Introduction
Retinoblastoma (RB) stands as the most prevalent
intraocular malignant tumor in children, with an incidence ranging
from 1/20,000 to 1/15,000, predominantly affecting children aged
<3 years (1,2). Globally, it accounts for 2–4% of
childhood malignant tumors, where there are 8,000-9,000 new cases
of RB each year (3,4). RB can be typically classified into
hereditary (35–45%) and non-hereditary (55–65%) types (5). The former frequently develops into
bilateral or unilateral multifocal RB, characterized by mutations
in the RB1 gene, whilst the latter mainly consists of
unilateral or single lesions, with leukocoria and strabismus
(6). Ultrasonography is the
preferred imaging method for diagnosing RB (7). Early detection and diagnosis are
crucial for the management of RB in children. With the widespread
implementation of genetic diagnosis for RB, novel avenues for
treatment have been provided, offering potential for future
clinical applications.
Ferroptosis is a type of regulated cell death that
was discovered by Dixon et al (8) in 2012. It is characterized by
intracellular iron overload, dependent on reactive oxygen species
production and intracellular lipid peroxidation accumulation. Under
electron microscopy, various changes, such as cell membrane
rupture, decreased mitochondrial cristae and mitochondrial outer
membrane wrinkling and rupture, can be observed (9). At present, pharmacological induction
or inhibition of ferroptosis offers therapeutic potential for
drug-resistant cancers, ischaemic organ injury and degenerative
diseases linked to extensive lipid peroxidation (10). For RB, it has been reported that
TP53 gene mutations and RB1 gene deletion are
important causes of RB, both of which participate in ferroptosis
(11). Therefore, ferroptosis may
be involved in the pathogenesis of RB and represents a potential
mechanism worthy of further investigation. Non-coding RNAs (ncRNAs)
are becoming increasingly recognized for their role in regulating
the expression of ferroptosis-related genes and influencing disease
prognosis (12). They can be
categorized into basic structural ncRNAs and regulatory ncRNAs
based on their function (13).
Amongst the ncRNAs implicated in ferroptosis regulation, regulatory
ncRNAs, particularly microRNAs (miRNAs or miR), circular RNAs and
long non-coding (lnc)RNAs, are the most prominent (14). Beyond directly influencing enzymes
involved in iron and lipid metabolism, these regulatory ncRNAs
participate in a wider regulatory network governing ferroptosis.
Their dysregulation in RB also highlights their potential as novel
biomarkers, paving the way for future therapeutic applications
(15).
In the present study, a bioinformatics analysis of
publicly available microarray datasets from RB tumor samples and
control samples was performed. Differentially expressed genes
(DEGs) were intersected with a list of ferroptosis-related (FR)
genes (FRGs) to identify the FR-DEGs. Gene Ontology (GO) and Kyoto
Encyclopedia of Genes and Genomes (KEGG) functional analysis,
protein-protein interaction (PPI) network and immune cell
infiltration analyses were then performed to explore the potential
functions and interconnections among these genes. miRNAs for these
hub genes were then predicted, which were used further to predict
relevant lncRNAs. An FR lncRNA-miRNA-mRNA competing endogenous
(ce)RNA network was constructed, providing novel insights into the
pathology and treatment of RB.
Materials and methods
Data collection
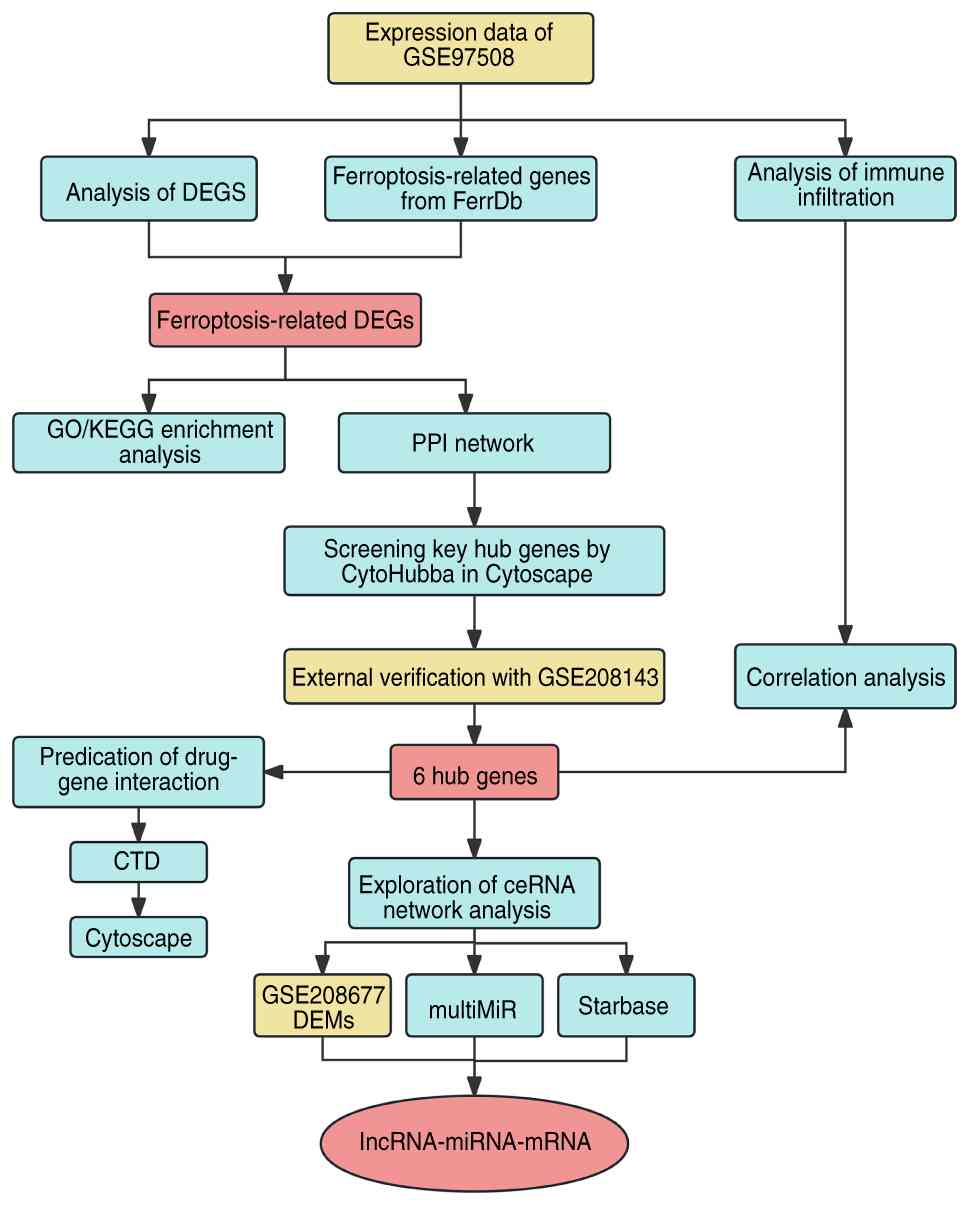
Fig. 1 illustrates
the flowchart of the present study design. The microarray
expression profiles for RB were obtained through the Gene
Expression Omnibus (GEO) database (https://www.ncbi.nlm.nih.gov/geo/) (16). GSE97508 represented mRNA expression
profiles in GPL15207, including 6 RB tumor samples and 3 control
samples. The validation dataset GSE208143 is based on GPL17077 and
includes 27 RB tumor samples and 6 control samples. It was used for
hub gene validation. GSE208677 represented miRNA expression
profiles in GPL18358, which contains 25 RB tumors and 5 control
pediatric retina samples. The FerrDb database (http://www.zhounan.org/ferrdb/) includes gene
regulatory factors such as driver, suppressor, marker and
unclassified regulatory factors.
Differential expression analyses
The raw data were background-corrected and
normalized using the R package ‘limma’ (17). In the R software environment, the
‘limma’ package was utilized to identify DEGs in GSE97508, with a
significance threshold set at adjusted P<0.05 and
|log2fold change|>1. As for the differentially
expressed miRNAs (DEMs), the criteria were adjusted P<0.05 and
|log2fold change|>2. In addition, a Venn diagram was
used to identify the overlapping sections of the DEGs and FRGs,
revealing the critical FR-DEGs.
Enrichment analysis of FR-DEGs
The FR-DEGs were submitted to GO function enrichment
analysis and the KEGG pathway enrichment analysis using the R
package ‘clusterProfiler’ (18). A
false discovery rate <0.05 was considered statistically
significant. The R package ‘ggplot2’ was adopted to visualize the
results (19). Furthermore, the
FR-DEGs were submitted to Metascape (http://metascape.org) for supplementary KEGG
analysis.
