Introduction
Multiple myeloma (MM) is a malignant hematological
tumor with monoclonal plasma cell hyperplasia. The incidence of MM
in European and American countries has already surpassed acute
leukemia and accounts for 12% of all hematologic malignancies
(1). Currently, the major clinical
treatment strategy for MM is chemotherapy (2). However, effective chemotherapeutic
drugs are still limited by the development of drug resistance
mechanisms. Therefore, the effects of chemotherapy are greatly
reduced (3). Bortezomib has
achieved great progress in MM treatment. As a proteasome inhibitor,
bortezomib inhibits the tumor activity of multiple myeloma cells
(MMCs) through various routes (4–7). As
the first-line drug in MM chemotherapy, bortezomib has significant
inhibitory functions in the biological progression and pathological
progression of myeloma. However, secondary drug resistance
accompanied by chemotherapy usually results in the poor
chemotherapeutic effects of bortezomib and poor prognosis in MM
(8–11). Thus, studies related to MM drug
resistance mechanism are a research ‘hotspot’ in the field of MM
chemotherapy.
Firstly, we found that the expression of Snail
family transcriptional repressor 1 (Snail1) was significantly
increased in MMCs from bortezomib-resistant MM patients. In this
study, an in vitro resistance model was constructed to
investigate the reasons for the abnormal expression of Snail1
during the development of bortezomib resistance and its
contribution to resistance. Data analyses showed that Snail1 had a
positive association and a negative association with the expression
of the MDR1 gene and P53 protein, respectively (12,13).
Subsequent studies concerning the mechanisms were based on the
following three confirmed facts. The Snail1 nuclear transcription
factor directly binds to the multi-drug resistance gene (MDR1)
promoter to positively regulate gene transcription. Snail1 binds to
the hsa-miRNA-22-3p precursor promoter to positively regulate its
transcription. Finally, hsa-miRNA-22-3p binds to the 3′UTR region
of the tumor protein p53 gene (P53) to negatively regulate its
protein expression. Pathway-specific analysis using Snail1 gene
silencing confirmed that the Snail1/MDR1 and
Snail1/hsa-miRNA-22-3p/P53 pathways dominated the reversal of drug
resistance in bortezomib-resistant MMCs. For the first time, this
study confirmed the critical role of the Snail1 nuclear
transcription factor in the regulation of the drug resistance
mechanisms in bortezomib-resistant MMCs and preliminarily
elucidated the pathways of the decreased drug resistance of MMCs to
bortezomib by Snail1 silencing. Research data showed that Snail1
affects the development of bortezomib drug resistance mechanisms in
MMCs through regulation of the expression of MDR1 and P53.
In addition, to the best of our knowledge, this was
the first study on the association between hsa-miRNA-22-3p and the
drug resistance mechanisms in MMCs. Based on the mechanistic study,
we aimed to improve the effect of bortezomib on MM by silencing
Snail1. In the long run, the study provides support for elucidating
the development of resistance of MM, and new insight on possible
solutions. For the clinical treatment of MM, this study may guide
the development of novel combination therapy including gene
intervention combined chemotherapeutics to minimize the resistance
of MM to bortezomib.
Materials and methods
Cell culture
The human multiple myeloma cell lines, XG-7 and
RPMI-8226, were purchased from the Cell Bank of the Chinese Academy
of Sciences (CBCS; Shanghai, China). The establishment of
bortezomib-resistant cell lines, XG-7/Bor and RPMI-8226/Bor, was
performed using the gradient induction method by our study group.
Drug-resistant cells were cultured in Invitrogen™ RPMI-1640 culture
medium (Thermo Fisher Scientific, Inc., Waltham, MA, USA)
containing 10% fetal bovine serum (FBS; Invitrogen; Thermo Fisher
Scientific, Inc.) and 10 nM bortezomib (Sigma-Aldrich; Merck KGaA,
Darmstadt, Germany) under culture conditions of 37°C and 5%
CO2. Cells showed semi-suspension growth, and cell
passage was performed using the centrifugation method. In addition,
293T cells were purchased from the American Type Culture Collection
(ATCC, Manassas, VA, USA) and were used as the tool and virus
package cell line, and they were cultured in high-glucose DMEM
culture medium (Invitrogen; Thermo Fisher Scientific, Inc.)
containing 10% FBS under culture conditions of 37°C and 5%
CO2. These cells showed adherent growth and were
passaged using trypsin digestion (Invitrogen; Thermo Fisher
Scientific, Inc.) method when cell confluence reached 70%.
Collection of MMCs from MM patients
with bortezomib-resistance and detection of related indictors
Primary MMCs were collected before and after
development of bortezomib resistance from 20 MM patients treated at
the First Affiliated Hospital of Anhui Medical University, China
(from January 2016 to December 2017). Eligible patients exhibited
refractory/relapse to their therapy after at least two cycles of
bortezomib (administered in combination with other agents), defined
as no response (less than partial response) or progression either
during therapy or ≤60 days after completion of therapy. The
baseline characteristics of the patients at diagnosis are shown in
Table I. Informed consent was
provided by all patients in writing and the study was approved by
the Medical Ethics Committee of the First Affiliated Hospital of
Anhui Medical University. MMCs were obtained by
fluorescence-activated cell sorting for
CD38+CD45+ mononuclear cells, about
1×107 cells from each sample, collected by
centrifugation after phosphate-buffered saline (PBS) resuspension,
placed in cell cryopreservation tubes, labeled and stored in liquid
nitrogen. Total RNA and protein were extracted, and the relative
levels of Snail1, hsa-miRNA-22-3p, MDR1, and P53 were detected
using real-time PCR, and the protein expression levels of Snail1,
P-gp (protein encoded by the MDR1 gene) and P53 were detected using
western blot analysis.
 | Table I.Baseline characteristics of the MM
patients (N=20) at diagnosis. |
Table I.
Baseline characteristics of the MM
patients (N=20) at diagnosis.
|
Characteristics | Data |
|---|
| Median age (range),
in years | 58.2 (35–76) |
| Sex |
|
| Male, n
(%) | 11 (55) |
| Female,
n (%) | 9 (45) |
| Serum heavy chain
at diagnosis, n (%) |
|
|
None | 3 (15) |
|
IgG | 11 (55) |
|
IgA | 5 (25) |
|
IgD | 1 (5) |
| ECOG score at
diagnosis, n (%) |
|
| 0 | 1 (5) |
| 1 | 3 (15) |
| 2 | 6 (30) |
| 3 | 7 (35) |
| 4 | 2 (10) |
| 5 | 1 (5) |
| ISS stage at
diagnosis, n (%) |
|
| Stage
I | 4 (20) |
| Stage
II | 6 (30) |
| Stage
III | 10 (50) |
| FISH at
diagnosis |
|
|
del17p | 4 (20) |
|
1q21 | 3 (15) |
|
t(4;14) | 1 (10) |
Construction of vectors and RNA
synthesis
Construction of cDNA expression vector. The coding
sequence (CDS) of human MDR1 (NM_001348945.1) was amplified by
using the primers 5′-GGAATTCGCCACCATGAGTGTCAACTTGCAA-3′ and
5′-CGGGATCCTCACTGGCGCTTTGTTCCAG-3′, which contain an EcoRI
cutting site and kozak sequence and a BamHI cutting site,
respectively, with the cDNA prepared by reverse transcription of
RNA isolated from XG-7/Bor cells. The PCR product was digested and
cloned into the pcDH1 lentiviral expressing vector (System
Biosciences, Palo Alto, CA, USA); the recombinant vector was named
pcDH1-MDR1.
Construction of the shRNA vector
A siRNA sequence complementarily binding to human
Snail1 (NM_005985.3) was chosen. The target sequences of siRNA
(5′-GCTCTGTGGCATTCACGCA-3′) were homologous to Snail1,
respectively. The oligonucleotide templates of these shRNAs were
chemically synthesized and cloned into the linear pshRNA-copGFP
shRNA vector (System Biosciences) which was obtained through
digestion by BamHI and EcoRI (Takara Biotechnology
Co., Ltd., Dalian, China) and purification by agarose gel
electrophoresis. An invalid siRNA sequence
(5′-TTGCAGCCAGTGGCTTACC-3′) was used as a negative control (NC).
Sequencing was used to confirm the vectors constructed
(pshRNA-Snail1 and pshRNA-NC), and pshRNA-P53 for the human P53
gene (NM_000546.5) was constructed by the same method (siRNA,
5′-GACTCCAGTGGTAATCTAC-3′).
Construction of the luciferase
reporter vector
The 3′-untranslated region (3′UTR, 212 bp) of human
P53 was amplified from cDNA obtained through the reverse
transcription of total RNA of XG-7/Bor cells, with the following
primers: 5′-GCTCTAGAGGCATTTGCACCTACCTC-3′ and
5′-GCTCTAGATTGGCTGGGCCAGCAGAGAC-3′. The amplification parameters
were as follows: 32 cycles of denaturation at 95°C for 10 sec,
annealing at 58°C for 30 sec and extension at 72°C for 30 sec. The
product was digested with XbaI and inserted into the
pGL3-promotor vector (Promega, Madison, WI, USA). TargetScan
(http://www.targetscan.org/vert_71/)
was used to predict the theoretic target (seed region) of
hsa-miRNA-22-3p in the 3′UTR of the P53 mRNA sequence. The seed
region was mutated from 5′-GGCAGCT-3′ to 5′-TCGAGCG-3 by point
mutation. The resultant vectors were termed pGL3-wt- P53 and
pGL3-mt-P53 respectively.
