Introduction
More than 350 million people are infected with
chronic hepatitis B virus (HBV) worldwide, and are at risk of
developing liver diseases, such as chronic hepatitis, cirrhosis and
hepatocellular carcinoma (HCC) (1).
Due to widespread use of the HBV vaccine, the global pandemic of
HBV has decreased in prevalence, but the absolute number of
HBsAg-positive individuals is still increasing (2). In addition, there are huge populations
with chronic HBV infection, particularly in East Asia, and
HBV-related liver diseases, particularly HCC. Hence, HBV-related
liver diseases will continue to be a public health burden for
decades.
Due to the rapid growth of cancer cells and limited
oxygen from the environment, hypoxia is a common feature in cancer
(3). In solid tumors such as HCC,
the cancer cells encounter low oxygen pressure stress, particularly
those cancer cells encompassed in tumor tissues (4). In order to adapt to hypoxic stress,
cancer cells induce neovasculature, breaking the natural barriers
to tumor angiogenesis (3), thus
pushing cancer to be more aggressive. Hypoxic stress induces the
expression of hypoxia-induced factor 1α (HIF-1α), which
translocates into the nucleus and acts as a transcription factor to
initiate the expression of downstream genes and one master hypoxic
microRNA (miRNA), miR-210 (5).
miR-210 regulates hypoxia-induced intracellular pathways, including
cell cycle progression (6–9), cell survival (10,11),
genome stabilization (12), cell
differentiation (13,14), angiogenesis (15–24)
and cell metabolism (25–27). The hypothesis that upregulated
miR-210 may help cancer cells adapt to neoplastic stress has been
partly proved by the increased potential of metastasis in HCC cells
(28), and the increased cell
growth, survival, genome destabilization and angiogenesis in other
human tumor models (29). Moreover,
miR-210 is a regulator of multiple function in terms of
carcinogenesis and progression, and it can act as both oncogene and
tumor suppressor under different conditions or in different types
of cancer (30).
miRNAs are special non-coding RNAs, which are
involved in a wide array of cellular processes, including
differentiation, proliferation, apoptosis, stress, carcinogenesis
and progression (31,32). In recent years, miRNAs have been
identified as important regulators in liver diseases, including HCC
(33). Among these miRNAs,
upregulation of miR-210 in HCC tumor tissues was first reported in
genomic screening studies (34–36),
and its upregulation in the serum of HCC patients was also reported
(37). Functional studies showed
that the upregulation of hypoxic miR-210 in HCC induced cell
metastasis and radiotherapy resistance, although its role in cell
proliferation was not consistent possibly due to different
experimental approaches (28,38,39).
However, it remains unclear whether miR-210 contributes to HCC
angiogenesis, and whether deregulation of miR-210 can be a new
prognosis predictor for HCC patients.
In the present study, we examined the roles of
miR-210 in the prognosis and angiogenesis of HBV-related HCC, using
both clinical samples and cell models. We found that miR-210
expression is upregulated in HCC tumor tissues, and high expression
of miR-210 in HCC tissues is an independent risk factor for both
tumor-free and overall survival of HCC patients. In addition, the
relative miR-210 expression in tumor/non-tumor of HCC patients may
serve as a prognostic factor. Moreover, the present study provides
evidence to show that miR-210 promoted HCC angiogenesis, and the
corresponding mechanism was identified to be the direct targeting
and inhibition of fibroblast growth factor receptor-like 1 (FGFRL1)
expression. Thus, we suggested a new prognosis predictor for HCC
patients, and determined the roles of hypoxic miR-210 in HCC
angiogenesis.
Materials and methods
Patients
All clinical samples were obtained with informed
consent from the HCC surgery undertaken in Eastern Hepatobiliary
Surgery Hospital, Shanghai, China. From 2007 to 2010, 212 paired
HCC and non-tumor samples from HBV-related HCC patients were
examined, who were HBV-positive (HBsAg, HBV DNA or HBeAg
serum-positive) and HCV-negative (anti-HCV serum-negative)
(Table I). Thirty-one non-tumor
tissues served as healthy liver controls, and were obtained from
hepatic hemangioma patients, without HBV infection (serum HBsAg and
anti-HBc-negative). The present study was approved by the Ethics
Committee of Eastern Hepatobiliary Surgery Hospital.
 | Table IComparison of clinicopathological and
demographic characteristics of patients with high and low miR-210
expression level. |
Table I
Comparison of clinicopathological and
demographic characteristics of patients with high and low miR-210
expression level.
| miR-210 high
expression
(n=106) | miR-210 low
expression
(n=106) | P-value |
|---|
| Gender | | | 0.369 |
| Male | 97 (91.51) | 93 (87.73) | |
| Female | 9 (8.49) | 13 (12.27) | |
| Age (years)a | 49.77±10.63 | 50.89±10.14 | 0.936 |
| Cirrhosis | | | 0.411 |
| Yes | 51 (48.11) | 57 (53.77) | |
| No | 55 (51.89) | 49 (46.23) | |
| HBeAg | | | 0.434 |
| Positive | 30 (28.30) | 25 (23.58) | |
| Negative | 76 (71.70) | 81 (76.42) | |
| AFP (ng/ml) | | | 0.674 |
| ≥20 | 66 (62.26) | 63 (59.43) | |
| <20 | 40 (37.74) | 43 (40.57) | |
| Alanine
aminotransferase (U/l) | | | 0.088 |
| ≥40 | 45 (42.45) | 33 (31.13) | |
| <40 | 61 (57.55) | 73 (68.87) | |
| Aspartate
aminotransferase (U/l) | | | 0.059 |
| ≥40 | 42 (39.62) | 29 (27.36) | |
| <40 | 64 (60.38) | 77 (72.64) | |
| Total bilirubin
(ummol/ml) | | | 0.465 |
| ≥17.1 | 37 (34.90) | 32 (30.19) | |
| <17.1 | 69 (65.10) | 74 (69.81) | |
| Albumin (g/l) | | | 0.361 |
| ≥35 | 73 (68.87) | 79 (74.52) | |
| <35 | 33 (31.13) | 27 (25.48) | |
| HBV DNA
(IU/ml) | | | 0.321 |
| ≥2,000 | 63 (59.43) | 70 (66.04) | |
| <2,000 | 43 (40.57) | 36 (33.96) | |
| Tumor diameter
(cm)a | 7.28±4.07 | 6.29±3.89 | 0.035b |
| Tumor
encapsulation | | | 0.022b |
| None | 76 (71.70) | 60 (56.60) | |
| Complete | 30 (28.30) | 46 (43.40) | |
| Vascular
invasion | | | 0.008b |
| Yes | 52 (49.06) | 33 (31.13) | |
| No | 54 (50.94) | 73 (68.87) | |
| Tumor number | | | 0,208 |
| Single | 83 (78.30) | 75 (70.75) | |
| Multiple | 23 (21.70) | 31 (29.25) | |
| Tumor
differentiation (BCLC) | | | 0.014b |
| I/II | 10 (9.43) | 23 (21.70) | |
| III/IV | 96 (90.57) | 83 (78.30) | |
Cells, plasmids, reagents and
antibodies
Cells: HL-7702 and SMMC-7721 cells were maintained
in RPMI-1640 medium (HyClone, Logan, UT, USA) supplemented with 10%
fetal bovine serum (FBS; Gibco, Carlsbad, CA, USA). Huh7 and 293T
cells were maintained in Dulbecco's modified Eagle's medium (DMEM)
(HyClone) supplemented with 10% FBS. The cells were purchased from
the Cell Bank of Institutes for Biological Sciences, Chinese
Academy of Sciences, Shanghai. Primary human umbilical vein
endothelial cells (HUVECs) were purchased from Lonza and were
maintained in EGM2 medium (Lonza, Walkersville, MD, USA). Plasmids:
the pre-miR-210 fragment was cloned from human hepatocyte genome
into the pCDH-CMV-MCS-EF1-puro vector (SBI), for miR-210 lentivirus
production and infection. Lentivirus package plasmids, CMV-VSVG and
δ-8.9, were purchased from SBI. Lentivirus production, infection
and establishment of stable transduced cell lines were performed
according to the manufacturer's protocol. Reagents: TRIzol agent
(Invitrogen, Carlsbad, CA, USA), puromycin, deferoxamine (DFX)
(both from Sigma, St. Louis, MO, USA), Matrigel™ Matrix (BD
Biosciences, Bedford, MA, USA), recombinant mouse FGF R5/FGFRL1 Fc
Chimera (R&D Systems, Minneapolis, MN, USA). miR-210 mimic,
mimic control, miR-210 inhibitor and inhibitor control (RiboBio,
Guangzhou, China). BCA protein assay kit (Beyotime, Jiangsu, China)
and Complete Protease Inhibitor Cocktail Tablets (Roche).
