Introduction
The use of antisense oligonucleotides (ASOs) to
control functions in diseased cells was first recognized more than
20 years ago (1). ASOs,
downregulating the expression of target molecules via mRNA
silencing, exhibit tissue specificity and can be used to ameliorate
the symptoms of various clinical conditions and to significantly
improve the quality of life of patients (2–4). In
the fight against malignant neoplasms, the strategy employing
antisense therapy can block cancer genes, which produce malicious
proteins; hence, ASOs have the potential to become a powerful
weapon in this field (5). Moreover,
it has been demonstrated that the use of a single treatment
strategy against cancer is generally ineffective due to the
multifactorial nature of this disease (6). Therefore, increased attention is being
given to the combination of antisense therapy and chemotherapy
(6,7).
Although the promising properties of ASOs have been
recognized, the lack of methods applicable for ASO delivery
avoiding the degradation of ASOs through the delivery routes is the
major challenge in the wide utilization of these oligonucleotides
in clinical practice. Thereby, the development of this new class of
drugs has been delayed because of the difficulties in DNA, or RNA
delivery, to cells (8), where
ubiquitous nucleases may attack a cargo (9). Moreover, the transfer across membranes
is also complicated due to the hydrophilic character and anionic
backbone contained in the ASO structure (10).
Therefore, an effective delivery system is essential
to protect a cargo against premature degradation. Regarding this
issue, cationic liposomes have been demonstrated as a promising
tool with which to protect ASOs from the extracellular matrix with
potential to transport these molecules into cells (11–13).
Liposomes can be safely administered to humans, because of their
low toxicity and immunogenicity (14). However, this method still has the
disadvantage of low efficacy of cargo transfer and therefore,
improvement of transfer is crucial for their further utilization
(15).
This study focused on the construction of cationic
liposomes containing E2 in the lipid bilayer and
antisense oligonucleotides designed against the human transcription
factors estrogen receptor α (ERα) and β (ERβ) enclosed in their
inner cavities. The E2 was employed to replace
cholesterol, constituting a lipid bilayer of liposomes, to increase
the efficiency of transporters. To study the effects of
liposome-based transporters on proliferation, gene expression and
redox status, a subclone of the well-characterized human estrogen
receptor-positive breast cancer cell line MCF-7 was employed, as
one of the E2-liposome transporters specifically
inhibits MCF-7 cell proliferation and affects gene expression.
Moreover, the levels of oxidative indicators were determined.
Materials and methods
Chemicals and pH measurement
The working solutions such as buffers or the
standard solution of 17β-estradiol (E2) were prepared
daily by diluting the stock solutions. Estradiol, oligonucleotides
and other agents were purchased from Sigma-Aldrich (St. Louis, MO,
USA) at ACS purity, unless noted otherwise. The antisense
oligonucleotides against estrogen receptor α and β (Sigma-Aldrich)
were designed and the sequences are shown in Table I.
 | Table IAntisense oligonucleotide
sequences. |
Table I
Antisense oligonucleotide
sequences.
| Antisense1α |
5′-GACCATGACCATGACCCT-3′ |
| Antisense1β |
5′-TGTGCCTGGCTAGAGATC-3′ |
Preparation of liposomes
Liposomes were prepared following the lipidic
thin-film hydration method. Briefly, 100 mg of cholesterol, 100 mg
of 1,2-dioleoyl-sn-glycero-3-phosphorac-(1-glycerol) sodium salt
and 100 mg of phosphatidylcholine were dissolved in chloroform (4.5
ml). A lipid film was obtained by rotary evaporation of solvent and
the residual chloroform was blown out by nitrogen to remove the
organic solvent until a thin film was formed. For encapsulation
into liposomes, 2 ml of solution containing 17β-estradiol and ASOs
was used. The samples were homogenized in ultrasonic bath Sonorex
Digital 10P (Bandelin, Berlin, Germany) for 15 min. The homogenized
mixtures were then heated and shaken for 15 min at 60°C using the
Thermomixer comfort (Eppendorf AG, Hamburg, Germany). The samples
were then washed several times with Britton-Robinson buffer (pH,
10.0) using Amicon 3K (Millipore, Billerica, MA, USA).
Determination of 17β-estradiol content in
the liposomes
The hormone content in the liposomes was determined
using AIA-Pack E2 sta assay (Tosoh Corp., Tokyo, Japan) according
to the manufacturer’s instructions, using immunoanalyzer AIA 600 II
(Tosoh).
Determination of total RNA content in the
liposomes
The total amount of RNA in the liposomes was
determined using an electrochemical analyzer 663 VA Stand (Metrohm,
Herissau, Switzerland) using square wave voltammetry with a
three-electrode system, according to our previous study (16). For each electrochemical analysis, 10
μl of antisense RNA solution was used.
MCF-7 cell culture
MCF-7 epithelial breast cancer cells were maintained
in Dulbecco’s modified Eagle’s medium (DMEM; Gibco-BRL, Grand
Island, NY, USA), supplemented with 10% (v/v) fetal bovine serum
(Sigma-Aldrich), 1 mM sodium pyruvate, non-essential amino acids
and 100 μg/ml penicillin and streptomycin (Sigma-Aldrich).
Liposomes, containing antisense oligonucleotides at different
concentrations, were applied directly into the medium to treat the
cells and were labeled as L2, L3 and L4 and contained 1 μM and/or
10 μM of single or both types of oligonucleotides. L1 did not
contain any ASOs and served as a control. The treatment of MCF-7
cells was conducted for 12 h.
Cell proliferation assessment using
real-time cell-based assay (RTCA)
Real-time cell-based assay xCELLigence (Roche
Applied Science and ACEA Biosciences, San Diego, CA, USA) consisted
of four main components: RTCA analyser, RTCA station, RTCA computer
with integrated software and disposable E-plate 16. First, the
optimal seeding concentration for the proliferation and cytotoxic
assays was determined. After seeding, the total number of cells (in
200 μl medium of each well in the E-plate 16, the attachment,
proliferation and spreading of the cells) was monitored every 15
min. All the experiments were carried out for 65 h. The results are
expressed as cell viability using the software of the manufacturers
(Roche Applied Science and ACEA Biosciences).
Microscopy
Eclipse TS100 inverted phase contrast microscope
(Nikon Instruments, Amsterdam, The Netherlands) with a ×20
magnification lens was used to view the cell cultures in
cultivation flasks. The images were obtained directly using the
C-5060 camera (Olympus, Tokyo, Japan).