The promoters of MDR1 and hsa-miRNA-22-3p were
amplified by using human genomic DNA as a template and cloned into
a luciferase reporter vector pGL3-Enhancer (Promega) upstream of
the luciferase gene. We constructed the wild-type transcription
factor binding site (TFBS) vectors, pGL3-wt-pro-P53 and
pGL3-wt-pro-miRNA-22. We then constructed the mutant-type TFBS
vectors pGL3-mt-pro-P53 and pGL3-mt-pro-miRNA-22 through the point
mutation, 5′-CCGCTG-3′ to 5′-CAGGCT-3′. The products of the vectors
were confirmed by DNA sequencing. Endotoxin-free DNA was prepared
in all cases.
RNA synthesis
Chemically synthesized hsa-miRNA-22-3p mimics
(5′-AAGCUGCC AGUUGAAGAACUGUtt-3′), inhibitor
(5′-ACAGUUCUUCAACUGGCAGCUUtt-3′) and NC
(5′-CGAAUACAGAAAGGCGUUCUGUtt-3′) were obtained from Sangon
(Shanghai, China), ‘tt’ protective bases being added to the RNA
end.
Packaging of the recombinant lentivirus and
infection of XG-7/Bor and RPMI-8226/Bor cells with the lentivirus.
One day before transfection, 293T cells were seeded into 10-cm
dishes. Two micrograms of each vector (pshRNA-Snail1 or pcDH1-MDR1
or pshRNA-P53) and 10 µg pPACK Packaging Plasmid Mix (System
Biosciences) were co-transfected using Lipofectamine 2000
(Invitrogen; Thermo Fisher Scientific, Inc.) in accordance with the
manufacturer's protocol. The medium was replaced with DMEM plus 1%
FBS. Forty-eight hours later, the supernatant was harvested and
then cleared by centrifugation at 5000 × g at 4°C for 5 min and
passed through a 0.45-µm polyvinylidene fluoride (PVDF) membrane
(Millipore). The titer of the virus was determined by gradient
dilution. The packaged lentiviruses were named as Lv-shRNA-Snail1
or Lv-MDR1 or Lv-shRNA-P53.
One day before viral infection, XG-7/Bor and
RPMI-8226/Bor cells in the logarithmic phase were made into a
suspension, and the number of viable cells was counted by using
trypan blue staining. Cells were collected by centrifugation at
1000 × g and re-suspended in RPMI-1640 medium (10% FBS and 10 nM
bortezomib) at a concentration of 5×105 cells/ml. Cells
were seeded on 6-well plates, 2 ml/well, and cultured overnight
under normal conditions. One day after seeding, the cells were
infected with lentiviruses diluted with dPBS at an MOI
(multiplicity of infection) of 20, and the medium was refreshed
after 24 h. The infection efficiency was assessed by fluorescence
microscopy 72 h after infection. Total protein was extracted from
the cells using the M-PER Mammalian Protein Extraction Reagent kit
(Thermo Fisher Scientific, Inc.) and subjected to western blotting
to measure the levels of Snail1, P-gp and P53.
Luciferase assay
Verification of the binding site of
hsa-miRNA-22-3p on P53 3′UTR
293T cells were transfected with the
hsa-miRNA-22-3p-mimics or inhibitor or NC, and pGL-wt-P53 or
pGL-mt-P53 using Lipofectamine 3000 (Invitrogen; Thermo Fisher
Scientific, Inc.) according to the manufacturer's instructions.
Forty-eight hours after transient transfection, the cells were
harvested, and luciferase assays were performed.
Verification of the binding site of
the transcription factor in MDR1 and the pri-miRNA-22 promoter
293T cells were co-transfected with the wild-type or
mutant promoter reporter vectors and pshRNA-snail1, and harvested
for luciferase activity assay 48 h later. The transfection
experiment was carried out in 24-well plates, following the
instructions for Lipofectamine 3000. PGL-TK (100 ng) was
transfected for each well as the internal reference for the
luciferase assay. The relative luciferase activities (ratios of
firefly and Renilla luciferase activity) of the lysates were
measured by dual luciferase reporter assay system (Promega).
Assessment of cell viability and
IC50 values
The two genetically engineered cell lines were
seeded in 96-well plates at 5×104 cells/well, and bortezomib was
added to a final concentration of 1, 2, 4, 8, 16 or 32 nM for a
period of 48 h, followed by a CCK-8 (Cell Counting Kit-8) assay for
cell viability. Briefly, 10 µl CCK-8 solution was added, and the
cells were cultured under normal conditions for an additional 4 h
before measuring absorbance at 450 nm. The cell inhibition ratio
was calculated, based on the IC50 values at 48 h.
Detection of apoptosis
Seventy-two hours after viral infection, XG-7/Bor
cells were seeded in 6-well plates at 1×105 cells per
well in medium containing 10 nM bortezomib and were cultured for 48
h. The cells were collected and measured for apoptosis using flow
cytometry (FACS Calibur; BD Biosciences, Franklin Lakes, NJ, USA)
after treatment using Annexin V:FITC Apoptosis Detection Kit II
(cat. no. 556570; BD Biosciences). Cells were made into suspensions
by trypsinization and the cells were washed with dPBS and suspended
in 500 µl binding buffer and 5 µl Annexin V-FITC was added and the
cells were maintained in the dark for 10 min. Cells were then
stained with 5 µl propidium iodide (PI) for 5 min. Apoptosis was
analyzed on BD-FACSCalibur using FITC (FL1) channel and PI (FL2)
channel at an excitation wavelength at 488 nm.
Real-time PCR
Total RNA was isolated with TRIzol reagent
(Invitrogen; Thermo Fisher Scientific, Inc.) according to the
manufacturer's instructions and was reversely transcribed into cDNA
using M-MLV Reverse Transcriptase and oligo(dT)18 primer (Takara).
The following specific primers were used for the quantitative PCR
of human Snail1, MDR1, P53 and β-actin: Snail1,
5′-ATGCCGCGCTCTTTGCTC-3′ and 5′-GGTGGGCCTGGTCGTAG-3′; MDR1,
5′-CCGTGGGGCAAGTCAGTTCA-3′ and 5′-CCGGTCGGGTGGGATAGTTG-3′; P53,
5′-CGTACTCCCCTGCCCTCAACAAGA-3′ and 5′-GCAGCGCCTCACAACCTCCGTCAT-3′;
and β-actin, 5′-CCTGTACGCCAACACAGTGC-3′ and
5′-ATACTCCTGCTTGCTGATCC-3′. The lengths of the amplified products
were 118, 186, 155 and 211 bp, respectively. Real-time PCR was
performed using SYBR Premix Ex Taq™ kit and TP800 System (Takara
Biotechnology). cDNA from 200 ng total RNA was used as the
template. The PCR reactions were carried out under the following
conditions: 40 cycles of denaturation at 95°C for 10 sec, annealing
at 60°C for 20 sec and extension at 72°C for 20 sec. The mRNA
levels of twist1 were normalized using the ∆∆Ct method, to the
expression of an endogenous housekeeping gene, β-actin. The
specific primers for hsa-miRNA-22-3p detection, U6 snRNA
(NM_001101. 3), 5′-TACCTTGCGAAGTGCTTAAAC-3′ and hsa-miRNA22-3p,
5′-GTCGTATCCAGTGCGTGTCGTGGAGTCGGCAATTGCACTGGATACGAACAGT-3′ were
used for cDNA preparation. The following primers were used for
quantification of human U6 snRNA and hsa-miRNA22-3p: U6 snRNA,
5′-GTGCTCGCTTCGGCAGCACAT-3′ and 5′-TACCTTGCGAAGTGCTTAAAC-3′, which
produced a segment of 112 bp; and hsa-miRNA-22-3p,
5′-GCCGGCGCCCGAGCTCTGGCTC-3′ and 5′-AAGCTGCCAGTTGAAGAACTGT-3′,
which produced a segment of 72 bp.
Detection of protein contents
The total protein was extracted from the cells using
M-PER mammalian protein extraction reagent (Pierce; Thermo Fisher
Scientific, Inc.). Equal amounts of protein (20 µg per lane)
estimated by a bicinchoninic acid (BCA) protein assay kit (Pierce)
were loaded onto (11%) SDS-PAGE gels and transferred onto
nitrocellulose membranes. The blots were probed with a monoclonal
antibody against human Snail1 (dilution 1:400; cat. no. sc-393172),
P-gp (cat. no. 1:600; cat. no. sc-13131), P53 (dilution 1:500; cat.
no. sc-393031) and β-actin (dilution 1:1,000; cat. no. sc-130065)
(Santa Cruz Biotechnology, Inc., Dallas, TX, USA), followed by the
secondary HRP-conjugated anti-mouse antibody (cat. no. sc-516102;
Santa Cruz Biotechnology). After washing, the bands were detected
by chemiluminescence and imaged with X-ray film. The relative
optical densities were analyzed using Image Processing software
Totallab (Nonlinear Dynamics Ltd. Newcastle, UK) and β-actin was
used as an endogenous reference for normalization.
