Introduction
Esophageal cancer is among the leading causes of
cancer-related mortality worldwide (1). Each year, ~16,120 individuals in the
United States are diagnosed with esophageal cancer (2). Despite advances in diagnostic
techniques and treatment options, the prognosis for individuals
with esophageal cancer remains unfavorable, with 5-year survival
rates typically ranging between 15 and 25% (3,4).
Notably, esophageal squamous cell carcinoma (ESCC) accounts for
~90% of cases of esophageal cancer (5). However, despite advances in screening
and therapeutic interventions, patient outcomes remain poor, and a
considerable subset of patients continues to experience disease
recurrence or progression (6,7).
Therefore, the discovery of novel targets for the management of
ESCC are required.
In the search for such novel targets, dysregulation
of fundamental cellular processes such as alternative splicing has
emerged as a key area of investigation in oncology. Alternative
splicing is a pivotal regulatory mechanism that governs gene
expression and increases protein diversity (8). This process is carried out by the
spliceosome, a complex ribonucleoprotein machine. Small nuclear
ribonucleoprotein polypeptides B (SNRPB) and B' are essential
components of this machinery (9).
Accumulating evidence has associated SNRPB to
various human diseases. For instance, aberrations in SNRPB
contribute to the pathogenesis of cerebrocostomandibular syndrome
(10,11) and carry out a role in regulating
bone and cartilage differentiation (12). In the context of cancer, SNRPB has
been identified as a key oncogenic driver. It has been shown to
promote cell proliferation in lung cancer (13) and regulate the cell cycle to drive
the progression of liver cancer (14). Furthermore, in cervical and thyroid
cancer, SNRPB exerts its effects by inhibiting the tumor suppressor
protein p53 (15,16). Although previous studies have
demonstrated that SNRPB plays a tumor-promoting role in various
types of cancer, including effects on cell proliferation and
apoptosis (13,17–19),
there is currently a lack of data regarding its expression,
functional role and clinical relevance in ESCC. Therefore, the
present study focused on elucidating the role of SNRPB in ESCC.
The present study aimed to characterize SNRPB
expression in ESCC and clarify its functional role in tumor
progression; SNRPB expression and clinical relevance in ESCC was
first analyzed using bioinformatics tools and immunohistochemical
staining of clinical tissue samples, subsequently the biological
effects of SNRPB silencing on ESCC cells were investigated using
lentivirus-mediated shRNA knockdown in vitro. Thus, we
hypothesized that SNRPB may promote ESCC progression through
impacting cell proliferation and apoptosis, positioning it as a
potential prognostic biomarker and therapeutic target.
Materials and methods
Prognostic analysis of SNRPB mRNA
In the present study, Gene Expression Profiling
Interactive Analysis (GEPIA; (http://gepia.cancer-pku.cn/)) was used to assess the
differential expression of SNRPB between ESCC and matched healthy
tissues (20). Transcriptome data
and clinical information of 161 patients with esophageal cancer and
12 healthy esophageal tissue samples were downloaded from The
Cancer Genome Atlas (TCGA; http://www.cancer.gov/ccg/research/genome-sequencing/tcga).
Data were obtained from The Cancer Genome Atlas esophageal
carcinoma (ESCA) cohort, which includes both esophageal squamous
cell carcinoma (ESCC) and esophageal adenocarcinoma (EAC). The
SNRPB mRNA expression matrix was extracted from the dataset for
subsequent analysis. The association between the expression levels
of SNRPB and the survival outcomes of patients with ESCC was
assessed using Kaplan-Meier survival analysis. Univariate and
multivariate Cox regression analyses were performed via the
‘survival’ R software package (version 3.8–3) (21) to explore the independent prognostic
factors of patients with ESCC (21).
Clinical sample collection
Samples of both neoplastic and pair-matched adjacent
non-cancerous tissues, preserved in paraffin blocks, were collected
from a cohort of 50 patients diagnosed with ESCC at the Affiliated
Hospital of North Sichuan Medical College, Sichuan, China. The
adjacent tissues were obtained from sites at least 5 cm away from
the tumor margin and were histologically confirmed to be free of
cancer cells by a certified pathologist. These patients received
surgical interventions at the Affiliated Hospital of North Sichuan
Medical College (Sichuan, China) from January 2019 to December
2021. The present study was approved by the Medical Ethics
Committee of the Affiliated Hospital of North Sichuan Medical
College, Sichuan, China (approval no. 2023ER344-1) and followed the
guidelines outlined in the Declaration of Helsinki. As the present
study was retrospective and patient data was anonymized, the
Medical Ethics Committee of the Affiliated Hospital of North
Sichuan Medical College waived the requirement for informed
consent. In total, 50 patients with ESCC (median age, 60 years;
range, 45–77; 38 men and 12 women) were included, with tumor stages
I (n=15), II (n=9), III (n=22) and IV (n=4). Characteristics of the
50 patients are summarized in Table
SI.
Inclusion criteria were as follows: i)
Histologically confirmed primary ESCC; ii) a history of curative
resection between January 2019 and December 2021; and iii) no
history of neoadjuvant radiotherapy or chemotherapy prior to
surgery. Patients were excluded from the present study according to
the following criteria: i) Incomplete clinical or follow-up data;
and ii) a history of other synchronous or metachronous malignant
tumors.
Immunohistochemical analysis
Immunohistochemistry was performed to assess the
expression of SNRPB in tissues. Tissue samples were formalin-fixed
and paraffin-embedded, with 4-µm-thick serial sections prepared for
staining. Permeabilization was not required for the staining
procedure. Prior to antibody incubation, sections were blocked with
5% bovine serum albumin in phosphate-buffered saline (PBS) at room
temperature for 1 h. The primary antibody against SNRPB (cat. no.
10278-1-AP; Proteintech Group, Inc.) was diluted at 1:100 and
incubated with the sections at 4°C overnight (12 h). The secondary
antibody used was HRP-conjugated goat anti-rabbit IgG (cat. no.
SA00001-2; Proteintech Group, Inc.), diluted at 1:500 and incubated
at room temperature for 1 h. All stained sections were observed and
imaged using a Leica DM4 B upright light microscope (Leica
Microsystems). To ensure the specificity and reliability of the
staining, positive and negative controls were included in each
staining batch. Colon cancer tissue sections, known to have high
expression of SNRPB, were used as a positive control. For the
negative control, the primary antibody was replaced with PBS. The
immunoreactive score (IRS) scoring system was used to evaluate the
expression of SNRPB (22).
