Introduction
Human neural glioma, the most lethal brain tumor and
the most frequent type of primary adult brain neoplasms, has dismal
outcome for patient (1–3). Patients diagnosed with glioma have a
very low 5-year survival, compared with patients affected by
glioblastoma multiforme (GBM) (4,5).
Despite significant advances in neurosurgery and
chemo-radiotherapy, glioma remains highly resistant to conventional
treatments and improvements in patient outcome have been modest
(6). For this reason, it is vital
and urgent to investigate thoroughly the molecular pathological
mechanism, for some more specific biomarkers and potential remedy
targets.
Protein phosphatase 1 (PP1) is a member of the
well-known mammalian protein phosphatases, serine/threonine
phosphatases, which catalyzes the majority of protein
dephosphorylation events that regulate diverse cellular processes,
such as neuronal signaling, muscle contraction, glycogen synthesis,
and cell proliferation. Mammals have three PP1 catalytic genes,
PP1α, γ and δ, which are encoded by separate genes. These isoforms
are >89% identical in amino acid sequence, with minor
differences primarily at their NH2 and COOH termini (7). The equivalent T residue is conserved
in all three PP1 isoforms (T316 in PP1β and T311 in PP1γ), and PP1γ
can be inactivated by Cdk-dependent T311 phosphorylation. PP1α
and/or -β could compensate for the depletion of PP1γ in
development, but not in the specific function of spermiogenesis.
PP1γ has two isoforms, γ1 and γ2, generated by differential
splicing of PP1γ. PP1 isoforms are expressed in all tissues and are
widely distributed, except for PP1γ2, which is found only in the
testes. Mice with depleted PP1γ are viable, but males show
defective spermiogenesis and are infertile (8). Thus, in this study, we focused on
PP1γ1.
The transcription factor, nuclear factor-kappa B
(NF-κB), plays an important role in tumor cells. Constitutive
activation of NF-κB is responsible for proliferation, because
inhibition of NF-κB leads to abrogation of proliferation (9). In normal cells, most of NF-κB
complexes are kept predominantly cytoplasmic, and stay in an
inactive form, by binding to a family of inhibitor proteins, the
IκBs. Generally, the inactive NF-κB-IκBα complex is activated by
phosphorylation on two conserved serine residues within the
N-terminal domain of the IκB proteins (10). PP1γ enhances TRAF6 E3 ubiquitin
ligase activity (11) and the
induction of NF-κB activation (12). Subsequently, TRAF6 catalyzes the
ubiquitin of IKK/NEMO (NF-κB essential modulator) (11). IKK proteins coordinate the
phosphorylation, ubiquitination, and degradation of inhibitory IκBα
proteins, liberating NF-κB heterodimers to translocate into the
nucleus. Activation of the IκB kinase (IKK) complex results in the
phosphorylation and subsequent proteasomal degradation of IκBα
(13). Activation of the NF-κB
signaling cascade results in complete degradation of IκB, allowing
translocation of NF-κB to the nucleus, where it induces
transcription (14). Activated
NF-κB binds to specific DNA sequences in target genes involved in
tumor cell proliferation, invasion, metastasis, angiogenesis,
chemoresistance, radioresistance, inflammation and immunoregulation
(10).
According to previous studies, PP1γ might promote
tumor progression in colorectal cancer (15). Here, we identified the paralleled
positive correlation between the expression of PP1γ and Ki-67 in
human glioma tissue and cell lines, and the depletion of PP1γ
reduced cell proliferation distinctly. Besides, we demonstrated
PP1γ might accelerate the glioma cell proliferation via NF-κB
pathway by enhancing the transportation of p65 into the nucleus.
Based on the results, it might be a promising target for human
glioma diagnosis and therapy in future.
Materials and methods
Patients and tissue specimens
The human glioma tissues were collected from 100
glioma surgical specimens without any therapy, and the
clinicopathological data were provided by the Department of
Pathology, Affiliated Hospital of Nantong University. Normal brain
specimens obtained from 10 patients who received epilepsy surgery
were verified for the absence of any tumor. Specimens were
immediately frozen in liquid nitrogen and stored at −80°C until
use. Some parts of the specimens (including WHO grade II, III and
IV) were fixed in 10% formalin and embedded in paraffin for
immunohistochemical analysis. All the tissues were collected and
applied in accordance with The Code of Ethics of the World Medical
Association. Ethics approval was given by the medical ethics
committee of the Affiliated Hospital of Nantong University.
Cell lines and cell culture
The human glioblastoma cell lines H4, SHG44, U87MG,
U251, A172 and U373 were purchased from Shanghai Institute of Cell
Biology. All cells were cultured in the DMEM high-glucose medium
(Gibco BRL, Grand Island, NY, USA) with 10% heat-inactivated fetal
bovine serum at 37°C with 5% CO2.
Antibodies
The antibodies applied to immunohistochemistry
included: anti-PP1γ (diluted 1:400, Santa Cruz Biotechnology),
anti-Ki-67 (diluted 1:400, Santa Cruz Biotechnology). The
antibodies applied to western blot analysis included: anti-PP1γ
(diluted 1:2,000, Santa Cruz Biotechnology), anti-p65 (diluted
1:2,000, Santa Cruz Biotechnology), anti-p-p65 (S536-p, diluted
1:2,000, Cell Signaling Technology), anti-IκBα (diluted 1:2,000,
Santa Cruz Biotechnology), anti-proliferating cell nuclear antigen
(PCNA, diluted 1:1,000, Santa Cruz Biotechnology), anti-cyclin D1
(diluted 1:1,000, Santa Cruz Biotechnology), anti-α-tubulin
(diluted 1:1,000, Santa Cruz Biotechnology), anti-Lamin B (diluted
1:1,000, Santa Cruz Biotechnology), anti-glyceraldehyde-3-phosphate
dehydrogenase (GAPDH, diluted 1:1,500, Santa Cruz
Biotechnology).
Immunohistochemistry (IHC)
The glioma tissues sections were deparaffinized in
xylene, rehydrated in graded ethanol solutions, and then we block
the endogenous peroxidase activity in 0.3% hydrogen peroxide. Then
the sections were heated at 105°C in 0.1 M citrate buffer, pH 6.0
for 10 min and incubated at room temperature for 1 h to retrieve
the antigen. Afterwards, the section were incubated with 5% bovine
serum in PBS (phosphate-buffered saline) (pH 7.2) for blocking
nonspecific protein binding, followed by overnight incubation with
appropriate antibodies at 4°C. Negative-control groups were treated
with non-specific immunoglobulin IgG (Sigma Chemical Co., St.
Louis, MO, USA) as the first antibody. Then the sections were
incubated. According to the manufacturer's instructions, the slides
were rinsed in PBS and incubated with hematoxylin, dehydrated and
mounted successively in resin mount (16).