Screening and validation of hub
genes
The Search Tool for the Recovery of Interacting
Genes and proteins (STRING) online database serves as a valuable
platform for the exploration and analysis of gene or protein
interactions (https://string-db.org). A PPI network
was constructed by selecting gene pairs with confidence scores
>0.4. To determine the hub genes, the CytoHubba (version 0.1)
plug-in of Cytoscape (version 3.7.1) software was employed
(20). In total, 5 algorithms
[BottleNeck, Degree, Edge Percolated Component (EPC), Maximal
Clique Centrality (MCC) and Maximum Neighborhood Component (MNC)]
were applied to calculate scores for the FR-DEGs and generated a
Venn diagram to identify hub genes.
Furthermore, a validation was conducted using
GSE208143. A bar chart illustrating the expression levels of hub
genes in RB and normal tissues was generated using the R package
‘ggpubr’. In the ROC curve analysis, the area under the curve (AUC)
for each hub gene was calculated using the R package ‘pROC’ to
evaluate their standalone diagnostic performance. Statistical
significance was assessed using an unpaired t-test with a
significance threshold of P<0.05.
Immune infiltration analysis
The R package ‘IOBR’ was utilized to employ the
CIBERSORT algorithm for detecting the expression of 22 different
immune cell populations in the GSE97508 dataset (21). Box plots were then generated to
compare the expression of various immune cell types between the RB
group and the normal group. The association between hub genes and
the immune cell populations was assessed using Spearman correlation
analysis, where the results were visually presented using the R
package ‘ggplot’.
Prediction of miRNAs and construction
of FR miRNA-mRNA regulatory network
The R package ‘multiMiR’ was utilized to predict the
miRNAs for the hub genes (22).
Subsequently, the predicted miRNA was filtered by overlapping them
with the identified DEMs in the GSE208677 dataset. To further
refine the selection, only miRNAs showing a significant negative
correlation with the expression of their target hub genes in the
GSE97508 dataset were retained for network construction. In
addition, an FR miRNA-mRNA regulatory network was constructed and
visualized using Cytoscape.
Prediction of lncRNA and construction
of ferroptosis-related lncRNA-miRNA-mRNA ceRNA network
The Starbase online tool (https://starbase.sysu.edu.cn/) was utilized to input
the three miRNAs negatively regulating hub genes and predict
potential upstream lncRNAs involved in their interactions. To
refine the focus, lncRNAs that overlapped with all three identified
miRNAs were focused upon.
Predication of drug-gene
interaction
To explore the interaction between drugs and genes,
the Comparative Toxicogenomics Database (CTD; http://ctdbase.org/) was utilized to identify existing
or potential relevant drugs. The selected hub genes with potential
therapeutic relevance were queried in the CTD to identify existing
or potential drug-gene interactions, which were visualized using
the Cytoscape software.
Cell lines and culture
The human retinal pigment epithelium cell line
ARPE-19 and the human RB cell line Y79 were sourced from the
American Type Culture Collection. ARPE-19 cells and Y79 cells were
