Introduction
Glioblastoma multiforme (GBM), which is classified
by the World Health Organization as a grade IV glioma, exhibits a
high morbidity and mortality, comprising 47.1% of all malignant
tumors of the central nervous system (1,2). In
total, ~13,000 people in America are diagnosed with GBM each year
(3). The main treatment of GBM is
surgical resection in combination with radiotherapy or
chemotherapy. However, the majority of patients relapse within the
7 months following their original diagnoses (4). Furthermore, a resistance to current
chemotherapy leads to a heavy tumor burden for patients with GBM.
Although novel treatments, including immunotherapy and molecular
targeted therapy, have been in development for several years
(5,6), the 5-year survival rate is relatively
low, with a median survival time of 15 months (4), indicating the urgency of determining
novel therapies.
MicroRNAs (miRNAs) are a class of small non-coding
RNAs comprising ~19–23 nucleotides (7). By binding to target mRNAs, miRNAs
regulate gene expression at the transcriptional or
posttranscriptional level, which enables them to serve pivotal
roles in various biological processes, including cell growth,
apoptosis, invasion and metastasis (8,9). In
terms of GBM, it is widely reported that miRNAs participate in
various molecular pathways associated with cancer development
(10–12). miRNA-targeted therapy has been
utilized in cancer treatment by developing miRNA mimetic and
anti-miRNA agents (13). The
present study aimed to determine latent miRNA-based therapeutic
agents for GBM by employing a Connectivity Map (CMap) method. Using
>7,000 expression profiles (representing 1,309 compounds), the
CMap reveals connections among genes, chemicals and diseases
(14). It is extensively used in
the exploitation of novel drugs or existing drug applications
(15,16). Drugs that are available in CMap are
all licensed for human use by the Food and Drug Administration
(17). It is an ideal database for
probing chemicals that may be applied in the therapy of GBM.
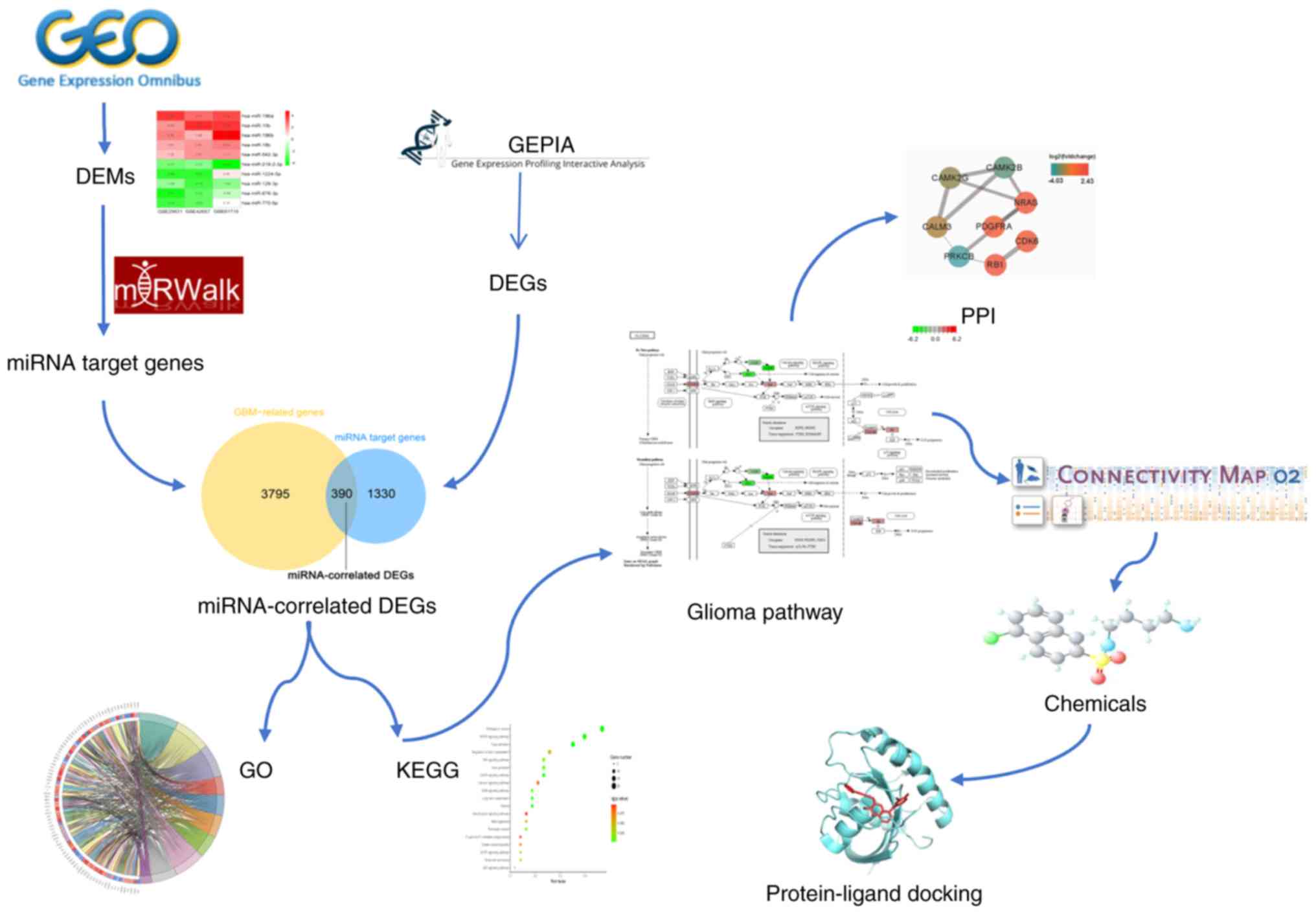
The experimental processes of the present study were
as follows (Fig. 1): i) A
statistical analysis of differentially expressed miRNAs (DEMs) in
GBM was performed using the expression data of gene chips collected
from the Gene Expression Omnibus (GEO); ii) the target genes of
DEMs were predicted and intersected with the GBM-associated genes
and the resulting genes were defined as miRNA-associated
differentially expressed genes (DEGs); iii) Gene Ontology (GO)
functional annotations and a Kyoto Encyclopedia of Genes and
Genomes (KEGG) pathway analysis were performed, and the genes
participating in the ‘glioma pathway’ were selected for further
analysis; iv) a CMap analysis was applied to probe for potential
therapeutic chemicals for GBM; and v) a molecular docking approach
was adopted to assess the affinity of the selected chemicals to
their target genes.
Materials and methods
Identification of DEMs in GBM
A search of miRNA-associated microarray datasets in
GBM was performed using the GEO database (18) with the following screening criteria:
i) the organism must be restricted to ‘homo sapiens’; ii) a
complete expression profile determination of the miRNA had to have
been performed; and iii) cancerous and noncancerous samples needed
to contain at least five samples. A Bioconductor package ‘Limma’
(19) was applied to screen for
DEMs in the available gene chip. The DEMs in the individual gene
chips were then integrated and ranked with RobustRankAggreg
(20), which is an R package for
the comprehensive integration of the gene list.
Collection of miRNA-associated
DEGs
With filter conditions of [log2 (fold
change)]≥1.5 and an adjusted P<0.05, DEGs in GBM were collected
from Gene Expression Profiling Interactive Analysis (GEPIA)
(21), an online interactive server
for gene sequence analysis that provides a gene expression profile
comparison between 163 GBM specimens and 207 normal brain samples
from the Cancer Genome Atlas and Genotype-Tissue Expression
projects.
The target genes of DEMs were predicted using
miRWalk (22), an integrated
resource that provides predictions of miRNA- target interactions.
Only the target genes that were predicted by at least eight
target-predicting algorithms were considered to intersect with the
DEGs gathered from GEPIA. Overlapping genes were regarded as
miRNA-associated DEGs in GBM.
GO and KEGG enrichment analyses and
protein-protein interaction (PPI) network construction
GO and KEGG enrichment analyses were performed using
the ClusterProfiler (23), an R
package that is used for the systematic analysis of gene clusters.
A PPI network was constructed using STRING (24), a database that provides functional
interactions among proteins.
Immunohistochemistry
The Human Protein Atlas (https://www.proteinatlas.org/), an interactive web
tool that assesses proteins in all major tissues and organs in the
human body (25) was used to
determine the expression level of proteins. The main clinical
characteristic of the patients are presented in Table SII.
Drug discovery in CMap
Query genes were uploaded to the CMap web tool
(26), comparing >7,000 gene
expression profiles following treatment with 1,309 active chemicals
in human cell lines. The link between the query genes and the 1,309
chemicals was measured via a connectivity score provided by the
CMap tool, which ranged valued from −1 to 1. A positive score
implied a stimulative effect, while a negative score indicated a
suppressive effect of the chemical on the given signatures.
Molecular docking analysis
Molecular docking between the proteins encoded by
miRNA-associated DEGs and filtered chemicals was performed using
Sybyl-X (27). Protein crystal
structures were downloaded from the Research Collaboratory for
Structural Bioinformatics Protein Data Bank (PDB) (28) and chemical structures were obtained
from PubChem (29). First, the
protein crystal structure was imported into the Sybyl-X 2.1.1
software on the Surflex-Dock interface. Following the removal of
irrelevant water molecules and ions, the repair of side chains, the
charging of terminal groups and the addition of polar hydrogen
atoms, the proteins were prepared for docking. Protomols, which are
active pockets that ligands are aligned to, were generated with the
use of an automatic mode. Compounds in the mol2 format were then
imported into the software on the Docking interface and
protein-ligand docking was run under the surflex-dock geom mode,
after which a total score was exported, with these scores being
directly proportional to the binding affinity. Other parameters
were set by default. The binding mode was visualized by the use of
PyMOL (30).
