Introduction
Ovarian cancer (OC) is a prevalent malignancy
affecting >320,000 women globally (1). Epithelial OC, constituting 90% of
cases, is highly heterogeneous and poses challenges for early
detection (2). The absence of
sensitive diagnostic tools often leads to late-stage diagnoses,
resulting in a low 5-year survival rate of 30% for advanced cases
(3). Therefore, identifying precise
biomarkers in early screening and predicting overall survival in
high-risk OC populations is imperative to address the challenges of
late-stage diagnosis and poor prognosis.
Carbohydrate antigen 125 (CA125) is a conventional
tumor marker for OC, albeit with limited sensitivity (50–62%) and
specificity (73–77%) (4). Elevated
CA125 levels have been observed in pregnancy, pelvic inflammatory
disease and various other types of cancer [including breast
(5), uterine (6) and lung (7) cancer]. Previous research has
highlighted the utility of cell-free DNA (cfDNA) methylation
analysis in early cancer detection. Aberrant cfDNA methylation,
detected in the initial stages of OC, modulates gene expression by
methylating specific gene sites (8–10). For
example, hypermethylation of the tumor suppressor genes homeobox A9
(HOXA9) and hypermethylated in cancer-zinc finger and BTB
domain-containing protein transcriptional repressor 1 (HIC1)
facilitates uncontrolled cell proliferation, contributing to OC
development (11). Furthermore,
hypermethylation of bromodomain-containing 4 enhances the invasive
potential of OC (12). These gene
methylation alterations represent promising biomarkers for early
cancer diagnosis.
cfDNA harbors tumor-specific genetic alterations and
can be utilized for non-invasive cancer detection by analyzing
aberrant methylation patterns (13). Previous research has highlighted the
significance of nidogen-2 (NID2) and cysteine dioxygenase 1 (CDO1)
as tumor suppressor genes (14,15).
NID2, a basement membrane component, serves a key role in
maintaining membrane integrity, with its deficiency promoting
cancer development (16). By
contrast, upregulation of NID2 in lung cancer cells suppresses
proliferation and invasiveness (17). CDO1 facilitates apoptosis by
modulating antioxidant levels and is frequently epigenetically
altered in various types of cancer, commonly through promoter
methylation (18). Elevated CDO1
promoter methylation has been observed in solid tumors such as
breast (19) and lung cancer
(20). However, the methylation
status of NID2 and CDO1 in OC remains poorly understood. The
present study aimed to assess the methylation profiles of NID2 and
CDO1 in OC, to establish associations with clinical data and to
investigate their potential as specific biomarkers for OC
screening.
Patients and methods
Study population and blood sample
collection
The present study recruited 147 participants who
attended Hangzhou Cancer Hospital (Hangzhou, China) between
February 2023 and June 2024. Among them, 72 were patients with a
clear clinical diagnosis of OC and 75 were normal control (NC)
group. Ovarian cancer staging was determined according to the
International Federation of Gynecology and Obstetrics (FIGO) 2018
staging system (21). Inclusion
criteria were as follows: i) Female patients with a confirmed
diagnosis of OC; ii) aged between 35 and 75 years; iii) Eastern
Cooperative Oncology Group (ECOG) performance status 0–4 (22); iv) hospitalized patients with a life
expectancy of >2 months; v) participants voluntarily joined the
present study and provided signed informed consent; and vi)
patients were conscious and able to manage their own dietary
control. Exclusion criteria were as follows: i) Expected hospital
stay of <3 days; ii) underwent major surgery recently or any
surgery lasting >2 h; iii) participation in any other clinical
trial within 4 weeks prior to the start of the present study; iv)
respiratory depression, (pulmonary) airway obstruction or tissue
hypoxia; v) hematological diseases; vi) cardiac diseases [namely,
cardiac function of New York Heart Association class II or higher
(23)]; vii) pregnant or lactating
women; viii) brain disorders with impaired judgment; and ix) unable
to provide signed informed consent. A 1.5-ml peripheral blood
sample was collected from each individual for methylation testing,
with blood from patients with OC collected prior to antitumor
therapy. Clinical data and the results of tumor marker CA125
detection were collected from all subjects. The cut-off value of
CA125 was 35 U/ml. The present study was approved by the
Institutional Review Board of Hangzhou Cancer Hospital (approval
no. HZCH-2023-006; Hangzhou, China). Written informed consent to
participate was obtained from all individual participants included
in the present study prior to their enrolment. The present study
was conducted in accordance with the principles of The Declaration
of Helsinki.
DNA isolation and bisulfite
conversion
cfDNA was extracted from serum samples using the
QIAamp® DNA Mini Kit (Qiagen GmbH). The concentration
and purity of the nucleic acids were measured using the NanoDrop™
One microvolume UV–Vis spectrophotometer (Thermo Fisher Scientific,
Inc.). For bisulfite conversion, 500 ng of the extracted cfDNA was
used with the EZ DNA Methylation™ Kit (Zymo Research Corp.), and
the reaction was carried out according to the manufacturer's
protocol.
Quantitative methylation-specific PCR
(qMSP)
Methylation levels of the NID2 and CDO1 genes were
analyzed using qMSP with the Applied Biosystems 7500 real-time PCR
system (cat. no. ABI7500; Thermo Fisher Scientific, Inc.). Each
reaction contained a fluorogenic probe labeled with VIC™
fluorescent dye (Thermo Fisher Scientific, Inc.) and conjugated
with a minor groove binder. The primers were designed to adhere to
the principle of sequence specificity following bisulfite
conversion and pre-validated using PCR amplification using DNA
templates with known methylation status that had undergone
bisulfite treatment. The sequences of the primers used for qMSP are
detailed in Table SI.
Each PCR was performed in a 96-well plate with
β-actin as an internal reference gene and three parallel wells per
reaction. Each assay rigorously incorporated both negative and
positive controls. The thermal cycling conditions were as follows:
i) 95°C for 5 min; ii) 15 cycles at 95°C for 15 sec and 64°C for 30
sec; and iii) 30 cycles at 95°C for 15 sec and 62°C for 32 sec. For
analysis, the quantification cycle (Cq) value for each sample was
used, ΔCq was calculated using the following formula: ΔCq=average
Cq target gene - average Cq internal reference. Gene methylation
levels for each sample were expressed as 2−ΔΔCq,
ΔΔCq=ΔCq (OC) - ΔCq (NC average) (24).
Statistical analysis
Statistical analyses were performed using SPSS
(version 26.0; IBM Corp.). The Shapiro-Wilk test was employed to
analyze the distribution pattern of the data. As the data exhibited
significant dispersion, non-parametric tests were employed, and the
results for each index were expressed as the median and
interquartile range [M (25th percentile, 75th percentile)]. The
Mann-Whitney U test was used to compare continuous variables
between two groups. Spearman's correlation analysis was used to
analyze correlations between variables. Categorical variables were
expressed as n (%). The χ2 test was used to compare
count data. When the expected frequency in >20% of the cells was
<5, Fisher's exact test was used instead. To adjust for
potential confounding factors (hypertension and performance
status), a propensity score matching (PSM) analysis was performed
using a 1:1 nearest neighbor matching algorithm without
replacement. Receiver operating characteristic (ROC) curve analysis
was used to determine the best key value in predicting OC. Each
qMSP assay was performed in triplicate, and the mean values were
used for analysis. The χ2 test and logistic regression
analysis were used to determine the risk factors for OC. P<0.05
was considered to indicate a statistically significant
difference.