cultured at 37°C with 5% CO2 in Dulbecco's Modified
Eagle's Medium (cat. no. 12491015; Gibco; Thermo Fisher Scientific,
Inc.) supplemented with 10% fetal bovine serum (cat. no. A5256701;
Gibco; Thermo Fisher Scientific, Inc.), 100 µg/ml penicillin and
100 µg/ml streptomycin (cat. no. C0222; Beyotime Institute of
Biotechnology).
Reverse transcription-quantitative PCR
(RT-qPCR)
Total RNA was extracted from ARPE-19 and Y79 cells
using the RNA isolater Total RNA Extraction Reagent (cat. no.
R401-01; Vazyme Biotech Co., Ltd.). cDNA was synthesized with
PrimeScript™ RT Master Mix (cat. no. RR036Q; TaKaRa Bio, Inc.)
according to the manufacturer's instructions. qPCR was conducted
using TB Green® Premix Ex Taq™ II (cat. no. RR820A;
TaKaRa Bio, Inc.) according to the manufacturer's instructions. The
qPCR thermocycling conditions were as follows: Initial denaturation
at 95°C for 30 sec, followed by 40 cycles of 95°C for 5 sec and
60°C for 30 sec. A melting curve analysis was conducted from 65 to
95°C to verify amplification specificity. Data were normalized to
GAPDH expression using the 2−ΔΔCq method (23), with each experiment performed in
triplicate. Primers used were as follows: IDH2 forward,
5′-CGCCACTATGCCGACAAAAG-3′ and reverse,
5′-ACTGCCAGATAATACGGGTCA-3′; CDKN2A forward,
5′-GATCCAGGTGGGTAGAAGGTC-3′ and reverse, 5′-CCCCTGCAAACTTCGTCCT-3′;
and GAPDH forward, 5′-ACAACTTTGGTATCGTGGAAGG-3′ and reverse,
5′-GCCATCACGCCACAGTTTC-3′.
Statistical analysis
Statistical analysis was performed using R software
(version 4.2.1). A two-tailed unpaired Student's t-test was used to
determine the statistical significance of differences between two
groups, where P<0.05 was considered to indicate a statistically
significant difference. The correlation between hub genes and
immune cells was assessed using the Spearman correlation
coefficient, where corrections were applied using the
Benjamini-Hochberg multiple testing correction method.
Results
Identification of ferroptosis-related
DEGs
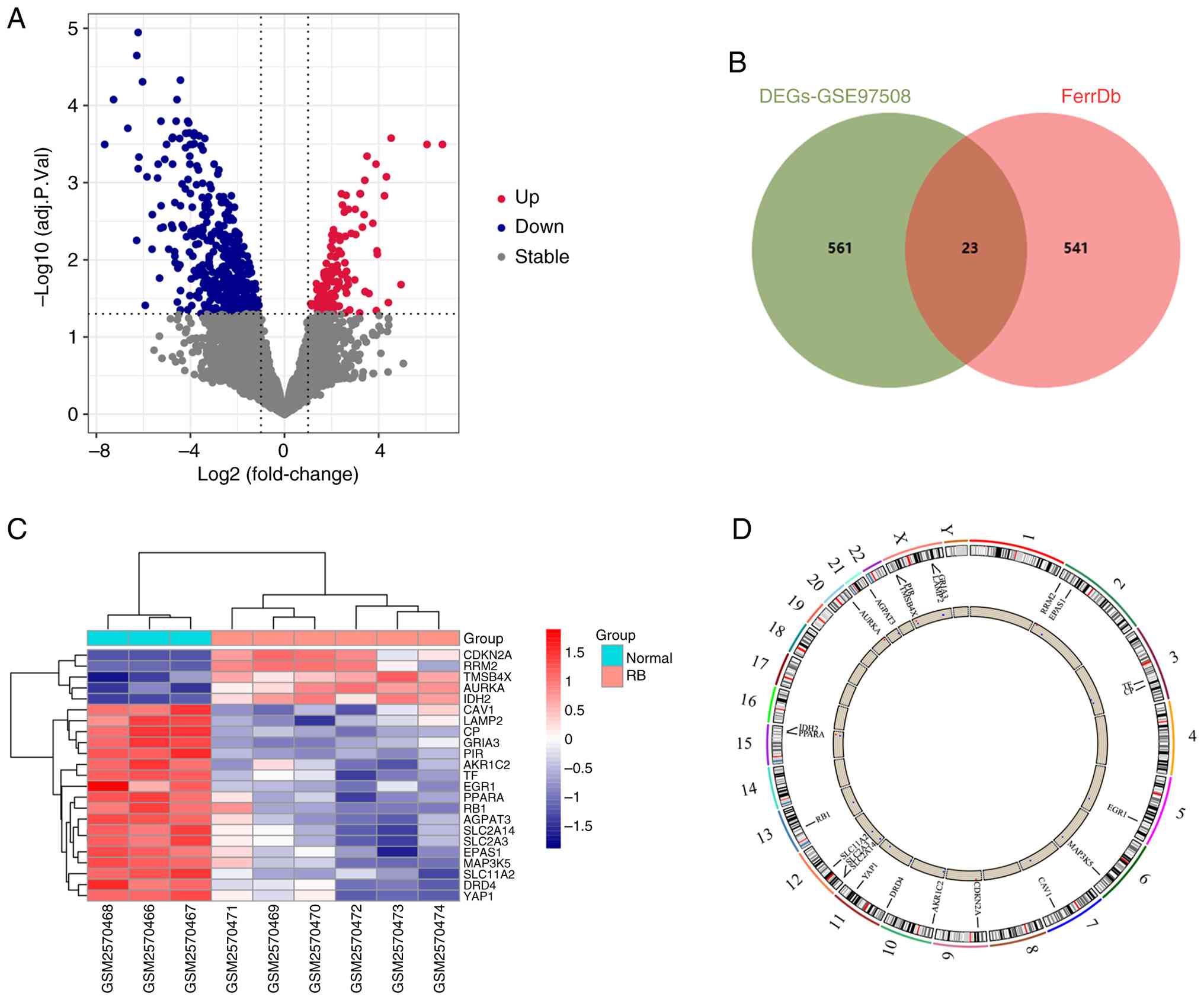
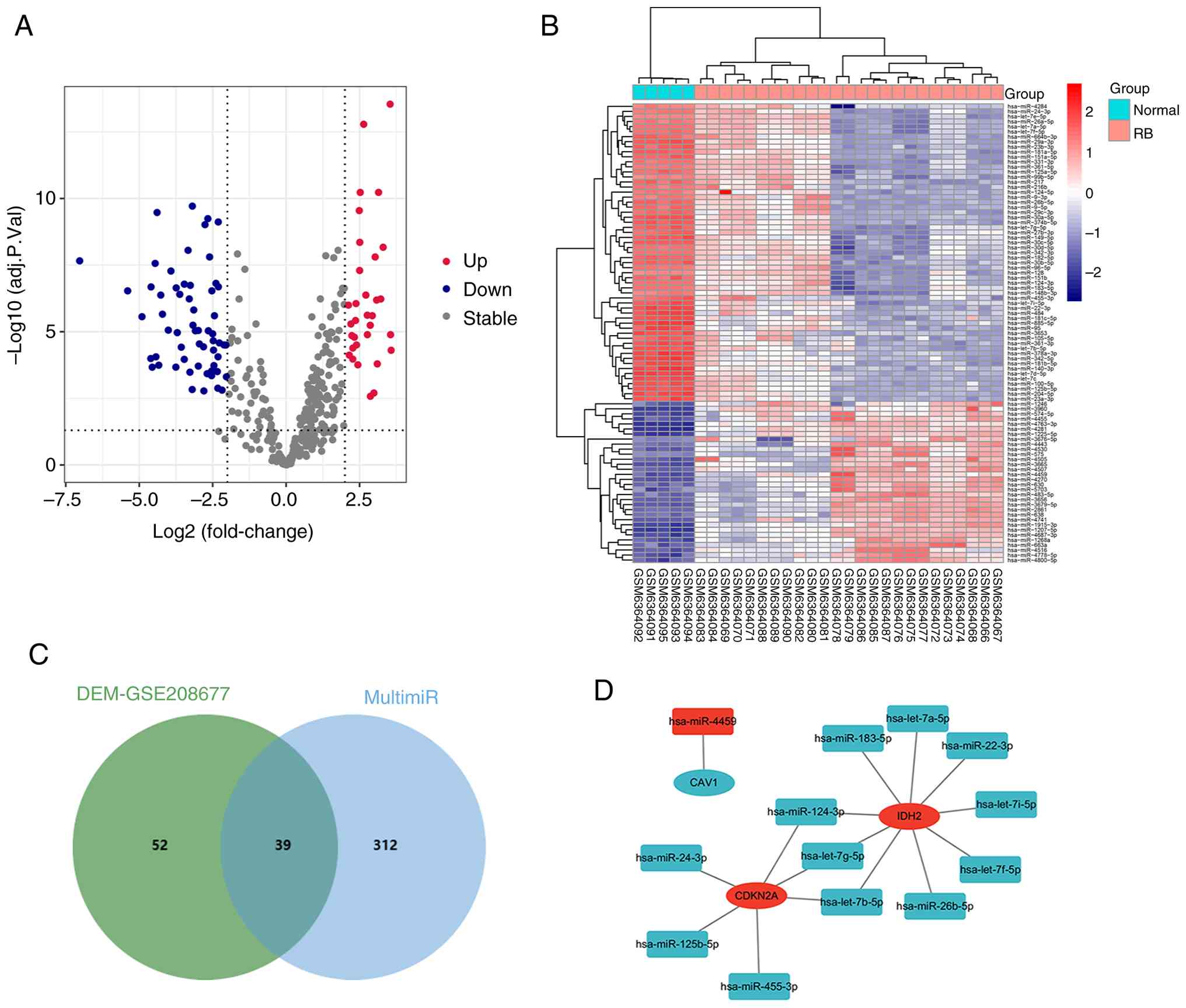
In total, 584 DEGs (135 upregulated and 449
downregulated genes) were obtained from the GSE97508 dataset using
the R package ‘limma’ (Fig. 2A).
Subsequently, an analysis was conducted by generating a Venn
diagram to identify the intersection between the FRGs extracted
from the ‘FerrDb’ database and the DEGs in GSE97508. The
overlapping set, consisting of 23 genes, represents the FR-DEGs
(Fig. 2B). The heatmap in Fig. 2C displays the standardized
expression of FR-DEGs (5 upregulated and 18 downregulated).
Fig. 2D displays the chromosomal
positions of these 23 FR-DEGs, represented in a circular plot for
visualization.
Function and pathway enrichment
analysis of FR-DEGs
The ‘clusterProfiler’ package was used to conduct GO
and KEGG functional analysis on the 23 FR-DEGs, aiming to gain a
comprehensive understanding of their cellular functions and pathway
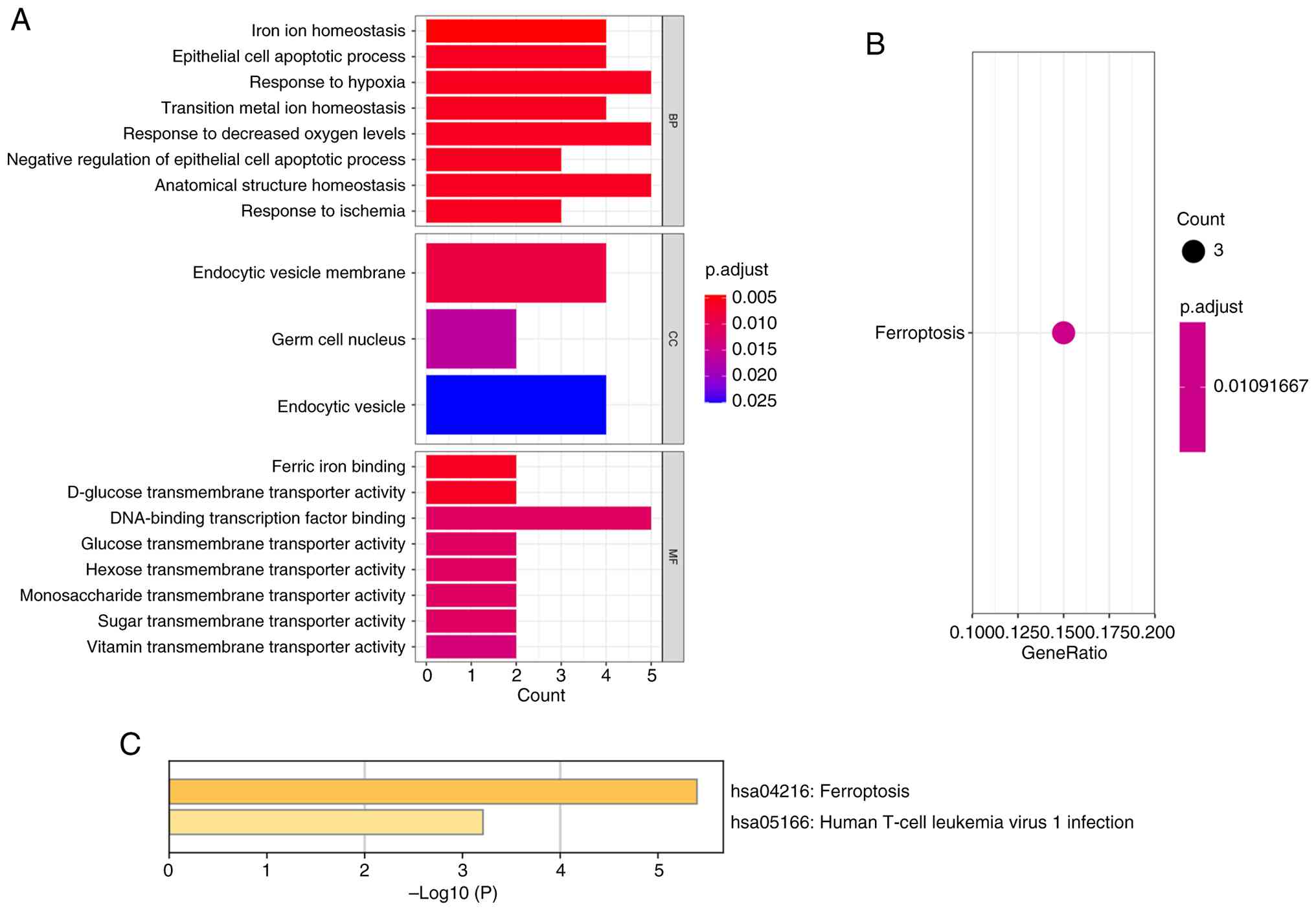
involvement in RB. The enrichment results revealed significant
involvement of FRGs in the regulation of ‘iron ion homeostasis’ and
‘transition metal ion homeostasis’, in the ‘response to hypoxia’
and ‘response to ischemia’, and in the modulation of ‘epithelial
cell apoptotic processes’ (Fig.