Results
Screening of 10 DEMs in GBM by the
RobustRankAggreg method
Three gene chips [GSE25631 (31), GSE42657 (32) and GSE61710 (33)], were collected from GEO. The basic
information of the three microarray datasets are presented in
Table I and the main
clinicopathological characteristics of the samples in the three
gene chips are presented in supplementary Table SI with the screening criteria of
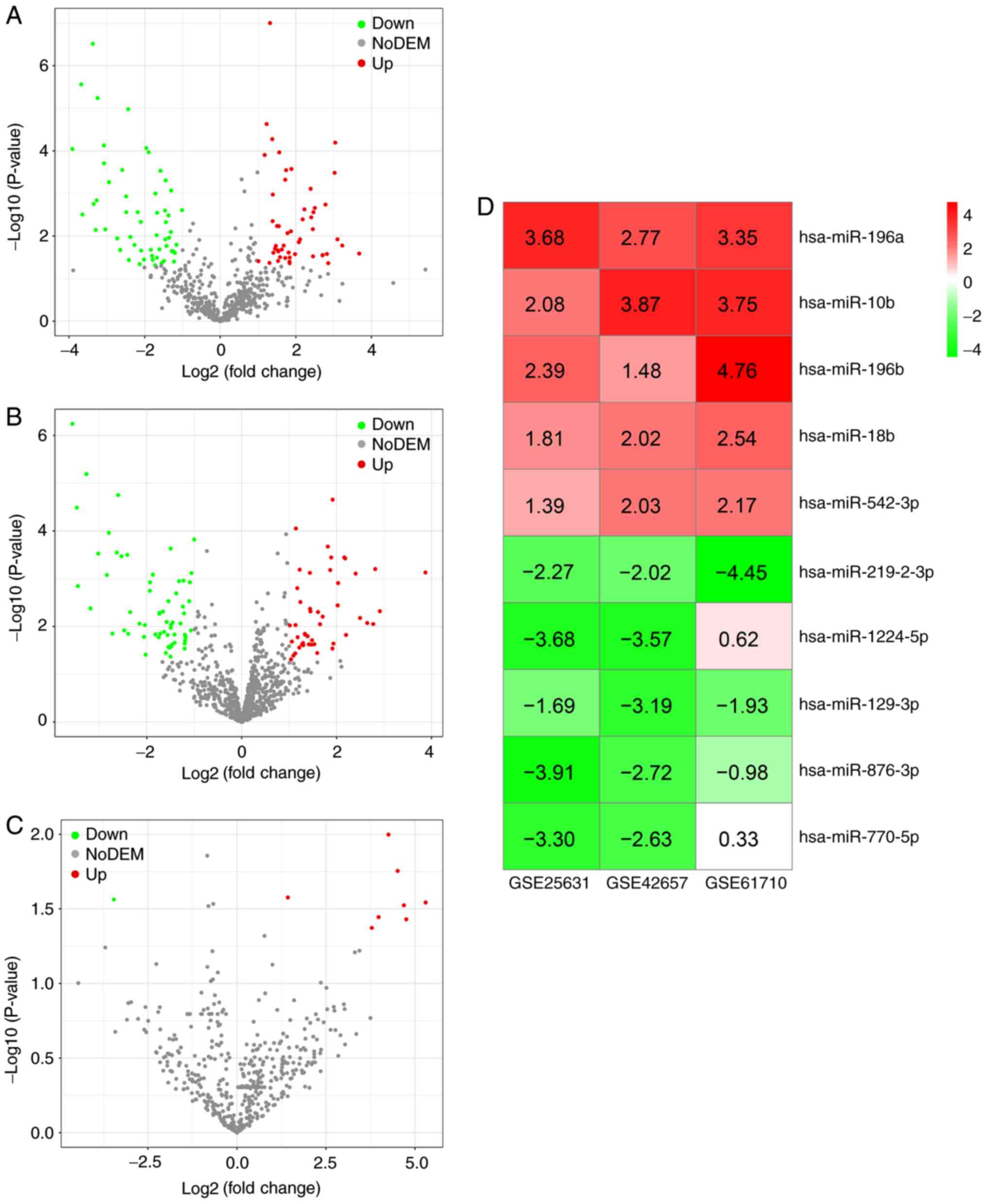
[log2 (fold change)] >1 and P<0.05, 51 upregulated miRNAs and
56 downregulated miRNAs were obtained in gene chip GSE25631
(Fig. 2A); 47 upregulated miRNAs
and 61 downregulated miRNAs were obtained in gene chip GSE42657
(Fig. 2B); and eight upregulated
miRNAs and one downregulated miRNA were identified in gene chip
GSE61710 (Fig. 2C). Following
robust rank aggregation, an ordered list of all of the DEMs in the
three datasets was acquired, among which the top five upregulated
DEMs (hsa-miR-196a, hsa-miR-10b, hsa-miR-196b, hsa-miR-18b and
hsa-miR-542-3p) and the top five downregulated DEMs
(hsa-miR-219-2-3p, hsa-miR-1224-5p, hsa-miR-129-3p, hsa-miR-876-3p
and hsa-miR-770-5p) were determined as being miRNAs associated with
GBM (Fig. 2D). The expression of
the 10 DEMs in each dataset is presented in Fig. 3. Published studies focusing on the
associations between the 10 DEMs and GBM are included in Table II.
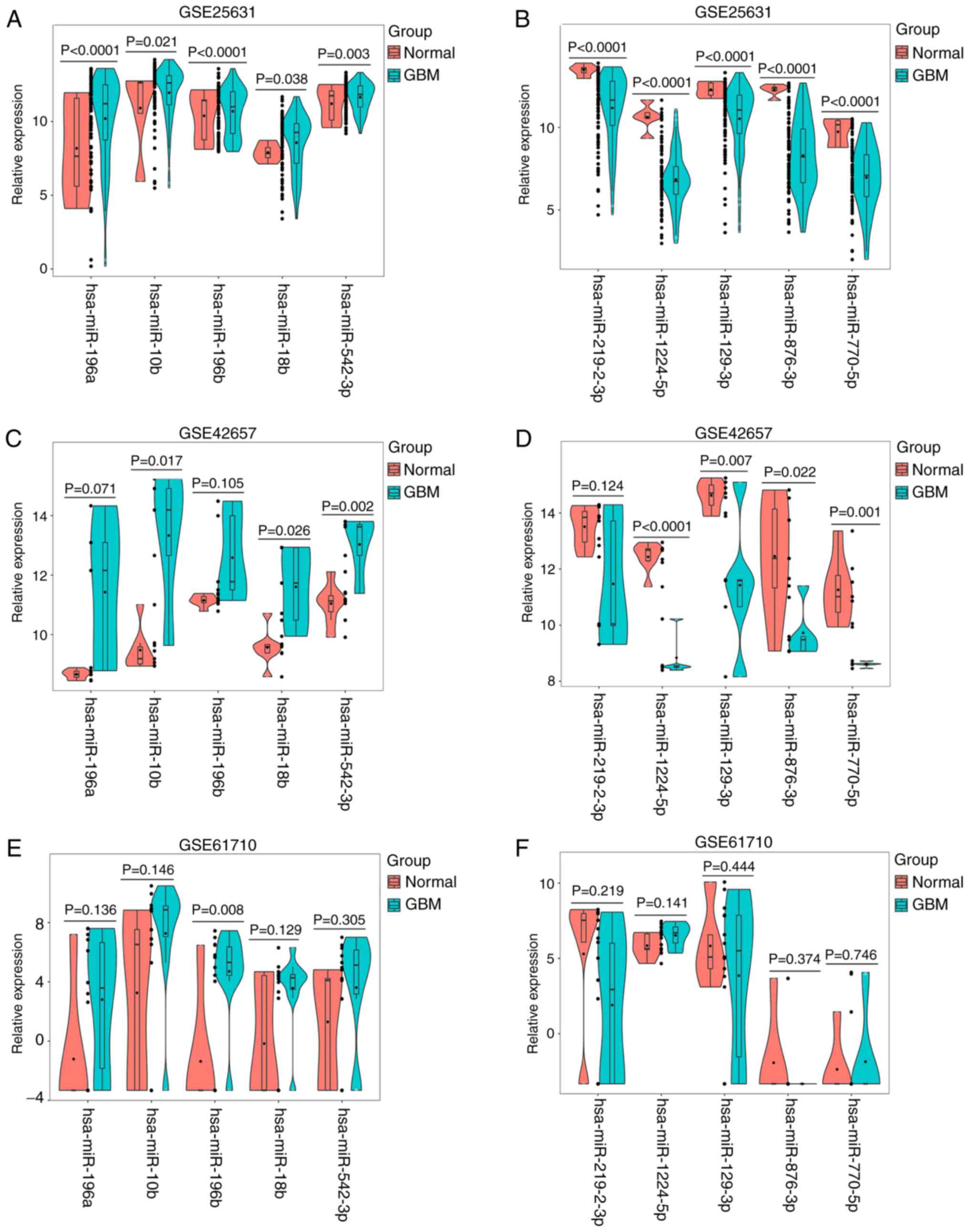 | Figure 3.Violin plots for the expression of
the 10 DEMs in the 3 gene chips. (A) Expression of upregulated
miRNAs (hsa-miR-196a, hsa-miR-10b, hsa-miR-196b, hsa-miR-18b and
hsa-miR-542-3p) in GSE25631. (B) Expression of five downregulated
miRNAs (hsa-miR-219-2-3p, hsa-miR-1224-5p, hsa-miR-129-3p,
hsa-miR-876-3p and hsa-miR-770-5p) in GSE25631. (C) Expression of
upregulated miRNAs (hsa-miR-196a, hsa-miR-10b, hsa-miR-196b,
hsa-miR-18b and hsa-miR-542-3p) in GSE42657. (D) Expression of
downregulated miRNAs (hsa-miR-219-2-3p, hsa-miR-1224-5p,
hsa-miR-129-3p, hsa-miR-876-3p and hsa-miR-770-5p) in GSE42657. (E)
Expression of upregulated miRNAs (hsa-miR-196a, hsa-miR-10b,
hsa-miR-196b, hsa-miR-18b and hsa-miR-542-3p) in GSE61710. (F)
Expression of downregulated miRNAs (hsa-miR-219-2-3p,
hsa-miR-1224-5p, hsa-miR-129-3p, hsa-miR-876-3p and hsa-miR-770-5p)
in GSE61710. DEMs, differentially expressed miRNAs; miR, microRNA;
GBM, glioblastoma multiforme. |
 | Table I.Basic information of the three gene
chips obtained from Gene Expression Omnibus. |
Table I.
Basic information of the three gene
chips obtained from Gene Expression Omnibus.
| First author
(publication year) | Country | Data source | Platform | Sample size
(T/N) | (Refs.) |
|---|
| Zhang et al
(2012) | China | GSE25631 | GPL8179 | 82/5 | (31) |
| Jones et al
(2015) | United Kingdom | GSE42657 | GPL8179 | 5/7 | (32) |
| Piwecka et
al (2015) | Poland | GSE61710 | GPL10656 | 10/5 | (33) |
 | Table II.Five top upregulated miRNAs and five
top downregulated miRNAs from Gene Expression Omnibus. |
Table II.