Results
Clinical characteristics
The clinical baseline characteristics of the
participants are presented in Table
I. There were 72 individuals in the OC group, with a median age
of 59.5 years, and the tissue typing was mainly high-grade serous
carcinoma [HGSC; 88.9% (64/72)]. The majority of patients were in
the advanced stages [81.9% (59/72)] of OC, with chemotherapy [88.9%
(64/72)] being the primary treatment method. During
hospitalization, disease progression occurred in 25/72 (34.7%)
patients and 5/72 (6.9%) patients died. The NC group consisted of
75 healthy individuals, with a median age of 65.0 years. Analysis
of CA125 levels in the two groups revealed that the median CA125
level in the OC group was 178.5 U/ml, whereas the median CA125
level in the NC group was 12.8 U/ml (normal reference value: <35
U/ml; P<0.001), with a statistically significant difference.
Furthermore, patients with OC had a significantly higher prevalence
of hypertension and worse performance status scores, whereas there
was no significant difference in the prevalence of type 2 diabetes
mellitus compared with that in healthy individuals.
 | Table I.Characteristics of participants in
the present study. |
Table I.
Characteristics of participants in
the present study.
|
Characteristics | NC (n=75) | OC (n=72) | P-value |
|---|
| Age, years [median
(range)] | 65.0
(55.0–71.0) | 59.5
(54.0–66.8) | 0.105 |
| Histology |
|
| - |
|
HGSC | - | 64 (88.9) |
|
|
OCCC | - | 8 (11.1) |
|
| FIGO stage |
|
| - |
|
I–II | - | 13 (18.1) |
|
|
III–IV | - | 59 (81.9) |
|
| No metastasis | - | 13 (18.1) | - |
| Metastasis |
|
| - |
| Lymph
nodes | - | 21 (29.2) |
|
|
Abdominal cavity | - | 9 (12.5) |
|
|
Pelvis | - | 14 (19.4) |
|
|
Liver | - | 2 (2.8) |
|
|
Lungs | - | 1 (1.4) |
|
|
Multiple metastases | - | 12 (16.7) |
|
| ECOG PS score |
|
| 0.012 |
|
0-2 | 70 (93.3) | 57 (79.2) |
|
|
3-4 | 5 (6.7) | 15 (20.8) |
|
| Hypertension | 16 (21.3) | 27 (37.5) | 0.031 |
| Type 2 diabetes
mellitus | 13 (17.3) | 21 (29.2) | 0.089 |
| Family history of
cancer | - | 22 (30.6) | - |
| Treatment
method |
|
| - |
|
Chemotherapy | - | 64 (88.9) |
|
|
Nutritional support | - | 8 (11.1) |
|
| CA125, U/ml
(range) | 12.8
(8.9–19.1) | 178.5
(34.0–582.2) | <0.01 |
| Primary therapy
outcome |
|
| - |
|
CR-PR | - | 11 (15.3) |
|
| SD | - | 36 (50.0) |
|
| PD | - | 25 (34.7) |
|
| Mortality | - | 5 (6.9) | - |
DNA methylation levels of NID2 and
CDO1 genes
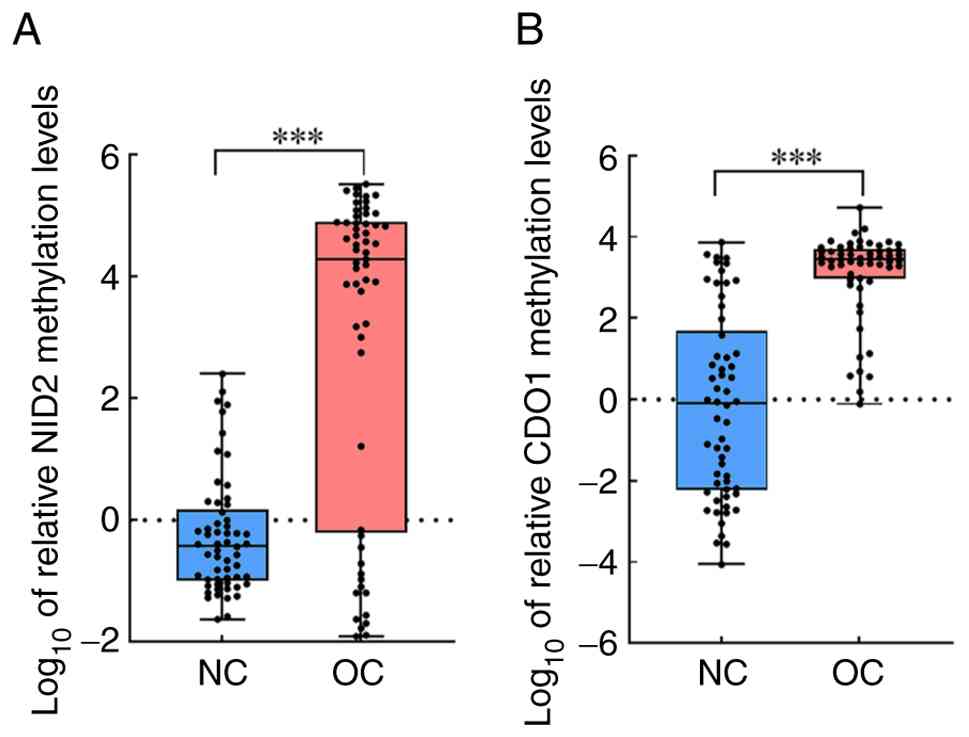
Methylation levels were determined using qMSP
(Fig. 1A and B). The results
demonstrated that the methylation levels of the NID2 gene in the
serum of the NC and OC groups were 0.41 (0.11–2.26) and
1.90×104 (1.49–7.75×104), respectively,
whereas the methylation levels of the CDO1 gene were 0.91
(0.01–180.1) and 2.83×103 (836.3–4.67×103),
respectively. Compared with the NC group, the methylation levels of
NID2 and CDO1 in patients with OC were significantly higher
(P<0.001). Furthermore, to evaluate the potential confounding
effects of baseline hypertension status and physical fitness on
methylation outcomes, PSM analysis was performed (Table SII; Fig. S1). Using a 1:1 nearest neighbor
matching algorithm without replacement, 57 participants in the NC
group were successfully matched to 57 patients in the OC group.
Post-PSM analyses revealed that the methylation levels of both NID2
and CDO1 genes remained significantly elevated in the OC cohort
compared with that in the matched controls (P<0.05), which is
consistent with pre-matched observations.
