Introduction
Colorectal cancer (CRC) is one of the most commonly
diagnosed cancer types, and it accounts for 10% of all cancer
cases, with an occurrence of >1.8 million cases every year
worldwide (1). Treatment strategies
involving surgery, chemotherapy and radiotherapy have increased the
overall survival rates for patients with early stage CRC (2,3). However,
~60% of patients with CRC develop metastasis, which is the major
cause of mortality in patients with CRC (4,5). Thus, it
is important to elucidate the mechanisms underlying the metastasis
of CRC, and to investigate new CRC treatments based on the
discoveries.
The activation of pro-oncogenes KRAS, c-Src
and c-Myc, and inactivation of tumor suppressor genes, such
as the loss of adenomatous polyposis coli (APC) and tumor
suppressor p53 (TP53), all contribute to CRC tumorigenesis
and development (2). Exosomes, as
mediators of intercellular communication, have been reported to be
involved in CRC progression (6,7). Exosomes
are extracellular vesicles (EVs) with a diameter of 30–180 nm,
secreted by various cells (8–10). A recent study demonstrated that
exosomes derived from the HCT-8 CRC cell line contributed to the
EMT process via miR-210 (6). Exosomes
derived from CRC cells (HT29, SW480 and SW620) were revealed to
increase the migration and invasion capacities of colonic
mesenchymal cells (7,11). In addition, drug resistance during
tumor treatment could also be attributed to exosomes. miR-145 and
miR-34a released via exosomes are hypothesized to enhance the
5-fluorouracil resistance in DLD-1 cells (12). Epidermal growth factor receptor has
been identified to be expressed on the surface of exosomes and was
able to bind to Cetuximab, which reduced its therapeutic efficacy
in Caco2 and HCT-116 cells (13).
Exosomes form as intraluminal vesicles (ILVs) by
budding into early endosomes and multi-vesicular bodies (MVBs).
Subsequently, MVBs are released upon fusion with the plasma
membrane (PM) (14,15). The SNARE complex may serve a role in
the fusion of MVBs with the PM. However, the underlying mechanism
remains to be fully elucidated (16).
On the other hand, the intracellular adaptor protein syntenin,
endosomal sorting complex required for transport (ESCRT),
heparanase and tetraspanins demonstrate key roles in the loading of
exosomes (7). In addition,
sumoylation is considered to be a mechanism involved in the loading
of miRNAs into exosomes (17).
Ca2+-dependent activator protein for
secretion 1 (CAPS1) is characterized as a 145-kDa brain protein
containing a central pleckstrin homology domain, a Munc-13 homology
(MH) domain, a C-terminal domain, a C-terminal membrane-association
domain and a C2 domain (18). As a
special regulator of the Ca2+-dependent large dense-core
vesicle fusion process, CAPS1 promotes the assembly of the SNARE
complex via the MH domain, which regulates vesicle exocytosis
(19,20). Our previous study demonstrated that
CAPS1 was upregulated in CRC, and that increased CAPS1 expression
was associated with frequent metastasis and poor prognosis in
patients with CRC (18). Furthermore,
CAPS1 was revealed to promote CRC cell migration and invasion in
vitro and facilitate CRC liver metastasis in vivo
(18). However, whether exosomes are
required for CAPS1-induced CRC metastasis requires further
investigation.
BMP4 is a member of bone morphogenetic proteins
(BMPs), which are multi-functional cytokines belonging to the
transforming growth factor-β (TGF-β) family (21). Previous studies suggest that BMP4 is
closely associated with tumorigenesis (22,23). In
CRC, BMP4 was revealed to be frequently upregulated due to aberrant
activation of Wnt-β-catenin signaling, and promoted cell migration
and invasion (22,24). Knockdown of BMP4 inhibited tumor
formation of CRC cells in vivo through apoptosis induction
(22). The expression of BMP4 was
significantly increased in hepatocellular carcinoma (HCC) tissues
(25). Increased BMP4 was correlated
with high metastasis of HCC cells (25). BMP4 facilitated HCC cell invasion and
metastasis though ID2-mediated EMT and promoted HCC cell
proliferation via autophagy activation (23,25). In
breast cancer, BMP4 promoted cell migration and invasion possibly
via induction of MMP-1 and CXCR4 expression (26). The function of BMP4 appears to be
divergent but with clear evidence supporting tumor suppressing
functions in lung squamous cell carcinoma (SQC). For example,
decreased BMP4 induced by SOX2 enhanced lung SQC cell growth
(27). This finding indicated a
tissue context specific role of BMP4.
The present study revealed that CAPS1 promoted FHC
cell migration by altering the protein expression profile of
exosomes derived from CRC cells. GW4869, an exosome inhibitor,
inhibited CAPS1-induced cell migration.
Materials and methods
Cell culture and conditioned medium
preparation
The cell lines, HT29, SW480, FHC and 293T, used in
the present study were purchased from The American Type Culture
Collection. The cells were cultured in complete DMEM containing 10%
fetal bovine serum (FBS; Gibco; Thermo Fisher Scientific, Inc.),
100 mg/ml penicillin, and 10 mg/ml streptomycin (Gibco; Thermo
Fisher Scientific, Inc.), at 37°C and 5% CO2.
Conditioned medium (CM) was harvested at 48 h from confluent
cultures with exosome-depleted medium and centrifuged at 1,400 × g
for 2 min at 4°C to remove cellular debris. To inhibit exosome
secretion, cells were treated with 10 µM GW4869 (MedChemExpress)
before collecting the CM.
Exosome isolation and
characterization
Exosomes were isolated from HT29/SW480 CM by serial
centrifugation. The medium was subjected to ultracentrifugation at
100,000 × g for 6 h at 4°C and washed with PBS (100,000 × g for 20
min) (7,28). Subsequently, the exosomes were
re-suspended in PBS. The presence of exosomes was confirmed by
particle size with a Nanoparticle Tracking Analysis (NTA) system
(NTA 3.2 Dev Build 3.2.16, Malvern Panalytical Ltd., Malvern, UK),
and the expression of exosome-specific markers such as tumor
susceptibility gene 101 protein (TSG101) and CD81 was evaluated by
western blot analysis.
