|
1
|
Miller AJ and Mihm MC Jr: Melanoma. N Engl
J Med. 355:51–65. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Wu Y, Wang Y, Wang L, Yin P, Lin Y and
Zhou M: Burden of melanoma in China, 1990–2017: Findings from the
2017 global burden of disease study. Int J Cancer. 147:692–701.
2020. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Aubuchon MM, Bolt LJ, Janssen-Heijnen ML,
Verleisdonk-Bolhaar ST, van Marion A and van Berlo CL:
Epidemiology, management and survival outcomes of primary cutaneous
melanoma: A ten-year overview. Acta Chir Belg. 117:29–35. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Karlsson P, Boeryd B, Sander S, Westermark
P and Rosdahl I: Increasing incidence of cutaneous malignant
melanoma in children and adolescents 12–19 years of age in Sweden
1973–92. Acta Derm Venereol. 78:289–292. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Veierød MB, Weiderpass E, Thörn M, Hansson
J, Lund E, Armstrong B and Adami HO: A prospective study of
pigmentation, sun exposure, and risk of cutaneous malignant
melanoma in women. J Natl Cancer Inst. 95:1530–1538. 2003.
View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Lees VC and Briggs JC: Effect of initial
biopsy procedure on prognosis in Stage 1 invasive cutaneous
malignant melanoma: Review of 1086 patients. Br J Surg.
78:1108–1110. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Tseng WW, Fadaki N and Leong SP:
Metastatic tumor dormancy in cutaneous melanoma: Does surgery
induce escape? Cancers (Basel). 3:730–746. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Kim JH, Hahn EW and Tokita N: Combination
hyperthermia and radiation therapy for cutaneous malignant
melanoma. Cancer. 41:2143–2148. 1978. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Li PF, Chen SC, Xia T, Jiang XM, Shao YF,
Xiao BX and Guo JM: Non-coding RNAs and gastric cancer. World J
Gastroenterol. 20:5411–5419. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Moran VA, Perera RJ and Khalil AM:
Emerging functional and mechanistic paradigms of mammalian long
non-coding RNAs. Nucleic Acids Res. 40:6391–6400. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Chen X, Gao J, Yu Y, Zhao Z and Pan Y:
LncRNA FOXD3-AS1 promotes proliferation, invasion and migration of
cutaneous malignant melanoma via regulating miR-325/MAP3K2. Biomed
Pharmacother. 120:1094382019. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Chen XE, Chen P, Chen S, Lu J, Ma T, Shi G
and Sheng L: Long non-coding RNA FENDRR inhibits migration and
invasion of cutaneous malignant melanoma cells. Biosci Rep.
40:BSR201911942020. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Yu S, Wang D, Shao Y, Zhang T, Xie H,
Jiang X, Deng Q, Jiao Y, Yang J, Cai C and Sun L: SP1-induced
lncRNA TINCR overexpression contributes to colorectal cancer
progression by sponging miR-7-5p. Aging. 11:1389–1403. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Dong H, Hu J, Zou K, Ye M, Chen Y, Wu C,
Chen X and Han M: Activation of LncRNA TINCR by H3K27 acetylation
promotes Trastuzumab resistance and epithelial-mesenchymal
transition by targeting MicroRNA-125b in breast cancer. Mol Cancer.
18:32019. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Dong L, Ding H, Li Y, Xue D and Liu Y:
LncRNA TINCR is associated with clinical progression and serves as
tumor suppressive role in prostate cancer. Cancer Manag Res.