RNA isolation and reverse
transcription
High pure total-RNA isolation kit (Roche, Basel,
Switzerland) was used to isolate RNA from the treated cells. The
medium was removed and the samples were washed twice with 5 ml of
ice-cold PBS. The cells were scraped off, transferred to clean
tubes and centrifuged at 20,800 × g for 5 min at 4°C
(Microcentrifuge 5417R; Eppendorf AG, Hamburg, Germany). After this
step, lysis buffer was added and RNA isolation was carried out
according to the manufacturer’s instructions. The isolated RNA was
used for cDNA synthesis. The transcription of RNA (600 ng) was
carried out using Transcriptor First Strand cDNA synthesis kit
(Roche, Basel, Switzerland) according to the manufacturer’s
instructions. Prepared cDNA (20 μl) from total-RNA was diluted with
RNase-free water to 100 μl and subsequently 5 μl was directly
analyzed using the LightCycler 480 real-time PCR system (Roche
Applied Science).
Quantitative polymerase chain reaction
(q-PCR)
q-PCR was performed in triplicate using the TaqMan
gene expression assay system with the LightCycler 480 real-time PCR
system, and the amplified DNA was analyzed by the comparative Ct
method using β-actin as an endogenous control for metallothionein
MT1A, K-ras, TFF1 and NF-κB1 gene
expression quantification. The primer and probe sets for
β-actin (assay ID: Hs99999903_m1), MT1A
(Hs00831826_s1), K-ras (Hs00364284_g1), NF-κB1
(Hs00765730_m1) and TTF1 (Hs00907239_m1) were selected from
TaqMan gene expression assays (Life Technologies, Grand Island, NY,
USA). The thermal profile for q-PCR was as follows: initial
incubation at 50°C for 2 min followed by denaturation at 95°C for
10 min and then 45 cycles at 95°C for 15 sec and 60°C for 1 min.
The total volume of the reaction mixture was 20 μl.
Preparation of MCF-7 cells for
analyses
The harvested MCF-7 cells were frozen by liquid
nitrogen to disrupt their structure. The frozen sample was further
homogenized using ultrasonic homogenizer SONOPLUS mini20 (Bandelin
electronic, Berlin, Germany). Subsequently, 1 ml of 0.2 M phosphate
buffer (pH, 7.0) was added and the sample was homogenized for 5
min. The homogenate was further centrifuged using Microcentrifuge
5417R under the following conditions at 4°C for 15 min. Finally,
the supernatant was filtered through a membrane filter (0.45-μm
nylon filter disk; Millipore, Billerica, MA, USA) and analyzed.
Determination of the total protein
content
The total protein content was determined for results
standardization and was performed using SKALAB CBT 600T kit
(Skalab, Svitavy, Czech Republic) according to the manufacturer’s
instructions. For analysis, the BS-400 automated spectrophotometer
(Mindray, Schenzhen, China) was used.
Determination of the metallothionein
content with differential pulse voltammetry (DPV)
Differential pulse voltammetric (DPV) measurements
were performed with the 747 VA Stand instrument connected to the
693 VA Processor and 695 Autosampler (Metrohm, Herissau,
Switzerland), using a standard cell with three electrodes,
according to our previous study (17).
High performance liquid chromatography
coupled with electrochemical detector for determination of
glutathiones
Contents of reduced and oxidized glutathiones (GSH
and GSSG, respectively) were determined in the MCF-7 cell culture
after liposome treatment using high performance liquid
chromatography coupled with an electrochemical detector (HPLC-ED)
system under the conditions used in our previous study (18).
Spectrophotometric determination of
malondialdehyde (MDA)
Three hundred milliliters of the sample was mixed
with 10 μl of 0.5 M butylated hydroxytoluene in 96% ethanol (v/v)
and 310 μl of 20% trichloroacetic acid (v/v) prepared in 0.6 M HCl.
After 20 min of incubation on ice, the mixture was centrifuged at
11,000 rpm for 15 min (Microcentrifuge 5417R). Subsequently, 400 μl
of the supernatant was mixed with 800 μl of 30 mM thiobarbituric
acid and the mixture was incubated at 90°C for 30 min. After
cooling to room temperature, the absorbance of MDA was measured
using the BS-400 spectrophotometer at a wavelength of 535 nm.
Descriptive statistics
Mathematical analysis of the data and their
graphical interpretation were carried out using Microsoft
Excel®, Word® and PowerPoint®
(Microsoft, Redmond, WA, USA). The results are expressed as mean ±
standard deviation (SD) unless noted otherwise. Statistical
significance of the differences in gene expression was evaluated at
P=0.05.
Results and Discussion
The main challenge facing the approach using the
delivery of antisense oligonucleotides (ASO) to target tumor cells
is how to keep them intact to ubiquitous nucleases. Moreover, the
hydrophilic character and anionic backbone of antisense
oligonucleotides reduce the probability of their transfer across
membranes (10). Thereby, liposomes
were employed to protect ASOs in our study. Zhong et al
reported that, due to their properties, liposomes continue to
evolve as a promising tool for the delivery of potentially useful
therapeutic agents (11).
Therefore, we suggested estradiol-based liposomes operating on the
principle of natural affinity of hormone and estrogen receptor
proteins. Cholesterol (precursor of 17β-estradiol or
E2), which is commonly applied as the hydrophilic part
of the lipid bilayers of liposomes (19), was replaced by E2. This
complex offers similar hydrophilic properties and moreover the
possibility to target the liposomes towards transcription factors
(ERα and ERβ). Furthermore, it was shown that E2
enhances ASO uptake and therefore, the transfection efficiency due
to the interaction with fundamental components of the plasma
membrane, which causes the alteration in its physical properties
(20).
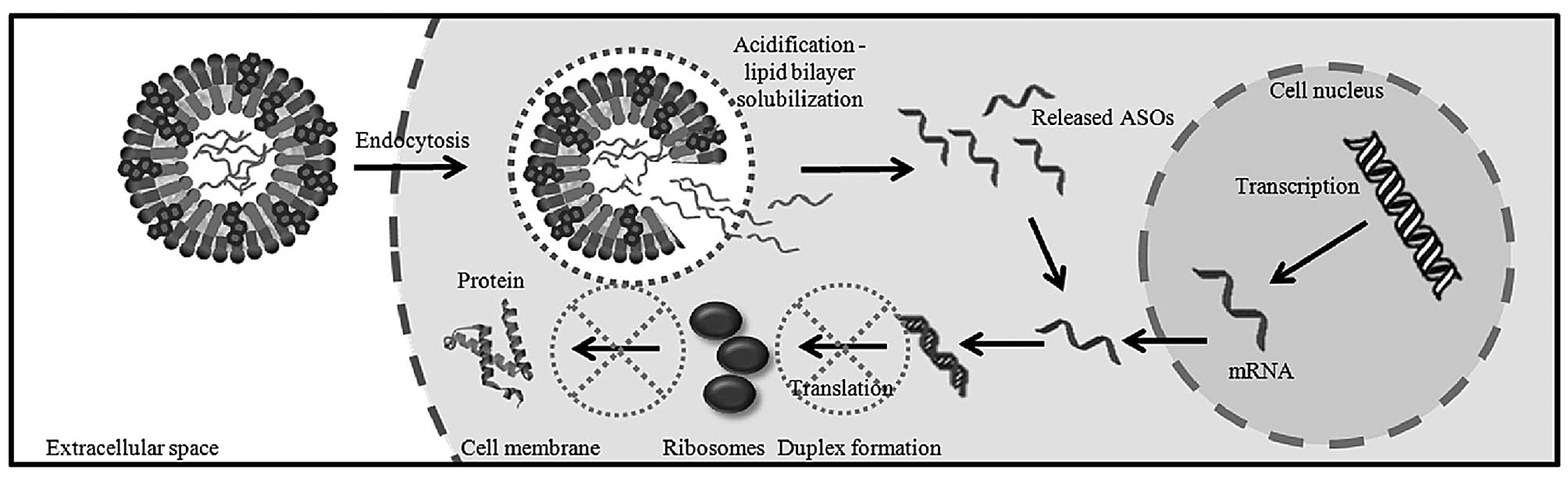
The scheme of E2-based liposome
interaction with cells is shown in Fig.