Electron microscopy
For electron microscopy, exosomes were fixed with 2%
paraformaldehyde and loaded on carbon-coated copper grids. The
grids were placed on 2% gelatin for 20 min at 37°C and washed with
0.15 M glycine in PBS. Subsequently, the sections were blocked with
1% cold water fish-skin gelatin (11,29,30). The
grids were viewed under a Philips CM120 transmission electron
microscope (Philips Research).
Exosome uptake assay
The exosomes were fluorescently labeled using an
ExoGlow-Protein EV Labeling kit (System Biosciences), according to
the manufacturer's instructions. Approximately 100–500 µg of
labeled exosomes were added to 1×105 293T/FHC cells. The
red fluorescent signal was observed at 24 and 48 h.
Cell migration assay
Transwell migration assays were performed in 24-well
chambers (pore size, 0.8 µm; EMD Milipore). FHC cells were treated
with HT29 cell-derived exosomes, HT29/SW480-CM or
HT29/SW480-CM+GW4869 for 24 h. FHC cells were then re-suspended in
FBS-free DMEM and added to the upper chamber. DMEM containing 10%
FBS was added to the lower chamber. The cells were incubated for 5
days, and subsequently, the cells on the undersurface of the
membrane were fixed by 4% paraformaldehyde for 30 min at room
temperature and stained with 0.1% crystal violet overnight at room
temperature. The cells that had migrated through the filter were
counted in four randomly selected regions per filter.
Western blot analysis
The cells or exosomes were homogenized in RIPA lysis
buffer supplemented with protease inhibitors (Roche Diagnostics).
The concentration of proteins was measured using the BCA Protein
assay kit (Beyotime Institute of Biotechnology). Western blot
analysis was performed as previously described (18). A total of 40 µg proteins per lane were
loaded. The cell or exosome lysates were immunoblotted with
antibodies against CAPS1 (cat. no. ab32011; Abcam), β-actin (cat.
no. ab8227), CD63 (cat. no. ab59479), CD81 (cat. no. ab35026),
TSG101 (cat. no. ab133586; all from Abcam) or BMP4 (cat. no. A0425;
ABclonal).
Quantitative proteomics and
bioinformatics analyses of exosomal proteins
Proteins were extracted from the isolated exosomes,
digested with trypsin, and analyzed by LC-MS. The mass spectra were
searched against the Swiss-Prot Human Proteome database (https://www.uniprot.org/). Gene Ontology (GO)
(http://www.omicsbean.com:88/) and
Ingenuity Pathway Analysis (IPA) analysis software were used to
analyze differentially expressed proteins.
Statistical analysis
The data are presented as the means ± standard
deviation. GraphPad Prism 7 software (GraphPad Prism, Inc.) was
used for statistical analysis. Student's t-test was used for
comparisons between groups. P<0.05 (two-sided) was considered to
indicate a statistically significant difference.
Results
Characterization of exosomes isolated
from CAPS1-overexpressing CRC cells
Our previous study indicated that CAPS1 was
upregulated in CRC, and that increased CAPS1 facilitated CRC
metastasis (18). The previous study
generated stable CAPS1-overexpressing HT29 and SW480 cell lines
(CAPS1-HT29 and CAPS1-SW480) by lentiviral infection, as well as
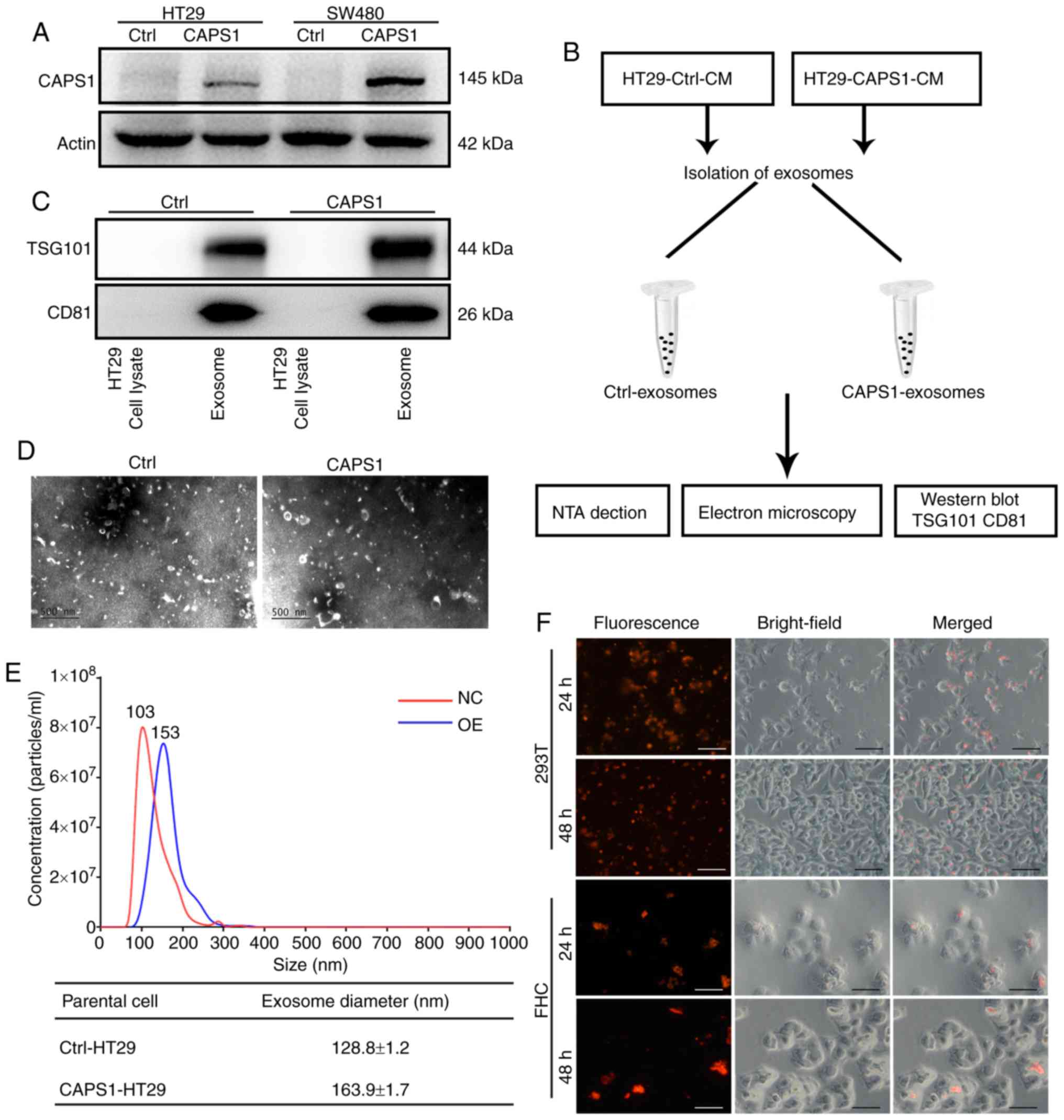
corresponding control cell lines (18). The protein level of CAPS1 was detected
in CAPS1-HT29 and CAPS1-SW480 cells in the present study (Fig. 1A). In order to confirm the role of
exosomes in the process of CAPS1-induced CRC metastasis, exosomes
were isolated from the CM of CAPS1-overexpressing HT29 cells
(CAPS1-HT29-CM) and control cells (Fig.
