Introduction
Breast cancer is the most common and the leading
cause of death from all cancers among women in China (1). Mortality from breast cancer has been
rising in recent years. Chemotherapy is still the preferred
strategy applied for breast cancer along with surgery and
radiotherapy. However, the non-targeted delivery of
chemotherapeutic drugs greatly affects normal cells (2), and to a great extent limits the drug
dosage to tumor cells altogether compromising the therapeutic
effects of the drugs.
Nanoparticles (NPs) are increasingly considered as a
powerful tool for chemotherapeutic drug delivery (3–5).
Targeted drug delivery using NPs functionalized with targeting
ligands is one of the most promising means which augments the
selectivity and drug dosage to tumor cells leading to an enhanced
therapeutic index of drugs. There are many types of NPs for drug
delivery, such as polymeric micelles (6,7),
liposomes (8–10), and lipoprotein-based carriers
(11,12). The most common vehicle currently
used for targeted drug delivery is the liposome. They are
biocompatible and biodegradable and can be designed to avoid
clearance.
Systematic evolution of ligands by exponential
enrichment (SELEX) is an efficient strategy for high throughput
in vitro selection of bio-library pool (13,14).
In recent years, a modified SELEX process using whole living cells
as targets was developed and designated as cell-SELEX (15,16),
which can select aptamers from unknown complex targets.
In the present study, we aimed to achieve targeted
delivery of doxorubicin to 4T1 tumors by conjugating 4T1 tumor
cell-specific aptamers which were selected using 4T1 cells
(positive selection) and 67NR cells (negative selection). The
aptamers with the highest binding affinity and specificity as
detected by flow cytometry were carried by doxorubicin-loaded
DOTAP:DOPE NPs. Various in vitro and in vivo studies
conducted to date have demonstrated that the AS1411 aptamer
enhances the uptake of NPs in 4T1 tumor cells. Hence, these
anti-4T1 cell aptamers can be effectively used to target NPs to 4T1
cancer cells.
Materials and methods
Cell culture
Mouse mammary breast cancer 4T1, 4TO7 and 67NR cells
were purchased from the American Type Culture Collection (ATCC) and
maintained in our laboratory. Normal mouse mammary gland NMuMG
cells were purchased from ATCC and cultured in Dulbecco's modified
Eagle's medium (DMEM) with 10 µg/ml of insulin and 10% fetal
bovine serum (FBS). Mouse Lewis lung carcinoma cell line LL/2,
mouse hepatoma cell line Hepa 1–6 and mouse B cell lymphoma cell
line WEHI-231 were purchased from ATCC and cultured in DMEM with
10% FBS. Mouse colon cancer CT26 cells were cultured in RPMI-1640
medium with 10% FBS.
Random single-strand DNA library and
primers
A 60-mer ssDNA library containing molecules with a
30-base random sequence flanked by two primer hybridization sites
was chosen as the initial library (5′-CCT GAC AGT CGA GAC-N30-CAC
CGG GGT CCT AGG-3′). The FAM-labeled forward primer (5′-FAM-CCT GAC
AGT CGA GAC-3′), and biotin-labeled reverse primer (5′-biotin-CCT
AGG ACC CCG GTG-3′) were used in PCR to obtain the double-labeled
DNA and to separate the single-stranded DNA by streptavidin-coated
magnetic particles. All sequences were synthesized by Sangon and
purified by reverse phase HPLC.
Cell-SELEX
The 4T1 cells with high invasive and metastatic
potential were used as target cells, and non-invasive and
non-metastatic 67NR cells were used as the negative selection cells
in the present study. ssDNA (10 nmol) library was denatured at
100°C for 5 min and kept on ice for 10 min and dissolved in 500
µl phosphate-buffered saline (PBS) buffer with 1 M
MgCl2, 0.1 mg/ml yeast tRNA, 1 mg/ml BSA and 0.1 mg/ml
salmon sperm DNA. The 4T1 cells (5×106) were washed,
dissociated and incubated with the ssDNA library on ice for 1 h.
After 3 washings, cells were re-suspended in 200 µl
DNase-free water. The cell-ssDNA mixture was heated at 100°C for 5
min, centrifuged at 4,000 rpm for 5 min, and the supernatant
containing eluted ssDNA was collected and ssDNA was amplified by
PCR using FAM- and biotin-labeled primers. Both positive and
negative selections were performed after 5 rounds of selection. The
ssDNA pools were initially incubated with 67NR cells, and the
supernatant containing unbound ssDNA was collected and incubated
with 4T1 cells.
Determination of binding efficiency
To determine the binding efficiency of the selected
ssDNA pools with target cells, FAM-labeled ssDNA pools were
respectively incubated with 1×106 of the 4T1 target
cells or subtractive 67NR cells in 500 µl binding buffer at
4°C for 30 min. Cells were washed 3 times at 1,000 rpm for 5 min,
and the fluorescence intensity was determined by flow
cytometry.
Cloning and sequencing of the selected
ssDNA pool
Based on the binding efficiency results, the 10th
round of enriched ssDNA pool was amplified using unmodified primers
and cloned into Escherichia coli using the TA cloning kit
(Invitrogen). The candidate aptamer sequences were determined by
Invitrogen Co., Ltd. (Shanghai, China). Nine aptamers were
selected, and the binding affinity was detected by amplifying
FAM-labeled ssDNA aptamers using FAM-conjugated primers. In brief,
the target cells (1×106) were incubated with varying
concentrations of FITC-labeled aptamers in 500 µl binding
buffer on ice for 30 min. Cells were washed twice after incubation,
and the fluorescence intensity was determined by flow cytometry.
Three aptamers with high binding affinity were sequenced, and the
structure of aptamer SRZ1 with best binding affinity was predicted
by RNA structure software.
Binding specificity
To study the binding specificity, FAM-conjugated
SRZ1 was synthesized by Sangon, Shanghai. To determine the cell
specificity of SRZ1, mouse mammary breast cancer 4T1, 4TO7 and 67NR
cells, normal mouse mammary gland NMuMG cells, mouse Lewis lung
carcinoma cell line LL/2, mouse hepatoma cell line Hepa 1–6 and
mouse B cell lymphoma cell line WEHI-231 were used in binding
assays by flow cytometry.
Preparation of NPs
The particles employed for the present study were
DOTAP:DOPE (molar ratio 50:50) based liposome preparations. DIR or
PKH26 dye was added into the liposome formulations at a ratio of 5
mol% if fluorescence indicators were needed. Liposomes were
prepared by thin film and hydration method as reported. Briefly, a
rotary evaporator was used to remove solvent from a glass tube
containing lipid mixed at the appropriate ratios in a water bath at
42°C until a thin film was deposited. The lipid film was hydrated
with ddH2O, ddH2O containing doxorubicin or
ddH2O containing both doxorubicin and aptamers in a
water bath at 37°C, and then placed on ice at 15-min intervals for
10 cycles.
Characterization of the NPs
Particle size and ζ-potential of the DOTAP:DOPE
liposomes, doxorubicin-loaded DOTAP:DOPE liposomes (DOTAP:DOPE/DOX)
or doxorubicin and aptamer-carried DOTAP:DOPE liposomes
(DOTAP:DOPE/DOX/aptamer) were determined by Zetasizer.
Uptake of particles
For the uptake study, doxorubicin-loaded DOTAP:DOPE
liposomes (DOTAP:DOPE/DOX) or doxorubicin and aptamer-carried
DOTAP:DOPE liposomes (DOTAP:DOPE/DOX/aptamer) were respectively
incubated with 4T1 cells at 37°C for 12 h, and then the cells were
washed 3 times, and the uptake efficiency was detected by flow
cytometry.