Statistical analysis
The data are shown as the mean ± SD of three
independent experiments. All statistical data were analyzed using
the SPSS GradPack, version 20.0, statistical software (IBM Corp,
Armonk, NY, USA) and GraphPad Prism 7.0 (GraphPad Software, Inc, La
Jolla, CA, USA). Comparisons between groups were analyzed using
two-tailed Student's t-test or one-way ANOVA with post hoc Tukey's
test. All differences were considered to be statistically
significant at P<0.05.
Results
Expression of Snail1, hsa-miRNA-22-3p,
MDR1 and P53 in MMCs from clinical bortezomib-resistant MM
patients
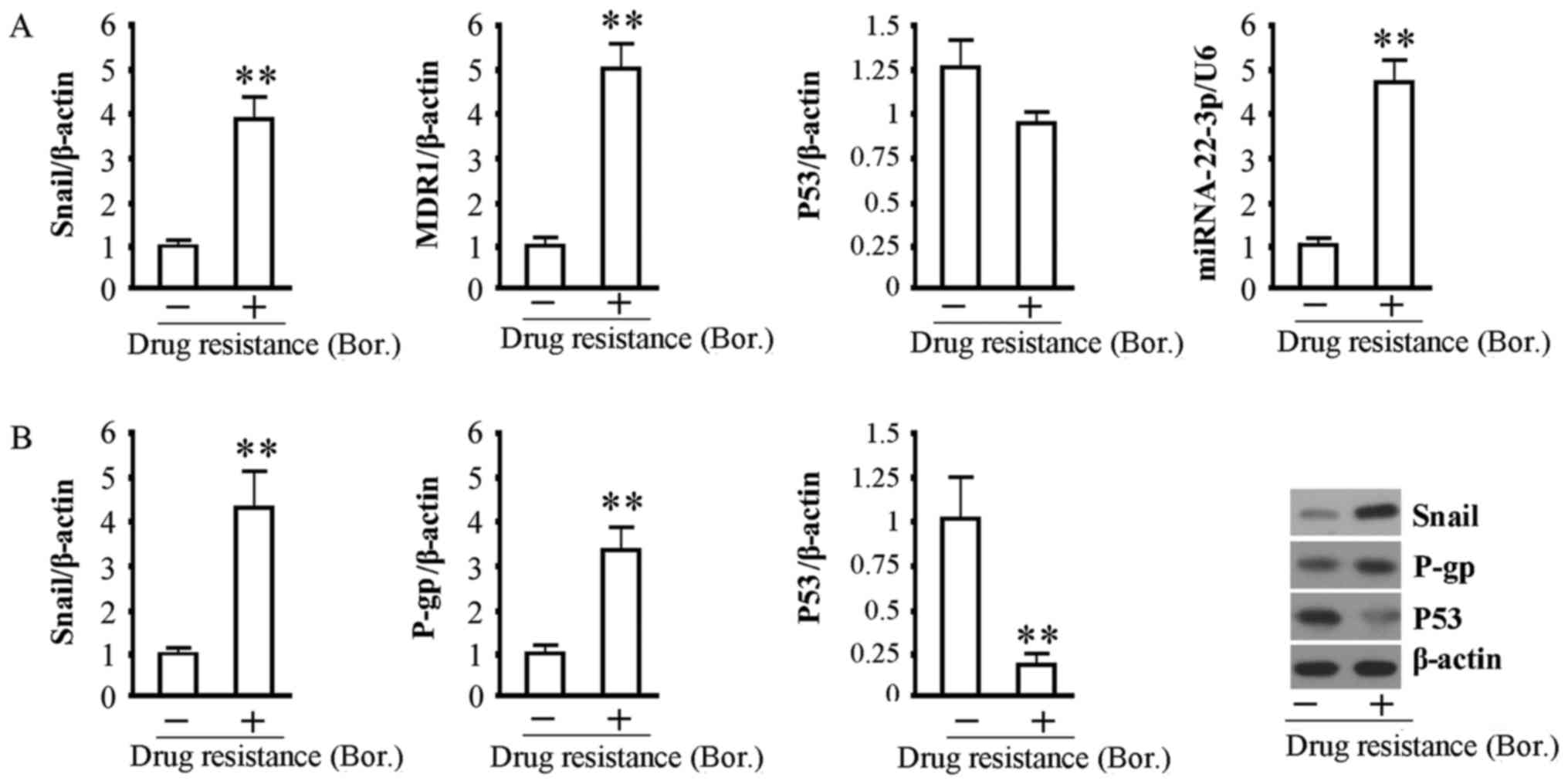
The relative levels of Snail1, MDR1 and
hsa-miRNA-22-3p in MMCs were all significantly increased in MM
patients after the development of bortezomib resistance, and the
mRNA level of P53 was slightly lower compared with MMCs from MM
patients before development of bortezomib resistance (Fig. 1A). The western blot results showed
that the protein expression levels of Snail1 and P-gp were
significantly increased and that the protein expression level of
P53 was significantly decreased in MMCs with drug resistance
compared with MMCs from MM patients before development of
bortezomib resistance (Fig. 1B).
Comprehensive analyses of these indicators showed that the abnormal
expression of MMC drug resistance and apoptosis-related functional
genes, namely MDR1 and P53, in MMCs with drug resistance to
bortezomib resulted from inactivation of the transcription and
post-transcription regulatory mechanisms, respectively. In
addition, the association between hsa-miRNA-22-3p and the
expression of Snail1 and P53 proteins was also consistent with the
transcription regulation of hsa-miRNA-22-3p by Snail1 and the
negative regulation of the target gene P53 by hsa-miRNA-22-3p.
Gene intervention experiment using a
lentiviral approach
Using a lentiviral system, the silencing viruses
targeting the Snail1 and P53 genes, namely Lv-shRNA-Snail1 and
Lv-shRNA-P53, as well as the MDR1-expressing recombinant virus,
Lv-MDR1, were constructed. In addition, the gene intervention
effect was validated in XG-7/Bor cells. After 72 h of treatment
with the viruses at a multiplicity of infection (MOI) of 20, the
estimated virus infection efficiency was no lower than 90% in cells
based on the expression of the GFP fluorescent marker (Fig. 2A). After 72 h of virus infection,
the fluorescence quantitative detection results showed that the
Snail1 and P53 gene silencing rates were ~81.4 and 83.2%,
respectively, and that MDR1 gene expression was ~4.2-fold that of
the control group (Fig. 2B). The
protein expression results showed that the trends in the different
groups were consistent with those of mRNA expression Fig. 2C). These results were consistent
with the relationship among these proteins. In other words, Snail1
silencing significantly decreased Snail1 gene expression in cells,
inhibited MDR1 expression and promoted P53 gene expression, while
exogenous MDR1 overexpression and P53 gene silencing did not have
significant effects on the expression of the Snail1 gene in cells.
These results directly explain the upstream and downstream
regulatory relationship among these genes and proteins.