Cell culture and transfection
ESCC cell lines, KYSE150, KYSE510, KYSE30 and
KYSE410, were purchased from Saiku Biotechnology. All cell lines
were cultivated in RPMI-1640 medium enriched with 10% fetal bovine
serum, and maintained in an incubator at 37°C with 5%
CO2. After confirming the high expression of SNRPB in
ESCC tissues (Fig. 1), western blot
analysis was performed to detect its expression in ESCC cell lines,
which showed markedly higher endogenous SNRPB levels in KYSE30 and
KYSE510 cells compared to the other tested lines (Fig. 2A). These two cell lines were thus
selected for subsequent functional knockdown experiments. ShRNA
constructs were integrated into the
hU6-MCS-CBh-gcGFP-IRES-puromycin vector, and lentiviral particles
with a titer of 1×108 TU/ml were used for transduction;
2 µl of lentiviral particles were added per well in a 6-well plate,
combined with HiTransG A Lentiviral Transduction Reagent (GeneChem,
Inc.). The transfection was performed at 37°C with 5%
CO2 for 8 h, and all subsequent functional assays were
conducted at a 48 h time interval post-transfection. The lentiviral
infection was performed at a multiplicity of infection of 10, which
was determined as optimal in preliminary experiments to achieve
high transduction efficiency with minimal cytotoxicity. Design and
production of lentiviral constructs were carried out by Shanghai
GeneChem Co., Ltd. All procedures involving lentiviral particles
were performed in a Biosafety Level 2 (BSL-2) containment facility.
The specific shRNA sequences used for SNRPB knockdown were as
follows: shSNRPB#1, CCACAAGGAAGAGGTACTGTT; shSNRPB#2,
CCTCCCAAAGATACTGGTATT; and shSNRPB#3, GCAGCATATTGATTACAGGAT. The
sequence for the negative control (NC) was as follows: shNC,
TTCTCCGAACGTGTCACGT.
Colony formation assay
Following transfection with shSNRPB#1, KYSE 30 and
KYSE 510 cells were maintained in RPMI-1640 medium with 10% fetal
bovine serum at 37°C with 5% CO2 for 12 days
post-transfection, with the medium replaced every three days and no
additional drug treatment applied in this experiment. Subsequently,
cells were digested to prepare single-cell suspensions, and 1,000
cells per well were inoculated in 6-well plates, with control cells
treated with a blank vector. After ~10 days, colonies were fixed
with 4% paraformaldehyde at room temperature for 15 min, then
stained with 0.1% crystal violet solution at room temperature for
20 min. Colonies containing >50 cells were counted to calculate
the clonogenic survival rate. Colony counting was performed
manually by two independent investigators who were blinded to the
experimental group assignments.
Cell cycle analysis
SNRPB-knockdown KYSE30 and KYSE510 cells
(transfected with shSNRPB#1) and negative control cells
(transfected with shNC) were used for this assay, and analysis was
performed 48 h after lentiviral transfection (consistent with the
time interval for other subsequent functional experiments). For
sample preparation, the cells were harvested by trypsinization with
0.25% trypsin-EDTA (Gibco; Thermo Fisher Scientific, Inc.), washed
twice with pre-chilled PBS, and fixed with 70% cold ethanol at 4°C
for 24 h for cell cycle arrest. A Cell Cycle Detection kit (Nanjing
KeyGen Biotech Co., Ltd.) was then used to assess cell cycle
distribution according to the manufacturer's protocol. Analysis was
conducted via a NovoCyte flow cytometer (NovoCyte 3130; ACEA
Biosciences, Inc.), which allowed precise cell cycle phase
determination.
Apoptosis analysis
SNRPB-knockdown KYSE30 and KYSE510 cells
(transfected with shSNRPB#1) and negative control cells
(transfected with shNC) were used for this assay. For apoptosis
analysis, the Annexin V-APC/PI Apoptosis Detection kit (cat. no.
KGA1030-100; Nanjing KeyGen Biotech Co., Ltd.) was utilized
according to the manufacturer's protocol. A NovoCyte flow cytometer
(ACEA Biosciences, Inc.) was used to quantify the proportion of
apoptotic cells.
Western blot analysis
Intracellular proteins were obtained by lysing
SNRPB-knockdown and negative control KYSE30 and KYSE510 cells using
RIPA buffer (Beijing Applygen Technologies Inc.), with the addition
of phosphatase inhibitors. Total proteins were quantified using a
BCA Protein Assay Kit (Beyotime Biotechnology) to determine the
concentration, and 30 µg of total protein was loaded into each lane
for separation by SDS-PAGE on a 12% gel and transferred onto a PVDF
membrane (Sigma-Aldrich; Merck KGaA). Membranes were subsequently
blocked with a solution containing 5% skimmed milk powder (Beyotime
Biotechnology) for 2 h at room temperature. Following blocking,
membranes were incubated with the following primary antibodies
overnight at 4°C: Anti-SNRPB (cat. no. FNab08070; 1:1,000;
FineTest), anti-GAPDH (cat. no. EM1101; 1:50,000; HUABIO),
anti-CDK1 (cat. no. ET1607-54; 1:1,000; HUABIO), anti-CCNA2 (cat.
no. : ET1612-26; 1:1,000; HUABIO), anti-BAX (cat. no. ET1603-34;
1:5,000; HUABIO) and anti-Bcl-2 (cat. no. : HA721235; 1:2,000;
HUABIO). Following primary incubation, membranes were incubated
with a secondary antibody (cat. no. FNSA-0004; 1:5,000; FineTest,
goat anti-rabbit IgG HRP-conjugated) for 1 h at room temperature.
Protein bands were visualized using a ChemiDoc™ XRS+ system
(Universal Hood II; Bio-Rad Laboratories, Inc.). Protein expression
was quantified using ImageJ software (version 1.54f; National
Institutes of Health) with GAPDH serving as the loading
control.
Statistical analysis
R software (version, 4.2.2; Foundation for
Statistical Computing) and GraphPad Prism 8 (Dotmatics) were used
for data analysis and visualization. Differences between two groups
were evaluated using paired or unpaired Student's t tests for
pairwise comparisons of independent or matched samples (specified
in corresponding figure legends). One-way ANOVA followed by
Dunnett's post hoc test was used for multiple group comparisons.
The associations between SNRPB expression and clinicopathological
characteristics were assessed using the chi-square (χ2)
test for variables meeting the expected count assumption; Fisher's
exact test (or the Freeman-Halton extension for >2×2 contingency
tables) was used for Histological grade, Location and M stage
(23). Cox regression analysis
identified independent prognostic factors for esophageal cancer.
