|
1
|
Singh AK, Kumar R and Pandey AK:
Hepatocellular carcinoma: Causes, mechanism of progression and
biomarkers. Curr Chem Genom Transl Med. 12:9–26. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Zhu X, Tang Z and Sun HC: Targeting
angiogenesis for liver cancer: Past, present, and future. Genes
Dis. 7:328–335. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Sagnelli E, Potenza N, Onorato L, Sagnelli
C, Coppola N and Russo A: Micro-RNAs in hepatitis B virus-related
chronic liver diseases and hepatocellular carcinoma. World J
Hepatol. 10:558–570. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Sengupta S and Parikh ND: Biomarker
development for hepatocellular carcinoma early detection: Current
and future perspectives. Hepat Oncol. 4:111–122. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Rupaimoole R, Calin GA, Lopez-Berestein G
and Sood AK: miRNA deregulation in cancer cells and the tumor
microenvironment. Cancer Discov. 6:235–246. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
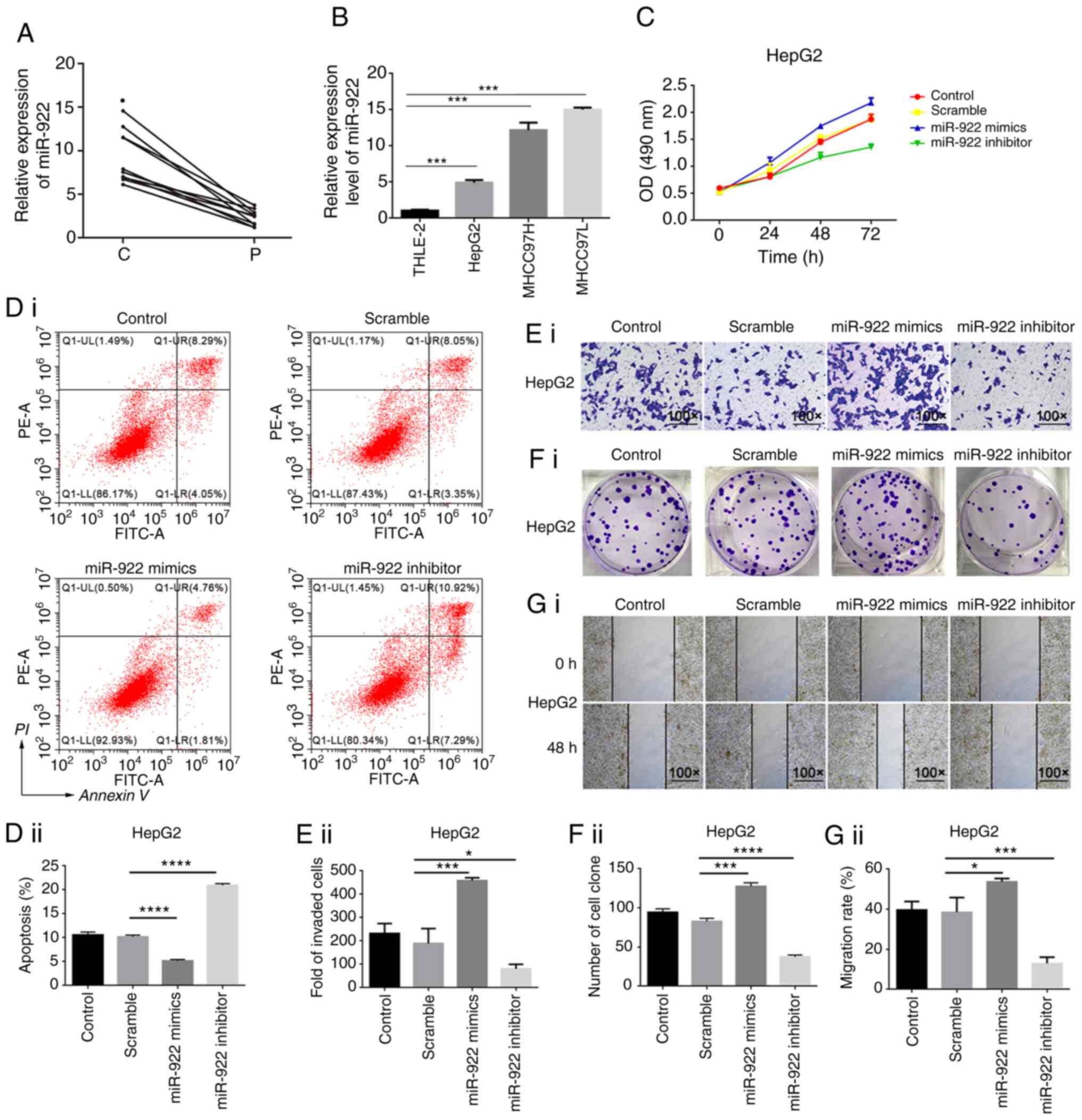
Shayimu P, Wang JB, Liu L, Tuerdi R, Yu CG
and Yusufu A: miR-922 regulates apoptosis, migration, and invasion
by targeting SOCS1 in gastric cancer. Kaohsiung J Med Sci.