Immunohistochemical evaluation
All of the stained sections were evaluated in a
blinded manner by two independent senior pathologists without any
clinicopathological variables of the patients (17). Five high-power fields (Leica
microscope Germany) were selected randomly and >300 cells in
each field were counted to determine the labeling index (LI), which
means the percentage of immunostained cells relative to the total
number of cells (18). Intensity
was evaluated in comparison with the control and scored as follows:
0, negative staining; 1, weak staining; 2, moderate staining; 3,
strong staining. The percentage of tumor cells stained positive was
scored as follows: 0, <1% positive tumor cells; 1, 1–10%
positive tumor cells; 2, 10–50% positive tumor cells; 3, 50–75%
positive tumor cells; 4, >75% positive tumor cells. Then we
added the scores of intensity and percentage as 0 and 2–7. 0,
negative; 2–3, weak stained; 4–5, moderate stained; 6–7, strong
stained. For statistical analysis, 0–3 were counted as low
expression, while 4–7 were counted as overexpression (19).
Cellular fractionation
Cells were washed twice with phosphate-buffered
saline (PBS), and resuspended in buffer A (50 mM NaCl, 10 mM HEPES,
pH 8.0, 500 mM sucrose, 1 mM EDTA, 0.2% Triton X-100, 1 mM NaF, 0.5
mM Na3VO4, 1 mM PMSF, and 2 µg/ml
aprotinin, 0.5 mM 2-mercaptoethanol) for 15 min on ice. Cells were
homogenized with 20 strokes using a Dounce homogenizer. After brief
centrifugation, the supernantant was collected as a cytoplasmic
fraction and the pelleted nuclei was further washed three times
with isotonic sucrose buffer (250 mM sucrose, 6 mM
MgCl2, 10 mM Tris-HCl, pH 7.4) containing 0.5% non-ionic
detergent Triton X-100 to dissolve any cytoplasmic membrane
contaminants (20). To extract
nuclear proteins, the isolated nuclei were resuspended in buffer C
(350 mM NaCl, 10 mM HEPES, pH 8.0, 25% glycerol, 0.1 mM EDTA, 1 mM
PMSF, and 2 µg/ml aprotinin, 0.5 mM 2-mercaptoethanol) with
gentle rocking for 30 min at 4°C. After centrifugation, the
supernatant was collected as a nuclear fraction (21).
Western blot assay
Glioma tissue samples were homogenized in lysis
buffer (1% Nonidet P-40, 50 mmol/l Tris, pH 7.5, 5 mmol/l EDTA, 1%
SDS, 1% sodium deoxycholate, 1% Triton X-100, 1 mmol/l PMSF, 10
mg/ml aprotinin, and 1 mg/ml leupeptin). Cell samples were washed
three times with PBS, suspended in 2X lysis buffer (50 mM Tris-HCl,
120 mM NaCl, 0.5% Nonidet P-40, 100 mM NaF, 200 M
Na3VO4, protease inhibitor mixture). Then,
samples were denatured at 100°C for 15 min and evaluated with
Bio-Rad protein assay (Bio-Rad, Hercules, CA, USA) for the total
protein concentration, and then stored at −20°C. The protein
samples were separated via SDS-polyacrylamide gel electrophoresis
(SDS-PAGE) and transferred to polyvinylidene difluoride filter
(PVDF) membranes (Millipore, Bedford, MA, USA). The membranes were
blocked in TBST (20 mM Tris, 150 mM NaCl, 0.05% Tween-20) with 5%
evaporated milk for 2 h at room temperature and then incubated in
appropriate antibodies at 4°C for 6–8 h. The membranes were washed
with TBST for three times, 5 min/each time and incubated with
horseradish peroxidase-linked IgG for 2 h at room temperature, and
then detected by infrared imaging system (Odyssey, USA). The blot
band intensity was quantified by ImageJ analysis file (Wayne
Rasband, National Institutes of Health, USA) (22).
RNA interference of PP1γ
The PP1γ-shRNAs were synthesized by GeneChem.
PP1γ-shRNAs target sequences: shRNA#1, 5′-AATGCCACGAGACCTGTAA-3′;
shRNA#2, 5′-GAATT ATGCGACCAACTGA-3′; shRNA#3, 5′-GACCGATTATGC
TTTCTTT-3′; shRNA#4, 5′-TGCTGTCATGGAGGTTTAT-3′. U251 cells were
transfected with 100 nmol/l of shRNA#2 performed with Lipofectamine
2000 (Invitrogen, Carlsbad, CA, USA) and U87MG cells with shRNA#4.
The cells were identified as negative-control groups treated with
control shRNA, and the mock non-treated groups.
Flow cytometry analysis of cell
cycle
Following the indicated pre-treatment, cells were
fixed with 70% methanol in PBS at −20°C for 48 h, and then with 1
mg/ml RNase A for 20 min at 37°C. Afterwards, the cells were
stained with 0.5 mg/ml of propidium iodide (PI). The DNA contents
were analyzed by a Becton-Dickinson flow cytometer, BD FACScan (San
Jose, CA, USA).
Cell Counting Kit-8 (CCK)-8
The cells transfected by shRNAs, with the untreated
groups were seeded on a 96-well cell culture cluster (Corning Inc.,
Corning, NY, USA) at 2×104 cells/well in 100 µl
medium and incubated overnight. Cell Counting Kit-8 (Dojindo,
Kumamoto, Japan) reagents were added to a subset of wells with
different treatments, after which absorbance was measured at a test
wavelength of 450 nm on an automated plate reader with the 630 nm
wavelength as a reference group.
Colony formation assay
The selected and stably-transfected cells were
plated in 6-cm culture plates at 100 cells/well. After incubation
for 12 days at 37°C, the cells were washed twice with PBS and
stained with Giemsa solution. The number of colonies containing
>50 cells was counted under a microscope.
NF-κB activity assay
Nuclear extracts from U251 and U87MG cell lines,
treated with conditioned media were prepared using the nuclear
extract kit (Active Motif). Total protein amounts were measured by
BCA-assay (Pierce, Rockford, IL, USA), NF-κB activity was measured
using the TransAM NF-κB p65 assay kit according to the
manufacturer's instructions. Briefly, 5 µg of nuclear
extract per sample was added to the 96-well plate, in which
oligonucleotide containing the NF-κB consensus site
(5′-GGGACTTTCC-3′) was immobilized. The active form of NF-κB bound
to the oligonucleotide was detected using an antibody against NF-κB
p65 subunit. An HRP-conjugated secondary antibody provided a
sensitive colorimetric readout that was quantified by
spectrophotometry.
Statistical analysis
Statistical analysis was performed by the SPSS17.0
statistical analysis software. The statistical correlations between
PP1γ and Ki-67 expression and the clinicopathological features were
analyzed using the χ2 test. Survival analysis was
carried out using the Kaplan-Meier method, and curves were compared
using the log-rank test. All of values were expressed as mean± SEM
(standard error of the mean), and P<0.05 was considered
statistically significant.