Predictive value of NID2 and CDO1 gene
DNA methylation for OC
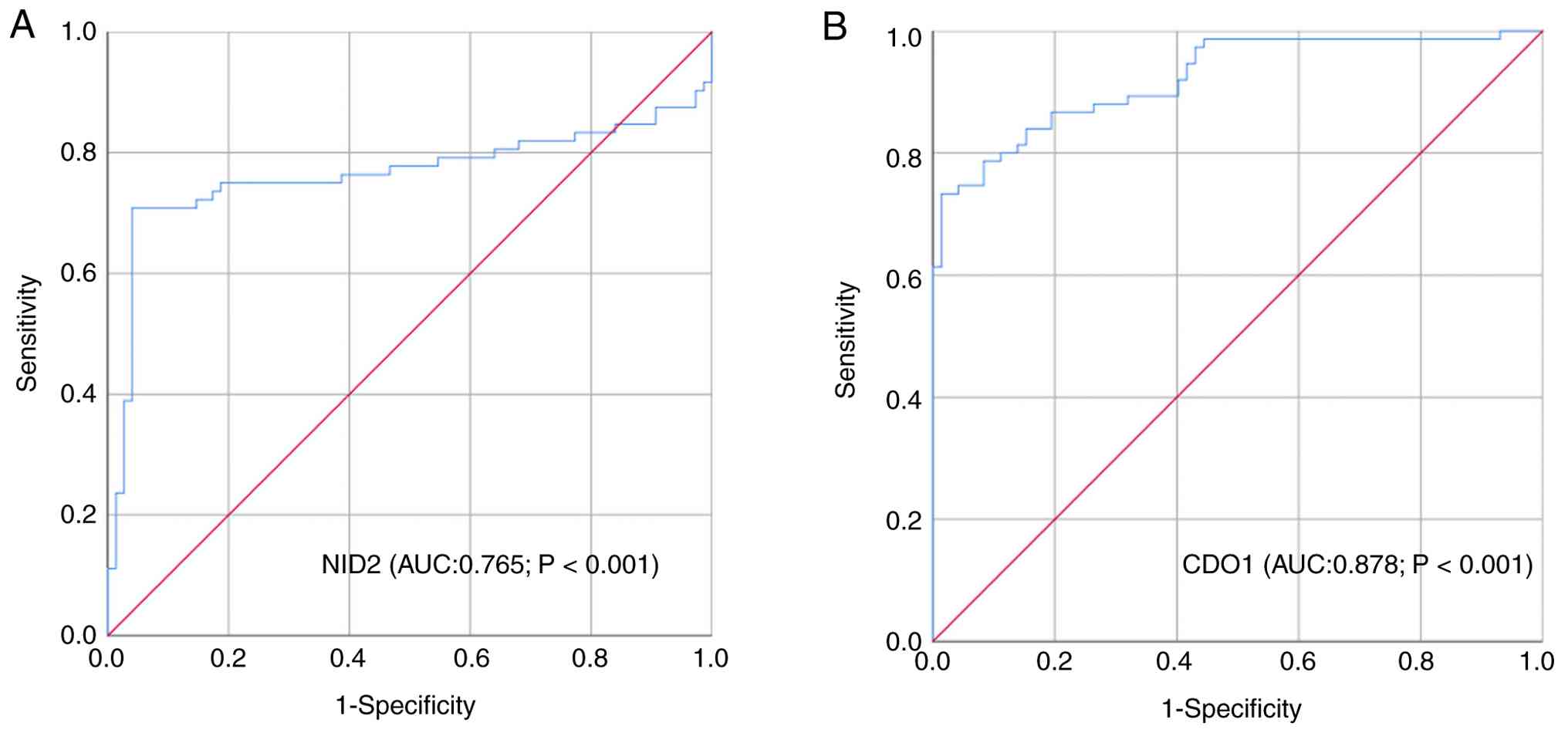
To investigate the predictive value and detection
cut-off values of the NID2 and CDO1 genes for OC, ROC curves were
plotted using the ΔCq values. As presented in Fig. 2A and B, and Table II, the area under the curve (AUC)
values in predicting OC with NID2 and CDO1 were 0.765 and 0.878,
respectively. When the Youden index was maximized, the optimal ΔCq
values in predicting OC with these markers were 16.95 for NID2 and
15.99 for CDO1. Among these, CDO1 exhibited a higher detection
efficacy for OC, with an AUC of 0.878 (P<0.001).
 | Table II.Diagnostic values of serum gene
methylation makers for patients with ovarian cancer. |
Table II.
Diagnostic values of serum gene
methylation makers for patients with ovarian cancer.
| Markers | Cut-off | Sn, % | Sp, % | Youden index | AUC | P-value | 95% CI |
|---|
| NID2 | 16.95 | 70.83 | 96.00 | 0.668 | 0.765 | <0.001 | 0.677–0.853 |
| CDO1 | 15.99 | 90.28 | 69.33 | 0.596 | 0.878 | <0.001 | 0.880–0.965 |
Detection rates of the NID2 and CDO1
methylation test
Based on the cut-off values of NID2 and CDO1 gene
methylation (Table II), the
positive detection rates were calculated in different FIGO stages
and pathological types (Fig. 3A-D;
Table III). The results
demonstrated that the positive detection rates of NID2 and CDO1
methylation in OC were 70.8% (51/72) and 90.3% (65/72),
respectively. In stages I–II, the positive detection rates for NID2
and CDO1 methylation were 46.2% (6/13) and 84.6% (11/13),
respectively, and in stages III–IV, the positive detection rates
were 76.3% (45/59) and 91.5% (54/59), respectively. Furthermore,
the positive detection rates of NID2 and CDO1 methylation in HGSC
were 71.9% (46/64) and 90.6% (58/64), respectively, whereas in
ovarian clear cell carcinoma, the positive detection rates were
62.5% (5/8) and 87.5% (7/8), respectively. The clinical data
analysis indicated that patients with OC ≥65 years of age and with
FIGO stages III–IV had a higher positive detection rate for NID2
methylation, with statistically significant differences (both
P<0.05), whereas there were no significant differences in
different histological subtypes and various prognoses among the
patients. Additionally, no significant changes were observed in the
positive detection rate of CDO1 methylation among these variables
(all P>0.05).
 | Table III.Detection rates of NID2 and CDO1
methylation. |
Table III.
