Introduction
Gastric cancer is the fourth most common malignancy
in the world and the second leading cause of cancer-related
mortality (1). At present, surgical
resection remains the main therapeutic strategy for gastric cancer,
supplemented with perioperative chemotherapy, chemoradiotherapy
and/or immunotherapy (2–5). However, most patients are diagnosed
with advanced gastric cancer which may have progressed beyond the
curative potential of surgical operation (6,7). In
addition, previous studies have demonstrated that a considerable
proportion of patients receiving potentially curative resection
experienced recurrences which lead to unfavorable prognosis
(8,9). Adjuvant therapy, such as chemotherapy,
provides rather limited survival advantage (10,11).
These facts attest to the deficiency in the current strategies for
treating gastric cancer and the demand for novel approaches to the
management of gastric cancer.
Among various ingredients eligible for adjuvant
therapy for gastric cancer, the significance of natural products,
particularly the essence extracted from Chinese herbs are gaining
increasing attention in basic and clinical research (12). β-elemene
(1-methyl-1-vinyl-2,4-diisopropenyl-cyclohexane) is a novel
anticancer agent extracted from the Chinese medicinal herb
Curcuma wenyujin (13). In
recent studies, β-elemene was shown to have diverse anticancer
potential, such as inhibiting proliferation and inducing apoptosis
of cancer cells, and interacting with multiple oncogenic or tumor
suppressing signaling pathways in a broad spectrum of cancers
(14–16). Other studies found that β-elemene
could enhance tumor chemosensitivity or overcome drug resistance
(17,18). In addition, β-elemene has been
approved by the China Food and Drug Administration as a therapeutic
drug in clinical practice where its efficacy has been exhibited
when combined with first-line chemotherapy for malignant tumors
(19,20). However, the mechanisms by which
β-elemene is involved in tumor suppressing activities remain
largely unknown.
In the present study, we examined the anticancer
potential of β-elemene in the proliferation, clonogenic survival
and apoptosis in SGC7901 and MKN45 gastric cancer cells. Then, in
order to investigate the molecules through which β-elemene
exhibited its anticancer effects and to obtain a better
understanding of its therapeutic role in gastric cancer, we
employed isobaric tags for relative and absolute quantitation
(iTRAQ), a high-throughput proteomic approach, to profile proteins
that were differentially expressed following β-elemene treatment in
gastric cancer cells.
Materials and methods
Reagents
β-elemene was obtained from Jingang Pharmaceutical
Co. (Dalian, China). Annexin V-FITC/PI apoptosis detection kit was
purchased from 7 Sea Pharmacy Technology (Shanghai, China). iTRAQ
reagents were from Applied Biosystems (New York, NY, USA).
Anti-PAK1IP1 antibody was purchased from Abcam (#ab67348; UK).
Anti-TOPIIα antibody was obtained from Proteintech (#20233-1-AP;
USA). Anti-BTF antibody was from BD Biosciences Pharmingen
(#611726; San Diego, CA, USA).
Cell culture
The SGC7901 and MKN45 human gastric cancer cell
lines were obtained from the Lab Animal Centre of the Fourth
Military Medical University (Xi’an, China). Cells were cultured in
RPMI-1640 medium (HyClone, USA), supplemented with 10% fetal bovine
serum (Sijiqing, Huzhou City, China) at 37°C with 5% CO2
in a humidified atmosphere.
MTT assay
Cell viability was measured using
3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT)
assay. The cells were seeded in 24-well plates at
5–10×104/well. After overnight incubation, cells were
exposed to different concentrations of β-elemene for 24–72 h. Then,
50 μl MTT (5 mg/ml) was added to each well and the cells were
incubated for another 4 h at 37°C. After gentle removal of the
supernatant, 500 μl dimethyl sulfoxide (DMSO) was added to each
well to solubilize the purple formazan crystal. The optical density
(OD) was measured using a microplate reader at 490 nm and then
transformed into cell viability using the following formula: Cell
viability = (OD of the experimental sample)/(OD of the control
sample) × 100%.
Annexin V-FITC/PI apoptosis detection
assay
To explore the effect of β-elemene on apoptotic cell
death, Annexin V-FITC/PI apoptosis detection assay was used. The
cells were seeded in 6-well plates at 3×105/well. After
overnight incubation, cells were exposed to different
concentrations of β-elemene for 24 h. Then, cells were collected
and manipulated following the manufacturer’s instructions,
incubated with Annexin V-FITC and propidium iodide (PI), then
analyzed using flow cytometry (FCM; BD Biosciences-Clontech, Palo
Alto, CA, USA) within 30 min.
Clonogenic survival assay
Cells were trypsinized and counted. Then, 200 cells
were seeded into each well of 6-well plates. After overnight
incubation for attachment and recovery, the cells were treated with
different concentrations of β-elemene. Ten to fourteen days after
seeding, cells were washed with PBS twice, fixed with methyl
alcohol for 15 min and stained with 1% crystal violet for 20 min.
Colonies containing >50 cells was counted and the surviving
fractions were calculated as follows: Plating efficiency (PE) =
colony number of the control group/the number of cells seeding.
Surviving fraction = colony number of the treated-group/(the number
of cells seeding × PE). This assay was carried out in
duplicate.
Protein preparation
After SGC7901 cells were treated with or without
β-elemene at 30 μg/ml for 48 h, cell total protein was extracted.
Protein concentration was determined using Pierce™ BCA Protein
Assay (Thermo Scientific, Rockford, IL, USA). Total protein
extracted from two separate experiments was mixed together for use
in the subsequent proteomic analysis. Protein samples were reduced
and alkylated, then added into 5-fold volume of ice-cold acetone
and put in −20°C condition overnight. Then, the precipitate was
harvested by centrifugation at 25,000 × g at 4°C for 20 min and
dried in the air for 5 min. The precipitate was dissolved in 200 μl
0.5 M tetraethylammonium bromide (TEAB) and dealt with sonicate for
15 min. Finally, the supernatant was harvested and quantified.
iTRAQ proteomic analysis
One hundred micrograms of protein were taken out
from each sample and digested with trypsin. Thereafter, the
peptides of each sample were labeled with iTRAQ reagents
respectively, according to the manufacturer’s protocol (Applied
Biosystems) (SGC7901-control-114 tag and SGC7901-β-elemene
treated-115 tag). Then, the labeled samples were pooled and sent to
fractionating using strong cation exchange (SCX) chromatography
(Shimadzu LC-20AB HPLC Pump system and the 4.6×250 mm Ultremex SCX
column). After elution, 20 fractions of peptides were obtained.
Each fraction was then desalted by Strata-X C18 column (Phenomenex)
and vacuum-dried. Each fraction of peptides was resuspended in
buffer A (5% ACN, 0.1% FA) and centrifuged at 20,000 × g for 10 min
to remove the insoluble substances. In each fraction, the final
concentration of peptides was ~0.5 μg/μl. Five microlitres (~2.5
μg) of supernatant was loaded onto a Shimadzu LC-20AD nano HPLC by
the autosampler for separation. Mass spectrometric analysis of the
iTRAQ labelled peptides was performed using a Q Exactive (Thermo
Fisher Scientific, San Jose, CA, USA) coupled online to the HPLC.
Data processing of LC-MS/MS samples was searched against the
International Protein Index (IPI) human protein database version
3.87 FASTA (91,464 sequences) using Mascot 2.3.02 software (Matrix
Science, UK). When the fold-change of protein abundance was >1.2
and the P-value was <0.05, we defined this protein as
differentially expressed. The identified proteins were categorized
according to the Gene Ontology (GO) classification terms
(http://www.geneontology.org/). GO
enrichment analysis was performed to display the GO terms which the
differentially expressed proteins enriched in all identified
proteins.
Western blot analyses
Equal amounts of protein samples were subjected to
SDS-PAGE. Proteins were transferred to the nitrocellulose (NC)
membranes followed by 2 h blocking with 5% skimmed milk at room
temperature. The NC membranes were sequentially incubated with
primary antibodies at 4°C overnight and secondary antibody for 1 h
at room temperature. The reactions were visualized using
electrochemiluminescence (ECL) detection kit (CWBIO, Beijing,
China). The band quantification was performed using Image-Pro Plus
6.0 software (Media Cybernetics).
Statistical analysis
Data are presented as the means ± SD. Statistical
analysis was performed using a two-tailed Student’s t-test through
SPSS 17.0 software (Chicago, USA). P-value <0.05 was considered
to indicate a statistically significant difference.
