Exosomes are membrane-bound microvesicles, ranging
from 30–150 nm in diameter, that carry nucleic acids, proteins,
lipids and metabolites. These vesicles are widely distributed in
various biological fluids and are involved in diverse physiological
and pathological processes (14).
The heterogeneity of exosomes is determined by their size, cargo
composition and cellular origin (15). Exosomes derived from different
tissues and cell types exhibit distinct characteristics, which can
exert divergent effects on CRC (16). Previous studies have highlighted
their therapeutic potential in treating CRC (17–19).
However, it has been reported that tumor-derived exosomes can
promote CRC progression by modulating the activity of diverse
molecular pathways (20).
Therefore, elucidating the dual roles of exosome subpopulations
from different cellular sources in CRC is of clinical
importance.
The vast majority of CRC arise from colorectal
polyps that initially develop into early adenomas. In turn, these
adenomas can progress into advanced adenoma, and ultimately, into
CRC. This process is commonly caused by gene mutations and
typically occurs over a period of 10–15 years. The most commonly
mutated genes in CRC include APC, CTNNB1, KRAS, BRAF, SMAD4,
TGFBR2 and TP53 (21,22).
Mutations in the aforementioned genes can promote the occurrence
and metastasis of CRC by disrupting the functions of key signaling
pathways, including those of Wnt/β-catenin, epidermal growth
factor/MAPK, phosphoinositide-3 kinase (PI3K) and TGF-β (23,24).
Although alterations in the Wnt signaling pathway are commonly
associated with CRC progression, subsequent dysregulation of other
signaling pathways serve a key role in promoting tumor progression
and metastasis (25).