36:178–185. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Liu J, Su Z, Zeng Y, Zhang H, Yang S and
Liu G: miR-922 regulates CYLD expression and promotes the cell
proliferation of human hepatocellular carcinoma. Oncol Rep.
37:1445–1450. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Sevinc ED, Cecener G, Ak S, Tunca B, Egeli
U, Gokgoz S, Tolunay S and Tasdelen I: Expression and clinical
significance of miRNAs that may be associated with the FHIT gene in
breast cancer. Gene. 590:278–284. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
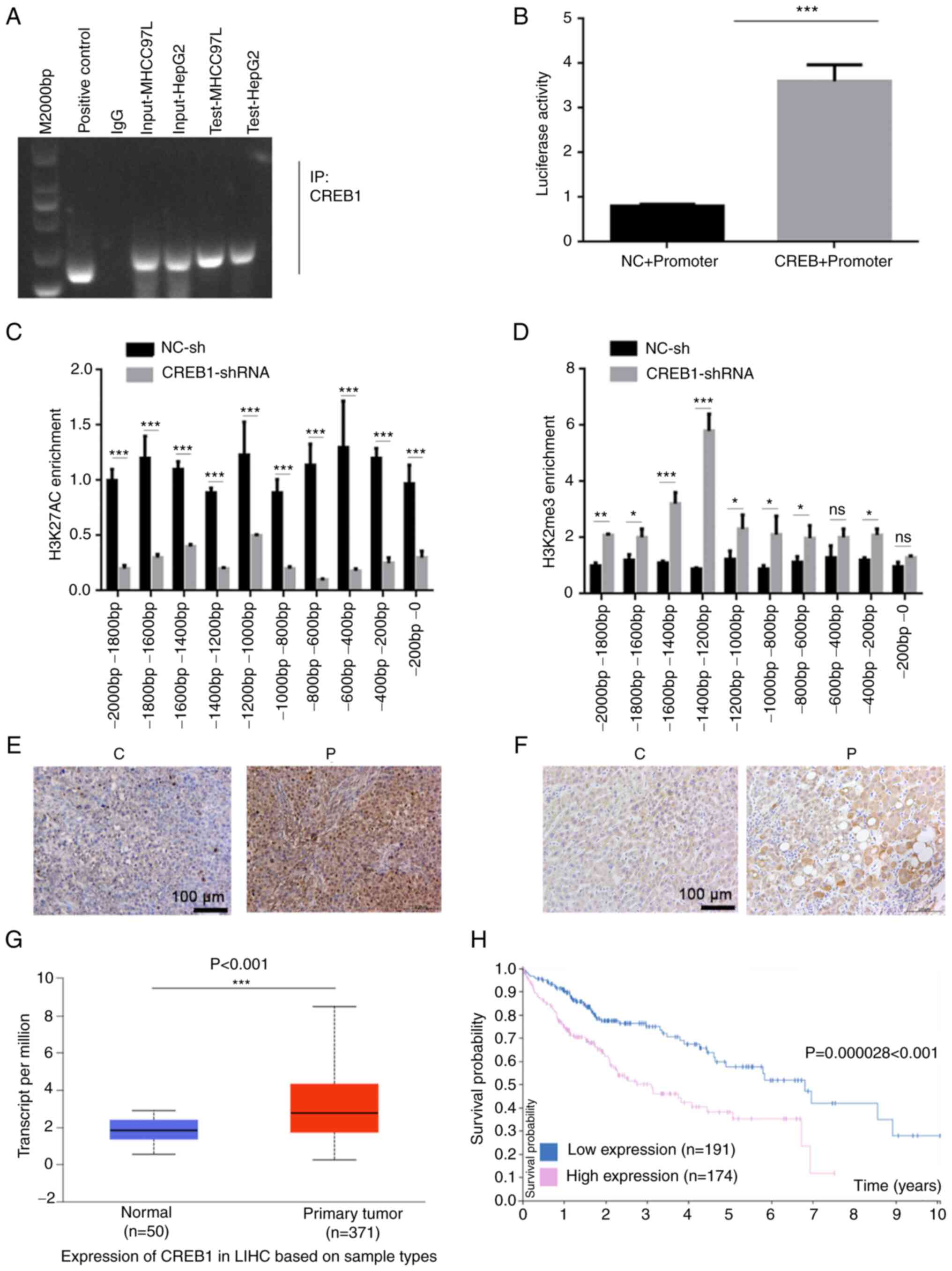
Abramovitch R, Tavor E, Jacob-Hirsch J,
Zeira E, Amariglio N, Pappo O, Rechavi G, Galun E and Honigman A: A
pivotal role of cyclic AMP-responsive element binding protein in
tumor progression. Cancer Res. 64:1338–1346. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Kovach SJ, Price JA, Shaw CM, Theodorakis
NG and McKillop IH: Role of cyclic-AMP responsive element binding
(CREB) proteins in cell proliferation in a rat model of
hepatocellular carcinoma. J Cell Physiol. 206:411–419. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Li Y, Fu Y, Hu X, Sun L, Tang D, Li N,
Peng F and Fan XG: The HBx-CTTN interaction promotes cell