1. First, endocytosis occurs, since liposomes have been shown
to be preferentially internalized through endocytosis (21). Acidification of endosomes causes
release of ASOs into the cytoplasm and the formation of duplexes
with mRNA encoding target protein due to Watson-Crick hybridization
thus leading to mRNA degradation and/or inhibition of translation
(10).
17β-estradiol-based liposomes loaded with
ASOs
Following the lipidic thin-film hydration method
(22), we synthesized four
different types of 17β-estradiol-based liposomes loaded with ASOs
offering a powerful approach with which to selectively target and
inhibit gene expression, critical in disease progression. As shown
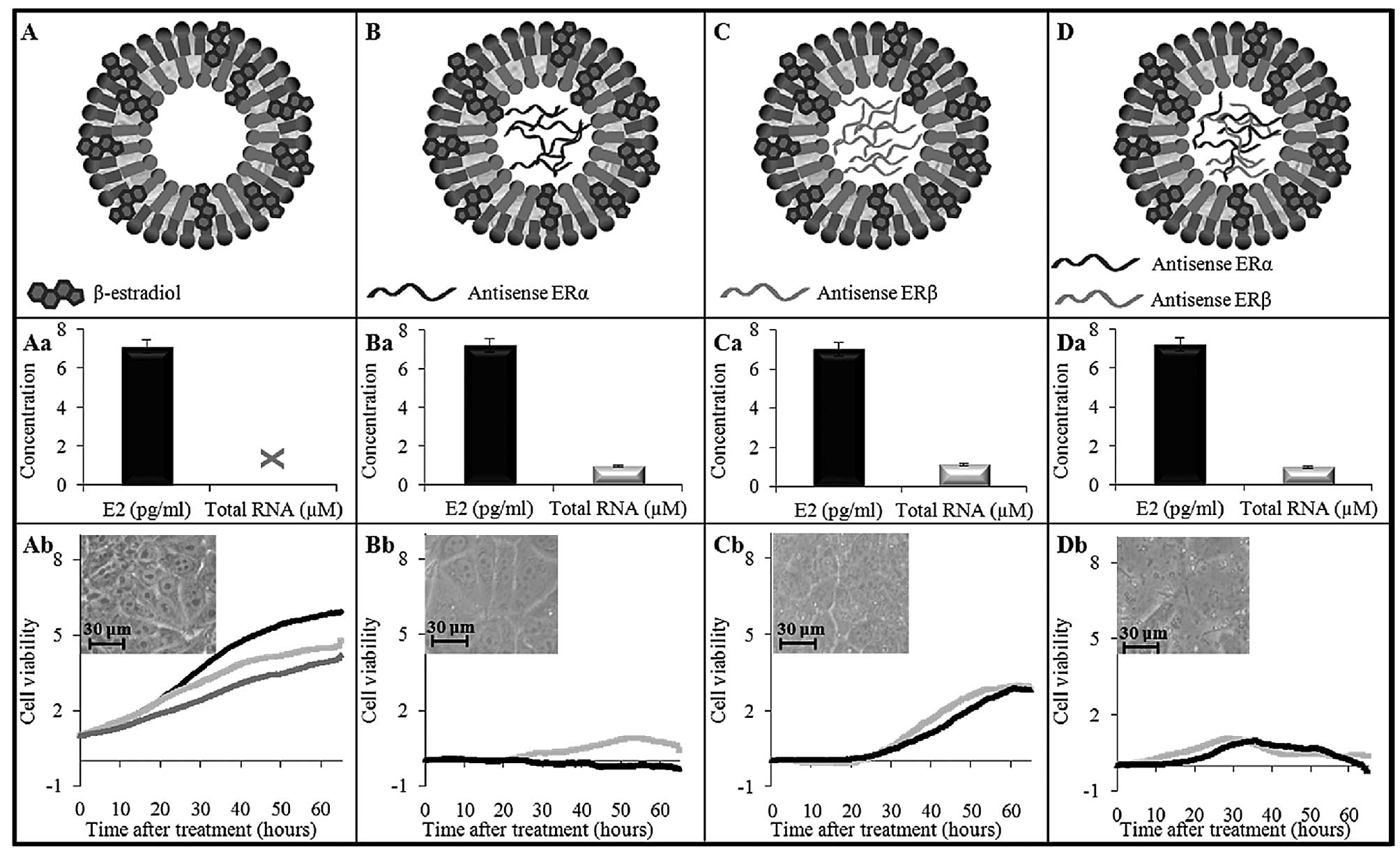
in Fig. 2A, liposomes L1 were
prepared with no ASO cargo and were considered as the control.
Other types of liposomes were prepared in the same manner,
containing E2 in their lipidic envelopes, but moreover
they were loaded with antisense ERα (Fig. 2B) named liposome L2, antisense ERβ
(Fig. 2C) named liposome L3 and
with both types of antisense sequences (Fig. 2D) named liposome L4. The antisense
sequences used in our study are complementary to mRNAs encoding
human transcription factors (ERα and/or β) and thus they are able
to target and modulate the receptor’s responses.
To obtain the basic characteristics of liposome
transporters, the hormone content was evaluated using
immunochemistry and total RNA content using electrochemistry. It
clearly follows from the results obtained that there was no
distinct variation in 17β-estradiol content among the four types of
liposomes ranging between 7–7.2 pg/ml with an average of 7.125
pg/ml (Fig. 2Aa–Da). Subsequently,
electrochemistry was employed for determination of the total RNA
amount in the estradiol-based liposomes and it was revealed that
unlike L1 (Fig. 2Aa), the other
liposomes (L2, L3 and L4, respectively) contained concentrations
from 0.91 to 1.12 μM (average of 0.993 μM) of total-RNA (volume of
solution, used for measurement: 100 μl). This finding approximately
corresponds with the concentrations calculated prior to liposome
synthesis (1 μM). Thus, we confirmed that our lipidic thin-film
hydration method of synthesis works properly.