1B). The harvested exosomes were detected by the presence of
exosome-specific markers, TSG101 and CD81, using western blot
analysis (Fig. 1C). The morphology of
the exosomes was characterized by transmission electron microscopy,
which revealed no noticeable differences (Fig. 1D). The particles of exosomes were
measured by NTA system, revealing a diameter range of 30–180 nm. In
comparison, exosomes derived from CAPS1-overexpressing cells
exhibited a larger mean diameter. (Fig.
1E). To investigate whether HT29-derived exosomes (HT29-exo)
could be internalized by target cells, the exosomes were
pre-labeled fluorescently and co-cultured with 293T and FHC cells.
Red fluorescence was observed within target cells at 24 and 48 h,
indicating that exosomes were endocytosed successfully by target
cells (Fig. 1F).
Exosomes derived from
CAPS1-overexpressing CRC cells enhance FHC migration
To investigate whether CAPS1 promotes CRC metastasis
via exosomes. CAPS1-HT29-derived exosomes were co-cultured with FHC
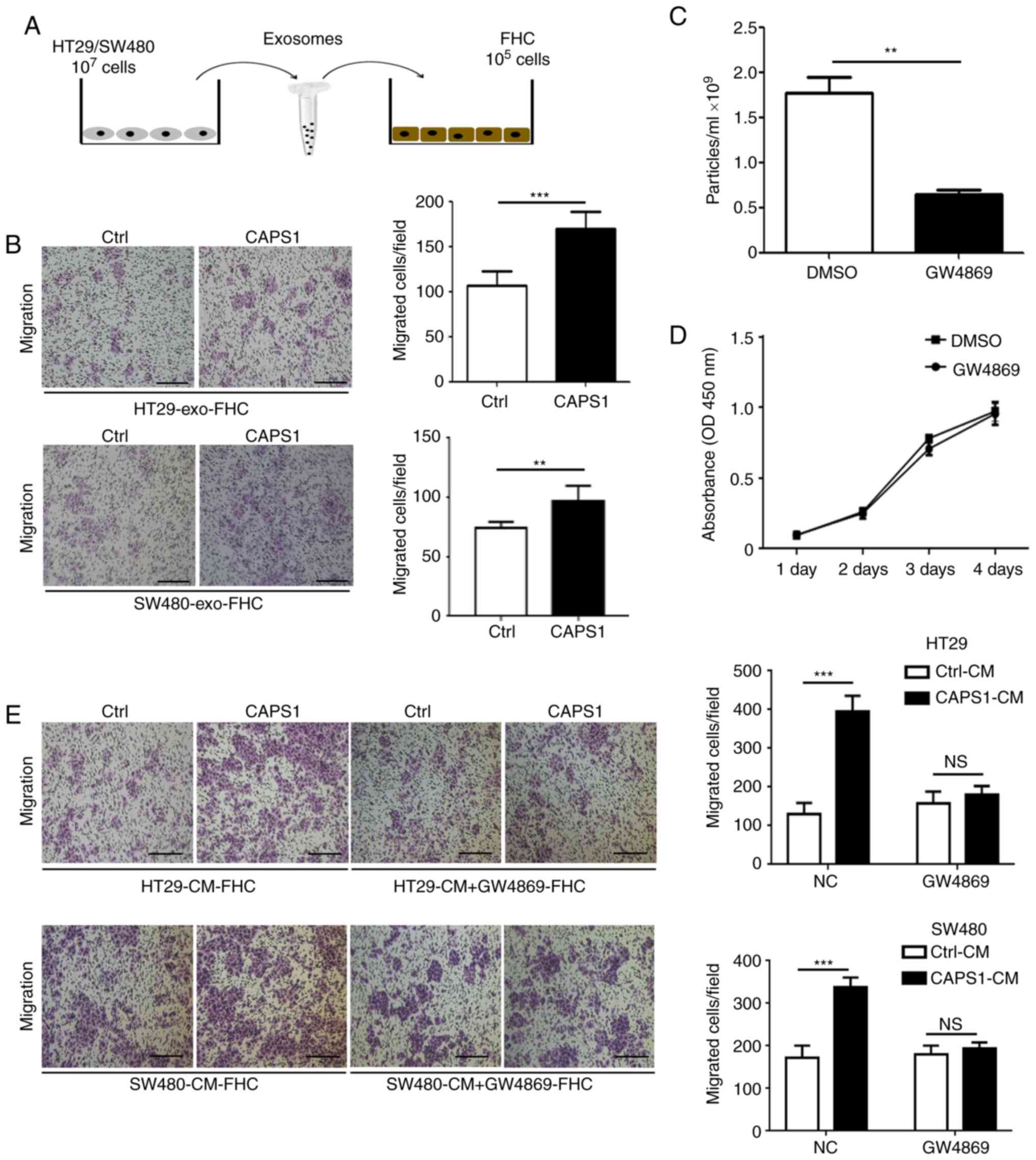
cells for 24 h (Fig. 2A). The FHC
cell line is from normal fetal colonic mucosa, which exhibits a
tumorigenic phenotype and has a metastatic potential in vivo
(31,32). A Transwell migration assay was
performed to detect the migration of FHC cells, which demonstrated
that exosomes derived from CAPS1-overexpressing CRC cells (HT29 and
SW480) significantly enhanced the migration of FHC cells (HT29,
P<0.001 SW480, P=0.0067; Fig.
