Introduction
Lung cancer remains a life-threatening malignancy
worldwide, with >2 million new cases and ~1.76 million
mortalities reported annually (1).
Non-small cell lung cancer (NSCLC) is the most predominant subtype,
accounting for 85% of all annual lung cancer diagnoses (2). Despite advancements in surgical
techniques, precision treatment approaches and immunotherapeutic
interventions, the 5-year survival rate of patients with NSCLC has
exhibited limited improvement (3,4).
Therefore, it is necessary to identify novel biomarkers and develop
targeted pharmacological interventions to improve the prognosis of
NSCLC (5,6).
A defining characteristic of malignant tumors is
reprogrammed energy metabolism, which facilitates tumor development
and progression. However, the precise underlying molecular
mechanisms warrant further investigation (7,8). The
Warburg effect, characterized by enhanced lactate production
instead of mitochondrial oxidation under oxygen-rich conditions,
represents a fundamental metabolic adaptation in cancer (9,10).
This glycolytic pathway serves a dual purpose, as it immediately
generates energy substrates while producing key biosynthetic
precursors, such as glucose-6-phosphate, pyruvate derivatives and
lactate, for macromolecule synthesis. Notably, the lactate
accumulated in the tumor microenvironment induces extracellular
acidification, creating favorable conditions for the metastatic
spread and immune evasion of tumor cells (11–13).
Studies have shown that lactate can impair immune cell function. In
particular, it can suppress the cytotoxic function of natural
killer cells and T lymphocytes, facilitating tumor immune evasion
by impairing immune surveillance mechanisms (14–16).
In addition, lactate can promote the proliferation and
differentiation of regulatory T cells, inhibiting antitumor immune
responses (17). A number of
studies have shown that targeting the glycolytic pathway in cancer
cells is a promising therapeutic strategy. For example,
phosphoglycerate mutase 1 (PGAM1), a key glycolytic enzyme, has
emerged as a key therapeutic target for cancer, and marked
advancements have been made in designing PGAM1 inhibitors (18,19).
N6-methyladenosine (m6A)
modification, characterized by the addition of a methyl group to
the sixth nitrogen of adenine in RNA molecules, is a prevalent
post-transcriptional modification in eukaryotic organisms (20,21).
This dynamic modification influences the stability, localization
and translational efficiency of RNA through enzymatic regulation.
m6A modification is a reversible process regulated by
three functional protein groups, namely methyltransferases
(writers), demethylases (erasers) and methyl recognition proteins
(readers). The methyltransferase complex involved in m6A
modification is composed of core catalytic subunits such as
methyltransferase (METTL)-3, METTL5 and Wilms' tumor 1-associating
protein (WTAP) and auxiliary components such as vir-like M6A
methyltransferase associated (VIRMA), RNA binding motif protein 15
(RBM15) and METTL16 (22–24). Fat mass and obesity-associated
protein and α-ketoglutarate-dependent dioxygenase homolog 5
function as key m6A erasers (demethylases) by catalyzing
the removal of methyl groups from modified nucleotides (25,26).
YTH protein family members, including YTH
N6-methyladenosine RNA binding protein (YTHD)-F1, YTHDF2
and YTHDC2, serve as primary m6A readers that modulate
RNA stability and translational efficiency through selective
binding to methylated transcripts (27–29).
m6A modification serves as an important
regulatory mechanism in cancer initiation and progression. In
gastric cancer, this epigenetic modification accelerates malignant
progression by stimulating key oncogenic pathways, such as the Wnt
and PI3K/AKT/mTOR pathways (30–32).
METTL3 knockout can markedly inhibit the proliferation and
migration of oxaliplatin-resistant gastric cancer cells and induce
their apoptosis, thereby enhancing oxaliplatin sensitivity. These
findings indicate that m6A modification may affect
chemotherapy resistance in gastric cancer (33). Therefore, small-molecule inhibitors
targeting m6A modification-related enzymes are promising
therapeutic agents for cancer (34).
METTL5 is an emerging RNA methyltransferase that
catalyzes m6A modification of target mRNAs, often in
complex with regulatory proteins such as WTAP (35). Dysregulation of METTL5 has been
associated with tumorigenesis across numerous cancer types,
highlighting its context-dependent oncogenic roles. For example, in
liver hepatocellular carcinoma (LIHC), the upregulation of METTL5
enhances the stability of c-Myc, which in turn activates glycolytic
genes, leading to abnormal glucose metabolism and tumor growth
(36). In renal clear cell
carcinoma (KIRC), METTL5 participates in regulation of the tumor
immune microenvironment and may promote tumor progression by
facilitating the infiltration of immunosuppressive cells (37). High METTL5 expression has been shown
to be negatively associated with the prognosis of patients with
stomach adenocarcinoma. The mechanism involves increased stability
of nuclear factor (erythroid-derived 2)-like 2 mRNA, inhibition of
Fe2+ accumulation and ferroptosis and therefore,
suppression of the antitumor immunity mediated by peripheral blood
mononuclear cells (38). However,
the functional roles and regulatory mechanisms of METTL5 in NSCLC
remain poorly understood. In addition, to the best of our
knowledge, the mechanisms by which METTL5-mediated m6A
methylation regulates PGAM1 mRNA expression in NSCLC remain
unexplored. The potential interplay among METTL5, PGAM1 and glucose
transporter type 1 (GLUT1), another key protein in metabolic
adaptation (39,40), has not been elucidated. Notably, to
the best of our knowledge, the clinical relevance of this
regulatory axis and its impact on patient prognosis remain
unexplored.
Given these gaps, the present study aimed to
determine whether METTL5 is expressed in NSCLC and assess its
prognostic importance, investigate the functional role of METTL5 in
regulating glycolytic metabolism and tumor progression, elucidate
the molecular mechanism by which METTL5 modulates PGAM1 expression
through m6A modification, and examine the downstream
effects on GLUT1 expression and metabolic reprogramming. The
present investigation of this regulatory axis is considered to
provide novel insights for the development of precision therapeutic
strategies for NSCLC.
Materials and methods
Cell culture and cell lines
The A549 human lung adenocarcinoma (LUAD) cell line,
as well as the BEAS-2B bronchial epithelial cell line, the PC9 LUAD
cell line, the H520 lung squamous cell carcinoma cell line and the
NCI-H1299 NSCLC cell line were obtained from ZFdows Biotech Co.,
Ltd. All cell lines were authenticated through short tandem repeat
profiling, which determined the absence of Mycoplasma
contamination. Detailed specifications of all cell lines are
provided in Table I, with data
derived from the Catalogue of Somatic Mutations in Cancer (Sanger
Institute; http://cancer.sanger.ac.uk/cosmic) and the Cancer Cell
Line Encyclopedia (American Type Culture Collection; http://www.atcc.org/) databases. All cells were
cultured in DMEM/F12 (1:1 ratio; cat. no. BL1917A; Biosharp Life
Sciences) enriched with 10% FBS (cat. no. BL305A; Biosharp Life
Sciences) under standard conditions (37°C; 5% CO2) for
optimal cell proliferation.
 | Table I.Cell line characteristics. |
Table I.
Cell line characteristics.
| Cell line | Histologic
origin | Key driver
mutation(s) (COSMIC/ATCC) | TP53 status | Other recurrent
alterations | Doubling time,
h | Tumorigenicity in
mice | Typical
applications |
|---|
| A549 | Adenocarcinoma
(peripheral) | KRAS G12S | Wild-type | STK11 loss, KEAP1
mutation | 22-24 | ++ (s.c. and
orthotopic) | KRAS-mutant model;
chemo-/radio-/immuno-therapy studies |
| H1299 | Large-cell
carcinoma | None (EGFR/KRAS/ALK
WT) | Homozygous
deletion | NF1 loss, PTEN
loss | 20-22 | ++ | p53-null
background; high transfection efficiency; generescue assays |
| PC9 | Adenocarcinoma | EGFR E746_A750
deletion (exon 19) | Wild-type | BIM deletion
(subset) | 24-26 | ++ | Prototype
EGFR/TKI-sensitive line; parental strain for resistant clones |
| H520 | Squamous-cell
carcinoma | KRAS G12C + BRAF
G466V | Mutant | - | 28-32 | + | Squamous KRAS
model; immune-checkpoint studies |
| BEAS-2B | Immortalized
(non-cancerous) bronchial epithelium | None | Wild-type | SV40 large
T-antigen | 30-34 | - | Normal control for
transformation studies; chemical- or oncogene-induced
carcinogenesis models; MSC-like properties reported |
Cell transfection
METTL5- and PGAM1-overexpression plasmids were
purchased from Heyuan Liji (Shanghai) Biotechnology Co., Ltd. The
open reading frames of METTL5 and PGAM1 were cloned into the
pcDNA3.1(+) vector. Specific small interfering RNAs (siRNAs), along
with a non-targeting negative control siRNA, were obtained from
Guangzhou RiboBio Co., Ltd. For transient transfection, cells were
seeded in 6-well plates at a density of 3×105 cells per
well 24 h prior to transfection. Plasmid DNA (2 µg) or siRNA (50
nM) were transfected into cells using Lipo8000™ reagent (cat. no.