P<0.05 was considered to indicate a statistically significant
difference. All in vitro experiments were performed in
triplicate, as independent biological replicates.
Results
SNRPB is highly expressed in ESCC
tissues
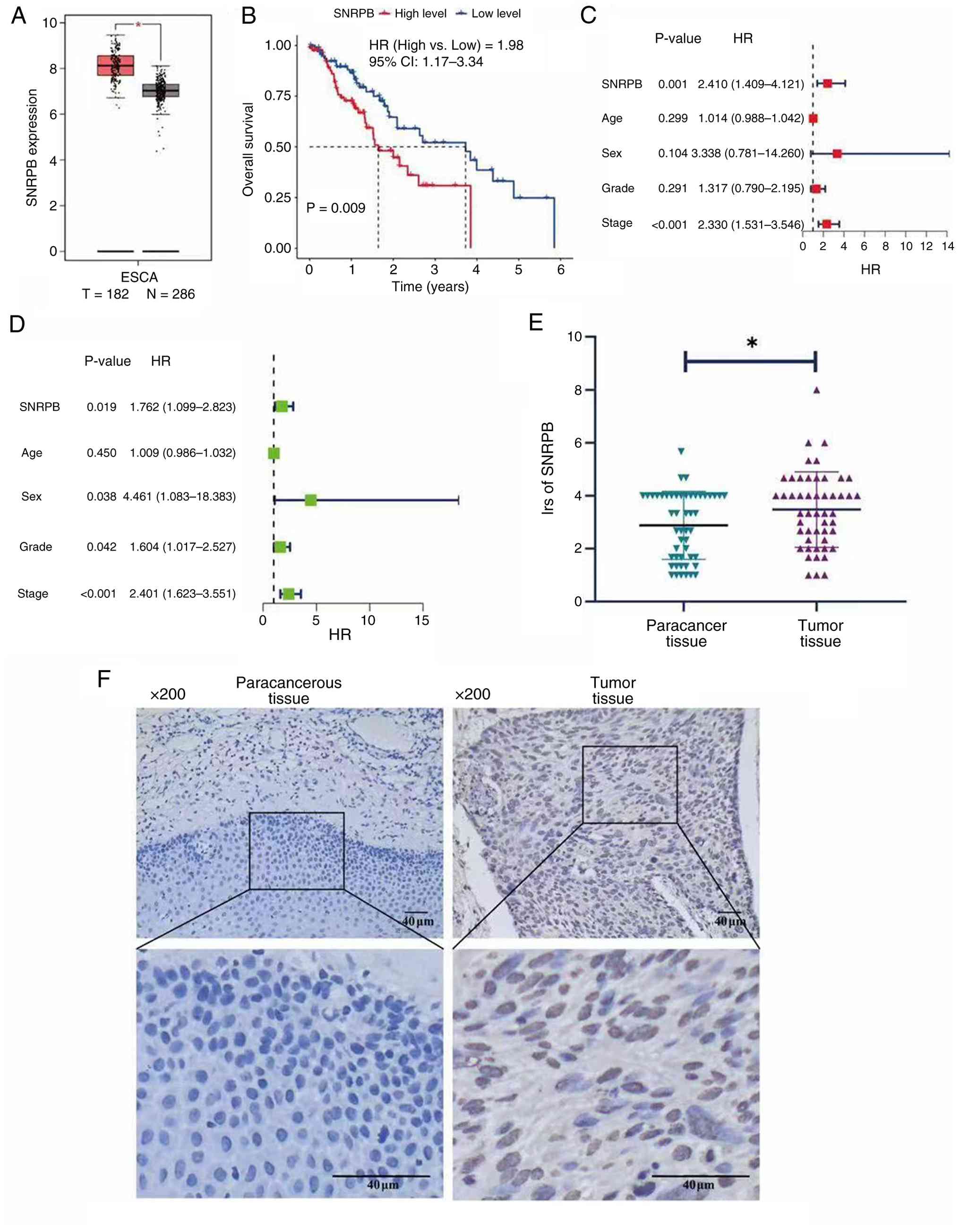
Online analysis using the GEPIA database confirmed
the overexpression of SNRPB in esophageal cancer tissues compared
with normal tissues (Fig. 1A).
Survival analysis revealed a significant correlation between high
SNRPB expression and reduced overall survival [hazard ratio (HR),
1.98; 95% confidence interval (CI), 1.17–3.34; P=0.009], as shown
in Fig. 1B. Both univariate and
multivariate analyses further demonstrated the prognostic value of
SNRPB expression levels, along with tumor stage, in predicting
patient outcomes (P<0.05 for both; Fig. 1C and D).
Immunohistochemical analysis further corroborated
these findings, revealing elevated SNRPB protein expression levels
in ESCC tissues (Fig. 1E). The IRS
for SNRPB was significantly greater in tumor tissues when compared
with that in adjacent non-cancerous tissues (P=0.012; Fig. 1F; paired Student's t test). Thus,
results of the present study suggested that SNRPB expression may be
associated with the clinicopathological features of patients with
ESCC. A retrospective analysis of clinical data obtained from a
cohort of 50 patients diagnosed with ESCC was conducted. The cohort
comprised 38 male (76%) and 12 female (24%) patients, with a median
age of 60 years (range, 45–77 years). Participants were divided
into two groups, ‘low’ and ‘high’ expression, according to the
median IRS for SNRPB obtained through immunohistochemical analysis.
The full clinicopathological characteristics are detailed in
Table I.
 | Table I.Correlations between SNRPB expression
and the clinicopathological characteristics of patients with
ESCC. |
Table I.