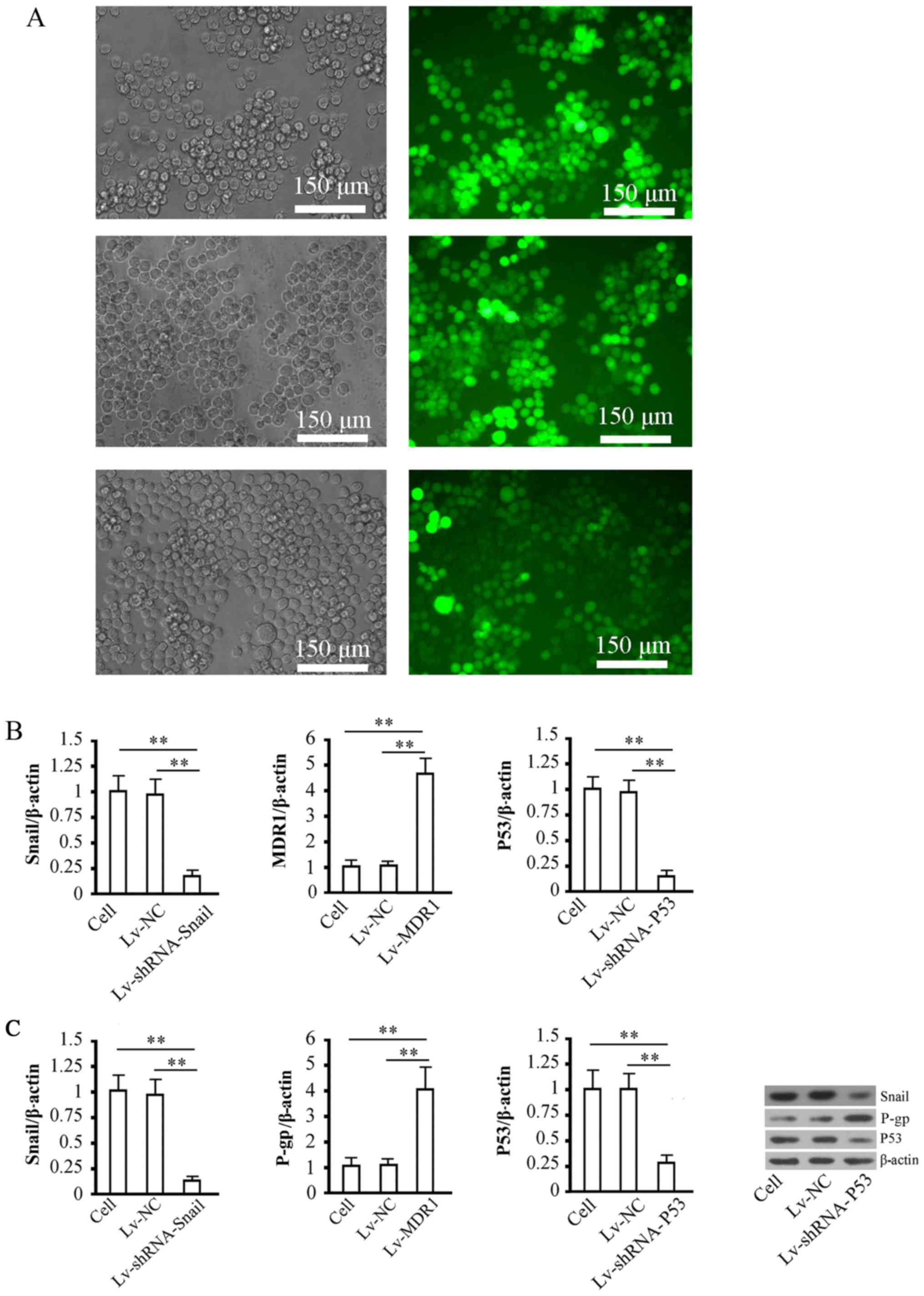 | Figure 2.Detection of virus infection
efficiency and gene intervention efficiency in cells. (A) Detection
of infection efficiency after 72 h of infection with 3 groups of
viruses, namely Lv-shRNA-Snail1 (upper), Lv-MDR1 (middle) and
Lv-shRNA-P53 (lower) in XG-7/Bor cells. The left panels present
images of cells under visible light, and the right panels present
images of cells under UV excitation in the corresponding field. The
virus infection efficiency in cells was estimated by dividing the
number of cells with fluorescence expression by the total number of
cells in the same field. In the statistical analysis, 5 fields were
randomly selected to calculate the virus infection rate, and the
mean value was obtained. (B) Quantitative analyses of mRNA levels
of Snail1, MDR1, and P53 genes in all groups of cells after 72 h of
recombinant virus infection. β-actin was used as the internal
control. The relative level of genes was calculated using the
2∆∆Cq method. (C) Detection of the expression levels of
Snail1, P-gp, and P53 proteins in all groups of cells after 72 h of
recombinant virus infection. The left panels show the target
protein bands, and the right panel shows the relative optical
densities of target proteins in cells among the different groups.
β-actin was used as the internal control. **P<0.01. Cell, cell
control group; Lv-NC, NC control group. |
Validation of the negative regulation
of MDR1 gene expression by hsa-miRNA-22-3p
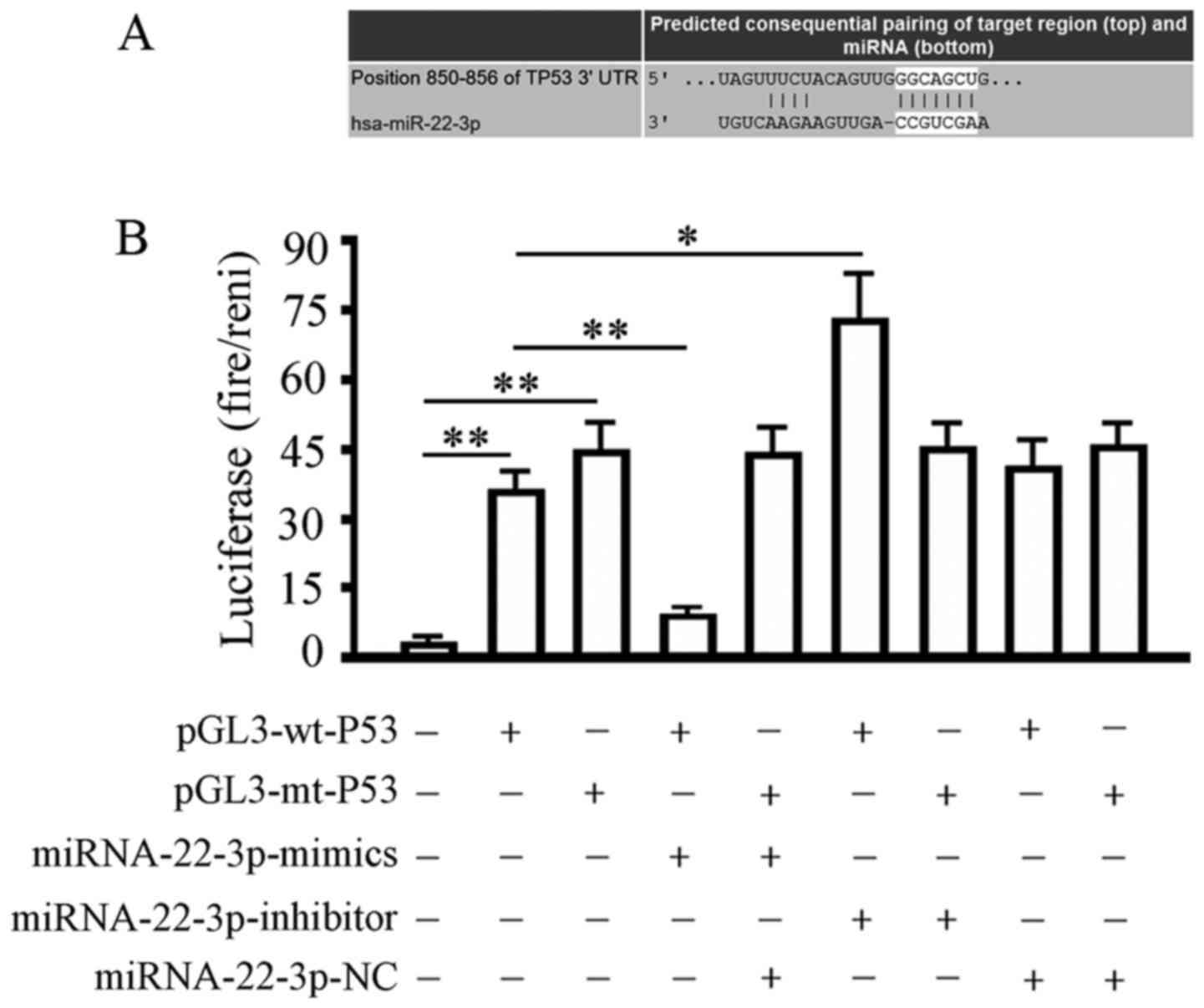
TargetScan Human 7.1 analysis showed that the 3′UTR
region of the P53 gene contained a 6-base seed region, namely
5′-CACUCCA-3′, of hsa-miRNA-22-3p (Fig.
3A). After co-transfection in 293T cells for 48 h, relative
luciferase activity showed that the hsa-miRNA-22-3p-mimic
significantly decreased the luciferase activity of the wild-type
(wt) luciferase reporter gene from 31.02±5.16 to 5.38 ±0.20. The
relative luciferase activity in the hsa-miRNA-22-3p-inhibitor
transfection group increased from 31.02±5.16 to 72.15±14.31. For
the mutant luciferase reporter gene, the difference between all
co-transfection groups and the pGL3-mt-P53 alone transfection group
was not significant (Fig. 3B). The
above data and results indicated the presence of the predicted
target site between hsa-miRNA-22-3p and the P53 gene.