Cell viability assay
In vitro cytotoxicities of DOTAP:DOPE, free
doxorubicin, DOTAP:DOPE/DOX and DOTAP:DOPE/DOX/SRZ1 were analyzed
in the 4T1, 67NR and NMuMG cells using the CCK-8 method. Briefly,
the cells (100 µl) were cultured in a 96-well culture plate
at a density of 104/well and were subsequently incubated
for 24 h. Then the samples were added to each group (3-wells) for
24 h. The cells were incubated in 110 µl of DMEM containing
10 µl CCK-8 solution for 1 h after removing previous
nutrient solution. Absorbance of the suspension was measured at 450
nm on an ELISA reader.
In vitro DOX release
DOTAP:DOPE/DOX, DOTAP:DOPE/DOX/SRZ1 and free
doxorubicin (500 µg) were suspended in 2 ml of PBS buffer
with a pH of 7.4. The suspensions were placed at 37°C with constant
stirring at 100 rpm. At a desired time point, 1 ml of suspension
was removed and centrifuged. The supernatant was collected for
UV-vis measurement at a wavelength of 480 nm to determine the
release of doxorubicin.
Animals
Female BALB/c mice, 6–8 weeks of age, purchased from
the Laboratory Animal Center of Nanjing Medical University were
employed in this study. The present study complied with standards
for the Care and Use of Laboratory Animals and was approved by the
Animal Care and Use Committee of Nanjing Medical University. All of
the mice were monitored every day. All surgery was performed under
isoflurane anesthesia. All efforts were made to minimize suffering,
and all of the mice were finally sacrificed by CO2
asphyxiation.
In vivo image and biodistribution
To study the in vivo targets of
DOTAP:DOPE/DOX/aptamer in 4T1 tumor-bearing mice, DIR dye-labeled
DOTAP:DOPE/DOX/aptamer liposomes were prepared and intravenously
injected into the mice. Twenty-four hours following the injection,
mice were anesthetized and mouse bodies were scanned using Kodak
image system. The 4T1 tumors, livers, lungs, spleens, kidneys,
thymus, heart and gut were removed 48 h following the injection,
and the DIR dye signals in each organ were collected using the
Kodak image system.
Tumor model
Female BALB/c mice (4 weeks, 18–20 g, 10 mice/group)
were used. The 4T1 cells were harvested and resuspended in PBS to a
final density of 1×107 cells/ml. Before injection, cells
were resuspended in PBS and analyzed by 0.4% trypan blue exclusion
assay (viable cells, >90%). For cancer cell injection,
~5×105 4T1 cells in 100 µl of PBS were injected
into the mammary fat pad of each mouse. Treatment was initiated
when the primary tumors reached a mean diameter of 3–4 mm.
Tumor-bearing mice were divided into 5 groups and treated with PBS,
DOTAP:DOPE, free DOX, DOTAP:DOPE/DOX and DOTAP:DOPE/DOX/SRZ1 every
3 days for 10 times. Tumor volume was measured, and the survival
rate of the mice was monitored.
Results
Selection and binding affinity
identification of the aptamer pools
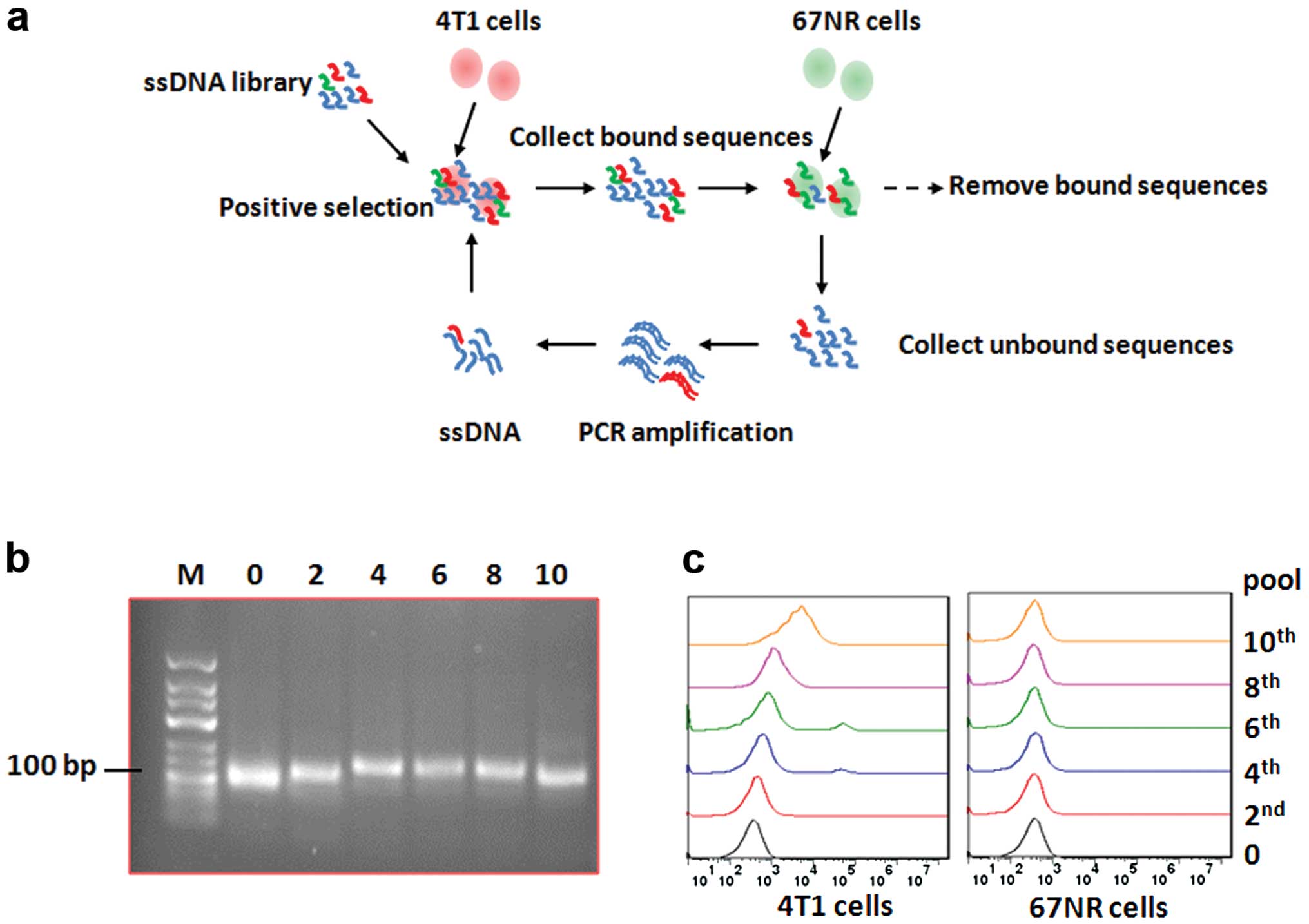
Cell-SELEX strategy was applied in the present study
to enrich specific anti-4T1 cell aptamers. As shown in the
schematic (Fig. 1a), 4T1 cells with
high invasive and metastatic potential were used as target cells,
while non-invasive and non-metastatic 67NR cells were used as
subtracted cells. After 10 rounds of selection, 10 ssDNA pools were
enriched, and the original ssDNA pool (0), 2nd, 4th, 6th, 8th and
10th pools were identified by running agarose gel electrophoresis
(Fig. 1b). Then the binding
affinity of the aptamer pools and 4T1 cells was analyzed by flow
cytometry. With increasing rounds of selection, the fluorescence
intensity bound on 4T1 cells was significantly increased (Fig. 1c, left), while there was no change
in the fluorescence intensity of the 67NR cells (Fig. 1c, right).