Results
The correlation of the expression of
PP1γ, p-p65, p65 and Ki-67 in human glioma tissues with the WHO
grades of human glioma
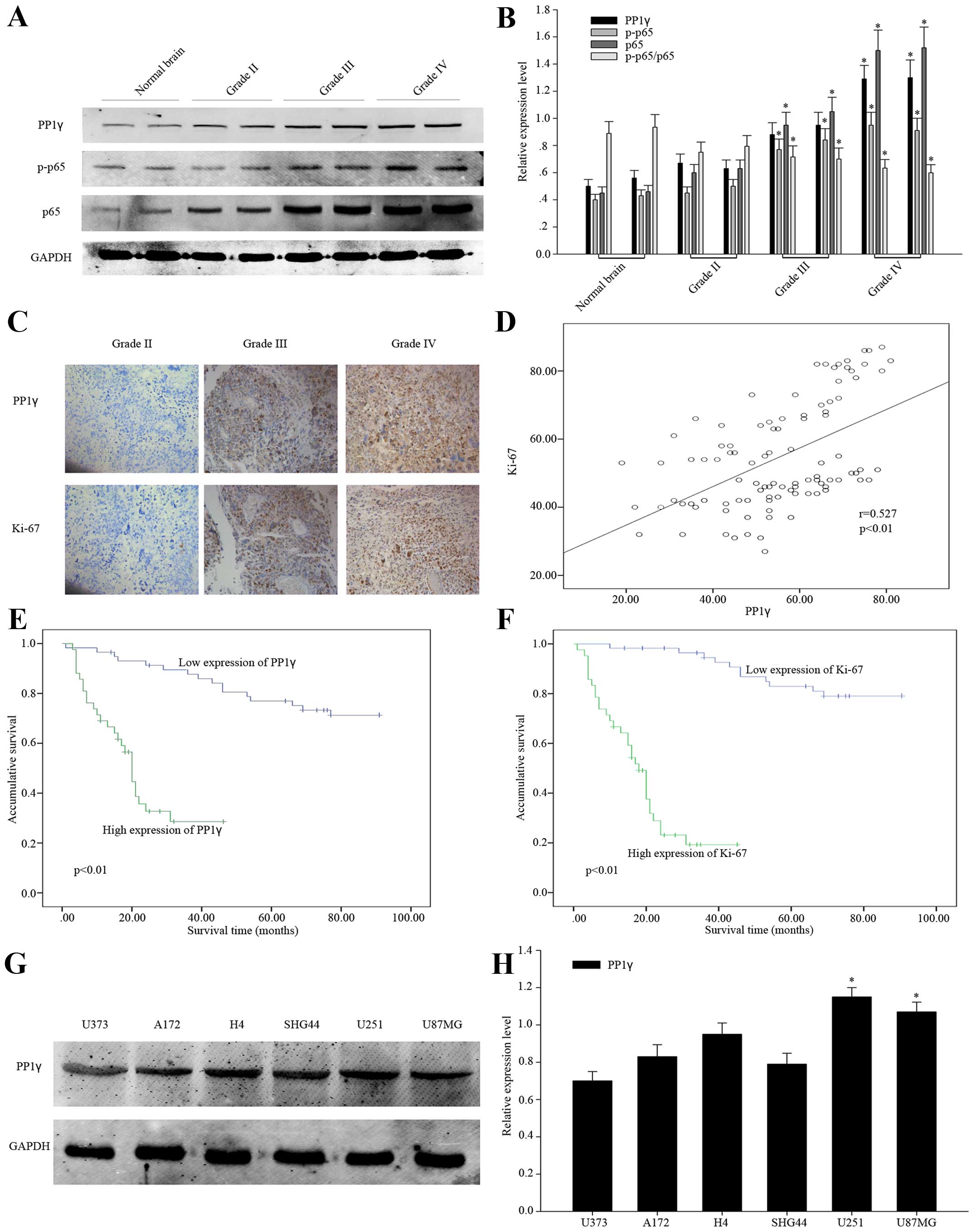
For investigating the possible role of PP1γ in the
development of glioma, we detected and compared the expression of
PP1γ, p-p65, p65 in normal brain tissues and three couples of
glioma tissues in the rank of WHO grade II, III and IV by western
blot analysis. According to statistical analysis, we observed
significant rising tendency of PP1γ in normal brain tissues,
low-grade and high-grade glioma tissues (Fig. 1A and B), as well as p-p65 and p65.
Especially, the relative expression of p-p65 is decreased after
being normalized to total p65. As an important reported
proliferation marker, Ki-67 was examined in 100 specimens of human
glioma tissues with PP1γ by immunohistochemical staining. It showed
consistent correlation of the expression of Ki-67 and PP1γ
(Fig. 1C and D), indicating that
PP1γ was related to the proliferation of glioma cells. Then, we
summarized the clinicopathological statistics and found the close
relationship of PP1γ to WHO grades of glioma (P<0.01), and it
was the same as Ki-67 used as cross reference (Table I). There was no obvious relevance to
other clinicopathological factors. Based on this, we considered
that PP1γ might have a great influence on the progression of glioma
relatively, and its overexpression might lead to poor
prognosis.
 | Table IThe correlation between PP1γ, and
Ki-67 expression and clinicopathological parameters in 100 glioma
specimens. |
Table I
The correlation between PP1γ, and
Ki-67 expression and clinicopathological parameters in 100 glioma
specimens.
| Variable | Total | PP1γ expression
| P-value | Ki-67 expression
| P-value |
|---|
| Low | High | Low | High |
|---|
| Age | | | | | | | |
| <40 | 17 | 12 | 5 | 0.291 | 12 | 5 | 0.291 |
| ≥40 | 83 | 46 | 37 | | 46 | 37 | |
| Grender | | | | | | | |
| Female | 37 | 22 | 15 | 0.494 | 24 | 13 | 0.304 |
| Male | 63 | 36 | 27 | | 34 | 29 | |
| Tumor location | | | | | | | |
| Frontal | 26 | 17 | 9 | 0.550 | 18 | 8 | 0.387 |
| Parietal | 25 | 14 | 11 | | 13 | 12 | |
| Occipital | 20 | 10 | 10 | | 10 | 10 | |
| Temporal | 23 | 12 | 11 | | 12 | 11 | |
| Unknown | 6 | 5 | 1 | | 5 | 1 | |
| Surgery | | | | | | | |
| Biopsy | 17 | 11 | 6 | 0.824 | 11 | 6 | 0.779 |
| Partial
resection | 41 | 23 | 18 | | 24 | 17 | |
| Gross total
resection | 42 | 24 | 18 | | 23 | 19 | |
| Vessel density | | | | | | | |
| Normal | 23 | 16 | 7 | 0.235 | 15 | 8 | 0.478 |
| Increase | 77 | 42 | 35 | | 43 | 34 | |
| Tumor diameter | | | | | | | |
| <4 cm | 41 | 24 | 17 | 0.547 | 24 | 17 | 0.547 |
| ≥4 cm | 59 | 34 | 25 | | 34 | 25 | |
| Necrosis | | | | | | | |
| Absence | 40 | 28 | 12 | 0.077 | 28 | 12 | 0.063 |
| Presence | 60 | 30 | 30 | | 30 | 30 | |
| WHO grade | | | | | | | |
| II | 36 | 27 | 9 | 0.004a | 28 | 8 | 0.001a |
| III | 35 | 21 | 14 | | 21 | 14 | |
| IV | 29 | 10 | 19 | | 9 | 20 | |
The relationship between the expression
of PP1γ in the specimens and the patient 5-year survival or the
prognosis of human glioma
To verify the presumption, we carried out
statistical analysis via SPSS. According to Kaplan-Meier survival
curves, the patients with high expression of PP1γ had lower
accumulative 5-year survival ratio (P<0.01). It was just the
same as the impact of overexpression of Ki-67 (Fig. 1E and F). Subsequently, we employed
the univariate survival analysis to compare the impact of other
clinicopathological factors on the 5-year survive to that of PP1γ
(P<0.05) and Ki-67 (P<0.01), and we concluded that the
overexpression of the two molecules was closely related to the poor
prognosis of the glioma patients (Table II).