proliferation and migration of hepatocellular carcinoma via CREB1.
Cell Death Dis. 10:4052019. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Oba A, Shimada S, Akiyama Y, Nishikawaji
T, Mogushi K, Ito H, Matsumura S, Aihara A, Mitsunori Y, Ban D, et
al: ARID2 modulates DNA damage response in human hepatocellular
carcinoma cells. J Hepatol. 66:942–951. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Marrero JA, Kulik LM, Sirlin CB, Zhu AX,
Finn RS, Abecassis MM, Roberts LR and Heimbach JK: Diagnosis,
staging, and management of hepatocellular carcinoma: 2018 practice
guidance by the American association for the study of liver
diseases. Hepatology. 68:723–750. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Tellapuri S, Sutphin PD, Beg MS, Singal AG
and Kalva SP: Staging systems of hepatocellular carcinoma: A
review. Indian J Gastroenterol. 37:481–491. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Zhu P, Wang Y, Wu J, Huang G, Liu B, Ye B,
Du Y, Gao G, Tian Y, He L and Fan Z: LncBRM initiates YAP1
signalling activation to drive self-renewal of liver cancer stem
cells. Nat Commun. 7:136082016. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Wang YW, Chen X, Ma R and Gao P:
Understanding the CREB1-miRNA feedback loop in human malignancies.