Five top upregulated miRNAs and five
top downregulated miRNAs from Gene Expression Omnibus.
| Literature
retrieval |
|---|
|
|---|
| Author | miRNA |
RobustRankAggreg | Expression | Function | (Refs.) |
|---|
| Dou et
al | hsa-miR-196a | Up |
| Polymorphism | (37) |
| Yang et
al |
|
| Up | Tumor growth | (38) |
| Guan et
al |
|
| Up | None | (39) |
| Yang et
al |
|
| Up | None | (40) |
| Sasayama et
al | hsa-miR-10b | Up | Up | Cell invasion | (41) |
| Guessous et
al |
|
| Up | Cell invasion, cell
migration, tumor growth | (42) |
| Gabriely et
al |
|
| Up | Cell proliferation,
cell death | (43) |
| Ji et
al |
|
| Up | None | (44) |
| Guan et
al | hsa-miR-196b | Up | Up | None | (45) |
| Lakomy et
al |
|
| Up | None | (46) |
| Ma et
al |
|
| Up | None | (47) |
| You et
al |
|
| Up | None | (48) |
| Karsy M et
al |
|
| Up | None | (49) |
|
| hsa-miR-18b | Up |
| None | No ref. |
| Cai et
al | hsa-miR-543-3p | Up | Down | Cell invasion | (50)a |
|
|
hsa-miR-219-2-3p | Down |
| None | No ref. |
| Qian et
al |
hsa-miR-1224-5p | Down | Down | Cell proliferation,
cell invasion, cell apoptosis | (51) |
| Ouyang et
al | hsa-miR-129-3p | Down | Down | Cell proliferation,
tumor growth | (52) |
| Fang et
al |
|
| Down | Cell viability,
cell growth | (53) |
|
| hsa-miR-876-3p | Down |
| None | No ref. |
|
| hsa-miR-770-5p | Down |
| None | No ref. |
Collection of 390 miRNA-associated
DEGs in GBM
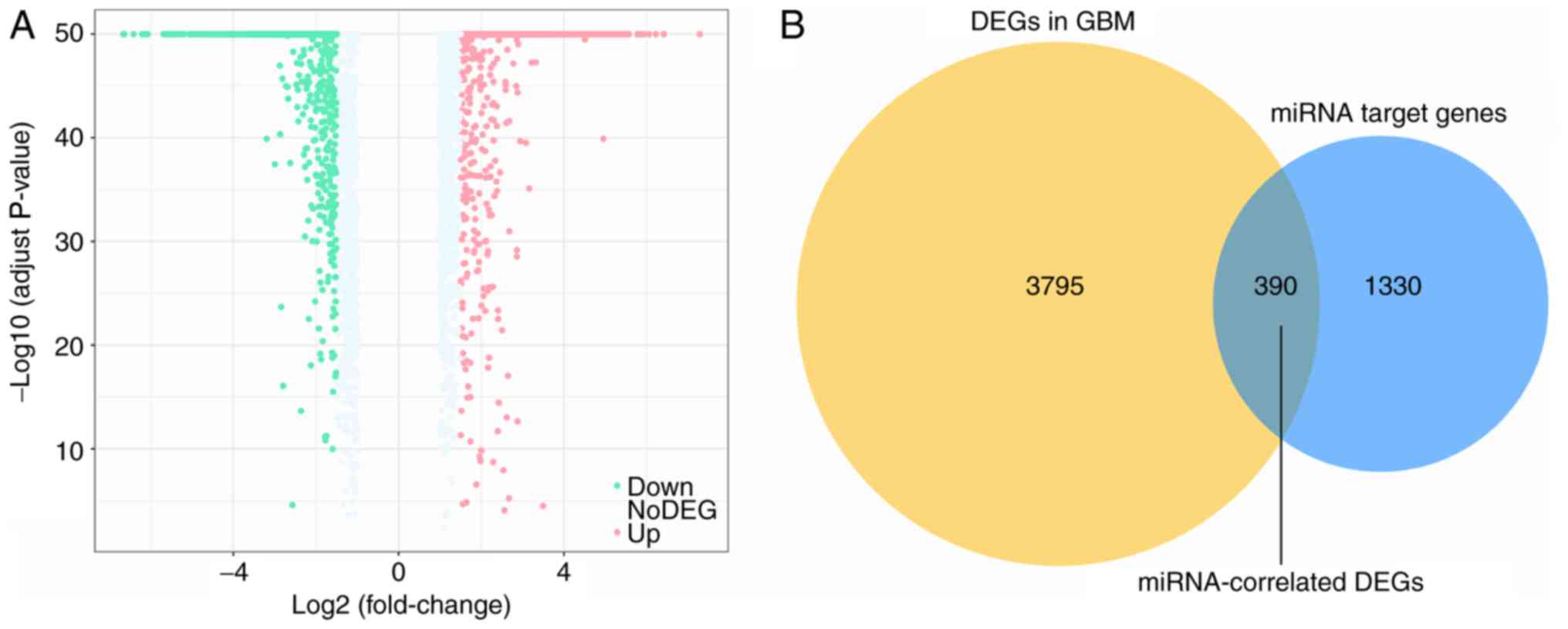
A total of 1,720 genes were identified as targets of
the aforementioned 10 DEMs. Additionally, 4,185 DEGs containing
2,719 upregulated genes and 1,466 downregulated genes in GBM were
collected from GEPIA (Fig. 4A). An
intersection between the 1,720 target genes and the 4,185 DEGs was
performed, revealing 390 overlapping miRNA-associated DEGs
(Fig. 4B).
GO and KEGG enrichment analyses of 390
miRNA-associated DEGs, and PPI analysis of eight GBM-associated
genes
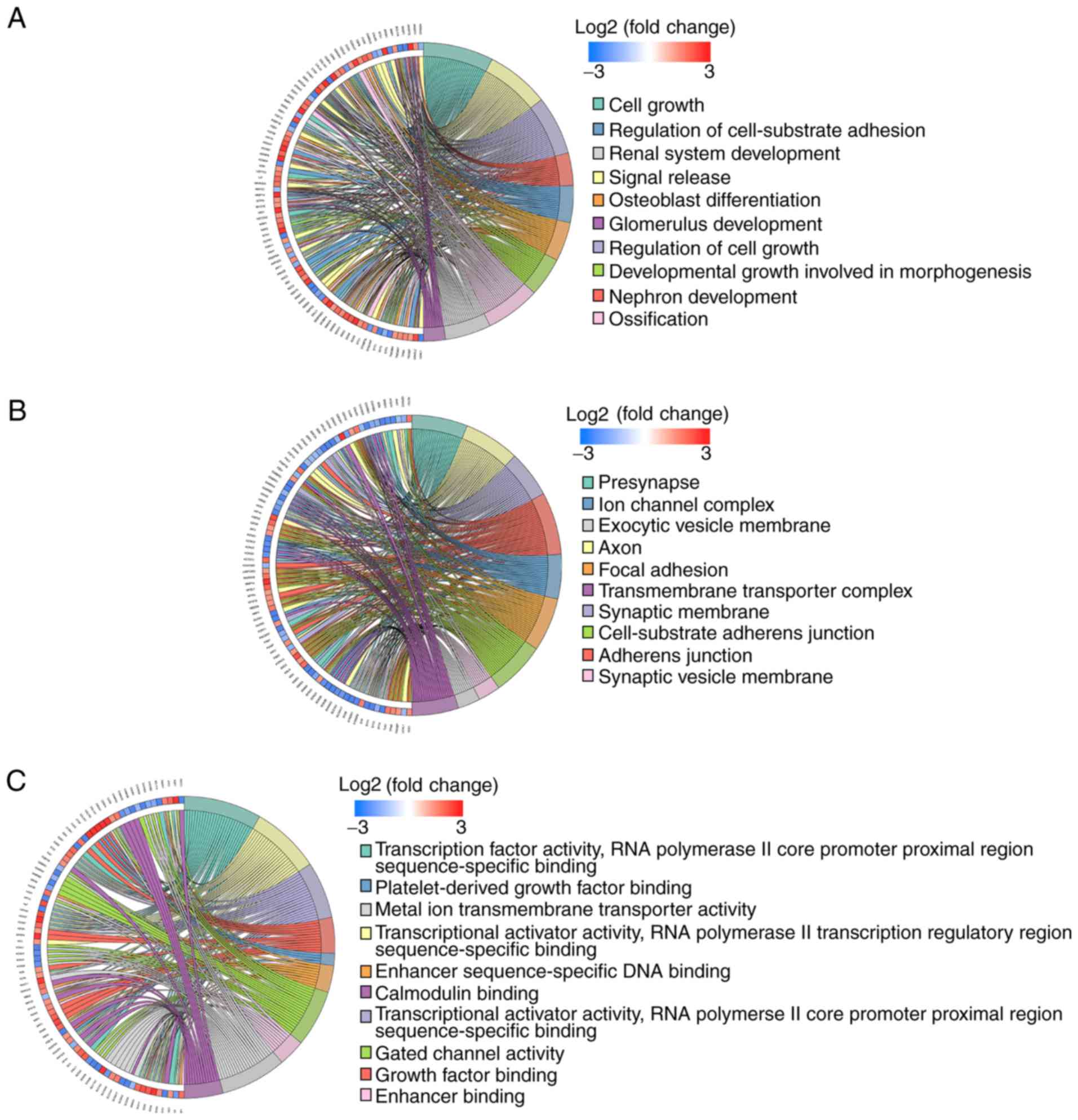
GO functional annotations were performed to
determine the potential molecular mechanisms employed by the 390
miRNA-associated DEGs. The top 10 biological processes (BP),
cellular components (CC) and molecular functions (MF) are listed in
Fig. 5A-C. The highly enriched BP
terms were ‘cell growth’, ‘signal release’ and ‘regulation of cell
growth’. The markedly enriched CC terms were ‘presynapse’, ‘axon’
and ‘synaptic membrane’. The predominantly enriched MF terms were
‘transcription factor activity, RNA polymerase II core promoter
proximal region sequence-specific binding’, ‘transcriptional
activator activity, RNA polymerase II transcription regulatory
region sequence-specific binding’ and ‘transcriptional activator
activity, RNA polymerase II core promoter proximal region
sequence-specific binding’.