3A). Furthermore, the genes exhibit enrichment in specific
cellular component terms, emphasizing their roles in ‘endocytic
vesicles’ and ‘germ cell nucleus’. Molecularly, these genes
demonstrated diverse transmembrane transporter activities,
suggesting their influence on the transport of various ions and
substances across cellular membranes. In the KEGG pathway
enrichment analysis, a significant enrichment of genes related to
iron death in the ‘Ferroptosis’ pathway was observed (Fig. 3B and C).
Identification and verification of hub
genes
To investigate potential relationships among
proteins encoded by FR-DEGs and identify hub genes, STRING was
utilized to screen the PPI network of FR-DEGs (Fig. 4A). Subsequently, multiple
topological analysis algorithms (BottleNeck, EPC, Degree, MCC and
MNC) were utilized to predict and explore the top 10 significant
hub genes in the PPI network. The intersection of these five
algorithms led to the identification of CAV1, CDKN2A, EPAS1,
IDH2, RB1 and SLC2A3 as the hub genes (Fig. 4B). Amongst them, CAV1, EPAS1,
RB1 and SLC2A3 were significantly downregulated in RB
tumors, whilst CDKN2A and IDH2 were significantly
upregulated in GSE97508.
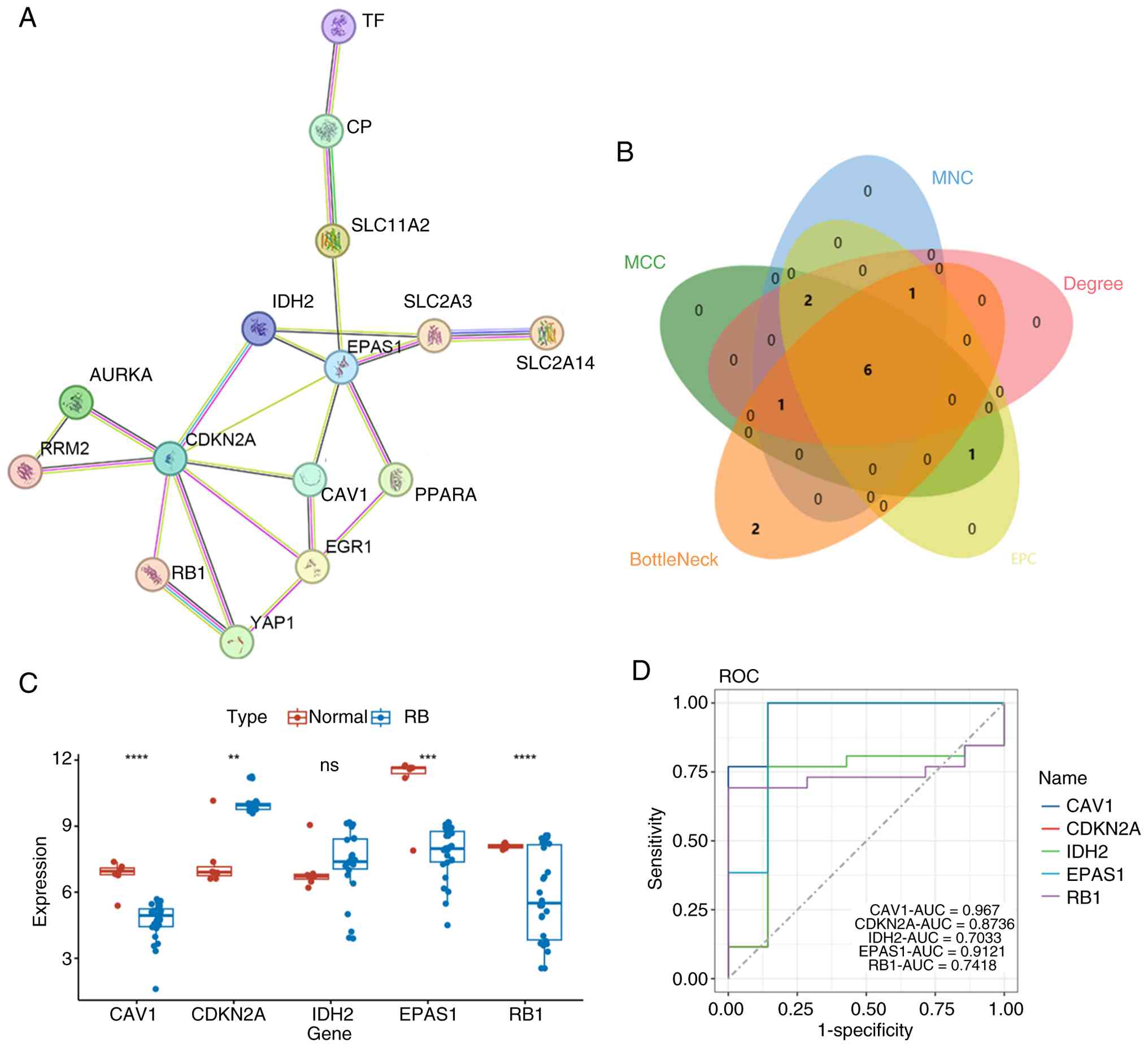 | Figure 4.Construction of PPI network and
identification of hub genes. (A) PPI network of ferroptosis-related
differentially expressed genes. Each node in the network represents
a protein. The small 3D structural models inside the circles are
schematic representations of the protein conformations. TF,
Transferrin; CP, Ceruloplasmin; SLC11A2, Solute Carrier Family 11
Member 2; SLC2A3, Solute Carrier Family 2 Member 3; SLC2A14, Solute
Carrier Family 2 Member 14; IDH2, Isocitrate Dehydrogenase 2;
EPAS1, Endothelial PAS Domain Protein 1; PPARA, Peroxisome
Proliferator-Activated Receptor Alpha. (B) Venn diagram
illustrating the hub genes identified by five algorithms
(BottleNeck, Degree, EPC, MCC and MNC) using Cytoscape software and
CytoHubba. (C) Validation of hub genes in the GSE208143 dataset.
The box represents the interquartile range, the line inside the box
denotes the median and the dots represent individual sample data
points. (D) ROC curve of hub genes in the GSE208143 dataset.
**P<0.01, ***P<0.001 and ****P<0.0001. ns, no
significance; PPI, protein-protein interaction; EPC, Edge
Percolated Component; MCC, Maximal Clique Centrality; MNC, Maximum
Neighborhood Component; ROC, receiver operating characteristic;
AUC, area under the ROC curve; RB, retinoblastoma. |
The expression of the 6 hub genes was next validated
using the GSE208143 dataset. However, SLC2A3 was not
represented in this dataset-a common limitation when reanalyzing
public genomic data due to the inherent differences in microarray
platform designs (GSE97508 used GPL15207, while GSE208143 used
GPL17077). SLC2A3 was not solely dependent on the validation
dataset for its significance. Its identification as a hub gene was
based on multiple topological algorithms (BottleNeck, EPC, Degree,
MCC and MNC) applied to the PPI network, and its differential
expression was initially observed in the discovery dataset
GSE97508. While it was not possible to validate its expression in
GSE208143, its biological relevance in ferroptosis and cancer
metabolism is supported by existing literature (24). Therefore, validation was possible
for the remaining 5 hub genes. Despite this limitation, the
successful validation of 5 out of 6 hub genes (an 83% validation
rate) provides robust support for the hub gene selection in the
present study. CDKN2A and IDH2 exhibited higher
expression levels in the RB group, whilst CAV1, EPAS1 and
RB1 showed significantly lower expression levels in patients
with RB (Fig. 4C). Subsequent ROC
curve analysis revealed an AUC value of 0.967 for CAV1,
0.8736 for CDKN2A, 0.9121 for EPAS1, 0.7033 for
IDH2 and 0.7418 for RB1 (Fig. 4D). These results indicate that CAV1
and EPAS1 possess excellent diagnostic accuracy, while CDKN2A shows
good diagnostic performance, and IDH2 and RB1 demonstrate moderate
diagnostic value. Collectively, these 5 genes represent promising
diagnostic biomarkers for RB.