C0533; Beyotime Biotechnology) according to the manufacturer's
protocol. The transfection mixture was incubated with cells for 6 h
at 37°C before replacing with fresh complete medium. After 48 h
transfection, cell samples were collected for protein
quantification.
For stable gene knockdown, short hairpin RNA
(shRNA/sh)-YTHDF1 and sh-PGAM1 lentiviral particles were obtained
from Guangzhou RiboBio Co., Ltd. These lentiviral particles were
constructed in the pLKO.1-puro vector system, which drives shRNA
expression under the human U6 promoter and contains a puromycin
resistance gene for selection of transduced cells. A549 and PC9
cells were infected with lentiviral particles encoding either
sh-YTHDF1 or sh-PGAM1, or corresponding control shRNA, in the
presence of polybrene (8 µg/ml) to enhance infection efficiency. At
48 h post-infection, cells were selected with puromycin (2 µg/ml)
for 7–10 days to establish stable cell lines. The knockdown
efficiency was validated by western blot analysis 72 h
post-infection and prior to use in downstream assays.
Detailed information on the plasmid constructs,
siRNA sequences and shRNA sequences is provided in Tables II and III. Validation of transfection
efficiency for YTHDF1 and PGAM1 modulations in A549 and PC9 cells
is shown in Fig. S1.
 | Table II.Overexpression constructs generated
in the present study. |
Table II.
Overexpression constructs generated
in the present study.
| Plasmid | Backbone | Selection | 5′MCS | 3′MCS | Insert
(codon-optimized) | Insert length,
bp | Plasmid length,
bp |
|---|
|
pcDNA3.1(+)-PGAM1-HA | pcDNA3.1(+) | Ampicillin | NheI | BamHI | Homo sapiens
PGAM1 (NM_002629.3) | 810 | 6,213 |
|
pcDNA3.1(+)-METTL5-FLAG | pcDNA3.1(+) | Ampicillin | NheI | BamHI | Homo sapiens
METTL5 (NM_001318165.2) | 672 | 6,075 |
 | Table III.Sequences of the siRNAs and shRNAs
used in cell transfection. |
Table III.
Sequences of the siRNAs and shRNAs
used in cell transfection.
| Name | Sequence
(5′-3′) |
|---|
| si-PGAM1 |
|
|
Sense |
CCACAUCUGUAGACAUCUU |
|
Antisense |
AAGAUGUCUACAGAUGUGG |
| si-METTL5 |
|
|
Sense |
GCAUGUAUGCUCUAUACAA |
|
Antisense |
UUGUAUAGAGCAUACAUGC |
| si-YTHDF1 |
|
|
Sense |
CCGCGUCUAGUUGUUCAUGAA |
|
Antisense |
UUCAUGAACAACUAGACGCGG |
| sh-YTHDF1 |
GTTCGTTACATCAGAAGGATATCAAGAGTATCCTTCTGATGTAACGAACTTTTTT |
| sh-PGAM1 |
CCCTTCTGGAATGAAGAAATATCAAGAGTATTTCTTCATTCCAGAAGGGTTTTTT |
Reverse transcription-quantitative PCR
(RT-qPCR)
Total RNA was extracted from cells using the
SteadyPure Mag Tissue & Cell RNA Extraction Kit (cat. no.
AG21207; Hunan Accurate Bio-Medical Technology Co., Ltd.). cDNA was
synthesized using the Evo M-MLV Reverse Transcription Premix Kit
(cat. no. AG11728; Hunan Accurate Bio-Medical Technology Co., Ltd.)
according to the manufacturer's instructions. qPCR was performed
using the ChamQ Universal SYBR qPCR Master Mix (cat. no. Q711;
Vazyme Biotech Co., Ltd.), a ROX reference dye-containing SYBR
Green I-based fluorescent dye system. The reactions were performed
using an Applied Biosystems™ PCR thermocycler (Thermo Fisher
Scientific, Inc.) with three technical replicates (41). The thermal cycling conditions were
as follows: Initial denaturation at 95°C for 30 sec, followed by 40
cycles of denaturation at 95°C for 5 sec, and annealing/extension
at 60°C for 30 sec. GAPDH was used as the internal reference gene
and the relative expression levels of target genes were calculated
using the 2−ΔΔCq method. The specific oligonucleotide
sequences used for PCR are shown in Table IV.
 | Table IV.Primer sequences used in reverse
transcription-quantitative PCR. |
Table IV.
Primer sequences used in reverse
transcription-quantitative PCR.
| Name | Primer | Sequence
(5′-3′) |
|---|
| METTL5 | Forward |
TGTTAGGAGCAGGGTTGTGTG |
|
| Reverse |
AAGCACACATCACATTGAACCAT |
| PGAM1 | Forward |
ATGATGTCCCACCACCTCCGAT |
|
| Reverse |
ATCCTTCAGACTCTCACAGGAG |
| GAPDH | Forward |
ATGAGAAGTATGACAACAGCCTCA |
|
| Reverse |
GAGTCCTTCCACGATACCAAAG |
Western blotting
Total proteins were extracted from cells using RIPA
lysis buffer (cat. no. 89901; ThermoFisher Scientific, Inc.).
Protein concentration was determined using the BCA Protein Assay
Kit (cat. no. 23227; Thermo Fisher Scientific, Inc.) according to
the manufacturer's instructions. Equal amounts of protein (30 µg
per lane) were separated by 10% SDS-PAGE and subsequently
transferred onto PVDF membranes (MilliporeSigma). The membranes
were blocked with 5% non-fat dry milk (cat. no. P0216; Beyotime
Biotechnology) in TBST containing 0.1% Tween-20 (cat. no. ST671;
Beyotime Biotechnology) for 1 h at room temperature. The membranes
were incubated with specific primary antibodies at 4°C overnight.
Subsequently, the membranes were incubated with HRP-conjugated goat
anti-rabbit IgG (H+L) secondary antibody (1:5,000; cat. no.
SA00001-2; Proteintech Group, Inc.) at room temperature for 1 h.
Protein bands were visualized using sensitive ECL detection kit
(cat. no. PK10002; Proteintech Group, Inc.) and the band intensity
was semi-quantified using ImageJ software (version 1.54; National
Institutes of Health). The experiment was performed using three
independent biological replicates (n=3), with β-actin serving as
the loading control. The following primary antibodies were used for
western blotting: Anti-METTL5 (1:1,000; cat. no. CL488-16791;
Proteintech Group, Inc.), anti-PGAM1 (1:1,000; cat. no. 16126-1-AP;
Proteintech Group, Inc.), anti-YTHDF1 (1:1,000; cat. no.
17479-1-AP; Proteintech Group, Inc.), anti-GLUT1 (1:1,000; cat. no.
21829-1-AP; Proteintech Group, Inc.) and anti-β-actin (1:1,000;
cat. no. 60008-1-Ig; Proteintech Group, Inc.).
Colony formation assay
Cells were seeded in 6-well plates at a density of
200 cells/well and maintained under standard conditions (37°C; 5%
CO2; humidified atmosphere). After 14 days culture,
during which no additional treatments were applied, cell colonies
were fixed with 95% ethanol at room temperature for 15 min and
stained with 0.1% crystal violet at room temperature for 20 min.
The colonies were visualized using low-magnification microscopy
(CKX53; Olympus Corporation) and quantified using ImageJ software
(version 1.54; National Institutes of Health). Briefly, digital
images were captured and analyzed using the ‘Cell Counter’
(https://imagej.nih.gov/ij/plugins/cell-counter.html)
plugin to mark and count clusters containing ≥50 cells, which were
defined as viable colonies. Quantification was performed in a
blinded manner by two independent observers. Three independent
replicates (n=3) were prepared for each sample to ensure
reproducibility.
Transwell migration assay
For the migration assay, a total of 5×104
cells were cultured in serum-free DMEM/F12 (1:1) liquid medium(cat.
no. BL1917A; Biosharp Life Sciences) in the upper Transwell
chamber, whereas 600 µl of DMEM/F12 (1:1) medium supplemented with
10% FBS was added to the lower chamber. After 12–24 h of incubation
at 37°C in a humidified atmosphere containing 5% CO2,
the migrated cells were fixed with methanol at room temperature for
15 min, stained with 0.1% crystal violet for 20 min at room
temperature and observed under a phase-contrast light microscope
(CKX53; Olympus Corporation) at a magnification of ×400. The cells
were manually counted in in five randomly selected visual fields
per membrane. All experimental groups and biological replicates
were seeded with an identical initial density of 5×104
cells per well. Three independent replicates were prepared for each
sample to ensure reproducibility.