The KEGG pathway was utilized to further probe the
underlying pathological pathways that the 390 genes are involved in
during the inception and progression of GBM. With an adjusted value
of P<0.05, 32 pathways were enriched by the 390 genes (Fig. 5D), many of which were
tumor-associated pathways, including the ‘mitogen activated protein
kinase (MAPK) signaling pathway’, the ‘gonadotropin-releasing
hormone (GnRH) signaling pathway’ and the ‘oxytocin signaling
pathway’. In addition, eight genes [cyclin-dependent kinase-6
(CDK6), retinoblastoma-associated protein (RB1),
calcium/calmodulin-dependent protein kinase type II subunit gamma
(CAMK2G), calcium/calmodulin-dependent protein kinase type II
subunit beta (CAMK2B), GTPase NRas (NRAS), protein kinase C beta
type (PRKCB), platelet derived growth factor receptor alpha
(PDGFRA) and calmodulin 3 (CALM3)] were determined to be
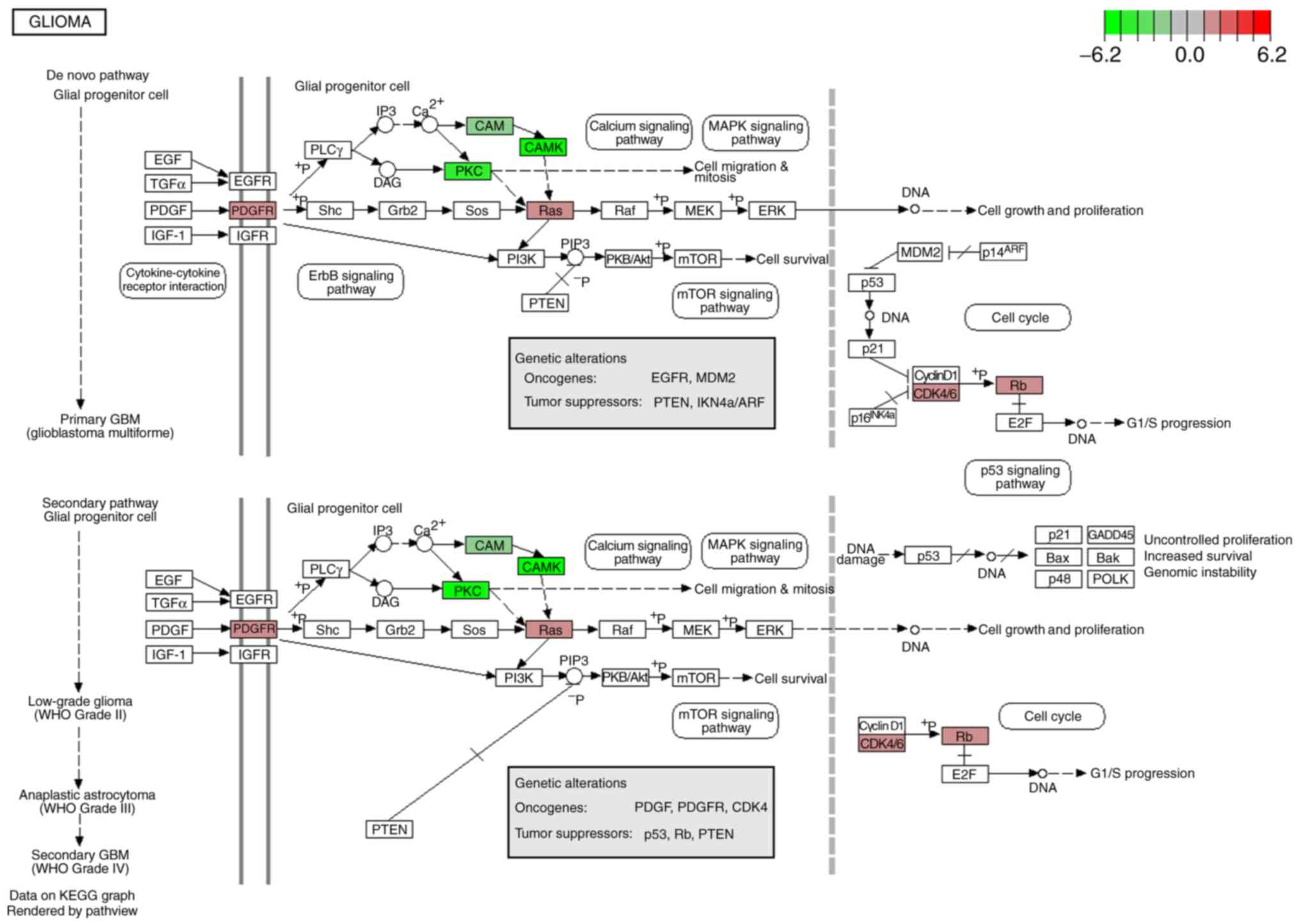
centralized in the glioma pathway (Fig.
6). In this pathway, the eight genes were revealed to
participate in cell growth and proliferation, G1/S progression,
cell migration and mitosis by interacting with upstream or
downstream genes. The expression of the eight genes in GBM and
normal controls were compared based on the data from GEPIA using a
Student's t-test presented as scatter-box plots (Fig. 7). Among them, CDK6, RB1, NRAS, and
PDGFRA were upregulated in GBM, and CAMK2G, CAMK2B, PRKCB, and
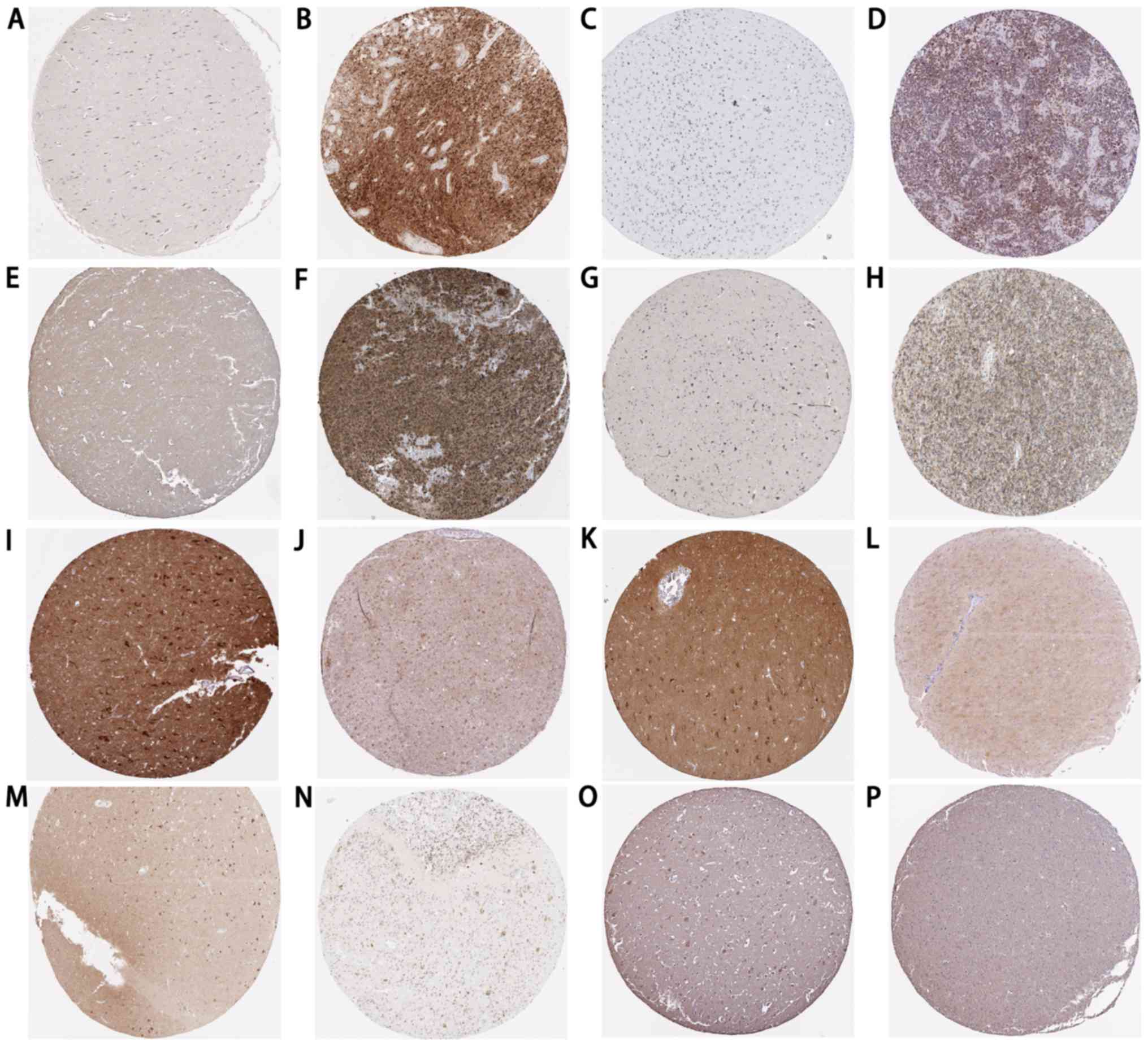
CALM3 were downregulated in GBM tissues. Additionally, the
expression patterns of their encoded proteins in GBM and normal
brain tissues were validated using The Human Protein Atlas
(https://www.proteinatlas.org/) as
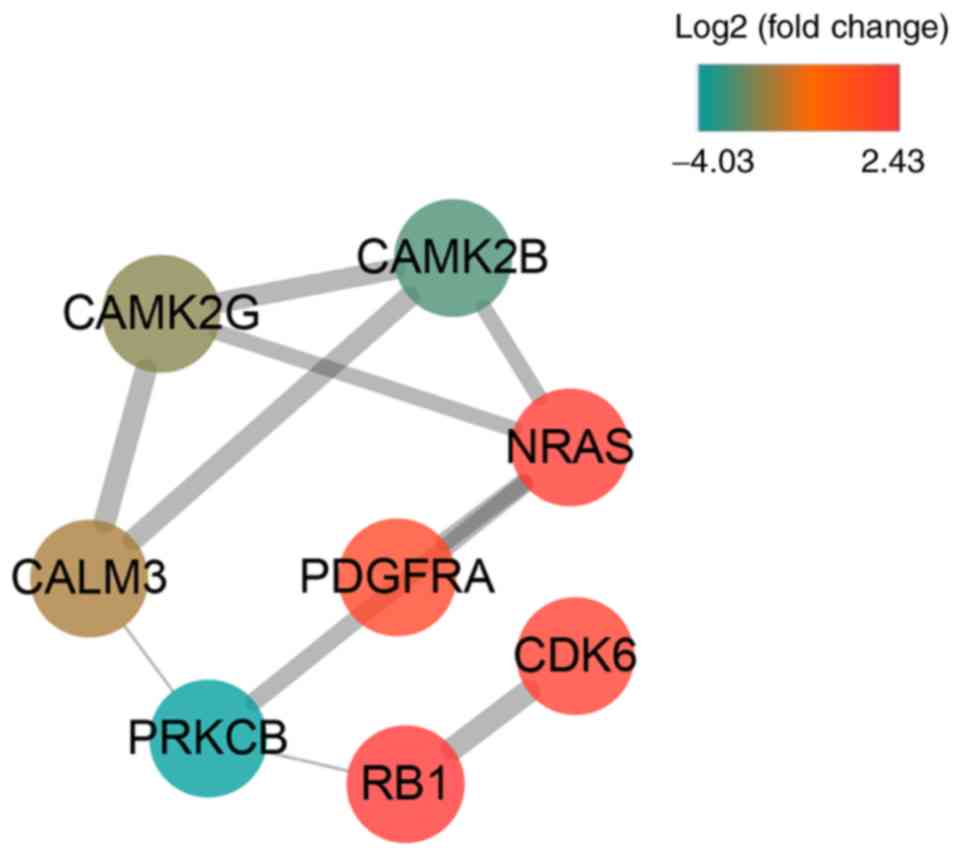
presented in Fig. 8. The PPI
network indicated close associations among the eight genes
(Fig. 9).
 | Figure 7.Box plots for the expression of the
eight genes in glioblastoma multiforme determined from the Gene
Expression Profiling Interactive Analysis: (A) CDK6, (B) RB1, (C)
NRAS, (D) PDGFRA, (E) CAMK2G, (F) CAMK2B, (G) PRKCB and (H) CALM3.