Correlations between SNRPB expression
and the clinicopathological characteristics of patients with
ESCC.
|
Characteristics | No. of
patients | Low SNRPB
expression | High SNRPB
expression | Test
method/statistic | P-value |
|---|
| Age |
|
|
|
χ2=0.045 | 0.832 |
|
<60 | 18 | 9 | 9 |
|
|
|
≥60 | 32 | 17 | 15 |
|
|
| Sex |
|
|
|
χ2=1.937 | 0.164 |
|
Male | 32 | 19 | 13 |
|
|
|
Female | 18 | 7 | 11 |
|
|
| History of
smoking |
|
|
|
χ2=0.855 | 0.355 |
|
Yes | 20 | 12 | 8 |
|
|
| No | 30 | 14 | 16 |
|
|
| History of alcohol
consumption |
|
|
|
χ2=0.142 | 0.706 |
|
Yes | 18 | 10 | 8 |
|
|
| No | 32 | 16 | 16 |
|
|
| Histological
grade |
|
|
| F-H | 0.987 |
| G1 | 17 | 9 | 8 |
|
|
| G2 | 31 | 16 | 15 |
|
|
| G3 | 2 | 1 | 1 |
|
|
| Location |
|
|
| F-H | 0.742 |
|
Upper | 7 | 3 | 4 |
|
|
|
Middle | 35 | 18 | 17 |
|
|
|
Lower | 8 | 5 | 3 |
|
|
| TNM stage |
|
|
|
χ2=2.039 | 0.153 |
| IA + IB
+ IIA + IIB | 24 | 15 | 9 |
|
|
| IIIA +
IIIB + IVA + IVB | 26 | 11 | 15 |
|
|
| T stage |
|
|
|
χ2=5.059 | 0.025a |
| T1 +
T2 | 27 | 18 | 9 |
|
|
| T3 +
T4 | 23 | 8 | 15 |
|
|
| N stage |
|
|
|
χ2=0.349 | 0.555 |
| N0 | 23 | 13 | 10 |
|
|
| N1 + N2
+ N3 | 27 | 13 | 14 |
|
|
| M stage |
|
|
| Fisher's exact | 0.225 |
| M0 | 48 | 26 | 22 |
|
|
| M1 | 2 | 0 | 2 |
|
|
Specifically, individuals with an IRS ≤3.33 were
placed in the low-expression category, and those individuals with
an IRS >3.33 were placed in the high-expression category. The
associations between SNRPB expression levels and a range of
clinical characteristics were evaluated using a χ2 test.
This analysis revealed a statistically significant correlation
between high SNRPB protein expression and advanced T stage in
patients with ESCC (P=0.025; Table
I).
Knockdown of SNRPB inhibits cell
proliferation in ESCC
To further investigate the impact of SNRPB on the
biological behaviors of ESCC, previous research (13–15)
has established SNRPB as a significant prognostic gene in multiple
human malignancies. The expression levels of SNRPB were assessed in
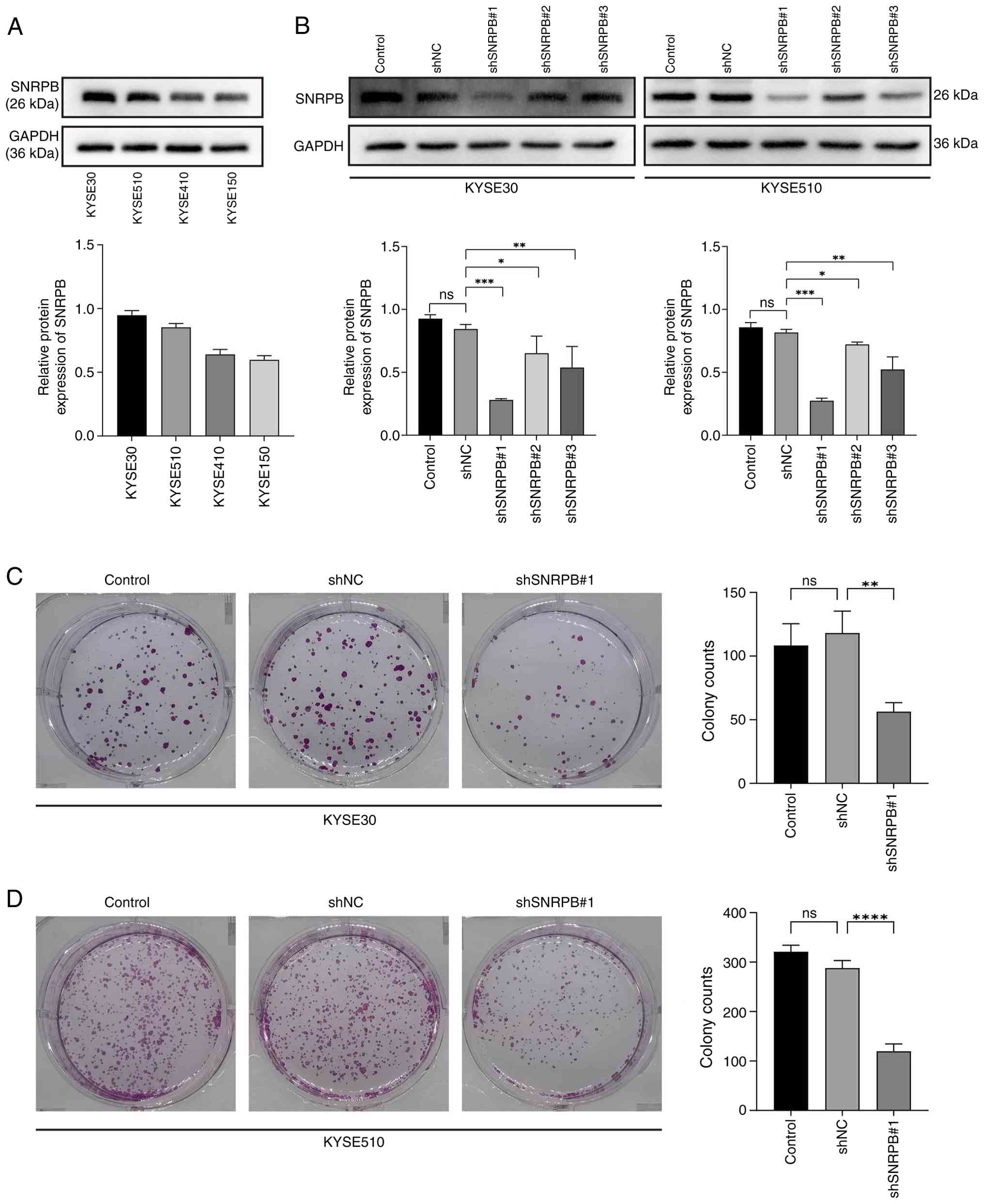
four ESCC cell lines, KYSE150, KYSE510, KYSE30 and KYSE410. Results
of the western blot analysis revealed notably high SNRPB expression
levels in KYSE30 and KYSE510 cell lines compared with KYSE150 and
KYSE410 cells (Fig. 2A).