Antibodies: anti-HIF1α, anti-FGFRL1 and anti-GAPDH (Sigma),
anti-human and mouse CD34 (Abcam, Cambridge, MA, USA).
RNA extraction and miRNA qRT-PCR
Total RNA was isolated from clinical samples or
cultured cells by TRIzol according to the manufacturer's protocol.
The quality of the RNA was assessed by NanoDrop 2000 (NanoDrop
Technologies, Wilmington, DE, USA). miRNA qRT-PCR was performed
using the TaqMan Human miRNA Assay kit (Applied Biosystems, Foster
City, CA, USA) according to the manufacturer's instructions. The
expression of miR-210 was normalized relative to the expression of
small nuclear RNA U6, as an internal control. Data were analyzed
using the comparative Ct method (2−ΔΔCt).
Protein purification and western blot
(WB) assay
Secreted FGFRL1 protein in the culture medium was
purified by heparin-sepharose as previously described (40). Briefly, heparin-sepharose was washed
with phosphate-buffered saline (PBS) twice, and then added to PBS
1:1 diluted culture medium, the culture medium was filtered using a
0.45 μm filter. Heparin-sepharose and the medium mixture
were rotated at 4°C for 4 h. Heparin-sepharose was pelleted using a
centrifuge and washed 5 times with PBS. Heparin binding proteins
were eluted with Laemmli buffer at 95°C, and then subjected to
SDS-PAGE for WB assay. For WB assay, the cells were lysed in lysis
buffer (50 mM Tris, pH 8.0, 150 mM NaCl, 1% Nonidet P-40 and
complete protease inhibitor mixture), for 30 min at 4°C. The cell
lysates were centrifuged at 12,000 × g and 4°C for 10 min, and the
supernatant was used in the next step. The cell lysates were
quantified using the BCA kit and adjusted to the same level in each
experiment. The protein samples were then subjected to SDS-PAGE and
transferred to nitrocellulose membranes for WB assay.
Tube formation assay
The protocol for endothelial cell tube formation
assay was based on the procedure outlined by Corning (Corning, NY,
USA). Briefly, for pre-treatment, HUVEC cells were serum-starved 12
h before the tube formation assay, and the Matrigel™ Matrix was
thawed on ice overnight. Thawed Matrigel Matrix (50 μl) was
added to each well of a pre-chilled 96-well sterile plate. The
plate was incubated for 30 min at 37°C to allow the Matrigel Matrix
to form a gel. HUVEC cells were harvested and re-suspended in the
indicated medium at 1.5×105 cells/ml. Cell suspension
(100 μl) (1.5×104 cells) was added to each well
containing solidified Matrigel Matrix. The assay plate was
incubated at 37°C for 4 h. The endothelial tubes were pictured for
calculation using the angiogenesis-analyzer tool plugin for
ImageJ.
Microvessel density (MVD) CD34 assay
Immunohistochemical staining for CD34 in both human
and mouse liver sections was performed in the Immunohistochemistry
Department of Eastern Hepatobiliary Surgery Hospital. The MVD CD34
value was assessed using the method developed by Weidner et
al (41). Briefly, the CD34
immunohistochemically stained tissue sections were examined in the
200× field, 5 hotspots were pictured for MVD assessment. Any
endothelial cell or endothelial-cell cluster stained brown which
was clearly separate from adjacent microvessels, tumor cells and
other connective tissue elements was considered a single countable
microvessel. The average of the 5 hotspots in each sample was
considered the MVD CD34 value.
Xenograft growth
miR-210 stably transduced SMMC-7721 and control
cells were trypsinized and re-suspended in the medium. Male BALB/c
nude mice (6-weeks-old) were subcutaneously inoculated in the
dorsal area with 2.0×106 cells in 0.1 ml. Six weeks
after inoculation, the mice were sacrificed and the tumors were
dissected and fixed for CD34 immunohistochemical staining.
Statistical analysis
Data are expressed as the mean ± SEM. Differences
between groups were assessed using the t-test or Mann-Whitney U
test. Time-to-recurrence was calculated according to the
Kaplan-Meier method and compared using the log-rank test. All
statistical analyses were performed using Prism GraphPad software
(GraphPad Software, La Jolla, CA, USA). A P-value <0.05 denoted
the presence of a statistically significant difference.