Analyses of transcription regulation
of MDR1 and hsa-miRNA-22-3p by Snail1
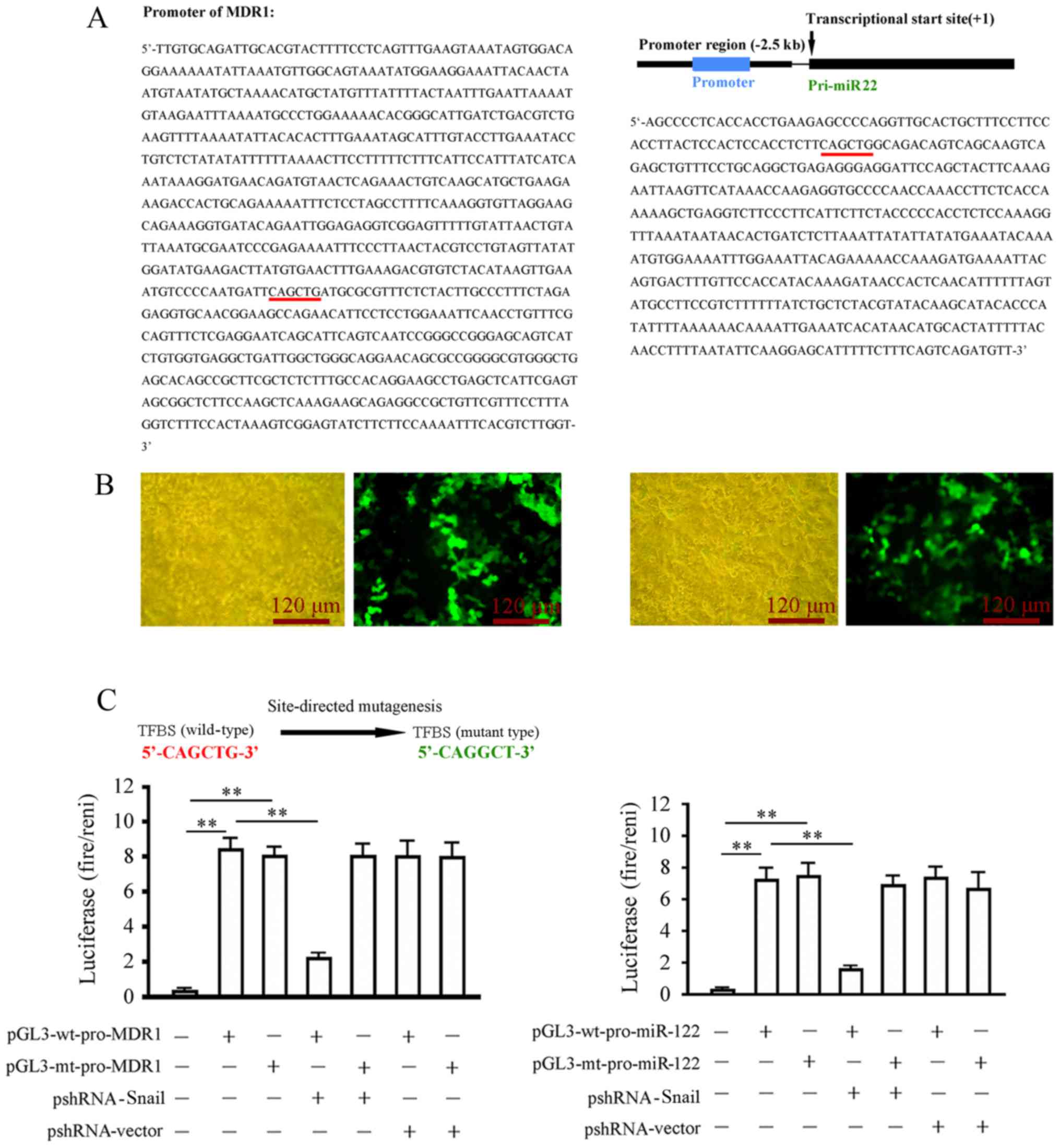
Through literature review and database search, the
promoter sequence of the MDR1 (ACBC1) gene was obtained (Fig. 4A). To identify the hsa-miRNA-22-3p
promoter, the hsa-miRNA22-3p precursor sequence, the pri-miRNA22
gene, in the human genome was first acquired. The 2.5 kb promoter
sequence upstream of the transcription initiation sites was
obtained. Promoter 2.0 prediction software was used to predict the
location of the promoter sequence. The prediction results showed
that the promoter was a 443 bp DNA sequence (Fig. 4A). A GFP-expressing plasmid
containing the hsa-miRNA-22-3p promoter was then constructed for
related tests of promoter activity. The hsa-miRNA-22-3p promoter
effectively induced GFP expression in 293T cells (Fig. 4B). The Snail1 binding sites in these
two promoters were analyzed and predicted using the JASPAR
transcription factor binding site prediction software (http://jaspar.genereg.net/). The results showed that
Snail1 had theoretical transcription factor binding site (TFBS)
(Fig. 4A). The TFBS binding sites
in the two promoters were validated using a luciferase reporter
gene assay. After 48 h of co-transfection in 293T cells, Snail1
silencing (pshRNA-Snail1 transfection) significantly inhibited
luciferase activity of the wild-type reporter gene
(pGL3-wt-pro-MDR1/hsa-miRNA-22-3p) in the transfected cells
compared with the wild-type reporter gene alone transfection group,
but did not have a significant effect on the luciferase activity in
cells transfected with the mutant reporter gene compared with
mutant reporter gene alone transfection group (Fig. 4C). These results indicated that
Snail1 binds to the promoters of MDR1 and hsa-miRNA-22-3p through
the TFBS to positively regulate the transcription of these two
genes.
Analyses of the pathways of
Snail1/MDR1 and Snail1/hsa-miRNA-22-3p/P53 in bortezomib-resistant
MMCs
We performed Snail1 gene silencing experiments in
two drug-resistant cell lines, XG-7/Bor and RPMI-8226/Bor, using
the lentiviral approach. The IC50 values of bortezomib
in these two drug-resistant cell lines after 48 h of gene
intervention were detected and analyzed. The results showed that
the IC50 values of bortezomib in XG-7/Bor and
RPMI-8226/Bor cells after 48 h of Snail1 gene silencing decreased
significantly from 68.52 nM and 81.24 nM to 12.13 nM and 9.28 nM,
respectively (Fig. 5A). The cell
apoptosis rate in the Snail1-silencing group significantly
increased after XG-7/Bor cells in all groups were treated with 10
nM bortezomib for 48 h. The cell apoptosis rates in the
Snail1-silencing combined with MDR1 expression group and the
Snail1-silencing combined with P53-silencing group both
significantly decreased (Fig. 5B).
After cells were infected with Lv-shRNA-Snail1 for 72 h, Snail1 and
P-gp protein expression was significantly decreased, and P53
protein expression was significantly increased. In the
Lv-shRNA-Snail1 infection combined with Lv-MDR1 infection group,
the Snail1 protein expression did not significantly change. In the
same group, P-gp protein expression was significantly increased,
and P53 protein expression was significantly decreased. In the
Lv-shRNA-Snail1 infection combined with Lv-shRNA-P53 infection
group, Snail1 protein expression did not significantly change. In
the same group, P-gp and P53 proteins expression were significantly
decreased. The combination of the above results showed that Snail1
gene silencing significantly enhanced the sensitivity of XG-7/Bor
cells to bortezomib and increased cell apoptosis. MDR1
overexpression or P53 silencing significantly reversed the function
of Snail1 gene silencing in the promotion of bortezomib-induced
apoptosis in XG-7/Bor cells. Exogenous MDR1 did not contain the
wild-type promoter and was not regulated by Snail1. Similarly, P53
gene silencing was transcriptionally regulated and was not
negatively regulated by hsa-miRNA-22-3p. Therefore, Snail1 gene
silencing inhibited MDR1 and upregulated P53 expression to promote
the sensitivity of XG-7/Bor to bortezomib. These results indicate
that although the Snail1/MDR1 and Snail1/hsa-miRNA-22-3p/P53
pathways are regulated by an upstream factor, Snail1, they regulate
multi-drug resistance and apoptosis of MMCs via two independent
pathways and thus co-dominate the development of bortezomib
resistance in MM.
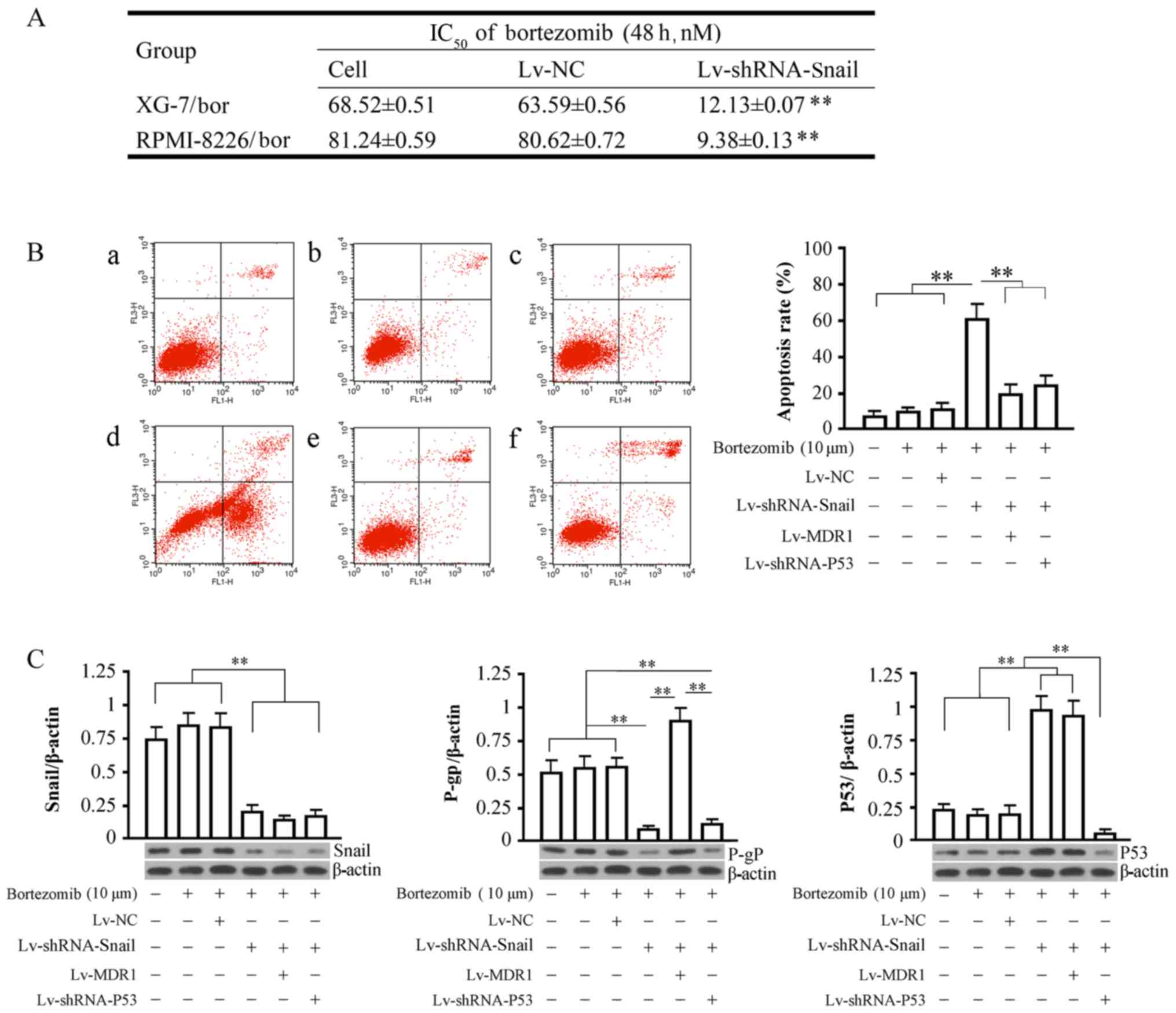 | Figure 5.Analyses of IC50,
apoptosis and protein expression detection. (A) The effect of
Snail1 gene silencing on the IC50 values of bortezomib
in XG-7/Bor and RPMI-8226/Bor cells after 48 h. The groups are
indicated as follows: Cell, cell control group; Lv-NC, NC control
group; Lv-shRNA-Snail1, virus infection group. Cells were infected
with recombinant viruses for 72 h. (B) Detection of cell apoptosis
in all groups. Left: Raw data of cell detection. The horizontal
axis represents fluorescein isothiocyanate (FITC), and the vertical
axis represents propidium iodide (PI). The upper right and lower
right quandrants indicate late-stage and early-stage apoptosis,
respectively. Right: Analyses of apoptosis rates among the groups.