10:2799–2807. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
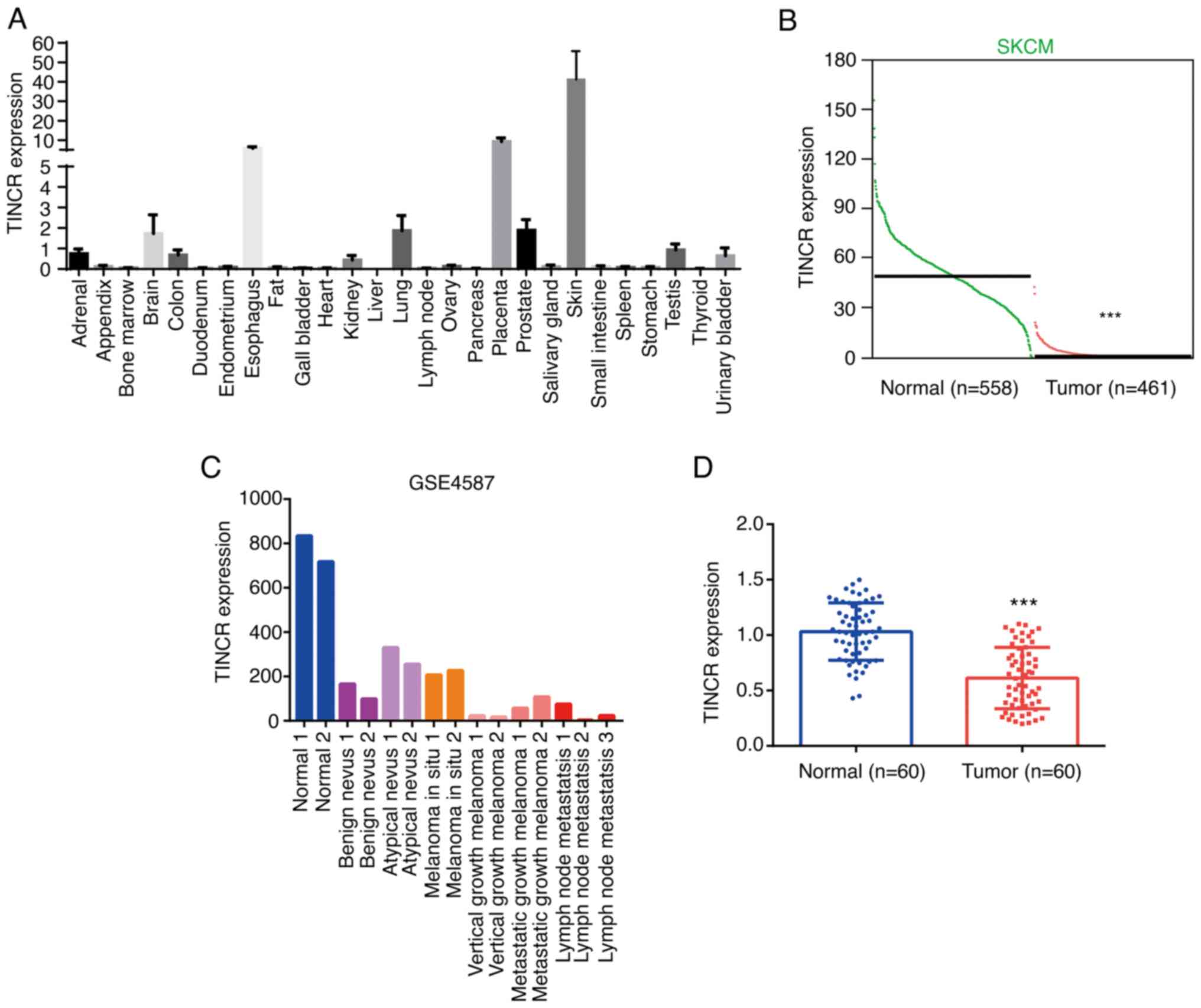
Smith AP, Hoek K and Becker D:
Whole-genome expression profiling of the melanoma progression
pathway reveals marked molecular differences between nevi/melanoma
in situ and advanced-stage melanomas. Cancer Biol Ther. 9:1018–29.
2005. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Fagerberg L, Hallström BM, Oksvold P,
Kampf C, Djureinovic D, Odeberg J, Habuka M, Tahmasebpoor S,
Danielsson A, Edlund K, et al: Analysis of the human
tissue-specific expression by genome-wide integration of
transcriptomics and antibody-based proteomics. Mol Cell Proteomics.