Results
Expression of miR-210 is upregulated in
HBV-related HCC
In order to examine the expression of hypoxic
miR-210 in HCC, miR-210 expression was measured by qRT-PCR in 212
paired HCC tumor and non-tumor samples obtained from surgery
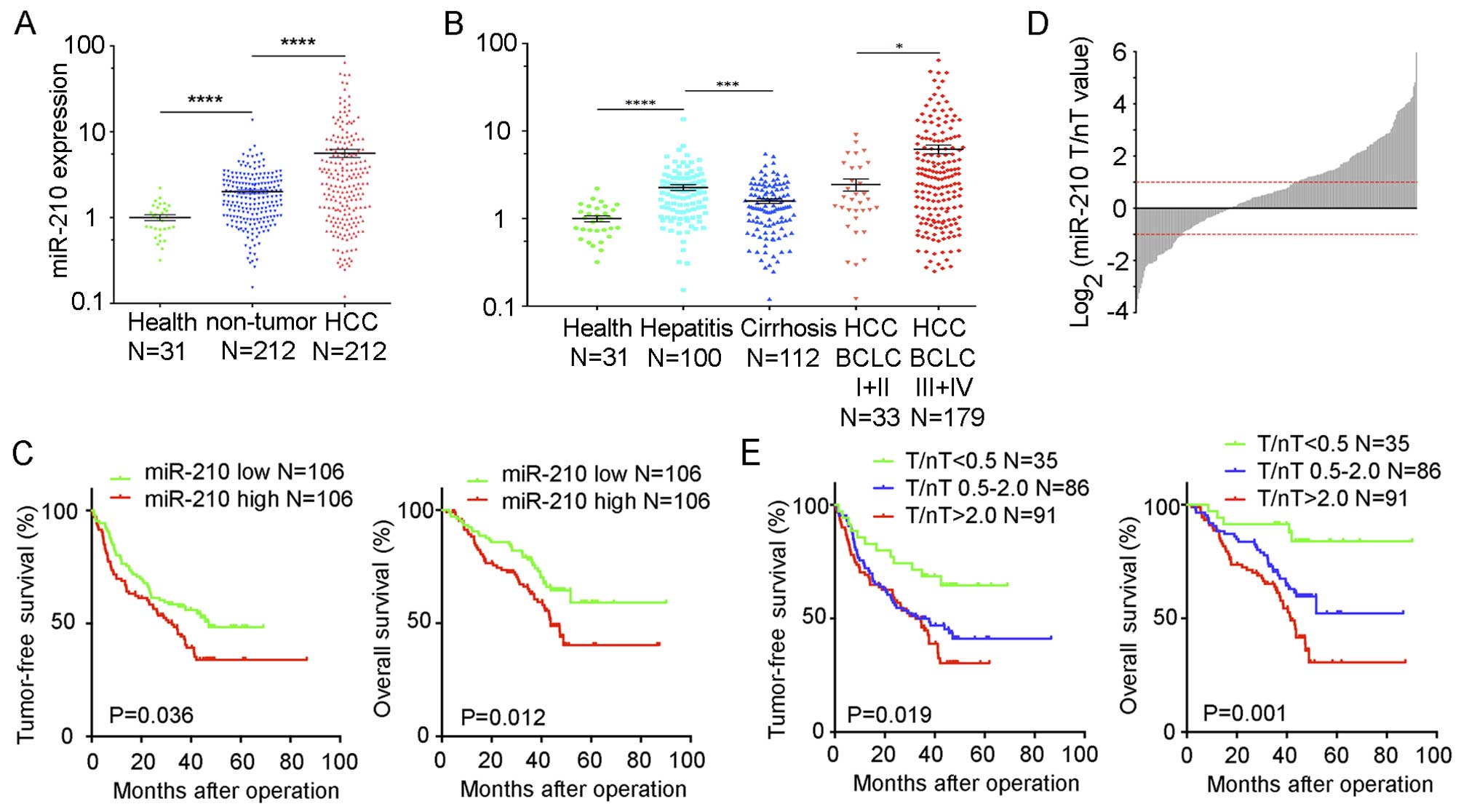
(Fig. 1A). miR-210 expression in
HCC tissues was significantly higher than that in non-tumor
tissues. In addition, to determine whether hepatitis stress induced
miR-210 expression, 31 healthy liver samples obtained from
HBV-negative hepatic hemangioma patients were examined, and we
found that miR-210 expression level in healthy liver was
significantly lower than that in the HCC non-tumor tissues
(Fig. 1A). Thus, miR-210 expression
was increased progressively from normal liver and adjacent
non-tumor tissues, to HCC tissues.
We also determined whether miR-210 expression was
correlated with the progression of liver disease. According to the
pathological reports, we divided non-tumor samples into the
hepatitis group (100 patients) and cirrhosis group (112 patients),
the HCC samples were divided into the BCLC I+II group (33 patients)
and the BCLC III+IV group (179 patients). The hepatitis group had a
higher miR-210 expression level than that in the cirrhosis group
(P=0.0093), and the BCLC III+IV group had a higher miR-210
expression level than that in the BCLC I+II group (P=0.029)
(Fig. 1B). The decrease of miR-210
in the cirrhosis group suggests that miR-210 expression may be
correlated with liver damage (42),
and the increase of miR-210 in the BCLC III+IV group suggests that
miR-210 induction is correlated with HCC progression.
miR-210 expression is correlated with the
prognosis of HCC patients
To analyze the correlation between miR-210
expression and HCC prognosis, we divided the patients into two
groups according to miR-210 expression in HCC using the median
value as the cut-off. The clinicopathological and demographic
characteristics of the patients in these two groups are listed in
Table I. Compared to the miR-210
low group, miR-210 high group had significantly less tumor
encapsulation (P=0.020), larger tumor size (7.82±4.07 vs. 6.29±3.89
cm; P=0.035), more vascular invasion (P=0.008) and a higher
proportion of undifferentiated tumors (P=0.014). Furthermore,
patients with high miR-210 expression in HCC had significantly
shorter tumor-free survival (P=0.036; Fig. 1C) and overall survival (P=0.012;
Fig. 1C), as compared to that of
the miR-210 low patients. Moreover, multivariate analysis showed
that HCC miR-210 expression (P=0.03; Table II) and vascular invasion (P=0.013;
Table II) were significant
independent factors for tumor-free survival, and HCC miR-210
expression (P=0.012; Table III)
and tumor diameter (P=0.001; Table
III) were significant independent factors for overall survival.
Thus, miR-210 expression is significantly correlated with the
prognosis of HCC patients.
 | Table IIUnivariate and multivariate analysis
of factors associated with tumor-free survival of HCC patients. |
Table II
Univariate and multivariate analysis
of factors associated with tumor-free survival of HCC patients.
| Factors | Univariate analysis
| Multivariate
analysis
|
|---|
| HR (95% CI) | P-value | HR (95% CI) | P-value |
|---|
| Age (years) | 0.987
(0.620–1.571) |
0.957 | – | – |
| Gender | 0.688
(0.359–1.316) |
0.258 | – | – |
| Liver cirrhosis
(yes vs. no) | 0.946
(0.657–1.362) |
0.765 | – | – |
| Tumor
differentiation (III+IV vs. I+II) | 1.217
(0.717–2.065) |
0.468 | – | – |
| Tumor diameter (≥5
vs. <5 cm) | 1.375
(0.947–1.997) |
0.094 | – | – |
| Tumor encapsulation
(no vs. yes) | 1.374
(0.938–2.015) |
0.103 | – | – |
| Alanine
aminotransferase (≥40 vs. <40 U/l) | 1.393
(0.964–2.012) |
0.078 | – | – |
| Total bilirubin
(≥17.1 vs. <17.1 ummol/ml) | 1.014
(0.852–1.206) |
0.879 | – | – |
| HBV DNA (≥2,000 vs.
<2,000 IU/ml) | 1.354
(0.916–2.002) |
0.129 | – | – |
| HBeAg (positive vs.
negative) | 1.130
(0.755–1.693) |
0.552 | – | – |
| AFP (≥20 vs. <20
ng/ml) | 1.323
(0.895–1.956) |
0.161 | – | – |
| Aspartate
aminotransferase (≥40 vs. <40 U/l) | 1.631
(1.130–2.355) |
0.009a | 1.363
(0.933–1.991) | 0.110 |
| Tumor number
(multiple vs. single) | 1.834
(1.236–2.722) |
0.003a | 1.504
(0.988–2.290) | 0.057 |
| miR-210 expression
(high vs. low) | 1.549
(1.071–2.239) |
0.02a | 1.515
(1.041–2.203) | 0.03a |
| Vascular invasion
(yes vs. no) | 1.963
(1.345–2.865) | <0.001a | 1.675
(1.116–2.513) | 0.013a |
 | Table IIIUnivariate and multivariate analysis
of factors associated with overall survival of HCC patients. |
Table III
Univariate and multivariate analysis
of factors associated with overall survival of HCC patients.
| Factors | Univariate analysis
| Multivariate
analysis
|
|---|
| HR (95% CI) | P-value | HR (95% CI) | P-value |
|---|
| Age (years) | 1.029
(0.597–1.774) |
0.918 | – | – |
| Gender | 0.691
(0.319–1.498) |
0.349 | – | – |
| Liver cirrhosis
(yes vs. no) | 0.927
(0.606–1.419) |
0.728 | – | – |
| Tumor
differentiation (III+IV vs. I+II) | 1.987
(0.994–3.974) |
0.052 | – | – |
| Tumor encapsulation
(no vs. yes) | 1.386
(0.886–2.168) |
0.153 | – | – |
| Alanine
aminotransferase (≥40 vs. <40 U/l) | 1.345
(0.876–2.066) |
0.176 | – | – |
| Total bilirubin
(≥17.1 vs. <17.1 ummol/ml) | 0.863
(0.619–1.205) |
0.387 | – | – |
| HBV DNA (≥2,000 vs.