The apoptosis rate was the sum of early-stage apoptosis and
late-stage apoptosis. a, cell control group without bortezomib; b,
cell control group; c, NC control group; d, Snail1-silencing group;
e, MDR1-silencing group; f, P53-silencing group. (C) Detection of
the expression of Snail1, P-gp, and P53 proteins in all groups of
cells. The bottom panels show the target protein bands, and the top
panels show the analysis of the relative optical density of the
target proteins in all groups of cells. β-actin was used at the
internal control. **P<0.01. IC50, half maximal
inhibitory concentration. |
Discussion
Multiple myeloma (MM) is one of the most common
malignant tumors in the hematological system, and its incidence
accounts for 10% of the total number of malignant tumors in the
hematological system (1,11). Currently, there is no effective
clinical measure that can completely cure MM. The commonly used
treatment methods for MM include chemotherapy, radiotherapy,
hematopoietic stem cell transplantation, and combined treatment
using these methods. Various targeted formulations are used for the
chemotherapy of MM. The approval of the clinical use of the
proteasome inhibitor, bortezomib, was important for the
chemotherapy strategy for MM. Bortezomib not only directly inhibits
the proliferation of multiple myeloma cells (MMCs) and induces
apoptosis but also improves the chemotherapy sensitivity of
drug-resistant MMCs to dexamethasone, melphalan and thalidomide
(14). Bortezomib is the first
proteasome inhibitor approved by the FDA to treat recurrent and
refractory MM. The single-drug effective rate of bortezomib in the
treatment of MM is approximately 30%. When bortezomib is used in
combination with dexamethasone or immunomodulators, the effective
rate increases to more than 60%. The 2008 National Comprehensive
Cancer Network (NCCN) guideline listed bortezomib single-drug and
its combination with liposomal doxorubicin as the category I
preferred regimen and bortezomib + dexamethasone as the category 2A
preferred regimen (15). As the
first-line drug for MM treatment, bortezomib has its own advantages
(16). Although the chemotherapy
regimen based on bortezomib significantly improves the related
symptoms of MM patients within a short time period and delays tumor
progression, almost all MM patients with bortezomib administration
exhibit drug resistance features after bortezomib medication for a
period of time, which greatly limits the long-term efficacy of
bortezomib in MM (17). Currently,
basic studies on drug resistance in MM chemotherapy indicate that
the mechanisms underlying the development of drug resistance in MM
chemotherapy might be associated with NF-κB activity as well as
overexpression of heat shock proteins and BCL protein family
members (18–20). However, these previous studies did
not provide complete, in-depth, and systematic elucidation of the
development of MM clinical drug resistance mechanisms. Thus,
additional studies on the mechanisms of drug resistance in MM
chemotherapy are required. Using the drug resistance theory studies
as guidance, improvement of drug resistance mechanisms, screening
and demonstration of key gene targets to improve drug resistance as
well as searching for effective regimens to prevent the development
of drug resistance or eradicate malignant plasma cells are
important directions for studies on the treatment theory of MM in
the future. At present, there are few studies on the drug
resistance mechanism of MM to bortezomib. Thus, the drug resistance
mechanism remains unclear, and there is no effective clinical
solution.
The Snail superfamily has been highly conserved
during the evolution of vertebrates. The human Snail1 protein is
composed of 264 amino acids. The Snail1 protein has an
amino-terminal basic amino acid-rich domain and a carboxyl-terminal
DNA-binding domain, and these two special domains regulate its
interaction with co-repressors and the epigenetic remodeling
complex and DNA binding, respectively. A serine rich domain (SRD)
and nuclear export sequence (NSE) control the stability and
subcellular location of the Snail1 protein (21). As a transcription factor, snail1
functions through binding to the E-box element of the 5′-CANNTG-3′
core base sequence in the promoter region of many tumor-related
genes (22). Snail1 can promote or
inhibit the transcription of E-cadherin, IL-8 and CD147 to regulate
the epithelial-mesenchymal transition (EMT) process of many tumors
(23–25). The Snail family contains three
family members, namely Snail1, Snail2 and Snail3. Studies in recent
years have indicated that Snail1 plays a critical role in the
development and regulation of drug resistance mechanisms in tumors.
The expression of Snail1 protein in non-small cell lung cancer
(NSCLC) positively correlates with the expression of drug
resistance-related genes. Blocking Snail1 in the A549 lung cancer
cell line effectively reverses the multiple drug resistance
characteristics of lung cancer. Snail1 expression was found to be
significantly increased in the H446/CDDP NSCLC cell line compared
to that in the H446 cell line, indicating that the drug resistance
of NSCLC to cisplatin might be associated with the high expression
of Snail1. Therefore, Snail1 can be used as a new target for
clinical treatment of drug-resistant lung cancer patients (26,27).
The high Snail1 expression in bladder cancer tissues has a
significant positive correlation with ERCC1 (28). Furthermore, Snail1 has also been
confirmed to be a promotion factor (29) for the development of drug resistance
mechanisms in breast cancer (30)
and prostate cancer (31). P53 is
currently one of the most extensively studied genes. Wild-type P53
is a tumor-suppressor gene, and it is a negative regulatory factor
in cell growth cycles and inhibits tumor development (32). Studies on P53 gene functions in
recent years have indicated that P53 and many factors have mutual
regulation functions and can regulate and improve the sensitivity
of tumor cells to the treatment of various chemotherapeutic drugs
(33). One recent study has
indicated that loss of P53 causes the abnormal expression of the
RNA-binding protein HnRNPA0 to eventually result in the development
of drug resistance in various tumor cells to chemotherapy that can
kill them (34). In addition, P53
has also been confirmed to directly or indirectly participate in
the development of drug resistance mechanisms in many types of
tumors (35).
The multidrug resistance (MDR) phenomenon refers to
the development of multiple drug resistance in tumor cells after
development of drug resistance to one chemotherapeutic drug. In
other words, tumor cells will develop different levels of drug
resistance to other antitumor drugs that have different structures
or different targets. The MDR of tumor cells is an important reason
causing the ineffectiveness of antitumor drug treatment. The
abnormal expression of MDR and its expression product,
P-glycoprotein, is considered one of the major reasons for the
development of MDR in tumor cells. The human MDR gene family
includes MDR1 and MDR2. MDR1 is associated with drug resistance in
various types of malignant tumors, but MDR2 cannot cause tumor
cells to develop drug resistance (36). P-gp is an important expression
product of the MDR gene. P-gp is an adenosine triphosphate
(ATP)-dependent efflux pump, and it has an ATP-binding site. When
tumor cells come into contact with antitumor drugs, lipid-soluble
drugs enter into cells with the concentration gradient. P-gp binds
to drug molecules and connects the molecules to the ATP-binding
site to form a phosphorylated and glycosylated ATP-binding cassette
of drug macromolecules. During ATP hydrolysis, the released energy
pumps drugs that enter into cells outside cells to decrease the
drug concentration in tumor cells. Therefore, MDR1 expression in
the body of tumor patients is associated with MDR of tumor cells.