Results
β-elemene inhibits the viability of
gastric cancer cells
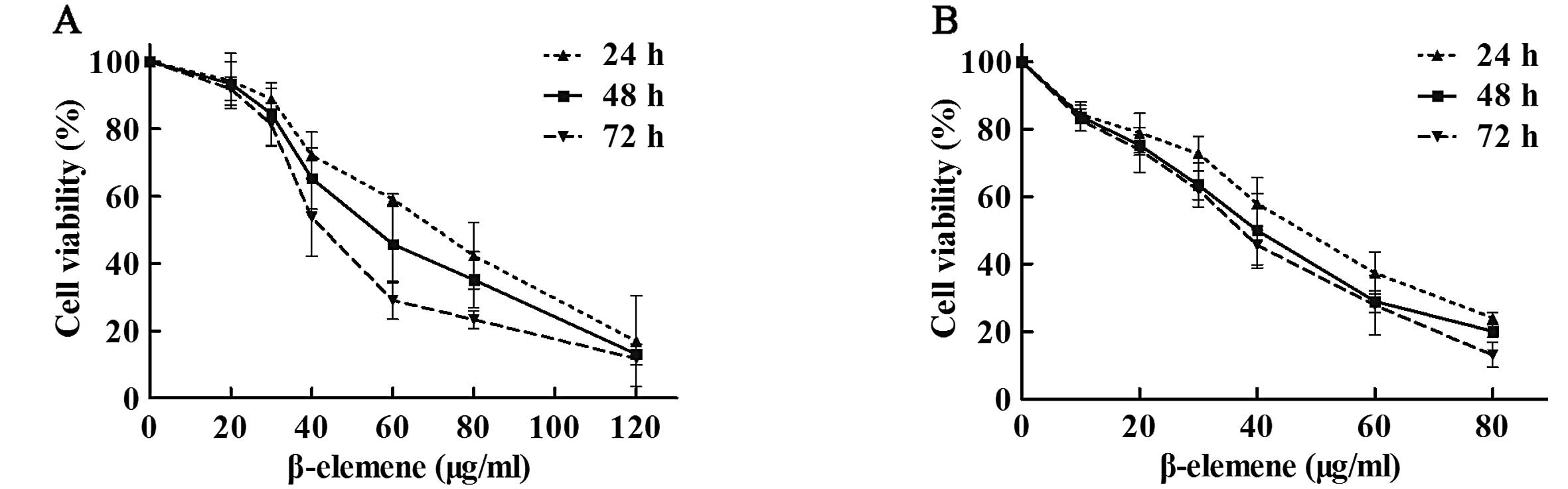
To investigate the antiproliferative effect of
β-elemene in gastric cancer cells, SGC7901 and MKN45 cells were
exposed to different concentrations of β-elemene (0, 10, 20, 30,
40, 60, 80 and 120 μg/ml) for 24, 48 or 72 h. Cell viability
analysis showed that β-elemene suppressed the viability of gastric
cancer cells in a dose-dependent manner (Fig. 1). The 50% inhibitory concentration
(IC50) values of β-elemene for SGC7901 gastric cancer
cells at 24, 48 and 72 h were 67.15, 56.89 and 46.05 μg/ml,
respectively. The IC50 values for MKN45 cells at 24, 48
and 72 h were 45.57, 37.97 and 35.29 μg/ml, respectively. These
results indicate that β-elemene inhibits the viability of gastric
cancer cells in a dose-dependent manner.
β-elemene induces apoptotic cell death in
gastric cancer cells
FCM analysis showed that the apoptotic rate
gradually increased after 24-h exposure to increased concentrations
of β-elemene. The percentage of apoptotic cells in SGC7901 cells in
the control group and β-elemene-treated groups (20, 30, 40, 60 and
80 μg/ml) was 8.5±0.5, 10.4±1.0, 22.2±1.6, 32.6±5.4, 42.8±4.3 and
52.1±4.3%, respectively (Fig. 2A and
B). Compared with the control group, the increased rate of
apoptosis reached statistical significance when the concentrations
of β-elemene were >30 μg/ml in SGC7901 cells. Similar trends of
apoptosis induction effects were observed in MKN45 cells (Fig. 2C and D). These data suggest that
β-elemene induces apoptotic cell death in gastric cancer cells in a
dose-dependent manner.
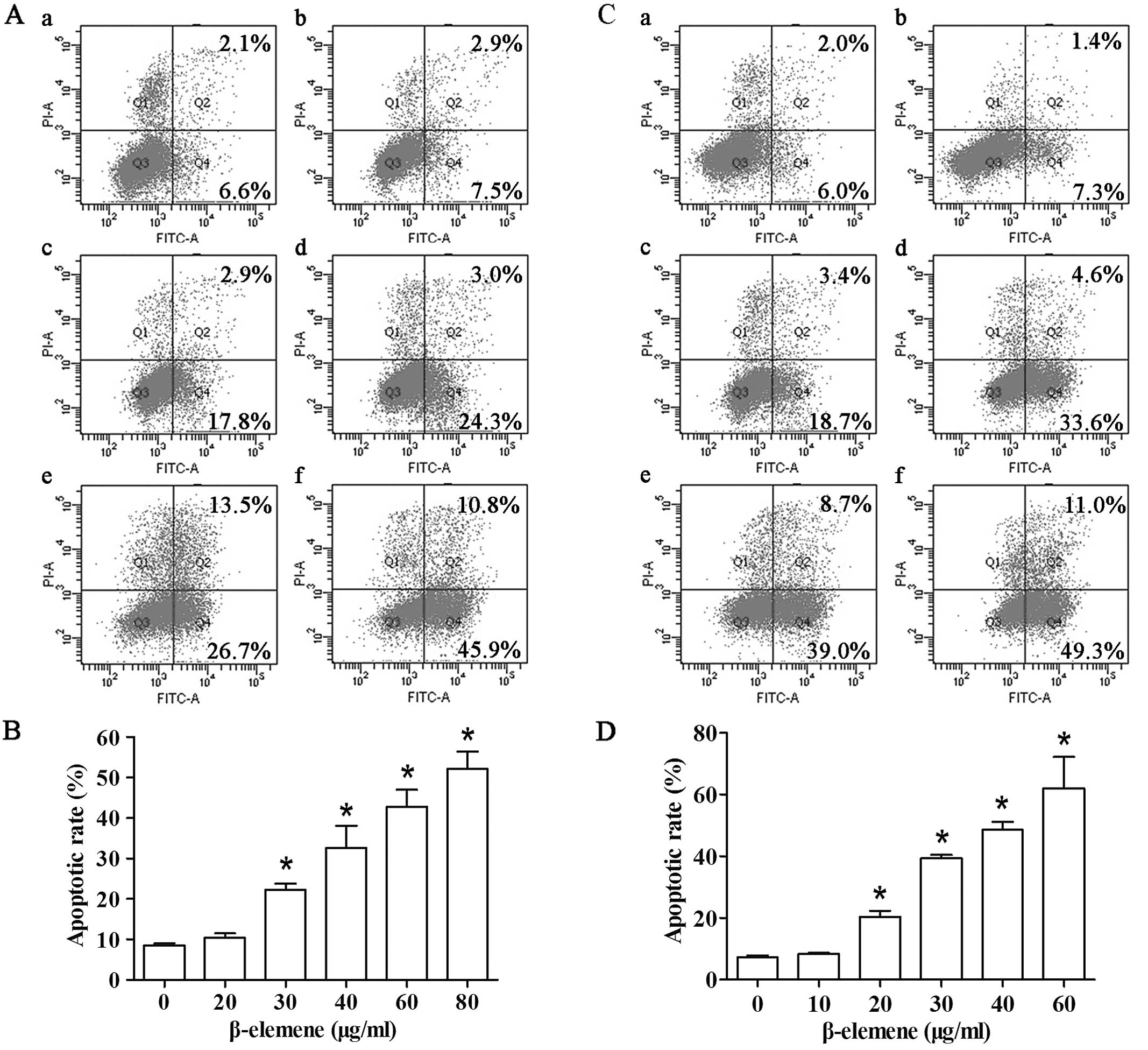 | Figure 2β-elemene induces apoptotic cell
death in gastric cancer cells. Cells were treated with different
concentrations of β-elemene for 24 h. Apoptotic rate was analyzed
by FCM following the manufacturer’s instructions. (A)
Representative FCM results of SGC7901 cells. a, control; b, 20
μg/ml; c, 30 μg/ml; d, 40 μg/ml; e, 60 μg/ml; and f, 80 μg/ml. (B)
Student’s t-test of apoptotic rate was performed between the
control group and groups treated with β-elemene in SGC7901 cells.