<2,000 IU/ml) | 1.447
(0.911–2.300) |
0.118 | – | – |
| HBeAg (positive vs.
negative) | 1.069
(0.661–1.727) |
0.786 | – | – |
| Vascular invasion
(yes vs. no) | 2.505
(1.616–3.881) | <0.001a | 1.365
(0.916–2.213) | 0.163 |
| AFP (≥20 vs. <20
ng/ml) | 1.645
(1.025–2.641) |
0.039a | 1.364
(0.867–2.145) | 0.179 |
| Aspartate
aminotransferase (≥40 vs. <40 U/l) | 1.800
(1.176–2.758) |
0.007a | 1.426
(0.958–2.195) | 0.152 |
| Tumor number
(multiple vs. single) | 1.732
(1.088–2.757) |
0.021a | 1.278
(0.767–2.131) | 0.346 |
| miR-210 expression
(high vs. low) | 1.817
(1.173–2.814) |
0.007a | 1.781
(1.136–2.794) | 0.012a |
| Tumor diameter (≥5
vs. <5 cm) | 1.575
(1.015–2.446) |
0.043a | 2.181
(1.349–3.527) | 0.001a |
To further assess the possible prognostic value of
miR-210, we evaluated miR-210 expression in non-tumor liver
tissues. The expression of miR-210 in non-tumor tissues was not
correlated with tumor-free survival (P=0.479) or overall survival
(P=0.290) of HCC patients (data not shown). However, the
distribution of relative tumor/non-tumor (T/nT) miR-210 expression
suggested the complicated regulation of miR-210 expression in the
tumor microenvironment (Fig. 1D).
According to the T/nT miR-210 expression fold-change, we used one
fold as the cut-off value (35) and
further divided all patients into 3 groups, T/nT >2.0 (91
patients), T/nT <0.5 (35 patients), and T/nT 0.5–2.0 (86
patients). The Kaplan-Meier analysis of both tumor-free and overall
survival showed that the 3 groups of patients were clearly
distinguished (P=0.0001; Fig. 1E),
suggesting that miR-210 expression T/nT <0.5 may serve as a
favorable prognostic factor for HBV-related HCC. Thus, miR-210
expression in HCC, particularly the T/nT value, is a new prognosis
predictor for HCC patients.
miR-210 promotes HCC angiogenesis
As the role of miR-210 in HCC angiogenesis remains
unclear, we then investigated the correlation between miR-210
expression in HCC and intratumor MVD. The results of CD34
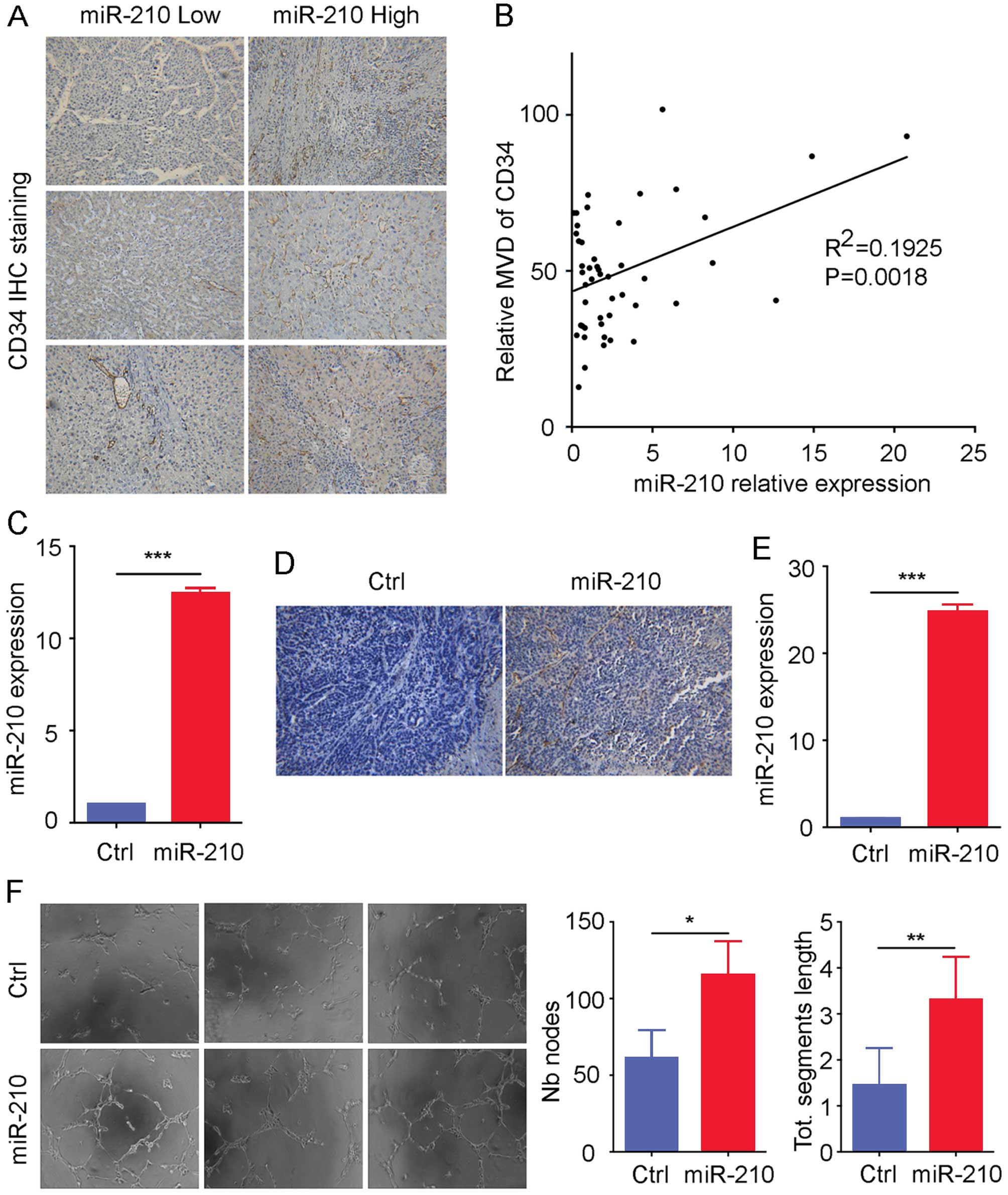
immunohistochemistry showed that HCC tissues with high miR-210
expression had high CD34 staining (Fig.