Analysis of the extracellular
acidification rate (ECAR) and oxygen consumption rate (OCR)
The cellular metabolic parameters ECAR and OCR were
measured on a Seahorse XFe96 metabolic analyzer (Agilent
Technologies, Inc.). The glycolytic rate and mitochondrial function
were assessed using the Agilent Seahorse XF Glycolytic Rate Assay
Kit (cat. no. 103344-100) and the Seahorse XF Cell Mito Stress Test
Kit (cat. no. 103015-100), respectively. The assay solution was
prepared according to the manufacturer's instructions. Briefly,
glucose, pyruvate, glutamine, oligomycin, rotenone/antimycin A and
carbonyl cyanide 4-(trifluoromethoxy)phenylhydrazone (FCCP) were
used at final concentrations of 10 mmol/l, 1 mmol/l, 2 mmol/l, 1.5
µmol/l, 0.5 µmol/l and 1 µmol/l, respectively. Injections were
automatically performed by the instrument at the following
timepoints: For the OCR assay, oligomycin was injected at 14 min,
FCCP was injected at 35 min and Rotenone/Antimycin A was injected
at 55 min. For the ECAR assay, glucose was injected at 14 min,
oligomycin was injected at 35 min and 2-DG was injected at 55 min.
Measurements were recorded after achieving thermal equilibrium and
stable pH conditions.
RNA stability assay
RNA stability was assessed by exposing cells to 5
µg/ml actinomycin D at 37°C in a humidified incubator with 5%
CO2 and extracting total RNA at specified intervals (0,
4 and 8 h). Temporal mRNA expression was quantified using
RT-qPCR.
Treatment with methylation inhibitor
3-deazaadenosine (DAA)
For functional validation, METTL5-overexpressing
cell lines were established and treated with the methylation
inhibitor 3-deazaadenosine (DAA; cat. no. HY-W013332;
MedChemExpress). Cells were seeded at a density of 1×105
cells per well in 6-well plates and allowed to adhere overnight
under standard culture conditions (37°C; 5% CO2). The
following day, cells were treated with 50 µM DAA, a concentration
previously established to effectively inhibit methyltransferase
activity without inducing significant cytotoxicity (42). DAA was dissolved in dimethyl
sulfoxide (DMSO; final concentration ≤0.1%; cat. no. HY-Y0320C;
MedChemExpress) and added directly to the culture medium. Control
cells were treated with an equivalent volume of DMSO vehicle. Cells
were incubated at 37°C in a humidified atmosphere with 5%
CO2 for 48 h prior to harvest for downstream assays.
Bioinformatics analysis
Data (TCGA-LUSC, TCGA-LUAD) used for bioinformatics
analysis were obtained from The Cancer Genome Atlas (TCGA)
repository (https://portal.gdc.cancer.gov/). The mRNA expression
patterns and prognostic importance of METTL5 and PGAM1 in NSCLC
were examined using R software (https://www.r-project.org/; version 4.3.2).
Differentially expressed genes (DEGs) were identified using the
‘limma’ package with a significance threshold of adjusted P-value
<0.05 and log2 fold change (log2FC) >1.
Gene expression correlation analysis was performed using the Gene
Expression Profiling Interactive Analysis (GEPIA) platform
(43). Potential targets of METTL5
were identified using the M6A2Target database (44).
Statistical analysis
All statistical analyses were performed using
GraphPad Prism (version 9.5; Dotmatics) and R (https://www.r-project.org/; version 4.3). Data are
presented as the mean ± standard deviation (SD). Comparisons
between two groups were performed using unpaired Student's t-tests
(for independent samples) or paired t-tests (for matched samples,
as indicated in figure legends), whereas multi-group comparisons
were performed using one-way ANOVA followed by the Bonferroni post
hoc test. Survival probabilities were calculated using Kaplan-Meier
analysis and differences in survival curves were assessed using the
log-rank test. Gene expression correlations were quantified using
Spearman's rank coefficients. All experiments were performed in
triplicate. P<0.05 was considered to indicate a statistically
significant difference.
Results
METTL5 is highly expressed in NSCLC
and contributes to tumor progression
Analysis of transcriptomic data obtained from TCGA
revealed significantly elevated METTL5 mRNA expression levels in
NSCLC (Fig. 1A). Furthermore,
METTL5 mRNA expression was significantly upregulated in colon
adenocarcinoma, stomach and esophageal carcinoma, pan-kidney
cohort, stomach adenocarcinoma, uterine corpus endometrial
carcinoma, head and neck squamous cell carcinoma, kidney renal
clear cell carcinoma, LIHC, kidney chromophobe and
cholangiocarcinoma (Fig. 1B).
RT-qPCR and western blotting revealed consistent upregulation
patterns, with lung cancer cell lines (A549, PC9 and H520)
exhibiting significantly higher METTL5 mRNA and protein expression
levels compared with normal pulmonary epithelial cells (Fig. 1D). Prognostic evaluation through
Kaplan-Meier analysis indicated that elevated METTL5 expression was
associated with reduced survival rates in patients with NSCLC
(Fig. 1C).
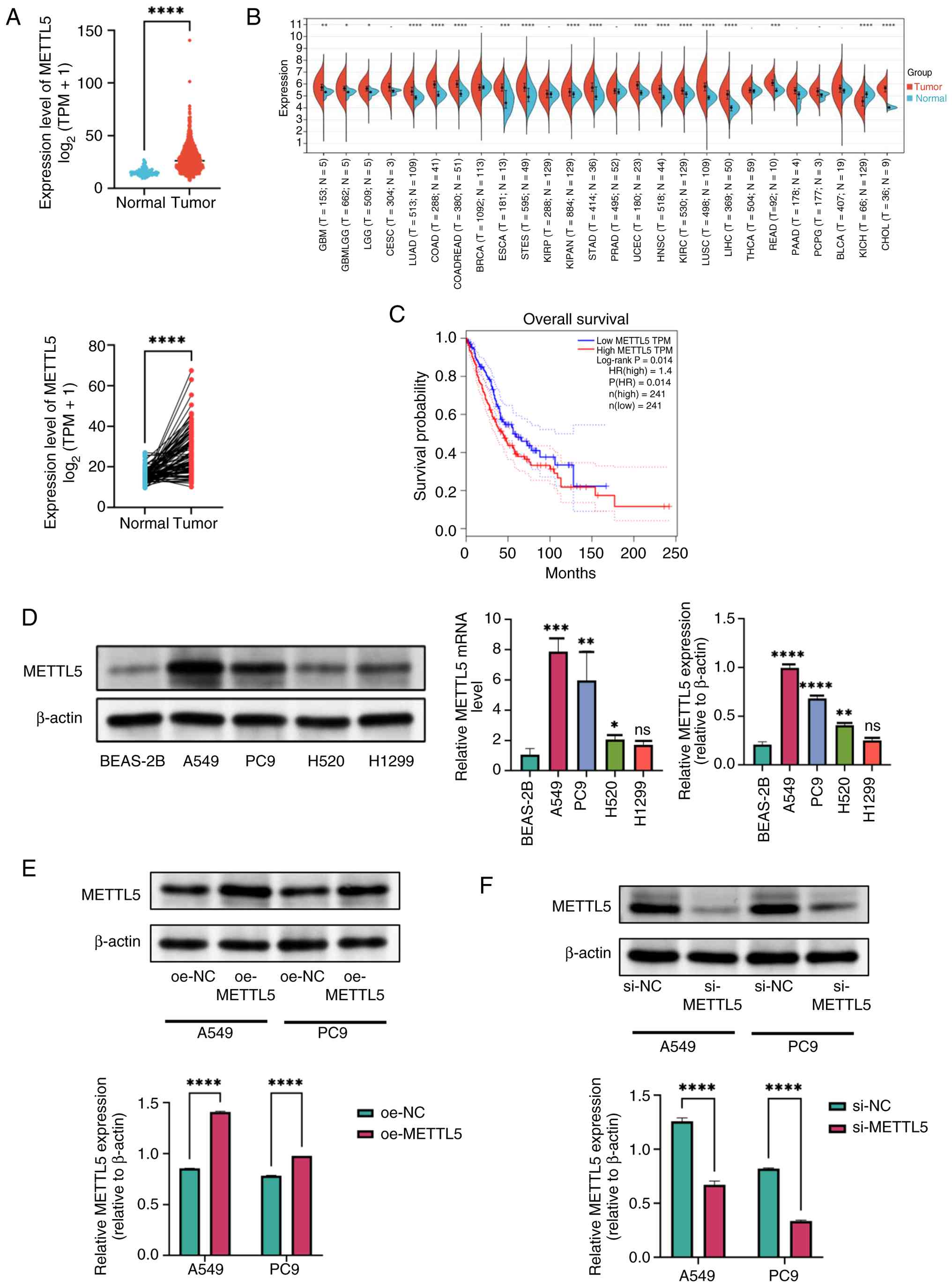 | Figure 1.METTL5 is upregulated in NSCLC
tissues. (A) Expression levels of METTL5 in NSCLC tissues and
healthy tissues (unpaired samples) and in paired samples. (B)