Num, number; T, tumor; N, normal; CDK6, cyclin-dependent kinase 6;
RB1, retinoblastoma-associated protein; NRAS, GTPase NRas; PDGFRA,
platelet-derived growth factor receptor alpha; CAMK2G,
calcium/calmodulin-dependent protein kinase type II subunit gamma;
CAMK2B, calcium/calmodulin-dependent protein kinase type II subunit
beta; PRKCB, protein kinase C beta type; CALM3, calmodulin 3. |
Identification of three potential
chemicals for GBM treatment using CMap analysis
The eight genes, including four that were
upregulated (CDK6, RB1, NRAS and PDGFRA) and four that were
downregulated (CAMK2G, CAMK2B, PRKCB and CALM3) were submitted to
the CMap web tool as up and down tags to acquire latent drugs in
the therapy for GBM. By ranking the connectivity score in
descending order, the top three chemicals (W-13, gefitinib and
exemestane) were identified as being potential treatment options
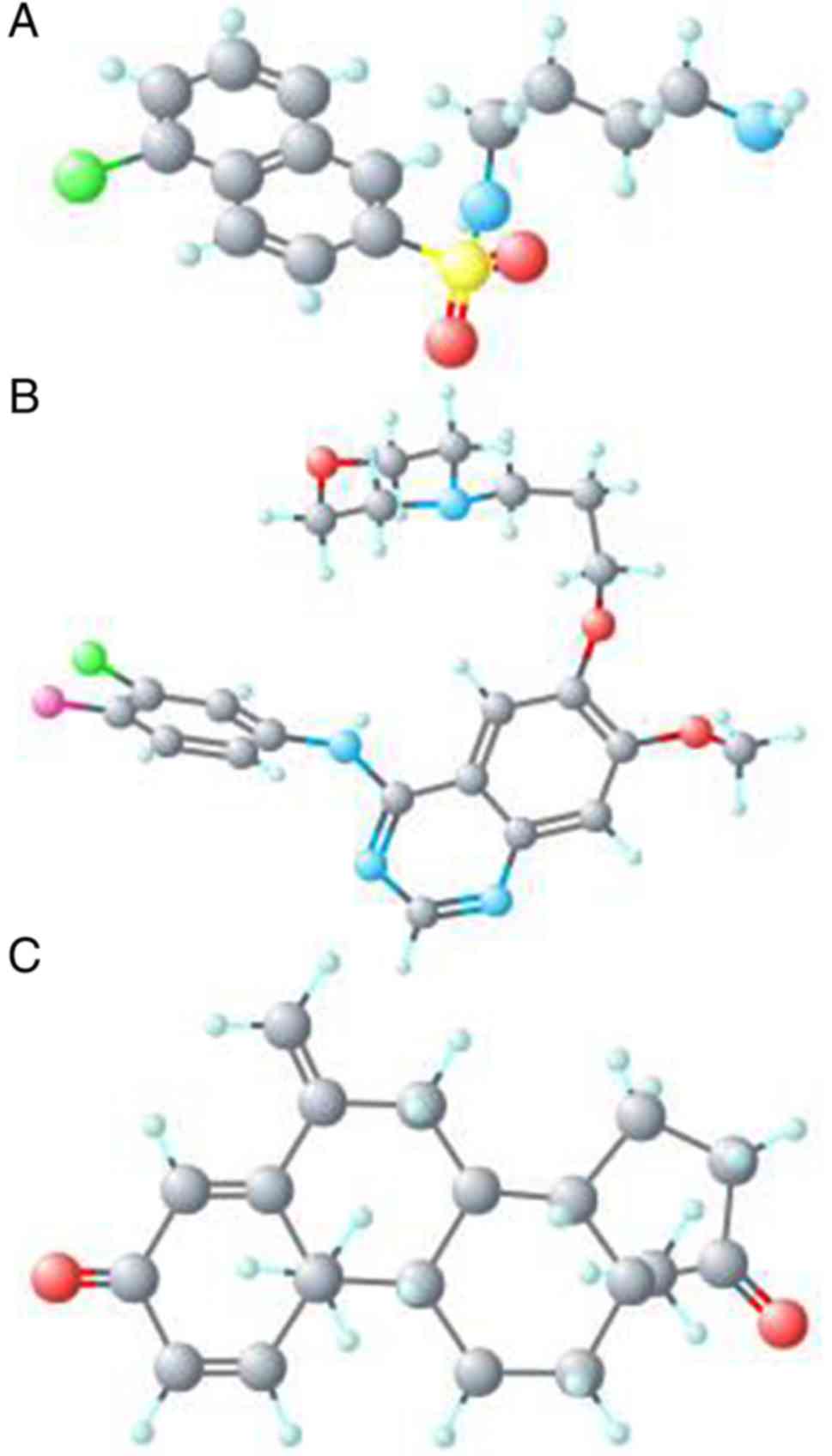
for GBM (Table III). The chemical
structures of the three chemicals are presented in Fig. 10.
 | Table III.Three chemicals identified as
therapeutic agents for glioblastoma multiforme from CMap
analysis. |
Table III.
Three chemicals identified as
therapeutic agents for glioblastoma multiforme from CMap
analysis.
| CMap name | Enrichment | Dose | Cell lines | Up score | Down score |
|---|
| W-13 | −0.989 | 10 µM | MCF7 | −0.356 | 0.532 |
| Gefitinib | −0.989 | 10 µM | HL60 | −0.289 | 0.479 |
| Exemestane | −0.981 | 10 nM | MCF7 | −0.382 | 0.351 |
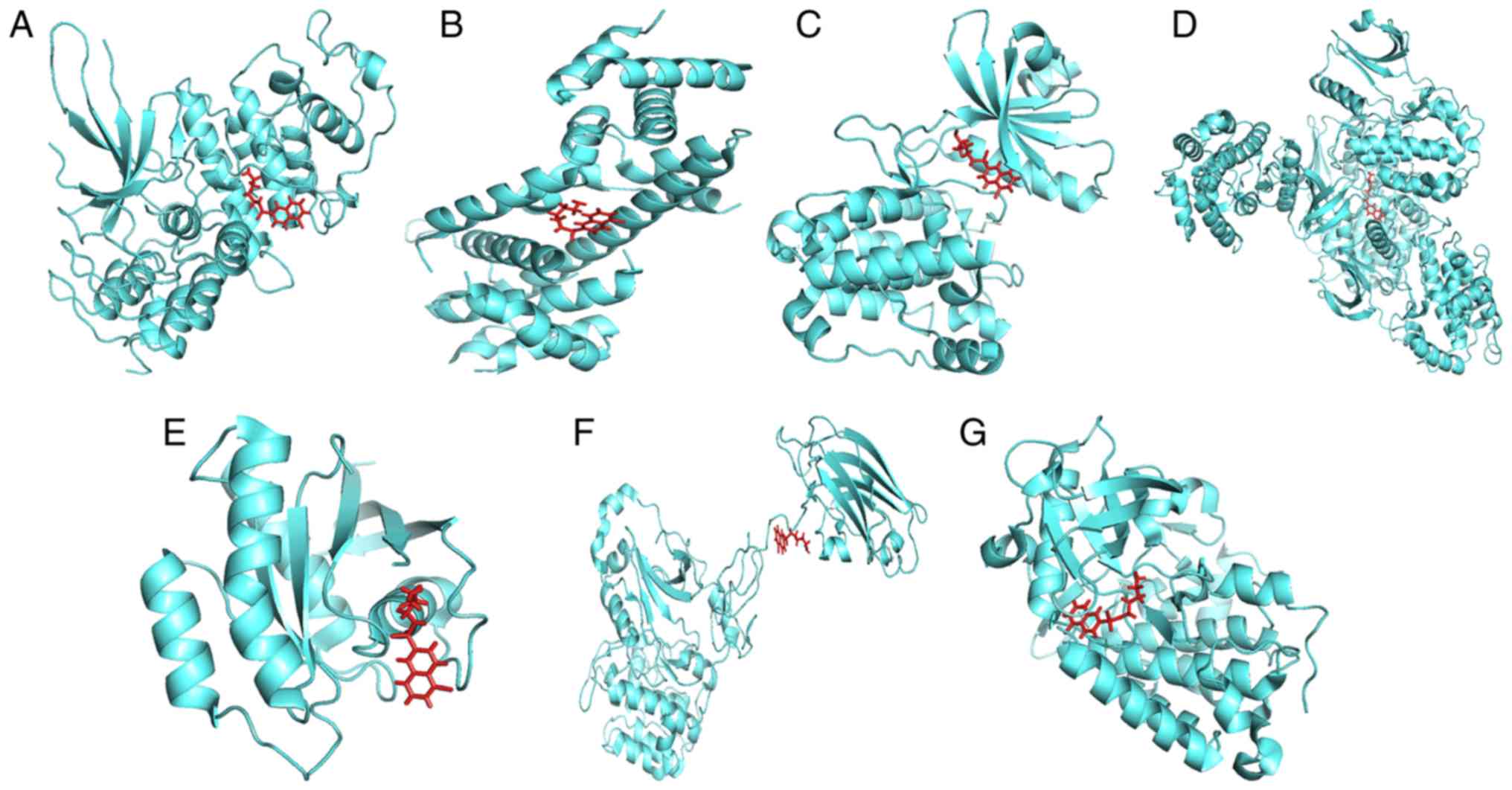
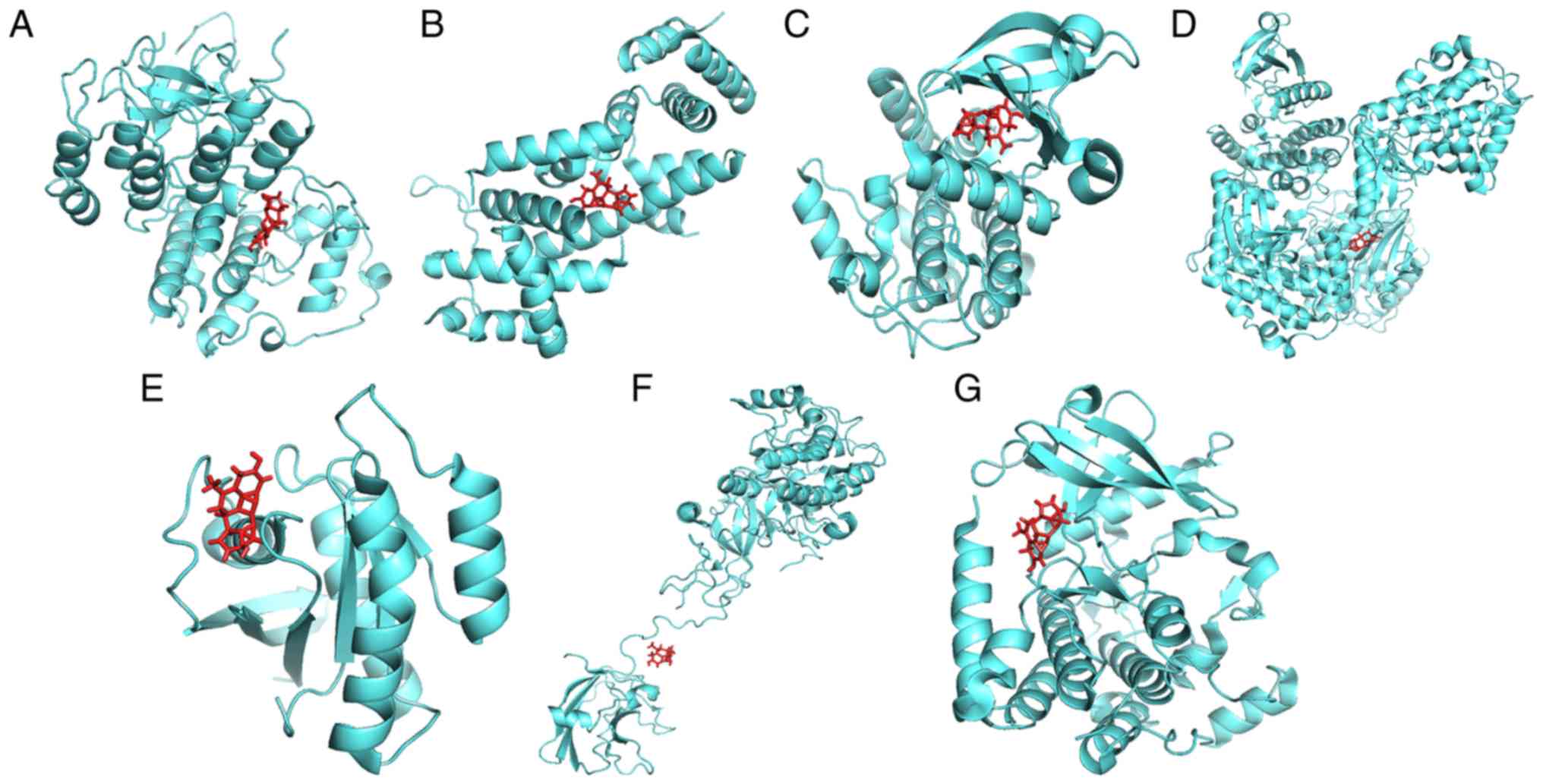
Exploration of the interactions
between the three chemicals and the eight genes
To ascertain whether the three chemicals directly
bind to the proteins encoded by the eight genes, a protein-ligand
docking analysis was performed. As the crystal structure of protein
CALM3 was not identified in the PDB database, it was removed from
the molecular docking analysis. The docking results are presented
in Table IV and Figs. 11–13. The total docking score ranged from
3.5943 to 6.9781, indicating interactions between the three
chemicals and the seven proteins. However, further experiments are
required to verify these associations.
 | Table IV.Results of protein-ligand
docking. |
Table IV.
Results of protein-ligand
docking.
| Gene symbol | Protein name | PDB ID | Chemical | Total score | Crash | Polar |
|---|
| CDK6 | Cyclin-dependent
kinase 6 | 1BI7 | W-13 | 6.9158 | −0.6018 | 1.0852 |
|
|
|
| Gefitinib | 4.5461 | −1.5497 | 1.0374 |
|
|
|
| Exemestane | 4.2617 | −0.6835 | 1.2109 |
| RB1 |
Retinoblastoma-associated protein | 2QDJ | W-13 | 6.1141 | −0.7408 | 2.1589 |
|
|
|
| Gefitinib | 5.1361 | −3.0039 | 1.0429 |
|
|
|
| Exemestane | 4.5673 | −1.2303 | 2.2315 |
| CAMK2G |
Calcium/calmodulin-dependent protein
kinase type II subunit gamma | 2V7O | W-13 | 6.5899 | −0.9631 | 2.1155 |
|
|
|
| Gefitinib | 5.5314 | −0.8239 | 4.6009 |
|
|
|
| Exemestane | 5.5085 | −2.4882 | 2.8500 |
| CAMK2B |
Calcium/calmodulin-dependent protein
kinase type II subunit beta | 3BHH | W-13 | 6.9781 | −0.8004 | 2.9012 |
|
|
|
| Gefitinib | 5.4665 | −1.7396 | 3.6928 |
|
|
|
| Exemestane | 4.7782 | −1.8678 | 4.6573 |
| NRAS | GTPase NRas | 3CON | W-13 | 6.9580 | −0.7966 | 3.5558 |
|
|
|
| Gefitinib | 5.1191 | −1.6695 | 2.1113 |
|
|
|
| Exemestane | 3.7527 | −1.3243 | 1.6902 |
| PRKCB | Protein kinase C
beta type | 3PFQ | W-13 | 5.9087 | −0.7642 | 4.3292 |
|
|
|
| Gefitinib | 5.9886 | −0.9356 | 2.5291 |
|
|
|
| Exemestane | 3.5943 | −0.6226 | 2.3138 |
| PDGFRA | Platelet-derived
growth factor receptor alpha | 5K5X | W-13 | 6.7711 | −1.3263 | 4.9966 |
|
|
|
| Gefitinib | 4.0157 | −1.0638 | 1.1911 |
|
|
|
| Exemestane | 4.8780 | −0.9005 | 2.6185 |
Discussion
Since the identification of the first miRNA, an
increasing number of studies have focused on the action and
clinical application of miRNAs, particularly in terms of neoplasm
treatment (34). It has been
demonstrated that miRNAs participate in multiple biological
processes involved in tumorigenesis (35). With the advantage of targeting genes
that are involved in multiple pathological pathways, drug
developments based on miRNAs have received increasing attention
(36).
The present study obtained 10 DEMs (hsa-miR-196a,