Detection rates of NID2 and CDO1
methylation.
|
| NID2 (n=72) | CDO1 (n=72) |
|---|
|
|
|
|
|---|
|
Characteristics | Negative
(n=21) | Positive
(n=51) | P-value | Negative (n=7) | Positive
(n=65) | P-value |
|---|
| Age, years |
|
| 0.015 |
|
| 0.413 |
|
≥65 | 18 (85.7) | 27 (52.9) |
| 3 (42.9) | 42 (64.6) |
|
|
<65 | 3 (14.3) | 24 (47.1) |
| 4 (57.1) | 23 (35.4) |
|
| FIGO stage |
|
| 0.031 |
|
| 0.602 |
|
I–II | 7 (33.3) | 6 (11.8) |
| 2 (28.6) | 11 (16.9) |
|
|
III–IV | 14 (66.7) | 45 (88.2) |
| 5 (71.4) | 54 (83.1) |
|
| Histology |
|
| 0.684 |
|
| 0.578 |
|
HGSC | 18 (85.7) | 46 (90.2) |
| 6 (85.7) | 58 (89.2) |
|
|
OCCC | 3 (14.3) | 5 (9.8) |
| 1 (14.3) | 7 (10.8) |
|
| Prognosis |
|
| 0.495 |
|
| >0.999 |
|
DCB | 15 (71.4) | 32 (62.7) |
| 5 (71.4) | 42 (64.6) |
|
|
Non-DCB | 6 (28.6) | 19 (37.3) |
| 2 (28.6) | 23 (35.4) |
|
Combined detection of NID2 and CDO1
gene DNA methylation and CA125
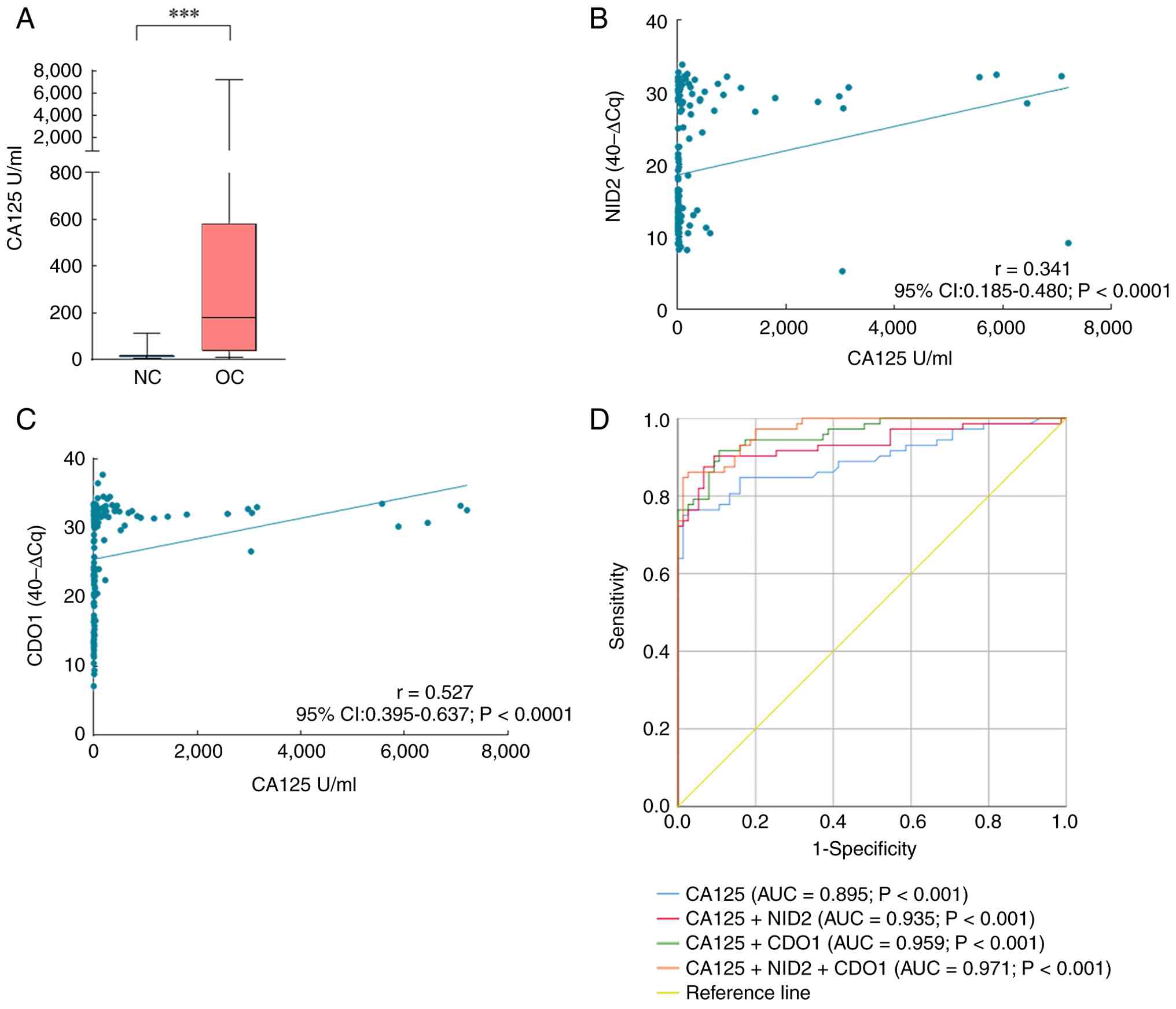
To further evaluate the predictive value of NID2 and
CDO1 gene DNA methylation for OC, the serum levels of the
traditional OC marker CA125 were also analyzed (Fig. 4A). The results indicated that the
serum CA125 level in the NC group was 12.8 (8.90–19.10) U/ml,
whereas in the OC group it was 178.45 (34.00–582.23) U/ml, which
was significantly higher compared with that in the NC group
(P<0.01). Correlation analysis demonstrated that serum CA125 was
positively correlated with NID2 (r=0.341; P<0.0001) and CDO1
(r=0.527; P<0.0001) (Fig. 4B and
C). Furthermore, ROC curves for the three markers across
different FIGO stages of OC demonstrated that the AUC values of
NID2 and CDO1 methylation, and CA125 were 0.560 (P=0.491), 0.916
(P<0.001) and 0.893 (P<0.001), respectively, in patients with
FIGO stage I–II OC (Fig. S2;
Table SIII). Notably, CDO1
methylation exhibited the highest sensitivity (84.6%), whereas
CA125 exhibited the highest specificity (97.3%). In patients with
stage III–IV OC (Fig. S2; Table SIV), the AUC values were 0.810
(P<0.001), 0.870 (P<0.001) and 0.878 (P<0.001),
respectively, with CDO1 methylation achieving the highest
sensitivity (91.5%) and NID2 methylation demonstrating the highest
specificity (96.0%). The three markers were further arranged and
combined, and according to the ROC curve results, among the
multiple combined detections, the combination of CA125 + NID2 +
CDO1 exhibited the highest efficacy, with an AUC value of 0.971
(P<0.001), a sensitivity of 86.1% and a specificity of 97.3%
(Fig. 4D; Table IV).
 | Table IV.Predictive value of combined
detection of NID2 and CDO1 gene DNA methylation and CA125 for
ovarian cancer. |
Table IV.