METTL5 expression in pan-cancer. (C) Kaplan-Meier overall survival
analysis showing the survival probability of patients with NSCLC
and high or low METTL5 expression (HR=1.4; log-rank P=0.014). (D)
mRNA levels of METTL5 measured by reverse
transcription-quantitative PCR and protein levels detected by
western blot analysis in NSCLC cell lines (A549, PC9, H520 and
NCI-H1299) compared with normal human lung epithelial cell line
(BEAS-2B). β-actin was used as a loading control. Western blot
analysis demonstrating the efficient (E) overexpression or (F)
knockdown of METTL5 in A549 and PC9 cells transfected with
si-METTL5 or si-NC. *P<0.05, **P<0.01, ***P<0.001 and
****P<0.0001. Data are presented as the mean ± SD. METTL5,
methyltransferase 5; NSCLC, non-small cell lung cancer; HR, hazard
ratio; TPM, transcripts per million; NC, negative control; oe,
overexpression; ns, not significant; si, small interfering RNA. |
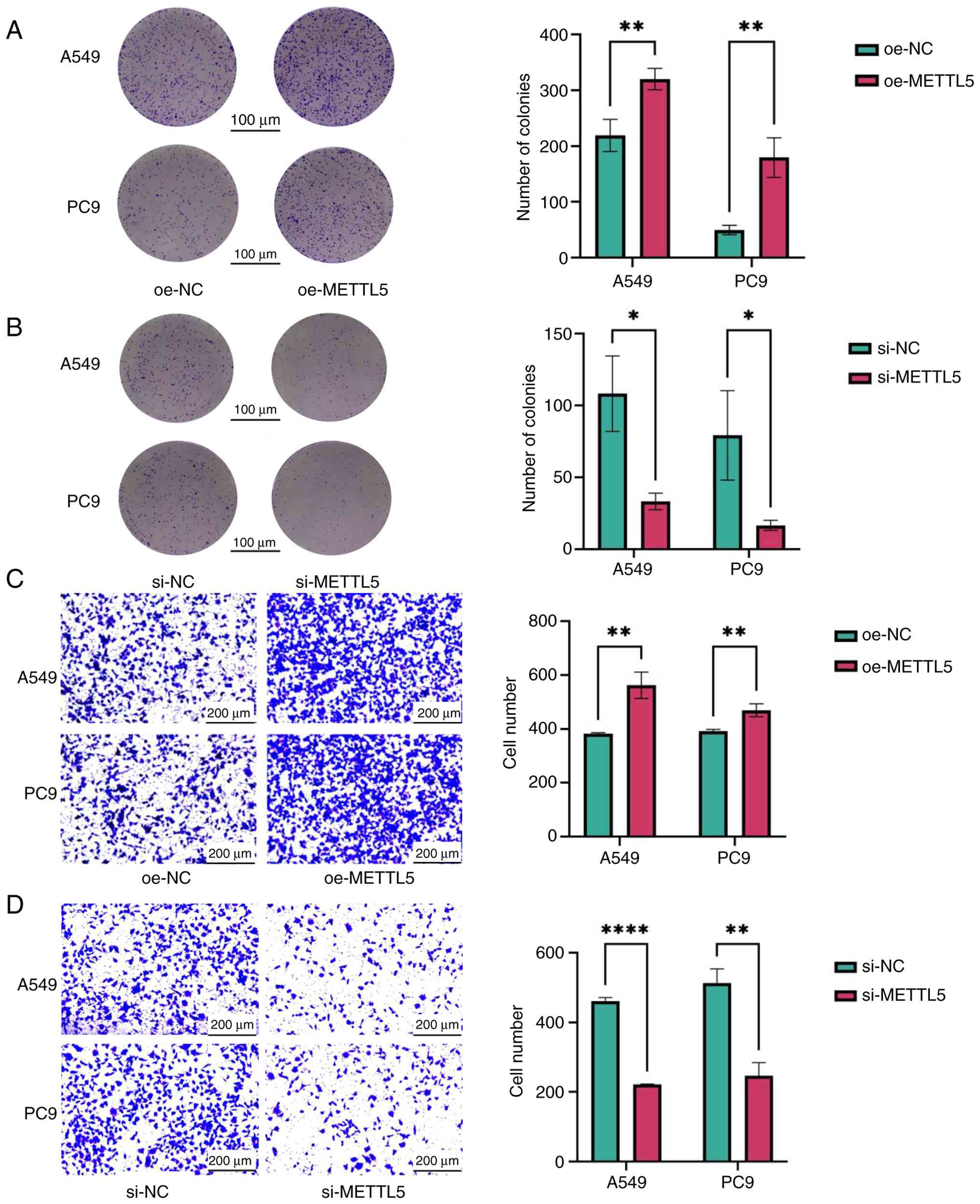
To investigate the role of METTL5 in NSCLC
pathogenesis, cell proliferation and migration were evaluated after
METTL5 knockdown or overexpression. Western blotting demonstrated
the successful knockdown or overexpression of METTL5 in
experimental models constructed through transfection (Fig. 1E and F). The colony formation assay
showed that silencing of METTL5 significantly suppressed the
proliferation of A549 and PC9 lung cancer cells, whereas ectopic
overexpression significantly enhanced cell proliferation (Fig. 2A and B). The Transwell migration
assay showed that silencing of METTL5 significantly inhibited the
migration of lung cancer cells, whereas its overexpression
significantly stimulated cell migration (Fig. 2C and D). These findings collectively
indicated that METTL5 serves an oncogenic role in NSCLC,
positioning it as a promising diagnostic biomarker and therapeutic
target.
Identification of PGAM1 as a direct
target of METTL5
To investigate the molecular mechanisms via which
METTL5 contributes to NSCLC progression, the M6A2Target database
was used to predict the targets of METTL5. PGAM1 mRNA was
identified as a key potential target (45). Subsequent analysis using the GEPIA
database showed a significant positive correlation between the mRNA
expression levels of METTL5 and PGAM1 (R=0.45;
P=5.4×10−56; Fig. 3A).
Analysis of transcriptomic data obtained from TCGA showed that the
mRNA expression levels of PGAM1 were higher in NSCLC tissues
compared with corresponding healthy tissues (Fig. 3B). For functional validation,
METTL5-overexpressing cell lines were established and treated with
the methylation inhibitor 3-deazaadenosine (DAA). Western blot
analysis revealed that overexpression of METTL5 significantly
increased PGAM1 protein levels compared with the negative control,
while treatment with the methylation inhibitor DAA reversed this
effect (Fig. 3C), indicating that
PGAM1 expression is dependent on METTL5-mediated methylation. These
findings collectively suggested that PGAM1 mRNA was methylated by
METTL5 under physiological conditions.
 | Figure 3.METTL5 regulates PGAM1 through
N6-methyladenosine methylation of PGAM1 mRNA. (A)
Correlation analysis between METTL5 and PGAM1 expression in NSCLC
tissues from The Cancer Genome Atlas database. (B) Expression
levels of PGAM1 in NSCLC tissues and normal tissues and in paired
samples. (C) Western blot analysis of PGAM1 protein levels in A549
and PC9 cells under different experimental conditions compared with
oe-NC group. (D) PGAM1 protein expression in A549 and PC9 cells
with YTHDF1 knockdown. The stability of PGAM1 mRNA was detected in
YTHDF1-konckdown compared with the shNC group through reverse
transcription-quantitative PCR at the indicated time after
actinomycin D (5 µg/ml) treatment in (E) A549 and (F) PC9 cells.
*P<0.05, **P<0.01, ***P<0.001, ****P<0.0001,
##P<0.01 and ###P<0.001. METTL5,
methyltransferase 5; PGAM1, phosphoglycerate mutase 1; TPM,
transcripts per million; NSCLC, non-small cell lung cancer; YTHDF1,
YTH N6-methyladenosine RNA binding protein 1; oe,
overexpression; NC, negative control; DAA, 3-deazaadenosine; si,
small interfering RNA; sh, short hairpin RNA. |
Identification of YTHDF1 as an
m6A reader of PGAM1
Studies have shown that m6A readers serve
an important role in methylation (46,47).
Based on the predictions of the M6A2Target database and Shi et
al (48) (GSE ID: GSE136433),
YTHDF1 was identified as an m6A reader that recognizes
and binds to methylated PGAM1 mRNA. Western blotting further
confirmed these findings, indicating that PGAM1 protein expression
was significantly lower in YTHDF1-silenced cells compared with
control cells (Fig. 3D). These
findings align with those of RNA immunoprecipitation (RIP)
sequencing in previous studies (48,49),
determining direct molecular interactions between YTHDF1 and PGAM1
mRNA. Furthermore, RNA stability analysis revealed that the
silencing of YTHDF1 accelerated PGAM1 mRNA decay rates in NSCLC
cells, indicating enhanced mRNA degradation (Fig. 3E and F) These findings collectively
indicated that YTHDF1 directly recognized m6A-modified
PGAM1 mRNA to regulate its stability and expression.