Antiproliferative effect of
E2-based liposomes loaded with ASOs
The real-time cell-based assay (RTCA) was carried
out to monitor the proliferation of MCF-7 epithelial breast cancer
cells after application of the liposomes. The MCF-7 cells were
derived from estrogen receptor-positive ductal carcinoma (23) and thus we chose this cell line for
monitoring cell viability influenced by the estradiol-based
liposomes. As shown in Fig. 2Ab,
the application of increasing volumes of L1 (0, 90 or 900 μl)
increased the proliferative effect (expressed as cell viability) of
MCF-7 cells. This finding was expected due to the proliferative
properties of MCF-7 cells in response to E2 as
previously described by Lacroix and Leclercq (23) and Resende et al (24).
To reflect the real antiproliferative effect of
ASOs, we subtracted the cell viability of the controls (L1) from
the test samples (Fig. 2Bb–Db).
Antisense against ERα was shown to decrease the proliferative
ability of MCF-7 cells, since their proliferation is controlled by
ERα-mediated gene regulation as described previously (25). While E2 stimulates
proliferation via ERα, the signaling via ERβ inhibits proliferation
and promotes apoptosis (26). It
was shown in transfection studies in breast and colon cancer cells
that the lack of ERβ proteins leads to increased cell proliferation
both in culture and in vivo xenografts (27,28).
Based on this information, ERβ was suggested to be a pro-apoptotic
tumor suppressor (29) and thus
alteration of protein translation subsequently elevated MCF-7 cell
proliferation (Fig. 2Cb). Notably,
the synergistic effect of both ASOs was shown to also have
antiproliferative effects, but after a relatively long exposure
time (Fig. 2Db). This phenomenon
may be caused by a strong duplexing effect of antisense
oligonucleotides complementary to mRNA encoding ERα translation and
conversely only a weak complementary affinity to mRNA encoding ERβ.
Therefore, despite the treatment with ASOs, the translation pathway
was still able to produce ERβ proteins at threshold
concentrations.
Liposomes are preferentially internalized through
caveolae-dependent endocytosis (21), where the main role is played by
acidification of the endocytic vesicle permitting release of
antisense oligonucleotides rather than liposome membrane fusion.
Therefore, in the micrographs of MCF-7 cells after liposome
application, it can be observed that the cell size was increased
due to endosome formation (insets in Fig. 2Ab–Db).
Influence of E2-based
liposomes on MCF-7 cell selected gene expression
First, we decided to monitor the expression activity
of the MT1A gene encoding metallothionein (MT) protein. MT
is primarily responsible for metal sequestration and
detoxification. It was previously shown that metallothionein
expression is modulated also by E2 in in vivo
studies using fish experimental models (30,31),
where a positive correlation between MT1A expression and
E2 levels was described in muscle tissue in contrast to
liver and intestines. Our results revealed that the application of
E2-based liposomes with no cargo elevated the
MT1A expression by more than 60% (90 μl of L1 added) or 100%
(900 μl of L1 added), when compared with the control (Fig. 3A), determined by expression of the
gene without application of liposomes standardized to β-actin
(labeled as 0 μl). The increase in gene expression may be
attributed to a defense mechanism in response to oxidative stress
caused by the presence of E2 (32). Importantly, after the application of
liposomes containing ASOs (1 and 10 μM) against ERα protein (L2),
it was shown that MT1A expression was downregulated (by
>15 and/or 55%, respectively, Fig.
3B). This phenomenon points to a successful steric blockade of
the ERα translational start site resulting in protein depletion and
therefore downregulation of transcription related to
E2-ER binding with subsequent effect on the MT1A
gene. After the application of ASOs (10 μM) against ERβ (L3), a
significant increase in expression of the metallothionein gene was
observed (Fig. 3C). Proper
translation of the ERβ protein seems to be important for the
regulation of the MT1A gene. Although the role of ERβ is
still under investigation in breast cancer, it was previously shown
that ERβ acts as a tumor suppressor (33). Moreover, we offer evidence that ERβ
downregulation directly influences the expression of the
MT1A gene in MCF-7 cells. After treatment with the liposomes
carrying both ASOs (L4), we observed a similar effect on
MT1A (~75 and/or 85% downregulation after addition of 1 and
10 μM ASOs, respectively) as in the case of treatment with ASOs
against ERα (Fig. 3D). Low
threshold concentrations were required to trigger an antisense
effect of ERα ASOs. On the other hand, ERβ functions were not
significantly influenced when compared with ERα and thus the
desired effect was observed.