Predictive value of combined
detection of NID2 and CDO1 gene DNA methylation and CA125 for
ovarian cancer.
| Markers | Sn, % | Sp, % | Youden index | AUC | P-value | 95% CI |
|---|
| CA125 | 76.4 | 97.3 | 0.737 | 0.895 | <0.001 | 0.840–0.949 |
| CA125 + NID2 | 90.3 | 90.7 | 0.809 | 0.935 | <0.001 | 0.891–0.979 |
| CA125 + CDO1 | 91.7 | 89.3 | 0.810 | 0.959 | <0.001 | 0.931–0.987 |
| CA125 + NID2 +
CDO1 | 86.1 | 97.3 | 0.834 | 0.971 | <0.001 | 0.950–0.992 |
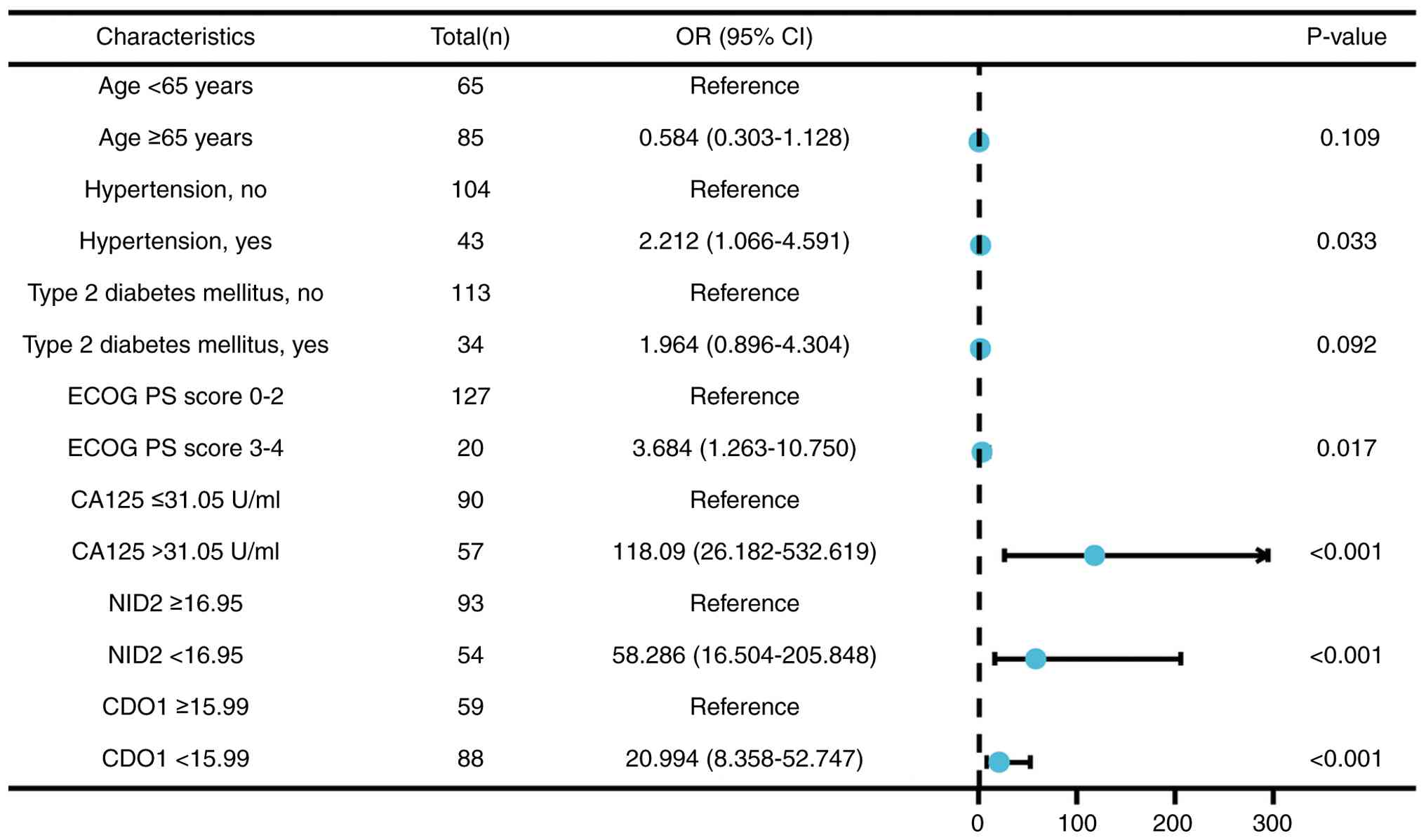
Risk factor analysis for the
occurrence of OC
The present study combined the predictive serum
levels of NID2, CDO1 and CA125 for OC with clinical data to perform
a risk factor analysis (Fig. 5).
The results indicated that serum CA125 level >31.05 U/ml [odds
ratio (OR), 118.09; 95% CI, 26.182–532.619; P<0.001], NID2 ΔCq
<16.95 (OR, 58.286; 95% CI, 16.504–205.848; P<0.001) and CDO1
ΔCq <15.99 (OR, 20.994; 95% CI, 8.358–52.747; P<0.001) are
potential risk factors for the occurrence of OC.
Discussion
DNA methylation is a pivotal early event in cancer
initiation, offering potential for cancer diagnosis and therapy.
Recent studies have highlighted the key involvement of DNA
methylation in the onset (25),
progression (26) and drug
resistance (27) of OC. Serum
methylation markers exhibited promise in detecting OC within the
initial 2 years of diagnosis, enabling early detection and
personalized treatment strategies for OC (28,29).
The present study revealed that patients with OC
exhibited elevated methylation levels of NID2 and CDO1 compared
with healthy individuals. This finding was further validated in the
post-PSM analyses, confirming the robustness of the observed
association. Specifically, older patients with advanced-stage OC
exhibited a higher prevalence of NID2 methylation, whereas its
efficacy was diminished in early-stage OC. This may be associated
with the case cohort that was primarily composed of patients with
late-stage disease. However, since the majority of patients with OC
are already in the advanced stages of the disease at their first
clinical visit, the results of methylation testing in recent years
have also been generally influenced by late-stage bias, with
markedly reduced sensitivity in early asymptomatic patients
(30–32). Notably, in the present study, the
positive rate of CDO1 methylation did not demonstrate a significant
difference across different FIGO stages, and CDO1 methylation
demonstrated high detection sensitivity in patients with both
early-stage (84.6%) and advanced-stage (91.5%) OC. These findings
indicated that CDO1 methylation may become an important marker for
the early screening of OC. Despite exhibiting significantly lower
diagnostic capability in early-stage OC compared with that in
late-stage OC, NID2 methylation maintained consistently higher
detection specificity compared with CDO1 (96.00%) throughout the
analysis.
Compared with existing diagnostic methods, CA125,
currently relied upon for OC diagnosis, has insufficient
sensitivity and specificity in early-stage disease and certain
pathological types, while imaging techniques are insensitive to
microscopic lesions (33). In the
present study, CDO1 methylation maintained a high positive rate
(84.6%; 11/13) and strong discriminatory ability (AUC=0.916) even
in patients with FIGO stage I–II, suggesting its potential to
address the shortcomings of current methods in early screening.