Selection, sequencing and structure
prediction of the 4T1-specific aptamers
After 10 rounds of enrichment, the selected ssDNA
library was PCR-amplified and cloned into Escherichia coli.
Clones were subjected to sequence analysis, and definite results
were obtained from 51 clones, including 12 clones yielding the same
sequence named SRZ1, 11 clones yielding the same sequence named
SRZ2, 8 clones yielding the same sequence named SRZ3, 6 clones
yielding the same sequence named SRZ4, 5 clones yielding the same
sequence named SRZ5, 4 clones yielding the same sequence named
SRZ6, 3 clones yielding the same sequence named SRZ7, another 2
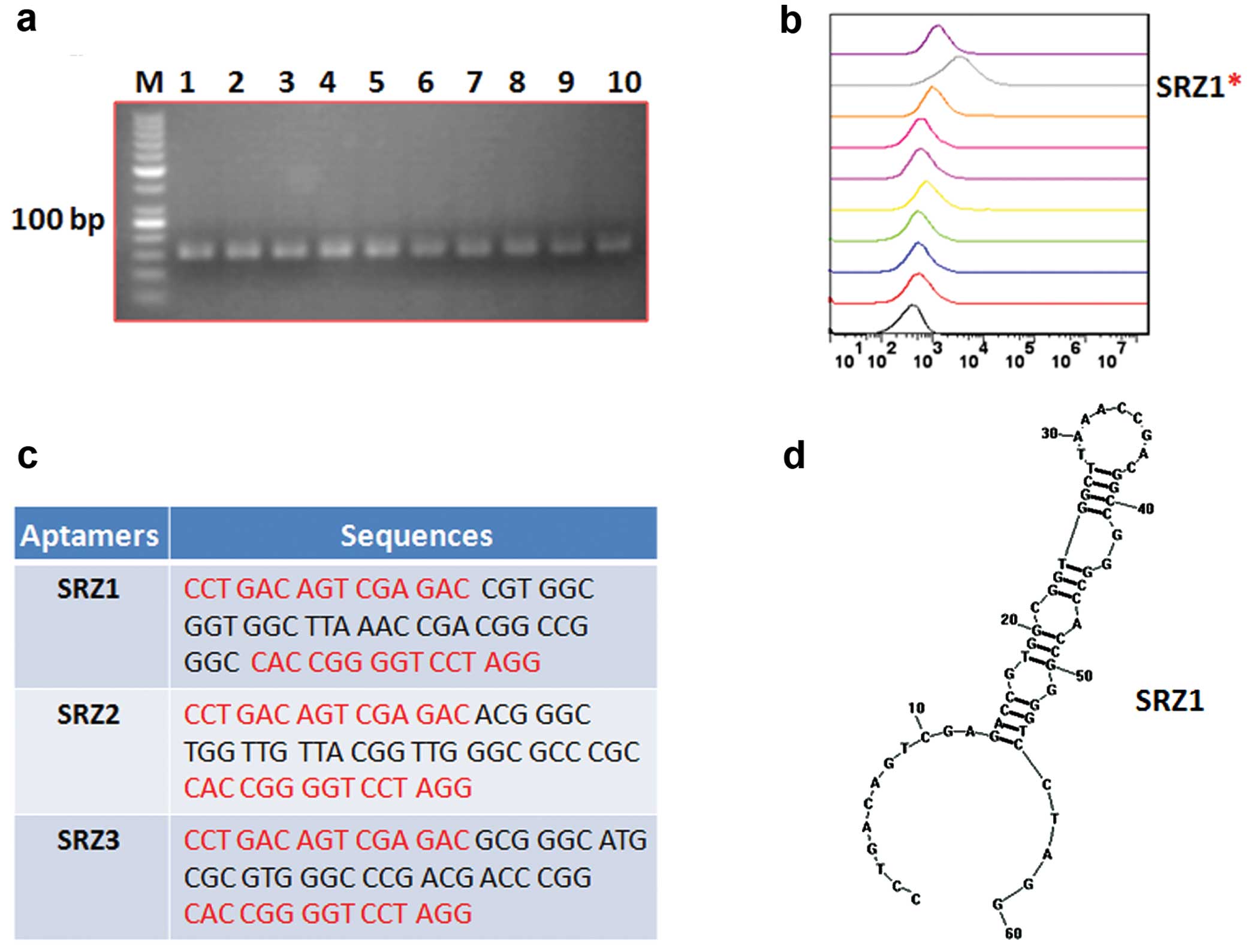
clones named SRZ8 and SRZ9. ssDNA was obtained by asymmetry PCR
from plasmid DNA and identified using agarose gel electrophoresis
(Fig. 2a). FAM-labeled SRZ1-9
aptamers were then synthesized, and the binding affinity was
evaluated. Data from the flow cytometry indicated that SRZ1 had the
best binding affinity to 4T1 cells (Fig. 2b). The sequences (Fig. 2c) were analyzed, and the structure
(Fig. 2d) was predicted by RNA
structure software. SRZ1 was selected for detailed study.