Consequently, these two cell lines were selected for SNRPB
knockdown. In total, three distinct shRNAs targeting SNRPB,
shSNRPB#1, shSNRPB#2 and shSNRPB#3, were transfected into KYSE30
and KYSE510 cells. Moreover, western blot analysis was performed to
determine knockdown efficiency, and the results demonstrated that
shSNRPB#1 exhibited the highest level of knockdown efficiency
compared with shSNRPB#2, shSNRPB#3 and shNC (Fig. 2B). Thus, shSNRPB1#1 shRNA was
selected for use in subsequent experiments. As previous findings
revealed that SNRPB expression was associated with tumor T stage
(13,15,24,25), a
colony formation assay was performed to assess the impact of SNRPB
knockdown on the proliferation of cells. Based on three independent
experiments, the results indicated that SNRPB knockdown markedly
reduced colony formation capacity. Specifically, the number of
colonies was significantly lower in the knockdown group compared
with the shNC control group (Fig. 2C
and D).
SNRPB knockdown inhibits ESCC cell
cycle progression
To further elucidate the impact of SNRPB on the
growth of ESCC cells, cell cycle analysis was conducted using
KYSE30 and KYSE510 cell lines. Analysis suggested cell cycle arrest
as flow cytometry analysis demonstrated that SNRPB knockdown
increased the percentage of cells arrested in the G2/M phase from
12.3 to 28.7% in KYSE30 cells and from 10.5 to 25.2% in KYSE510
cells (Fig. 3A). To investigate the
molecular mechanism underlying this arrest, western blot analysis
was performed, which revealed that SNRPB knockdown led to
significant reductions in the expression levels of Cyclin A2 and
CDK1 in both KYSE30 and KYSE510 cell lines compared with the shNC
group (KYSE30-CDK1, P<0.01; KYSE30-CCNA2, P<0.01;
KYSE510-CDK1, P<0.01; KYSE510-CCNA2, P<0.01; one-way ANOVA
followed by Dunnett's post hoc test; Fig. 3B and C).
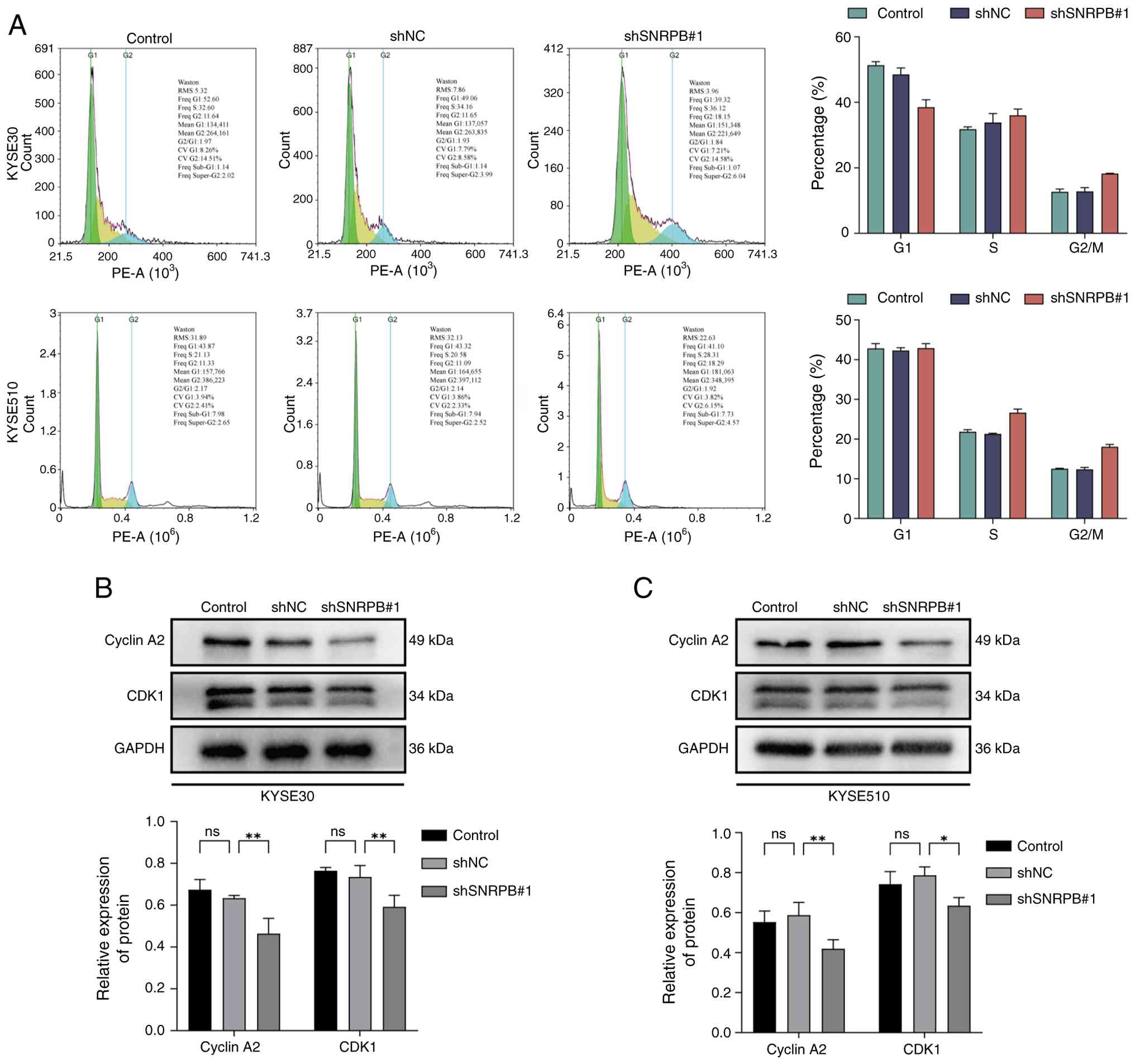 | Figure 3.Silencing SNRPB arrests the cell
cycle of esophageal squamous cell carcinoma cells. (A) Cell cycle
distribution of the KYSE30 and KYSE510 cell lines after
transduction with lentiviral shNC or SNRPB1#1 shRNA. Cell cycle
analysis. SNRPB-knockdown KYSE30 and KYSE510 cells (transfected
with shSNRPB#1) and negative control cells (transfected with shNC)
