|
1
|
Hanahan D and Weinberg RA: Hallmarks of
cancer: The next generation. Cell. 144:646–674. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Boshuizen J and Peeper DS: Rational cancer
treatment combinations: An urgent clinical need. Mol Cell.
78:1002–1018. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Mun EJ, Babiker HM, Weinberg U, Kirson ED
and Von Hoff DD: Tumor-treating fields: A fourth modality in cancer
treatment. Clin Cancer Res. 24:266–275. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Ilves I, Petojevic T, Pesavento JJ and
Botchan MR: Activation of the MCM2-7 helicase by association with
Cdc45 and GINS proteins. Mol Cell. 37:247–258. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Boos D, Frigola J and Diffley JF:
Activation of the replicative DNA helicase: Breaking up is hard to
do. Curr Opin Cell Biol. 24:423–430. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Tanaka S and Araki H: Helicase activation
and establishment of replication forks at chromosomal origins of
replication. Cold Spring Harb Perspect Biol. 5:a0103712013.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Owens JC, Detweiler CS and Li JJ: CDC45 is
required in conjunction with CDC7/DBF4 to trigger the initiation of
DNA replication. Proc Natl Acad Sci USA. 94:12521–12526. 1997.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Feng D, Tu Z, Wu W and Liang C: Inhibiting
the expression of DNA replication-initiation proteins induces
apoptosis in human cancer cells. Cancer Res. 63:7356–7364.
2003.PubMed/NCBI
|
|
9
|
Pollok S, Bauerschmidt C, Sänger J,
Nasheuer HP and Grosse F: Human Cdc45 is a proliferation-associated
antigen. FEBS J. 274:3669–3684. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Srinivasan SV, Dominguez-Sola D, Wang LC,
Hyrien O and Gautier J: Cdc45 is a critical effector of
myc-dependent DNA replication stress. Cell Rep. 3:1629–1639. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Hu Y, Wang L, Li Z, Wan Z, Shao M, Wu S
and Wang G: Potential prognostic and diagnostic values of CDC6,
CDC45, ORC6 and SNHG7 in colorectal cancer. Onco Targets Ther.
12:11609–11621. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Huang J, Li Y, Lu Z, Che Y, Sun S, Mao S,
Lei Y, Zang R, Li N, Zheng S, et al: Analysis of functional hub
genes identifies CDC45 as an oncogene in non-small cell lung
cancer-a short report. Cell Oncol (Dordr). 42:571–578. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Piao J, Sun J, Yang Y, Jin T, Chen L and
Lin Z: Target gene screening and evaluation of prognostic values in
non-small cell lung cancers by bioinformatics analysis. Gene.
647:306–311. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Ke Y, Guo W, Huang S, Li Y, Guo Y, Liu X,
Jin Y and Ma H: RYBP inhibits esophageal squamous cell carcinoma
proliferation through downregulating CDC6 and CDC45 in G1-S phase
transition process. Life Sci. 250:1175782020. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Lu HP, Du XF, Li JD, Huang SN, He RQ, Wu
HY, Li MF, Wu WZ, Chen JT, Mo WJ and Chen G: Expression of cell
division cycle protein 45 in tissue microarrays and the CDC45 gene
by bioinformatics analysis in human hepatocellular carcinoma and
patient outcomes. Med Sci Monit. 27:e9288002021.PubMed/NCBI
|
|
16
|
Sang L, Wang XM, Xu DY and Zhao WJ:
Bioinformatics analysis of aberrantly methylated-differentially
expressed genes and pathways in hepatocellular carcinoma. World J
Gastroenterol. 24:2605–2616. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Xiang XH, Yang L, Zhang X, Ma XH, Miao RC,
Gu JX, Fu YN, Yao Q, Zhang JY, Liu C, et al:
Seven-senescence-associated gene signature predicts overall
survival for asian patients with hepatocellular carcinoma. World J
Gastroenterol. 25:1715–1728. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Sun J, Shi R, Zhao S, Li X, Lu S, Bu H and
Ma X: Cell division cycle 45 promotes papillary thyroid cancer
progression via regulating cell cycle. Tumour Biol.