Prior research has demonstrated that integrating methylation data
from multiple genes can markedly enhance the accuracy of OC
detection in comparison with the utilization of individual genes
(34,35). A previous meta-analysis focusing on
OC revealed that out of 16 methylated genes, the sensitivity for OC
detection varied from 41.4 to 100%, with 12 genes exhibiting a
sensitivity of >75%. Furthermore, the specificity for OC
detection ranged from 55.0 to 100%, with seven genes demonstrating
a specificity of >95% (30). The
present study revealed a significantly positive correlation between
the methylation levels of NID2 and CDO1 and the conventional OC
serological marker CA125. The combined assay of the three detection
markers further improved diagnostic sensitivity (86.1%) and
specificity (97.3%) for OC, thereby partially addressing the
limitations in sensitivity and specificity associated with CA125 as
a single biomarker. However, compared with mature biomarker
detection systems, methylation testing currently incurs higher
costs and requires standardized experimental procedures and
threshold definition, which presents a challenge for its broader
clinical translation in the future (36).
The clinical utility of NID2 and CDO1 as methylation
biomarkers requires evaluation within the broader context of
methylation screening technologies. As a key tumor suppressor gene,
promoter hypermethylation of CDO1 is a common epigenetic event in
various solid tumors, including breast (19), endometrial (37) and lung cancer (38). This aligns biologically with the
notable hypermethylation observed in the serum of patients with OC
in the present study. More notably, recent whole-genome sequencing
studies have identified CDO1 hypermethylation in both urine and
tissue samples from patients with OC (39). The present study has further
confirmed this phenomenon in blood samples and demonstrated its
diagnostic performance using quantitative analysis, collectively
reinforcing the scientific foundation and translational potential
of CDO1 hypermethylation as a cross-cancer biomarker for OC. In
contrast to CDO1, although direct evidence for NID2 promoter
methylation in OC cfDNA is limited, NID2, as a basement
membrane-associated gene, has been repeatedly reported to exhibit
promoter hypermethylation and functional silencing in other
malignancies such as gastrointestinal (40) and lung cancer (17). Mechanistic studies have demonstrated
that NID2 hypermethylation leads to reduced expression, while
demethylation or upregulation can inhibit tumor cell proliferation,
migration and invasion, supporting the hypothesis that NID2 may act
as a tumor suppressor silenced via methylation (17,41,42).
In the field of OC, previous research has largely focused on NID2
protein levels, revealing elevated serum levels associated with
CA125 (43,44). The present study findings further
confirmed that NID2 promoter hypermethylation can be detected in
serum cfDNA with high specificity, providing novel evidence from an
epigenetic perspective in understanding its involvement in OC
development and progression (44,45).
In a recent study on OC epigenetics, Faaborg et al (46) detected HOXA9 gene methylation in 93%
(82/88) of patients with OC. In addition, Singh et al
(11) developed a novel epigenetic
panel (combining HOXA9 and HIC1 methylation) achieving 88.9%
sensitivity with an AUC value of 0.950. The combined detection
model assessed in the present study demonstrated comparable
performance to these extensively researched epigenetic panels, with
a sensitivity of 86.1%, a specificity of 97.3% and an AUC value of
0.971.
Enhancing early screening and diagnosis of patients
with OC may improve clinical outcomes and decrease mortality rates.
Liquid biopsy has emerged as a viable option for broad early
screening, with higher patient acceptability compared with tissue
biopsy due to its minimally invasive nature (47,48).
Previous studies have indicated that cfDNA methylation assay
outcomes are typically consistent with tumor tissue methylation
assay results and encompass a notable portion of solid tumor
genetic data, rendering the findings of the present study
clinically applicable (4,49,50).
In addition, cfDNA methylation could potentially associate with
treatment outcomes and prognosis in patients (51). In a previous study analyzing data
from 2,636 participants across 15 studies, Kalachand et al
(52) observed that BRCA1
methylation displayed similar clinicopathological characteristics
to BRCA1 mutations in patients with OC. However, the predictive
capability of BRCA1 methylation for platinum sensitivity and
survival prognosis remains to be elucidated. Furthermore, Tserpeli
et al (53) observed that
the predictive capacity for platinum resistance in OC could be
evaluated via Schlafen family member 11 methylation, an aspect that
warrants further exploration.
To the best of our knowledge, the novelty of the
present study is reflected in the evaluation of NID2 promoter
methylation in serum cfDNA from patients with OC, which
demonstrated its complementary diagnostic utility to CDO1, and in
the development of a clinically applicable multi-marker model that
combines these methylation markers with CA125. Several limitations
of the present study warrant consideration. Firstly, the present
study predominantly comprised patients with advanced-stage OC,
necessitating further prospective studies to evaluate the
predictive potential of NID2 and CDO1 in early-stage OC. The
clinical translation of the present findings warrants validation
through multi-center collaborations that recruit larger cohorts of
patients with FIGO stage I–II OC alongside comparative groups,
including benign ovarian lesions and high-risk populations, with
longitudinal monitoring of NID2 and CDO1 methylation dynamics to
establish their early-warning potential. Furthermore, the
methylation cut-off values determined by the Youden index in the
present study require prospective validation in broader populations
to confirm generalizability. Another key limitation lies in the
suboptimal specificity of CDO1 methylation, necessitating
systematic characterization of its methylation kinetics in
non-malignant conditions to establish disease-specific thresholds.
Integrating these thresholds with traditional imaging modalities
could optimize diagnostic accuracy, targeting the high sensitivity
of CDO1 for primary screening. Furthermore, its utility in
recurrence monitoring warrants investigation.
In conclusion, the present study indicated that the
methylation status of NID2 and CDO1 may function as serological
biomarkers for the detection and identification of OC and could
have potential clinical significance in the future.
Supplementary Material
Supporting Data
Supporting Data
Acknowledgements
Not applicable.
Funding
The present study was partially supported by the Hangzhou
Municipal Science and Technology Bureau Guided Projects, China
(grant no. 20220919Y048).
Availability of data and materials
The data generated in the present study may be
requested from the corresponding author.
Authors' contributions
MJ conceived the present study. SW carried out the
design of the present study. SW, YC and JJ performed the
experiments. JJ and QH collected clinical samples. SW, YC, QH and
XH analyzed the data and wrote the manuscript. SW and MJ cconfirm
the authenticity of all the raw data. MJ and XH revised the
manuscript. All authors read and approved the final manuscript.
Ethics approval and consent to
participate
The present study was approved by the Institutional
Review Board of Hangzhou Cancer Hospital (approval no.
HZCH-2023-006; Hangzhou, China). Written informed consent to
participate was obtained from all individual participants included
in the present study prior to their enrolment. The present study
was conducted in accordance with the principles of the Declaration
of Helsinki.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Bray F, Laversanne M, Sung H, Ferlay J,
Siegel RL, Soerjomataram I and Jemal A: Global cancer statistics
2022: GLOBOCAN estimates of incidence and mortality worldwide for
36 cancers in 185 countries. CA Cancer J Clin. 74:229–263.