13:397–406. 2014. View Article : Google Scholar
|
|
18
|
Tang Z, Li C, Kang B, Gao G, Li C and
Zhang Z: GEPIA: A web server for cancer and normal gene expression
profiling and interactive analyses. Nucleic Acids Res. 45:W98–W102.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PcR and
the 2(-delta deltac(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Mo J, Zhao X, Dong X, Liu T, Zhao N, Zhang
D, Wang W, Zhang Y and Sun B: Effect of EphA2 knockdown on melanoma
metastasis depends on intrinsic ephrinA1 level. Cell Oncol (Dordr).
43:655–667. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Babapoor S, Wu R, Kozubek J, Auidi D,
Grant-Kels JM and Dadras SS: Identification of microRNAs associated
with invasive and aggressive phenotype in cutaneous melanoma by
next-generation sequencing. Lab Invest. 97:636–648. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Liu X, Fu Y, Zhang G, Zhang D, Liang N, Li
F, Li C, Sui C, Jiang J, Lu H, et al: miR-424-5p promotes anoikis
resistance and lung metastasis by inactivating Hippo signaling in
thyroid cancer. Mol Ther Oncolytics. 15:248–260. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Wei S, Li Q, Li Z, Wang L, Zhang L and Xu
Z: miR-424-5p promotes proliferation of gastric cancer by targeting
Smad3 through TGF-β signaling pathway. Oncotarget. 7:75185–75196.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Sahami-Fard MH, Kheirandish S and Sheikhha
MH: Expression levels of miR-143-3p and −424-5p in colorectal
cancer and their clinical significance. Cancer Biomark. 24:291–297.
2019. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Zhang J, Liu H, Hou L, Wang G, Zhang R,
Huang Y, Chen X and Zhu J: Circular RNA_LARP4 inhibits cell
proliferation and invasion of gastric cancer by sponging miR-424-5p
and regulating LATS1 expression. Mol Cancer. 16:1512017. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Furth N, Pateras IS, Rotkopf R, Vlachou V,
Rivkin I, Schmitt I, Bakaev D, Gershoni A, Ainbinder E, Leshkowitz
D, et al: LATS1 and LATS2 suppress breast cancer progression by
maintaining cell identity and metabolic state. Life Sci Alliance.
1:e2018001712018. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Zhang ML, Sun WH, Wu HQ, Liu ZD and Wang
P: Knockdown of microRNA-103a-3p inhibits the malignancy of thyroid
cancer cells through Hippo signaling pathway by upregulating LATS1.
Neoplasma. 67:1266–1278. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Yu T, Bachman J and Lai ZC: Mutation
analysis of large tumor suppressor genes LATS1 and LATS2 supports a
tumor suppressor role in human cancer. Protein Cell. 6:6–11. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Zhang J, Wang G, Chu SJ, Zhu JS, Zhang R,
Lu WW, Xia LQ, Lu YM, Da W and Sun Q: Loss of large tumor
suppressor 1 promotes growth and metastasis of gastric cancer cells
through upregulation of the YAP signaling. Oncotarget.
7:16180–16193. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Ehmer U and Sage J: Control of
proliferation and cancer growth by the hippo signaling pathway. Mol
Cancer Res. 14:127–140. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Janse van Rensburg HJ and Yang X: The
roles of the hippo pathway in cancer metastasis. Cell Signal.
28:1761–1772. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Yu S, Zhang M, Huang L, Ma Z, Gong X, Liu
W, Zhang J, Chen L, Yu Z, Zhao W and Liu Y: ERK1 indicates good
prognosis and inhibits breast cancer progression by suppressing
YAP1 signaling. Aging. 11:12295–12314. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Brodowska K, Al-Moujahed A, Marmalidou A,
Horste M, Cichy J, Miller J, Evangelos E and Vavvas D: The
clinically used photosensitizer Verteporfin (VP) inhibits YAP-TEAD
and human retinoblastoma cell growth in vitro without light
activation. Exp Eye Res. 124:67–73. 2014. View Article : Google Scholar : PubMed/NCBI
|