Relationship between the immune cell
population and hub FR-DEGs
The CIBERSORT algorithm was used to calculate the
infiltration of 22 immune cell types in GSE97508 (6 RB and 3 normal
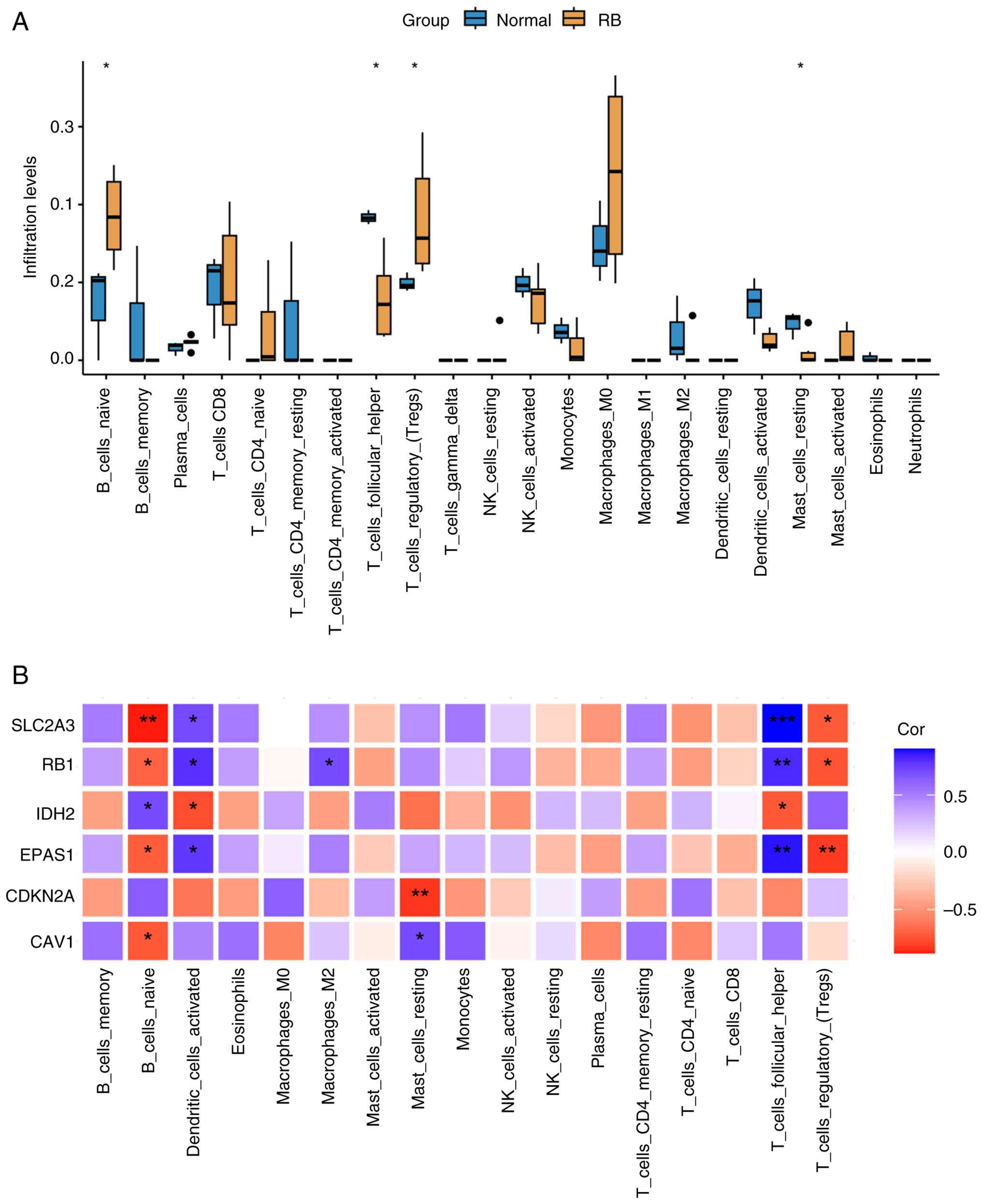
samples). The differences in the immune cell infiltration ratios
between the RB and normal groups are shown in Fig. 5A. Compared with those in the control
group, the infiltration levels of naive B cells and regulatory T
cells (Tregs) were higher, whilst the infiltration levels of T
follicular helper (Tfh) cells and resting mast cells were
significantly reduced.
Correlation analysis between hub genes and immune
cell populations revealed that EPAS1, RB1 and SLC2A3
exhibited negative correlations with naive B cells and Tregs,
whilst showing positive correlations with Tfh cells and activated
dendritic cells (Fig. 5B).
IDH2 demonstrated significant positive correlations with Tfh
cells and activated dendritic cells, whilst showing a significant
negative correlation with naive B cells. Resting mast cells
displayed a significant positive correlation with CDKN2A but
a significant negative correlation with CAV1. Additionally,
CAV1 exhibited a significant positive correlation with naive
B cells.
Construction of the miRNA-mRNA
network
From the GSE208677 dataset, DEMs (32 upregulated and
59 downregulated) were identified (Fig.
6A and B). For the six FR hub genes identified from the PPT
network analysis, namely CAV1, CDKN2A, EPAS1, IDH2, RB1 and SLC2A3,
multiMiR was utilized to predict 351 potential miRNAs.
Subsequently, an intersection analysis between these predicted
miRNAs and DEMs from the GSE208677 dataset was performed, resulting
in the identification of 39 FR miRNAs (Fig. 6C).
Given that miRNA expression levels are typically
negatively correlated with mRNA expression, to enhance the
reliability of the present findings, the predicted relationships
were then integrated with the corresponding expression data.
Subsequently, the miRNA-mRNA regulatory network was visualized
using Cytoscape software to provide a comprehensive and insightful
representation of the regulatory interactions (Fig. 6D).
Construction of a predicted FR
lncRNA-miRNA-mRNA ceRNA network
Among the 39 ferroptosis-related miRNAs identified
in the miRNA-mRNA network, those that were predicted to target both
CDKN2A and IDH2 and that showed negatively correlated expression
patterns with these mRNAs were further selected. A total of 3
miRNAs met these criteria: hsa-let-7b-5p, hsa-let-7g-5p and
hsa-miR-124-3p. These were selected as the core miRNAs for
subsequent ceRNA network construction. To explore potential lncRNAs
in RB, the Starbase tool was used to computationally predict the
target lncRNAs of these miRNAs. Applying a stringent filtering
criterion, only 4 lncRNAs predicted to have binding sites for all
miRNAs were retained for further analysis. These lncRNAs were long
intergenic non-coding RNA (LINC)00963, nuclear enriched abundant
transcript 1 (NEAT1), small nucleolar RNA host gene 16 (SNHG16) and
X-inactive specific transcript (XIST).
Based on these computational predictions, a putative
lncRNA-miRNA-mRNA ceRNA network for RB was successfully
established. This predicted network includes 2 mRNAs (CDKN2A and
IDH2), 3 miRNAs (hsa-let-7b-5p, hsa-let-7g-5p and hsa-miR-124-3p)
and 4 lncRNAs (LINC00963, NEAT1, SNHG16 and XIST). It is important
to note that these interactions are based solely on bioinformatic
predictions and have not been experimentally validated. This ceRNA
network may be closely associated with the progression of RB, where
some of these molecules may serve as key therapeutic targets for RB
(Fig. 7A), warranting further
experimental investigation.