2B).
To further investigate whether CRC cell-derived
exosomes participate in FHC cell migration, GW4869, an inhibitor of
exosomes, was used to inhibit exosome secretion in HT29 cells
(P=0.0038; Fig. 2C). It was
determined that GW4869 had no effect on HT29 cell viability
(P=0.4652; Fig. 2D). FHC cells were
cultured with CM derived from CAPS1-overexpressing CRC cells (HT29
and SW480), as well as control cells, for 24 h. CM derived from
CAPS1-overexpressing cells significantly increased FHC migration in
the Transwell migration assay (HT29, P<0.001; SW480,
P<0.001), an effect that was attenuated following
GW4869-treatment (HT29, P=0.2275; SW480, P=0.227; Fig. 2E), which indicated that CAPS1 promoted
FHC cell migration via exosomes.
Proteomic profiling of exosomes
derived from CAPS1-overexpressing CRC cells
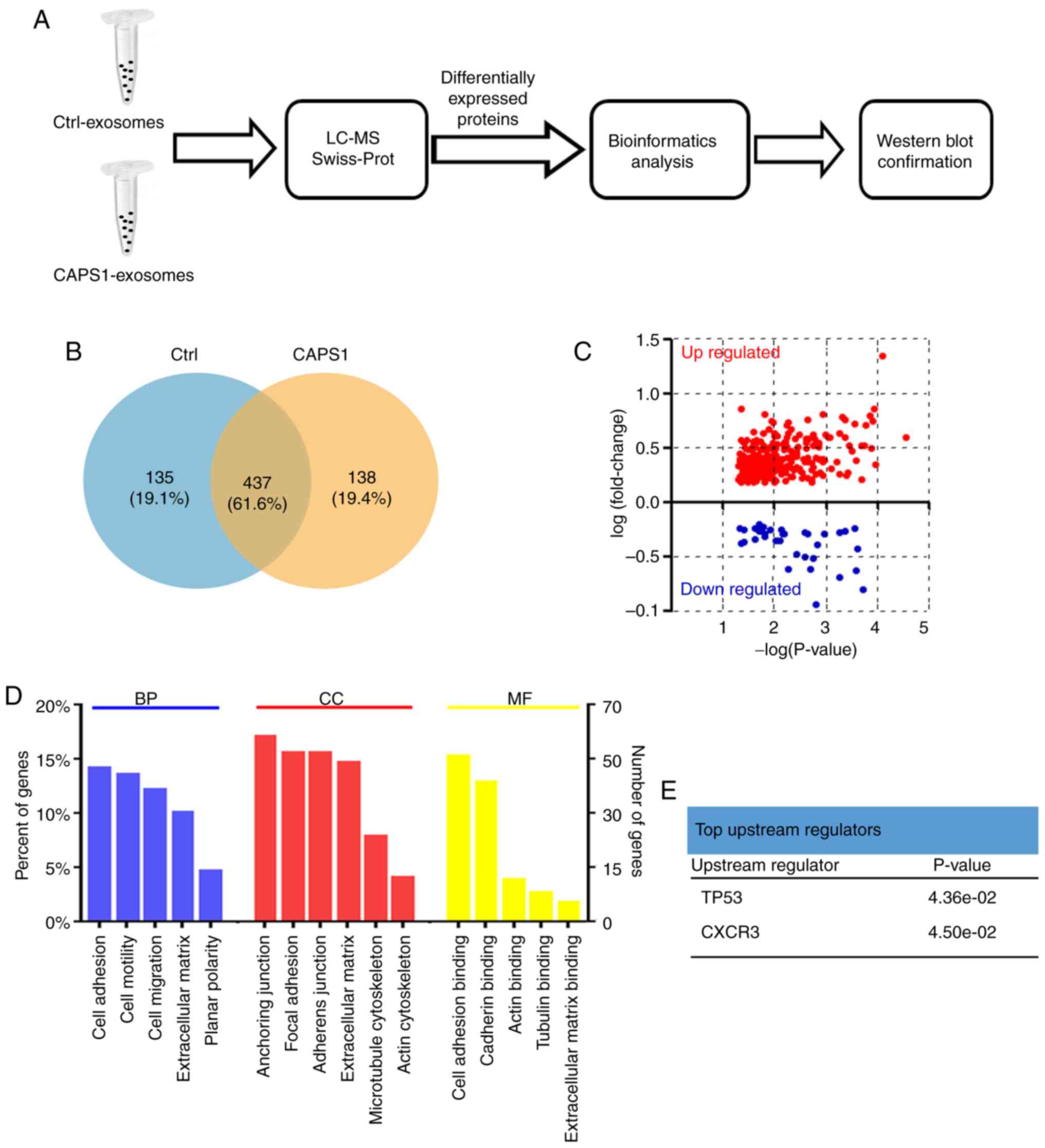
The present study further characterized the changes
in the exosome protein profile induced by CAPS1 overexpression
(Fig. 3A). Proteins were extracted
from exosomes derived from CAPS1-HT29 and control cells. LC-MS was
applied to assess the proteomics of exosomes, and the raw LC-MS
data files were searched against the Swiss-Prot Human Proteome
database. A total of 1,024 proteins were identified and quantified
(Table SI) (As aforementioned a
total of 1035 proteins was identified, but only 1,024 proteins
could be quantified since the intensity value of the remaining 11
proteins was extremely low and had to be ruled out for further
analysis). CAPS1-HT29 and control cells had in total 437 exosomal
proteins in common, while 138 exosomal proteins from CAPS1-HT29
differed from 135 proteins from control exosomes (Fig. 3B; Table
SII). Further analysis revealed that 297 proteins were
upregulated and 45 proteins were downregulated, with the cut-off
threshold set at fold-change=1.5 and P<0.05 (Fig. 3C; Table
SIII).
GO enrichment analysis and the UniProt-GOA database
were utilized to further classify the differentially expressed
proteins (Fig. 3D). The enriched
terms were cell adhesion, anchoring junction and cell adhesion
binding for biological processes, cell components and molecular
functions, respectively. Using IPA based on the Ingenuity Knowledge
Base, p53 and CXCR3 were identified as the top upstream regulators
(Fig. 3E).