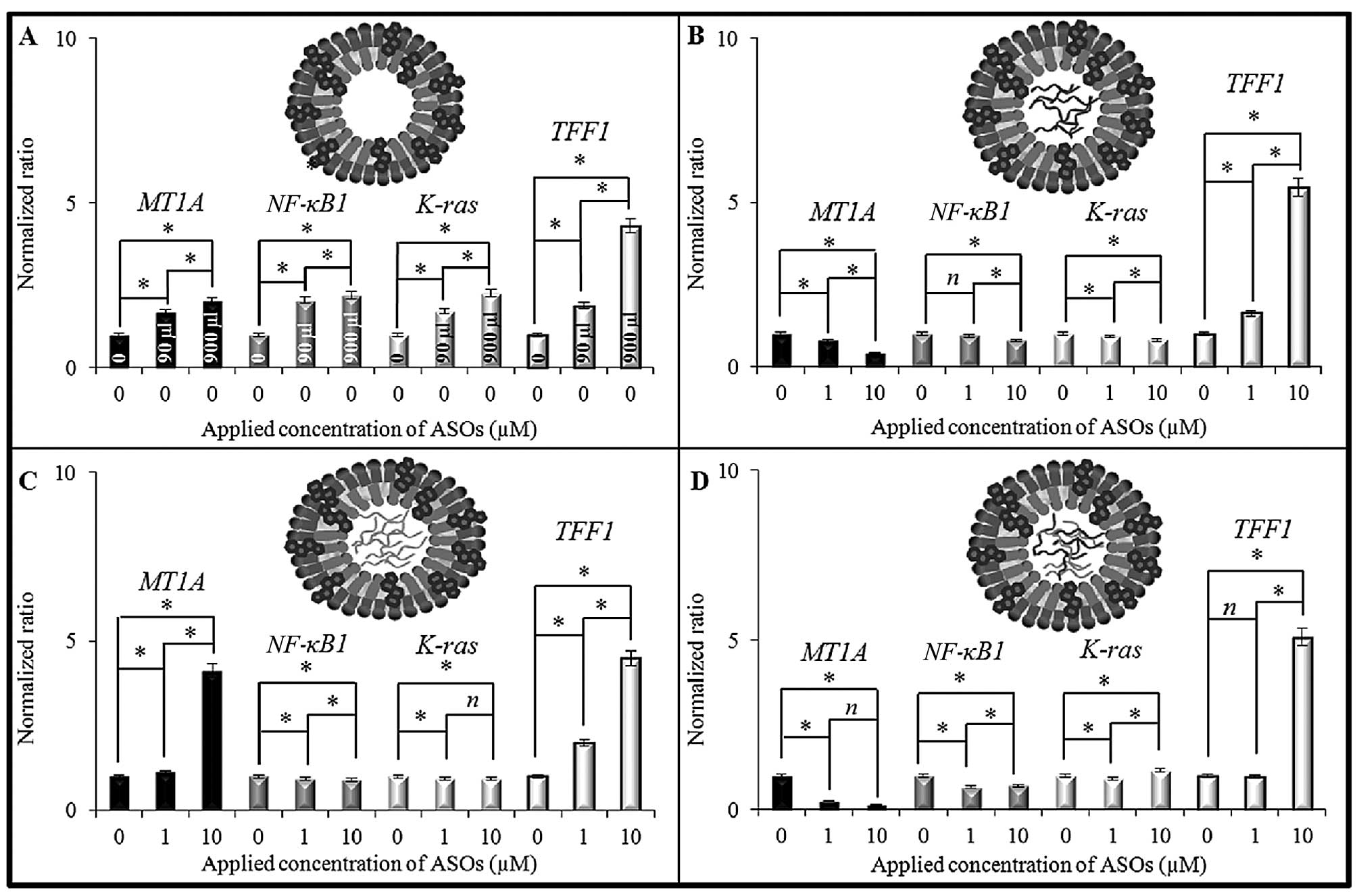 | Figure 3Effect of various 17β-estradiol
liposomes containing ER antisense sequences on MCF-7 cell culture.
The expression levels of MT1A, NF-κB1, K-ras
and TFF1 genes were observed. Schematic representations are
connected to the gene expression and show the scheme of
E2-based liposomes: (A) with no antisense
oligonucleotide (control), (B) containing ERα ASO, (C) containing
ERβ ASO and (D) containing both ERα and ERβ ASOs. Liposomes were
applied at concentrations of 0, 1 and/or 10 μM into the medium. In
the case of L1 (no ASOs) 0, 90 and 900 μl of the liposome solution
was applied, corresponding to the amounts of
E2-liposomes present in the case of 0, 1 and 10 μM ASOs
in L2, L3 and L4. *Statistically significant differences between
gene expression at P=0.05 level; n indicates no statistical
significance. ASOs, antisense oligonucleotides. |
Furthermore, the expression activity of the
NF-κB1 gene was determined, as the blocking of NF-κB1
can cause tumor cells to stop proliferation, to die, or to become
more sensitive to the action of chemotherapeutics (34). NF-κB1 transcription factor
influences various molecules involved in oncogenesis such as matrix
metalloproteinases (MMPs), their activators and/or inhibitors, cell
adhesion molecules and angiogenic factors (35). In the present study, NF-κB1
expression was upregulated in MFC-7 cells after treatment with L1
(~100 and 120% after the application of 90 and 900 μl,
respectively) (Fig. 3). This
indicates the ability of E2 to influence MCF-7 cells
undesirably. When compared with the application of ASOs, the
highest effect was observed after treatment with ERα, where
downregulation of ~5% was observed at 1 μM ASOs and ~20% at 10 μM
ASOs (Fig. 3B). The effect of both
ASOs was also shown to trigger downregulation of NF-κB1 by
~35% at 1 μM ASOs or 30% at 10 μM ASOs (Fig. 3D). The ASOs against ERα were found
to influence NF-κB1, because a downregulation ~9% was
observed in the case of 1 μM ASOs and further 10% in the case of 10
μM ASOs (Fig. 3C), when compared
with the L1 liposome. Nevertheless, the best results were obtained
using liposome L2 and L4. This phenomenon suggests that the
presence of ASOs against ERα is more crucial for the downregulation
of expression of NF-κB1 than ERβ. Based on these results, it
can be concluded that ASOs may offer the possibility to inhibit
NF-κB1 gene activity and thus increase the efficiency of
chemotherapeutics applicable in breast cancer treatment.
Moreover, we determined K-ras gene
expression, which was influenced in a similar way as NF-κB1.
The expression of K-ras was elevated with increasings
amounts of L1 applied (~70 and 125% at 1 and 10 μM of ASOs,
respectively) (Fig. 3A). The
application of ASOs exhibited a downregulatory influence on
K-ras expression (Fig.
3B–D). The greatest effect was determined at 10 μM antisense
against ERα (Fig. 3B). In the case
of ASOs against ERβ, no significant differences between 0 and 1 μM
application were observed at P<0.05; and moreover, a 10 μM
concentration resulted in a downregulation of only 7.5% when
compared to 0 μM (Fig. 3C). The ERα
ASO influence on K-ras gene expression may serve as a
potential tool to inhibit K-ras activity, as K-ras and ER proteins
are linked in K-ras 4B-mediated cell transformation by
p53-independent modulation of MDM2 functions as identified by Kato
et al (36). Because of the
fact that ER can be considered as one of the effectors of the
Ras/Raf signaling pathway involving in tumorigenesis (37), the inhibition of estrogen receptor
activity subsequently triggers the downregulation of the expression
of K-ras. However, the relationship of these two factors is still
not satisfactorily elucidated.
TFF1 gene encoding pS2 protein was also
determined as the only one that was not downregulated by our
antisense strategy. When comparing control liposome L1 (Fig. 3A) with liposomes loaded with ASOs
(Fig. 3B–D), it is obvious that
E2, but not ASOs, plays a crucial role in TFF1
expression due to the significant differences (P<0.05) between
0, 1 and 10 μM concentrations of ASOs observed for all liposomes
including L1. The induction phenomenon of TFF1 expression
was found to be primarily due to a response to estrogens and was
mediated by the binding of the ER-E2 complex to a 13-bp
near-palindromic ERE located 400 bases upstream of the TFF1
transcription start site (38,39).
Hence, the presence of E2, which forms a complex with
transcription factors, directly influences also the TFF1
transcription start site. As numerous studies have shown that
TFF1 does not act as an oncogene in the mammary gland, but
conversely, exerts a beneficial function during malignant processes
in ER-positive breast tumors (38,40–42),
our results point to the potential positive effect caused by our
antisense strategy.