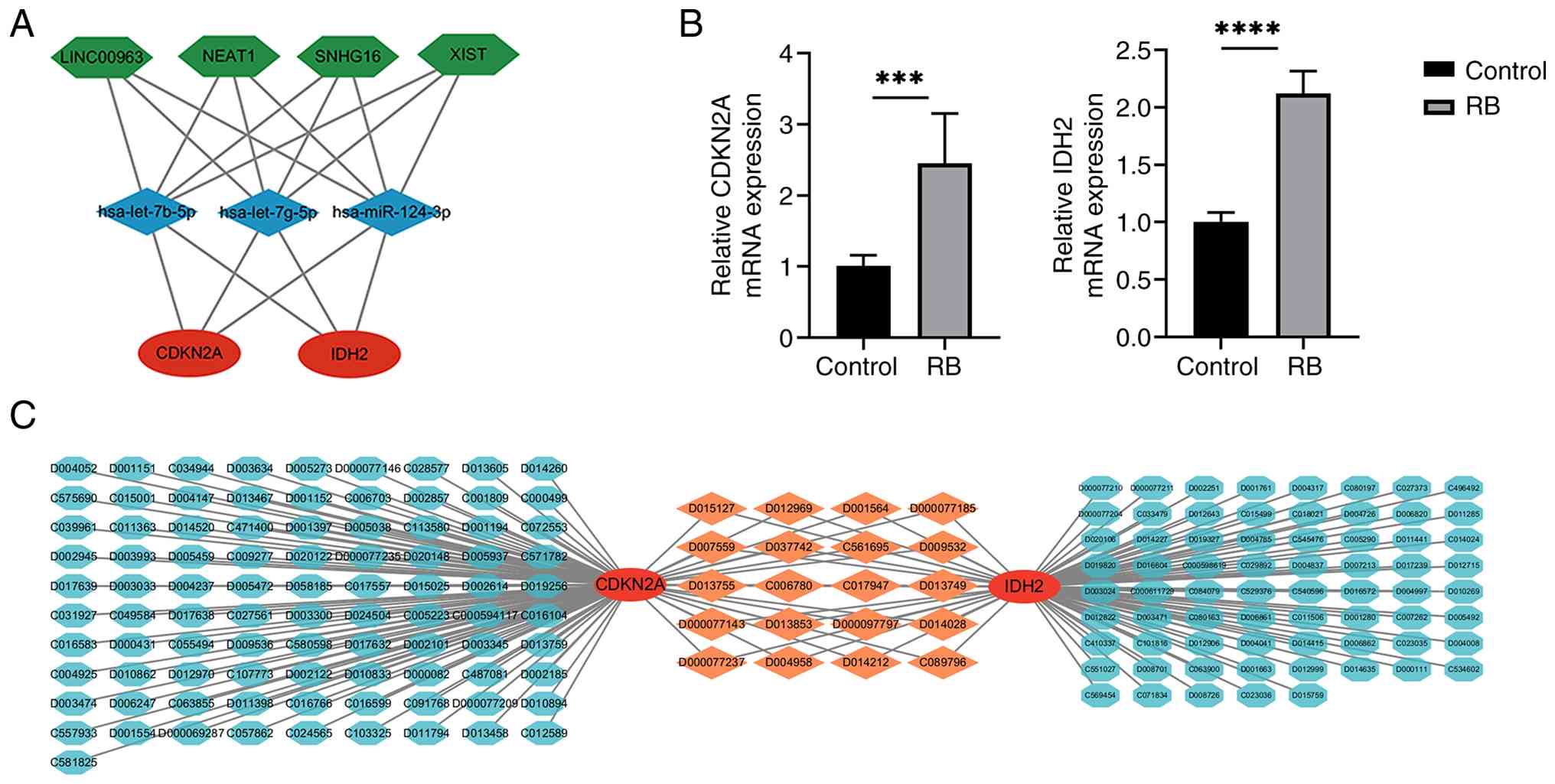 | Figure 7.ceRNA networks, drug-gene
interactions and PCR validation. (A) The ceRNA-regulating networks
illustrate protein-coding genes (red circles), miRNAs (blue
diamonds) and lncRNAs (green hexagons), with black lines indicating
interactions among lncRNA, miRNA and mRNA, where ceRNA refers to
ceRNAs. (B) Validation through reverse transcription-quantitative
PCR analysis confirms that CDKN2A and IDH2 expression
levels are significantly higher in Y79 cells compared to ARPE-19
cells. Data were normalized to GAPDH expression using the
2−ΔΔCq method, with each experiment performed in
triplicate. ***P<0.001 and ****P<0.0001. (C) The drug-gene
interaction analysis shows the linkage map of two hub genes,
CDKN2A and IDH2, along with their potential target
drugs, including bisphenol A (code: C006780), sodium arsenite
(code: C017947), docetaxel (code: D000077143) and
hexabromocyclododecane (code: C089796). ceRNA, competing endogenous
RNAs; miRNA, microRNA; lncRNA, long non-coding RNA; RB,
retinoblastoma. |
Drug-gene interactions and expression
levels of CDKN2A and IDH2 in RB
RT-qPCR analysis confirmed that CDKN2A and IDH2 mRNA
expression levels were significantly higher in Y79 retinoblastoma
cells than in ARPE-19 cells (Fig.
7B). Based on these findings, their potential clinical
implications were further explored by constructing a drug-gene
interaction network using the CTD database. As shown in Fig. 7C, this analysis identified 111
potential therapeutic agents targeting CDKN2A and 89 targeting
IDH2. Notably, several drugs, including bisphenol A (code:
C006780), sodium arsenite (code: C017947), docetaxel (code:
D000077143) and hexabromocyclododecane (code: C089796), were found
to target both genes, suggesting they may serve as candidate
compounds for further investigation in RB therapy.
Discussion
RB treatment requires early detection, diagnosis and
treatment. However, severe outcomes, such as blindness, proptosis
and intracranial metastasis, can occur (25). The present study delved into the
intricate molecular landscape of RB, focusing on FR ceRNA networks.
Ferroptosis is a form of regulated cell death that is driven by
iron-dependent lipid peroxidation. It has gained prominence in
cancer research (26). In the
context of RB, a set of FRGs that exhibited differential expression
was identified, forming a unique FR-DEGs signature.
Malignant tumor characteristics encompass persistent
proliferative signals, immune evasion and resistance to programmed
cell death (PCD) (27). Among
these, PCD resistance is a significant feature. Ferroptosis has
been associated with the development of various tumors and drug
resistance (28,29). Coordinated control over iron levels,
lipid oxidation and antioxidants in tumors can prevent ferroptosis,
fostering uncontrolled proliferation and tumor expansion.
Abnormalities in FR proteins can cause intracellular iron
accumulation, leading to increased iron-associated reactive oxygen
species and lipid peroxidation, ultimately disrupting cellular
membrane integrity and triggering ferroptosis (30). The RB1 gene mutation is
closely associated with the development of RB and liver
carcinogenesis. It has been demonstrated that RB1 loss can increase
cancer cell susceptibility to ferroptosis inducers by elevating
acyl-CoA synthetase long chain family member 4, a key enzyme that
promotes lipid peroxidation and triggers ferroptosis (31). p53 serves a bidirectional role in
regulating ferroptosis, either through transcriptional or
posttranslational mechanisms (32).
These findings suggest that intervening in the occurrence of
ferroptosis may provide a potential avenue for influencing the
development and progression of RB.
GO and KEGG pathway analyses were conducted on the
23 FR-DEGs. KEGG analysis revealed significant enrichment only in
the ferroptosis pathway. Apart from iron ion homeostasis, GO
analysis revealed that these 23 FR-DEGs are also involved in other
biological processes, including epithelial cell apoptosis process.
Epithelial cell apoptosis shares similarities with other cell
types, but the unique characteristics of epithelial cells equip
them with specific determinants for survival (33). Cell-matrix and cell-cell
interactions are crucial to prevent epithelial cells from entering
apoptosis. In RB tumors, the destruction of these structures may
occur alongside the apoptotic process of epithelial cells (34). This enrichment analysis provides
valuable insights into the functional implications of DEGs in
ferroptosis in RB, enhancing the understanding of the molecular
mechanisms involved in these processes.
The importance of the
immune system in tumor progression and therapy has been widely
recognized. Immune cells interact with malignant tumor cells
through a complex network (35).
Therefore, the present study aimed to identify immune cell
infiltration as a biomarker for diagnosis and prognosis. Analysis
revealed a significant upregulation of naive B cells and Tregs,
along with a decrease in Tfh cells and resting mast cells in RB
tumor tissues. Tumor cells may recruit naive B cells into the
microenvironment, potentially transforming them into Breg cells
through various mechanisms (36).
Tregs have been shown to impede anticancer immune responses,
suppressing the beneficial immunosurveillance of tumors and
reducing effective anti-tumor immune protection (37). By contrast, Tfh cells are generally
associated with a superior prognosis in tumor entities, whereas
resting mast cells tended to be enriched in patients with cancer
with a low risk of recurrence (38). These findings suggest that immune
checkpoint inhibitors may offer a novel possibility for the
treatment of RB.
Central to the present findings are the hub genes
CAV1, CDKN2A, EPAS1, IDH2, RB1 and SLC2A3, identified
through PPI network analysis. These genes not only serve as key
players in the FR-DEG signature but also exhibit significant
dysregulation in RB. Validation in an independent dataset
(GSE208143) confirmed the expression patterns of these hub genes
and their AUC values highlight their diagnostic potential as RB
biomarkers. The sensitivity and specificity of these hub genes can
be further evaluated in larger independent cohorts in the future to
better assess their clinical diagnostic utility. It is noteworthy
that IDH2 and RB1 exhibited only moderate diagnostic performance as
individual biomarkers, suggesting limited standalone clinical
utility. However, diagnostic performance is not the only measure of
gene significance. IDH2 upregulation was validated by RT-qPCR and
its role in ferroptosis resistance supports its potential as a
therapeutic target. RB1 remains central to RB pathogenesis despite
its moderate AUC. Therefore, these genes may be more valuable for
functional studies or multi-gene panels than as standalone
diagnostic markers. CDKN2A is a tumor suppressor gene
encoding p16 and p14, the deletion of which can lead to melanoma
progression and can cause alterations in the way lipids are
metabolized and distributed in the cancer cells, thereby triggering
ferroptosis in glioblastoma cells (39,40).