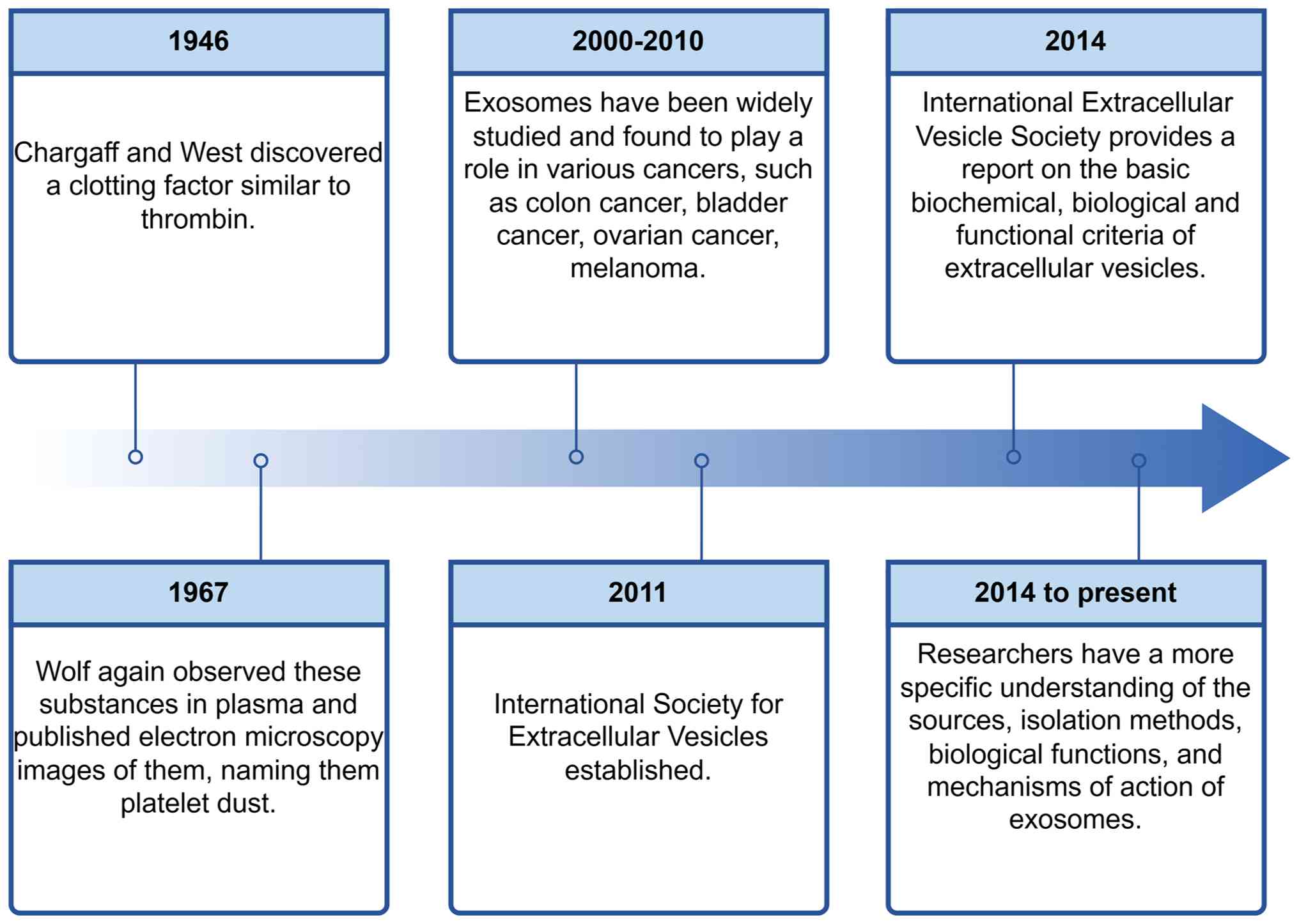
The field of extracellular vesicle (EV) biology
dates to 1946, with Chargaff and West's (26) identification of a thromboplastic
factor in hemophilic blood, now considered the first documented EV
observation. In 1967, Wolf (27)
further characterized these particles in plasma and published their
electron micrographs, coining the term ‘platelet dust’. Although
nomenclature evolved over decades, these entities are now
collectively referred to as EVs. Several conventional markers, such
as CD9, CD63, CD81, TSG101, Alix, Flotillin-1, heat shock cognate
70 kDa protein, actin, major histocompatibility complex (MHC) I and
MHCII are distributed across different EV subtypes, making
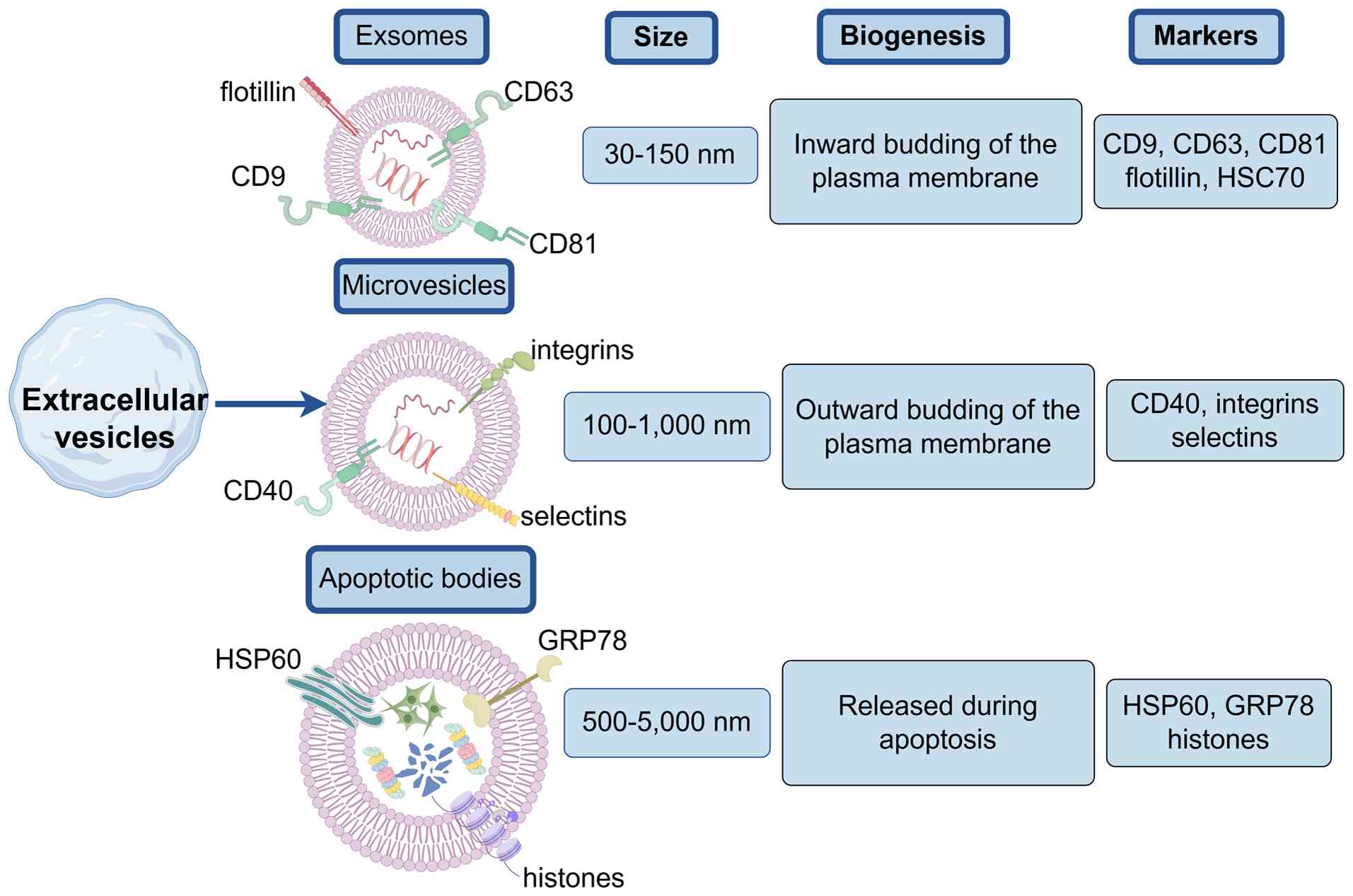
classification based on surface markers difficult. EVs are commonly
classified by size and biogenesis mechanisms into the following
three subtypes: Exosomes (diameter, 30–150 nm), microvesicles
(diameter, 100–1,000 nm) and apoptotic bodies (diameter, 500–5,000
nm; Fig. 1). The experiments by
Harding et al (28) and Pan
et al (29), successively
demonstrated that exosomes are formed by budding inwardly through
the cytoplasmic membrane and subsequently forming polycystic
vesicles. By contrast, microvesicles are released via outward
budding of the plasma membrane, while apoptotic bodies arise during
programmed cell death, the present review only focuses on exosomes.
Between 2000 and 2010, exosomes became a pronounced focus of
investigation. A study conducted in vitro experiments
revealed that exosomes are rich in proteins, lipids and RNA,
including mRNA and microRNA, which enables exosomes to mediate
diverse biological functions (30).
Exosomes are secreted by various cells and are present in almost