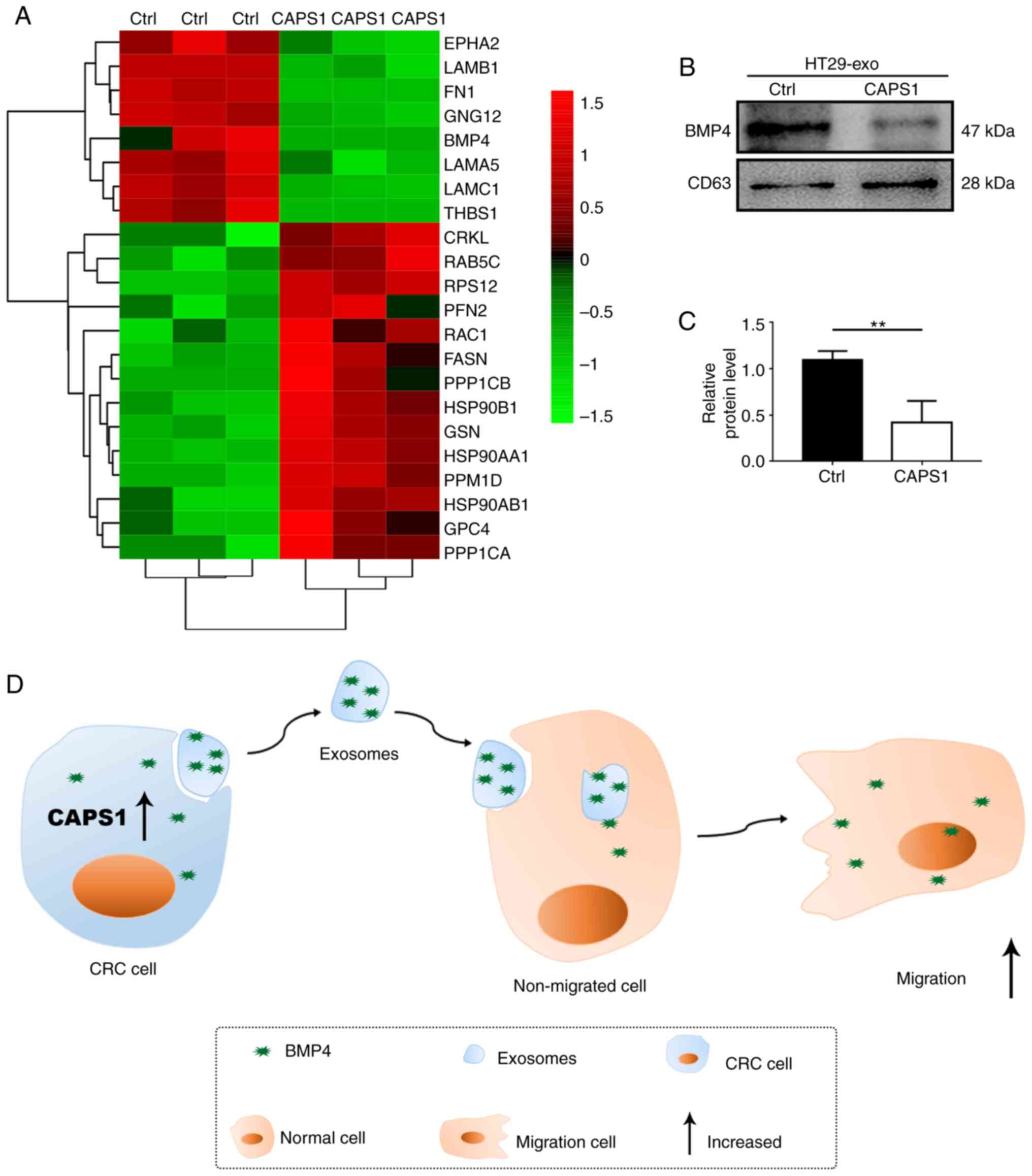
According to the bioinformatics analysis, the
candidate proteins that may participate in cell migration were
identified (Fig. 4A). Previously, it
has been reported that BMP4 is involved in metastasis in breast
cancer, prostate cancer, HCC and CRC (24–26,33). Thus,
BMP4 may serve vital roles in the process of CAPS1-induced cell
migration. To investigate the variation of BMP4 in the exosomes
derived from CAPS1-overexpressing CRC cells, western blot analysis
was performed. It was revealed that BMP4 was downregulated
~0.5-fold compared with Ctrl, which was consistent with the LC-MS
results (Fig. 4B and C). These data
indicated that exosomes are required for CAPS1-mediated cell
migration (Fig. 4D).
Discussion
The present study demonstrated that exosomes derived
from CAPS1-overexpressing CRC cells can enhance cell migration.
Overexpression of CAPS1 led to alterations in the expression
pattern of exosomal proteins, including the downregulation of BMP4,
which was further confirmed.
CAPS1 is the vertebrate homologue of the
Caenorhabditis elegans UNC-31 protein, which is involved in
vesicle secretion (34). Previously,
the essential role of CAPS1 in the development of cancer was
investigated. Our previous study revealed that CAPS1 promoted CRC
metastasis via the PI3K/Akt/GSK3β/Snail signaling pathway-mediated
EMT process (18). In addition, it
was demonstrated that CAPS1 serves as a biomarker of HCC, and the
expression of CAPS1 is decreased in patients with aggressive HCC
(35). Further investigation revealed
that CAPS1 likely inhibits HCC development via alteration of the
exocytosis-associated tumor microenvironment (36). In addition, decreased CAPS1 expression
was revealed to be correlated with poor prognosis of patients with
central nervous system primitive neuro-ectodermal tumors (37).
Metastasis is the predominant pathological reason
for the mortality of patients with CRC. Notably, increasing
evidence suggests that exosomes serve as key mediators in tumor
metastasis (11,38). It was reported that exosomes from
highly liver metastatic CRC cell lines could significantly enhance
the migration ability of Caco-2 cells, a CRC cell line with poor
liver metastatic potential (11).
Similarly, the present study revealed that exosomes derived from
CAPS1-overexpressing CRC cells facilitated FHC migration. The
function of exosomes in tumors predominantly depends on their
communication between cells. For example, exosomes can transfer
oncogenes, such as lncRNA H19 and miR-193a, to CRC, which promotes
cancer progression (39,40). Exosomes isolated from the serum of CRC
patients were revealed to transfer malignant traits and confer the
same phenotype of the primary tumor cells to the target cells
(41). Exosomes, as a communication
media for genetic exchange between cells, likely contribute to the
process of metastasis mediated by CAPS1 in CRC.