IDH2 is the most frequently mutated metabolic gene in human
cancers, interfering with cellular metabolism and epigenetic
regulation to promote cancer development (41). Hsa-let-7b-5p serves several
different roles in inhibiting tumor cell proliferation, migration,
invasion and progression by targeting hexokinase 2, high mobility
group AT-hook 2, IGF1R and KIAA1377 in various tumor types
(42). Hsa-let-7g-5p is a
circulating miRNA and may serve as a biomarker for Alzheimer's
disease (43). As a
tumor-associated miRNA, hsa-miR-124-3p inhibits lung cancer by
regulating the ITGB1/PI3K/AKT axis (44). Abnormal LINC00963 expression
frequently promotes oncogenic activity by regulating key cellular
processes, such as proliferation, migration, invasion,
epithelial-mesenchymal transition and apoptosis (45). The lncRNA NEAT1 facilitates glioma
progression through stabilization of PGK1 (46). LncRNA XIST participates in the
development of tumors and other human diseases, underscoring its
role as an important regulator of cell growth and development
(47). In the present study, an FR
ceRNA network based on the aforementioned mRNAs, miRNAs and lncRNAs
was established, providing a novel therapeutic approach for the
treatment of RB. It should be noted that the ceRNA network is based
entirely on computational predictions. While these predicted
interactions offer valuable hypotheses for regulatory mechanisms in
RB, they require relevant experimental validation to confirm direct
binding and their roles in ferroptosis regulation.
The present study suggests a potential link between
RB and ferroptosis, constructed an lncRNA-miRNA-mRNA network
associated with ferroptosis and verified the expression of hub
genes by RT-qPCR, which was consistent with predicted results.
These findings provide a novel perspective for studying the role
and mechanism of ferroptosis in RB. However, several limitations
should be acknowledged. First, the sample sizes of GSE208143 and
GSE97508 gene sets are relatively small, with 33 and 9 samples.
second, the lack of clinical and prognostic data precluded clinical
association studies or prognostic analyses. Third, while normal
human retinal progenitor cells or mature retinal neurons (such as
photoreceptors) would be ideal controls, obtaining and culturing
primary human retinal neuronal or progenitor cells is technically
challenging due to their limited availability and difficulty in
maintaining them in vitro (48,49).
Therefore, ARPE-19 cells-one of the most commonly used and
well-characterized ‘normal’ retinal cell lines in ophthalmic
research-were employed as a surrogate control. ARPE-19 cells are
readily available, stable in culture and enable reproducible
experimental conditions (50,51).
Although derived from a different retinal lineage, they represent
the best available immortalized cell line from the human retina and
have been widely used as controls in numerous published RB studies
(52–57). Finally, the present ceRNA network
construction employed a stringent filter requiring lncRNAs to bind
all 3 hub gene-targeting miRNAs. While this approach minimized
false positives and identified core regulatory axes, it may have
omitted biologically relevant lncRNAs that interact with only one
or two of these miRNAs. Future experimental studies should explore
these partial interactions to fully elucidate the regulatory
landscape.
In conclusion, the present study unraveled the
ferroptotic landscape in RB, shedding light on the molecular
intricacies that govern this aggressive eye cancer. The
identification of key hub genes, functional insights and immune
modulation provide a foundation for future research directions and
therapeutic interventions targeting ferroptosis in RB.
Acknowledgements
Not applicable.
Funding
The present study was supported by the National Natural Science
Foundation in China (grant no. 81970830).
Availability of data and materials
Publicly available datasets were analyzed in this
study, which can be found in the Gene Expression Omnibus
(https://www.ncbi.nlm.nih.gov), FerrDb
database (http://www.zhounan.org/ferrdb/) and StarBase database
(http://StarBase.sysu.edu.cn/). The data
generated in the present study may be requested from the
corresponding author.
Authors' contributions
ZK contributed to the data collection and analysis,
as well as drafting and writing of the manuscript. GL conceived and
designed the manuscript, and critically reviewed the manuscript for
important intellectual content. Both authors have read and approved
the final manuscript. ZK and GL confirm the authenticity of all the
raw data.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Manukonda R, Narayana RV, Kaliki S, Mishra
DK and Vemuganti GK: Emerging therapeutic targets for
retinoblastoma. Expert Opin Ther Targets. 26:937–947. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Maçin A, Duman E, Özdemir İ, Öztürk Ş and
Tuncer MC: Hesperidin enhances doxorubicin efficacy by modulating
apoptosis- and migration-associated processes in human
retinoblastoma cells. Biology. 15:3052026. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Ancona-Lezama D, Dalvin L and Shields C:
Modern treatment of retinoblastoma: A 2020 review. Indian J
Ophthalmol. 68:23562020. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Zhao J, Feng ZX, Wei M, Liu G, Solarte CE,
Li B, Wang YZ, Zhang CY and Gallie BL: Impact of systemic
chemotherapy and delayed enucleation on survival of children with
advanced intraocular retinoblastoma. Ophthalmol Retina. 4:630–639.
2020. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Kumari A, Singh SP, Kumar P, Kondaveeti
SB, Garg VK, Kaur R, Buttar HS, Sak K, Yadav K and Yadav V: A
comprehensive review of the epidemiology, pathophysiology, risk
factors, and treatment strategies for retinoblastoma. Diseases.
13:3072025. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Kamihara J, Bourdeaut F, Foulkes WD,
Molenaar JJ, Mossé YP, Nakagawara A, Parareda A, Scollon SR,
Schneider KW, Skalet AH, et al: Retinoblastoma and neuroblastoma
predisposition and surveillance. Clin Cancer Res. 23:e98–e106.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Hosokawa T, Kuntaro D, Takei H, Arakawa Y,
Kambe T, Kurihara J, Mochizuki N, Sato Y and Tanami Y: Assessing
the usefulness of ultrasonography for the diagnosis and evaluation
of Intra-orbital lesions in pediatric patients: A retrospective
analysis. J Ultrasound Med. 43:573–585. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Dixon SJ, Lemberg KM, Lamprecht MR, Skouta
R, Zaitsev EM, Gleason CE, Patel DN, Bauer AJ, Cantley AM, Yang WS,
et al: Ferroptosis: An Iron-dependent form of nonapoptotic cell
death. Cell. 149:1060–1072. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Tang D, Chen X, Kang R and Kroemer G:
Ferroptosis: Molecular mechanisms and health implications. Cell
Res. 31:107–125. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Jiang X, Stockwell BR and Conrad M:
Ferroptosis: Mechanisms, biology and role in disease. Nat Rev Mol
Cell Biol. 22:266–282. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Abu-Amero KK, Kondkar AA, Almontashiri
NAM, Khan AM, Maktabi AMY, Hameed S and AlMesfer S: Genetics of
retinoblastoma: An overview and significance of genetic testing in
clinical practice. Genes. 16:10312025. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Zuo YB, Zhang YF, Zhang R, Tian JW, Lv XB,
Li R, Li SP, Cheng MD, Shan J, Zhao Z and Xin H: Ferroptosis in
cancer progression: Role of noncoding RNAs. Int J Biol Sci.
18:1829–1843. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Barbi V, De Martino S, Aiello A, Gottardi
Zamperla M, Negri S, Cis L, Pecci V, Nanni S, Farsetti A, Martelli
F, et al: Non-coding RNAs as novel biomarkers and therapeutic
targets in breast cancer. Oncol Rev. 19:16211442025. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Zheng X and Zhang C: The regulation of
Ferroptosis by noncoding RNAs. Int J Mol Sci. 24:133362023.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Wu J, Qian D and Sun X: Long noncoding
RNAs as potential biomarkers in retinoblastoma: A systematic review
and meta-analysis. Cancer Cell Int. 20:2012020. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Barrett T, Wilhite SE, Ledoux P,
Evangelista C, Kim IF, Tomashevsky M, Marshall KA, Phillippy KH,
Sherman PM, Holko M, et al: NCBI GEO: Archive for functional
genomics data sets-update. Nucleic Acids Res. 41:D991–D995. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW,
Shi W and Smyth GK: limma powers differential expression analyses
for RNA-sequencing and microarray studies. Nucleic Acids Res.
43:e47. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Yu G, Wang LG, Han Y and He QY:
clusterProfiler: An R package for comparing biological themes among
gene clusters. OMICS J Integr Biol. 16:284–287. 2012. View Article : Google Scholar
|
|
19
|
Ito K and Murphy D: Application of ggplot2
to pharmacometric graphics. CPT Pharmacomet Syst Pharmacol.