2A). In addition, miR-210 expression in HCC was positively
correlated with the MVD CD34 value (R2=0.1925; P=0.0018; Fig. 2B), in a randomly selected group of
48 HCC patients. To determine whether miR-210 is an independent
angiogenesis inducer, we established miR-210 stably overexpressed
HCC cell lines SMMC-7721 and HL-7702 and the corresponding control
cell lines for in vivo and in vitro assays. The
expression level of miR-210 was confirmed by qRT-PCR in each cell
line (Fig. 2C and E), and we
inoculated both control and miR-210 stably overexpressed SMMC-7721
cells into nude mice. CD34 staining of the xenograft showed that
miR-210 stably overexpressed SMMC-7721 cells had higher CD34
staining as compared to that of control cells (Fig. 2D), suggesting that HCC cells with
increased miR-210 may send a signal to recruit more endothelial
cells for vascular formation. Moreover, we used the medium
supernatants from control and miR-210 stably overexpressed HL-7702
cells for the tube formation assay, and the result showed that the
supernatants from HL-7702 of increased miR-210 expression induced
greater tube formation (Fig. 2F).
Thus, we conclude that increased miR-210 expression in HCC promotes
angiogenesis, which may be dependent on the secreted cytokines.
miR-210 targets FGFRL1 expression and
secretion in HCC
We next investigated the target genes of miR-210,
which are responsible for miR-210 mediated promotion of HCC
angiogenesis. As the target genes of miR-210 have been studied in
several genomic approaches (8,43), we
only screened for the experimentally proved target genes, which may
regulate angiogenesis with extracellular secretion capability and
which were relatively conserved between human and mouse. One of the
target genes of miR-210, FGFRL1, was found to fulfill all the
criteria listed above. FGFRL1, also known as FGFR5, and belongs to
the fibroblast growth factor receptor (FGFR) family. FGFRs consist
of an extracellular region of 3 immunoglobulin-like domains, a
single hydrophobic membrane-spanning segment, and a cytoplasmic
tyrosine kinase domain for cascading downstream signals. The
extracellular portion of FGFRs binds to fibroblast growth factors
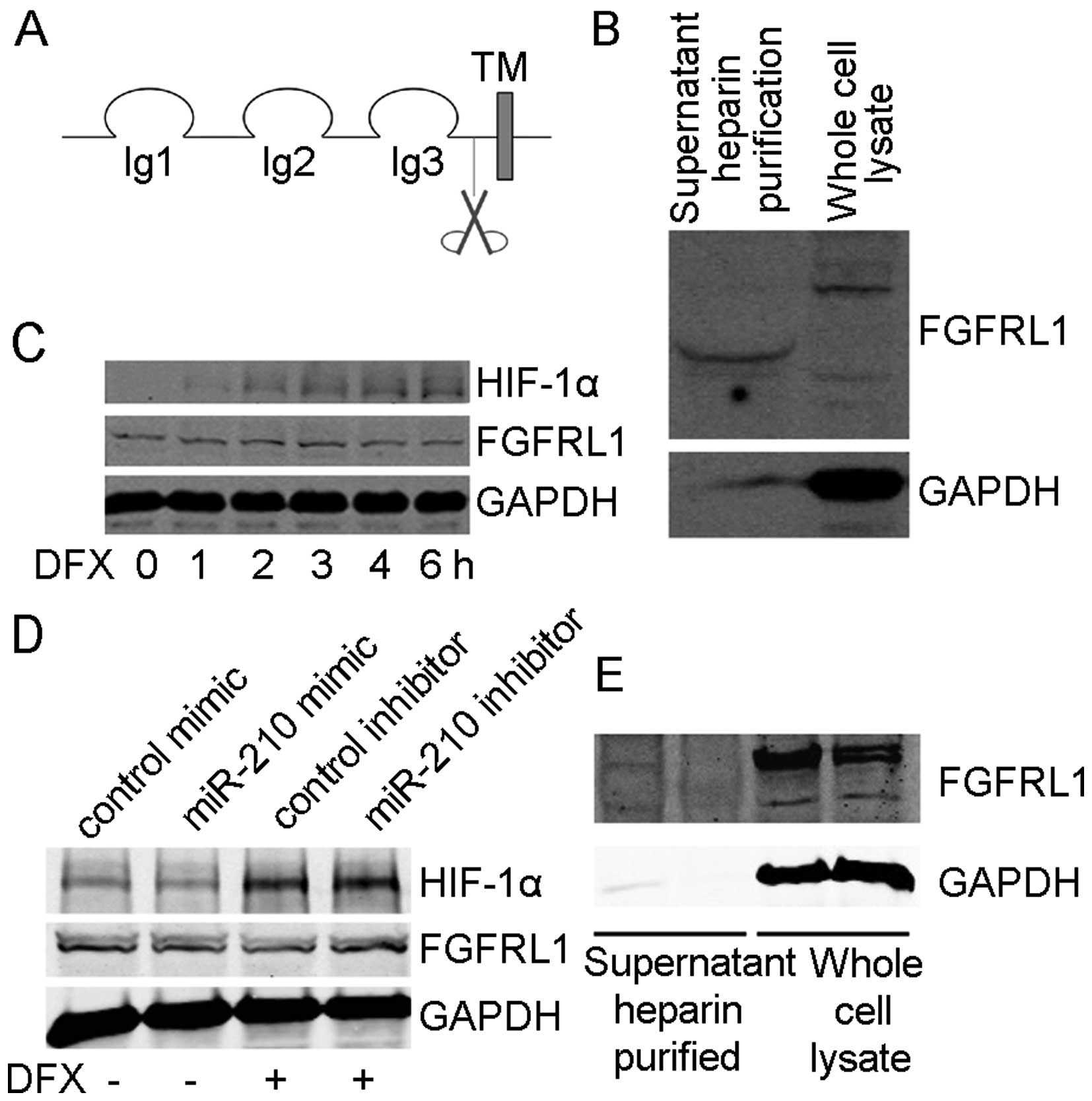
(FGFs). However, as a decoy FGF receptor (44), FGFRL1 lacks the cytoplasmic tyrosine
kinase domain (Fig. 3A).
It has been reported that FGFRL1 can be shed from
the cell membrane and secreted in myoblasts and HEK293 cells
(40), but it is unclear whether
hepatocytes can secrete FGFRL1. We established an FGFRL1 stably
transduced SMMC-7721 cell line to evaluate the shedding of FGFRL1
from hepatocytes. As previously mentioned, FGFRL1 binds to heparin
(40), therefore, the cell culture
supernatant was subjected to heparin purification and then was
analyzed by WB, while the whole cell lysate was served as a control
(Fig. 3B). We obtained a WB signal
for shed FGFRL1 in the cell culture supernatant, with a lower
molecular weight due to loss of transmembrane and intracellular
domains. This finding showed that FGFRL1 is released from the cell
membrane in a soluble form, suggesting that FGFRL1 acts as a
secreted regulator of FGF signaling.
Next, we determined whether hypoxia treatment
changed the expression of FGFRL1. SMMC-7721 cells were treated with
DFX for different time periods. HIF1α expression was induced while
the expression of FGFRL1 declined (Fig.
3C). To assess whether the decrease of FGFRL1 following hypoxia
was miR-210-dependent, we transiently transfected SMMC-7721 cells
with miR-210 mimic, miR-210 inhibitor and their respective
controls. The results showed that compared with mimic-control
transfected cells, miR-210 mimic transfected cells had lower
expression of FGFRL1, which is similar to the decrease seen in
DFX-treated cells (Fig. 3D).
miR-210 inhibitor transfected cells showed no decrease of FGFRL1
when treated with DFX (Fig. 3D).