 | Table IIContribution of various potential
prognostic factors to survival by univariate analysis in 100 glioma
specimens. |
Table II
Contribution of various potential
prognostic factors to survival by univariate analysis in 100 glioma
specimens.
|
Characteristics | Total | Survival status
| P-value |
|---|
| <5 years | ≥5 years |
|---|
| Age | | | | |
| <40 | 17 | 9 | 8 | 0.601 |
| ≥40 | 83 | 44 | 39 | |
| Grender | | | | |
| Female | 37 | 14 | 23 | 0.054 |
| Male | 63 | 39 | 24 | |
| Tumor location | | | | |
| Frontal | 26 | 13 | 13 | 0.821 |
| Parietal | 25 | 13 | 12 | |
| Occipital | 20 | 13 | 7 | |
| Temporal | 23 | 11 | 12 | |
| Unknown | 6 | 3 | 3 | |
| Surgery | | | | |
| Biopsy | 17 | 10 | 7 | 0.865 |
| Partial
resection | 41 | 21 | 20 | |
| Gross total
resection | 42 | 22 | 20 | |
| Vessel density | | | | |
| Normal | 23 | 13 | 10 | 0.813 |
| Increase | 77 | 40 | 37 | |
| Tumor diameter | | | | |
| <4 cm | 41 | 20 | 21 | 0.544 |
| ≥4 cm | 59 | 33 | 26 | |
| Necrosis | | | | |
| Absence | 40 | 18 | 22 | 0.223 |
| Presence | 60 | 35 | 25 | |
| WHO grade | | | | |
| II | 36 | 8 | 28 | 0.000a |
| III | 35 | 20 | 15 | |
| IV | 29 | 25 | 4 | |
| PP1γ
expression | | | | |
| Low | 58 | 25 | 33 | 0.016a |
| High | 42 | 28 | 14 | |
| Ki-67
expression | | | | |
| Low | 58 | 21 | 37 | 0.000a |
| High | 42 | 32 | 10 | |
The expression of PP1γ in glioma cell
lines and the relationship to cell proliferation
To identify the function of PP1γ in pathological
procession of glioma, we selected two glioma cell lines, U251 and
U87MG, for the following experiments, because PP1γ was shown to be
the highest expressed in these cells (Fig. 1G and H).
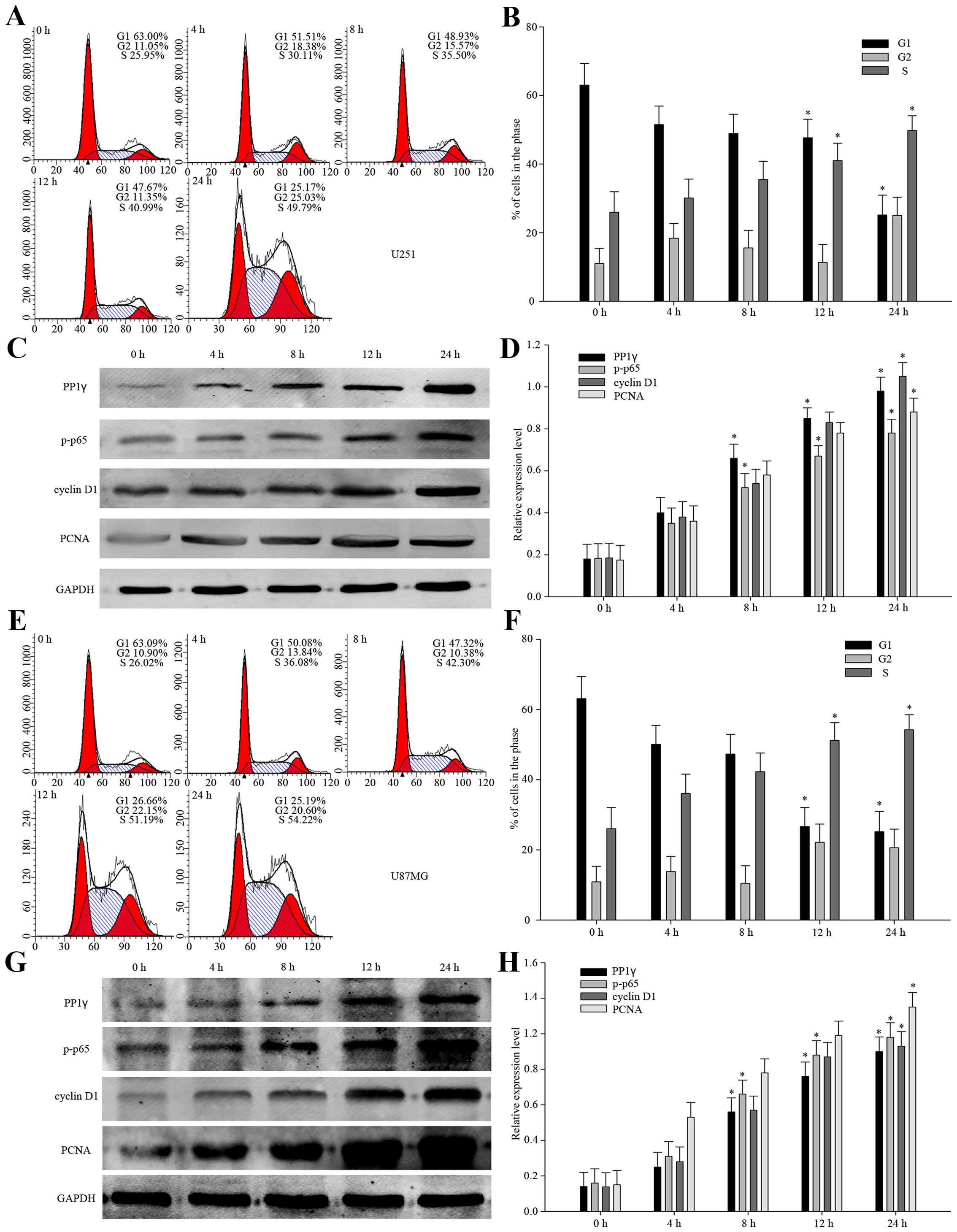
In view of the positive relationship between PP1γ
and Ki-67, overexpression of PP1γ was likely to accelerate cell
proliferation. To make it clear, we designed a serum-starvation and
re-feeding models with U251 and U87MG cell lines. The flow
cytometry analysis indicated that when the cells were controlled in
the serum deprivation environment for 72 h, the proportion of the
cells arrested in the G1 phase increased to 63.00 and 63.09%,
respectively. After re-feeding, the proportion of the cells in the
G1 phase reduced and that in S phase increased gradually (Fig. 2A, B, E and F). At the same time, we
detected the levels of the expression of PP1γ, p-p65 and reported
cell proliferation protein markers, such as PCNA, and cyclin D1.
Based on the results provided above (Fig. 2C, D, G and H), we could make the
tentative conclusion that PP1γ might be related to the
proliferation of glioma cells, and that it accelerates the
pathological process of glioma.