Exogenous substances, such as natural hydrophobic antitumor drugs
including anthracyclines, vincristine alkaloids,
epipodophyllotoxin, actinomycin D and paclitaxel, are easily pumped
out by MDR1 to reduce the drug concentration in cells, decrease the
antitumor cytotoxicity, and promote drug resistance. It has also
been shown that the drug resistance levels of tumor cells,
intracellular drug concentrations, and MDR expression on the
surface of the cell membrane are closely associated (37). Blockade of MDR1 using calcium
channel blockers significantly inhibits the efflux of drugs to
increase the aggregation of drugs in cells, reverse drug resistance
of cells, and increase the sensitivity of tumor cells to
chemotherapeutic drugs (38). The
function of miRNAs in drug resistance mechanisms in tumors is
always an important direction in tumor drug resistance studies
(39–41).
Hsa-miRNA22 localizes to human chromosome 17.
Current studies on hsa-miRNA22 in tumors have mainly focused on its
regulation in tumor progression (42,43).
However, there is no report on the association of hsa-miRNA-22 with
blood tumors and their drug resistance mechanisms.
This study is the first systematic study to focus on
the critical role of the Snail1 nuclear transcription factor in the
development of drug resistance mechanisms to bortezomib in MM and
the first to elucidate the Snail1/MDR1 and
Snail1/hsa-miRNA-22-3p/P53 drug resistance mechanism regulatory
pathways. The results have extraordinary significance in the
development of highly efficient therapeutic methods in the clinical
treatment of MM using bortezomib. Taking into consideration the
significant association between Snail1 and bortezomib-resistance in
MM, we think that understanding the action mechanism of Snail1 in
bortezomib-resistance is critical to the development of the
clinical therapy of MM by using gene therapy combined with
bortezomib. Therefore, to ascertain whether the increase in Snail1
expression is directly associated with the development of
bortezomib drug resistance mechanisms in MM, we silenced the Snail1
gene using a lentiviral experimental system in two
bortezomib-resistant MM cell lines, XG-7/Bor and RPMI-8226/Bor. The
IC50 values (48 h) of bortezomib in the drug-resistant
XG-7/Bor and RPMI-8226/Bor cell lines decreased from 68.5 nM and
81.2 nM to 12.1 nM and 9.4 nM, respectively, indicating that Snail1
silencing significantly reversed the drug resistance of MMCs to
bortezomib. We next performed deep mining on transcriptome data.
Combined with association analysis of drug resistance-related
genes, the pathways of Snail1 that influenced drug resistance were
investigated. Data analyses showed that Snail1 was associated with
the expression of the MDR1 gene and P53 protein, respectively. MDR1
and P53 are functional genes that are directly associated with drug
resistance (44). Therefore, this
study investigated whether the expression of MDR1 and P53 are
directly associated with Snail1 and whether they are in the
pathways of drug resistance improvement by Snail1 silencing. The
transcription and protein expression levels of MDR1 and P53 suggest
that the association between Snail1 and MDR1 was at the
transcription level (its mRNA and protein levels were consistently
changed) at that P53 was post-transcriptionally regulated (its mRNA
level was not changed but the protein content was changed). Since
Snail1 is a transcription factor, we have a reason to think that
Snail1 may regulate MDR1 transcription by binding to its promoter,
and may bind to various promoters of miRNAs and thus negatively
regulate P53 expression by stimulating the transcription of this
miRNA, that is, the former is direct regulation, and the latter is
indirect regulation. A series of bioinformatic prediction data
indicated that the MDR1 gene promoter contains a TFBS of Snail1 and
that the regulation of P53 expression by Snail1 might be mediated
by hsa-miRNA-22-3p. The findings also suggest that there is a TFBS
of Snail1 in the promoter region of the hsa-miRNA-22-3p precursor
sequence and that the 3′UTR region of the P53 gene has a
hsa-miRNA-22-3p-binding site (seed region). Therefore, we
hypothesize that inactivation of the regulation of the Snail1/MDR1
and Snail1/hsa-miRNA-22-3p/P53 pathways occurs during development
of bortezomib drug resistance mechanisms in MM and that silencing
of Snail1 effectively reverses the drug resistance of
bortezomib-resistant MMCs through the inhibition of MDR1
transcription and promotion of P53 protein expression. Elucidation
of the above-mentioned drug resistance mechanisms and regulatory
pathways will provide a solid theoretical basis for the treatment
of MM by Snail1-based gene intervention combined with
bortezomib.
The development of drug resistance mechanisms
results from the combination of the comprehensive functions of
multiple factors. After confirmation of the leading functional
routes and regulatory pathways, we can start at the upstream
pathways to look for more high-efficient gene targets to improve
the clinical efficacy of bortezomib using gene intervention
methods. The most significant conclusion of this study is that
Snail1 gene silencing can effectively increase the sensitivity of
two groups of bortezomib-resistant MMCs to bortezomib, which
provides a theoretical basis for high-efficient therapy using genes
combined with bortezomib to target MM using Snail1 as the gene
target. The most evident feature of this high-efficient therapy is
to improve the rapid development of bortezomib drug resistance
mechanisms in MMCs to the maximum extent. The reasons for the
enhancement of Snail1 expression in bortezomib-resistant MMCs were
not further analyzed, but future studies in this direction are
planned. However, that does not affect the significance of this
study at this stage. In addition, this study on hsa-miRNA-22-3p
first demonstrated its association with drug resistance in tumors.
These results suggest that hsa-miRNA-22-3p has the potential to
become a biomarker for screening the bortezomib drug-resistant
population in MM patients, which has important significance in the
development of drug administration methods and comprehensive
therapy methods for treatment of MM patients. Therefore, related
studies on hsa-miRNA-22-3p in bortezomib drug resistance is also an
important direction for our future studies.
Overall, this study elucidated the function of
Snail1 in the development of bortezomib drug resistance mechanisms
in MM, confirmed the drug resistance regulatory pathways of
Snail1/MDR1 and Snail1/hsa-miRNA-22-3p/P53, and preliminarily
established a method to improve the sensitivity of
bortezomib-resistant MMCs to bortezomib treatment by silencing the
Snail1 gene. These findings shed light on the development of drug
resistance mechanisms to bortezomib in MM and provide information
that may increase the clinical efficacy of bortezomib and improve
the poor prognosis of MM patients.
Acknowledgements
This research was supported by the Central
Laboratory of the First Affiliated Hospital of Anhui Medical
University.
Funding
The present study was supported by the Anhui
Provincial Natural Science Foundation of China (grant no.
1208085QH154 and 1708085MH224).
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
ZH was responsible for the formulation of the
research plan, the analysis of the results and the writing of the
manuscript. ZH, XL, WW and XC were specifically responsible for the
implementation of the plan, including cell culture, collection of
patient data, testing of drug resistance genes and other tests and
the statistical analysis of the results. QZ, MY, JG and RX were
responsible for the revision of the plan, guidance of the tests,
the evaluation of the results and the revision of the manuscript.
All authors read and approved the manuscript and agree to be
accountable for all aspects of the research in ensuring that the
accuracy or integrity of any part of the work are appropriately
investigated and resolved.
Ethics approval and consent to
participate
The study was approved by the Medical Ethics
Committee of the First Affiliated Hospital of Anhui Medical
University.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Meister S, Schubert U, Neubert K, Herrmann
K, Burger R, Gramatzki M, Hahn S, Schreiber S, Wilhelm S, Herrmann
M, et al: Extensive immunoglobulin production sensitizes myeloma
cells for proteasome inhibition. Cancer Res. 67:1783–1792. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Griffin PT, Ho VQ, Fulp W, Nishihori T,
Shain KH, Alsina M and Baz RC: A comparison of salvage infusional
chemotherapy regimens for recurrent/refractory multiple myeloma.
Cancer. 121:3622–3630. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Nikesitch N and Ling SC: Molecular
mechanisms in multiple myeloma drug resistance. J Clin Pathol.
69:97–101. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Adams J: The proteasome: A suitable
antineoplastic target. Nat Rev Cancer. 4:349–360. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Chauhan D and Anderson KC: Mechanisms of
cell death and survival in multiple myeloma (MM): Therapeutic
implications. Apoptosis. 8:337–343. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Qin JZ, Ziffra J, Stennett L, Bodner B,
Bonish BK, Chaturvedi V, Bennett F, Pollock PM, Trent JM, Hendrix
MJ, et al: Proteasome inhibitors trigger NOXA-mediated apoptosis in
melanoma and myeloma cells. Cancer Res. 65:6282–6293. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Roccaro AM, Hideshima T, Raje N, Kumar S,
Ishitsuka K, Yasui H, Shiraishi N, Ribatti D, Nico B, Vacca A, et
al: Bortezomib mediates antiangiogenesis in multiple myeloma via
direct and indirect effects on endothelial cells. Cancer Res.