These results indicated that hypoxia-induced downregulation of
FGFRL1 was mainly through miR-210. We further assessed whether
expression of miR-210 downregulated FGFRL1 secretion. The heparin
extract of the supernatant and whole cell lysate of miR-210 stably
transfected and control cells were examined by WB. The results
showed that miR-210 expression downregulated FGFRL1 both inside and
outside the cells (Fig. 3E). Hence,
we conclude that hypoxia-induced miR-210 targets FGFRL1 expression
and secretion in HCC.
FGFRL1 is a negative regulator of HCC
angiogenesis
Although there are no reports on the angiogenesis
functions of FGFRL1, we propose that FGFRL1 may inhibit
angiogenesis by blocking FGF2 (45). To prove our hypothesis, we used a
commercially available recombinant FGFRL1 protein in an in
vitro angiogenesis assay. An in vitro HUVEC tube
formation assay was performed to evaluate the effect of FGFRL1 on
the anti-angiogenic response. HUVEC cells were seeded on Matrigel
support using different stimulations: complete EGM2 medium without
FGF2 (EGM2 - FGF2), complete EGM2 medium (EGM2), complete EGM2
medium with FGFRL1 (EGM2 + FGFRL1), and complete EGM2 medium with
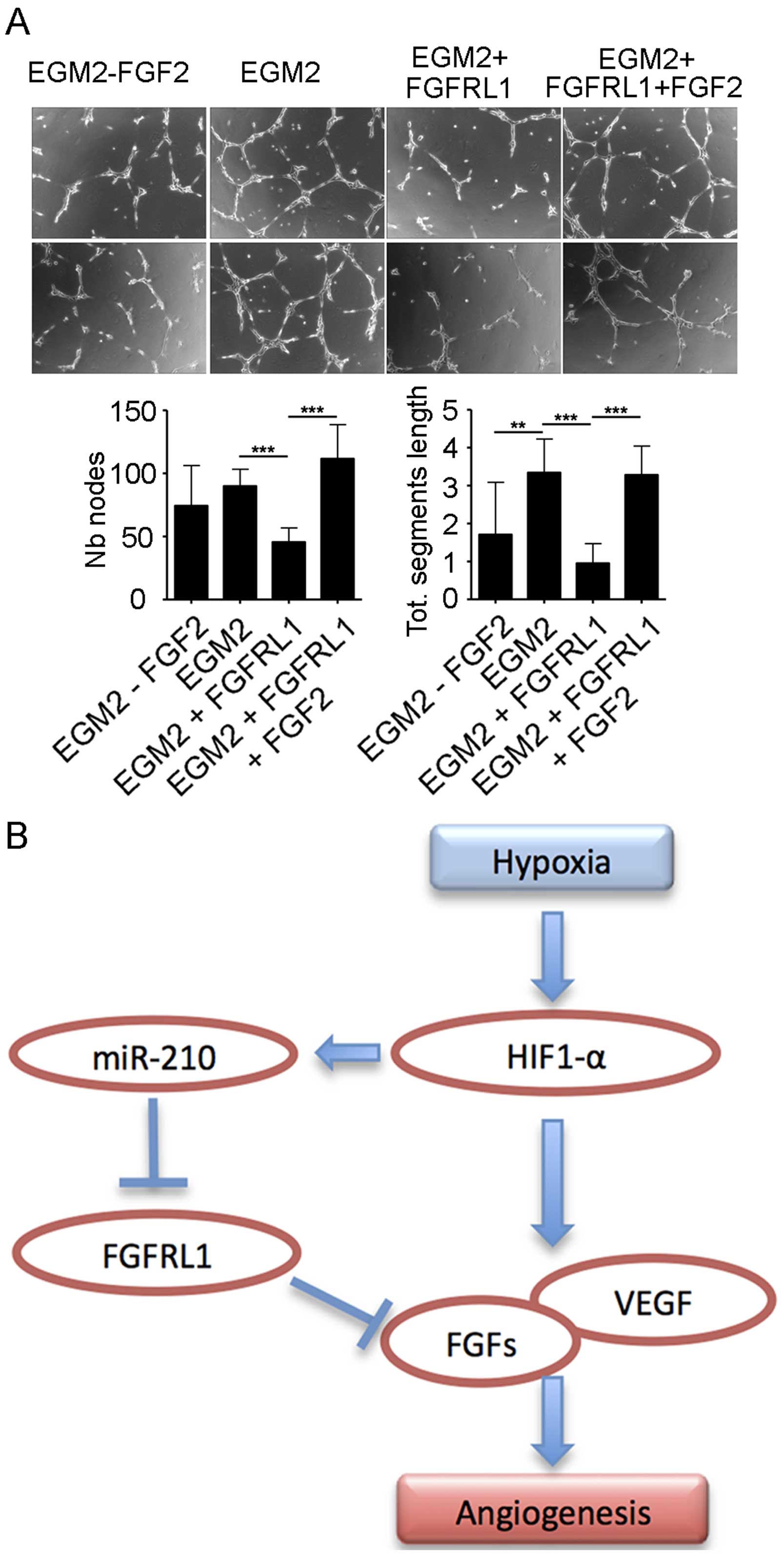
FGFRL1 plus FGF2 (EGM2 + FGFRL1 + FGF2). As shown in Fig. 4A, EGM2 + FGFRL1 significantly
inhibited in vitro tube formation as compared to EGM2, while
additional FGF2 rescued inhibition shown in the EGM2 + FGFRL1 +
FGF2 group. Thus, FGFRL1 is a negative regulator of HCC
angiogenesis, and hypoxia-induced miR-210 promotes HCC angiogenesis
by targeting FGFRL1 expression and secretion.
Based on our findings, we propose the following
working model to explain how induced miR-210 expression promotes
HCC angiogenesis. Hypoxia induced the transcription factor HIF1α
expression and activation in HCC cells. HIF1α then promotes the
secretion of FGFs and VEGF to enhance HCC angiogenesis.
Alternatively, HIF1α induced the expression of hypoxic miR-210 to
target FGFRL1 expression and secretion. As FGFRL1 is a decoy FGF
receptor and an inhibitor of HCC angiogenesis, hypoxia-induced
miR-210 promotes HCC angiogenesis by targeting FGFRL1 expression
(Fig. 4B).
Discussion
In the present study, we demonstrated that miR-210
expression level was significantly upregulated in HCC tumor
tissues, as compared to that of non-tumor liver tissues. In
addition, miR-210 expression level altered with different stages of
liver disease, and was correlated with prognosis of HCC patients.
We also introduced a new working model of miR-210 promoted HCC
angiogenesis through the targeting of FGFRL1. These data suggested
that miR-210 may be an important regulator in HCC carcinogenesis
and progression.
The relative expression level of miR-210 T/nT value
was determined to be a predictor of HCC prognosis. Previous studies
suggested that miR-210 was not a specific HCC diagnostic biomarker
(46), however, our data showed
that miR-210 could be a predictor of HCC recurrence and final
outcome, although more evidence from multicenter clinical studies
are required to confirm these findings. Tumors with high expression
of miR-210 showed more aggressive phenotypes, and the expression of
miR-210 was not always higher in tumor tissues. As miR-210 T/nT
value classifies the prognosis of HCC patients more precisely, the
regulation of miR-210 expression in HCC cells may be more
complicated under hypoxic conditions. For example, the epigenetic
regulation of miR-210 expression may be interesting, considering
that the high GC content in the promoter region of miR-210
(43).