Downregulation of PP1γ inhibits the
proliferation of U251 and U87MG cell lines in vitro
Since we had uncovered that PP1γ was upregulated in
the starve-released cells, it was necessary to verify its
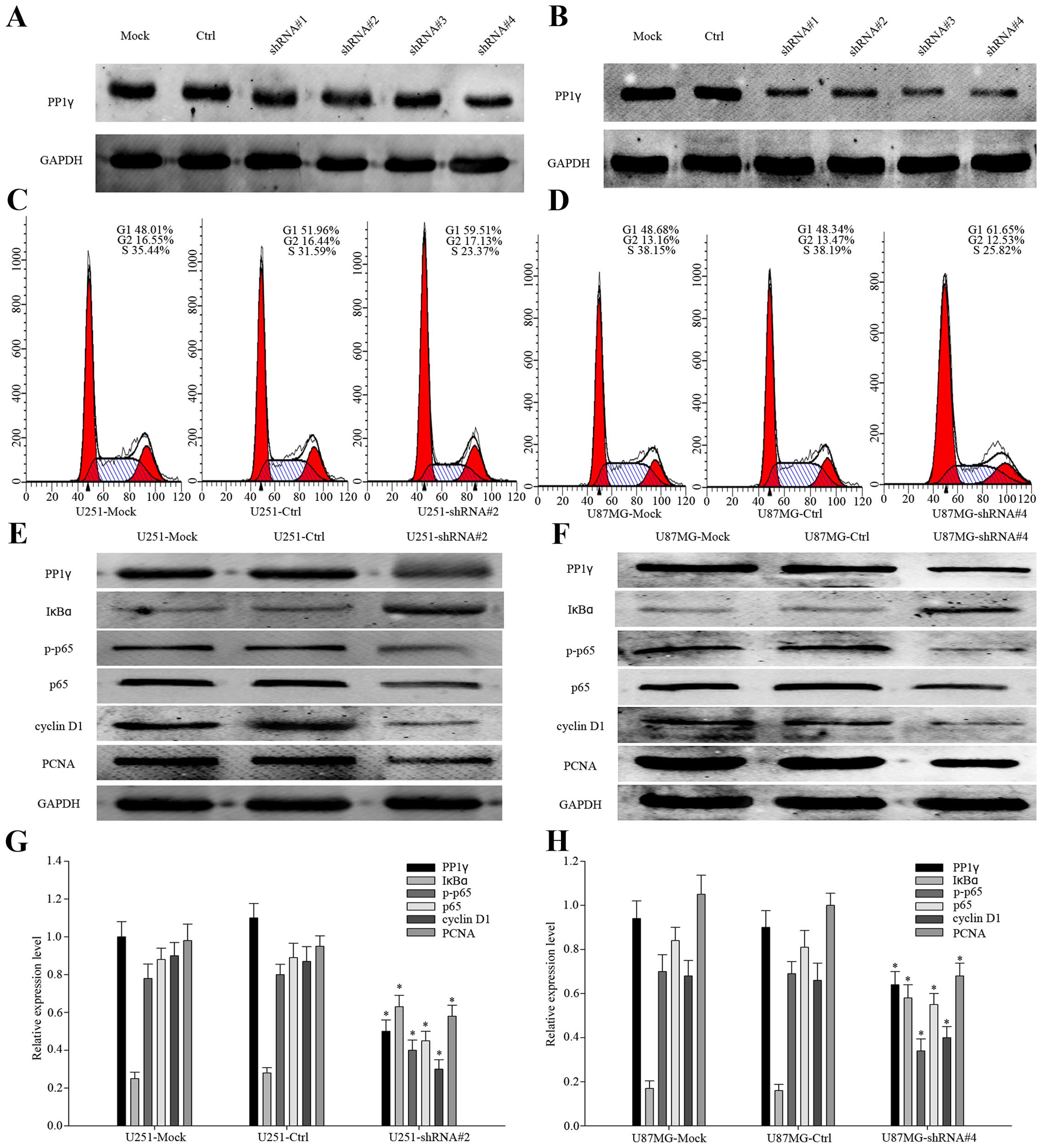
characteristics in the silencing of PP1γ. We transfected U251 and
U87MG cells with control-shRNA, PP1γ-shRNA#1, PP1γ-shRNA#2,
PP1γ-shRNA#3 and PP1γ-shRNA#4, respectively. The efficiency of all
the shRNAs was compared by western blot analysis (Fig. 3A and B). As shown, PP1γ-shRNA#2
achieved the best knock-down efficiency in U251 cell line, as
PP1γ-shRNA#4 to U87MG. We performed flow cytometry analyses to show
the proportion of the transfected cells in G1 and S phases
(Fig. 3C and D, P<0.01).
Obviously, on the one hand the cells of G1 phase increased from
48.01 to 59.51% in U251, 48.34 to 61.65% in U87MG, on the other
hand the cells of S phase decreased from 35.44 to 23.37% in U251,
38.19 to 25.82% in U87MG, which indicated that down-expression of
PP1γ delayed the cell cycle. Moreover, we detected the expression
of p-p65, p65, IκBα, cyclin D1, and PCNA (Fig. 3E and F). According to previous
studies, the activation of the NF-κB signaling cascade results in
complete degradation of IκB, allowing translocation of NF-κB to the
nucleus, where it induces transcription (23). It revealed the declined expression
of PP1γ and p-p65 accompanied by the increased IκBα, which meant
that PP1γ did play an important role in the cell cycle and cell
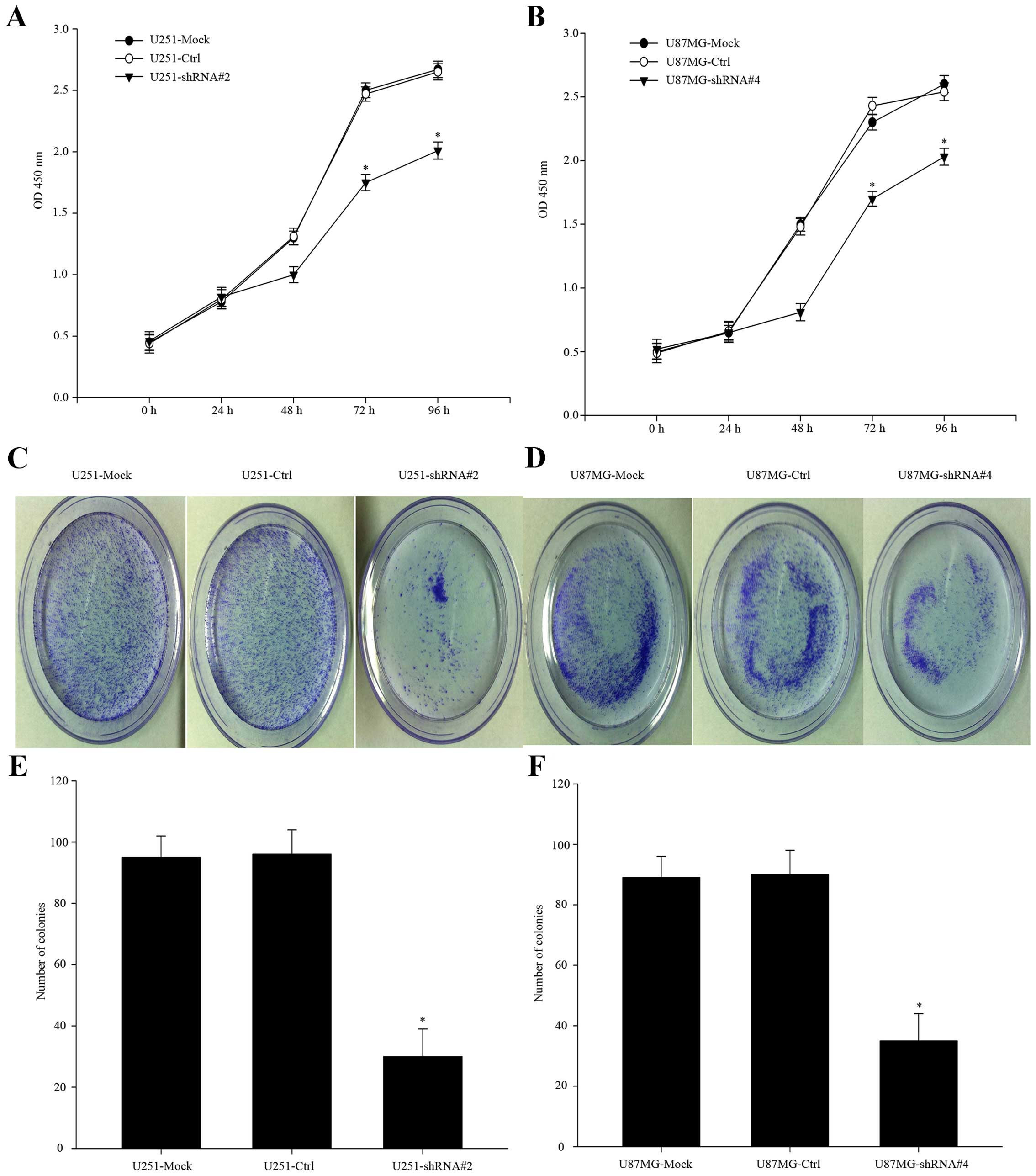
proliferation via the NF-κB pathway. To complete the evidence for
conclusion, we carried out CCK-8 assay to compare the proliferation