2024.PubMed/NCBI
|
|
2
|
Konstantinopoulos PA and Matulonis UA:
Clinical and translational advances in ovarian cancer therapy. Nat
Cancer. 4:1239–1257. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Armstrong DK, Alvarez RD, Backes FJ,
Bakkum-Gamez JN, Barroilhet L, Behbakht K, Berchuck A, Chen LM,
Chitiyo VC, Cristea M, et al: NCCN Guidelines® Insights:
Ovarian cancer, version 3.2022. J Natl Compr Canc Netw. 20:972–980.
2022. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Zhang R, Siu MKY, Ngan HYS and Chan KKL:
Molecular biomarkers for the early detection of ovarian cancer. Int
J Mol Sci. 23:120412022. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Einama T, Yamagishi Y, Takihata Y, Suzuki
T, Yamasaki T, Hirose Y, Kobayashi K, Yonamine N, Fujinuma I,
Tsunenari T, et al: Co-expression of mesothelin and CA125/MUC16 is
a prognostic factor for breast cancer, especially in luminal-type
breast cancer patients. Biomark Res. 9:782021. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Zheng L, Shan L and Cai F: Uterine
adenomyotic cyst with markedly elevated serum CA19-9 and CA125
levels: A case report. Exp Ther Med. 24:6652022. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Saad HM, Tourky GF, Al-Kuraishy HM,
Al-Gareeb AI, Khattab AM, Elmasry SA, Alsayegh AA, Hakami ZH,
Alsulimani A, Sabatier JM, et al: The potential role of MUC16
(CA125) biomarker in lung cancer: A magic biomarker but with
adversity. Diagnostics (Basel). 12:29852022. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Mattei AL, Bailly N and Meissner A: DNA
methylation: A historical perspective. Trends Genet. 38:676–707.
2022. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Lee AV, Nestler KA and Chiappinelli KB:
Therapeutic targeting of DNA methylation alterations in cancer.
Pharmacol Ther. 258:1086402024. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Cristalli C and Scotlandi K: Targeting DNA
methylation machinery in pediatric solid tumors. Cells.
13:12092024. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Singh A, Gupta S, Badarukhiya JA and
Sachan M: Detection of aberrant methylation of HOXA9 and HIC1
through multiplex MethyLight assay in serum DNA for the early
detection of epithelial ovarian cancer. Int J Cancer.
147:1740–1752. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Liu Y, Liu H, Ye H, Jiang M, Chen X, Song
G, Ji H, Wang ZW and Zhu X: Methylation of BRD4 by PRMT1 regulates
BRD4 phosphorylation and promotes ovarian cancer invasion. Cell
Death Dis. 14:6242023. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Liu MC, Oxnard GR, Klein EA, Swanton C and
Seiden MV; CCGA Consortium, : Sensitive and specific multi-cancer
detection and localization using methylation signatures in
cell-free DNA. Ann Oncol. 31:745–759. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Pereira BA, Ritchie S, Chambers CR, Gordon
KA, Magenau A, Murphy KJ, Nobis M, Tyma VM, Liew YF, Lucas MC, et
al: Temporally resolved proteomics identifies nidogen-2 as a
cotarget in pancreatic cancer that modulates fibrosis and therapy
response. Sci Adv. 10:eadl11972024. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Chen M, Zhu JY, Mu WJ and Guo L: Cysteine
dioxygenase type 1 (CDO1): Its functional role in physiological and
pathophysiological processes. Genes Dis. 10:877–890. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Chettiankandy TJ, Sachdev SS, Khandekar
SP, Dive A, Nagpal D and Tupkari JV: Role of Nidogen-2 in diagnosis
and prognosis of head and neck squamous cell carcinoma: A
systematic review. J Oral Maxillofac Pathol. 26:382–388. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Wang J, Zhao Y, Xu H, Ma J, Liang F, Zou Q
and Lin F: Silencing NID2 by DNA hypermethylation promotes lung
cancer. Pathol Oncol Res. 26:801–811. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Chen X and Poetsch A: The role of Cdo1 in
ferroptosis and apoptosis in cancer. Biomedicines. 12:9182024.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Yang J, Sun L, Liu XY, Huang C, Peng J,
Zeng X, Zheng H, Cen W, Xu YX, Zhu W, et al: Targeted demethylation
of the CDO1 promoter based on CRISPR system inhibits the malignant
potential of breast cancer cells. Clin Transl Med. 13:e14232023.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Park SJ, Kang D, Lee M, Lee SY, Park YG,
Oh T, Jang S, Hwang WJ, Kwon SJ, An S, et al: Combination analysis
of PCDHGA12 and CDO1 DNA methylation in bronchial washing fluid for
lung cancer diagnosis. J Korean Med Sci. 39:e282024. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Bhatla N, Aoki D, Sharma DN and
Sankaranarayanan R: Cancer of the cervix uteri: 2021 update. Int J
Gynaecol Obstet. 155 (Suppl 1):S28–S44. 2021. View Article : Google Scholar
|
|
22
|
Oken MM, Creech RH, Tormey DC, Horton J,
Davis TE, McFadden ET and Carbone PP: Toxicity and response
criteria of the Eastern Cooperative Oncology Group. Am J Clin
Oncol. 5:649–656. 1982. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Heidenreich PA, Bozkurt B, Aguilar D,
Allen LA, Byun JJ, Colvin MM, Deswal A, Drazner MH, Dunlay SM,
Evers LR, et al: 2022 AHA/ACC/HFSA guideline for the management of
heart failure: Executive summary: A report of the American college
of cardiology/American heart association joint committee on
clinical practice guidelines. Circulation. 145:e876–e894. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−Delta Delta C(T)) Method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Fu M, Deng F, Chen J, Fu L, Lei J, Xu T,
Chen Y, Zhou J, Gao Q and Ding H: Current data and future
perspectives on DNA methylation in ovarian cancer (Review). Int J
Oncol. 64:622024. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Xu X, Zhuang X, Yu H, Li P, Li X, Lin H,
Teoh JP, Chen Y, Yang Y, Cheng Y, et al: FSH induces EMT in ovarian
cancer via ALKBH5-regulated Snail m6A demethylation. Theranostics.
14:2151–2166. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Duan C, Yan Z, Wu C, Zhou X and Bao W: DNA
methylation characteristics associated with chemotherapy resistance
in epithelial ovarian cancer. Heliyon. 10:e272122024. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Widschwendter M, Zikan M, Wahl B,
Lempiäinen H, Paprotka T, Evans I, Jones A, Ghazali S, Reisel D,
Eichner J, et al: The potential of circulating tumor DNA
methylation analysis for the early detection and management of
ovarian cancer. Genome Med. 9:1162017. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Herzog C, Jones A, Evans I, Reisel D,
Olaitan A, Doufekas K, MacDonald N, Rådestad AF, Gemzell-Danielsson
K, Zikan M, et al: Plasma cell-free DNA methylation analysis for
ovarian cancer detection: Analysis of samples from a case-control
study and an ovarian cancer screening trial. Int J Cancer.