all body fluids (31), including
blood, saliva, urine, cerebrospinal fluid and breast milk. However,
exosomes derived from different sources exhibit their own unique
advantages and have been associated with the onset of several types
of cancer, such as colon cancer (32), bladder cancer (33), ovarian cancer (34) and melanoma (35), all of which have been confirmed
in vitro. Exosomes also carry out a notable role in other
diseases, such as neurodegenerative diseases, diabetic
cardiomyopathy, metabolic disorders and ischemic stroke (36–39). A
pivotal advancement occurred in 2011 with the establishment of the
International Society for Extracellular Vesicles (ISEV), which
accelerated exosome research. In 2014, the ISEV issued seminal
guidelines standardizing EV characterization based on biochemical,
biological and functional criteria (40). Since then, substantial progress has
been made in elucidating exosome biogenesis, refining isolation
methodologies and clarifying their biological and mechanistic
functions (Fig. 2).
Different isolation approaches can substantially
affect exosome purity, yield and physicochemical properties
(41). Furthermore, the intrinsic
heterogeneity of exosomes in size, cargo and function presents
challenges for their separation and extraction (42). Therefore, achieving high-purity
exosome isolation with high efficiency remains a major technical
challenge. Based on research, the present review systematically
summarizes the specific separation methods: Ultracentrifugation
(43–45), size-based separation (46–48),
flush separation (49), polymer
precipitation (43,50,51),
ultrafiltration (52–54), immunoaffinity capture (55–57)
and microfluidics technology (58,59),
as well as their respective advantages and disadvantages (Table SI) (43–59).