hsa-miR-10b, hsa-miR-196b, hsa-miR-18b, hsa-miR-542-3p,
hsa-miR-219-2-3p, hsa-miR-1224-5p, hsa-miR-129-3p, hsa-miR-876-3p
and hsa-miR-770-5p) using a robust rank aggregation, which is a
recognized method for integrating genes from diverse resources that
are free of outliers, noise and errors (20). Among these literatures, four studies
corroborated overexpressed hsa-miR-196a in GBM and its contribution
to the development of GBM (37–40).
Additionally, four studies indicated that a high expression of
hsa-miR-10b promoted the progression of GBM (41–44).
Five studies focused on hsa-miR-196b and demonstrated its high
expression in GBM (45–49) and one study, performed by Cai et
al (50), focused on
hsa-miR-542-3p. The authors of these studies determined that
hsa-miR-542-3p was downregulated in glioblastoma cell lines, which
was not consistent with the gene chip results from the present
study. Given the differences in sample sources, RNA extraction and
detection, more studies are required to further assess the role of
hsa-miR-542-3p in GBM. Additionally, the decreased expression of
hsa-miR-1224-5p (51) and
hsa-miR-129-3p (52,53), as well as their inhibitory effects
on GBM, have been verified in previous studies. Furthermore, the
association of hsa-miR-18b, hsa-miR-219-2-3p, hsa-miR-876-3p and
hsa-miR-770-5p with GBM have not yet been reported. Since miRNAs
control tumor development by regulating their downstream target
genes (54), the present study
collected the target genes of the aforementioned 10 DEMs to
elucidate how these DEMs mediate the pathophysiological processes
of GBM. The DEGs in GBM were also obtained and an intersection
between the target genes and the DEGs was performed to determine
miRNA-associated DEGs. Gene functional and pathway enrichment
analyses of the miRNA-associated DEGs revealed that these genes
were involved with multiple tumor-associated biological processes
and signaling pathways, including the ‘cell growth’, the ‘MAPK
signaling pathway’, the ‘GnRH signaling pathway’ and the ‘oxytocin
signaling pathway’. Furthermore, eight genes (CDK6, RB1, CAMK2G,
CAMK2B, NRAS, PRKCB, PDGFRA and CALM3) of the miRNA-associated DEGs
were enriched in the glioma pathway, indicating their important
roles in GBM. In the glioma pathway, the eight genes primarily
participated in cell growth and proliferation, G1/S progression,
and cell migration and mitosis, indicating that these genes
participate in the development of GBM by mediating cell
proliferation and metastasis. Each of the eight genes was targeted
by more than one miRNA and one miRNA targeted more than one gene.
For example, the PDGFRA gene was targeted by hsa-miR-770-5p and
hsa-miR-196a. hsa-miR-196a targeted NRAS, PDGFRA and CALM3. miRNAs
primarily exert effects via destabilization or translational
repression by targeting the 3′ untranslated region of mRNA
transcripts in the cytoplasm (7).
However, an increasing number of studies have indicated that miRNAs
positively regulate gene transcription by targeting promoter
elements (55–57). The present study revealed that
certain miRNAs were negatively associated with their target genes
(hsa-miR-1224-5p and CDK6; hsa-miR-196a and CALM3), while other
miRNAs were positively associated with their target genes
(hsa-miR-196a and NRAS; hsa-miR-1224-5p and PRKCB). However,
further studies are required to assess the regulatory mechanisms of
the 10 miRNAs and their target genes.
CMap is a practical tool for the exploration of
novel drugs and for the repurposing of existing drugs, and its
efficiency has been supported by numerous studies (58,59).
Aramadhaka et al (58)
identified Gila monster venom and Byetta® as being
therapeutic drugs for the treatment of type-2 diabetes using CMap
analysis. Wang et al (59)
demonstrated that via cell apoptosis, prenylamine could be a
candidate agent for the treatment of hepatocellular carcinoma. The
present study selected the aforementioned eight genes for CMap
analysis. Following this analysis, three chemicals (W-13, gefitinib
and exemestane) were determined as latent therapeutic agents for
GBM. As a calmodulin antagonist, W-13 has been demonstrated to
inhibit cell growth (60) and to
induce cell apoptosis (61).