39:10104283177053422017. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Deng M, Brägelmann J, Schultze JL and
Perner S: Web-TCGA: An online platform for integrated analysis of
molecular cancer data sets. BMC Bioinformatics. 17:722016.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
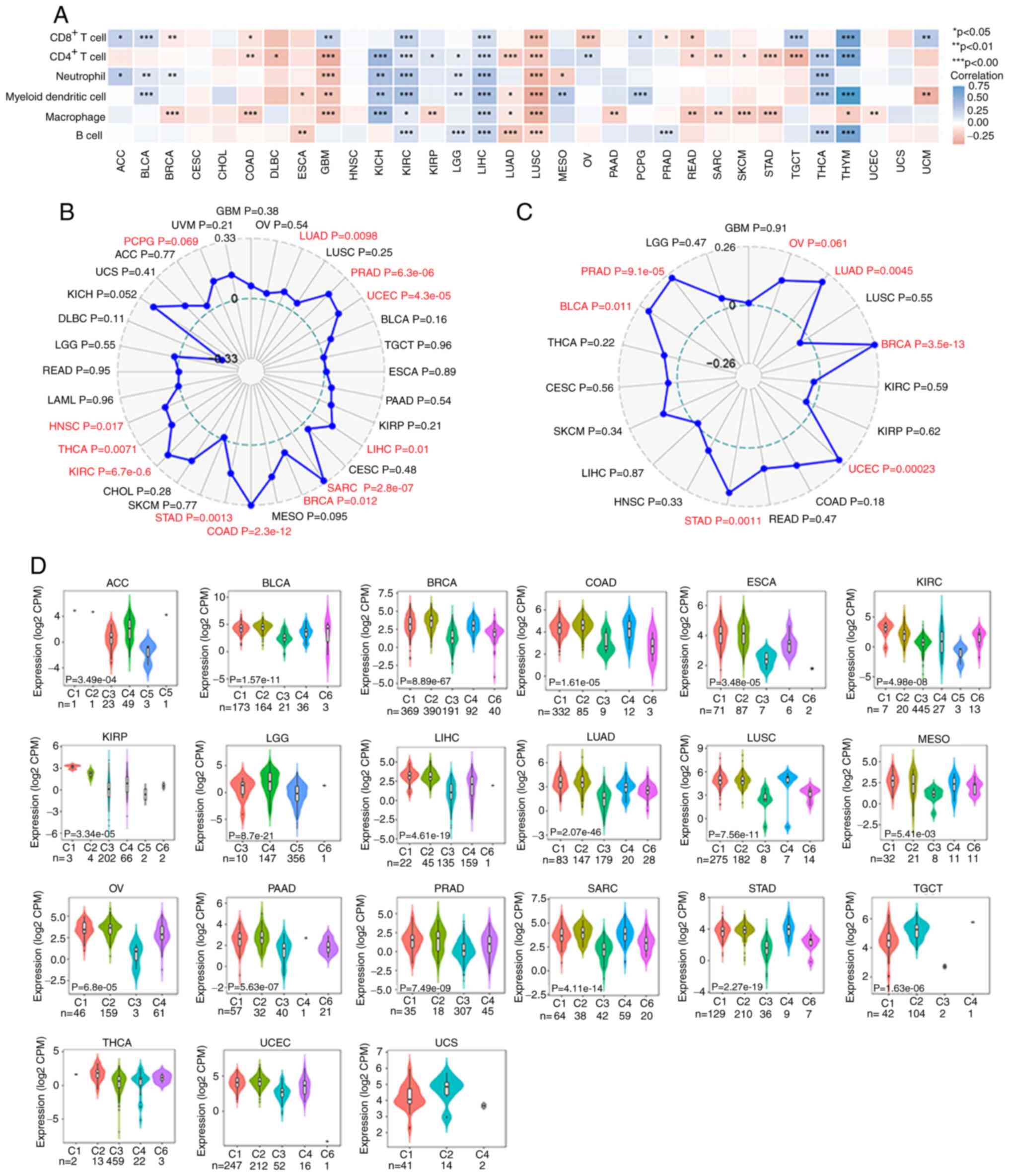
Ru B, Wong CN, Tong Y, Zhong JY, Zhong SS,
Wu WC, Chu KC, Wong CY, Lau CY, Chen I, et al: TISIDB: An
integrated repository portal for tumor-immune system interactions.