were used for this assay; the analysis was performed 48 h after
lentiviral transfection (the same time interval as subsequent
functional experiments). For sample preparation, the cells were
harvested by trypsinization with 0.25% trypsin-EDTA (Gibco; Thermo
Fisher Scientific, Inc.), washed twice with pre-chilled
phosphate-buffered saline (PBS), and fixed with 70% cold ethanol at
4°C for 24 h for cell cycle arrest. After fixation, the cells were
processed for cell cycle staining and detection according to the
instructions of the Cell Cycle Detection kit (Nanjing KeyGen
Biotech Co., Ltd.). Analysis was conducted via a NovoCyte flow
cytometer (NovoCyte 3130; ACEA Biosciences, Inc.), which allowed
precise cell cycle phase determination. The expression levels of
cyclin proteins in (B) KYSE30 cells and (C) KYSE510 cells. Data are
presented as the mean ± standard deviation. Statistical
significance was determined using Student's t-test. *P<0.05,
**P<0.01; one-way ANOVA followed by Dunnett's post hoc test.
shRNA, short hairpin RNA; ns, non-significant; NC, negative
control; SNRPB, small nuclear ribonucleoprotein polypeptide B. |
SNRPB knockdown promotes ESCC cell
apoptosis
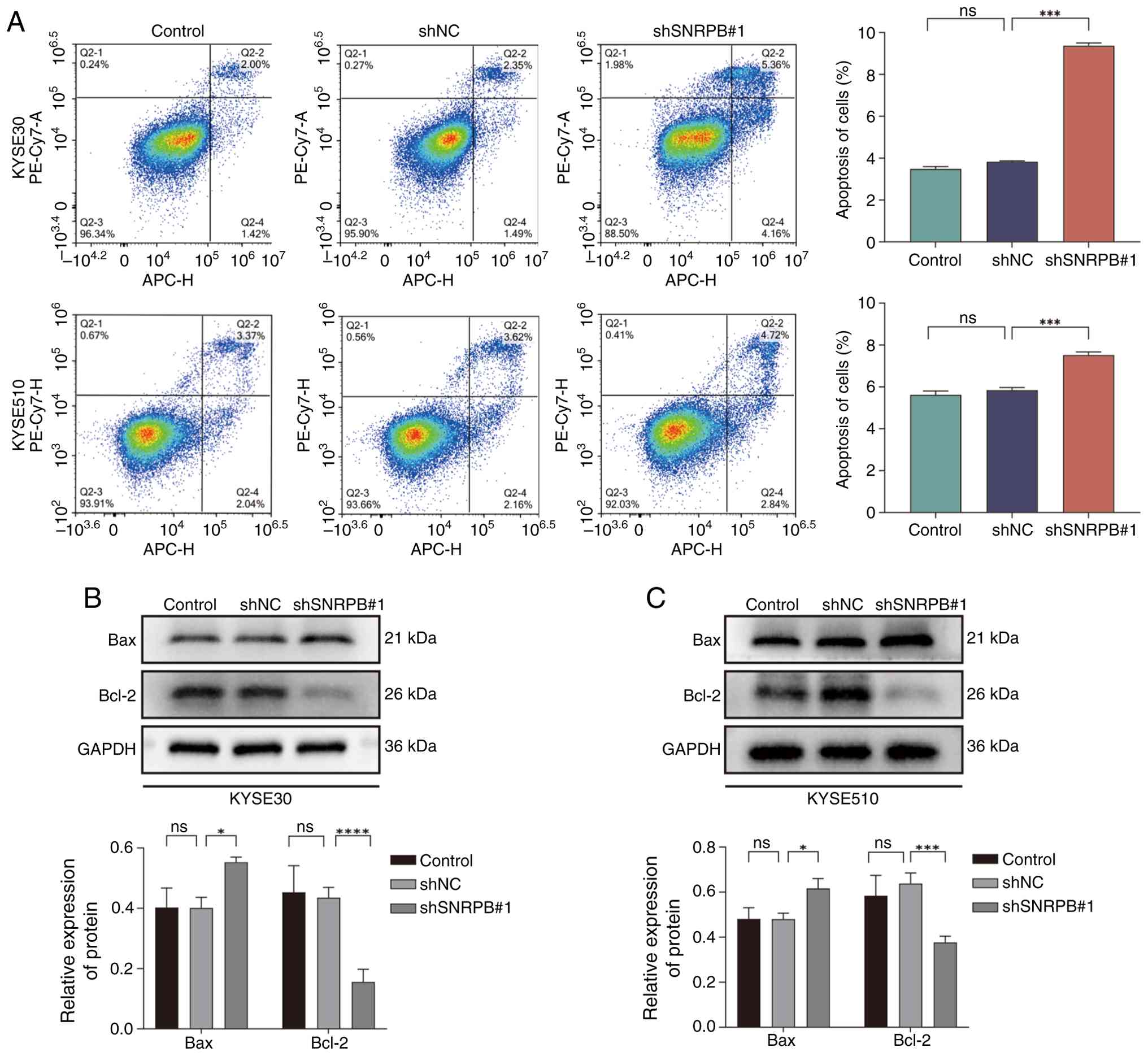
The potential effects of SNRPB knockdown on
apoptosis were determined in KYSE30 and KYSE510 cell lines. Results
of the present study revealed a significant increase in the
percentage of apoptotic cells in the SNRPB knockdown cells compared
with shNC in both cell lines (Fig.
4A). As shown in Fig. 4A, flow
cytometry analysis revealed that SNRPB knockdown led to a
significant increase in the percentage of apoptotic cells in both
KYSE30 and KYSE510 cell lines (KYSE30, P<0.001; KYSE510,
P<0.001; determined by one-way ANOVA followed by Dunnett's post
hoc test). To explore the molecular basis for this pro-apoptotic
effect, the expression levels of key apoptosis-regulating proteins,
Bax and Bcl-2, were determined by western blot analysis. Results of
the present study revealed that shSNRPB1#1 shRNA-mediated SNRPB
knockdown led to significant upregulation of Bax and significant
downregulation of Bcl-2 protein expression levels, compared with
those in the shNC group (KYSE30-Bax, P<0.001; KYSE30-Bcl-2,
P<0.001; KYSE510-Bax, P<0.001; KYSE510-Bcl-2, P<0.001;
one-way ANOVA followed by Dunnett's post hoc test; Fig. 4B and C).
Discussion
Results of previous studies have indicated that
SNRPB is overexpressed in a variety of tumor types and is
implicated in tumor progression (15,16,26).
However, its specific role in ESCC has remained largely unexplored.
To the best of our knowledge, the present study is the first to
demonstrate the oncogenic role of SNRPB in ESCC through an
integrated approach of bioinformatic analysis, clinical validation
and in vitro mechanistic investigations. The findings of the
present study revealed high expression of SNRPB in ESCC tissues and
revealed that its knockdown may inhibit cell proliferation and
promote apoptosis.
Dysregulated alternative pre-mRNA splicing
represents a hallmark molecular characteristic of the majority of
neoplasms (27), and markedly
contributes to the progression of ESCC and other malignancies
(24). SNRPB, a core component of
the spliceosome, carries out a key role in this process by
mediating alternative splicing and regulating gene expression
(18,28–30).
