, Fang X and Xu S: RNAi-mediated SYT14 knockdown inhibits the
growth of human glioma cell line U87MG. Brain Res Bull. 140:60–64.
2018. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Taylor KR, Barron T, Hui A, Spitzer A,
Yalçin B, Ivec AE, Geraghty AC, Hartmann GG, Arzt M, Gillespie SM,
et al: Glioma synapses recruit mechanisms of adaptive plasticity.
Nature. 623:366–374. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Venkatesh HS, Morishita W, Geraghty AC,
Silverbush D, Gillespie SM, Arzt M, Tam LT, Espenel C, Ponnuswami
A, Ni L, et al: Electrical and synaptic integration of glioma into
neural circuits. Nature. 573:539–545. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Venkataramani V, Tanev DI, Strahle C,
Studier-Fischer A, Fankhauser L, Kessler T, Körber C, Kardorff M,
Ratliff M, Xie R, et al: Glutamatergic synaptic input to glioma
cells drives brain tumour progression. Nature. 573:532–538. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Winkler F, Venkatesh HS, Amit M, Batchelor
T, Demir IE, Deneen B, Gutmann DH, Hervey-Jumper S, Kuner T,
Mabbott D, et al: Cancer neuroscience: State of the field, emerging
directions. Cell. 186:1689–1707. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Greengard P, Valtorta F, Czernik AJ and
Benfenati F: Synaptic vesicle phosphoproteins and regulation of
synaptic function. Science. 259:780–785. 1993. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Baba T, Sakisaka T, Mochida S and Takai Y:
PKA-catalyzed phosphorylation of tomosyn and its implication in
Ca2+-dependent exocytosis of neurotransmitter. J Cell Biol.
170:1113–1125. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Chheda MG, Ashery U, Thakur P, Rettig J
and Sheng ZH: Phosphorylation of Snapin by PKA modulates its
interaction with the SNARE complex. Nat Cell Biol. 3:331–338. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Clementi F, Fesce R, Meldolesi J, Valtorta
F, Hilfiker S, Pieribone VA, Czernik AJ, Kao HT, Augustine GJ and
Greengard P: Synapsins as regulators of neurotransmitter release.
Philosophical Transactions of the Royal Society of London Series B:
Biological Sciences. 354:269–279. 1999. View Article : Google Scholar
|
|
46
|
Raja SA, Abbas S, Shah STA, Tariq A, Bibi
N, Yousuf A, Khawaja A, Nawaz M, Mehmood A, Khan MJ and Hussain A:
Increased expression levels of Syntaxin 1A and Synaptobrevin
2/vesicle-associated membrane protein-2 are associated with the
progression of bladder cancer. Genet Mol Biol. 42:40–47. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Hasan N and Hu C: Vesicle-associated
membrane protein 2 mediates trafficking of alpha5beta1 integrin to
the plasma membrane. Exp Cell Res. 316:12–23. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Trisciuoglio D and Degrassi F: The tubulin
code and tubulin-modifying enzymes in autophagy and cancer. Cancers
(Basel). 14:62021. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Peters A, Rabe P, Krumbholz P, Kalwa H,
Kraft R, Schöneberg T and Stäubert C: Natural biased signaling of
hydroxycarboxylic acid receptor 3 and G protein-coupled receptor
84. Cell Commun Signal. 18:312020. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Cho-Chung YS and Nesterova MV: Tumor
reversion: Protein kinase A isozyme switching. Ann N Y Acad Sci.
1058:76–86. 2005. View Article : Google Scholar : PubMed/NCBI
|