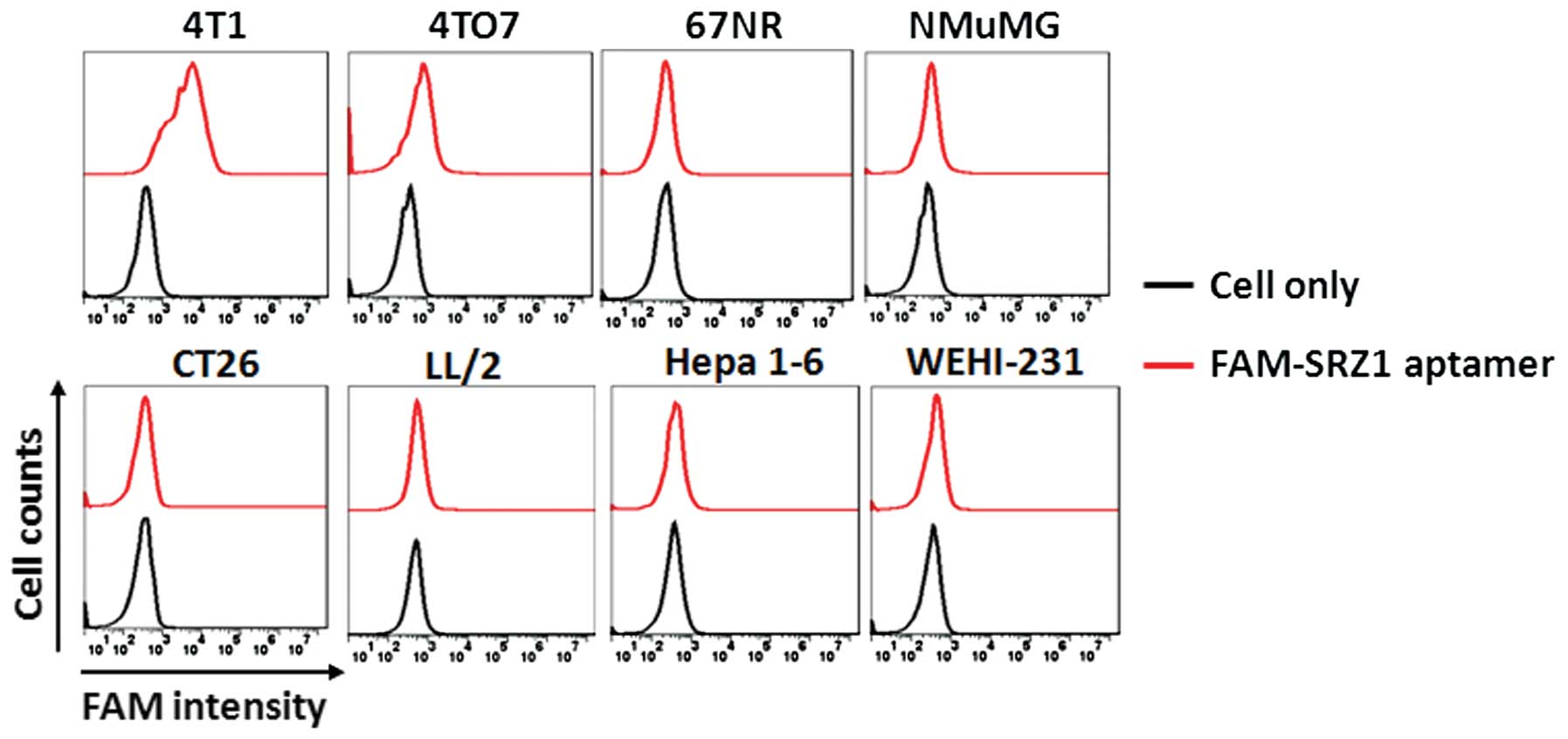
SRZ1 specifically binds to 4T1 cells
Mouse mammary breast cancer cell lines (4T1, 4TO7
and 67NR), the mouse normal mammary gland cell line NMuMG, the
human colon carcinoma cell line CT26, the mouse Lewis lung
carcinoma cell line LL/2, the mouse hepatoma cell line Hepa 1–6 and
the mouse B cell lymphoma cell line WEHI-231 were used to test the
binding specificity of aptamer SRZ1. As shown in Fig. 3, SRZ1 had highly specific binding
capacity to 4T1 cells and low specific binding capacity to 4TO7
cells, yet no or little binding to other cancer cells was
noted.
Characterization of DOTAP:DOPE liposomes
and liposome-based particles
We successfully selected the 4T1 cell-specific SRZ1
aptamer as indicated above. Thus, we developed a novel 4T1
tumor-targeted chemotherapeutic drug delivery system by conjugating
the SRZ1 aptamer on DOTAP:DOPE liposomes. Free DOTAP:DOPE,
doxorubicin-loaded DOTAP:DOPE (DOTAP:DOPE/DOX) and
doxorubicin-loaded SRZ1-conjugated DOTAP:DOPE (DOTAP:DOPE/DOX/SRZ1)
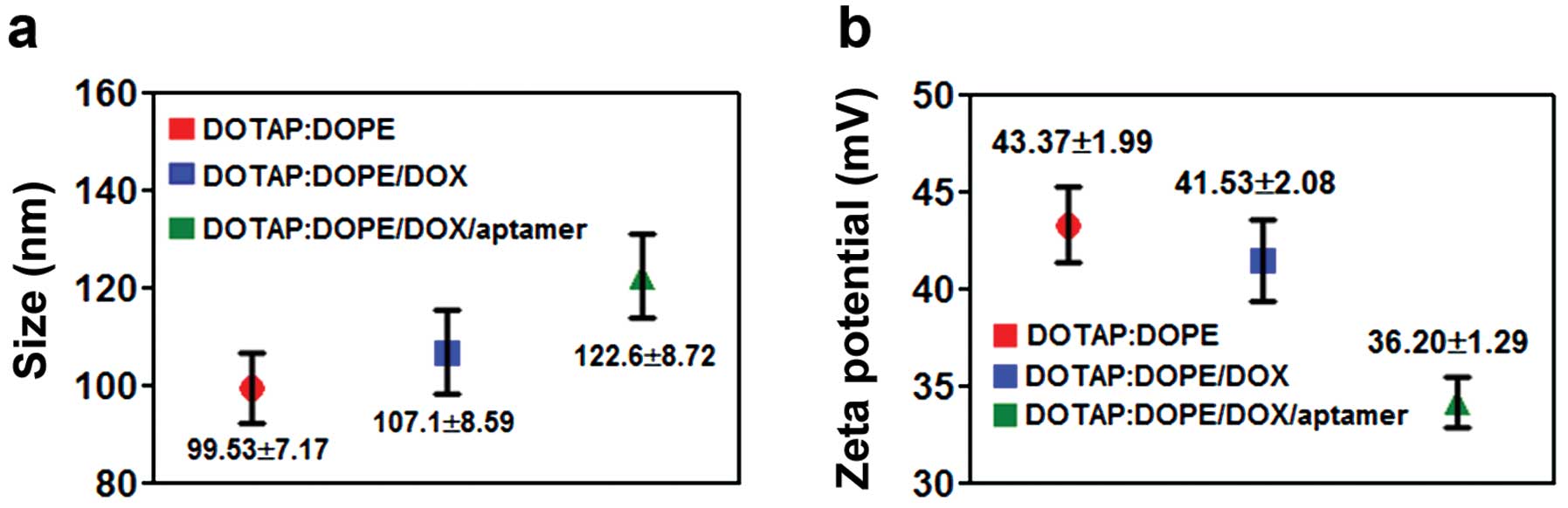
liposomes were prepared by thin film and hydration method. The size
distribution (Fig. 4a) and surface
ζ-potential (Fig. 4b) were measured
using Zetasizer. The size of all of the particles was ~100 nm, and
the charge of the particles was positive.