The oncogenic role of SNRPB is supported by previous studies that
reported high SNRPB expression levels in cervical (15), liver (19) and ovarian cancer (18), as well as its ability to promote
proliferation in various cellular contexts (13). Consistent with these reports, the
present study demonstrated that SNRPB knockdown reduced the
proliferation of ESCC cells. This anti-proliferative effect is
mechanistically underpinned by two key events. First, arrest in the
G2/M phase of the cell cycle was observed (31), which associated with the
downregulation of key regulatory proteins CDK1 and Cyclin A2
(32–34). Second, SNRPB knockdown promoted
apoptosis, which was associated with a disruption of the key
balance between pro- and anti-apoptotic proteins, as evidenced by
the upregulation of Bax (35) and
concomitant downregulation of Bcl-2 (25).
Mechanistically, while the present study revealed
that SNRPB depletion leads to these downstream molecular changes
(including CDK1/Cyclin A2 downregulation and the Bax/Bcl-2 shift),
the precise upstream link remains to be fully elucidated. As a core
component of the spliceosome, the primary function of SNRPB is to
regulate pre-mRNA splicing (10,18).
It is plausible that the observed phenotypes are a consequence of
altered alternative splicing of key regulatory genes. For example,
previous studies in cervical and thyroid cancer have associated
SNRPB to the inhibition of the tumor suppressor p53 (15,16).
Given that p53 is a master regulator of both the G2/M checkpoint
and the intrinsic apoptosis pathway (via Bax), it is a potential
candidate for being a downstream target of SNRPB-mediated splicing
in ESCC. Future research should aim to identify the specific
splicing events regulated by SNRPB to fully unravel its oncogenic
signaling network
However, the present study has several limitations
that must be acknowledged. First, and most importantly, the
findings of the present study are based entirely on in vitro
experiments. In vivo validation using animal models, such as
xenograft studies, is required to confirm the clinical relevance of
the results. Second, the present study focused on cell
proliferation and apoptosis; the potential role of SNRPB in other
cancer hallmarks, such as migration and invasion, was not
investigated. Third, a survival analysis was not performed on the
institutional cohort of 50 patients due to the limited sample size
and short follow-up period. Finally, a formal power analysis was
not conducted prior to the present study to determine the optimal
clinical sample size; the sample size was based on patient
availability. This should be considered when interpreting the
clinical correlation results. Future studies are therefore needed
to address these limitations and build upon the findings of the
present study.
The clinical implications of these findings may hold
relevance. First, the correlation between high SNRPB expression and
poor overall survival, as demonstrated in the TCGA cohort,
substantiates its potential as a prognostic biomarker. Patients
with SNRPB-high tumors could potentially be stratified for more
aggressive treatment regimens or closer monitoring. Second, the
results highlight SNRPB as a potential therapeutic target.
Targeting the spliceosome machinery is an emerging anti-cancer
strategy (26–28), and several small-molecule inhibitors
of splicing are under clinical investigation. These findings
suggest that patients with ESCC that have tumors overexpressing
SNRPB might be particularly vulnerable to such therapies.
Furthermore, given that altered splicing is frequently associated
with treatment resistance (13,17,29,36),
future studies should explore whether SNRPB contributes to
chemoresistance in ESCC, potentially opening avenues for
combination therapies.
Collectively, the molecular analyses of the present
study demonstrated that SNRPB knockdown impacts key proteins
involved in cell cycle progression and apoptosis, such as CDK1,
Cyclin A2, Bax and Bcl-2. These findings suggested that SNRPB may
play a key role in ESCC progression and may exhibit potential as a
biomarker for clinical applications in ESCC.
Supplementary Material
Supporting Data
Acknowledgements
Not applicable.
Funding
This research was funded by the Nanchong Science and Technology
Program (grant no. 23JCYJPT0060) and the Key Project of Research
and Development Program of the Affiliated Hospital of North Sichuan
Medical College (grant no. 2023ZD009).
Availability of data and materials
The data generated in the present study may be
requested from the corresponding author.
Authors' contributions
XFW contributed to conception and design of the
study, project administration, funding acquisition, analysis and
interpretation of clinical and experimental data, writing-reviewing
and editing of the manuscript. HYG contributed to acquisition of
experimental data, data curation, formal analysis, writing-original
draft of the manuscript. JZ, GBH and YT contributed to acquisition
of experimental data, visualization, investigation. LYL and ZCL
contributed to conception of the study, resources, sample
collection. XRC and XBW contributed to validation, formal analysis
and interpretation of experimental data. All authors read and
approved the final manuscript. XFW and HYG confirm the authenticity
of all the raw data.
Ethics approval and consent to
participate
The present study was approved by the Medical Ethics
Committee of the Affiliated Hospital of North Sichuan Medical
College, Sichuan, China (approval no. 2023ER344-1). Due to the
retrospective nature of the present study and the anonymization of
patient data, the Medical Ethics Committee of the Affiliated
Hospital of North Sichuan Medical College, Sichuan, China waived
the need of obtaining informed consent.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Sung H, Ferlay J, Siegel RL, Laversanne M,
Soerjomataram I, Jemal A and Bray F: Global cancer statistics 2020:
GLOBOCAN estimates of incidence and mortality worldwide for 36
cancers in 185 countries. CA Cancer J Clin. 71:209–249.
2021.PubMed/NCBI
|
|
2
|
Siegel RL, Miller KD, Wagle NS and Jemal
A: Cancer statistics, 2023. CA Cancer J Clin. 73:17–48.
2023.PubMed/NCBI
|
|
3
|
Pennathur A, Gibson MK, Jobe BA and
Luketich JD: Oesophageal carcinoma. Lancet. 381:400–412. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Siegel RL, Miller KD and Jemal A: Cancer
statistics, 2019. CA Cancer J Clin. 69:7–34. 2019.PubMed/NCBI
|
|
5
|
Abnet CC, Arnold M and Wei WQ:
Epidemiology of esophageal squamous cell carcinoma.
Gastroenterology. 154:360–373. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Koshy M, Esiashvilli N, Landry JC, Thomas
CR Jr and Matthews RH: Multiple management modalities in esophageal
cancer: Epidemiology, presentation and progression, work-up, and
surgical approaches. Oncologist. 9:137–146. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Huang FL and Yu SJ: Esophageal cancer:
Risk factors, genetic association, and treatment. Asian J Surg.
41:210–215. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Nilsen TW and Graveley BR: Expansion of
the eukaryotic proteome by alternative splicing. Nature.