The induction of angiogenesis is a hallmark of
cancer (3). Hypoxia regulates
angiogenesis through different mechanisms (47), and miR-210 plays an important role
in these mechanisms. It was reported that increased miR-210 in
endothelial cells induced angiogenesis through activation of Notch
signaling (19), and miR-210 could
be exported out of the cell through exosome mediated mechanism to
enhance angiogenesis (22,23). In the present study, we describe a
new mechanism for enhanced angiogenesis by miR-210 through
downregulation of FGFRL1. As miRNAs can target multiple mRNAs, it
is not surprising to find several angiogenesis associated
mechanisms which miR-210 may use (15–24).
Possible synchronization of different pro-angiogenesis mechanisms
of miR-210 could be an economic way for cells to deal with hypoxic
stress. Considering the other oncogenic effects of miR-210, those
tumor cells with better vascular formation and a high expression
level of miR-210 adapt better.
It is proposed that miR-210 can act both as an
oncogene and a tumor suppressor, depending on specific cellular
conditions (30). In general, our
data suggest that miR-210 acts as an oncogene in the latter stages,
at least in HCC. The genetic events which occurred in HCC cells
during hypoxia adaptation are essential in the functional switch of
miR-210. Those HCC cells which overcome hypoxic stress and maintain
expression of miR-210 are more aggressive. In summary, the present
study show higher miR-210 expression in HCC, which is correlated
with poor prognosis of HCC patients, and hypoxic miR-210 promotes
HCC angiogenesis.
Acknowledgments
We thank Dr Qiang Deng and Dr Yuan Gao from the
Institute Pasteur of Shanghai for extensive assistance in our
research projects, and Zhide Zhang for the excellent technical
assistance. The present study was supported by the founding of
National Key Basic Research Program of China (2014CB542102), the
National High Technology Research and Development Program of China
(2013AA032202), the Science Fund for Creative Research Groups,
NSFC, China (81221061 and 81372207), the National Natural Science
Foundation of China (81502416 and 81572791), the Innovation Program
of Shanghai Municipal Education Commission (11ZZ76), and the
Shanghai Municipal Commission of Health and Family Planning Project
(140822111537488).
References
|
1
|
El-Serag HB: Hepatocellular carcinoma. N
Engl J Med. 365:1118–1127. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Ott JJ, Stevens GA, Groeger J and Wiersma
ST: Global epidemiology of hepatitis B virus infection: New
estimates of age-specific HBsAg seroprevalence and endemicity.
Vaccine. 30:2212–2219. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Hanahan D and Weinberg RA: Hallmarks of
cancer: The next generation. Cell. 144:646–674. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Carmeliet P and Jain RK: Angiogenesis in
cancer and other diseases. Nature. 407:249–257. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Chan YC, Banerjee J, Choi SY and Sen CK:
miR-210: The master hypoxamir. Microcirculation. 19:215–223. 2012.
View Article : Google Scholar :
|
|
6
|
Giannakakis A, Sandaltzopoulos R, Greshock
J, Liang S, Huang J, Hasegawa K, Li C, O'Brien-Jenkins A, Katsaros
D, Weber BL, et al: miR-210 links hypoxia with cell cycle
regulation and is deleted in human epithelial ovarian cancer.
Cancer Biol Ther. 7:255–264. 2008. View Article : Google Scholar
|
|
7
|
Zhang Z, Sun H, Dai H, Walsh RM, Imakura
M, Schelter J, Burchard J, Dai X, Chang AN, Diaz RL, et al:
MicroRNA miR-210 modulates cellular response to hypoxia through the
MYC antagonist MNT. Cell Cycle. 8:2756–2768. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Tsuchiya S, Fujiwara T, Sato F, Shimada Y,
Tanaka E, Sakai Y, Shimizu K and Tsujimoto G: MicroRNA-210
regulates cancer cell proliferation through targeting fibroblast
growth factor receptor-like 1 (FGFRL1). J Biol Chem. 286:420–428.
2011. View Article : Google Scholar :
|
|
9
|
Biswas S, Roy S, Banerjee J, Hussain SR,
Khanna S, Meenakshisundaram G, Kuppusamy P, Friedman A and Sen CK:
Hypoxia inducible microRNA 210 attenuates keratinocyte
proliferation and impairs closure in a murine model of ischemic
wounds. Proc Natl Acad Sci USA. 107:6976–6981. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Kim HW, Haider HK, Jiang S and Ashraf M:
Ischemic preconditioning augments survival of stem cells via
miR-210 expression by targeting caspase-8-associated protein 2. J
Biol Chem. 284:33161–33168. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Hu S, Huang M, Li Z, Jia F, Ghosh Z,
Lijkwan MA, Fasanaro P, Sun N, Wang X, Martelli F, et al:
MicroRNA-210 as a novel therapy for treatment of ischemic heart
disease. Circulation. 122(Suppl 11): S124–S131. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Crosby ME, Kulshreshtha R, Ivan M and
Glazer PM: MicroRNA regulation of DNA repair gene expression in
hypoxic stress. Cancer Res. 69:1221–1229. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Cicchillitti L, Di Stefano V, Isaia E,
Crimaldi L, Fasanaro P, Ambrosino V, Antonini A, Capogrossi MC,
Gaetano C, Piaggio G, et al: Hypoxia-inducible factor 1-α induces
miR-210 in normoxic differentiating myoblasts. J Biol Chem.
287:44761–44771. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Gong Y, Xu F, Zhang L, Qian Y, Chen J,
Huang H and Yu Y: MicroRNA expression signature for Satb2-induced
osteogenic differentiation in bone marrow stromal cells. Mol Cell
Biochem. 387:227–239. 2014. View Article : Google Scholar
|
|
15
|
Pulkkinen K, Malm T, Turunen M, Koistinaho
J and Ylä-Herttuala S: Hypoxia induces microRNA miR-210 in vitro
and in vivo ephrin-A3 and neuronal pentraxin 1 are potentially
regulated by miR-210. FEBS Lett. 582:2397–2401. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Alaiti MA, Ishikawa M, Masuda H, Simon DI,
Jain MK, Asahara T and Costa MA: Up-regulation of miR-210 by
vascular endothelial growth factor in ex vivo expanded
CD34+ cells enhances cell-mediated angiogenesis. J Cell
Mol Med. 16:2413–2421. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Donnem T, Fenton CG, Lonvik K, Berg T,
Eklo K, Andersen S, Stenvold H, Al-Shibli K, Al-Saad S, Bremnes RM,
et al: MicroRNA signatures in tumor tissue related to angiogenesis
in non-small cell lung cancer. PLoS One. 7:e296712012. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Liu F, Lou YL, Wu J, Ruan QF, Xie A, Guo
F, Cui SP, Deng ZF and Wang Y: Upregulation of microRNA-210
regulates renal angiogenesis mediated by activation of VEGF
signaling pathway under ischemia/perfusion injury in vivo and in
vitro. Kidney Blood Press Res. 35:182–191. 2012. View Article : Google Scholar
|
|
19
|
Lou YL, Guo F, Liu F, Gao FL, Zhang PQ,
Niu X, Guo SC, Yin JH, Wang Y and Deng ZF: miR-210 activates notch
signaling pathway in angiogenesis induced by cerebral ischemia. Mol
Cell Biochem. 370:45–51. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Shoji T, Nakasa T, Yamasaki K, Kodama A,
Miyaki S, Niimoto T, Okuhara A, Kamei N, Adachi N and Ochi M: The
effect of intra-articular injection of microRNA-210 on ligament
healing in a rat model. Am J Sports Med. 40:2470–2478. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Yamasaki K, Nakasa T, Miyaki S, Yamasaki
T, Yasunaga Y and Ochi M: Angiogenic microRNA-210 is present in
cells surrounding osteonecrosis. J Orthop Res. 30:1263–1270. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Kosaka N, Iguchi H, Hagiwara K, Yoshioka
Y, Takeshita F and Ochiya T: Neutral sphingomyelinase 2
(nSMase2)-dependent exosomal transfer of angiogenic microRNAs
regulate cancer cell metastasis. J Biol Chem. 288:10849–10859.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Tadokoro H, Umezu T, Ohyashiki K, Hirano T
and Ohyashiki JH: Exosomes derived from hypoxic leukemia cells
enhance tube formation in endothelial cells. J Biol Chem.