METTL5 and PGAM1 promote glycolysis by
regulating GLUT1 expression
As an important enzyme in glycolysis, PGAM1 mediates
the interconversion between 3-phosphoglycerate (3-PGA) and
2-phosphoglycerate (2-PGA), thereby enhancing cellular energy
metabolism. Through its regulatory role in maintaining equilibrium
between these metabolites, PGAM1 affects ancillary metabolic
processes. This enzymatic activity influences a number of
biosynthetic processes, not only channeling carbon flux towards
biosynthetic pathways for macromolecule production but also
maintaining cellular redox homeostasis. Furthermore, emerging
evidence indicates that PGAM1 regulates mitochondrial function and
shapes the tumor immune microenvironment, collectively driving
malignant proliferation and metastatic spread in neoplastic tissues
(50–52). In the present study, the functional
importance of the METTL5/PGAM1 axis in NSCLC pathogenesis and
energy metabolism was examined. Metabolic flux analysis showed that
PGAM1-knockdown cells exhibited a decreased ECAR but an increased
OCR (Fig. 4F and G).
 | Figure 4.Correlation analysis and metabolic
flux analysis. Correlation analysis between PGAM1 expression and
(A) GLUT1, (B) PFKP, (C) PFKFB3, (D) HK2 and (E) GLUT3 expression
in non-small cell lung cancer tissues from The Cancer Genome Atlas
database. (F) ECAR and (G) OCR profiles were measured in
PGAM1-knockdown A549 cells. The metabolic inhibitors were injected
sequentially at different time points as indicated. TPM,
transcripts per million; PGAM1, phosphoglycerate mutase 1; SLC2A1,
solute carrier family 2 member 1; GLUT, glucose transporter; PFKP,
phosphofructokinase, platelet; PFKFB,
6-phosphofructo-2-kinase/fructose-2,6-bisphosphatases; HK2,
hexokinase 2; ECAR, extracellular acidification rate; OCR, oxygen
consumption rate; sh, short hairpin RNA; Rot-AA, rotenone/antimycin
A; 2-DG, 2-deoxy-D-glucose; FCCP, carbonyl cyanide
4-(trifluoromethoxy)phenylhydrazone. |
Studies have shown that overexpression of GLUT1 and
GLUT3 can promote glucose uptake in tumor cells and lead to
chemoresistance by activating oncogenic signaling pathways, such as
the PI3K/AKT pathway (53,54). Hexokinase 2 (HK2) is a key enzyme in
glycolysis initiation, and its activation can drive tumor
metabolism toward glycolysis (55,56).
Phosphofructokinase, platelet (PFKP) and
6-phosphofructo-2-kinase/fructose-2,6-bisphosphatases (PFKFB)-3
serve important roles in tumor glycolysis, inducing high glycolytic
activity, which may lead to drug resistance (57). To determine the molecular mechanism
of METTL5/PGAM1-mediated glycolysis, correlation analysis was
performed between PGAM1 expression and GLUT1, GLUT3, HK2, PFKP or
PFKFB3 expression (Fig. 4A-E). The
correlation between PGAM1 and GLUT1 expression exhibited the
highest significance (P=4.12ⅹ10−183; R=0.6). This
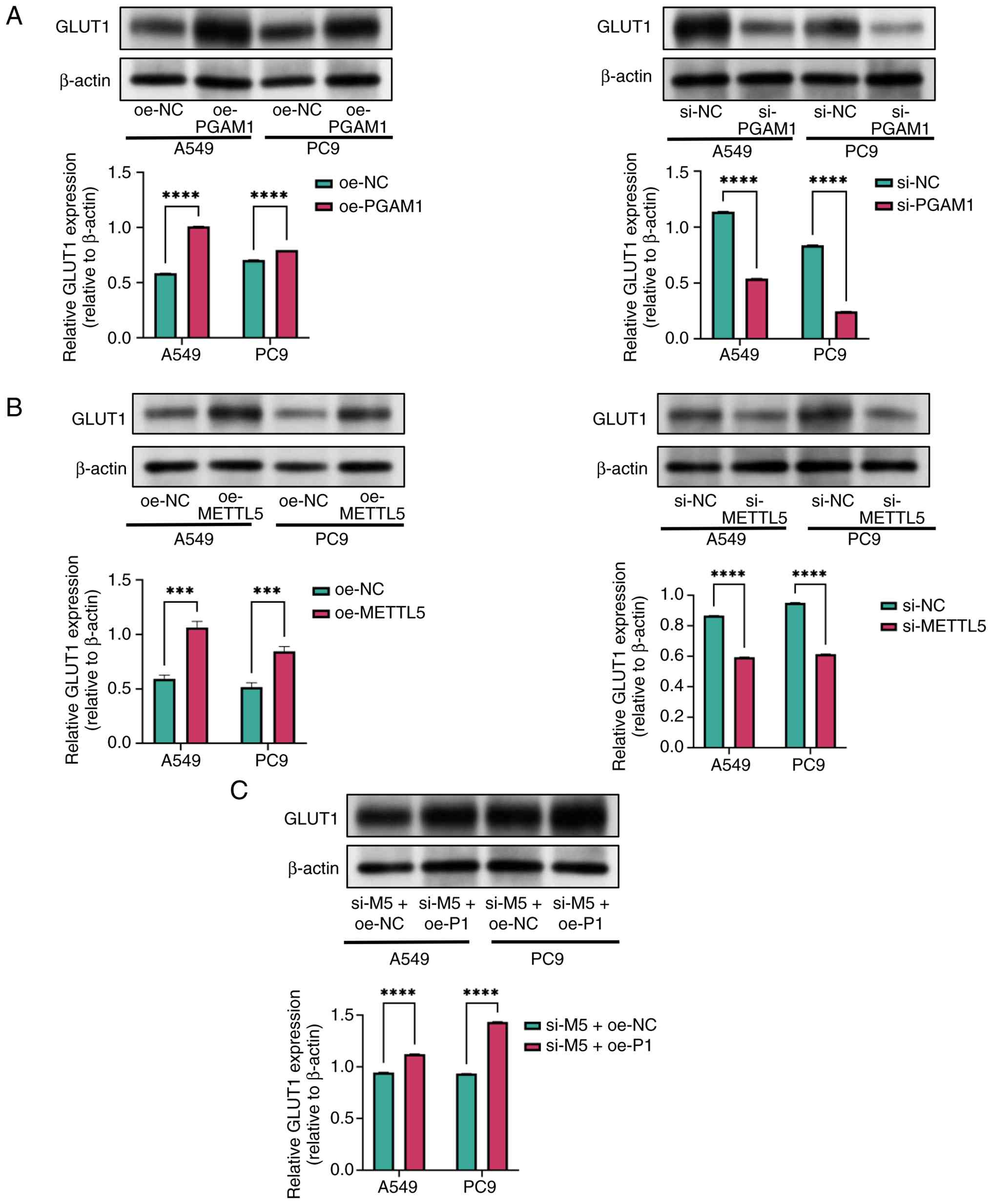
finding was validated by detecting protein expression. Knockdown of
PGAM1 significantly decreased GLUT1 protein expression in NSCLC
cells, whereas its overexpression had the opposite effect (Fig. 5A).
Furthermore, the effects of METTL5 on GLUT1
expression were investigated. Overexpression of METTL5
significantly upregulated GLUT1 expression in NSCLC cells, whereas
its knockdown significantly downregulated GLUT1 expression
(Fig. 5B). In rescue experiments,
compared with the METTL5-knockdown group, GLUT1 protein expression
was elevated in cells with METTL5 knockdown combined with PGAM1
overexpression, implying that PGAM1 overexpression partially
reversed the downregulation of GLUT1 caused by METTL5 deficiency.
This supported the present hypothesis that PGAM1 acts downstream of
METTL5 in regulating GLUT1 expression (Fig. 5C).
Discussion
As a hallmark of malignant transformation, the
Warburg effect refers to tumor cells predominantly using glycolysis
rather than oxidative phosphorylation for energy production, even
under normoxic conditions (58).
Although the underlying mechanisms are multifaceted, studies have
indicated that this metabolic reprogramming supports the
proliferation, metastatic spread and invasive potential of tumor
cells (59–61). As a predominant RNA modification,
m6A serves an important role in regulating both coding
and non-coding RNAs. A previous study has highlighted its notable
role in carcinogenesis, particularly through the modulation of
metabolic pathways in a number of malignancies (62). For example, in colorectal carcinoma,
the methyltransferase VIRMA enhances m6A-mediated
methylation of HK2 mRNA, leading to elevated HK2 mRNA expression
and increased transcript stability. This process ultimately
promotes aerobic glycolysis in cancer cells and augments their
malignant potential (63).
Furthermore, in gastric cancer, the stability and expression levels
of heparin binding growth factor (HDGF) mRNA are enhanced through
m6A modification mediated by METTL3, followed by its
interaction with insulin-like growth factor 2 mRNA-binding protein
3. Nuclear HDGF binds to the promoter regions of GLUT4 and enolase
2, leading to elevated levels of glycolytic enzymes, thereby
stimulating glycolysis, tumor growth and hepatic metastasis in
gastric cancer (64). Yang et
al (65) demonstrated that
elevated hepatitis B virus X-interacting protein (HBXIP) expression
enhanced glycolytic activity in HCC cells, thereby increasing their
malignant potential. This effect was mediated through HBXIP-induced
upregulation of METTL3. In METTL3-overexpressing hepatocellular
carcinoma (HCC) cells, elevated m6A methylation of
hypoxia inducible factor 1-α was observed, and this led to the
activation of downstream glycolytic enzymes and a corresponding
increase in the invasiveness of HCC cells (65).
METTL5 is an 18S ribosomal RNA (rRNA)-specific
m6A methyltransferase. Its main function is to regulate
ribosomal translation by catalyzing m6A modification at
the A1832 site in 18S rRNA (35).