Tumour Biol. 37:8487–8502. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Rad R, Rad L, Wang W, Strong A, Ponstingl
H, Bronner IF, Mayho M, Steiger K, Weber J, Hieber M, et al: A
conditional piggyBac transposition system for genetic screening in
mice identifies oncogenic networks in pancreatic cancer. Nat Genet.
47:47–56. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Xu J, Li J, Zheng TH, Bai L and Liu ZJ:
MicroRNAs in the occurrence and development of primary
hepatocellular carcinoma. Adv Clin Exp Med. 25:971–975. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Pineau P, Volinia S, McJunkin K, Marchio
A, Battiston C, Terris B, Mazzaferro V, Lowe SW, Croce CM and
Dejean A: miR-221 overexpression contributes to liver
tumorigenesis. Proc Natl Acad Sci USA. 107:264–269. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Varnholt H: The role of microRNAs in
primary liver cancer. Ann Hepatol. 7:104–113. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
He XX, Chang Y, Meng FY, Wang MY, Xie QH,
Tang F, Li PY, Song YH and Lin JS: MicroRNA-375 targets AEG-1 in
hepatocellular carcinoma and suppresses liver cancer cell growth in
vitro and in vivo. Oncogene. 31:3357–3369. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Liu Y, Chen SH, Jin X and Li YM: Analysis