288:34343–34351. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Zeng L, He X, Wang Y, Tang Y, Zheng C, Cai
H, Liu J, Wang Y, Fu Y and Yang GY: MicroRNA-210 overexpression
induces angiogenesis and neurogenesis in the normal adult mouse
brain. Gene Ther. 21:37–43. 2014. View Article : Google Scholar
|
|
25
|
Chan SY, Zhang YY, Hemann C, Mahoney CE,
Zweier JL and Loscalzo J: MicroRNA-210 controls mitochondrial
metabolism during hypoxia by repressing the iron-sulfur cluster
assembly proteins ISCU1/2. Cell Metab. 10:273–284. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Chen Z, Li Y, Zhang H, Huang P and Luthra
R: Hypoxia-regulated microRNA-210 modulates mitochondrial function
and decreases ISCU and COX10 expression. Oncogene. 29:4362–4368.
2010. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Puisségur MP, Mazure NM, Bertero T,
Pradelli L, Grosso S, Robbe-Sermesant K, Maurin T, Lebrigand K,
Cardinaud B, Hofman V, et al: miR-210 is overexpressed in late
stages of lung cancer and mediates mitochondrial alterations
associated with modulation of HIF-1 activity. Cell Death Differ.
18:465–478. 2011. View Article : Google Scholar :
|
|
28
|
Ying Q, Liang L, Guo W, Zha R, Tian Q,
Huang S, Yao J, Ding J, Bao M, Ge C, et al: Hypoxia-inducible
microRNA-210 augments the metastatic potential of tumor cells by
targeting vacuole membrane protein 1 in hepatocellular carcinoma.
Hepatology. 54:2064–2075. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Hong L, Han Y, Zhang H, Zhao Q and Qiao Y:
miR-210: A therapeutic target in cancer. Expert Opin Ther Targets.
17:21–28. 2013. View Article : Google Scholar
|
|
30
|
Qin Q, Furong W and Baosheng L: Multiple
functions of hypoxia-regulated miR-210 in cancer. J Exp Clin Cancer
Res. 33:502014. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Leung AK and Sharp PA: MicroRNA functions
in stress responses. Mol Cell. 40:205–215. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Esquela-Kerscher A and Slack FJ: Oncomirs
- microRNAs with a role in cancer. Nat Rev Cancer. 6:259–269. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Haybaeck J, Zeller N and Heikenwalder M:
The parallel universe: microRNAs and their role in chronic
hepatitis, liver tissue damage and hepatocarcinogenesis. Swiss Med
Wkly. 141:w132872011.PubMed/NCBI
|
|
34
|
Meng F, Henson R, Wehbe-Janek H, Ghoshal
K, Jacob ST and Patel T: MicroRNA-21 regulates expression of the
PTEN tumor suppressor gene in human hepatocellular cancer.
Gastroenterology. 133:647–658. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Hou J, Lin L, Zhou W, Wang Z, Ding G, Dong
Q, Qin L, Wu X, Zheng Y, Yang Y, et al: Identification of miRNomes
in human liver and hepatocellular carcinoma reveals miR-199a/b-3p
as therapeutic target for hepatocellular carcinoma. Cancer Cell.
19:232–243. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Pineau P, Volinia S, McJunkin K, Marchio
A, Battiston C, Terris B, Mazzaferro V, Lowe SW, Croce CM and
Dejean A: miR-221 overexpression contributes to liver
tumorigenesis. Proc Natl Acad Sci USA. 107:264–269. 2010.
View Article : Google Scholar :
|
|
37
|
Zhan M, Li Y, Hu B, He X, Huang J, Zhao Y,
Fu S and Lu L: Serum microRNA-210 as a predictive biomarker for
treatment response and prognosis in patients with hepatocellular
carcinoma undergoing transarterial chemoembolization. J Vasc Interv
Radiol. 25:1279–1287.e1. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Yang W, Sun T, Cao J, Liu F, Tian Y and
Zhu W: Downregulation of miR-210 expression inhibits proliferation,
induces apoptosis and enhances radiosensitivity in hypoxic human
hepatoma cells in vitro. Exp Cell Res. 318:944–954. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Yang W, Wei J, Sun T and Liu F: Effects of
knockdown of miR-210 in combination with ionizing radiation on
human hepatoma xenograft in nude mice. Radiat Oncol. 8:1022013.
View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Steinberg F, Zhuang L, Beyeler M, Kälin
RE, Mullis PE, Brändli AW and Trueb B: The FGFRL1 receptor is shed
from cell membranes, binds fibroblast growth factors (FGFs), and
antagonizes FGF signaling in Xenopus embryos. J Biol Chem.
285:2193–2202. 2010. View Article : Google Scholar :
|
|
41
|
Weidner N, Semple JP, Welch WR and Folkman
J: Tumor angio-genesis and metastasis - correlation in invasive
breast carcinoma. N Engl J Med. 324:1–8. 1991. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Albanis E and Friedman SL: Hepatic
fibrosis. Pathogenesis and principles of therapy. Clin Liver Dis.
5:315–334. v–vi. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Huang X, Ding L, Bennewith KL, Tong RT,
Welford SM, Ang KK, Story M, Le QT and Giaccia AJ:
Hypoxia-inducible mir-210 regulates normoxic gene expression
involved in tumor initiation. Mol Cell. 35:856–867. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Bertrand S, Somorjai I, Garcia-Fernandez
J, Lamonerie T and Escriva H: FGFRL1 is a neglected putative actor
of the FGF signalling pathway present in all major metazoan phyla.
BMC Evol Biol. 9:2262009. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Trueb B, Zhuang L, Taeschler S and
Wiedemann M: Characterization of FGFRL1, a novel fibroblast growth
factor (FGF) receptor preferentially expressed in skeletal tissues.
J Biol Chem. 278:33857–33865. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Lu J, Xie F, Geng L, Shen W, Sui C and
Yang J: Potential role of microRNA-210 as biomarker in human
cancers detection: A meta-analysis. Biomed Res Int.
2015:3039872015. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Pugh CW and Ratcliffe PJ: Regulation of
angiogenesis by hypoxia: Role of the HIF system. Nat Med.
9:677–684. 2003. View Article : Google Scholar : PubMed/NCBI
|