METTL5 has been reported to be markedly upregulated in a number of
cancer types, including breast cancer and HCC (36,66,67).
Furthermore, METTL5 has been shown to regulate metabolic
reprogramming. In HCC, METTL5 promotes fatty acid β-oxidation and
the Warburg effect by targeting acyl-coA synthetase long chain
family member 4 (ACSL4). ACSL4 encodes a key metabolic enzyme that
activates fatty acids by catalyzing their conversion to acyl-CoA
esters, thereby supporting the energy demand of tumor cells
(66). The present study
demonstrated that METTL5 enhanced glycolytic activity and tumor
cell proliferation in NSCLC, thereby promoting NSCLC
progression.
PGAM1 serves as a key metabolic catalyst in
facilitating the interconversion between 3-PGA and 2-PGA during
glycolysis. By regulating the flux of these intermediates, PGAM1
supports the generation of energy and provides essential precursors
for serine synthesis, the pentose phosphate pathway and
phospholipid metabolism (68). In
certain tumors, PGAM1 not only exhibits enzymatic activity but also
participates in enzyme-independent metabolic regulation. For
example, in breast cancer, PGAM1 interacts with α-smooth muscle
actin through direct binding, enhancing the migration and
metastatic spread of cancer cells through non-catalytic mechanisms
involving protein-protein interfaces. This interaction is
independent of PGAM1 metabolic activity; the enzymatically inactive
H186R mutant retains ACTA2 binding, whereas a mutant lacking amino
acids 201–210 fails to interact despite maintaining full enzymatic
function. By acting as a structural adaptor, PGAM1 directly
modulates actin filament assembly and cytoskeletal dynamics,
thereby promoting cell motility (51). PGAM1 has emerged as a promising
therapeutic target for cancer and marked advancements have been
made in designing PGAM1 inhibitors. For example, HKB99 is a novel
allosteric PGAM1-targeted compound with potent suppressive effects
on both tumor progression and metastatic spread in NSCLC. In
addition, it has exhibited therapeutic potential in
erlotinib-resistant tumors (69).
As an important glucose transporter, GLUT1 regulates
cellular glucose uptake and frequently acts as a metabolic
bottleneck in malignant tumors. These key functions of GLUT1 are
observed in a number of malignancies. For example, GLUT1-enriched
cancer-associated fibroblasts can drive metastatic niche formation
through metabolic reprogramming in ovarian carcinoma (70). In LUAD, the long non-coding RNA
GAS6-AS1 blocks glucose metabolic reprogramming by inhibiting GLUT1
expression, thereby inhibiting tumor progression (71). In glioblastoma, GLUT1 inhibition can
reduce the excretion of lactate produced by tumor glycolysis,
thereby improving the immunosuppressive tumor microenvironment
(72). These findings collectively
indicate that the METTL5/PGAM1 signaling axis facilitates
glycolytic metabolism and tumor progression by regulating GLUT1 in
NSCLC.
The present study demonstrated that METTL5 was
markedly upregulated in NSCLC, with this upregulation being
associated with unfavorable clinical outcomes. METTL5 promoted the
proliferation, migration and glycolytic activity of NSCLC cells
in vitro, regulating the expression of PGAM1 mRNA through
m6A modification. PGAM1 mRNA could also be recognized
and bound by YTHDF1 to inhibit its degradation. PGAM1
overexpression results in the upregulation of GLUT1, thereby
enhancing glycolysis and lactate production in NSCLC cells. PGAM1
catalyzes the key reaction in glycolysis, converting 3-PGA to
2-PGA, increasing GLUT1 expression, accelerating glucose uptake
efficiency and providing sufficient substrates for cells (68). Therefore, we hypothesized that the
METTL5/PGAM1/GLUT1 axis promoted NSCLC progression by reprogramming
glucose metabolism.
Despite notable findings, the present study had
certain limitations. While the colony formation assays demonstrated
the impact of METTL5 on clonal expansion, the integration of MTT
analysis will be considered in future studies to validate these
effects across different proliferation metrics. The colony
formation assay was specifically selected to assess the
self-renewal and sustained proliferation capacities of cells, which
are important in understanding tumorigenicity. This approach
aligned with the objective of the present study to explore the role
of METTL5 in cancer stemness and metastatic potential. Furthermore,
due to the established association between clonogenicity and tumor
aggressiveness (73), the present
study prioritized this endpoint to provide mechanistic insights
into the contribution of METTL5 to NSCLC progression. Nevertheless,
it should be recognized that MTT assays may complement the present
findings by capturing shorter-term proliferation rates. Although
the biological function of METTL5 in NSCLC cells was investigated
using RT-qPCR, western blotting and cellular functional assays,
further validation in animal or clinical tissue samples is
required. While in vitro cell line experiments provide
valuable mechanistic insights, they exhibit inherent limitations in
fully recapitulating the complexity of human tumors, including
tumor heterogeneity, the tumor microenvironment and systemic
physiological regulation. In the future, the authors plan to
validate key findings in clinical NSCLC tissue samples and
establish animal models to evaluate in vivo tumorigenicity
and therapeutic potential. In addition, small-molecule inhibitors
targeting METTL5 or its downstream effectors (such as PGAM1) are
being screened in preclinical models, with the aim of identifying
potential strategies for metabolism-targeted therapy in NSCLC.
These complementary approaches will allow the gap between in
vitro observations and clinical reality to be closed, thereby
enhancing the translational importance of the present study.
Overall, the mechanistic interactions between METTL5
and PGAM1, particularly those contributing to m6A
modification, warrant further investigation. The m6A
levels of PGAM1 transcripts were not directly quantified in the
present study. Instead, inferences were drawn from protein
expression patterns after DAA treatment. In future studies,
methylated RIP sequencing may be used to directly quantify the
m6A levels of PGAM1 mRNA. To assess whether YTHDF1 is an
m6A reader of PGAM1, bioinformatics analysis and western
blotting were used to indirectly determine the association between
YTHDF1 and PGAM1. Although previous studies (48,49)
have reported the use of RIP sequencing, RNA pulldown experiments
may be used to elucidate the association between YTHDF1 and PGAM1.
In addition, the METTL5/PGAM1 axis may target proteins other than
GLUT1 to promote glycolysis. Notably, while HK2 and PFKP showed
more modest but statistically significant correlations with PGAM1
expression (R=0.37 and R=0.33, respectively) compared with GLUT1
(R=0.6), these findings suggested that HK2 and PFKP may also be
functionally associated with the METTL5/PGAM1 axis, potentially
through indirect mechanisms such as metabolic reprogramming or
transcriptional network modulation. These potential targets,
especially HK2 and PFKP, should be identified to further understand
the role of the METTL5/PGAM1 axis in NSCLC progression.
Supplementary Material
Supporting Data
Acknowledgements
Not applicable.
Funding
The present study was funded by the Changzhou High-Level Medical
Talents Training Project (grant no. 2022CZBJ069), the Changzhou Sci
& Tech Program (grant no. CZ20220025), the ‘333 Project’ of
Jiangsu Province (grant no. BRA2020157), the 333 High-Level Talent
Training Project (grant no. 2022-2) and the Science and Technology
Development Fund of Nanjing Medical University (grant no.
NMUB20250013).
Availability of data and materials
The data generated in the present study may be
requested from the corresponding author.
Authors' contributions
YS, ZG and XD conceived and designed the
experiments. Data collection and analysis was performed by KY, ML
and QW. ZG and ML also analyzed and interpreted the data and
prepared all figures for publication. KY and QW provided financial
and technical support. The first draft of the manuscript was
written by ML and ZG and revised by YS. All authors commented on
previous versions of the manuscript. All authors have read and
approved the final version of the manuscript. YS and ZG confirm the
authenticity of all the raw data.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Sung H, Ferlay J, Siegel RL, Laversanne M,
Soerjomataram I, Jemal A and Bray F: Global cancer statistics 2020:
GLOBOCAN estimates of incidence and mortality worldwide for 36
cancers in 185 countries. CA Cancer J Clin. 71:209–249.
2021.PubMed/NCBI
|
|
2
|
Molina JR, Yang P, Cassivi SD, Schild SE
and Adjei AA: Non-small cell lung cancer: Epidemiology, risk
factors, treatment, and survivorship. Mayo Clin Proc. 83:584–594.
2008. View
Article : Google Scholar : PubMed/NCBI
|
|
3
|
Lobb RJ, Visan KS, Wu LY, Norris EL,
Hastie ML, Everitt S, Yang IA, Bowman RV, Siva S, Larsen JE, et al:
An epithelial-to-mesenchymal transition induced extracellular
vesicle prognostic signature in non-small cell lung cancer. Commun
Biol. 6:682023. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Richards TB, Henley SJ, Puckett MC, Weir
HK, Huang B, Tucker TC and Allemani C: Lung cancer survival in the
United States by race and stage (2001–2009): Findings from the
CONCORD-2 study. Cancer. 123 (Suppl 24):S5079–S5099. 2017.