However, few studies on W-13 have assessed its anti-GBM effects.
The results from the molecular docking analysis performed in the
present study revealed that W-13 could bind to proteins CDK6, RB1,
NRAS, PDGFRA, CAMK2G, CAMK2B and PRKCB, exhibiting high binding
scores and indicating that W-13 could exert its anti-GBM effects by
acting on these GBM-associated genes. The present study provides a
theoretical basis for the application of W-13 in patients with GBM,
but further studies are required to corroborate this
conclusion.
The inhibitory effect of gefitinib on GBM, which is
an epidermal growth factor receptor (EGFR) tyrosine kinase
inhibitor, has been demonstrated in previous studies (62,63).
However, its clinical application is limited due to gefitinib
resistance (62). A study by
Aljohani et al (64)
revealed that PDGFRA was significantly upregulated in
gefitinib-resistant GBM cells and that overexpressed PDGFRA
regulated gefitinib resistance. The present study identified that
PDGFRA was a target gene of the 10 DEMs, suggesting the potential
of the 10 DEMs in enhancing gefitinib sensitivity in patients with
GBM.
Exemestane is a widely used drug in the prevention
and treatment of breast cancer due to its aromatase inhibitory role
in the production of oestrogen (65–67).
However, to the best of our knowledge, its antitumor effect on GBM
has not yet been elucidated. A study by Kritikou et al
(68) revealed that the combination
of exemestane and erlotinib significantly inhibited EGFR
mitochondrial translocation. The EGFR mitochondrial translocation
event serves important roles in tumor progression (69) and contributes to drug resistance
(70). A study by Dasari et
al (71) indicated that the
inhibition of the EGFR mitochondrial translocation event in GBM may
be a therapeutic strategy. Additionally, the translocation of EGFR
into mitochondria contributes to EGFR inhibitor drug resistance
(70). The combined use of
exemestane and the EGFR inhibitor, gefitinib, in patients with GBM
may therefore increase gefitinib sensitivity by inhibiting the
translocation of EGFR into the mitochondria. However, further
in-depth in vitro and in vivo experiments are
essential to verify the anti-GBM effects and the synergistic
antitumor effects of these compounds.
In the present study, the identification of W-13,
gefitinib and exemestane were made on the basis of the eight
GBM-associated genes (CDK6, RB1, CAMK2G, CAMK2B, NRAS, PRKCB,
PDGFRA and CALM3), which were the target genes of the 10 DEMs
(hsa-miR-196a, hsa-miR-10b, hsa-miR-196b, hsa-miR-18b,
hsa-miR-542-3p, hsa-miR-219-2-3p, hsa-miR-1224-5p, hsa-miR-129-3p,
hsa-miR-876-3p and hsa-miR-770-5p). Thus, the present study
hypothesizes that the 10 DEMs may produce synergistic or
antagonistic effects on the three chemicals by targeting these
genes. However, more experiments are necessary to validate this
conjecture.
In conclusion, by employing an integrated strategy
of data mining and computational biology, the present study
obtained 10 DEMs that may participate in the development of GBM.
Furthermore, three candidate agents (gefitinib, W-13 and
exemestane) in the treatment of GBM were identified following CMap
analysis. Since the results are based on In silico analysis,
further in-depth studies are necessary to add to the validity of
these results.
Supplementary Material
Supporting Data
Acknowledgements
Not applicable.
Funding
No funding was received.
Availability of data and materials
The datasets used and/or analyzed during the present
study are available from the corresponding author on reasonable
request.
Authors' contributions
DDX designed the present study, collected
miRNA-associated datasets, screened differently expressed miRNAs
and wrote the manuscript. WQX collected miRNA-associated datasets
and wrote the manuscript. RQH and YWD performed GO and KEGG
analyses and construed the PPI network. GC performed CMap analysis
and checked all data. DZL designed the experiments and wrote
manuscript. All authors read and approved the final manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Louis DN, Perry A, Reifenberger G, von
Deimling A, Figarella- Branger D, Cavenee WK, Ohgaki H, Wiestler
OD, Kleihues P and Ellison DW: The 2016 World health organization
classification of tumors of the central nervous system: A summary.
Acta Neuropathol. 131:803–820. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Ostrom QT, Gittleman H, Liao P,
Vecchione-Koval T, Wolinsky Y, Kruchko C and Barnholtz-Sloan JS:
CBTRUS statistical report: Primary brain and other central nervous
system tumors diagnosed in the United States in 2010–2014. Neuro
Oncol. 19 (Suppl_5):v1–v88. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Omuro A and DeAngelis LM: Glioblastoma and
other malignant gliomas: A clinical review. JAMA. 310:1842–1850.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Alifieris C and Trafalis DT: Glioblastoma
multiforme: Pathogenesis and treatment. Pharmacol Ther. 152:63–82.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Cloughesy TF, Cavenee WK and Mischel PS:
Glioblastoma: From molecular pathology to targeted treatment. Annu
Rev Pathol. 9:1–25. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Lim M, Xia Y, Bettegowda C and Weller M:
Current state of immunotherapy for glioblastoma. Nat Rev Clin
Oncol. 15:422–442. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Bartel BP: MicroRNAs: Genomics,
biogenesis, mechanism, and function. Cell. 116:281–297. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Dong H, Lei J, Ding L, Wen Y, Ju H and
Zhang X: MicroRNA: Function, detection, and bioanalysis. Chem Rev.
113:6207–6233. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Bracken CP, Scott HS and Goodall GJ: A
network-biology perspective of microRNA function and dysfunction in
cancer. Nat Rev Genet. 17:719–732. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Sumazin P, Yang X, Chiu HS, Chung WJ, Iyer
A, Llobet-Navas D, Rajbhandari P, Bansal M, Guarnieri P, Silva J,
et al: An extensive microRNA-mediated network of RNA-RNA
interactions regulates established oncogenic pathways in
glioblastoma. Cell. 147:370–381. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Godlewski J, Nowicki MO, Bronisz A, Nuovo
G, Palatini J, De Lay M, Van Brocklyn J, Ostrowski MC, Chiocca EA
and Lawler SE: MicroRNA-451 regulates LKB1/AMPK signaling and
allows adaptation to metabolic stress in glioma cells. Mol Cell.
37:620–632. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Hu J, Sun T, Wang H, Chen Z, Wang S, Yuan
L, Liu T, Li HR, Wang P, Feng Y, et al: MiR-215 is induced
post-transcriptionally via HIF-Drosha complex and mediates
glioma-initiating cell adaptation to hypoxia by targeting
KDM1B. Cancer Cell. 29:49–60. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Berindan-Neagoe I, Monroig Pdel C,
Pasculli B and Calin GA: MicroRNAome genome: A treasure for cancer
diagnosis and therapy. CA Cancer J Clin. 64:311–336. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Subramanian A, Narayan R, Corsello SM,
Peck DD, Natoli TE, Lu X, Gould J, Davis JF, Tubelli AA, Asiedu JK,
et al: A next generation connectivity map: L1000 platform and the
first 1,000,000 profiles. Cell. 171:1437–1452.e17. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Qu XA and Rajpal DK: Applications of
connectivity map in drug discovery and development. Drug Discov
Today. 17:1289–1298. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Chien W, Sun QY, Lee KL, Ding LW, Wuensche
P, Torres-Fernandez LA, Tan SZ, Tokatly I, Zaiden N, Poellinger L,
et al: Activation of protein phosphatase 2A tumor suppressor as
potential treatment of pancreatic cancer. Mol Oncol. 9:889–905.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Lamb J: The connectivity map: A new tool
for biomedical research. Nat Rev Cancer. 7:54–60. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Clough E and Barrett T: The gene
expression omnibus database. Methods Mol Biol. 1418:93–110. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW,
Shi W and Smyth GK: limma powers differential expression
analyses for RNA-sequencing and microarray studies. Nucleic Acids
Res. 43:e472015. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Kolde R, Laur S, Adler P and Vilo J:
Robust rank aggregation for gene list integration and
meta-analysis. Bioinformatics. 28:573–580. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Tang Z, Li C, Kang B, Gao G, Li C and
Zhang Z: GEPIA: A web server for cancer and normal gene expression
profiling and interactive analyses. Nucleic Acids Res. 45:W98–W102.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Dweep H, Gretz N and Sticht C: miRWalk
database for miRNA- target interactions. Methods Mol Biol.
1182:289–305. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Yu G, Wang LG, Han Y and He QY:
clusterProfiler: An R package for comparing biological themes among
gene clusters. OMICS. 16:284–287. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Szklarczyk D, Morris JH, Cook H, Kuhn M,
Wyder S, Simonovic M, Santos A, Doncheva NT, Roth A, Bork P, et al:
The STRING database in 2017: Quality-controlled protein-protein
association networks, made broadly accessible. Nucleic Acids Res.
45:D362–D368. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Uhlén M, Fagerberg L, Hallström BM,
Lindskog C, Oksvold P, Mardinoglu A, Sivertsson Å, Kampf C,
Sjöstedt E, Asplund A, et al: Proteomics. Tissue-based map of the
human proteome. Science. 347:12604192015. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Musa A, Ghoraie LS, Zhang SD, Glazko G,
Yli-Harja O, Dehmer M, Haibe-Kains B and Emmert-Streib F: A review
of connectivity map and computational approaches in
pharmacogenomics. Brief Bioinform. 18:9032017. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Guedes IA, de Magalhaes CS and Dardenne
LE: Receptor-ligand molecular docking. Biophys Rev. 6:75–87. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Rose PW, Prlic A, Bi C, Bluhm WF, Christie
CH, Dutta S, Green RK, Goodsell DS, Westbrook JD, Woo J, et al: The
RCSB protein data bank: Views of structural biology for basic and
applied research and education. Nucleic Acids Res. 43:D345–D356.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Kim S, Thiessen PA, Bolton EE, Chen J, Fu
G, Gindulyte A, Han L, He J, He S, Shoemaker BA, et al: PubChem
substance and compound databases. Nucleic Acids Res.