Ultracentrifugation is currently the most widely
used isolation method and is considered the gold standard for
exosome separation and extraction (60,61).
In 1955, Lathe and Ruthven (62)
proposed a size-based separation method, known as size-exclusion
chromatography (SEC). SEC offers the key advantage of preserving
the natural biological activity of exosomes (63). Cheng et al (49) suggested that although the rinsing
separation method was simple to operate and cost-effective, it
could inevitably lead to the loss of exosomes during processing,
thus resulting in a low yield of extracted exosomes. Polymer
precipitation represents another commonly used exosome isolation
strategy (50). Compared with
ultracentrifugation, ultrafiltration can markedly reduce
preparation time, while it does not require special equipment, thus
it is considered an ideal alternative to traditional
ultracentrifugation (64).
Immunoaffinity capture technology can enable the specific isolation
and extraction of exosomes with high purity and sensitivity
(65). The study by
Salieb-Beugelaar et al (66)
demonstrated that microfluidic technology could efficiently
separate exosomes through microfluidic systems.
Although several methods have been developed for
exosome isolation and extraction, each approach has certain
limitations and no approach can simultaneously achieve high
efficiency, high purity, operational simplicity and low cost.
Consequently, previous studies have increasingly focused on
combining multiple methods for exosome isolation and extraction,
thus yielding promising results (67,68).
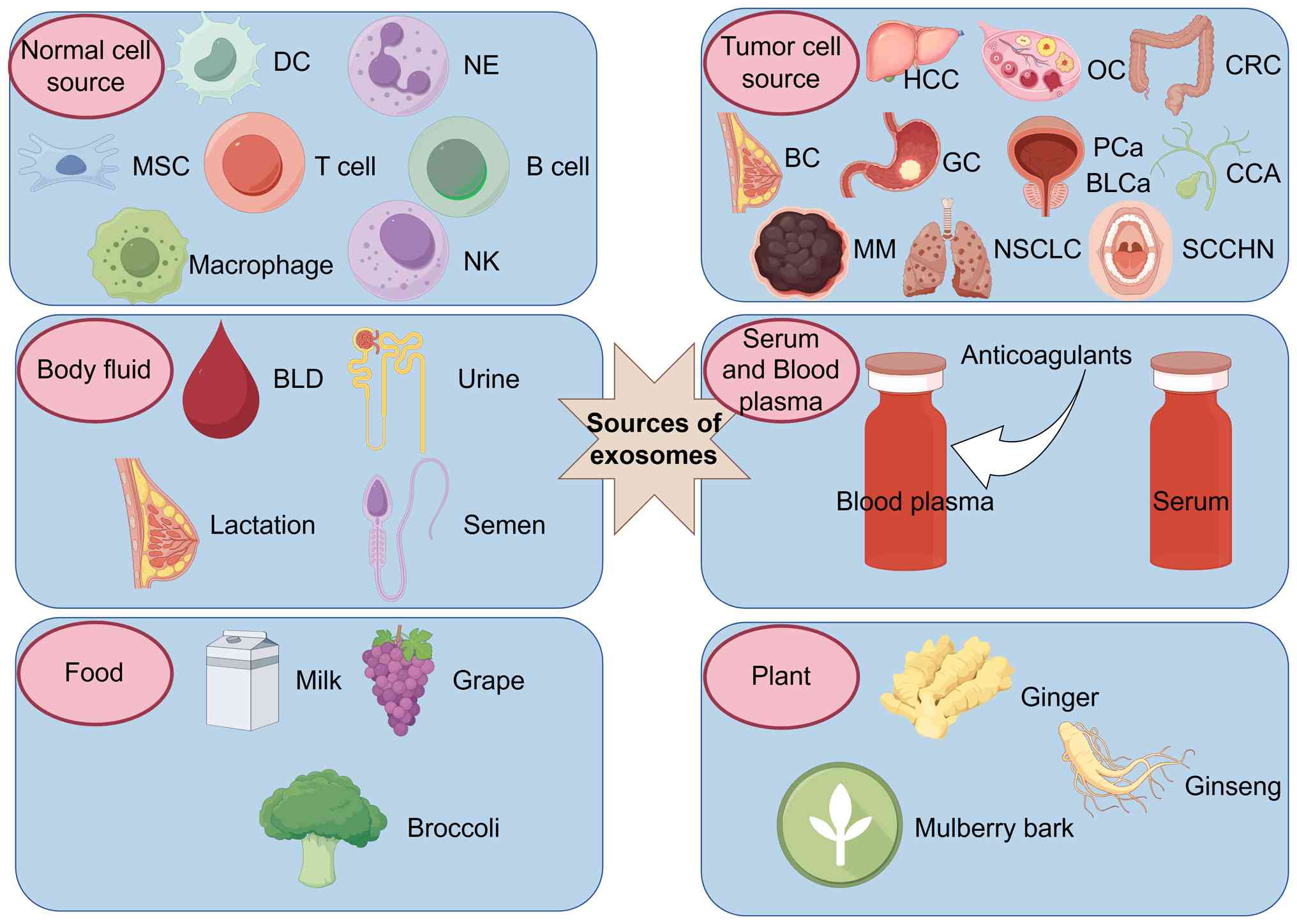
Exosomes arise from a wide range of biological
sources. The present review systematically categorized their
sources into the following: i) Normal cells, including mesenchymal