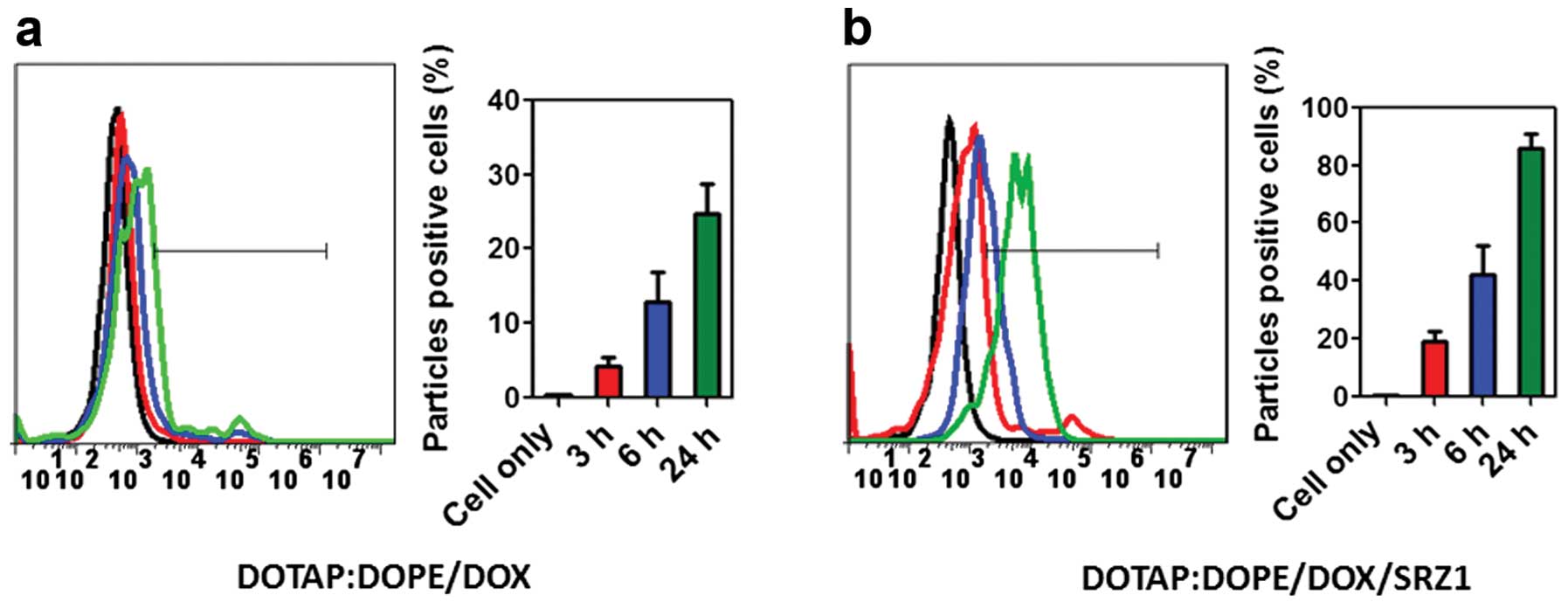
Aptamer SRZ1 promotes uptake of
DOTAP:DOPE/DOX by 4T1 cells
We next tested the uptake efficiency of
DOTAP:DOPE/DOX and aptamer SRZ1-conjugated DOTAP:DOPE/DOX
(DOTAP:DOPE/DOX/SRZ1) by 4T1 cells in vitro. DOTAP:DOPE/DOX
or DOTAP:DOPE/DOX/SRZ1 was respectively incubated with 4T1 cells
for 3, 6 and 24 h, and then the cells were washed and collected by
0.25% Trypsin-EDTA digestion. The doxorubicin signaling in the 4T1
cells was detected by flow cytometry. As shown in Fig. 5a, uptake of DOTAP:DOPE/DOX was
markedly increased by carrying 4T1 cell-specific aptamer SRZ1
compared with DOTAP:DOPE/DOX only (Fig.
5b).
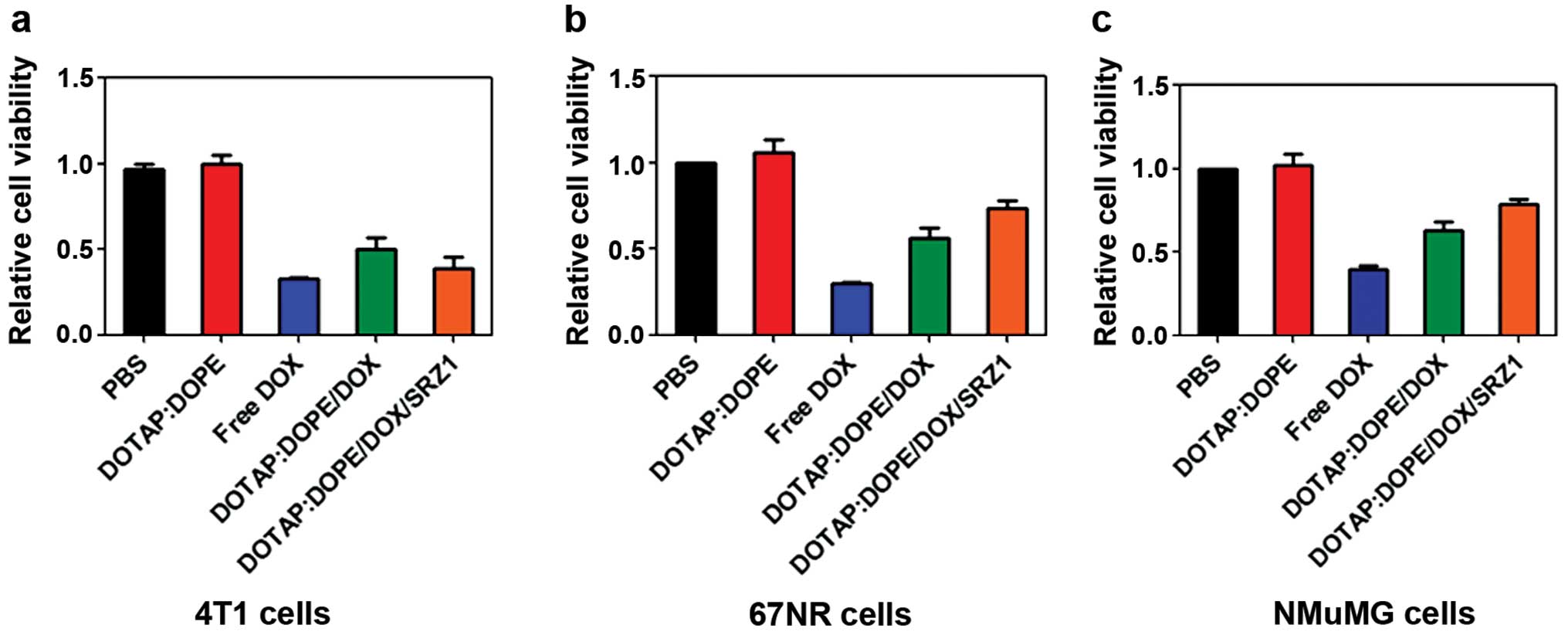
Cell toxicity of DOTAP:DOPE/DOX/SRZ1
Cell viability of the 4T1 cells treated with
DOTAP:DOPE, free doxorubicin, DOTAP:DOPE/DOX or DOTAP:DOPE/DOX/SRZ1
was analyzed (Fig. 6a). Data from
Fig. 3a showed that only the
DOTAP:DOPE liposome had no effect on 4T1 cell viability.
DOTAP:DOPE/DOX/SRZ1 significantly inhibited cell viability. Further
study demonstrated that free doxorubicin markedly suppressed the
viability of both 67NR (Fig. 6b)
and normal mouse mammary gland NMuMG cells (Fig. 6c), while DOTAP:DOPE/DOX/SRZ1 had
less toxicity to 67NR and NMuMG cells. Data from this panel
indicated that doxorubicin was targeted delivered to target cells
by the conjugated specific aptamers.