154:679–691. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Terp SK, Stoico MP, Dybkær K and Pedersen
IS: Early diagnosis of ovarian cancer based on methylation profiles
in peripheral blood cell-free DNA: A systematic review. Clin
Epigenetics. 15:242023. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Feng LY, Yan BB, Huang YZ and Li L:
Abnormal methylation characteristics predict chemoresistance and
poor prognosis in advanced high-grade serous ovarian cancer. Clin
Epigenetics. 13:1412021. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Panagopoulou M, Panou T, Gkountakos A,
Tarapatzi G, Karaglani M, Tsamardinos I and Chatzaki E: BRCA1 &
BRCA2 methylation as a prognostic and predictive biomarker in
cancer: Implementation in liquid biopsy in the era of precision
medicine. Clin Epigenetics. 16:1782024. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Zhang M, Cheng S, Jin Y, Zhao Y and Wang
Y: Roles of CA125 in diagnosis, prediction, and oncogenesis of
ovarian cancer. Biochim Biophys Acta Rev Cancer. 1875:1885032021.
View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Li G, Zhang Y, Li K, Liu X, Lu Y, Zhang Z,
Liu Z, Wu Y, Liu F, Huang H, et al: Transformer-based AI technology
improves early ovarian cancer diagnosis using cfDNA methylation
markers. Cell Rep Med. 5:1016662024. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Chen C, Zhu Y, Zhang H and Xiao L:
Prognostic effects of RASSF1A, BRCA1, APC, and p16 promoter
methylation in ovarian cancer: A meta-analysis. Gynecol Obstet
Invest. 89:363–375. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Kong C and Fu T: Value of methylation
markers in colorectal cancer (Review). Oncol Rep. 46:1772021.
View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Wang R, Yu X, Ye H, Ao M, Xi M and Hou M:
LncRNA FAM83H-AS1 inhibits ferroptosis of endometrial cancer by
promoting DNMT1-mediated CDO1 promoter hypermethylation. J Biol
Chem. 300:1076802024. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Yan Y, Xu Z, Liu F, Jia Y, Chu Q and Yuan
X: Diagnostic value of CDO1 promoter methylation in lung cancer via
liquid biopsy: A systematic review and meta-analysis. Front Biosci
(Landmark Ed). 30:439872025. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Wever BMM, Schaafsma M, Bleeker MCG, van
den Burgt Y, van den Helder R, Lok CAR, Dijk F, van der Pol Y,
Mouliere F, Moldovan N, et al: Molecular analysis for ovarian
cancer detection in patient-friendly samples. Commun Med (Lond).
4:882024. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Ulazzi L, Sabbioni S, Miotto E, Veronese
A, Angusti A, Gafà R, Manfredini S, Farinati F, Sasaki T, Lanza G
and Negrini M: Nidogen 1 and 2 gene promoters are aberrantly
methylated in human gastrointestinal cancer. Mol Cancer. 6:172007.
View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Rapado-González Ó, López-Cedrún JL,
López-López R, Rodríguez-Ces AM and Suárez-Cunqueiro MM: Saliva
gene promoter hypermethylation as a biomarker in oral cancer. J
Clin Med. 10:19312021. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Chai AW, Cheung AK, Dai W, Ko JM, Ip JC,
Chan KW, Kwong DL, Ng WT, Lee AW, Ngan RK, et al:
Metastasis-suppressing NID2, an epigenetically-silenced gene, in
the pathogenesis of nasopharyngeal carcinoma and esophageal
squamous cell carcinoma. Oncotarget. 7:78859–78871. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Kwiatkowska A, Krawczyk D and Kułak K: Can
Nidogen-1 and Nidogen-2 improve our preoperative cancer detection
rate? J Pre Clin Clin Res. 15:80–86. 2021. View Article : Google Scholar
|
|
44
|
Kuk C, Gunawardana CG, Soosaipillai A,
Kobayashi H, Li L, Zheng Y and Diamandis EP: Nidogen-2: A new serum
biomarker for ovarian cancer. Clin Biochem. 43:355–361. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Kraus FB, Xu M, Kumar R, Veillard I,
Manning WB, Bond TS, Kim E, Matoba Y, Azimi M, Czogalla B, et al:
Abstract B086: Anchored in Resistance: Overcoming Nidogen-2 (NID2)
- mediated Immunoresistance in Ovarian Cancer with Ras and Bcl-xl
Inhibition. Mol Cancer Ther. 24 (10_Supplement):B0862025.
View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Faaborg L, Jakobsen A, Waldstrøm M,
Petersen CB, Andersen RF and Steffensen KD: HOXA9-methylated DNA as
a diagnostic biomarker of ovarian malignancy. Biomark Med.
15:1309–1317. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Li W, Liu JB, Hou LK, Yu F, Zhang J, Wu W,
Tang XM, Sun F, Lu HM, Deng J, et al: Liquid biopsy in lung cancer:
Significance in diagnostics, prediction, and treatment monitoring.
Mol Cancer. 21:252022. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Nikanjam M, Kato S and Kurzrock R: Liquid
biopsy: Current technology and clinical applications. J Hematol
Oncol. 15:1312022. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Kim SY, Jeong S, Lee W, Jeon Y, Kim YJ,
Park S, Lee D, Go D, Song SH, Lee S, et al: Cancer signature
ensemble integrating cfDNA methylation, copy number, and
fragmentation facilitates multi-cancer early detection. Exp Mol
Med. 55:2445–2460. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Luo H, Wei W, Ye Z, Zheng J and Xu RH:
Liquid biopsy of methylation biomarkers in cell-free DNA. Trends
Mol Med. 27:482–500. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Yang X, Liu Q, Guo Z, Yang X, Li K, Han B,
Zhang M, Sun M, Huang L, Cai G and Wu Y: Promoter profiles in
plasma CfDNA exhibits a potential utility of predicting the
efficacy of neoadjuvant chemotherapy in breast cancer patients.
Breast Cancer Res. 26:1122024. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Kalachand RD, Stordal B, Madden S,
Chandler B, Cunningham J, Goode EL, Ruscito I, Braicu EI, Sehouli
J, Ignatov A, et al: BRCA1 promoter methylation and clinical
outcomes in ovarian cancer: An individual patient data
meta-analysis. J Natl Cancer Inst. 112:1190–1203. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Tserpeli V, Stergiopoulou D, Londra D,
Giannopoulou L, Buderath P, Balgkouranidou I, Xenidis N, Grech C,
Obermayr E, Zeillinger R, et al: Prognostic significance of SLFN11
methylation in plasma cell-free DNA in advanced high-grade serous
ovarian cancer. Cancers (Basel). 14:42021. View Article : Google Scholar : PubMed/NCBI
|