CAPS1 is multi-domain protein. Among these domains,
MH domain drives trans-SNARE complex formation and mediates
membrane fusion through syntaxin interactions (42,43).
Notably, SNARE complex protein YKT6 has been identified to
potentially be involved in the fusion of MVBs with the PM during
the biogenesis of exosomes (16).
Whether CAPS1 regulates the secretion and loading of exosomes via
the SNARE complex in CRC requires further investigation.
Bioinformatics strategies have frequently been used
to evaluate the function of exosomes (35,44). In
the present bioinformatics analysis, the differentially expressed
exosomal proteins involved in MAPK, Wnt, TGF-β, PI3K-AKT, or RAS
signaling, which were frequently dysregulated in CRC and were
associated with CRC metastasis (45–50), were
all considered as candidate proteins involved in cell migration
induced by CAPS1 (Fig. 4A). In view
of the role of BMP4 in metastasis, the present study examined the
expression level of BMP4 in the exosomes, and the downregulation of
BMP4 was further confirmed by western blot analysis (Fig. 4B and C). However, the role of BMP4 in
the process of CAPS1-induced cell migration should be further
investigated. Furthermore, p53 and CXCR3 were identified as
upstream regulators. p53 is activated by stress stimuli, and in
turn, it governs a complex anti-proliferative transcriptional
program to serve a tumor suppressive role (51,52).
However, TP53 is highly mutated and loses its function in
CRC (53). CXCR3 is a G
protein-coupled receptor that binds to chemokines to mediate
leukocyte trafficking, integrin activation, cytoskeletal changes
and chemotactic migration (54). A
previous study reported that CXCR3 was upregulated in CRC and
facilitated tumor metastasis via lymph nodes (55). Fibronectin 1 (FN1), which was
downregulated in the present LC-MS analysis, is a target molecule
of CXCR3. Furthermore, FN1 has been revealed to regulate numerous
molecules that are involved in cytoskeletal organization and
integrin signaling during tumor progression and metastasis
(56). Thrombospondin-1 (THBS-1), a
target gene of p53, is an endogenous inhibitor of angiogenesis and
was also downregulated in the current LC-MS analysis (57). BMP4, FN1 and THBS-1 transferred by
exosomes may serve an essential role in the process of
CAPS1-induced CRC cell migration.
CAPS1 had an effect on the diameter of the exosomes
according to the analysis of mean diameter by NTA detection. The
exosomes derived from CAPS1-HT29 cells exhibited a higher mean
diameter. Similarly, the majority of CAPS1-HT29-derived exosomes
presented with a diameter of 153 nm, whereas those from control
cells exhibited a diameter of 103 nm (Fig. 1E).
In summary, the present study demonstrated that
CAPS1 promoted cell migration by remodeling the protein profile of
exosomes. Furthermore, several candidate proteins were identified
that may participate in the process of CRC metastasis mediated by
CAPS1. However, the RNA profile of CAPS1-CRC-derived exosomes, and
the proteomics and transcriptomic profiles of target cells require
further analysis. The present data may improve understanding of a
new target for the treatment of patients with CRC metastasis via
the inhibition of exosome secretion.
Supplementary Material
Supporting Data
Supporting Data
Supporting Data
Acknowledgements
We thank Dr Xiaoxiao Wang and Dr Yaohua Li from Key
Laboratory of Glycoconjugate Research (Ministry of Public Health),
Department of Biochemistry and Molecular Biology, Shanghai Medical
college, Fudan University for technical assistance.
Funding
The present study was supported by grants from the
National Natural Science Foundation of China (grant nos. 81772615,
81672334, 81573423, 81770137 and 81802302) and the Shanghai Science
and Technology Commission (grant no. 15410710100).
Availability of data and materials
All data generated or analyzed during the present
study are included in this published article.
Authors' contributions
BW, DS, LD and SC designed and conceived this
project. BW, DS and LM developed methodology; BW, DS and YD
performed experiments and generated data; SC, SZ analyzed and
interpreted data; SC, BW wrote the manuscript. All authors
contributed to and approved the manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
There authors declare that they have no competing
interests.
References
|
1
|
Bray F, Ferlay J, Soerjomataram I, Siegel
RL, Torre LA and Jemal A: Global cancer statistics 2018: GLOBOCAN
estimates of incidence and mortality worldwide for 36 cancers in
185 countries. CA Cancer J Clin. 68:394–424. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Mundade R, Imperiale TF, Prabhu L, Loehrer
PJ and Lu T: Genetic pathways, prevention, and treatment of
sporadic colorectal cancer. Oncoscience. 1:400–406. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Brody H: Colorectal cancer. Nature.
521:S12015. View
Article : Google Scholar : PubMed/NCBI
|
|
4
|
Xu F, Tang B, Jin T and Dai C: Current
status of surgical treatment of colorectal liver metastases. World
J Clin Cases. 6:716–734. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Siegel RL, Miller KD and Jemal A: Cancer
statistics, 2018. CA Cancer J Clin. 68:7–30. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Bigagli E, Luceri C, Guasti D and Cinci L:
Exosomes secreted from human colon cancer cells influence the
adhesion of neighboring metastatic cells: Role of microRNA-210.
Cancer Biol Ther. 17:1062–1069. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Ruiz-López L, Blancas I, Garrido JM,
Mut-Salud N, Moya-Jódar M, Osuna A and Rodríguez-Serrano F: The
role of exosomes on colorectal cancer: A review. J Gastroenterol
Hepatol. 33:792–799. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Kreimer S, Belov AM, Ghiran I, Murthy SK,
Frank DA and Ivanov AR: Mass-spectrometry-based molecular
characterization of extracellular vesicles: Lipidomics and
proteomics. J Proteome Res. 14:2367–2384. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
van Niel G, Porto-Carreiro I, Simoes S and
Raposo G: Exosomes: A common pathway for a specialized function. J
Biochem. 140:13–21. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Schorey JS and Bhatnagar S: Exosome
function: From tumor immunology to pathogen biology. Traffic.
9:871–881. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Wang X, Ding X, Nan L, Wang Y, Wang J, Yan
Z, Zhang W, Sun J, Zhu W, Ni B, et al: Investigation of the roles
of exosomes in colorectal cancer liver metastasis. Oncol Rep.