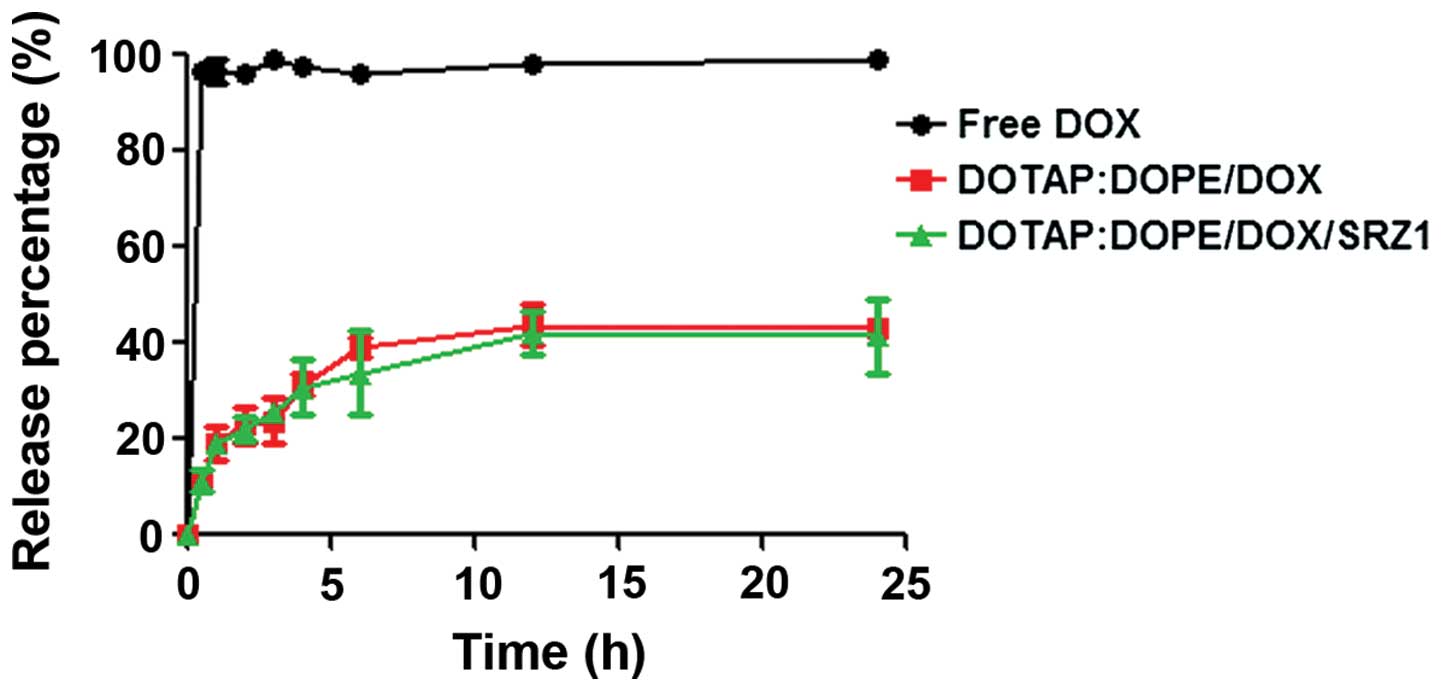
Release profile of doxorubicin
The release profile of DOX from DOTAP:DOPE/DOX and
DOTAP:DOPE/DOX/SRZ1 was investigated in PBS at pH 7.4 (Fig. 7). Both DOTAP:DOPE/DOX and
DOTAP:DOPE/DOX/SRZ1 resulted in sustained drug release, and the
release of DOX from DOTAP:DOPE/DOX was the same as that from
DOTAP:DOPE/DOX/SRZ1.
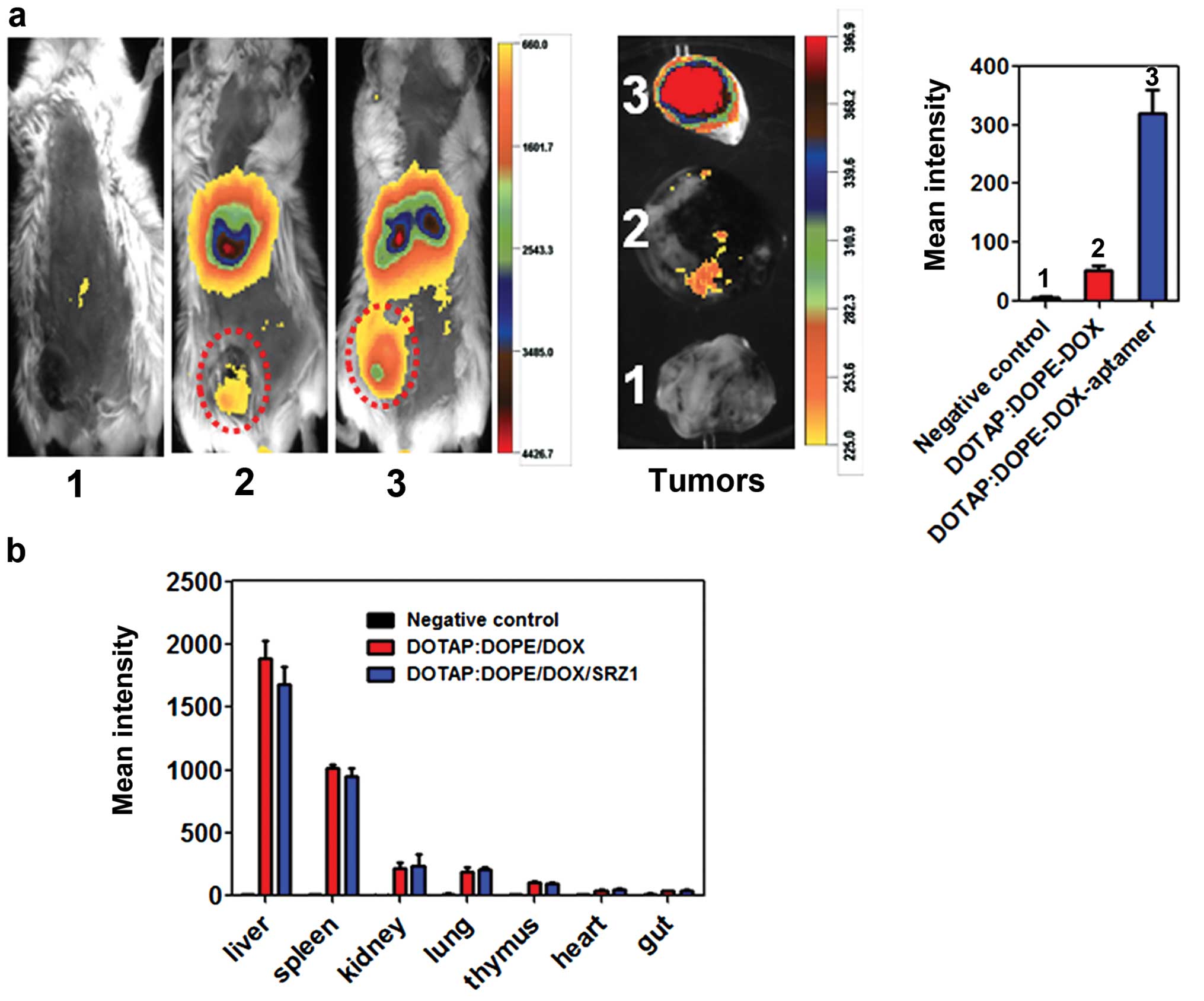
Targeted delivery and biodistribution in
the 4T1 tumor-bearing mice
To study the targeted deliver of DOTAP:DOPE/DOX by
aptamer SRZ1 in vivo, 4T1 tumor-bearing mice were
intravenously administered DIR dye-labeled DOTAP:DOPE/DOX or
DOTAP:DOPE/DOX/SRZ1 and body images of the mice were collected 24 h
following injection. As shown in Fig.
8a, more DOTAP:DOPE/DOX signals were detected in the 4T1 tumors
after conjugating with specific anti-4T1 cell SZR1 aptamer.
Seventy-two hours after injection, the mice were sacrificed, and
tumors, livers, lungs, spleens, kidneys, thymus and gut were
removed and DIR dye signals in each tissue were scanned. Consistent
with the body images, aptamer SZR1 significantly increased
accumulation of DOTAP:DOPE/DOX in the 4T1 tumor tissues (Fig. 8a, right). Organ scan results
indicated that most were DOTAP:DOPE/DOX and DOTAP:DOPE/DOX/SRZ1
distributed in the mouse liver, spleen, kidney and lung (Fig. 8b).