(C) Representative FCM results of MKN45 cells. a, control; b, 10
μg/ml; c, 20 μg/ml; d, 30 μg/ml; e, 40 μg/ml; and f, 60 μg/ml. (D)
Quantification of apoptosis of the control group and groups treated
with β-elemene in MKN45 cells. *P<0.05. Column and
bars, means ± SD. FCM, flow cytometry. |
β-elemene decreases clonogenic survival
of gastric cancer cells
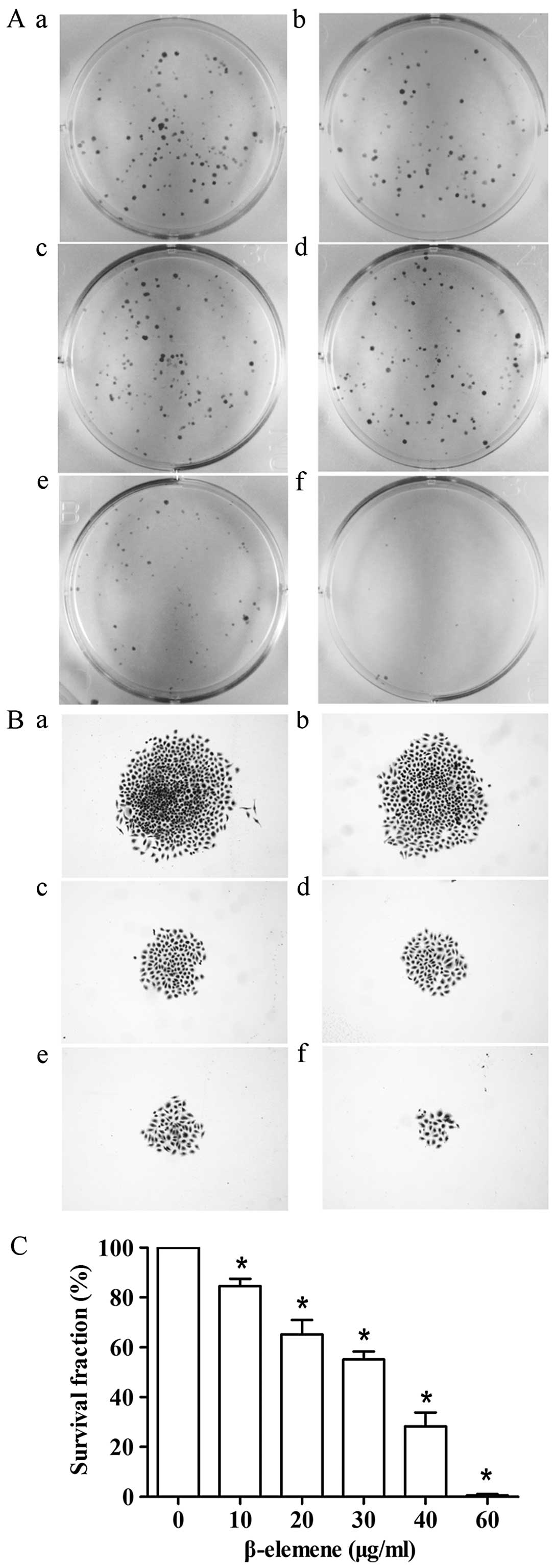
To determine whether β-elemene inhibits colony
forming efficiency, cells were exposed to different concentrations
of β-elemene (0, 10, 20, 30, 40 and 60 μg/ml) and grown in a cell
contact-independent manner. Consistent with the observed effects on
cell viability and apoptosis-induction activity, β-elemene
exhibited anti-clonogenic potential and led to a statistically
significant reduction in colony formation (Fig. 3). It is worth noting that the size
of the colonies tended to be smaller after treatment with β-elemene
(Fig. 3B). When the concentration
of β-elemene reached 60 μg/ml, the number of survived cells in each
cloned cell group barely reached the standard of a colony in
SGC7901 cells. These results suggest that β-elemene induces a
dose-dependent inhibition of clonogenicity in gastric cancer
cells.
iTRAQ identification and quantification
of differentially expressed proteins by β-elemene in SGC7901
gastric cancer cells
Through iTRAQ analysis, 17,154 unique peptides
corresponding to 4,267 proteins were identified in SGC7901 gastric
cancer cells (data not shown). According to our definition of
differentially expressed protein, a total of 233 identified
proteins were regulated by β-elemene intervention in SGC7901
gastric cancer cells, including 147 upregulated proteins and 86
downregulated proteins. The altered proteins of both lists are
shown in Tables I and II, respectively.
 | Table IUpregulated proteins in response to
β-elemene treatment in SGC7901 gastric cancer cells. |
Table I
Upregulated proteins in response to
β-elemene treatment in SGC7901 gastric cancer cells.
| No. | Accession | Gene symbol | Protein name | Fold-ratio
(115:114) |
|---|
| 1 | IPI00549540 | PAK1IP1 | p21-activated
protein kinase-interacting protein 1 | 3.967 |
| 2 | IPI00152429 | SPNS1 | Isoform 3 of
protein spinster homolog 1 | 2.979 |
| 3 | IPI00021885 | FGA | Isoform 1 of
fibrinogen α chain | 2.397 |
| 4 | IPI00011200 | PHGDH |
D-3-phosphoglycerate dehydrogenase | 2.319 |
| 5 | IPI00006213 | PCM1 | Isoform 1 of
pericentriolar material 1 protein | 2.316 |
| 6 | IPI00418426 | CNNM4 | Metal transporter
CNNM4 | 2.313 |
| 7 | IPI00013809 | GSTZ1 | Isoform 1 of
maleylacetoacetate isomerase | 2.292 |
| 8 | IPI00012433 | F8A | Factor VIII intron
22 protein | 2.172 |
| 9 | IPI00015856 | DNPEP | Aspartyl
aminopeptidase | 2.144 |
| 10 | IPI00002564 | XRCC1 | DNA repair protein
XRCC1 | 1.981 |
| 11 | IPI00909584 | HARS2 | Histidyl-tRNA
synthetase homolog | 1.922 |
| 12 | IPI00218116 | OASL | Isoform p30 of 59
kDa 2′–5′-oligoadenylate synthase-like protein | 1.915 |
| 13 | IPI00216138 | TAGLN | Transgelin | 1.904 |
| 14 | IPI00829741 | ANKLE2 | Isoform 1 of
ankyrin repeat and LEM domain-containing protein 2 | 1.872 |
| 15 | IPI00022145 | NUCKS1 | Isoform 1 of
nuclear ubiquitous casein and cyclin-dependent kinases
substrate | 1.771 |
| 16 | IPI00290979 | ABHD5 |
1-acylglycerol-3-phosphate
O-acyltransferase ABHD5 | 1.761 |
| 17 | IPI00554777 | ASNS | Asparagine
synthetase (glutamine-hydrolyzing) | 1.664 |
| 18 | IPI00021978 | PEX11B | Peroxisomal
membrane protein 11B | 1.661 |
| 19 | IPI00328737 | ZNF598 | Isoform 1 of Zinc
finger protein 598 | 1.644 |
| 20 | IPI00101782 | GMPPA | Isoform 1 of
mannose-1-phosphate guanyltransferase α | 1.641 |
| 21 | IPI00007320 | TCF25 | Transcription
factor 25 | 1.627 |
| 22 | IPI00031651 | C7orf50 | Uncharacterized
protein C7orf50 | 1.623 |
| 23 | IPI00247583 | RPL21 | 60S ribosomal
protein L21 | 1.622 |
| 24 | IPI00329596 | TMX2 | Uncharacterized
protein | 1.613 |
| 25 | IPI00740961 | INTS1 | DKFZP586J0619
protein | 1.568 |
| 26 | IPI00102752 | RBM15 | Isoform 1 of
putative RNA-binding protein 15 | 1.544 |
| 27 | IPI00024650 | SLC16A1 | Monocarboxylate
transporter 1 | 1.539 |
| 28 | IPI00026848 | LRPAP1 | α-2-macroglobulin
receptor-associated protein | 1.534 |
| 29 | IPI00019976 | CCDC85B | Coiled-coil
domain-containing protein 85B | 1.52 |
| 30 | IPI00021831 | PRKAR1A | cAMP-dependent
protein kinase type I-α regulatory subunit | 1.485 |
| 31 | IPI00145260 | IBA57 | Putative
transferase CAF17, mitochondrial | 1.474 |
| 32 | IPI00550021 | RPL3 | 60S ribosomal
protein L3 | 1.467 |
| 33 | IPI00853369 | PLXNB2 | Plexin-B2 | 1.465 |
| 34 | IPI00001734 | PSAT1 | Phosphoserine
aminotransferase | 1.463 |
| 35 | IPI00412607 | RPL35 | 60S ribosomal
protein L35 | 1.449 |
| 36 | IPI00027463 | S100A6 | Protein
S100-A6 | 1.447 |
| 37 | IPI00012772 | RPL8 | 60S ribosomal
protein L8 | 1.431 |
| 38 | IPI00299219 | CYR61 | Protein CYR61 | 1.425 |
| 39 | IPI00006362 | EDF1 | Isoform 2 of
endothelial differentiation-related factor 1 | 1.423 |
| 40 | IPI00007309 | TIMM23 | Mitochondrial
import inner membrane translocase subunit Tim23 | 1.422 |
| 41 | IPI00030362 | PLP2 | Isoform 1 of
proteolipid protein 2 | 1.421 |
| 42 | IPI00219840 | AP2S1 | Isoform 1 of AP-2
complex subunit sigma | 1.42 |
| 43 | IPI00980827 | Unknown | Uncharacterized
protein | 1.419 |
| 44 | IPI00296526 | NAGK |
N-acetyl-D-glucosamine kinase | 1.417 |
| 45 | IPI00465361 | RPL13 | 60S ribosomal
protein L13 | 1.416 |
| 46 | IPI00306749 | SLC4A1AP | Kanadaptin | 1.409 |
| 47 | IPI00168262 | GLT25D1 | Procollagen
galactosyltransferase 1 | 1.404 |
| 48 | IPI00410067 | ZC3HAV1 | Isoform 1 of Zinc
finger CCCH-type antiviral protein 1 | 1.397 |
| 49 | IPI00789805 | DIAPH3 | Isoform 3 of
protein diaphanous homolog 3 | 1.395 |
| 50 | IPI00293425 | FXN | Isoform 1 of
frataxin, mitochondrial | 1.394 |
| 51 | IPI00021389 | CCS | Copper chaperone
for superoxide dismutase | 1.376 |
| 52 | IPI00027096 | MRPL19 | 39S ribosomal
protein L19, mitochondrial | 1.376 |
| 53 | IPI00005040 | ACADM | Isoform 1 of
medium-chain specific acyl-CoA dehydrogenase, mitochondrial | 1.376 |
| 54 | IPI00556655 | LAMP1 | LAMP1 protein
variant (fragment) | 1.375 |
| 55 | IPI00101968 | DBNL | Isoform 3 of
drebrin-like protein | 1.37 |
| 56 | IPI00008418 | DIABLO | Diablo homolog,
mitochondrial precursor | 1.36 |
| 57 | IPI00396174 | CCDC25 | Coiled-coil
domain-containing protein 25 | 1.357 |
| 58 | IPI00010404 | SF3B5 | Splicing factor 3B
subunit 5 | 1.356 |
| 59 | IPI00216999 | C14orf21 | Pumilio
domain-containing protein C14orf21 | 1.354 |
| 60 | IPI00293276 | MIF | Macrophage
migration inhibitory factor | 1.352 |
| 61 | IPI00297241 | URB1 | Nucleolar
pre-ribosomal-associated protein 1 | 1.351 |
| 62 | IPI00004406 | UPP1 | Isoform 1 of
uridine phosphorylase 1 | 1.347 |
| 63 | IPI00023122 | PDLIM7 | Isoform 1 of PDZ