Bioinformatics. 35:4200–4202. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Li T, Fu J, Zeng Z, Cohen D, Li J, Chen Q,
Li B and Liu XS: TIMER2.0 for analysis of tumor-infiltrating immune
cells. Nucleic Acids Res. 48:W509–W514. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Thul PJ and Lindskog C: The human protein
atlas: A spatial map of the human proteome. Protein Sci.
27:233–244. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Cerami E, Gao J, Dogrusoz U, Gross BE,
Sumer SO, Aksoy BA, Jacobsen A, Byrne CJ, Heuer ML, Larsson E, et
al: The cBio cancer genomics portal: An open platform for exploring
multidimensional cancer genomics data. Cancer Discov. 2:401–404.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Tang Z, Kang B, Li C, Chen T and Zhang Z:
GEPIA2: An enhanced web server for large-scale expression profiling
and interactive analysis. Nucleic Acids Res. 47:W556–W560. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Chandrashekar DS, Bashel B, Balasubramanya
SA, Creighton CJ, Ponce-Rodriguez I, Chakravarthi BV and Varambally
S: UALCAN: A portal for facilitating tumor subgroup gene expression
and survival analyses. Neoplasia. 19:649–658. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Koch A, Jeschke J, Van Criekinge W, van
Engeland M and De Meyer T: MEXPRESS update 2019. Nucleic Acids Res.