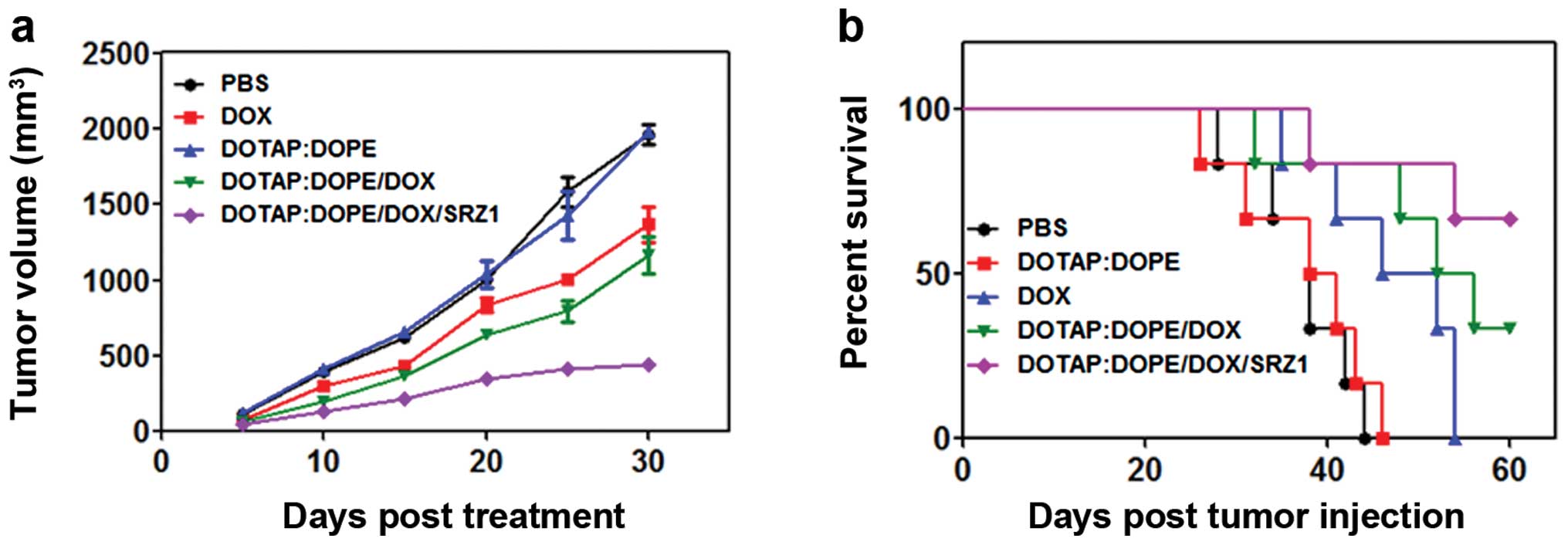
Tumor suppression is effectively enhanced
by conjugating SRZ1 aptamer with DOTAP:DOPE/DOX
To test the effects of DOTAP:DOPE/DOX on tumor
growth after conjugating with the SRZ1 aptamer, 4T1 tumor-bearing
mice were intravenously injected with PBS, free DOTAP:DOPE
particles, free DOX, DOTAP:DOPE/DOX or DOTAP:DOPE/DOX/SRZ1 every 3
days. The tumor size was measured every 5 days, and the volume data
revealed that DOX only and DOTAP:DOPE/DOX inhibited 4T1 tumor
growth, and the suppressive effect of DOTAP:DOPE/DOX was markedly
enhanced by the conjugated 4T1-specific aptamer SRZ1 (Fig. 9a). From the survival rate results of
the five groups, we determined that the survival time was prolonged
after treatment with free DOX and the DOX-loaded DOTAP:DOPE
particles, while optimal results were obtained from the aptamer
SRZ1-conjugated group (Fig. 9b),
which was consistent with the tumor inhibition results.
Discussion
Aptamer-mediated targeted delivery is a promising
strategy for improving the therapeutic index of cytotoxic drugs
that cause serious systemic toxicities (17,18),
such as doxorubicin. In the present study, we identified a novel
single-strand DNA aptamer to mouse breast cancer 4T1 cells by using
cell-SELEX. Our objective was to establish a more effective
chemotherapeutic drug target delivery platform. To our knowledge,
SRZ1 is the first aptamer to be identified as specific for mouse
mammary breast cancer 4T1 cells. Both in vitro and in
vivo data presented in the present study revealed that SRZ1 was
a powerful aptamer for targeted doxorubicin delivery in a mouse
breast cancer model therefore providing a novel strategy for human
breast cancer treatment.
Doxorubicin is used to treat a variety of tumors
including breast cancer. It is considered as the most useful
anticancer drug worldwide (19,20). A
large number of studies have reported that free doxorubicin can be
taken up by both tumor cells and normal cells via a passive
diffusion mechanism, and the proliferation of both tumor cells and
normal cells was suppressed by doxorubicin through interaction with
nuclear DNA (21,22). Thus, the acute toxicity of free
doxorubicin to normal tissues limits the therapeutic efficacy in
clinical use.
In recent years, liposomes have been extensively
employed as effective delivery systems to enhance the efficacy of
encapsulated chemotherapeutic drugs (23–25).
Liposomes provide a better accumulation of drugs in tumor tissues
through an enhanced permeability and retention (EPR) effect.
Liposomes composed of DOTAP and DOPE have been demonstrated as
great potential carriers for the delivery of anticancer agents
(26,27). To achieve an optimum therapeutic
effect of doxorubicin in a 4T1 breast cancer model, we generated an
efficient delivery system for doxorubicin by conjugating 4T1 tumor
cell-specific aptamers with doxorubicin-loaded DOTAP:DOPE
liposomes. Binding specificity was assessed in vitro by FACS
and proved that the aptamer SRZ1 was able to bind to mouse 4T1
cells, rather than mouse normal mammary gland cell line NMuMG,
human colon carcinoma cell line CT26, mouse Lewis lung carcinoma
cell line LL/2, mouse hepatoma cell line Hepa 1–6 and mouse B cell
lymphoma cell line WEHI-231. Meanwhile, we demonstrated that the
SRZ1 aptamer modified doxorubicin-loaded DOTAP:DOPE had a higher
internalization in 4T1 cells than in 67NR cells. This targeting was
achieved through the 4T1 cell-specific binding affinity of aptamer
SRZ1. Furthermore, we confirmed the specific toxicity of
DOTAP:DOPE/DOX/SRZ1 to 4T1 cells, when compared to the negative
screening NR67 and normal mouse mammary gland NMuMG cells, thereby
limiting the toxicity of doxorubicin to target cells. Additionally,
reduced release of doxorubicin in the DOTAP:DOPE/DOX and
DOTAP:DOPE/DOX/SRZ1 groups was detected by UV spectrometer.
Doxorubicin in free form can be uptaken by cells within 15 min
through a passive diffusion mechanism. However, after co-loading
doxorubicin and the 4T1-specific SRZ1 aptamer onto DOTAP:DOPE
liposomes, internalization of doxorubicin in the target cells was
mainly dependent on the interaction between the aptamer and the
cell membrane. Thus, less toxicity to 67NR or NMuMG cells was
observed in the DOTAP:DOPE/DOX/SRZ1 group than that in the free
doxorubicin group. However, due to the target recognition by the
SRZ1 aptamer, the local concentration of doxorubicin was markedly
increased, compared to the free doxorubicin and DOTAP:DOPE/DOX
group. Hence, maximum tumor suppression was achieved in the
DOTAP:DOPE/DOX/SRZ1-treated 4T1 tumor-bearing mouse.