stem cells, dendritic cells, neutrophils, natural killer cells,
immune cells and macrophages (69–71);
ii) tumor cells, which have been identified in several types of
cancer, such as in hepatocellular carcinoma, ovarian cancer, CRC,
breast cancer, gastric cancer, prostate cancer, bladder cancer,
melanoma, non-small cell lung cancer, cholangiocarcinoma and head
and neck squamous cell carcinoma (69,72,73);
iii) bodily fluids, including blood, urine, breast milk and semen
(74,75); iv) serum/plasma (76–78);
v) food sources, such as milk, grapes and broccoli (79,80);
and vi) plants, such as ginger, mulberry bark, ginseng and galangal
(81–83) (Fig.
3).
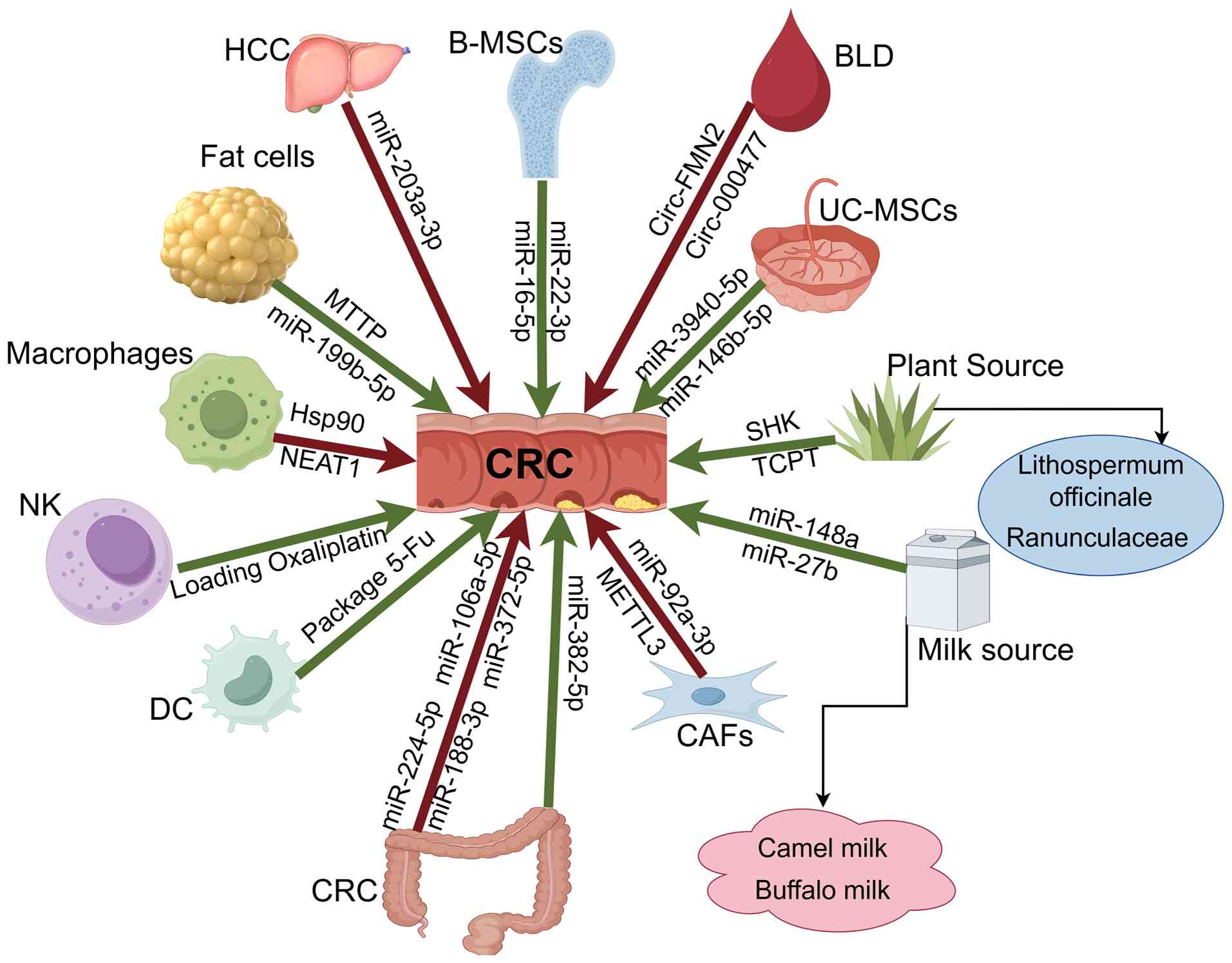
Exosomes, as key mediators of intercellular
communication, play multifaceted and complex roles in the
initiation, progression and metastasis of CRC. Emerging evidence
has suggested that exosomes derived from different cellular origins
exhibit dual functions in CRC (Fig.
4). Therefore, they can promote tumor growth, invasion and
immune evasion. Conversely, they can also suppress tumor growth or
enhance chemosensitivity. This functional duality supports the
important dependence of exosomal activity in CRC on their cellular
source.
In the following section, the bidirectional
regulatory mechanisms of exosomes derived from normal cells
(Table SII) (84–94),
tumor cells (Table SIII) (95–109),
blood (Table SIV) (110,111), food (Table SV) (112–114) and plants (Table SVI) (115,116) were systematically summarized, thus
providing a theoretical basis for the clinical development of novel
diagnostic and therapeutic strategies.
In the study on the effect of exosomes derived from
normal cells on CRC, the present review expounds that exosomes from