Oxidative stress determination after
E2-based liposome application
Numerous signaling pathways, which are linked to
tumorigenesis, can also regulate the metabolism of ROS through
various mechanisms. As mentioned in a review by Gorrini et
al, high ROS levels are generally harmful to cells, and the
redox status of cancer cells is usually higher than that in healthy
cells due to metabolic and signaling aberrations (43). For this reason, we assessed
oxidative stress indicators, particularly glutathione,
metallothionein and malondialdehyde to obtain a more detailed
insight into the manner that E2-based liposomes act on
MCF-7 cells at the molecular level.
It has been reported that elevated GSH levels are
present in various types of tumors and thus neoplastic tissues are
more resistant to chemotherapy (44). In malignant cells, an increased GSH
level is associated with a proliferative response and is essential
for cell cycle progression; yet, the molecular mechanism of how GSH
modulates cell proliferation remains largely speculative. As shown
in Fig. 4A, the addition of
liposomes L1, L3 and L4 caused an elevation in GSH levels due to
the presence of E2 in all cases and moreover ASOs
against ERβ in the case of L3 and L4. These data also corroborate
the RTCA results, where the presence of E2 and ASOs
towards ERβ resulted in the increased proliferation of MCF-7 cells.
On the other hand, the application of L2 (Fig. 4A) dramatically decreased the
glutathione redox ratio (similar trend to the cell index of MCF-7).
Hence, the application of E2-based liposomes carrying
ASOs may be helpful to decrease the resistance to chemotherapy
caused by high levels of GSH in tumor tissues.
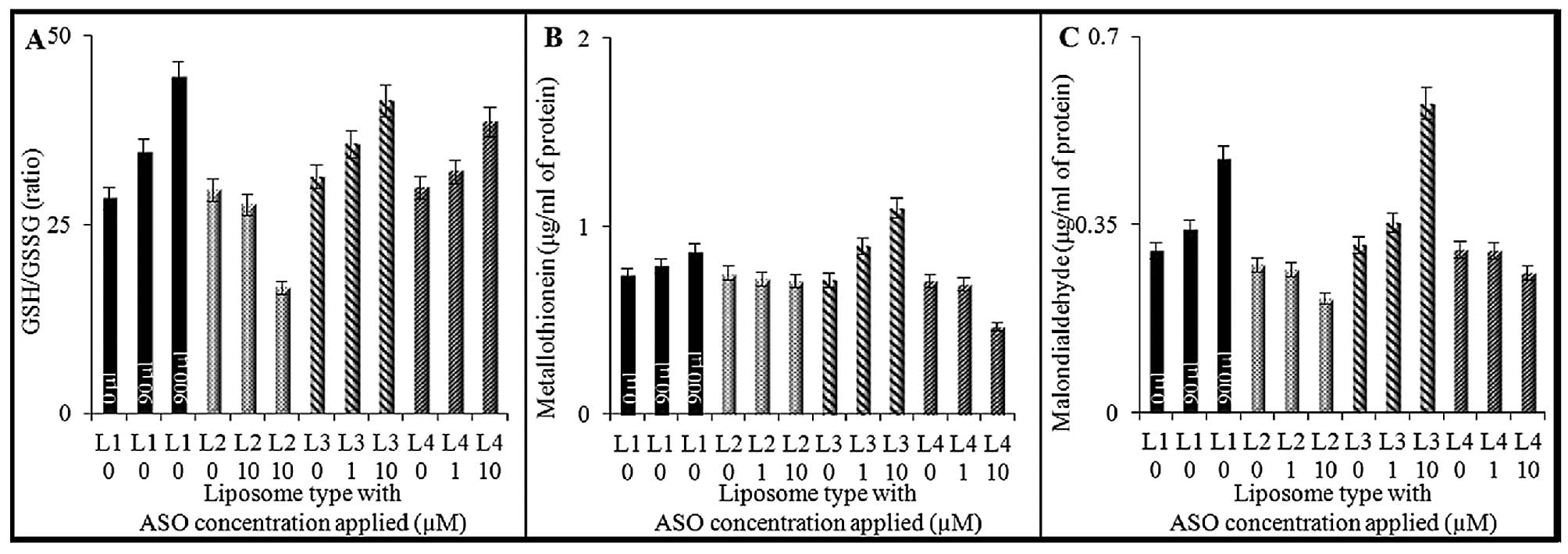 | Figure 4Expression of oxidative stress
markers determined in the MCF-7 cell line after treatment with the
different types of liposomes. (A) GSH/GSSG ratio, (B)
metallothionein and (C) malondialdehyde. Liposomes are indicated as
follows: L1, liposome containing E2 with no antisense sequence; L2,
liposome containing E2 with antisense sequence ERα; L3,
liposome containing E2 with antisense sequence ERβ and
L4, liposome containing E2 and both ERα and ERβ
antisense sequences. Liposomes were applied at concentrations of 0,
1 and/or 10 μM into the medium to obtain a final volume of 6 ml. In
the case of L1 (no ASOs), 0, 90 and/or 900 μl of liposome solution
was applied corresponding to the amount of liposomes present in the
case of 0, 1 and/or 10 μM ASOs in L2, L3 and L4. ASOs, antisense
oligonucleotides. |
Furthermore, metallothionein levels were assessed
(Fig. 4B). Liposomes L2 and L4 were
found to have a suppressive effect on protein formation. In the
case of L2, an application of 0 μM ASOs resulted in 0.75 μg of
metallothionein (standardized to 1 ml of total protein); 1 μM ASOs
resulted in 0.72 μg of metallothionein to 1 ml of total protein and
10 μM ASOs resulted in 0.7 μg of metallothionein to 1 ml of total
protein (Fig. 4B). The application
of L4 at a concentration of 0 μM ASOs resulted in 0.71 μg of
metallothionein to 1 ml of total protein; 1 μM ASOs resulted in
0.69 μg of metallothionein to 1 ml of total protein and 10 μM ASOs
resulted in 0.46 μg of metallothionein to 1 ml of total protein.
This corresponds to the results obtained from qRT-PCR. Surowiak
et al reported that the elevated level of
low-molecular-weight metallothionein in estrogen receptor-positive
breast tumors may be explained by endoplasmic reticulum damage or
by its malfunction and can be found typically in breast cancer
cases with less favorable prognosis (41). It was shown that the effective
application of ASOs may serve as a potential tool to mitigate the
damage of endoplasmic reticulum and thus decrease the levels of
metallothionein. Moreover, metallothionein was shown to be one of
the potential co-factors causing the chemoresistance of tumors to
platinum-based cytostatics and to anthracyclines (45–48).