47:W561–W565. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
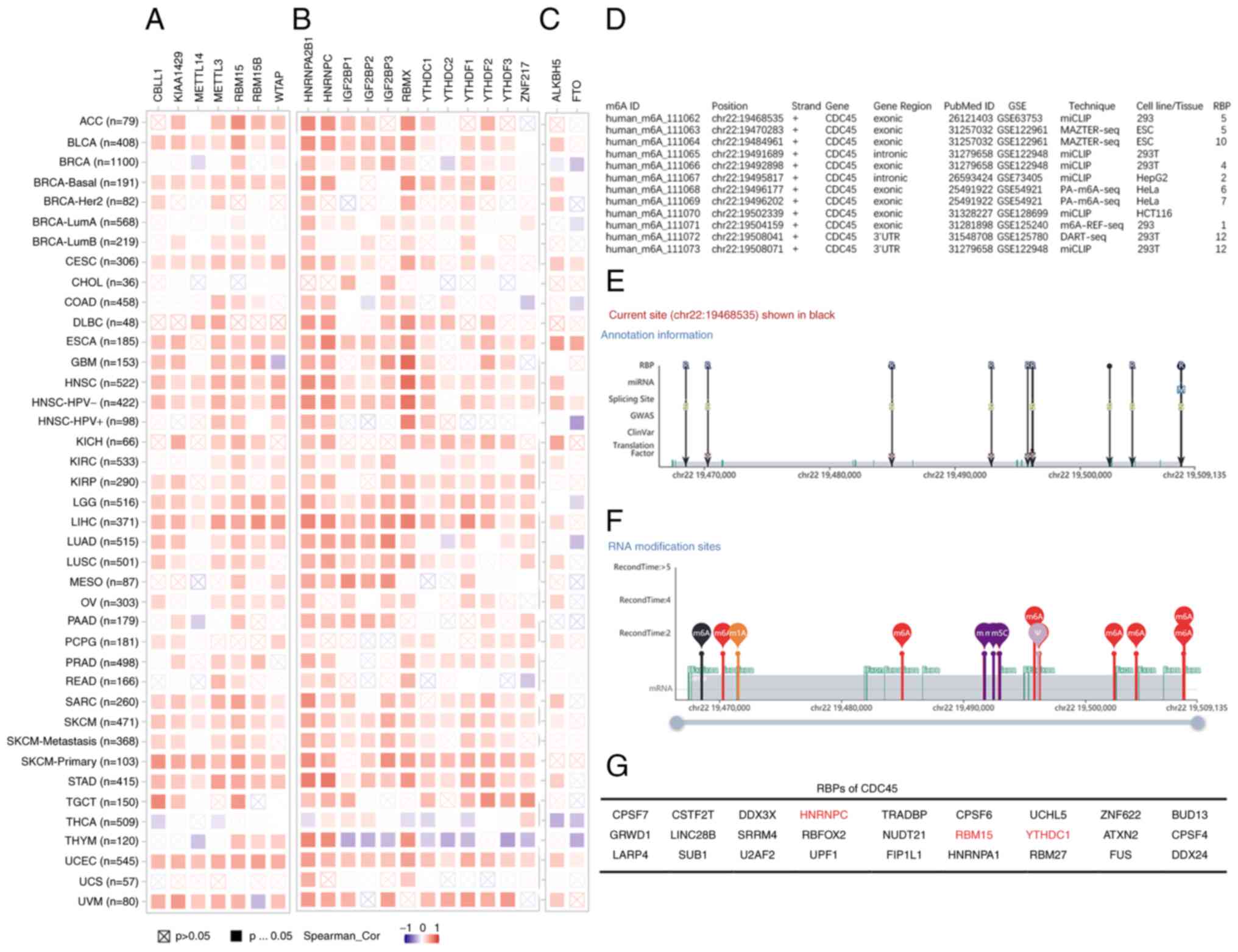
Tang Y, Chen K, Song B, Ma J, Wu X, Xu Q,
Wei Z, Su J, Liu G, Rong R, et al: m6A-Atlas: A comprehensive
knowledgebase for unraveling the N6-methyladenosine (m6A)
epitranscriptome. Nucleic Acids Res. 49:D134–D143. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Liu S, Zhu A, He C and Chen M: REPIC: A
database for exploring the N6-methyladenosine methylome. Genome
Biol. 21:1002020. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
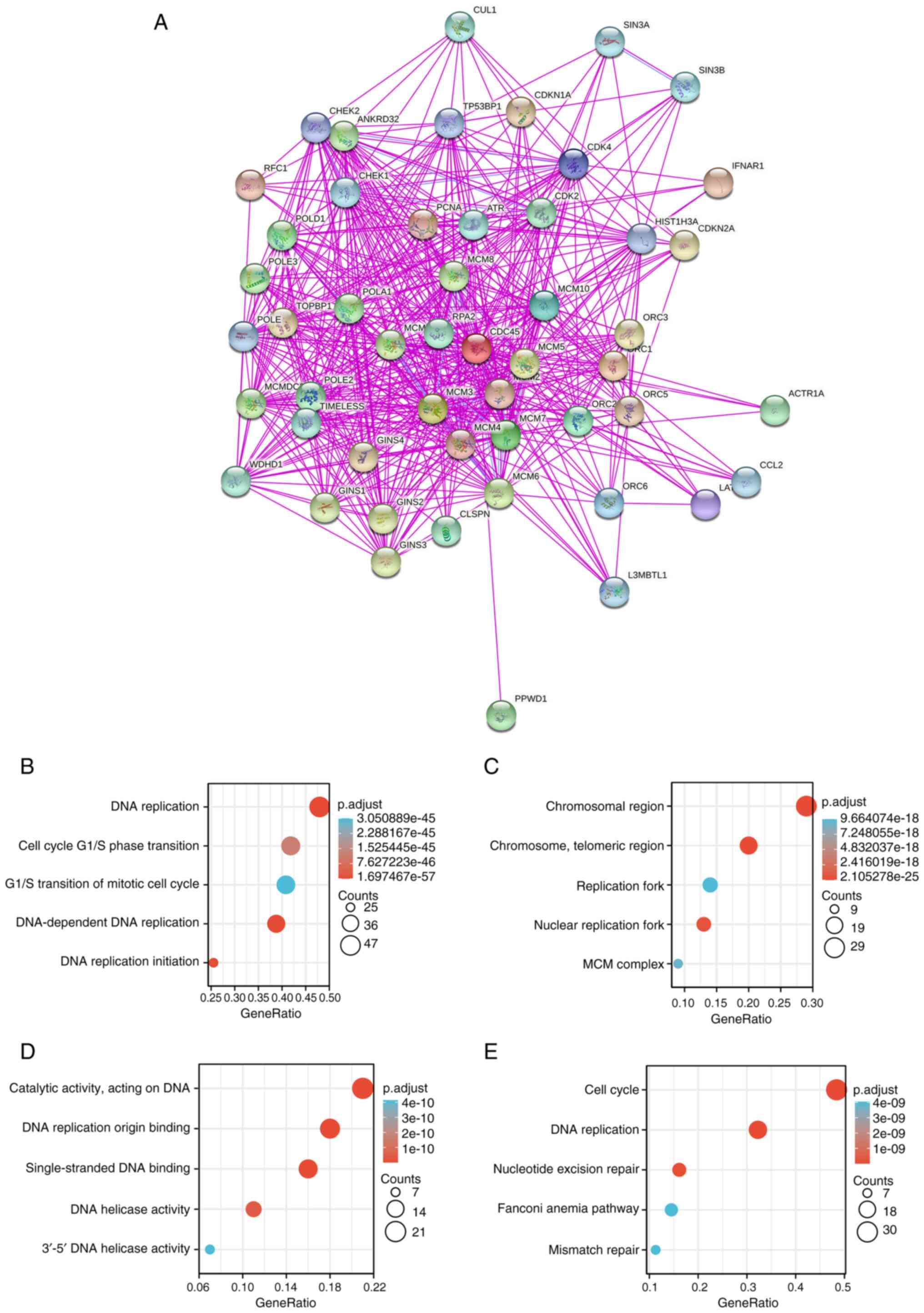
von Mering C, Huynen M, Jaeggi D, Schmidt
S, Bork P and Snel B: STRING: A database of predicted functional
associations between proteins. Nucleic Acids Res. 31:258–261. 2003.
View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Linder B, Grozhik AV, Olarerin-George AO,
Meydan C, Mason CE and Jaffrey SR: Single-nucleotide-resolution
mapping of m6A and m6Am throughout the transcriptome. Nat Methods.
12:767–772. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Herman JG and Baylin SB: Gene silencing in
cancer in association with promoter hypermethylation. N Eng J Med.
349:2042–2054. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Esteller M: Aberrant DNA methylation as a
cancer-inducing mechanism. Annu Rev Pharmacol Toxicol. 45:629–656.