of differentially expressed genes and microRNAs in alcoholic liver
disease. Int J Mol Med. 31:547–554. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Lee YS and Dutta A: MicroRNAs in cancer.
Annu Rev Pathol. 4:199–227. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Hu PC, Li K, Tian YH, Pan WT, Wang Y, Xu
XL, He YQ, Gao Y, Wei L and Zhang JW: CREB1/Lin28/miR-638/VASP
interactive network drives the development of breast cancer. Int J
Biol Sci. 15:2733–2749. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Hodges C, Kirkland JG and Crabtree GR: The
many roles of BAF (mSWI/SNF) and PBAF complexes in cancer. Cold
Spring Harb Perspect Med. 6:a0269302016. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Linhares AC, Stupka JA, Ciapponi A,
Bardach AE, Glujovsky D, Aruj PK, Mazzoni A, Rodriguez JA, Rearte
A, Lanzieri TM, et al: Burden and typing of rotavirus group A in
Latin America and the Caribbean: Systematic review and
meta-analysis. Rev Med Virol. 21:89–109. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
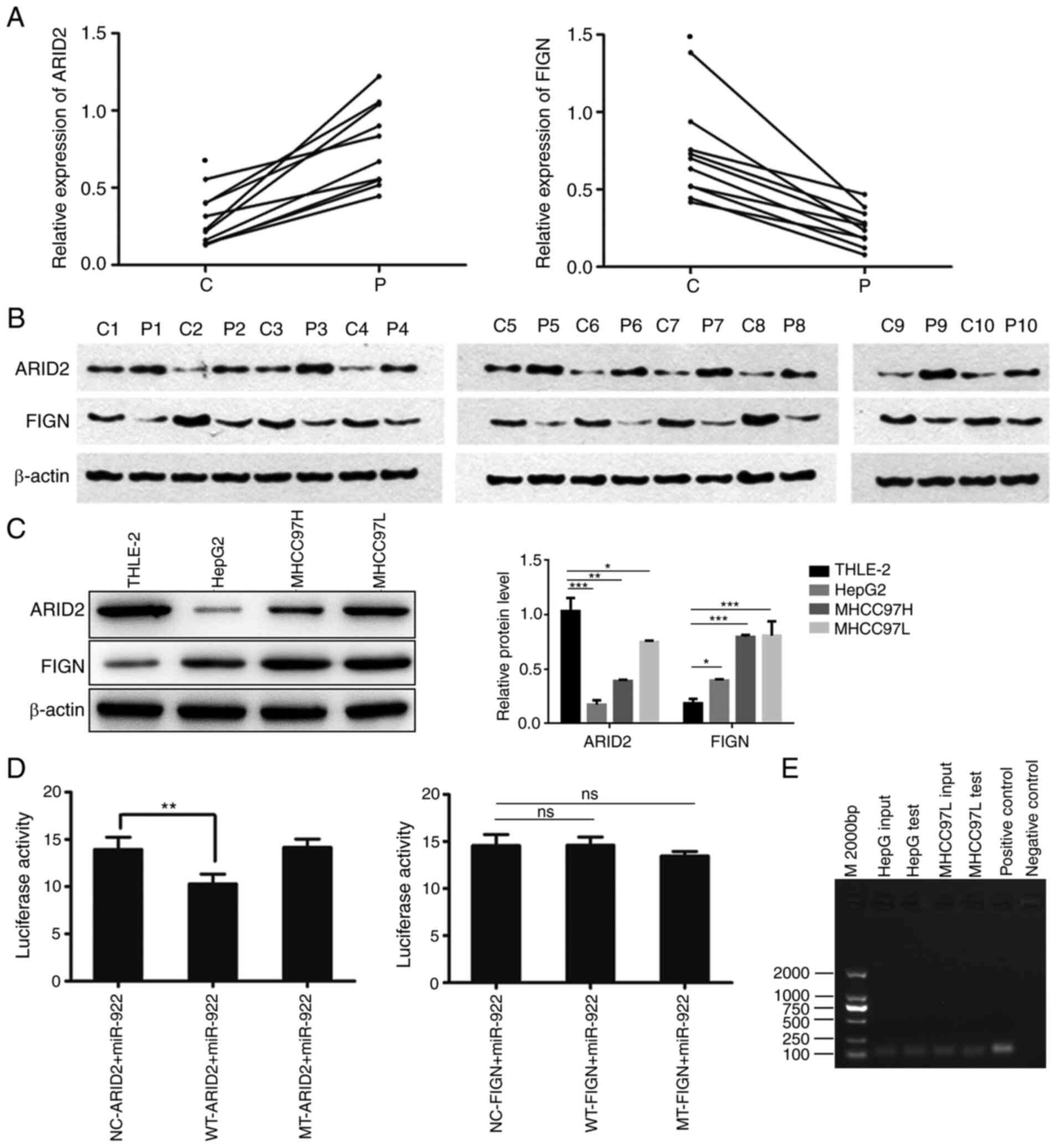
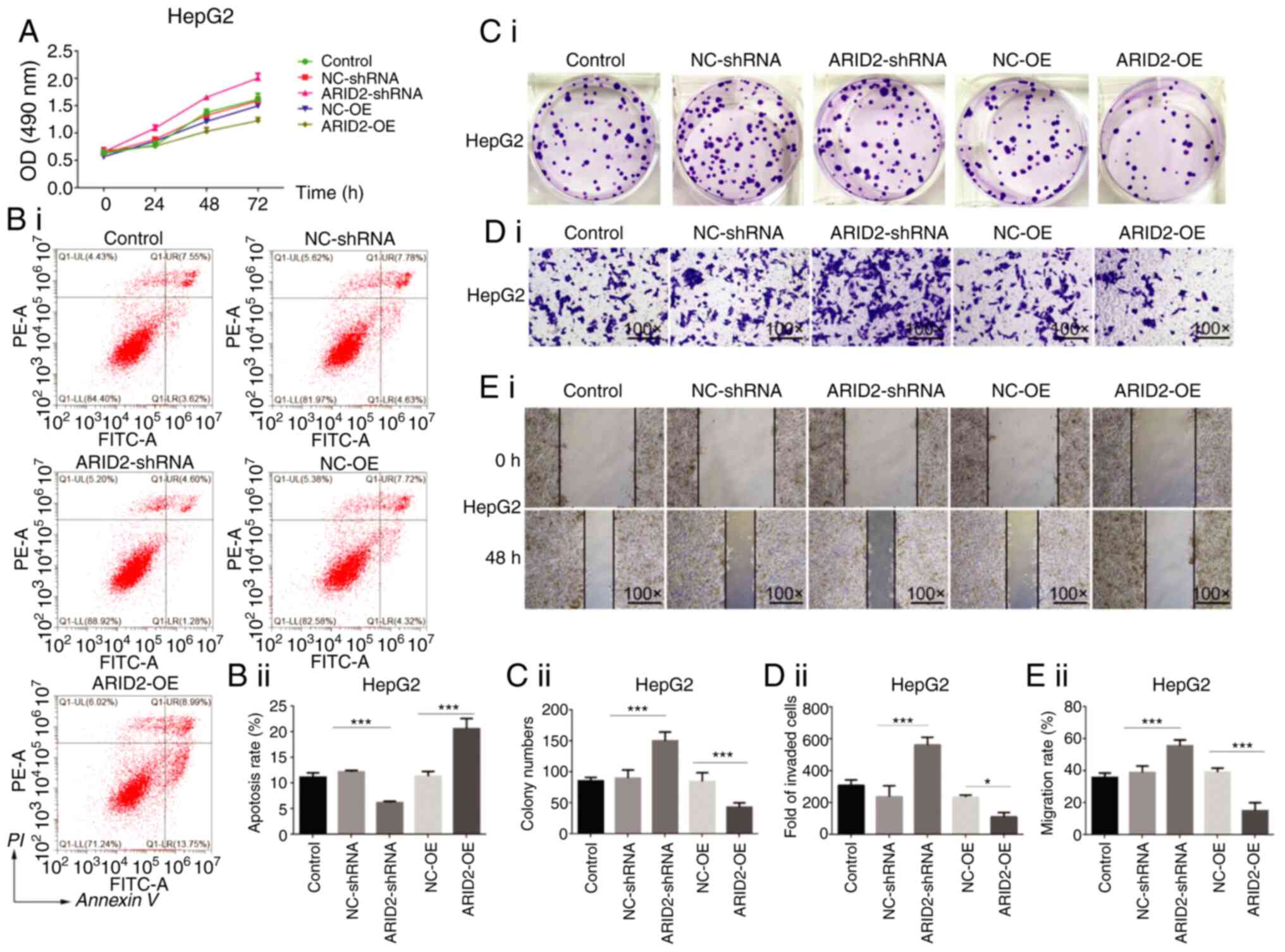
Zhao H, Wang J, Han Y, Huang Z, Ying J, Bi
X, Zhao J, Fang Y, Zhou H, Zhou J, et al: ARID2: A new tumor
suppressor gene in hepatocellular carcinoma. Oncotarget. 2:886–891.