Hence, the downregulation of MT expression via ASO targeting of the
MT1A gene may enhance tumor cell susceptibility to various
chemotherapeutic agents.
Malondialdehyde (MDA) is one of the final
decomposition products of lipid peroxidation and it is also formed
as a product of the cyclooxygenase reaction in prostaglandin
metabolism (49). It clearly
follows from the results obtained that our novel strategy for
antisense therapy was efficient in the case of MDA, as ASOs against
ERα (10 μM application) showed a decrease in its level; >0.06 μg
of MDA to 1 ml of total protein when compared to the control
(Fig. 4C). Similarly, a slightly
reducing effect was also observed in the case of the combination of
both ASOs. As MDA reacts with double-stranded DNA to form mutagenic
adducts and lipid peroxidation appears to be a major source of
endogenous DNA damage in humans significantly contributing to tumor
development (50), the reduction in
lipid peroxidation end products may be one of the very significant
effects of antisense therapy suggested by us.
In conclusion, we suggest four types of liposomes
based on E2 placed in their lipid bilayer substituting
commonly used cholesterol. As a cargo, antisense oligonucleotides
towards estrogen receptor α and β were applied, and it was revealed
that the liposomes synthesized by us showed a beneficial response
in regards to a decrease in proliferation of MCF-7 cells. Moreover,
it was shown that besides TTF1, the antisense strategy
downregulated evaluated genes (MT1A, K-ras and
NF-κB1) and also decreased the levels of oxidative stress
indicators (MT, GSH/GSSG and MDA). We conclude that antisense ERα
efficiently binds to ERα mRNA and thus the translation of nascent
ERα protein is inhibited. Furthermore, the therapeutic strategy
based on our ASOs entrapped in E2-based liposomes offers
the possibility to enhance tumor susceptibility to
chemotherapeutics via GSH oxidation and downregulation of MT
expression through ASO targeting of the MT1A gene.
Therefore, ASOs can be useful in cases where single treatment
therapy does not show adequate results and chemosensitization of
the tumor is required.
Acknowledgements
This study was supported by the Internal Grant
Agency of the University of Veterinary and Pharmaceutical Sciences
Brno, Czech Republic (Project 4/2013/FVHE) and CEITEC
CZ.1.05/1.1.00/02.0068.
References
|
1
|
Zamecnik PC and Stephenson ML: Inhibition
of Rous sarcoma virus replication and cell transformation by a
specific oligode-oxynucleotide. Proc Natl Acad Sci USA. 75:280–284.
1978. View Article : Google Scholar
|
|
2
|
Warzocha K: Antisense strategy in
hematological malignancies. Cytokines Cell Mol Ther. 5:15–23.
1999.PubMed/NCBI
|
|
3
|
Ruden M and Puri N: Novel anticancer
therapeutics targeting telomerase. Cancer Treat Rev. 39:444–456.
2013. View Article : Google Scholar
|
|
4
|
Wu Y, Zhang Y, Wang M, et al:
Downregulation of HER3 by a novel antisense oligonucleotide,
EZN-3920, improves the antitumor activity of EGFR and HER2 tyrosine
kinase inhibitors in animal models. Mol Cancer Ther. 12:427–437.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Panyam J and Labhasetwar V: Biodegradable
nanoparticles for drug and gene delivery to cells and tissue. Adv
Drug Deliv Rev. 55:329–347. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Jones SE: Metastatic breast cancer: the
treatment challenge. Clin Breast Cancer. 8:224–233. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Xie QC, Hu YD, Wang LL, et al: The
co-transfection of p16(INK4a) and p14(ARF) genes into human lung
cancer cell line A549 and the effects on cell growth and
chemosensitivity. Colloids Surf B Biointerfaces. 46:188–196. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Stein CA: Two problems in antisense
biotechnology: in vitro delivery and the design of antisense
experiments. Biochim Biophys Acta. 1489:45–52. 1999. View Article : Google Scholar
|
|
9
|
Balbino TA, Azzoni AR and de la Torre LG:
Microfluidic devices for continuous production of pDNA/cationic
liposome complexes for gene delivery and vaccine therapy. Colloid
Surf B Biointerfaces. 111:203–210. 2013. View Article : Google Scholar
|
|
10
|
Skoblov Mlu: Prospects of antisense
therapy technologies. Mol Biol. 43:984–998. 2009.(In Russian).
View Article : Google Scholar
|
|
11
|
Zhong J, Yao X, Li DL, et al: Large scale
preparation of midkine antisense oligonucleotides nanoliposomes by
a cross-flow injection technique combined with ultrafiltration and
high-pressure extrusion procedures. Int J Pharm. 441:712–720. 2013.
View Article : Google Scholar
|
|
12
|
Ye S, Yang W, Wang Y, et al: Cationic
liposome-mediated nitric oxide synthase gene therapy enhances the
antitumor effects of cisplatin in lung cancer. Int J Mol Med.
31:33–42. 2013.
|
|
13
|
Kim TI and Kim SW: Bioreducible polymers
for gene delivery. React Funct Polym. 71:344–349. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Xu M, Kumar D, Srinivas S, et al:
Parenteral gene therapy with p53 inhibits human breast tumors in
vivo through a bystander mechanism without evidence of toxicity.
Hum Gene Ther. 8:177–185. 1997. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Nakase M, Inui M, Okumura K, Kamei T,
Nakamura S and Tagawa T: p53 gene therapy of human osteosarcoma
using a transferrin-modified cationic liposome. Mol Cancer Ther.
4:625–631. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Krejcova L, Hynek D, Kopel P, et al:
Development of a magnetic electrochemical bar code array for point
mutation detection in the H5N1 neuraminidase gene. Viruses.
5:1719–1739. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Kensova R, Kremplova M, Smerkova K, et al:
Interactions of platinum-based cytostatics with metallothionein
revealed by electrochemistry. Int J Electrochem Sci. 8:4472–4484.
2013.
|
|
18
|
Krystofova O, Sochor J, Zitka O, et al:
Effect of magnetic nanoparticles on tobacco BY-2 cell suspension
culture. Int J Environ Res Public Health. 10:47–71. 2012.
View Article : Google Scholar
|
|
19
|
Misra SK, Naz S, Kondaiah P and
Bhattacharya S: A cationic cholesterol based nanocarrier for the
delivery of p53-EGFP-C3 plasmid to cancer cells. Biomaterials.
35:1334–1346. 2014. View Article : Google Scholar
|
|
20
|
Taylor AH, Pringle JH, Bell SC and
Al-Azzawi F: Specific inhibition of estrogen receptor alpha
function by antisense oligodeoxyribonucleotides. Antisense Nucleic
Acid Drug Dev. 11:219–231. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Ruyra A, Cano-Sarabia M, Mackenzie SA,
Maspoch D and Roher N: A novel liposome-based nanocarrier loaded
with an LPS-dsRNA cocktail for fish innate immune system
stimulation. PLoS One. 8:e763382013. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Dinu-Pirvu C, Ferdes M, Butu A, Ortan A
and Ghica MV: Physicochemical investigation of low soluble
biocompounds entrapped in lipid carriers. Farmacia. 61:182–192.