463:457–463. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Wu J, Lu F, Yu B, Wang W and Ye X: The
oncogenic role of SNRPB in human tumors: A pan-cancer analysis.
Front Mol Biosci. 9:9944402022. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Bacrot S, Doyard M, Huber C, Alibeu O,
Feldhahn N, Lehalle D, Lacombe D, Marlin S, Nitschke P, Petit F, et
al: Mutations in SNRPB, encoding components of the core splicing
machinery, cause cerebro-costo-mandibular syndrome. Hum Mutat.
36:187–190. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Lynch DC, Revil T, Schwartzentruber J,
Bhoj EJ, Innes AM, Lamont RE, Lemire EG, Chodirker BN, Taylor JP,
Zackai EH, et al: Disrupted auto-regulation of the spliceosomal
gene SNRPB causes cerebro-costo-mandibular syndrome. Nat Commun.
5:44832014. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Knill C, Henderson EJ, Johnson C, Wah VY,
Cheng K, Forster AJ and Itasaki N: Defects of the spliceosomal gene
SNRPB affect osteo- and chondro-differentiation. FEBS J.
291:272–291. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Liu N, Wu Z, Chen A, Wang Y, Cai D, Zheng
J, Liu Y and Zhang L: SNRPB promotes the tumorigenic potential of
NSCLC in part by regulating RAB26. Cell Death Dis. 10:6672019.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Wang X, Zhang H, Guo Z, Wang J, Lu C, Wang
J, Jin R and Mo Z: SNRPB promotes the progression of hepatocellular
carcinoma via regulating cell cycle, oxidative stress, and
ferroptosis. Aging (Albany NY). 16:348–366. 2024.PubMed/NCBI
|
|
15
|
Zhu L, Zhang X and Sun Z: SNRPB promotes
cervical cancer progression through repressing p53 expression.
Biomed Pharmacother. 125:1099482020. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Deng Y, Li X, Jiang W and Tang J: SNRPB
promotes cell cycle progression in thyroid carcinoma via inhibiting
p53. Open Med (Wars). 17:1623–1631. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Li Y, Chen Z, Xiao H, Liu Y, Zhao C, Yang
N, Yuan C, Yan S and Li P: Targeting the splicing factor SNRPB
inhibits endometrial cancer progression by retaining the POLD1
intron. Exp Mol Med. 57:420–435. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Li Y, Chen Z, Peng J, Yuan C, Yan S, Yang
N, Li P and Kong B: The splicing factor SNRPB promotes ovarian
cancer progression through regulating aberrant exon skipping of
POLA1 and BRCA2. Oncogene. 42:2386–2401. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Peng N, Li J, He J, Shi X, Huang H, Mo Y,
Ye H, Wu G, Wu F, Xiang B, et al: c-Myc-mediated SNRPB upregulation
functions as an oncogene in hepatocellular carcinoma. Cell Biol
Int. 44:1103–1111. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Tang Z, Li C, Kang B, Gao G, Li C and
Zhang Z: GEPIA: A web server for cancer and normal gene expression
profiling and interactive analyses. Nucleic Acids Res. 45:W98–W102.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Therneau T: A Package for Survival
Analysis in R. R Foundation for Statistical Computing. Vienna,
Austria: 2024
|
|
22
|
Dumitru CA, Bankfalvi A, Gu X, Zeidler R,
Brandau S and Lang S: AHNAK and inflammatory markers predict poor
survival in laryngeal carcinoma. PLoS One. 8:e564202013. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Bowker AH: A test for symmetry in
contingency tables. J Am Stat Assoc. 43:572–574. 1948. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Zhang Y, Qian J, Gu C and Yang Y:
Alternative splicing and cancer: A systematic review. Signal
Transduct Target Ther. 6:782021. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Maulik N, Engelman RM, Rousou JA, Flack JE
III, Deaton D and Das DK: Ischemic preconditioning reduces
apoptosis by upregulating anti-death gene Bcl-2. Circulation.
100:II369–II375. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Zhan YT, Li L, Zeng TT, Zhou NN, Guan XY
and Li Y: SNRPB-mediated RNA splicing drives tumor cell
proliferation and stemness in hepatocellular carcinoma. Aging
(Albany NY). 13:537–554. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Bradley RK and Anczuków O: RNA splicing
dysregulation and the hallmarks of cancer. Nat Rev Cancer.
23:135–155. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Gray TA, Smithwick MJ, Schaldach MA,
Martone DL, Graves JA, McCarrey JR and Nicholls RD: Concerted
regulation and molecular evolution of the duplicated SNRPB'/B and
SNRPN loci. Nucleic Acids Res. 27:4577–4584. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Zheng Y, Niu X, Xue W, Li L, Geng Q, Fan Z
and Zhao J: The role of alternative splicing factors hnRNP G and
Fox-2 in the progression and prognosis of esophageal cancer. Dis
Markers. 2022:30437372022. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Wu H, Zheng J, Deng J, Zhang L, Li N, Li
W, Li F, Lu J and Zhou Y: LincRNA-uc002yug.2 involves in
alternative splicing of RUNX1 and serves as a predictor for
esophageal cancer and prognosis. Oncogene. 34:4723–4734. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Evan GI and Vousden KH: Proliferation,
cell cycle and apoptosis in cancer. Nature. 411:342–348. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Lohberger B, Leithner A, Stuendl N,
Kaltenegger H, Kullich W and Steinecker-Frohnwieser B: Diacerein
retards cell growth of chondrosarcoma cells at the G2/M cell cycle
checkpoint via cyclin B1/CDK1 and CDK2 downregulation. BMC Cancer.
15:8912015. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Ma Q: MiR-219-5p suppresses cell
proliferation and cell cycle progression in esophageal squamous
cell carcinoma by targeting CCNA2. Cell Mol Biol Lett. 24:42019.
View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Fischer M, Quaas M, Steiner L and Engeland
K: The p53-p21-DREAM-CDE/CHR pathway regulates G2/M cell cycle
genes. Nucleic Acids Res. 44:164–174. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Schellenberg B, Wang P, Keeble JA,
Rodriguez-Enriquez R, Walker S, Owens TW, Foster F, Tanianis-Hughes
J, Brennan K, Streuli CH and Gilmore AP: Bax exists in a dynamic
equilibrium between the cytosol and mitochondria to control
apoptotic priming. Mol Cell. 49:959–971. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Wang E and Aifantis I: RNA splicing and
cancer. Trends Cancer. 6:631–644. 2020. View Article : Google Scholar : PubMed/NCBI
|