2005. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Fesnak AD, June CH and Levine BL:
Engineered T cells: The promise and challenges of cancer
immunotherapy. Nat Rev Cancer. 16:566–581. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Hu Z, Ott PA and Wu CJ: Towards
personalized, tumour-specific, therapeutic vaccines for cancer. Nat
Rev Immunol. 18:168–182. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
He L, Li H, Wu A, Peng Y, Shu G and Yin G:
Functions of N6-methyladenosine and its role in cancer. Mol Cancer.
18:1762019. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Garcia-Campos MA, Edelheit S, Toth U,
Safra M, Shachar R, Viukov S, Winkler R, Nir R, Lasman L, Brandis
A, et al: Deciphering the ‘m6A Code’ via antibody-independent
quantitative profiling. Cell. 178:731–747.e16. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Gene Ontology Consortium: Gene ontology
consortium: Going forward. Nucleic Acids Res. 43:D1049–D1056. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Broderick R and Nasheuer HP: Regulation of
Cdc45 in the cell cycle and after DNA damage. Biochem Soc Trans.
37:926–930. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Köhler C, Koalick D, Fabricius A, Parplys
AC, Borgmann K, Pospiech H and Grosse F: Cdc45 is limiting for
replication initiation in humans. Cell Cycle. 15:974–985. 2016.
View Article : Google Scholar : PubMed/NCBI
|