and LIM domain protein 7 | 1.346 |
| 64 | IPI00375731 | RBM10 | Isoform 1 of
RNA-binding protein 10 | 1.345 |
| 65 | IPI00170786 | WBP11 | WW domain-binding
protein 11 | 1.345 |
| 66 | IPI00010740 | SFPQ | Isoform long of
splicing factor, proline- and glutamine-rich | 1.343 |
| 67 | IPI00329321 | LYRM7 | LYR
motif-containing protein 7 | 1.341 |
| 68 | IPI00290857 | KRT3 | Keratin, type II
cytoskeletal 3 | 1.34 |
| 69 | IPI00456758 | RPL27A | 60S ribosomal
protein L27a | 1.339 |
| 70 | IPI00063673 | ISY1 | Isoform 1 of
pre-mRNA-splicing factor ISY1 homolog | 1.339 |
| 71 | IPI00217236 | TBCA | Tubulin-specific
chaperone A | 1.339 |
| 72 | IPI00301280 | TMEM43 | Transmembrane
protein 43 | 1.337 |
| 73 | IPI00006079 | BCLAF1 | Isoform 1 of
Bcl-2-associated transcription factor 1 | 1.336 |
| 74 | IPI00746351 | DIS3 | Isoform 1 of
exosome complex exonuclease RRP44 | 1.33 |
| 75 | IPI00180781 | MLKL | Isoform 1 of mixed
lineage kinase domain-like protein | 1.328 |
| 76 | IPI00012493 | RPS20 | 40S ribosomal
protein S20 | 1.321 |
| 77 | IPI00008922 | IFITM2 | Interferon-induced
transmembrane protein 2 | 1.32 |
| 78 | IPI00647650 | EIF3H | Eukaryotic
translation initiation factor 3 subunit 3 | 1.317 |
| 79 | IPI00299573 | RPL7A | 60S ribosomal
protein L7a | 1.317 |
| 80 | IPI01012991 | LUC7L2 | Isoform 1 of
putative RNA-binding protein Luc7-like 2 | 1.315 |
| 81 | IPI00300078 | PWP2 | Periodic tryptophan
protein 2 homolog | 1.311 |
| 82 | IPI00003016 | STRN4 | Striatin-4 | 1.31 |
| 83 | IPI00745955 | EBNA1BP2 | Probable
rRNA-processing protein EBP2 | 1.307 |
| 84 | IPI00220527 | SNX1 | Isoform 1A of
sorting nexin-1 | 1.305 |
| 85 | IPI00017448 | RPS21 | 40S ribosomal
protein S21 | 1.3 |
| 86 | IPI00002135 | TACC3 | Transforming acidic
coiled-coil-containing protein 3 | 1.296 |
| 87 | IPI00011635 | BCL2L13 | Isoform 2 of
Bcl-2-like protein 13 | 1.291 |
| 88 | IPI00157176 | MEA1 | Male-enhanced
antigen 1 | 1.288 |
| 89 | IPI00012756 | IFIT5 | Interferon-induced
protein with tetratricopeptide repeats 5 | 1.287 |
| 90 | IPI00022202 | SLC25A3 | Isoform A of
phosphate carrier protein, mitochondrial | 1.287 |
| 91 | IPI00064767 | ARHGAP17 | Isoform 1 of Rho
GTPase-activating protein 17 | 1.284 |
| 92 | IPI00063827 | ABHD14B | Isoform 1 of
abhydrolase domain-containing protein 14B | 1.282 |
| 93 | IPI00243221 | NRD1 | Nardilysin isoform
a | 1.274 |
| 94 | IPI00297160 | CD44 | Isoform 12 of CD44
antigen | 1.272 |
| 95 | IPI00014808 | PAFAH1B3 | Platelet-activating
factor acetylhydrolase IB subunit γ | 1.269 |
| 96 | IPI00022694 | PSMD4 | Isoform Rpn10A of
26S proteasome non-ATPase regulatory subunit 4 | 1.264 |
| 97 | IPI00011229 | CTSD | Cathepsin D | 1.261 |
| 98 | IPI00003856 | ATP6V1E1 | V-type proton
ATPase subunit E 1 | 1.255 |
| 99 | IPI00291669 | UBLCP1 | Ubiquitin-like
domain-containing CTD phosphatase 1 | 1.252 |
| 100 | IPI00029081 | LIG3 | Isoform α of DNA
ligase 3 | 1.248 |
| 101 | IPI00007797 | FABP5 | Fatty acid-binding
protein, epidermal | 1.248 |
| 102 | IPI00028387 | DDRGK1 | Isoform 1 of DDRGK
domain-containing protein 1 | 1.245 |
| 103 | IPI00007694 | PPME1 | Isoform 1 of
protein phosphatase methylesterase 1 | 1.243 |
| 104 | IPI00031519 | DNMT1 | Isoform 1 of DNA
(cytosine-5)-methyltransferase 1 | 1.241 |
| 105 | IPI00219684 | FABP3 | Fatty acid-binding
protein, heart | 1.24 |
| 106 | IPI00219153 | RPL22 | 60S ribosomal
protein L22 | 1.237 |
| 107 | IPI00219029 | GOT1 | Aspartate
aminotransferase, cytoplasmic | 1.236 |
| 108 | IPI00297178 | DHX16 | 119 kDa
protein | 1.234 |
| 109 | IPI00305267 | GOLGA3 | Isoform 1 of golgin
subfamily A member 3 | 1.234 |
| 110 | IPI00010697 | ITGA6 | Isoform α-6X1X2B of
integrin α-6 | 1.233 |
| 111 | IPI00396387 | GNL1 | Isoform 1 of
guanine nucleotide-binding protein-like 1 | 1.233 |
| 112 | IPI00000030 | PPP2R5D | Isoform δ-1 of
serine/threonine-protein phosphatase 2A 56 kDa regulatory subunit δ
isoform | 1.233 |
| 113 | IPI00007118 | SERPINE1 | Plasminogen
activator inhibitor 1 | 1.23 |
| 114 | IPI00021057 | SLC12A4 | Isoform 1 of solute
carrier family 12 member 4 | 1.229 |
| 115 | IPI00018206 | GOT2 | Aspartate
aminotransferase, mitochondrial | 1.229 |
| 116 | IPI00019472 | SLC1A5 | Neutral amino acid
transporter B(0) | 1.229 |
| 117 | IPI00306353 | DUSP23 | Dual specificity
protein phosphatase 23 | 1.227 |
| 118 | IPI00032825 | TMED7 | Transmembrane emp24
domain-containing protein 7 | 1.227 |
| 119 | IPI01010270 | WIZ | WIZ protein | 1.225 |
| 120 | IPI00006980 | C14orf166 | UPF0568 protein
C14orf166 | 1.224 |
| 121 | IPI00298625 | LYN | Isoform LYN A of
tyrosine-protein kinase Lyn | 1.223 |
| 122 | IPI00550523 | ATL3 | Atlastin-3 | 1.222 |
| 123 | IPI00333215 | TCEA1 | Isoform 1 of
transcription elongation factor A protein 1 | 1.222 |
| 124 | IPI00032038 | CPT1A | Isoform 1 of
carnitine O-palmitoyltransferase 1, liver isoform | 1.221 |
| 125 | IPI00025512 | HSPB1 | Heat shock protein
β-1 | 1.221 |
| 126 | IPI00013468 | BUB3 | Isoform 1 of
mitotic checkpoint protein BUB3 | 1.221 |
| 127 | IPI00307165 | TRIM47 | Tripartite
motif-containing protein 47 | 1.22 |
| 128 | IPI00181396 | VPRBP | Isoform 3 of
protein VPRBP | 1.217 |
| 129 | IPI00216237 | RPL36 | 60S ribosomal
protein L36 | 1.217 |
| 130 | IPI00550069 | RNH1 | Ribonuclease
inhibitor | 1.217 |
| 131 | IPI00215995 | ITGA3 | Isoform 1 of
integrin α-3 | 1.216 |
| 132 | IPI00902512 | PPP1CA |
Serine/threonine-protein phosphatase | 1.215 |
| 133 | IPI00003923 | UMPS | Isoform 1 of
uridine 5′-monophosphate synthase | 1.21 |
| 134 | IPI00010414 | PDLIM1 | PDZ and LIM domain
protein 1 | 1.21 |
| 135 | IPI01026021 | SHMT2 | SHMT2 protein | 1.21 |
| 136 | IPI00021828 | CSTB | Cystatin-B | 1.21 |
| 137 | IPI00060627 | CCDC124 | Coiled-coil
domain-containing protein 124 | 1.209 |
| 138 | IPI00218466 | SEC61A1 | Protein transport
protein Sec61 subunit α isoform 1 | 1.207 |
| 139 | IPI00026546 | PAFAH1B2 | Platelet-activating
factor acetylhydrolase IB subunit β | 1.207 |
| 140 | IPI00011592 | DYNC1LI2 | Cytoplasmic dynein
1 light intermediate chain 2 | 1.206 |
| 141 | IPI00216682 | CNN3 | Calponin-3 | 1.206 |
| 142 | IPI00297900 | DDX10 | Probable
ATP-dependent RNA helicase DDX10 | 1.206 |
| 143 | IPI00292657 | PTGR1 | Prostaglandin
reductase 1 | 1.204 |
| 144 | IPI00925046 | QARS | Glutaminyl-tRNA
synthetase | 1.204 |
| 145 | IPI00339379 | ARHGEF1 | Isoform 2 of Rho
guanine nucleotide exchange factor 1 | 1.202 |
| 146 | IPI00294486 | DUSP9 | Dual specificity
protein phosphatase 9 | 1.201 |
| 147 | IPI00159899 | ANKFY1 | Isoform 1 of
ankyrin repeat and FYVE domain-containing protein 1 | 1.201 |
 | Table IIDownregulated proteins in response to
β-elemene treatment in SGC7901 gastric cancer cells. |
Table II
Downregulated proteins in response to
β-elemene treatment in SGC7901 gastric cancer cells.