View Article : Google Scholar
|
|
5
|
Wang Q, Li J, Zhu J, Mao J, Duan C, Liang
X, Zhu L, Zhu M, Zhang Z, Lin F and Guo R: Genome-wide CRISPR/Cas9
screening for therapeutic targets in NSCLC carrying wild-type TP53
and receptor tyrosine kinase genes. Clin Transl Med. 12:e8822022.
View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Zhang J, Song Z, Zhang Y, Zhang C, Xue Q,
Zhang G and Tan F: Recent advances in biomarkers for predicting the
efficacy of immunotherapy in non-small cell lung cancer. Front
Immunol. 16:15548712025. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Sun L, Suo C, Li ST, Zhang H and Gao P:
Metabolic reprogramming for cancer cells and their
microenvironment: Beyond the Warburg Effect. Biochim Biophys Acta
Rev Cancer. 1870:51–66. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Yoshida GJ: Metabolic reprogramming: The
emerging concept and associated therapeutic strategies. J Exp Clin
Cancer Res. 34:1112015. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Lebelo MT, Joubert AM and Visagie MH:
Warburg effect and its role in tumourigenesis. Arch Pharm Res.
42:833–847. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Liberti MV and Locasale JW: The Warburg
effect: How does it benefit cancer cells? Trends Biochem Sci.
41:211–218. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Gao X, Zhou S, Qin Z, Li D, Zhu Y and Ma
D: Upregulation of HMGB1 in tumor-associated macrophages induced by
tumor cell-derived lactate further promotes colorectal cancer
progression. J Transl Med. 21:532023. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Jin X, Zhang N, Yan T, Wei J, Hao L, Sun
C, Zhao H and Jiang S: Lactate-mediated metabolic reprogramming of
tumor-associated macrophages: Implications for tumor progression
and therapeutic potential. Front Immunol. 16:15730392025.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Qian J, Gong ZC, Zhang YN, Wu HH, Zhao J,
Wang LT, Ye LJ, Liu D, Wang W, Kang X, et al: Lactic acid promotes
metastatic niche formation in bone metastasis of colorectal cancer.
Cell Commun Signal. 19:92021. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Gu XY, Yang JL, Lai R, Zhou ZJ, Tang D, Hu
L and Zhao LJ: Impact of lactate on immune cell function in the
tumor microenvironment: Mechanisms and therapeutic perspectives.
Front Immunol. 16:15633032025. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Watson MJ, Vignali PDA, Mullett SJ,
Overacre-Delgoffe AE, Peralta RM, Grebinoski S, Menk AV,
Rittenhouse NL, DePeaux K, Whetstone RD, et al: Metabolic support
of tumour-infiltrating regulatory T cells by lactic acid. Nature.
591:645–651. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Jin J, Yan P, Wang D, Bai L, Liang H, Zhu
X, Zhu H, Ding C, Wei H and Wang Y: Targeting lactylation
reinforces NK cell cytotoxicity within the tumor microenvironment.
Nat Immunol. 26L:1099–1112. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Gu J, Zhou J, Chen Q, Xu X, Gao J, Li X,
Shao Q, Zhou B, Zhou H, Wei S, et al: Tumor metabolite lactate
promotes tumorigenesis by modulating MOESIN lactylation and
enhancing TGF-β signaling in regulatory T cells. Cell Rep.
39:1109862022. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Xie Y, Wang M, Qiao L, Qian Y, Xu W, Sun
Q, Luo S and Li C: Photothermal-Enhanced Dual Inhibition of
Lactate/Kynurenine metabolism for promoting tumor immunotherapy.
Small Methods. 8:e23009452024. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Chen S, Xu Y, Zhuo W and Zhang L: The
emerging role of lactate in tumor microenvironment and its clinical
relevance. Cancer Lett. 590:2168372024. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Alarcón CR, Lee H, Goodarzi H, Halberg N
and Tavazoie SF: N6-methyladenosine marks primary microRNAs for
processing. Nature. 519:482–485. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Zhao BS, Roundtree IA and He C:
Post-transcriptional gene regulation by mRNA modifications. Nat Rev
Mol Cell Biol. 18:31–42. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Huang Q, Mo J, Liao Z, Chen X and Zhang B:
The RNA m6A writer WTAP in diseases: Structure, roles,
and mechanisms. Cell Death Dis. 13:8522022. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Zhou H, Yin K, Zhang Y, Tian J and Wang S:
The RNA m6A writer METTL14 in cancers: Roles, structures, and
applications. Biochim Biophys Acta Rev Cancer. 1876:1886092021.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Zeng C, Huang W, Li Y and Weng H: Roles of
METTL3 in cancer: Mechanisms and therapeutic targeting. J Hematol
Oncol. 13:1172020. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Gao Z, Zha X, Li M, Xia X and Wang S:
Insights into the m6A demethylases FTO and ALKBH5:
Structural, biological function, and inhibitor development. Cell
Biosci. 14:1082024. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Lan Q, Liu PY, Haase J, Bell JL,
Hüttelmaier S and Liu T: The critical role of RNA m6A
methylation in cancer. Cancer Res. 79:1285–1292. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Shi H, Wei J and He C: Where, When, and
How: Context-dependent functions of RNA methylation writers,
readers, and erasers. Mol Cell. 74:640–650. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Zaccara S, Ries RJ and Jaffrey SR:
Reading, writing and erasing mRNA methylation. Nat Rev Mol Cell
Biol. 20:608–624. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Wu X, Chen H, Li K, Zhang H, Li K and Tan
H: The biological function of the N6-Methyladenosine reader YTHDC2
and its role in diseases. J Transl Med. 22:4902024. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Huang H, Weng H and Chen J: m6A
Modification in coding and Non-coding RNAs: Roles and therapeutic
implications in cancer. Cancer Cell. 37:270–288. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Yi YC, Chen XY, Zhang J and Zhu JS: Novel
insights into the interplay between m6A modification and noncoding
RNAs in cancer. Mol Cancer. 19:1212020. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Zhang Y, Chen Y, Guo Q, Zhang Y and Liu A:
Fat mass and obesity-associated protein (FTO)-induced upregulation
of flotillin-2 (FLOT2) contributes to cancer aggressiveness in
diffuse large B-cell lymphoma (DLBCL) via activating the
PI3K/Akt/mTOR signal pathway. Arch Biochem Biophys. 758:1100722024.
View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Wang Y, Hong Z, Song J, Zhong P and Lin L:
METTL3 promotes drug resistance to oxaliplatin in gastric cancer
cells through DNA repair pathway. Front Pharmacol. 14:12574102023.
View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Deng LJ, Deng WQ, Fan SR, Chen MF, Qi M,
Lyu WY, Qi Q, Tiwari AK, Chen JX, Zhang DM and Chen ZS: m6A
modification: Recent advances, anticancer targeted drug discovery
and beyond. Mol Cancer. 21:522022. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
van Tran N, Ernst FGM, Hawley BR, Zorbas
C, Ulryck N, Hackert P, Bohnsack KE, Bohnsack MT, Jaffrey SR,
Graille M and Lafontaine DLJ: The human 18S rRNA m6A
methyltransferase METTL5 is stabilized by TRMT112. Nucleic Acids
Res. 47:7719–7733. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Xia P, Zhang H, Lu H, Xu K, Jiang X, Jiang
Y, Gongye X, Chen Z, Liu J, Chen X, et al: METTL5 stabilizes c-Myc
by facilitating USP5 translation to reprogram glucose metabolism
and promote hepatocellular carcinoma progression. Cancer Commun
(Lond). 43:338–364. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Zhang W, Chen Y, Zeng Z, Peng Y, Li L, Hu
N, Gao X, Cai W, Yin L, Xu Y, et al: The novel m6A writer METTL5 as
prognostic biomarker probably associating with the regulation of
immune microenvironment in kidney cancer. Heliyon. 8:e120782022.
View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Li X, Yang G, Ma L, Tang B and Tao T:
N6-methyladenosine (m6A) writer METTL5 represses the ferroptosis
and antitumor immunity of gastric cancer. Cell Death Discov.
10:4022024. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Zhou Z, Li Y, Chen S, Xie Z, Du Y, Liu Y,
Shi Y, Lin X, Zeng X, Zhao H and Chen G: GLUT1 promotes cell
proliferation via binds and stabilizes phosphorylated EGFR in lung
adenocarcinoma. Cell Commun Signal. 22:3032024. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Ren Z, Zhao J, Li S and Yuan H: Targeting
glucose transporter 1 (GLUT1) in cancer: Molecular mechanisms and
nanomedicine applications. Int J Nanomedicine. 20:11859–11879.
2025. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Lv YB, Chen C, Yu QM, Lyu L, Peng YF and
Tan XD: Synthesis and biological evaluation of novel pentanediamide
derivatives as S-adenosyl-l-homocysteine hydrolase inhibitors.