44:D1202–D1213. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Alexander N, Woetzel N and Meiler J:
bcl::Cluster: A method for clustering biological molecules coupled
with visualization in the Pymol Molecular Graphics System. IEEE Int
Conf Comput Adv Bio Med Sci. 2011:13–18. 2011.PubMed/NCBI
|
|
31
|
Zhang W, Zhang J, Hoadley K, Kushwaha D,
Ramakrishnan V, Li S, Kang C, You Y, Jiang C, Song SW, et al:
miR-181d: A predictive glioblastoma biomarker that downregulates
MGMT expression. Neuro Oncol. 14:712–719. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Jones TA, Jeyapalan JN, Forshew T,
Tatevossian RG, Lawson AR, Patel SN, Doctor GT, Mumin MA, Picker
SR, Phipps KP, et al: Molecular analysis of pediatric brain tumors
identifies microRNAs in pilocytic astrocytomas that target the MAPK
and NF-kB pathways. Acta Neuropathol Commun. 3:862015. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Piwecka M, Rolle K, Belter A, Barciszewska
AM, Żywicki M, Michalak M, Nowak S, Naskret-Barciszewska MZ and
Barciszewski J: Comprehensive analysis of microRNA expression
profile in malignant glioma tissues. Mol Oncol. 9:1324–1340. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Kong YW, Ferland-McCollough D, Jackson TJ
and Bushell M: microRNAs in cancer management. Lancet Onco.
13:e249–e258. 2012. View Article : Google Scholar
|
|
35
|
Ling H, Fabbri M and Calin GA: MicroRNAs
and other non-coding RNAs as targets for anticancer drug
development. Nat Rev Drug Discov. 12:847–865. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Garzon R, Marcucci G and Croce CM:
Targeting microRNAs in cancer: Rationale, strategies and
challenges. Nat Rev Drug Discov. 9:775–789. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Dou T, Wu Q, Chen X, Ribas J, Ni X, Tang
C, Huang F, Zhou L and Lu D: A polymorphism of microRNA196a genome
region was associated with decreased risk of glioma in Chinese
population. J Cancer Res Clin Oncol. 136:1853–1859. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Yang G, Han D, Chen X, Zhang D, Wang L,
Shi C, Zhang W, Li C, Chen X, Liu H, et al: MiR-196a exerts its
oncogenic effect in glioblastoma multiforme by inhibition of IκBα
both in vitro and in vivo. Neuro Oncol. 16:652–661. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Guan Y, Chen L, Bao Y, Qiu B, Pang C, Cui
R and Wang Y: High miR-196a and low miR-367 cooperatively correlate
with unfavorable prognosis of high-grade glioma. Int J Clin Exp
Pathol. 8:6576–6588. 2015.PubMed/NCBI
|
|
40
|
Yang JP, Yang JK, Li C, Cui ZQ, Liu HJ,
Sun XF, Geng SM, Lu SK, Song J, Guo CY and Jiao BH: Downregulation
of ZMYND11 induced by miR-196a-5p promotes the progression and
growth of GBM. Biochem Biophys Res Commun. 494:674–680. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Sasayama T, Nishihara M, Kondoh T, Hosoda
K and Kohmura E: MicroRNA-10b is overexpressed in malignant glioma
and associated with tumor invasive factors, uPAR and RhoC. Int J
Cancer. 125:1407–1413. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Guessous F, Alvarado-Velez M,
Marcinkiewicz L, Zhang Y, Kim J, Heister S, Kefas B, Godlewski J,
Schiff D, Purow B and Abounader R: Oncogenic effects of miR-10b in
glioblastoma stem cells. J Neurooncol. 112:153–163. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Gabriely G, Teplyuk NM and Krichevsky AM:
Context effect: microRNA-10b in cancer cell proliferation, spread
and death. Autophagy. 7:1384–1386. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Ji Y, Wei Y, Wang J, Gong K, Zhang Y and
Zuo H: Correlation of microRNA-10b upregulation and poor prognosis
in human gliomas. Tumour Biol. 36:6249–6254. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Guan Y, Mizoguchi M, Yoshimoto K, Hata N,
Shono T, Suzuki SO, Araki Y, Kuga D, Nakamizo A, Amano T, et al:
MiRNA-196 is upregulated in glioblastoma but not in anaplastic
astrocytoma and has prognostic significance. Clin Cancer Res.
16:4289–4297. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Lakomy R, Sana J, Hankeova S, Fadrus P,
Kren L, Lzicarova E, Svoboda M, Dolezelova H, Smrcka M, Vyzula R,
et al: MiR-195, miR-196b, miR-181c, miR-21 expression levels and
O−6-methylguanine-DNA methyltransferase methylation status
are associated with clinical outcome in glioblastoma patients.
Cancer Sci. 102:2186–2190. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Ma R, Yan W, Zhang G, Lv H, Liu Z, Fang F,
Zhang W, Zhang J, Tao T, You Y, et al: Upregulation of miR-196b
confers a poor prognosis in glioblastoma patients via inducing a
proliferative phenotype. PLoS One. 7:e380962012. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
You G, Yan W, Zhang W, Wang Y, Bao Z, Li
S, Li S, Li G, Song Y, Kang C, et al: Significance of miR-196b in
tumor-related epilepsy of patients with gliomas. PLoS One.
7:e462182012. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Karsy M, Arslan E and Moy F: Current
progress on understanding MicroRNAs in glioblastoma multiforme.
Genes Cancer. 3:3–15. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Cai J, Zhao J, Zhang N, Xu X, Li R, Yi Y,
Fang L, Zhang L, Li M, Wu J, et al: MicroRNA-542-3p suppresses
tumor cell invasion via targeting AKT pathway in human astrocytoma.
J Biol Chem. 290:24678–24688. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Qian J, Li R, Wang YY, Shi Y, Luan WK, Tao
T, Zhang JX, Xu YC and You YP: MiR-1224-5p acts as a tumor
suppressor by targeting CREB1 in malignant gliomas. Mol Cell
Biochem. 403:33–41. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Ouyang Q, Chen G, Zhou J, Li L, Dong Z,
Yang R, Xu L, Cui H, Xu M and Yi L: Neurotensin signaling
stimulates glioblastoma cell proliferation by upregulating c-Myc
and inhibiting miR-29b-1 and miR-129-3p. Neuro Oncol. 18:216–226.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Fang DZ, Wang YP, Liu J, Hui XB, Wang XD,
Chen X and Liu D: MicroRNA-129-3p suppresses tumor growth by
targeting E2F5 in glioblastoma. Eur Rev Med Pharmacol Sci.
22:1044–1050. 2018.PubMed/NCBI
|
|
54
|
Tutar Y: miRNA and cancer; computational
and experimental approaches. Curr Pharm Biotechnol. 15:4292014.
View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Matsui M, Chu Y, Zhang H, Gagnon KT,
Shaikh S, Kuchimanchi S, Manoharan M, Corey DR and Janowski BA:
Promoter RNA links transcriptional regulation of inflammatory
pathway genes. Nucleic Acids Res. 41:10086–10109. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Majid S, Dar AA, Saini S, Yamamura S,
Hirata H, Tanaka Y, Deng G and Dahiya R: MicroRNA-205-directed
transcriptional activation of tumor suppressor genes in prostate
cancer. Cancer. 116:5637–5649. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Xiao M, Li J, Li W, Wang Y, Wu F, Xi Y,
Zhang L, Ding C, Luo H, Li Y, et al: miRNA and cancer;
computational and experimental approaches. MicroRNAs activate gene
transcription epigenetically as an enhancer trigger. RNA Biol.
14:1326–1334. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Aramadhaka LR, Prorock A, Dragulev B, Bao
Y and Fox JW: Connectivity maps for biosimilar drug discovery in
venoms: The case of Gila monster venom and the anti-diabetes drug
Byetta®. Toxicon. 69:160–167. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
59
|
Wang J, Li M, Wang Y and Liu X:
Integrating subpathway analysis to identify candidate agents for
hepatocellular carcinoma. Onco Targets Ther. 9:1221–1230. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
60
|
Strobl JS and Peterson VA:
Tamoxifen-resistant human breast cancer cell growth: Inhibition by
thioridazine, pimozide and the calmodulin antagonist, W-13. J
Pharmacol Exp Ther. 263:186–193. 1992.PubMed/NCBI
|
|
61
|
Takadera T and Ohyashiki T: Calmodulin
inhibitor-induced apoptosis was prevented by glycogen synthase
kinase-3 inhibitors in PC12 cells. Cell Mol Neurobiol. 27:783–790.
2007. View Article : Google Scholar : PubMed/NCBI
|
|
62
|
Mu L, Wang T, Chen Y, Tang X, Yuan Y and
Zhao Y: β-Elemene enhances the efficacy of gefitinib on
glioblastoma multiforme cells through the inhibition of the EGFR
signaling pathway. Int J Oncol. 49:1427–1436. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
63
|
Parker JJ, Dionne KR, Massarwa R, Klaassen
M, Foreman NK, Niswander L, Canoll P, Kleinschmidt-Demasters BK and
Waziri A: Gefitinib selectively inhibits tumor cell migration in
EGFR-amplified human glioblastoma. Neuro Oncol.
15:1048–1057. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
64
|
Aljohani H, Koncar RF, Zarzour A, Park BS,
Lee SH and Bahassi el M: ROS1 amplification mediates resistance to
gefitinib in glioblastoma cells. Oncotarget. 6:20388–20395. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
65
|
Barton MK: Exemestane is effective for the
chemoprevention of breast cancer. CA Cancer J Clin. 61:363–364.
2011. View Article : Google Scholar : PubMed/NCBI
|
|
66
|
Pagani O, Regan MM and Francis PA:
Exemestane with ovarian suppression in premenopausal breast cancer.
N Engl J Med. 371:1358–1359. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
67
|
Van Asten K, Neven P, Lintermans A,
Wildiers H and Paridaens R: Aromatase inhibitors in the breast
cancer clinic: Focus on exemestane. Endocr Relat Cancer.
21:R31–R49. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
68
|
Kritikou I, Giannopoulou E, Koutras AK,
Labropoulou VT and Kalofonos HP: The combination of antitumor
drugs, exemestane and erlotinib, induced resistance mechanism in
H358 and A549 non-small cell lung cancer (NSCLC) cell lines. Pharm
Biol. Nov 5–2013.(Epub ahead of print). PubMed/NCBI
|
|
69
|
Che TF, Lin CW, Wu YY, Chen YJ, Han CL,
Chang YL, Wu CT, Hsiao TH, Hong TM and Yang PC: Mitochondrial
translocation of EGFR regulates mitochondria dynamics and promotes
metastasis in NSCLC. Oncotarget. 6:37349–37366. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
70
|
Cao X, Zhu H, Ali-Osman F and Lo HW: EGFR
and EGFRvIII undergo stress- and EGFR kinase inhibitor-induced
mitochondrial translocalization: A potential mechanism of
EGFR-driven antagonism of apoptosis. Mol Cancer. 10:262011.
View Article : Google Scholar : PubMed/NCBI
|
|
71
|
Dasari VR, Velpula KK, Alapati K, Gujrati
M and Tsung AJ: Cord blood stem cells inhibit epidermal growth
factor receptor translocation to mitochondria in glioblastoma. PLoS
One. 7:e318842012. View Article : Google Scholar : PubMed/NCBI
|