| No. | Accession | Gene symbol | Protein name | Fold-ratio
(115:114) |
|---|
| 1 | IPI00654755 | HBB | Hemoglobin subunit
β | 0.163 |
| 2 | IPI00290380 | ALPPL2 | Alkaline
phosphatase, placental-like | 0.531 |
| 3 | IPI00183695 | S100A10 | Protein
S100-A10 | 0.598 |
| 4 | IPI00419273 | CUL4A | Isoform 1 of
cullin-4A | 0.631 |
| 5 | IPI00410110 | DHX40 | Isoform 1 of
probable ATP-dependent RNA helicase DHX40 | 0.637 |
| 6 | IPI00011698 | SAP18 | Histone deacetylase
complex subunit SAP18 | 0.654 |
| 7 | IPI00022002 | MRPS27 | Mitochondrial 28S
ribosomal protein S27 | 0.664 |
| 8 | IPI00171438 | TXNDC5 | Thioredoxin
domain-containing protein 5 | 0.675 |
| 9 | IPI00023704 | LPP | Lipoma-preferred
partner | 0.701 |
| 10 | IPI00301503 | TRA2B | Isoform 1 of
transformer-2 protein homolog β | 0.706 |
| 11 | IPI00414836 | OSTF1 |
Osteoclast-stimulating factor 1 | 0.706 |
| 12 | IPI00019896 | MYCBP2 | Homo sapiens
protein associated with Myc mRNA (fragment) | 0.712 |
| 13 | IPI00414101 | TOP2A | Isoform 2 of DNA
topoisomerase 2-α | 0.716 |
| 14 | IPI00412545 | NDUFA5 | NADH dehydrogenase
(ubiquinone) 1 α subcomplex subunit 5 | 0.717 |
| 15 | IPI00003935 | HIST2H2BE | Histone H2B type
2-E | 0.718 |
| 16 | IPI00291467 | SLC25A6 | ADP/ATP translocase
3 | 0.722 |
| 17 | IPI00374272 | C5orf51 | UPF0600 protein
C5orf51 | 0.725 |
| 18 | IPI00005087 | TMOD3 | Tropomodulin-3 | 0.732 |
| 19 | IPI00008475 | HMGCS1 |
Hydroxymethylglutaryl-CoA synthase,
cytoplasmic | 0.736 |
| 20 | IPI00017342 | RHOG | Rho-related
GTP-binding protein RhoG | 0.742 |
| 21 | IPI00009901 | NUTF2 | Nuclear transport
factor 2 | 0.745 |
| 22 | IPI00298731 | PPP1R10 |
Serine/threonine-protein phosphatase 1
regulatory subunit 10 | 0.746 |
| 23 | IPI00020602 | CSNK2A2 | Casein kinase II
subunit α | 0.753 |
| 24 | IPI00025347 | EMG1 | Ribosomal RNA small
subunit methyltransferase NEP1 | 0.755 |
| 25 | IPI00218962 | C20orf43 | UPF0549 protein
C20orf43 | 0.755 |
| 26 | IPI00014230 | C1QBP | Complement
component 1 Q subcomponent-binding protein, mitochondrial | 0.755 |
| 27 | IPI00219025 | GLRX | Glutaredoxin-1 | 0.757 |
| 28 | IPI00168479 | APOA1BP | Apolipoprotein A-I
binding protein | 0.758 |
| 29 | IPI00027705 | PRIM2 | Isoform 1 of DNA
primase large subunit | 0.758 |
| 30 | IPI00219483 | SNRNP70 | Isoform 2 of U1
small nuclear ribonucleoprotein 70 kDa | 0.765 |
| 31 | IPI00220014 | IDI1 | Isoform 2 of
isopentenyl-diphosphate δ-isomerase 1 | 0.766 |
| 32 | IPI00434580 | MYOM1 | Isoform 1 of
myomesin-1 | 0.771 |
| 33 | IPI00023729 | FN3K |
Fructosamine-3-kinase | 0.773 |
| 34 | IPI00013446 | PSCA | Prostate stem cell
antigen | 0.78 |
| 35 | IPI00924816 | MTPN | Myotrophin | 0.781 |
| 36 | IPI00033022 | DNM2 | Isoform 1 of
dynamin-2 | 0.781 |
| 37 | IPI00009032 | SSB | Lupus la
protein | 0.781 |
| 38 | IPI00073779 | MRPS35 | Isoform 1 of 28S
ribosomal protein S35, mitochondrial | 0.782 |
| 39 | IPI00163644 | OSBPL8 | Oxysterol-binding
protein | 0.783 |
| 40 | IPI00419626 | MRPL55 | Isoform 2 of 39S
ribosomal protein L55, mitochondrial | 0.789 |
| 41 | IPI00006440 | MRPS7 | 28S ribosomal
protein S7, mitochondrial | 0.79 |
| 42 | IPI00027704 | PRIM1 | DNA primase small
subunit | 0.792 |
| 43 | IPI00018768 | TSN | Translin | 0.795 |
| 44 | IPI00876962 | INF2 | Isoform 2 of
inverted formin-2 | 0.795 |
| 45 | IPI00156374 | IPO4 | Isoform 1 of
importin-4 | 0.795 |
| 46 | IPI00017344 | RAB5B | Ras-related protein
Rab-5B | 0.799 |
| 47 | IPI00418290 | MRPL14 | 39S ribosomal
protein L14, mitochondrial | 0.8 |
| 48 | IPI00022820 | GTF2B | Transcription
initiation factor IIB | 0.801 |
| 49 | IPI00006579 | COX4I1 | Cytochrome c
oxidase subunit 4 isoform 1, mitochondrial | 0.801 |
| 50 | IPI00217766 | SCARB2 | Lysosome membrane
protein 2 | 0.801 |
| 51 | IPI00013396 | SNRPC | U1 small nuclear
ribonucleoprotein C | 0.802 |
| 52 | IPI00022977 | CKB | Creatine kinase
B-type | 0.804 |
| 53 | IPI00012074 | HNRNPR | Isoform 1 of