Bioorg Med Chem Lett. 72:1288802022. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Tang Z, Li C, Kang B, Gao G, Li C and
Zhang Z: GEPIA: A web server for cancer and normal gene expression
profiling and interactive analyses. Nucleic Acids Res. 45:W98–W102.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Deng S, Zhang H, Zhu K, Li X, Ye Y, Li R,
Liu X, Lin D, Zuo Z and Zheng J: M6A2Target: A comprehensive
database for targets of m6A writers, erasers and readers. Brief
Bioinform. 22:bbaa0552021. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Sepich-Poore C, Zheng Z, Schmitt E, Wen K,
Zhang ZS, Cui XL, Dai Q, Zhu AC, Zhang L, Sanchez Castillo A, et
al: The METTL5-TRMT112 N6-methyladenosine methyltransferase complex
regulates mRNA translation via 18S rRNA methylation. J Biol Chem.
298:1015902022. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
He L, Li H, Wu A, Peng Y, Shu G and Yin G:
Functions of N6-methyladenosine and its role in cancer. Mol Cancer.
18:1762019. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Fang X, Li M, Yu T, Liu G and Wang J:
Reversible N6-methyladenosine of RNA: The regulatory mechanisms on
gene expression and implications in physiology and pathology. Genes
Dis. 7:585–597. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Shi Y, Fan S, Wu M, Zuo Z, Li X, Jiang L,
Shen Q, Xu P, Zeng L, Zhou Y, et al: YTHDF1 links hypoxia
adaptation and non-small cell lung cancer progression. Nat Commun.
10:48922019. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Zhang X, Su T, Wu Y, Cai Y, Wang L, Liang
C, Zhou L, Wang S, Li XX, Peng S, et al: N6-Methyladenosine reader
YTHDF1 promotes stemness and therapeutic resistance in
hepatocellular carcinoma by enhancing NOTCH1 expression. Cancer
Res. 84:827–840. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Sun Q, Li S, Wang Y, Peng H, Zhang X,
Zheng Y, Li C, Li L, Chen R, Chen X, et al: Phosphoglyceric acid
mutase-1 contributes to oncogenic mTOR-mediated tumor growth and
confers non-small cell lung cancer patients with poor prognosis.
Cell Death Differ. 25:1160–1173. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Zhang D, Jin N, Sun W, Li X, Liu B, Xie Z,
Qu J, Xu J, Yang X, Su Y, et al: Phosphoglycerate mutase 1 promotes
cancer cell migration independent of its metabolic activity.
Oncogene. 36:2900–2909. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Luo JQ, Yang TW, Wu J, Lai HH, Zou LB,
Chen WB, Zhou XM, Lv DJ, Cen SR, Long ZN, et al: Exosomal PGAM1
promotes prostate cancer angiogenesis and metastasis by interacting
with ACTG1. Cell Death Dis. 14:5022023. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Hu W, Li F, Liang Y, Liu S, Wang S, Shen
C, Zhao Y, Wang H and Zhang Y: Glut3 overexpression improves
environmental glucose uptake and antitumor efficacy of CAR-T cells
in solid tumors. J Immunother Cancer. 13:e0105402025. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Xu M, Zhong W, Yang C, Liu M, Yuan X, Lu
T, Li D, Zhang G, Liu H, Zeng Y, et al: Tiliroside disrupted iron
homeostasis and induced ferroptosis via directly targeting
calpain-2 in pancreatic cancer cells. Phytomedicine.
127:1553922024. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Wu H, Yang J, Yang Z, Xiao Y, Liu R, Jia
J, Zhang X, Zhang Y, Fu Z, Yao Z and Lv J: Targeting the
BCKDK/BCLAF1/MYC/HK2 axis to alter aerobic glycolysis and overcome
Trametinib resistance in lung cancer. Cell Death Differ.
32:2210–2224. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Wang L, Kong L, Zhang DQ, Ye L, Nao SC,
Chan DS, Li X, Peng Y, Yang L and Wong CY: Inhibiting glycolysis
and disrupting the mitochondrial HK2-VDAC1 Protein-protein
interaction using a bifunctional Lonidamine-conjugated metal probe
for combating Triple-negative breast cancer. J Am Chem Soc.
147:14824–14836. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Zhang W, Xia M, Li J, Liu G, Sun Y, Chen X
and Zhong J: Warburg effect and lactylation in cancer: Mechanisms
for chemoresistance. Mol Med. 31:1462025. View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Bononi G, Masoni S, Di Bussolo V,
Tuccinardi T, Granchi C and Minutolo F: Historical perspective of
tumor glycolysis: A century with Otto Warburg. Semin Cancer Biol.
86:325–333. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
59
|
Chen Y, Bei J, Liu M, Huang J, Xie L,
Huang W, Cai M, Guo Y, Lin L and Zhu K: Sublethal heat
stress-induced O-GlcNAcylation coordinates the Warburg effect to
promote hepatocellular carcinoma recurrence and metastasis after
thermal ablation. Cancer Lett. 518:23–34. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
60
|
Guo Y, Li X, Sun X, Wang J, Yang X, Zhou
X, Zhou X, Liu X, Liu W, Yuan J, et al: Combined aberrant
expression of NDRG2 and LDHA predicts hepatocellular carcinoma
prognosis and mediates the anti-tumor effect of gemcitabine. Int J
Biol Sci. 15:1771–1786. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
61
|
Yadav D, Yadav A, Bhattacharya S, Dagar A,
Kumar V and Rani R: GLUT and HK: Two primary and essential key
players in tumor glycolysis. Semin Cancer Biol. 100:17–27. 2024.
View Article : Google Scholar : PubMed/NCBI
|
|
62
|
Gu J, Cao H, Chen X, Zhang XD, Thorne RF
and Liu X: RNA m6A modifications regulate crosstalk between tumor
metabolism and immunity. Wiley Interdiscip Rev RNA. 15:e18292024.
View Article : Google Scholar : PubMed/NCBI
|
|
63
|
Li Y, He L, Wang Y, Tan Y and Zhang F:
N6-methyladenosine methyltransferase KIAA1429 elevates colorectal
cancer aerobic glycolysis via HK2-dependent manner. Bioengineered.
13:11923–11932. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
64
|
Wang Q, Chen C, Ding Q, Zhao Y, Wang Z,
Chen J, Jiang Z, Zhang Y, Xu G, Zhang J, et al: METTL3-mediated m6A
modification of HDGF mRNA promotes gastric cancer progression and
has prognostic significance. Gut. 69:1193–1205. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
65
|
Yang N, Wang T, Li Q, Han F, Wang Z, Zhu R
and Zhou J: HBXIP drives metabolic reprogramming in hepatocellular
carcinoma cells via METTL3-mediated m6A modification of HIF-1α. J
Cell Physiol. 236:3863–3880. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
66
|
Peng H, Chen B, Wei W, Guo S, Han H, Yang
C, Ma J, Wang L, Peng S, Kuang M and Lin S: N6-methyladenosine
(m6A) in 18S rRNA promotes fatty acid metabolism and oncogenic
transformation. Nat Metab. 4:1041–1054. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
67
|
Rong B, Zhang Q, Wan J, Xing S, Dai R, Li
Y, Cai J, Xie J, Song Y, Chen J, et al: Ribosome 18S m6A
Methyltransferase METTL5 promotes translation initiation and breast
cancer cell growth. Cell Rep. 33:1085442020. View Article : Google Scholar : PubMed/NCBI
|
|
68
|
Yang GJ, Tao F, Zhong HJ, Yang C and Chen
J: Targeting PGAM1 in cancer: An emerging therapeutic opportunity.
Eur J Med Chem. 244:1147982022. View Article : Google Scholar : PubMed/NCBI
|
|
69
|
Huang K, Liang Q, Zhou Y, Jiang LL, Gu WM,
Luo MY, Tang YB, Wang Y, Lu W, Huang M, et al: A novel allosteric
inhibitor of phosphoglycerate Mutase 1 suppresses growth and
metastasis of non-small-cell lung cancer. Cell Metab.
30:1107–1119.e8. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
70
|
Zhang D, Li J, Xu X, Wang X, Lou Y, Zhou X
and Zhang K: CAF-derived GLUT1 and its role in modulating ovarian
cancer progression: A multi-dimensional analysis of the tumor
microenvironment. Commun Biol. 8:10202025. View Article : Google Scholar : PubMed/NCBI
|
|
71
|
Luo J, Wang H, Wang L, Wang G, Yao Y, Xie
K, Li X, Xu L, Shen Y and Ren B: lncRNA GAS6-AS1 inhibits
progression and glucose metabolism reprogramming in LUAD via
repressing E2F1-mediated transcription of GLUT1. Mol Ther Nucleic
Acids. 25:11–24. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
72
|
Li T, Xu D, Ruan Z, Zhou J, Sun W, Rao B
and Xu H: Metabolism/Immunity Dual-Regulation thermogels
potentiating immunotherapy of glioblastoma through
Lactate-excretion inhibition and PD-1/PD-L1 blockade. Adv Sci
(Weinh). 11:e23101632024. View Article : Google Scholar : PubMed/NCBI
|
|
73
|
Kim JY, Hong N, Park S, Ham SW, Kim EJ,
Kim SO, Jang J, Kim Y, Kim JK, Kim SC, et al: Jagged1 intracellular
domain/SMAD3 complex transcriptionally regulates TWIST1 to drive
glioma invasion. Cell Death Dis. 14:8222023. View Article : Google Scholar : PubMed/NCBI
|