2011. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
You J, Yang H, Lai Y, Simon L, Au J and
Burkart AL: AT-rich interactive domain 2, p110α, p53, and β-catenin
protein expression in hepatocellular carcinoma and
clinicopathologic implications. Hum Pathol. 46:583–592. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Zhang L, Wang W, Li X, He S, Yao J, Wang
X, Zhang D and Sun X: MicroRNA-155 promotes tumor growth of human
hepatocellular carcinoma by targeting ARID2. Int J Oncol.
48:2425–2434. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Wang Y, Chang W, Chang W, Chang X, Zhai S,
Pan G and Dang S: MicroRNA-376c-3p facilitates human hepatocellular
carcinoma progression via repressing AT-rich interaction domain 2.
J Cancer. 9:4187–4196. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Yu P, Wu D, You Y, Sun J, Lu L, Tan J and
Bie P: miR-208-3p promotes hepatocellular carcinoma cell
proliferation and invasion through regulating ARID2 expression. Exp
Cell Res. 336:232–241. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Liu Q, Yang P, Tu K, Zhang H, Zheng X, Yao
Y and Liu Q: TPX2 knockdown suppressed hepatocellular carcinoma
cell invasion via inactivating AKT signaling and inhibiting MMP2
and MMP9 expression. Chin J Cancer Res. 26:410–417. 2014.PubMed/NCBI
|