heterogeneous nuclear ribonucleoprotein R | 0.808 |
| 54 | IPI00216171 | ENO2 | γ-enolase | 0.809 |
| 55 | IPI00014958 | PON2 | Isoform 1 of serum
paraoxonase/arylesterase 2 | 0.812 |
| 56 | IPI00018288 | POLR2C | DNA-directed RNA
polymerase II subunit RPB3 | 0.812 |
| 57 | IPI00010948 | TRIM26 | Tripartite
motif-containing protein 26 | 0.812 |
| 58 | IPI00007052 | FIS1 | Mitochondrial
fission 1 protein | 0.813 |
| 59 | IPI00329625 | TBRG4 | Transforming growth
factor β regulator 4 | 0.813 |
| 60 | IPI00017510 | MT-CO2 | Cytochrome c
oxidase subunit 2 | 0.814 |
| 61 | IPI00645898 | XPNPEP1 | X-prolyl
aminopeptidase (aminopeptidase P) 1, soluble | 0.814 |
| 62 | IPI00029054 | NT5C2 | Cytosolic purine
5′-nucleotidase | 0.816 |
| 63 | IPI00374970 | SEPT10 | Isoform 1 of
septin-10 | 0.817 |
| 64 | IPI00005948 | MRI1 | Isoform 1 of
methylthioribose-1-phosphate isomerase | 0.817 |
| 65 | IPI00017526 | S100P | Protein S100-P | 0.817 |
| 66 | IPI00026964 | UQCRFS1 | Cytochrome b-c1
complex subunit Rieske, mitochondrial | 0.817 |
| 67 | IPI00219673 | GSTK1 | Isoform 1 of
glutathione S-transferase κ 1 | 0.818 |
| 68 | IPI00296432 | IWS1 | Isoform 1 of
protein IWS1 homolog | 0.818 |
| 69 | IPI00029697 | EXOSC9 | Isoform 2 of
exosome complex component RRP45 | 0.82 |
| 70 | IPI00062336 | RPRD1A | Isoform 2 of
regulation of nuclear pre-mRNA domain-containing protein 1A | 0.822 |
| 71 | IPI00179172 | PPFIBP1 | Isoform 2 of
liprin-β-1 | 0.822 |
| 72 | IPI00015972 | COX6C | Cytochrome c
oxidase subunit 6C | 0.823 |
| 73 | IPI00410657 | RNMT | Isoform 2 of mRNA
cap guanine-N7 methyltransferase | 0.823 |
| 74 | IPI00029019 | UBAP2L | Isoform 2 of
ubiquitin-associated protein 2-like | 0.825 |
| 75 | IPI00007188 | SLC25A5 | ADP/ATP translocase
2 | 0.826 |
| 76 | IPI00292056 | PIK3C2B |
Phosphatidylinositol-4-phosphate 3-kinase
C2 domain-containing subunit β | 0.828 |
| 77 | IPI00013475 | TUBB2A | Tubulin β-2A
chain | 0.829 |
| 78 | IPI00293590 | MGLL | Monoglyceride
lipase isoform 1 | 0.829 |
| 79 | IPI00303568 | PTGES2 | Prostaglandin E
synthase 2 | 0.829 |
| 80 | IPI00027444 | SERPINB1 | Leukocyte elastase
inhibitor | 0.829 |
| 81 | IPI00003765 | CAPN7 | Calpain-7 | 0.83 |
| 82 | IPI00215920 | ARF6 | ADP-ribosylation
factor 6 | 0.831 |
| 83 | IPI00008449 | FIP1L1 | Isoform 3 of
pre-mRNA 3′-end-processing factor FIP1 | 0.831 |
| 84 | IPI00061178 | RBMXL1 | Heterogeneous
nuclear ribonucleoprotein G-like 1 | 0.831 |
| 85 | IPI00180292 | BAIAP2 | Isoform 5 of
brain-specific angiogenesis inhibitor 1-associated protein 2 | 0.832 |
| 86 | IPI00029468 | ACTR1A | α-centractin | 0.833 |
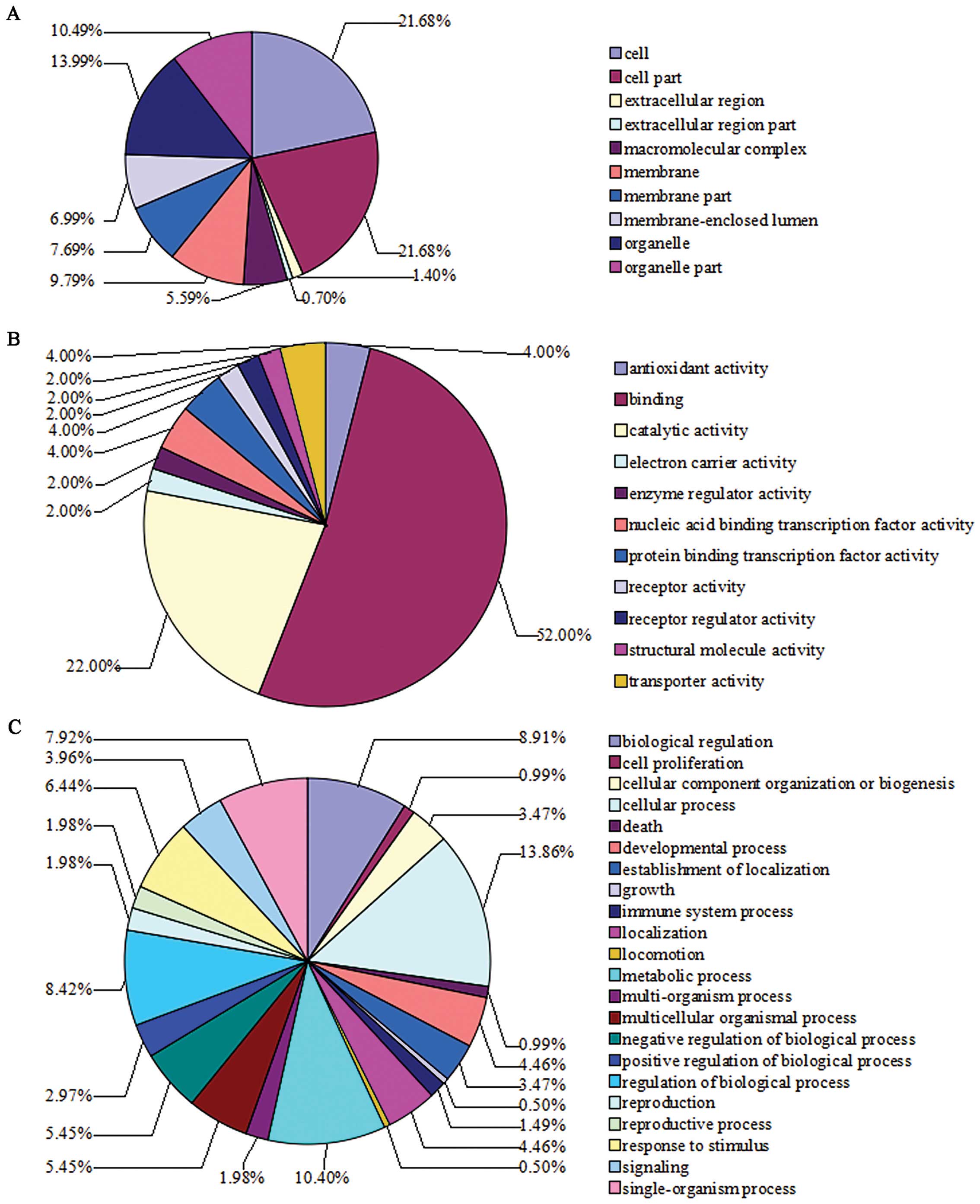
GO analysis of differentially expressed
proteins altered by β-elemene in SGC7901 gastric cancer cells
Based on the GO terms analysis, categorization of
these differentially expressed proteins according to cellular
component, molecular function and biological process is shown in
Fig. 4. GO enrichment analysis
shows the top pathways involved in the differentially expressed
proteins resulting from β-elemene treatment. They include
phenylalanine, tyrosine and tryptophan biosynthesis, ribosome
signaling, tyrosine metabolism, phenylalanine metabolism, PPAR
signaling pathway, regulation of actin cytoskeleton, cysteine and
methionine metabolism, ether lipid metabolism, hematopoietic cell
lineage and phagosome signaling pathways.
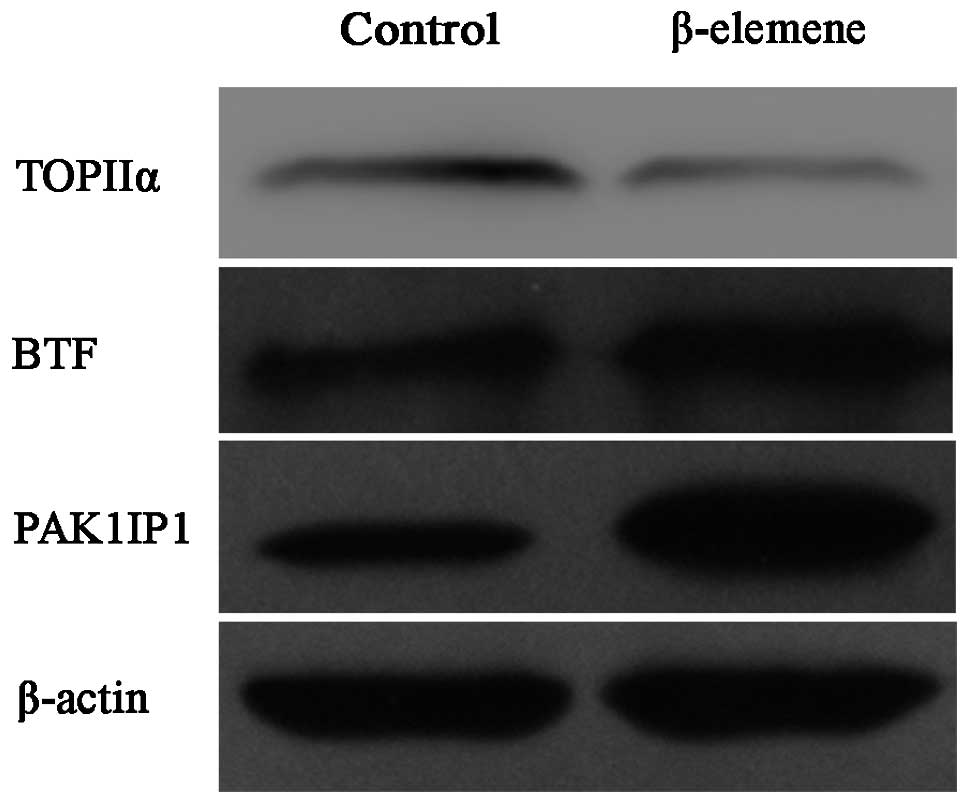
Validation of iTRAQ results by western
blot analyses
Western blot analyses were performed to validate the
differentially expressed proteins discovered by iTRAQ proteomic
analysis. The SGC7901 human gastric cancer cells were treated the
same as for the iTRAQ analysis and lysed for protein samples. Three
proteins were selected for validation purposes according to our
interests and the availability of antibodies. The results were
consistent with those found using iTRAQ (Fig. 5) and indicated the high reliability
of our iTRAQ results.