of PP1γ-depletion cells to the control-shRNA-transfected groups
(Fig. 4A and B, P<0.01). The
relative absorbance was less than that of the negative-control
group. In the colony formation assay, similar difference was
observed (Fig. 4C–F, P<0.01).
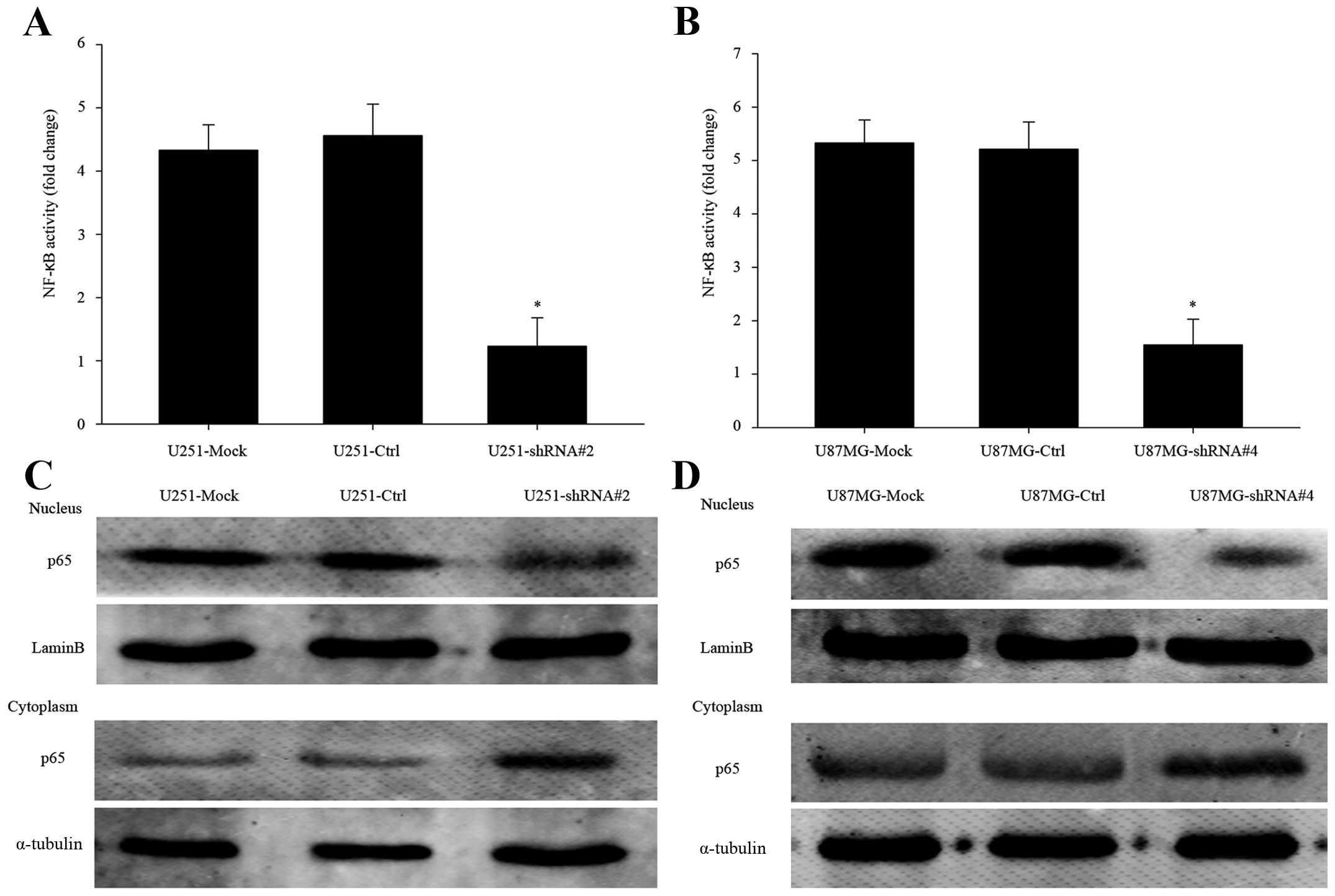
Besides, we extracted the nuclear of PP1γ-depletion cells for NF-κB
activity assay. The activity of NF-κB was obviously declined
comparing to negative groups (Fig. 5A
and B, P<0.01).
Depletion of PP1γ reduces p65 transport
into the nucleus
Via western blot analysis, we observed the clearly
decreased p65 expression in nucleus of PP1γ-depletion cells, and
its expression in cytoplasm was more than the control groups
(Fig. 5C and D). It showed that
knocking-down PP1γ actually declined p65 transport across the
nuclear envelope. In other words, PP1γ indeed played a role in
accelerating the NF-κB pathway, thus enhancing glioma cell
proliferation.
Discussion
Human glioma is the most common type of central
nervous system (CNS) malignancy (24). The current standard therapy includes
maximal safe resection followed by radiotherapy in combination with
temozolomide (25). It is still a
typical deadly cancer among the neurological oncology. Malignant
glioma cell proliferation and invasion are key stages in cancer
progression that affect mortality of the patients (26). Therefore, it is urgent to
investigate the underlying molecular pathological mechanism of
human glioma. In this study, we presented that PP1γ could be
involved in the molecule mechanisms for the proliferation of human
glioma, at least partially, via the NF-κB pathway.