the same source could have different effects on CRC through
different mechanisms. For instance, exosomes derived from
adipocytes have been reported to inhibit the progression of CRC by
delivering microsomal triglyceride transfer protein to suppress
lipid reactive oxygen species generation and reduce the
susceptibility of cells to ferroptosis (84). Adipocyte-derived exosomes can also
target jagged 1 by carrying miR-199b-5p to enhance the
radioresistance of CRC cells and promote their proliferation
(85). However, the latter
cancer-promoting effect has only been verified in in vitro
cell models to date (85). Its
in vivo pathological association and potential clinical
application still need to be further confirmed in animal models or
clinical samples that mimic the human physiological environment
more closely. Conversely, multiple studies have suggested that
mesenchymal stem cells (MSCs) [such as those derived from bone
marrow (B)-MSCs (86–88) and umbilical cord (UC)-MSCs (89,90)],
natural killer (NK) cells (91) and
dendritic cells (DCs) (92) exhibit
good anti-tumor potential in CRC models. The strength of the
evidence on which these conclusions are based varies: Among them,
the functions of UC-MSCs, NK cells, DC cells and macrophage-derived
exosomes have been verified in in vivo mouse models
(89–94); the mechanism of action of B-MSCs
exosomes is currently mainly based on in vitro experimental
data and the reliability of their in vivo behavior still
needs further exploration (86–88).
Exosomes derived from macrophages exhibit different functions from
those of other immune cells, such as NK cells mentioned above
(93,94). They can promote the occurrence and
development of CRC through mechanisms such as inhibiting the Hippo
signaling pathway and adsorbing miR-34a-5p (93,94).