Discussion
Previous studies have shown that β-elemene, a
promising anti-cancer drug extracted from natural plants, has
efficient growth inhibition effects in a broad range of cancer
cells, although with slight toxicity to normal tissue cells
(14,21,22).
In China, it has been used as a therapeutic candidate for certain
malignant tumors for several years (20). However, little is known about the
underlying molecules. In the present study, our data indicated that
β-elemene efficiently suppressed the proliferation and survival of
gastric cancer cells at least partly through the induction of
apoptosis. The effects are consistent with results in other
malignancies (15,21). Different from other studies, we
employed an iTRAQ proteomic method to explore the potential
proteins that may contribute to its anticancer effect. As a result,
the differentially expressed proteins in response to β-elemene
treatment in gastric cancer cells provided insight and supported
the results found at the cytological level. Furthermore, the
analysis provided some other molecules and signal pathways that may
predict other pharmacologic actions of β-elemene that had not been
studied in cancer therapy. In brief, our results provide evidence
that β-elemene may be a potential drug for gastric cancer. Some of
the key proteins are potential markers of gastric cancer treatment
and are briefly discussed below.
Bcl-2 family proteins were found to be modulated in
β-elemene-treated cancer cells in previous studies and make sense
in apoptosis induction (14,21,22).
The balance between pro-apoptotic and anti-apoptotic members of the
Bcl-2 family proteins decides the fates of cells, and the increased
proportion of pro-apoptotic proteins results in apoptotic cell
death. Therefore, they play vital role in cancer therapy (23). In the present study,
Bcl-2-associated transcription factor 1 (BTF) and Bcl-2-like
protein 13 (also known as Bcl-rambo) were upregulated to 1.34- and
1.29-fold in response to β-elemene treatment compared with the
control untreated gastric cancer cells, respectively. Both play
death-promoting roles in cancer cells. BTF was first identified as
a transcriptional repressor that interacted with Bcl-2 family
proteins and overexpression of BTF could induce apoptosis (24). Previous studies demonstrated the
roles of BTF in apoptosis promotion through the control of
transcription or correlations with Bcl-2 family members (25–27).
Moreover, BTF has been shown to inhibit DNA damage repair (25,27).
This may partly contribute to the fact that β-elemene enhances
tumor chemosensitivity or overcomes drug resistance (17,18).
The other molecule, Bcl-rambo, is also a pro-apoptotic member of
the Bcl-2 family (28,29). Although it was found to trigger cell
death in a way distinct from the traditional Bcl-2 family members,
Bcl-rambo-induced apoptotic signaling pathway eventually joined
other pro-apoptotic pathways at the level of caspase-3 (28). Taken together, these results
indicate that Bcl-2 family proteins play a critical role in
β-elemene-induced cell death in gastric cancer cells.
p21-activated protein kinase-interacting protein 1
(PAK1IP1) was the most influenced protein among the list of
upregulated proteins by β-elemene. P21-activated protein kinase 1
(PAK1) and PAK signal pathways have been shown to have multiple
roles in cancer cell biological behaviors, such as cytoskeletal
dynamics, survival, proliferation and transcription (30,31).
PAK1 and its PAK family members are overexpressed or hyperactivated
in several types of cancer and play a critical role in
tumorigenesis and metastasis. Inhibition of PAK1 may efficiently
block the transformation of cancer cells and act as a therapeutic
strategy in cancer treatment (30,32,33).
As a negative regulator of PAK1, PAK1IP1 specifically binds to the
N-terminal regulatory domain of PAK1 and inhibits the activation of
PAK1 by blocking the binding site of Rac and Cdc42, thus playing a
negative role in cancer development and progression (30,31).
However, little research has focused on PAK1IP1. In a recent study,
Yu et al found that PAK1IP1 was upregulated when cancer
cells were suffering from ribosomal stresses and overexpression of
PAK1IP1 could inhibit proliferation via p53-MDM2 loop (34). In the present study, PAK1IP1
expression was ~3-fold upregulated in β-elemene-treated gastric
cancer cells. Therefore, we hypothesize that β-elemene may
upregulate PAK1IP1 expression and thus inhibit the activation of
PAK1, which subsequently inhibits proliferation and induces
apoptosis in gastric cancer cells.
One of the major protein groups regulated by
β-elemene in SGC7901 gastric cancer cells was ribosomal proteins,
with 12 ribosomal proteins upregulated and 4 ribosomal proteins
downregulated. Over the last decade, some ribosomal proteins have
been linked with emerging functions in cancer, in addition to
protein synthesis. Ribosomal protein L5 (RPL5), along with RPL11
and RPL23, may form a complex with MDM2 oncoprotein and activate
p53 through the inhibition of MDM2-mediated p53 degradation
(35). RPS7 was also found to
interact with MDM2 and overexpression of RPS7 increased cell
apoptosis and suppressed cell proliferation after p53 activation
(36). RPS14 and RPL11 were
demonstrated to inhibit cell proliferation by negative regulation
of c-Myc activity (37,38). In some other studies, the
correlation between ribosomal proteins and drug resistance in
cancer therapy was established. RPS13 and RPL23 could suppress
drug-induced apoptosis and thus mediate multidrug resistance in
gastric cancer cells (39). RPL35a
was found overexpressed in many glioblastoma multiforme (GBM) brain
tumors and led to chemotherapy resistance in GBM (40). Together with the present study,
these results propose more roles of ribosomal proteins in cancer
and therefore merit further attention in cancer therapy
research.
S100A10 was a top molecule significantly
downregulated by β-elemene in the present study. S100A10, also
known as p11, is a unique member of the S100 protein family which
serves for intracellular calcium signaling and is characterized by
two EF hand motifs (41,42). The homodimer of S100A10 forms a
heterotetrameric complex with two molecules of Annexin A2, a type
of plasma membrane protein, to maintain stability and execute its
functions (43). Expression of
S100A10 has been detected in a broad spectrum of tissues and
cancers including gastric cancer (44–47).
Over the past years, increasing evidence has demonstrated the
promoting role of S100A10 in tumor invasion and metastasis and that
knockdown of S100A10 could efficiently suppress cancer progression
(46–48). Notably, PAK1IP1, the most influenced
protein of upregulated proteins by β-elemene, specifically targets
PAK1 which is also most associated with cytoskeletal dynamics.
Collectively, we deduced that β-elemene may inhibit invasion and
metastasis in gastric cancer therapy, which warrants further
investigation.
In conclusion, this is the first time that the iTRAQ
proteomic method has been employed in the study of β-elemene in
cancer cells. The present study indicated a promising anticancer
role of β-elemene in gastric cancer therapy. The expression of a
wide range of proteins was altered when gastric cancer cells were
exposed to β-elemene. The differentially expressed proteins
provided comprehensive insight into the potential underlying
molecular mechanisms of the anticancer effects of β-elemene in
gastric cancer cells. Furthermore, some of the proteins may act as
predictors regarding further therapeutic potential of β-elemene,
which merits further study in gastric cancer treatment. The present
study was a preliminary exploration into the anticancer potential
of β-elemene in gastric cancer cells. Based on the current results,
we expect more research to be carried out on β-elemene and other
traditional herbal medicine, in order to improve the management of
gastric cancer.
Acknowledgements
This study was funded by the National Natural
Science Foundation of China (grant no. 81172357). The field study
was conducted in the Center for Translational Medicine of the First
Affiliated Hospital of Xi’an Jiaotong University, and the
proteomics technology platform of BGI Technology Ltd. The authors
thank the staff workers for their practical assistance and
opinions.
References
|
1
|
Ferlay J, Shin HR, Bray F, Forman D,
Mathers C and Parkin DM: Estimates of worldwide burden of cancer in
2008: GLOBOCAN 2008. Int J Cancer. 127:2893–2917. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Saka M, Morita S, Fukagawa T and Katai H:
Present and future status of gastric cancer surgery. Jpn J Clin
Oncol. 41:307–313. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Sastre J, García-Saenz JA and Diaz-Rubio
E: Chemotherapy for gastric cancer. World J Gastroenterol.
12:204–213. 2006.
|
|
4
|
Smalley SR, Benedetti JK, Haller DG, et
al: Updated analysis of SWOG-directed intergroup study 0116: a
phase III trial of adjuvant radiochemotherapy versus observation
after curative gastric cancer resection. J Clin Oncol.
30:2327–2333. 2012. View Article : Google Scholar
|
|
5
|
Sakamoto J, Teramukai S, Nakazato H, et
al: Efficacy of adjuvant immunochemotherapy with OK-432 for
patients with curatively resected gastric cancer: a meta-analysis
of centrally randomized controlled clinical trials. J Immunother.