Mammals express three PP1 catalytic isoforms, PP1α,
PP1γ, and PP1β/σ, which show distinct subcellular localization
patterns (27). PP1 catalytic
isoforms do not exist freely in cells but rather associate with
regulatory subunits to form distinct multimeric holoenzymes. In
general, PP1 regulatory subunits function as signaling modules by
regulating the enzymatic activity or targeting of catalytic
subunits to specific substrates (28). According to some research, PP1α and
PP1β/σ facilitates pRB-dependent DNA repair in retinoblastoma
(29). Besides, PP1α has been
characterized as an inhibitor of TNFR-induced NF-κB signaling
(30), suggesting that the PP1
isoforms may be involved in DNA repair in human glioma. However,
there is no enough evidence to demonstrate that the two isoforms
are involved in the field of cell proliferation.
Protein phosphatase 1γ (PP1γ) is one of the
serine/threonine protein phosphatases in human body which take part
in the reversible phosphorylation reaction of nearly 70% of all
eukaryotic proteins (31). It has
been demonstrated that PP1γ executes positive regulation in the
TRAF6-dependent immune responses (11), in which, NF-κB (p65) is activated
and translocated into nucleus. The activated NF-κB binds to
specific DNA sequences in target genes involving tumor cell
proliferation (10). Thus, we
investigated whether PP1γ played a role in human glioma. According
to the World Health Organization (WHO), human glioma is classified
into four grades as follows: pilocytic astrocytoma, WHO grade I;
diffuse 'low grades' glioma, WHO grade II; anaplastic gliomas, WHO
grade III; glioblastoma (GBM), WHO grade IV (32). We found the decreased relative
expression of p-p65 hinting that p-p65 was the substrate of PP1γ.
On the basis of systemic analysis of the clinicopathological
statistics from experienced doctors of pathology department in the
Affiliated Hospital of Nantong University, we identified the great
positive correlation between the overexpression of PP1γ and the WHO
grades of human glioma, which indicated a poor prognosis with a low
5-year survival rate. Moreover, employing the serum-starve and
re-feeding models, we observed the positive correlation of PP1γ,
p-p65 and proliferation markers, cyclin D1, and PCNA, demonstrating
its function in proliferation. However, the molecule mechanism of
PP1γ was still unclear.
Based on previous studies, PP1γ specifically
dephosphorylate a phosphor-site on TRAF6, its E2 enzyme complex,
allowing for full E3 ubiquitin ligase activity. Such a
dephosphorylation event may expose nearby residues for modification
by ubiquitin and enhance the enzymatic activity of TRAF6 (11). Moreover, the ubiquitination of TRAF6
is required for the induction of NF-κB activation and
osteoclastogenesis (12). On the
basis of recent studies, we made the hypothesis that PP1γ interacts
with TRAF6 via residues 315–354 of the coiled-coil domain and
enhances its ubiquitin activity. Then IKKγ is activated leading to
the degradation of IκBα, which promotes the phosphorylation of p65
and the translocation into the nucleus. TRAF6 might be the key
upstream molecule of p65. In the PP1γ-depletion cell models, we
observed remarkably reduced expression of PP1γ and p-p65 and
increased expression of IκBα in the cell lysate comparing to the
negative-control groups (P<0.01). It uncovered the possible
inhibition to the NF-κB pathway, which needed to be identified.
The activation of NF-κB signaling cascade results in
complete degradation of IκB, allowing translocation of NF-κB to the
nucleus, where it induces transcription (33). Activated NF-κB binds to specific DNA
sequences in target genes involving tumor cell proliferation,
invasion, metastasis, angiogenesis, chemoresistance,
radioresistance, inflammation and immune-regulation (10). It means the translocation of p65 is
necessary for the activation of transcription. Via western blot
analysis, the reduced expression of p65 was detected in cell
nucleus comparing to the negative-control groups (P<0.01). In
addition, the inhibition of cell proliferation was observed by flow
cytometry analysis of cell cycle, Cell Counting Kit (CCK)-8 assay
and colony formation assay. Directly, the NF-κB activity assay
testified the inactivation of PP1γ-depletion cells in contract to
negative groups (P<0.01). The evidence was powerful
demonstrating that the upregulated PP1γ activated NF-κB (p65) and
enhanced the translocation into nucleus, leading to cell
proliferation in human glioma. Besides, the ubiquitin ligase
activity of TRAF6 can be diminished by the absence of PP1γ
phosphatase activity (11).
Nevertheless, the molecule mechanism is unknown. It is possible
that PP1γ may specifically dephosphorylate an unknown inhibitory
phosphosite on TRAF6, its E2 enzyme complex, or one of its
substrates, allowing for full E3 ubiquitin ligase activity. To
demonstrate the hypothesis and explore other possible upstream
molecules, more experiments will be carried out in the following
research plans.
In conclusion, PP1γ was significantly upregulated in
human glioma accompanied by the inhibition of p65 translocation
into the nucleus. We identified strong positive correlation of the
overexpression of PP1γ, p-p65, p65 and the WHO grades of human
glioma. Moreover, we observed the rising expression trend of PP1γ,
p-p65 and proliferation markers in serum-starved models. The
reduced expression of PP1γ and p-p65 and the increased expression
of IκBα were detected in cell lysate of PP1γ-depletion models
comparing to negative-control groups. Besides, the decreased
expression of p65 was detected in the cell nucleus of
PP1γ-deleption models. These data are compatible with the
hypothesis that increased levels of PP1γ promoted cell
proliferation through NF-κB pathway in human glioma. Nevertheless,
further research is necessary to elucidate the function of PP1γ in
essential biological events and to clarify possible novel
therapeutic strategies in human glioma.
Acknowledgments
This study was supported by National Natural Science
Foundation of China (no. 81372687).
References
|
1
|
Deorah S, Lynch CF, Sibenaller ZA and
Ryken TC: Trends in brain cancer incidence and survival in the
United States: Surveillance, Epidemiology, and End Results Program,
1973 to 2001. Neurosurg Focus. 20:E12006. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Reardon DA, Herndon JE II, Peters KB,
Desjardins A, Coan A, Lou E, Sumrall AL, Turner S, Lipp ES,
Sathornsumetee S, et al: Bevacizumab continuation beyond initial
bevacizumab progression among recurrent glioblastoma patients. Br J
Cancer. 107:1481–1487. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Bertling E, Hotulainen P, Mattila PK,
Matilainen T, Salminen M and Lappalainen P: Cyclase-associated
protein 1 (CAP1) promotes cofilin-induced actin dynamics in
mammalian nonmuscle cells. Mol Biol Cell. 15:2324–2334. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Maher EA, Furnari FB, Bachoo RM, Rowitch
DH, Louis DN, Cavenee WK and DePinho RA: Malignant glioma: Genetics
and biology of a grave matter. Genes Dev. 15:1311–1333. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Zhuang W, Qin Z and Liang Z: The role of
autophagy in sensitizing malignant glioma cells to radiation
therapy. Acta Biochim Biophys Sin (Shanghai). 41:341–351. 2009.