In CRC, exosomes from tumor cells mainly arise from
three types of cells: CRC cells, cancer associated fibroblasts
(CAFs) and hepatocellular carcinoma (HCC), exosomes from different
sources can promote the occurrence and metastasis of CRC through
multiple mechanisms in synergy (117–119).
Exosomes derived from CAFs also carry out a key role
in drug resistance and tumor progression. The mechanisms verified
in in vivo mouse models include: Exosome miR-92a-3p which
can activate the Wnt/β-catenin pathway and block mitochondrial
apoptosis by directly inhibiting F-box and WD repeat domain
containing 7 and modulator of apoptosis 1, thereby inducing
chemotherapy resistance in CRC cells (20). Methyltransferase-like 3 which can
induce Acyl-CoA synthetase long-chain family member 3 m6A
modification and stabilize its expression, promote the
proliferation and metastasis of CRC, and inhibit ferroptosis
(106). In addition, a study shows
that circ-0067557 not only participates in drug resistance, but
also promotes the occurrence, metastasis and chemical resistance of
CRC through the Lin28A/Lin28B pathway (107). Conversely, a study that used in
vitro cell models found that CAFs can also target miR-330-3p
through TP53 target 1 carried out by exosomes, enhance the activity
of CRC cells, promote epithelial-mesenchymal transition and inhibit
apoptosis, thereby promoting tumor progression (108).
In addition, exosomes derived from HCC are also
involved in regulating the progression of CRC. As demonstrated by a
study using an in vivo mouse animal model, HCC-derived
exosomes inhibit the expression of the sarcoma gene through
miR-203a-3p, thereby upregulating the level of E-cadherin and
ultimately promoting the occurrence and development of CRC
(109).
Exosomes derived from blood exhibit a promoting
effect on the development of CRC through different regulatory
mechanisms. For instance, a study using in vivo mouse
models, revealed that blood-derived exosomes promote the occurrence
and progression of CRC by mediating the miR-338-3p/musashi
RNA-binding protein-1 axis through circ-FMN2 (110). However, another study using only
in vitro cell models indicated that blood-derived exosomes
might regulate the miR-653/zinc finger E-box binding homeobox 2
(ZEB2) pathway through circ-0004771100 and be involved in the
occurrence of 5-FU resistance (111).
Food-derived exosomes have demonstrated excellent
potential to inhibit the development of CRC. For instance, a study
using an in vivo mouse model demonstrated that exosomes
derived from camel milk can reduce the expression of TNF-α and IL-6
genes in CT-26 cells (112).
Exosomes derived from milk target DNA methylation transferase-1
through miR-148a to inhibit the activity of CRC activators
(113). Another study using only
in vitro cell models demonstrated that exosomes derived from
buffalo milk exacerbate endoplasmic reticulum stress through
miR-27b, causing the death of CRC cells (114). To the best of our knowledge,
clinical evidence is still lacking regarding the role of
food-derived exosomes in CRC.
Currently, clinical treatment strategies for CRC
include early-stage endoscopic or surgical resection and for
advanced stages, radiotherapy, chemotherapy, immunotherapy and
targeted therapy (120,121). Patient survival for patients with
CRC is associated with early diagnosis and treatment. However,
since CRC is not easily detected in its early stages, several
patients are diagnosed at advanced stages (122). Colonoscopy still remains the gold
standard for the early diagnosis of CRC, however, due to its
invasive nature, several patients miss the opportunity for early
diagnosis. Therefore, there is an urgent need for reliable,
non-invasive biomarkers capable of predicting and diagnosing CRC at
an early stage. In recent years, exosomes have attracted attention
as potential diagnostic tools, since they carry different molecules
that reflect their origin (123–125).