2013.
|
|
23
|
Lacroix M and Leclercq G: Relevance of
breast cancer cell lines as models for breast tumours: an update.
Breast Cancer Res Treat. 83:249–289. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Resende FA, de Oliveira AP, de Camargo MS,
Vilegas W and Varanda EA: Evaluation of estrogenic potential of
flavonoids using a recombinant yeast strain and MCF7/BUS cell
proliferation assay. PLoS One. 8:e748812013. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Hu ZZ, Kagan BL, Ariazi EA, et al:
Proteomic analysis of pathways involved in estrogen-induced growth
and apoptosis of breast cancer cells. PLoS One. 6:e204102011.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Yakimchuk K, Jondal M and Okret S:
Estrogen receptor α and β in the normal immune system and in
lymphoid malignancies. Mol Cell Endocrinol. 375:121–129. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Tu ZZ, Ma Y, Tian J, et al: Estrogen
receptor β potentiates the antiproliferative effect of raloxifene
and affects the cell migration and invasion in HCT-116 colon cancer
cells. J Cancer Res Clin Oncol. 138:1091–1103. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Li H, Tu Z, An L, Qian Z, Achilefu S and
Gu Y: Inhibitory effects of ERβ on proliferation, invasion, and
tumor formation of MCF-7 breast cancer cells - prognostication for
the use of ERβ-selective therapy. Pharm Biol. 50:839–849. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Hartman J, Lindberg K, Morani A, Inzunza
J, Strom A and Gustafsson JA: Estrogen receptor beta inhibits
angiogenesis and growth of T47D breast cancer xenografts. Cancer
Res. 66:11207–11213. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Costa DD, Neto FF, Costa MD, et al:
Vitellogenesis and other physiological responses induced by
17-beta-estradiol in males of freshwater fish Rhamdia quelen. Comp
Biochem Physiol C Toxicol Pharmacol. 151:248–257. 2010. View Article : Google Scholar
|
|
31
|
Woo S, Won H, Lee A and Yum S: Oxidative
stress and gene expression in diverse tissues of Oryzias javanicus
exposed to 17β-estradiol. Mol Cell Toxicol. 8:263–269. 2012.
View Article : Google Scholar
|
|
32
|
Bhat HK, Calaf G, Hei TK, Loya T and
Vadgama JV: Critical role of oxidative stress in estrogen-induced
carcinogenesis. Proc Natl Acad Sci USA. 100:3913–3918. 2003.
View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Li W, Jia M, Qin X, Hu J, Zhang X and Zhou
G: Harmful effect of ERβ on BCRP-mediated drug resistance and cell
proliferation in ERα/PR-negative breast cancer. FEBS J.
280:6128–6140. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Garg A and Aggarwal BB: Nuclear
transcription factor-kappaB as a target for cancer drug
development. Leukemia. 16:1053–1068. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Chen F, Castranova V, Shi X and Demers LM:
New insights into the role of nuclear factor-kappaB, a ubiquitous
transcription factor in the initiation of diseases. Clin Chem.
45:7–17. 1999.PubMed/NCBI
|
|
36
|
Kato K, Horiuchi S, Takahashi A, et al:
Contribution of estrogen receptor alpha to oncogenic K-Ras-mediated
NIH3T3 cell transformation and its implication for escape from
senescence by modulating the p53 pathway. J Biol Chem.
277:11217–11224. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Tu Z, Gui L, Wang J, Li X, Sun P and Wei
L: Tumorigenesis of K-ras mutation in human endometrial carcinoma
via upregulation of estrogen receptor. Gynecol Oncol. 101:274–279.
2006. View Article : Google Scholar
|
|
38
|
Buache E, Etique N, Alpy F, et al:
Deficiency in trefoil factor 1 (TFF1) increases tumorigenicity of
human breast cancer cells and mammary tumor development in
TFF1-knockout mice. Oncogene. 30:3261–3273. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
May FE and Westley BR: Trefoil proteins:
their role in normal and malignant cells. J Pathol. 183:4–7. 1997.
View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Corte MD, Tamargo F, Alvarez A, et al:
Cytosolic levels of TFF1/pS2 in breast cancer: their relationship
with clinical-pathological parameters and their prognostic
significance. Breast Cancer Res Treat. 96:63–72. 2006. View Article : Google Scholar
|
|
41
|
Surowiak P, Matkowski R, Materna V, et al:
Elevated metallothionein (MT) expression in invasive ductal breast
cancers predicts tamoxifen resistance. Histol Histopathol.
20:1037–1044. 2005.PubMed/NCBI
|
|
42
|
Markicevic M, Petrovic A, Kanjer K,
Neskovic-Konstantinovic Z and Nikolic-Vukosavujevic D:
Estrogen-regulated cut-off values of pS2 and cathepsin D expression
in breast carcinomas. Adv Exp Med Biol. 617:341–348. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Gorrini C, Harris IS and Mak TW:
Modulation of oxidative stress as an anticancer strategy. Nat Rev
Drug Discov. 12:931–947. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Traverso N, Ricciarelli R, Nitti M, et al:
Role of glutathione in cancer progression and chemoresistance. Oxid
Med Cell Longev. 2013:9729132013. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Hishikawa Y, Abe S, Kinugasa S, et al:
Overexpression of metallothionein correlates with chemoresistance
to cisplatin and prognosis in esophageal cancer. Oncology.
54:342–347. 1997. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Endo T, Yoshikawa M, Ebara M, et al:
Immunohistochemical metallothionein expression in hepatocellular
carcinoma: relation to tumor progression and chemoresistance to
platinum agents. J Gastroenterol. 39:1196–1201. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Yap X, Tan HY, Huang J, et al:
Over-expression of metallothionein predicts chemoresistance in
breast cancer. J Pathol. 217:563–570. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Kizek R, Adam V, Hrabeta J, et al:
Anthracyclines and ellipticines as DNA-damaging anticancer drugs:
recent advances. Pharmacol Ther. 133:26–39. 2012. View Article : Google Scholar
|
|
49
|
Klein S, Dell’Arciprete ML, Wegmann M, et
al: Oxidized silicon nanoparticles for radiosensitization of cancer
and tissue cells. Biochem Biophys Res Commun. 434:217–222. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Marnett LJ: Oxy radicals, lipid
peroxidation and DNA damage. Toxicology. 181–182:219–222. 2002.
View Article : Google Scholar
|