View Article : Google Scholar
|
|
6
|
Wen PY and Kesari S: Malignant gliomas in
adults. N Engl J Med. 359:492–507. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Cohen PT: Protein phosphatase 1 - targeted
in many directions. J Cell Sci. 115:241–256. 2002.PubMed/NCBI
|
|
8
|
Varmuza S, Jurisicova A, Okano K, Hudson
J, Boekelheide K and Shipp EB: Spermiogenesis is impaired in mice
bearing a targeted mutation in the protein phosphatase 1cgamma
gene. Dev Biol. 205:98–110. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Bargou RC, Emmerich F, Krappmann D,
Bommert K, Mapara MY, Arnold W, Royer HD, Grinstein E, Greiner A,
Scheidereit C, et al: Constitutive nuclear factor-kappaB-RelA
activation is required for proliferation and survival of Hodgkin's
disease tumor cells. J Clin Invest. 100:2961–2969. 1997. View Article : Google Scholar
|
|
10
|
Borrello MG, Degl'Innocenti D and Pierotti
MA: Inflammation and cancer: The oncogene-driven connection. Cancer
Lett. 267:262–270. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Opaluch AM, Schneider M, Chiang CY, Nguyen
QT, Maestre AM, Mulder LC, Secundino I, De Jesus PD, König R, Simon
V, et al: Positive regulation of TRAF6-dependent innate immune
responses by protein phosphatase PP1-γ. PLoS One. 9:e892842014.
View Article : Google Scholar
|
|
12
|
Ang E, Pavlos NJ, Rea SL, Qi M, Chai T,
Walsh JP, Ratajczak T, Zheng MH and Xu J: Proteasome inhibitors
impair RANKL-induced NF-kappaB activity in osteoclast-like cells
via disruption of p62, TRAF6, CYLD, and IkappaBalpha signaling
cascades. J Cell Physiol. 220:450–459. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Yang Y, Blair HC, Shapiro IM and Wang B:
The proteasome inhibitor carfilzomib suppresses parathyroid
hormone-induced osteoclastogenesis through a RANKL-mediated
signaling pathway. J Biol Chem. 290:16918–16928. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Vallabhapurapu S and Karin M: Regulation
and function of NF-kappaB transcription factors in the immune
system. Annu Rev Immunol. 27:693–733. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Bettoun DJ, Buck DW II, Lu J, Khalifa B,
Chin WW and Nagpal S: A vitamin D receptor-Ser/Thr phosphatase-p70
S6 kinase complex and modulation of its enzymatic activities by the
ligand. J Biol Chem. 277:24847–24850. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Wang Y, Liu F, Mao F, Hang Q, Huang X, He
S, Wang Y, Cheng C, Wang H, Xu G, et al: Interaction with cyclin
H/cyclin-dependent kinase 7 (CCNH/CDK7) stabilizes C-terminal
binding protein 2 (CtBP2) and promotes cancer cell migration. J
Biol Chem. 288:9028–9034. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Tao T, Cheng C, Ji Y, Xu G, Zhang J, Zhang
L and Shen A: Numbl inhibits glioma cell migration and invasion by
suppressing TRAF5-mediated NF-κB activation. Mol Biol Cell.
23:2635–2644. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Huang X, Liu F, Zhu C, Cai J, Wang H, Wang
X, He S, Liu C, Yao L, Ding Z, et al: Suppression of KIF3B
expression inhibits human hepatocellular carcinoma proliferation.
Dig Dis Sci. 59:795–806. 2014. View Article : Google Scholar :
|
|
19
|
Ding Z, Liu X, Liu Y, Zhang J, Huang X,
Yang X, Yao L, Cui G and Wang D: Expression of far upstream element
(FUSE) binding protein 1 in human glioma is correlated with c-Myc
and cell proliferation. Mol Carcinog. 54:405–415. 2015. View Article : Google Scholar
|
|
20
|
Lin SY, Makino K, Xia W, Matin A, Wen Y,
Kwong KY, Bourguignon L and Hung MC: Nuclear localization of EGF
receptor and its potential new role as a transcription factor. Nat
Cell Biol. 3:802–808. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Giri DK, Ali-Seyed M, Li LY, Lee DF, Ling
P, Bartholomeusz G, Wang SC and Hung MC: Endosomal transport of
ErbB-2: Mechanism for nuclear entry of the cell surface receptor.
Mol Cell Biol. 25:11005–11018. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Liu Y, Wang Y, Cheng C, Chen Y, Shi S, Qin
J, Xiao F, Zhou D, Lu M, Lu Q, et al: A relationship between p27
(kip1) and Skp2 after adult brain injury: Implications for glial
proliferation. J Neurotrauma. 27:361–371. 2010. View Article : Google Scholar
|
|
23
|
Ahn KS, Sethi G and Aggarwal BB: Nuclear
factor-kappa B: From clone to clinic. Curr Mol Med. 7:619–637.
2007. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Ferguson SD: Malignant gliomas: Diagnosis
and treatment. Dis Mon. 57:558–569. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Stupp R, Hegi ME, van den Bent MJ, Mason
WP, Weller M, Mirimanoff RO and Cairncross JG; European
Organisation for Research and Treatment of Cancer Brain Tumor and
Radiotherapy Groups; National Cancer Institute of Canada Clinical
Trials Group: Changing paradigms - an update on the
multidisciplinary management of malignant glioma. Oncologist.
11:165–180. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Qi S, Song Y, Peng Y, Wang H, Long H, Yu
X, Li Z, Fang L, Wu A, Luo W, et al: ZEB2 mediates multiple
pathways regulating cell proliferation, migration, invasion, and
apoptosis in glioma. PLoS One. 7:e388422012. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Lee JH, You J, Dobrota E and Skalnik DG:
Identification and characterization of a novel human PP1
phosphatase complex. J Biol Chem. 285:24466–24476. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Virshup DM and Shenolikar S: From
promiscuity to precision: Protein phosphatases get a makeover. Mol
Cell. 33:537–545. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Lin CY, Tan BC, Liu H, Shih CJ, Chien KY,
Lin CL and Yung BY: Dephosphorylation of nucleophosmin by PP1β
facilitates pRB binding and consequent E2F1-dependent DNA repair.
Mol Biol Cell. 21:4409–4417. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Li HY, Liu H, Wang CH, Zhang JY, Man JH,
Gao YF, Zhang PJ, Li WH, Zhao J, Pan X, et al: Deactivation of the
kinase IKK by CUEDC2 through recruitment of the phosphatase PP1.
Nat Immunol. 9:533–541. 2008. View
Article : Google Scholar : PubMed/NCBI
|
|
31
|
Olsen JV, Vermeulen M, Santamaria A, Kumar
C, Miller ML, Jensen LJ, Gnad F, Cox J, Jensen TS, Nigg EA, et al:
Quantitative phosphoproteomics reveals widespread full
phosphorylation site occupancy during mitosis. Sci Signal.
3:ra32010. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Radner H, Blümcke I, Reifenberger G and
Wiestler OD: The new WHO classification of tumors of the nervous
system 2000. Pathology and genetics. Pathologe. 23:260–283. 2002.In
German. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Ahn KS and Aggarwal BB: Transcription
factor NF-kappaB: A sensor for smoke and stress signals. Ann NY
Acad Sci. 1056:218–233. 2005. View Article : Google Scholar
|