66:184–191. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Murray MY, Auger MJ and Bowles KM:
Overcoming bortezomib resistance in multiple myeloma. Biochem Soc
Trans. 42:804–808. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Zhou W, Yang Y, Xia J, Wang H, Salama ME,
Xiong W, Xu H, Shetty S, Chen T, Zeng Z, et al: NEK2 induces drug
resistance mainly through activation of efflux drug pumps and is
associated with poor prognosis in myeloma and other cancers. Cancer
Cell. 23:48–62. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Yu W, Chen Y, Xiang R, Xu W, Wang Y, Tong
J, Zhang N, Wu Y and Yan H: Novel phosphatidylinositol 3-kinase
inhibitor BKM120 enhances the sensitivity of multiple myeloma to
bortezomib and overcomes resistance. Leuk Lymphoma. 58:428–437.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Abe M: Multiple myeloma. Nihon Rinsho.
67:991–995. 2009.(In Japanese). PubMed/NCBI
|
|
12
|
Tomono T, Yano K and Ogihara T:
Snail-induced epithelial-to-mesenchymal transition enhances
P-gp-mediated multidrug resistance in HCC827 cells. J Pharm Sci.
106:2642–2649. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Cho JH, Lee SJ, Oh AY, Yoon MH, Woo TG and
Park BJ: NF2 blocks Snail-mediated p53 suppression in mesothelioma.
Oncotarget. 6:10073–10085. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Abdi J, Chen G and Chang H: Drug
resistance in multiple myeloma: Latest findings and new concepts on
molecular mechanisms. Oncotarget. 4:2186–2207. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Gaultney JG, Ng TW, Uyl-de Groot CA,
Sonneveld P, van Beers EH, van Vliet MH and Redekop WK: Potential
therapeutic and economic value of risk-stratified treatment as
initial treatment of multiple myeloma in Europe. Pharmacogenomics.
19:213–226. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Wu T, Zhou J, Wang C, Wang B, Zhang S and
Bai H: Bortezomib overcomes the negative prognostic impact of renal
impairment in a newly diagnosed elderly patient with multiple
myeloma: A case report. Oncol Lett. 14:7318–7322. 2017.PubMed/NCBI
|
|
17
|
Gavriatopoulou M, Terpos E, Kastritis E
and Dimopoulos MA: Efficacy and safety of elotuzumab for the
treatment of multiple myeloma. Expert Opin Drug Saf. 16:237–245.
2017.PubMed/NCBI
|
|
18
|
Adam Z, Sčudla V, Krejčí M, Cermáková Z,
Pour L and Král Z: Treatment of AL amyloidosis in 2012; the benefit
of new drugs (bortezomib, thalidomide, and lenalidomide). Summary
of published clinical trials. Vnitr Lek. 59:37–58. 2013.(In
Czech).
|
|
19
|
Abidi MH, Gul Z, Abrams J, Ayash L, Deol
A, Ventimiglia M, Lum L, Mellon-Reppen S, Al-Kadhimi Z,
Ratanatharathorn V, et al: Phase I trial of bortezomib during
maintenance phase after high dose melphalan and autologous stem
cell transplantation in patients with multiple myeloma. J
Chemother. 24:167–172. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Jacob P, Hirt H and Bendahmane A: The
heat-shock protein/chaperone network and multiple stress
resistance. Plant Biotechnol J. 15:405–414. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Carmichael CL and Haigh JJ: The Snail
family in normal and malignant haematopoiesis. Cells Tissues
Organs. 203:82–98. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Nieto MA: The snail superfamily of
zinc-finger transcription factors. Nat Rev Mol Cell Biol.
3:155–166. 2002. View
Article : Google Scholar : PubMed/NCBI
|
|
23
|
Cheng CW, Wu PE, Yu JC, Huang CS, Yue CT,
Wu CW and Shen CY: Mechanisms of inactivation of E-cadherin in
breast carcinoma: Modification of the two-hit hypothesis of tumor
suppressor gene. Oncogene. 20:pp. 3814–3823. 2001, View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Argast GM, Krueger JS, Thomson S,
Sujka-Kwok I, Carey K, Silva S, O'Connor M, Mercado P, Mulford IJ,
Young GD, et al: Inducible expression of TGFβ, snail and Zeb1
recapitulates EMT in vitro and in vivo in a NSCLC model. Clin Exp
Metastasis. 28:593–614. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Barnett P, Arnold RS, Mezencev R, Chung
LW, Zayzafoon M and Odero-Marah V: Snail-mediated regulation of
reactive oxygen species in ARCaP human prostate cancer cells.
Biochem Biophys Res Commun. 404:34–39. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Wang H, Zhang G, Zhang H, Zhang F, Zhou B,
Ning F, Wang HS, Cai SH and Du J: Acquisition of
epithelial-mesenchymal transition phenotype and cancer stem
cell-like properties in cisplatin-resistant lung cancer cells
through AKT/β-catenin/Snail signaling pathway. Eur J Pharmacol.
723:156–166. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Zhang K, Wang X and Wang H: Effect and
mechanism of Src tyrosine kinase inhibitor sunitinib on the
drug-resistance reversal of human A549/DDP cisplatin-resistant lung
cancer cell line. Mol Med Rep. 10:2065–2072. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Kawashima A, Takayama H, Kawamura N, Doi
N, Sato M, Hatano K, Nagahara A, Uemura M, Nakai Y, Nishimura K, et
al: Co-expression of ERCC1 and Snail is a prognostic but not
predictive factor of cisplatin-based neoadjuvant chemotherapy for
bladder cancer. Oncol Lett. 4:15–21. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Brozovic A: The relationship between
platinum drug resistance and epithelial-mesenchymal transition.
Arch Toxicol. 91:605–619. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Semina SE, Scherbakov AM, Kovalev SV,
Shevchenko VE and Krasil'nikov MA: Horizontal transfer of tamoxifen
resistance in MCF-7 cell derivates: Proteome study. Cancer Invest.
35:506–518. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Ware KE, Somarelli JA, Schaeffer D, Li J,
Zhang T, Park S, Patierno SR, Freedman J, Foo WC, Garcia-Blanco MA,
et al: Snail promotes resistance to enzalutamide through regulation
of androgen receptor activity in prostate cancer. Oncotarget.
7:50507–50521. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Zhan M, Yu D, Lang A, Li L and Pollock RE:
Wild type p53 sensitizes soft tissue sarcoma cells to doxorubicin
by down-regulating multidrug resistance-1 expression. Cancer.
92:1556–1566. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Hasna J, Hague F, Rodat-Despoix L, Geerts
D, Leroy C, Tulasne D, Ouadid-Ahidouch H and Kischel P: Orai3
calcium channel and resistance to chemotherapy in breast cancer
cells: The p53 connection. Cell Death Differ. 25:691–705.
2018.PubMed/NCBI
|
|
34
|
Cannell IG, Merrick KA, Morandell S, Zhu
CQ, Braun CJ, Grant RA, Cameron ER, Tsao MS, Hemann MT and Yaffe
MB: A pleiotropic RNA-binding protein controls distinct cell cycle
checkpoints to drive resistance of p53-defective tumors to
chemotherapy. Cancer Cell. 28:8312015. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Luanpitpong S, Angsutararux P, Samart P,
Chanthra N, Chanvorachote P and Issaragrisil S:
Hyper-O-GlcNAcylation induces cisplatin resistance via regulation
of p53 and c-Myc in human lung carcinoma. Sci Rep. 7:106072017.
View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Singh MS, Tammam SN, Shetab Boushehri MA
and Lamprecht A: MDR in cancer: Addressing the underlying cellular
alterations with the use of nanocarriers. Pharmacol Res. 126:2–30.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Montazami N, Aghapour M, Farajnia S and
Baradaran B: New insights into the mechanisms of multidrug
resistance in cancers. Cell Mol Biol (Noisy-le-grand). 61:70–80.
2015.PubMed/NCBI
|
|
38
|
Mancuso MR and Massarweh SA: Endocrine
therapy and strategies to overcome therapeutic resistance in breast
cancer. Curr Probl Cancer. 40:95–105. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Bach DH, Hong JY, Park HJ and Lee SK: The
role of exosomes and miRNAs in drug-resistance of cancer cells. Int
J Cancer. 141:220–230. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Çalışkan M, Güler H and Bozok Çetintaş V:
Current updates on microRNAs as regulators of chemoresistance.
Biomed Pharmacother. 95:1000–1012. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Leivonen SK, Icay K, Jäntti K, Siren I,
Liu C, Alkodsi A, Cervera A, Ludvigsen M, Hamilton-Dutoit SJ,
d'Amore F, et al: MicroRNAs regulate key cell survival pathways and
mediate chemosensitivity during progression of diffuse large B-cell
lymphoma. Blood Cancer J. 7:6542017. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Fan W, Huang J, Xiao H and Liang Z:
MicroRNA-22 is downregulated in clear cell renal cell carcinoma,
and inhibits cell growth, migration and invasion by targeting PTEN.
Mol Med Rep. 13:4800–4806. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Wang J, Li Y, Ding M, Zhang H, Xu X and
Tang J: Molecular mechanisms and clinical applications of miR-22 in
regulating malignant progression in human cancer (Review). Int J
Oncol. 50:345–355. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
JThottassery JV, Zambetti GP, Arimori K,
Schuetz EG and Schuetz JD: p53-dependent regulation of MDR1 gene
expression causes selective resistance to chemotherapeutic agents.
Proc Natl Acad Sci USA. 94:11037–11042. 1997. View Article : Google Scholar : PubMed/NCBI
|