25:405–412. 2002. View Article : Google Scholar
|
|
6
|
Dassen AE, Lemmens VE, van de Poll-Franse
LV, et al: Trends in incidence, treatment and survival of gastric
adenocarcinoma between 1990 and 2007: a population-based study in
the Netherlands. Eur J Cancer. 46:1101–1110. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Zhang H, Sun LL, Meng YL, et al: Survival
trends in gastric cancer patients of Northeast China. World J
Gastroenterol. 17:3257–3262. 2011.PubMed/NCBI
|
|
8
|
Yoo CH, Noh SH, Shin DW, Choi SH and Min
JS: Recurrence following curative resection for gastric carcinoma.
Br J Surg. 87:236–242. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
D’Angelica M, Gonen M, Brennan MF,
Turnbull AD, Bains M and Karpeh MS: Patterns of initial recurrence
in completely resected gastric adenocarcinoma. Ann Surg.
240:808–816. 2004.PubMed/NCBI
|
|
10
|
Di Costanzo F, Gasperoni S, Manzione L, et
al: Adjuvant chemotherapy in completely resected gastric cancer: a
randomized phase III trial conducted by GOIRC. J Natl Cancer Inst.
100:388–398. 2008.PubMed/NCBI
|
|
11
|
GASTRIC (Global Advanced/Adjuvant Stomach
Tumor Research International Collaboration) Group. Paoletti X, Oba
K, Burzykowski T, et al: Benefit of adjuvant chemotherapy for
resectable gastric cancer. JAMA. 303:1729–1737. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Gan T, Wu Z, Tian L and Wang Y: Chinese
herbal medicines for induction of remission in advanced or late
gastric cancer. Cochrane Database Syst Rev. 1:CD0050962010.
View Article : Google Scholar
|
|
13
|
Tan W, Lu J, Huang M, et al: Anti-cancer
natural products isolated from chinese medicinal herbs. Chin Med.
6:272011. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Lee RX, Li QQ and Reed E: β-elemene
effectively suppresses the growth and survival of both
platinum-sensitive and -resistant ovarian tumor cells. Anticancer
Res. 32:3103–3113. 2012.
|
|
15
|
Lu X, Wang Y, Luo H, et al: β-elemene
inhibits the proliferation of T24 bladder carcinoma cells through
upregulation of the expression of Smad4. Mol Med Rep. 7:513–518.
2013.
|
|
16
|
Zhan YH, Liu J, Qu XJ, et al: β-elemene
induces apoptosis in human renal-cell carcinoma 786-0 cells through
inhibition of MAPK/ERK and PI3K/Akt/mTOR signalling pathways. Asian
Pac J Cancer Prev. 13:2739–2744. 2012.
|
|
17
|
Xu HB, Li L, Fu J, Mao XP and Xu LZ:
Reversion of multidrug resistance in a chemoresistant human breast
cancer cell line by β-elemene. Pharmacology. 89:303–312.
2012.PubMed/NCBI
|
|
18
|
Li QQ, Lee RX, Liang H, Zhong Y and Reed
E: Enhancement of cisplatin-induced apoptosis by β-elemene in
resistant human ovarian cancer cells. Med Oncol. 30:1–13. 2013.
|
|
19
|
Wang B, Peng XX, Sun R, et al: Systematic
review of β-elemene injection as adjunctive treatment for lung
cancer. Chin J Integr Med. 18:813–823. 2012.
|
|
20
|
Xu HB, Zheng LP, Li L, Xu LZ and Fu J:
Elemene, one ingredient of a Chinese herb, against malignant
tumors: a literature-based meta-analysis. Cancer Invest.
31:156–166. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Li QQ, Wang G, Huang F, Banda M and Reed
E: Antineoplastic effect of β-elemene on prostate cancer cells and
other types of solid tumour cells. J Pharm Pharmacol. 62:1018–1027.
2010.
|
|
22
|
Wang G, Li X, Huang F, et al: Antitumor
effect of β-elemene in non-small-cell lung cancer cells is mediated
via induction of cell cycle arrest and apoptotic cell death. Cell
Mol Life Sci. 62:881–893. 2005.
|
|
23
|
García-Sáez AJ: The secrets of the Bcl-2
family. Cell Death Differ. 19:1733–1740. 2012.
|
|
24
|
Kasof GM, Goyal L and White E: Btf, a
novel death-promoting transcriptional repressor that interacts with
Bcl-2-related proteins. Mol Cell Biol. 19:4390–4404.
1999.PubMed/NCBI
|
|
25
|
Liu H, Lu ZG, Miki Y and Yoshida K:
Protein kinase C δ induces transcription of the TP53 tumor
suppressor gene by controlling death-promoting factor Btf in the
apoptotic response to DNA damage. Mol Cell Biol. 27:8480–8491.
2007.
|
|
26
|
Sarras H, Alizadeh Azami S and McPherson
JP: In search of a function for BCLAF1. Sci World J. 10:1450–1461.
2010. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Lee YY, Yu YB, Gunawardena HP, Xie L and
Chen X: BCLAF1 is a radiation-induced H2AX-interacting partner
involved in γH2AX-mediated regulation of apoptosis and DNA
repair. Cell Death Dis. 3:e3592012. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Kataoka T, Holler N, Micheau O, et al:
Bcl-rambo, a novel Bcl-2 homologue that induces apoptosis via its
unique C-terminal extension. J Biol Chem. 276:19548–19554. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Kim JY, So KJ, Lee S and Park JH:
Bcl-rambo induces apoptosis via interaction with the adenine
nucleotide translocator. FEBS Lett. 586:3142–3149. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Xia C, Ma W, Stafford LJ, Marcus S, Xiong
WC and Liu M: Regulation of the p21-activated kinase (PAK) by a
human Gβ-like WD-repeat protein, hPIP1. Proc Natl Acad Sci USA.
98:6174–6179. 2001.PubMed/NCBI
|
|
31
|
Dummler B, Ohshiro K, Kumar R and Field J:
Pak protein kinases and their role in cancer. Cancer Metastasis
Rev. 28:51–63. 2009. View Article : Google Scholar
|
|
32
|
Kissil JL, Wilker EW, Johnson KC, Eckman
MS, Yaffe MB and Jacks T: Merlin, the product of the Nf2 tumor
suppressor gene, is an inhibitor of the p21-activated kinase, Pak1.
Mol Cell. 12:841–849. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Hirokawa Y, Nakajima H, Hanemann CO, et
al: Signal therapy of NF1-deficient tumor xenograft in mice by the
anti-PAK1 drug FK228. Cancer Biol Ther. 4:379–381. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Yu W, Qiu Z, Gao N, et al: PAK1IP1, a
ribosomal stress-induced nucleolar protein, regulates cell
proliferation via the p53-MDM2 loop. Nucleic Acids Res.
39:2234–2248. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Dai MS and Lu H: Inhibition of
MDM2-mediated p53 ubiquitination and degradation by ribosomal
protein L5. J Biol Chem. 279:44475–44482. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Chen D, Zhang Z, Li M, et al: Ribosomal
protein S7 as a novel modulator of p53-MDM2 interaction: binding to
MDM2, stabilization of p53 protein, and activation of p53 function.
Oncogene. 26:5029–5037. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Dai MS, Sun XX and Lu H: Ribosomal protein
L11 associates with c-Myc at 5 S rRNA and tRNA genes and regulates
their expression. J Biol Chem. 285:12587–12594. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Zhou X, Hao Q, Liao JM, Liao P and Lu H:
Ribosomal protein S14 negatively regulates c-Myc activity. J Biol
Chem. 288:21793–21801. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Shi Y, Zhai H, Wang X, et al: Ribosomal
proteins S13 and L23 promote multidrug resistance in gastric cancer
cells by suppressing drug-induced apoptosis. Exp Cell Res.
296:337–346. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Lopez CD, Martinovsky G and Naumovski L:
Inhibition of cell death by ribosomal protein L35a. Cancer Lett.
180:195–202. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Marenholz I, Lovering RC and Heizmann CW:
An update of the S100 nomenclature. Biochim Biophys Acta.
1763:1282–1283. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Gross SR, Sin CG, Barraclough R and
Rudland PS: Joining S100 proteins and migration: for better or for
worse, in sickness and in health. Cell Mol Life Sci. 71:1551–1579.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Madureira PA, O’Connell PA, Surette AP,
Miller VA and Waisman DM: The biochemistry and regulation of
S100A10: a multifunctional plasminogen receptor involved in
oncogenesis. J Biomed Biotechnol. 2012:3536872012. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
El-Rifai W, Moskaluk CA, Abdrabbo MK, et
al: Gastric cancers overexpress S100A calcium-binding proteins.
Cancer Res. 62:6823–6826. 2002.PubMed/NCBI
|
|
45
|
Domoto T, Miyama Y, Suzuki H, et al:
Evaluation of S100A10, annexin II and B-FABP expression as markers
for renal cell carcinoma. Cancer Sci. 98:77–82. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Yang X, Popescu NC and Zimonjic DB: DLC1
interaction with S100A10 mediates inhibition of in vitro cell
invasion and tumorigenicity of lung cancer cells through a
RhoGAP-independent mechanism. Cancer Res. 71:2916–2925. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Shang J, Zhang Z, Song W, et al: S100A10
as a novel biomarker in colorectal cancer. Tumour Biol.
34:3785–3790. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Oue N, Hamai Y, Mitani Y, et al: Gene
expression profile of gastric carcinoma: identification of genes
and tags potentially involved in invasion, metastasis, and
carcinogenesis by serial analysis of gene expression. Cancer Res.
64:2397–2405. 2004. View Article : Google Scholar
|