Collectively, to maximize the tumor inhibitory
function of doxorubicin, we developed a novel delivery system by
conjugating a target tumor cell-specific aptamer with
doxorubicin-loaded DOTAP:DOPE liposomes. The binding specificity
and efficacy of this drug-delivery platform were further
investigated in vitro and in vivo. Nonspecific
cellular toxicity to normal cells was significantly reduced after
conjugating the target cell-specific aptamer with DOTAP:DOPE/DOX.
An aptamer is a type of small molecule and can be designed as a
targeting ligand, thus enabling the selective delivery of
therapeutic drugs to target cells providing a signifi-cant
potential for future clinical cancer therapy.
Acknowledgments
This study was supported in part by the National
Natural Science Foundation of China (nos. 81270952, 81070684,
81071753, 81172502, 81202077 and 81272916), the Natural Science
Foundation of Jiangsu Province (BK2010581, BK2011853 and
BK2011855), the Program for Development of Innovative Research Team
in the First Affiliated Hospital of NJMU (IRT-008), and a project
funded by the Priority Academic Program Development of Jiangsu
Higher Education Institutions (PAPD).
References
|
1
|
Wang B, He M and Wang L, Engelgau MM, Zhao
W and Wang L: Breast cancer screening among adult women in China,
2010. Prev Chronic Dis. 10:E1832013. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
MacDonald V: Chemotherapy: managing side
effects and safe handling. Can Vet J. 50:665–668. 2009.PubMed/NCBI
|
|
3
|
Lin X, Gao R, Zhang Y, Qi N, Zhang Y,
Zhang K, He H and Tang X: Lipid nanoparticles for chemotherapeutic
applications: strategies to improve anticancer efficacy. Expert
Opin Drug Deliv. 9:767–781. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
De Jong WH and Borm PJ: Drug delivery and
nanoparticles: applications and hazards. Int J Nanomed. 3:133–149.
2008. View
Article : Google Scholar
|
|
5
|
Shohdy KS and Alfaar AS: Nanoparticles
targeting mechanisms in cancer therapy: current limitations and
emerging solutions. Ther Deliv. 4:1197–1209. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Bastakoti BP, Torad NL and Yamauchi Y:
Polymeric micelle assembly for the direct synthesis of
platinum-decorated mesoporous TiO2 toward highly
selective sensing of acetaldehyde. ACS Appl Mater Interfaces.
6:854–860. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Quan L, Zhang Y, Crielaard BJ, Dusad A,
Lele SM, Rijcken CJ, Metselaar JM, Kostková H, Etrych T, Ulbrich K,
et al: Nano-medicines for inflammatory arthritis: head-to-head
comparison of glucocorticoid-containing polymers, micelles, and
liposomes. ACS Nano. 8:458–466. 2014. View Article : Google Scholar :
|
|
8
|
Chang DK, Li PC, Lu RM, Jane WN and Wu HC:
Peptide-mediated liposomal doxorubicin enhances drug delivery
efficiency and therapeutic efficacy in animal models. PLoS One.
8:e832392013. View Article : Google Scholar
|
|
9
|
Wang Q, Zhuang X, Mu J, Deng ZB, Jiang H,
Zhang L, Xiang X, Wang B, Yan J, Miller D, et al: Delivery of
therapeutic agents by nanoparticles made of grapefruit-derived
lipids. Nat Commun. 4:18672013. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Park K: Lessons learned from
thermosensitive liposomes for improved chemotherapy. J Control
Release. 174:2192014. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Sabnis N and Lacko AG: Drug delivery via
lipoprotein-based carriers: answering the challenges in systemic
therapeutics. Ther Deliv. 3:599–608. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Lacko AG, Nair M, Prokai L and McConathy
WJ: Prospects and challenges of the development of
lipoprotein-based formulations for anti-cancer drugs. Expert Opin
Drug Deliv. 4:665–675. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Lassalle HP, Marchal S, Guillemin F,
Reinhard A and Bezdetnaya L: Aptamers as remarkable diagnostic and
therapeutic agents in cancer treatment. Curr Drug Metab.
13:1130–1144. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Bunka DH, Platonova O and Stockley PG:
Development of aptamer therapeutics. Curr Opin Pharmacol.
10:557–562. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Shi H, Cui W, He X, Guo Q, Wang K, Ye X
and Tang J: Whole cell-SELEX aptamers for highly specific
fluorescence molecular imaging of carcinomas in vivo. PLoS One.
8:e704762013. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Meyer S, Maufort JP, Nie J, Stewart R,
McIntosh BE, Conti LR, Ahmad KM, Soh HT and Thomson JA: Development
of an efficient targeted cell-SELEX procedure for DNA aptamer
reagents. PLoS One. 8:e717982013. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Dassie JP and Giangrande PH: Current
progress on aptamer-targeted oligonucleotide therapeutics. Ther
Deliv. 4:1527–1546. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Subramanian N, Raghunathan V, Kanwar JR,
Kanwar RK, Elchuri SV, Khetan V and Krishnakumar S: Target-specific
delivery of doxorubicin to retinoblastoma using epithelial cell
adhesion molecule aptamer. Mol Vis. 18:2783–2795. 2012.PubMed/NCBI
|
|
19
|
Tacar O, Sriamornsak P and Dass CR:
Doxorubicin: an update on anticancer molecular action, toxicity and
novel drug delivery systems. J Pharm Pharmacol. 65:157–170. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Carvalho C, Santos RX, Cardoso S, Correia
S, Oliveira PJ, Santos MS and Moreira PI: Doxorubicin: the good,
the bad and the ugly effect. Curr Med Chem. 16:3267–3285. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Speelmans G, Staffhorst RW, De Kruijff B
and De Wolf FA: Transport studies of doxorubicin in model membranes
indicate a difference in passive diffusion across and binding at
the outer and inner leaflets of the plasma membrane. Biochemistry.
33:13761–13768. 1994. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Kamba SA, Ismail M, Hussein-Al-Ali SH,
Ibrahim TA and Zakaria ZA: In vitro delivery and controlled release
of doxorubicin for targeting osteosarcoma bone cancer. Molecules.
18:10580–10598. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Shaikh IM, Tan KB, Chaudhury A, Liu Y, Tan
BJ, Tan BM and Chiu GN: Liposome co-encapsulation of synergistic
combination of irinotecan and doxorubicin for the treatment of
intraperitoneally grown ovarian tumor xenograft. J Control Release.
172:852–861. 2013. View Article : Google Scholar
|
|
24
|
Zhaorigetu S, Rodriguez-Aguayo C, Sood AK,
Lopez-Berestein G and Walton BL: Delivery of negatively charged
liposomes into the atherosclerotic plaque of apolipoprotein
E-deficient mouse aortic tissue. J Liposome Res. 24:182–190. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Alavi SE, Esfahani MK, Alavi F, Movahedi F
and Akbarzadeh A: Drug delivery of hydroxyurea to breast cancer
using liposomes. Indian J Clin Biochem. 28:299–302. 2013.
View Article : Google Scholar :
|
|
26
|
Remaut K, Lucas B, Braeckmans K, Sanders
NN, Demeester J and De Smedt SC: Delivery of phosphodiester
oligonucleotides: can DOTAP/DOPE liposomes do the trick?
Biochemistry. 45:1755–1764. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Ciani L, Casini A, Gabbiani C, Ristori S,
Messori L and Martini G: DOTAP/DOPE and DC- Chol/DOPE lipoplexes
for gene delivery studied by circular dichroism and other
biophysical techniques. Biophys Chem. 127:213–220. 2007. View Article : Google Scholar : PubMed/NCBI
|