2:e792013. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Chin CH, Chen SH, Wu HH, Ho CW, Ko MT and
Lin CY: cytoHubba: Identifying hub objects and sub-networks from
complex interactome. BMC Syst Biol. 8 (Suppl 4):S112014. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Zeng D, Ye Z, Shen R, Yu G, Wu J, Xiong Y,
Zhou R, Qiu WJ, Huang N, Sun L, et al: IOBR: Multi-Omics
Immuno-oncology biological research to decode tumor
microenvironment and signatures. Front Immunol. 12:6879752021.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Ru Y, Kechris KJ, Tabakoff B, Hoffman P,
Radcliffe RA, Bowler R, Mahaffey S, Rossi S, Calin GA, Bemis L, et
al: The multiMiR R package and database: Integration of
microRNA-target interactions along with their disease and drug
associations. Nucleic Acids Res. 42:e133. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Xiang J, Chen H, Lin Z, Chen J and Luo L:
Identification and experimental validation of ferroptosis-related
gene SLC2A3 is involved in rheumatoid arthritis. Eur J Pharmacol.
943:1755682023. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Li H and Jin C: Retinoblastoma: Unveiling
molecular pathogenesis and pioneering organoid-driven therapeutic
innovations. Stem Cell Res Ther. 17:962026. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Fernández-Acosta R, Vintea I, Koeken I,
Hassannia B and Vanden Berghe T: Harnessing ferroptosis for
precision oncology: Challenges and prospects. BMC Biol. 23:572025.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Hanahan D: Hallmarks of cancer: New
dimensions. Cancer Discov. 12:31–46. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Zhao L, Zhou X, Xie F and Zhang L, Yan H,
Huang J, Zhang C, Zhou FF, Chen J and Zhang L: Ferroptosis in
cancer and cancer immunotherapy. Cancer Commun. 42:88–116. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Friedmann Angeli JP, Krysko DV and Conrad
M: Ferroptosis at the crossroads of cancer-acquired drug resistance
and immune evasion. Nat Rev Cancer. 19:405–414. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Yang L, Wang H, Yang X, Wu Q, An P, Jin X,
Liu W, Huang X, Li YZ, Yan S, et al: Auranofin mitigates systemic
iron overload and induces ferroptosis via distinct mechanisms.
Signal Transduct Target Ther. 5:1382020. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Wang ME, Chen J, Lu Y, Bawcom AR, Wu J, Ou
J, Asara JM, Armstrong AJ, Wang QB, Li L, et al: RB1-deficient
prostate tumor growth and metastasis are vulnerable to ferroptosis
induction via the E2F/ACSL4 axis. J Clin Invest. 133:e1666472023.
View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Ji H, Wang W, Li X, Han X, Zhang X, Wang
J, Liu C, Huang L and Gao W: p53: A double-edged sword in tumor
ferroptosis. Pharmacol Res. 177:1060132022. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Metcalfe A and Streuli C: Epithelial
apoptosis. Bioessays. 19:711–720. 1997. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Frisch SM and Francis H: Disruption of
epithelial cell-matrix interactions induces apoptosis. J Cell Biol.
124:619–626. 1994. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Zuo S, Wei M, Wang S, Dong J and Wei J:
Pan-cancer analysis of immune cell infiltration identifies a
prognostic immune-cell characteristic score (ICCS) in lung
adenocarcinoma. Front Immunol. 11:12182020. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Yang S, Yang Y, Fang Y, Zhou Q, Sun W,
Zhang Z, Yuan W and Li Z: Targeting tumour-infiltrating B cells:
Mechanisms and advances in cancer therapy. Cell Death Dis.
17:532025. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Li C, Jiang P, Wei S, Xu X and Wang J:
Regulatory T cells in tumor microenvironment: New mechanisms,
potential therapeutic strategies and future prospects. Mol Cancer.
19:1162020. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Baumjohann D and Brossart P: T follicular
helper cells: Linking cancer immunotherapy and immune-related
adverse events. J Immunother Cancer. 9:e0025882021. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Kreuger IZM, Slieker RC, Van Groningen T
and Van Doorn R: Therapeutic strategies for targeting CDKN2A loss
in melanoma. J Invest Dermatol. 143:18–25.e1. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Minami JK, Morrow D, Bayley NA, Fernandez
EG, Salinas JJ, Tse C, Zhu H, Su B, Plawat R, Jones A, et al:
CDKN2A deletion remodels lipid metabolism to prime glioblastoma for
ferroptosis. Cancer Cell. 41:1048–1060.e9. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Vuong HG, Ngo TNM and Dunn IF: Prognostic
importance of IDH mutations in chondrosarcoma: An individual
patient data meta-analysis. Cancer Med. 10:4415–4423. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Li L, Zhang X, Lin Y, Ren X, Xie T, Lin J,
Wu SM and Ye QN: Let-7b-5p inhibits breast cancer cell growth and
metastasis via repression of hexokinase 2-mediated aerobic
glycolysis. Cell Death Discov. 9:1142023. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Poursaei E, Abolghasemi M, Bornehdeli S,
Shanehbandi D, Asadi M, Sadeghzadeh M, Rahmanpour D and Sadeh RN:
Evaluation of hsa-let-7d-5p, hsa-let-7g-5p and hsa-miR-15b-5p
plasma levels in patients with Alzheimer's disease. Psychiatr
Genet. 32:25–29. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Han R, Wang S, Zhou Z, Huang D, Hou J,
Tian M, Ge R and Ma Y: MicroRNA-124-3p suppresses lung cancer by
targeting ITGB1/PI3K/p-AKT signal transduction pathway. Exp Cell
Res. 454:1148522026. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Xie Z, Zhong C, Shen J, Jia Y and Duan S:
LINC00963: A potential cancer diagnostic and therapeutic target.
Biomed Pharmacother. 150:1130192022. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Liang J, Liu C, Xu D, Xie K and Li A:
LncRNA NEAT1 facilitates glioma progression via stabilizing PGK1. J
Transl Med. 20:802022. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Wang W, Min L, Qiu X, Wu X, Liu C, Ma J,
Zhang DY and Zhu LY: Biological Function of Long Non-coding RNA
(LncRNA) Xist. Front Cell Dev Biol. 9:6456472021. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Coco-Martin RM, Pastor-Idoate S and Pastor
JC: Cell replacement therapy for retinal and optic nerve diseases:
Cell sources, clinical trials and challenges. Pharmaceutics.
13:8652021. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Zhang D, Shen B, Zhang Y, Ni N, Wang Y,
Fan X, Sun H and Gu P: Betacellulin regulates the proliferation and
differentiation of retinal progenitor cells in vitro. J Cell Mol
Med. 22:330–345. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Dunn KC, Aotaki-Keen AE, Putkey FR and
Hjelmeland LM: ARPE-19, A human retinal pigment epithelial cell
line with differentiated properties. Exp Eye Res. 62:155–170. 1996.
View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Hellinen L, Hagström M, Knuutila H,
Ruponen M, Urtti A and Reinisalo M: Characterization of
artificially re-pigmented ARPE-19 retinal pigment epithelial cell
model. Sci Rep. 9:137612019. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Jiang A, Wu W, Xu C, Mao L, Ao S, Guo H,
Sun X, Tao J, Sang Y and Huang G: SP2509, a Selective Inhibitor of
LSD1, suppresses retinoblastoma growth by downregulating β-catenin
signaling. Investig Opthalmology Vis Sci. 63:202022. View Article : Google Scholar
|
|
53
|
Fu K, Zhang K and Zhang X: LncRNA HOTAIR
facilitates proliferation and represses apoptosis of retinoblastoma
cells through the miR-20b-5p/RRM2/PI3K/AKT axis. Orphanet J Rare
Dis. 17:1192022. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Oshikawa M, Tsutsui C, Ikegami T, Fuchida
Y, Matsubara M, Toyama S, Usami R, Obtoko K and Kato S: Full-length
transcriptome analysis of human retina-derived cell lines ARPE-19
and Y79 using the Vector-capping method. Invest Ophthalmol Vis Sci.
52:6662–6670. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Bai S, Tian B, Li A, Yao Q, Zhang G and Li
F: MicroRNA-125b promotes tumor growth and suppresses apoptosis by
targeting DRAM2 in retinoblastoma. Eye (Lond). 30:1630–1638. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Sun QX, Wang RR, Liu N and Liu C:
Dysregulation of miR-204-3p driven by the viability and motility of
retinoblastoma via Wnt/β-catenin pathway in vitro and in vivo.
Pathol Oncol Res. 26:1549–1558. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Zhang H, Qiu X, Song Z, Lan L, Ren X and
Ye B: CircCUL2 suppresses retinoblastoma cells by regulating
miR-214-5p/E2F2 axis. Anticancer Drugs. 33:e218–e227. 2022.
View Article : Google Scholar : PubMed/NCBI
|