33:2445–2453. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Akao Y, Khoo F, Kumazaki M, Shinohara H,
Miki K and Yamada N: Extracellular disposal of tumor-suppressor
miRs-145 and −34a via microvesicles and 5-FU resistance of human
colon cancer cells. Int J Mol Sci. 15:1392–1401. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Ragusa M, Statello L, Maugeri M,
Barbagallo C, Passanisi R, Alhamdani MS, Li Destri G, Cappellani A,
Barbagallo D, Scalia M, et al: Highly skewed distribution of miRNAs
and proteins between colorectal cancer cells and their exosomes
following Cetuximab treatment: Biomolecular, genetic and
translational implications. Oncoscience. 1:132–157. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Kowal J, Tkach M and Théry C: Biogenesis
and secretion of exosomes. Curr Opin Cell Biol. 29:116–25. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Colombo M, Raposo G and Thery C:
Biogenesis, secretion, and intercellular interactions of exosomes
and other extracellular vesicles. Annu Rev Cell Dev Biol.
30:255–289. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Gross JC, Chaudhary V, Bartscherer K and
Boutros M: Active Wnt proteins are secreted on exosomes. Nat Cell
Biol. 14:1036–1045. 2012. View
Article : Google Scholar : PubMed/NCBI
|
|
17
|
Villarroya-Beltri C, Gutiérrez-Vázquez C,
Sánchez-Cabo F, Pérez-Hernández D, Vázquez J, Martin-Cofreces N,
Martinez-Herrera DJ, Pascual-Montano A, Mittelbrunn M and
Sánchez-Madrid F: Sumoylated hnRNPA2B1 controls the sorting of
miRNAs into exosomes through binding to specific motifs. Nat
Commun. 4:29802013. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Zhao GX, Xu YY, Weng SQ, Zhang S, Chen Y,
Shen XZ, Dong L and Chen S: CAPS1 promotes colorectal cancer
metastasis via Snail mediated epithelial mesenchymal
transformation. Oncogene. 38:4574–4589. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Speidel D, Varoqueaux F, Enk C, Nojiri M,
Grishanin RN, Martin TF, Hofmann K, Brose N and Reim K: A family of
Ca2+-dependent activator proteins for secretion: Comparative
analysis of structure, expression, localization, and function. J
Biol Chem. 278:52802–52809. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Grishanin RN, Kowalchyk JA, Klenchin VA,
Ann K, Earles CA, Chapman ER, Gerona RR and Martin TF: CAPS acts at
a prefusion step in dense-core vesicle exocytosis as a PIP2 binding
protein. Neuron. 43:551–562. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Ehata S, Yokoyama Y, Takahashi K and
Miyazono K: Bi-directional roles of bone morphogenetic proteins in
cancer: Another molecular Jekyll and Hyde? Pathol Int. 63:287–296.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Yokoyama Y, Watanabe T, Tamura Y,
Hashizume Y, Miyazono K and Ehata S: Autocrine BMP-4 signaling is a
therapeutic target in colorectal cancer. Cancer Res. 77:4026–4038.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Deng G, Zeng S, Qu Y, Luo Q, Guo C, Yin L,
Han Y, Li Y, Cai C, Fu Y and Shen H: BMP4 promotes hepatocellular
carcinoma proliferation by autophagy activation through
JNK1-mediated Bcl-2 phosphorylation. J Exp Clin Canc Res.
37:1562018. View Article : Google Scholar
|
|
24
|
Zhou J, Liu H, Zhang L, Liu X, Zhang C,
Wang Y, He Q, Zhang Y, Li Y, Chen Q, et al: DJ-1 promotes
colorectal cancer progression through activating PLAGL2/Wnt/BMP4
axis. Cell Death Dis. 9:8652018. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Zeng S, Zhang Y, Ma J, Deng G, Qu Y, Guo
C, Han Y, Yin L, Cai C, Li Y, et al: BMP4 promotes metastasis of
hepatocellular carcinoma by an induction of epithelial-mesenchymal
transition via upregulating ID2. Cancer Lett. 390:67–76. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Guo D, Huang J and Gong J: Bone
morphogenetic protein 4 (BMP4) is required for migration and
invasion of breast cancer. Mol Cell Biochem. 363:179–190. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Fang WT, Fan CC, Li SM, Jang TH, Lin HP,
Shih NY, Chen CH, Wang TY, Huang SF, Lee AY, et al: Downregulation
of a putative tumor suppressor BMP4 by SOX2 promotes growth of lung
squamous cell carcinoma. Int J Cancer. 135:809–819. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Ying W, Riopel M, Bandyopadhyay G, Dong Y,
Birmingham A, Seo JB, Ofrecio JM, Wollam J, Hernandez-Carretero A,
Fu W, et al: Adipose tissue macrophage-derived exosomal miRNAs can
modulate in vivo and in vitro insulin sensitivity. Cell.
171:372–384.e12. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Li XJ, Ren ZJ, Tang JH and Yu Q: Exosomal
MicroRNA MiR-1246 promotes cell proliferation, invasion and drug
resistance by targeting CCNG2 in breast cancer. Cell Physiol
Biochem. 44:1741–1748. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Singh R, Pochampally R, Watabe K, Lu Z and
Mo YY: Exosome-mediated transfer of miR-10b promotes cell invasion
in breast cancer. Mol Cancer. 13:2562014. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Soucek K, Gajdusková P, Brázdová M,
Hýzd'alová M, Kocí L, Vydra D, Trojanec R, Pernicová Z, Lentvorská
L, Hajdúch M, et al: Fetal colon cell line FHC exhibits tumorigenic
phenotype, complex karyotype, and TP53 gene mutation. Cancer Genet
Cytogenet. 197:107–116. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Siddiqui KM and Chopra DP: Primary and
long term epithelial cell cultures from human fetal normal colonic
mucosa. In Vitro. 20:859–868. 1984. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Lin SC, Lee YC, Yu G, Cheng CJ, Zhou X,
Chu K, Murshed M, Le NT, Baseler L, Abe JI, et al:
Endothelial-to-osteoblast conversion generates osteoblastic
metastasis of prostate cancer. Dev Cell. 41:467–480.e3. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Ann K, Kowalchyk JA, Loyet KM and Martin
TF: Novel Ca2+-binding protein (CAPS) related to UNC-31 required
for Ca2+-activated exocytosis. J Biol Chem. 272:19637–19640. 1997.
View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Liu T, Xue R, Huang X, Zhang D, Dong L, Wu
H and Shen X: Proteomic profiling of hepatitis B virus-related
hepatocellular carcinoma with magnetic bead-based matrix-assisted
laser desorption/ionization time-of-flight mass spectrometry. Acta
Biochim Biophys Sin (Shanghai). 43:542–550. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Xue R, Tang W, Dong P, Weng S, Ma L, Chen
S, Liu T, Shen X, Huang X, Zhang S, et al: CAPS1 negatively
regulates hepatocellular carcinoma development through alteration
of exocytosis-associated tumor microenvironment. Int J Mol Sci.
17(pii): E16262016. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Miller S, Rogers HA, Lyon P, Rand V,
Adamowicz-Brice M, Clifford SC, Hayden JT, Dyer S, Pfister S,
Korshunov A, et al: Genome-wide molecular characterization of
central nervous system primitive neuroectodermal tumor and
pineoblastoma. Neuro Oncol. 13:866–879. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Fang JH, Zhang ZJ, Shang LR, Luo YW, Lin
YF, Yuan Y and Zhuang SM: Hepatoma cell-secreted exosomal
microRNA-103 increases vascular permeability and promotes
metastasis by targeting junction proteins. Hepatology.
68:1459–1475. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Ren J, Ding L, Zhang D, Shi G, Xu Q, Shen
S, Wang Y, Wang T and Hou Y: Carcinoma-associated fibroblasts
promote the stemness and chemoresistance of colorectal cancer by
transferring exosomal lncRNA H19. Theranostics. 8:3932–3948. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Teng Y, Ren Y, Hu X, Mu J, Samykutty A,
Zhuang X, Deng Z, Kumar A, Zhang L, Merchant ML, et al:
MVP-mediated exosomal sorting of miR-193a promotes colon cancer
progression. Nat Commun. 8:144482017. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Abdouh M, Hamam D, Gao Z, Arena V, Arena M
and Arena GO: Exosomes isolated from cancer patients' sera transfer
malignant traits and confer the same phenotype of primary tumors to
oncosuppressor-mutated cells. J Exp Clin Canc Res. 36:1132017.
View Article : Google Scholar
|
|
42
|
James DJ, Kowalchyk J, Daily N, Petrie M
and Martin TF: CAPS drives trans-SNARE complex formation and
membrane fusion through syntaxin interactions. Proc Natl Acad Sci
USA. 106:17308–17313. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Koch H, Hofmann K and Brose N: Definition
of Munc13-homology-domains and characterization of a novel
ubiquitously expressed Munc13 isoform. Biochem J. 349:247–253.
2000. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Ji H, Greening DW, Barnes TW, Lim JW,
Tauro BJ, Rai A, Xu R, Adda C, Mathivanan S, Zhao W, et al:
Proteome profiling of exosomes derived from human primary and
metastatic colorectal cancer cells reveal differential expression
of key metastatic factors and signal transduction components.
Proteomics. 13:1672–1686. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Fang YJ, Lu ZH, Wang GQ, Pan ZZ, Zhou ZW,
Yun JP, Zhang MF and Wan DS: Elevated expressions of MMP7, TROP2,
and survivin are associated with survival, disease recurrence, and
liver metastasis of colon cancer. Int J Colorectal Dis. 24:875–884.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Urosevic J, Nebreda AR and Gomis RR: MAPK
signaling control of colon cancer metastasis. Cell Cycle.
13:2641–2642. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Jiang Y, Liu XQ, Rajput A, Geng L, Ongchin
M, Zeng Q, Taylor GS and Wang J: Phosphatase PRL-3 is a direct
regulatory target of TGFbeta in colon cancer metastasis. Cancer
Res. 71:234–244. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Pancione M, Forte N, Fucci A, Sabatino L,
Febbraro A, Di Blasi A, Daniele B, Parente D and Colantuoni V:
Prognostic role of beta-catenin and p53 expression in the
metastatic progression of sporadic colorectal cancer. Hum Pathol.
41:867–876. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Song G, Xu S, Zhang H, Wang Y, Xiao C,
Jiang T, Wu L, Zhang T, Sun X, Zhong L, et al: TIMP1 is a
prognostic marker for the progression and metastasis of colon
cancer through FAK-PI3K/AKT and MAPK pathway. J Exp Clin Canc Res.
35:1482016. View Article : Google Scholar
|
|
50
|
Bahrami A, Hassanian SM, ShahidSales S,
Farjami Z, Hasanzadeh M, Anvari K, Aledavood A, Maftouh M, Ferns
GA, Khazaei M and Avan A: Targeting RAS signaling pathway as a
potential therapeutic target in the treatment of colorectal cancer.
J Cell Physiol. 233:2058–2066. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Kastenhuber ER and Lowe SW: Putting p53 in
Context. Cell. 170:1062–1078. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Mills KD: Tumor suppression: Putting p53
in context. Cell Cycle. 12:3461–3462. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Iacopetta B: TP53 mutation in colorectal
cancer. Hum Mutat. 21:271–276. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Ma B, Khazali A and Wells A: CXCR3 in
carcinoma progression. Histol Histopathol. 30:781–792.
2015.PubMed/NCBI
|
|
55
|
Kawada K, Hosogi H, Sonoshita M, Sakashita
H, Manabe T, Shimahara Y, Sakai Y, Takabayashi A, Oshima M and
Taketo MM: Chemokine receptor CXCR3 promotes colon cancer
metastasis to lymph nodes. Oncogene. 26:4679–4688. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Chen Y, Xie Y, Xu L, Zhan S, Xiao Y, Gao
Y, Wu B and Ge W: Protein content and functional characteristics of
serum-purified exosomes from patients with colorectal cancer
revealed by quantitative proteomics. Int J Cancer. 140:900–913.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Jia L and Waxman DJ: Thrombospondin-1 and
pigment epithelium-derived factor enhance responsiveness of KM12
colon tumor to metronomic cyclophosphamide but have disparate
effects on tumor metastasis. Cancer Lett. 330:241–249. 2013.
View Article : Google Scholar : PubMed/NCBI
|