Previous studies demonstrated that the expression
levels of long non-coding RNAs NAMPT-AS (126) and LINC02418 (127), carried by serum exosomes, were
markedly increased in patients with CRC, thus highlighting their
potential as promising biomarkers for the diagnosis of CRC. In the
study by Fabijanec et al (128), plasma exosome-derived microRNA
(miR)-193a-3p was identified as a valuable biomarker for
distinguishing patients with CRC from those with colorectal
adenomas. In addition, the elevated levels of miR-461 (129), miR-205-5p (130), miR-6803-5p (131), miR-27a, miR-130a (132), as well as the reduced levels of
miR-377-3p, miR-381-3p (133),
miR-139-3p (134), miR-150-5p and
miR-99b-5p (135), were associated
with the initiation and progression of CRC. Notably, Wang et
al (136) revealed that
miR-125a-3p upregulation was particularly associated with the
occurrence of early-stage colon cancer. Additionally, the increased
levels of circular RNAs GAPVD1 (137) and PNN were also involved in CRC
development (138). Furthermore,
differential protein expression profiles of exosomes isolated from
ascites (139) and urine (140) could serve as effective diagnostic
indicators for CRC, consistent with the findings reported by Ma
et al (141).
Exosomes can carry several bioactive molecules and
are involved in intercellular communication. Targeted inhibition of
tumor-promoting exosome production, release or uptake can
effectively regulate tumor cell proliferation, invasion and
metastasis (142). Additionally,
exosomes exhibit excellent biocompatibility, high stability and
homing capacity, thus making them ideal nanocarriers for
efficiently delivering therapeutic drugs in CRC (143). Therefore, exosomes derived from
different sources hold considerable potential and value in CRC
treatment.
The present review systematically summarizes the key
molecules and mechanisms underlying the dual effects of exosomes
derived from different sources on CRC, with the aim of advancing
their diagnostic and therapeutic potential. By integrating current
evidence, clinicians could gain a deeper understanding of the
biological characteristics of exosomes, thereby providing
additional strategies for the diagnosis and treatment of CRC.
However, prior to their wide application in clinical practice, key
challenges remain. First, the lack of technology and standards, the
separation and purification techniques of exosomes still have
limitations. It is difficult to efficiently obtain exosomes from
specific cell sources from complex biological samples, and the
existing methods face difficulties in large-scale production. More
importantly, the lack of a unified standardized scheme, from
separation and identification to functional analysis, may lead to
poor comparability and repeatability among different research
results. Second, the extreme complexity of the biological functions
of exosomes. The function of exosomes is not static but highly
dependent on the tumor microenvironment, disease stage and the
physiological and pathological state of the body. This
characteristic brings great difficulties to the precise
interpretation of its function. Third, the insufficiency of
evidence for clinical transformation. At present, preclinical and
clinical research on the application of exosomes in the treatment
of CRC is still very limited, and their in vivo safety,
targeted delivery efficiency and long-term efficacy still need to
be systematically verified.
To accelerate the translation of exosome research in
clinical applications, several key steps should be considered.
First, to promote technical standardization and overcome the
problem of sample heterogeneity, the top priority in the future is
to establish recognized technical standards. This includes
formulating standard separation operation procedures for different
sample sources (such as plasma and tissue fluid) and utilizing
high-resolution sorting techniques (such as immunoaffinity capture
and microfluidic technology) to address the challenge of ‘sample
source variability’, thereby achieving high-purity acquisition of
specific exosome subpopulations. Second, deeply explore the
separation and purification technology of exosomes could ensure
efficient isolation and application. Third, in-depth studies should
be performed to evaluate the dynamic changes and precise mechanisms
of exosomes during different stages of CRC, including initiation,
progression, metastasis and drug resistance. Fourth, systematic
research should be conducted to identify and validate exosomal
biomarkers, thus promoting standardized classification and
functional analyses. Finally, preclinical and clinical trials are
urgently needed to evaluate the safety and efficacy of
exosome-based applications in CRC.
In summary, exosomes derived from different sources
play pivotal roles in CRC, thus providing promising opportunities
for their clinical application (149). However, their dual function on CRC
progression, both promoting and repressing the development of CRC,
is considerably associated with their heterogeneous origin
(84,93). Therefore, a comprehensive
understanding of this source-dependent functional heterogeneity is
of great importance for advancing CRC therapy. Despite their
notable potential, the clinical application of exosomes is hindered
by notable challenges, including their complex molecular mechanisms
and the lack of standardized isolation protocols. Collectively,
exosomes represent a powerful tool for deepening the understanding
of CRC biology and oncology, while offering novel directions and
strategies for the development of innovative diagnostic and
therapeutic approaches.
Not applicable.
Funding: No funding was received.
Not applicable.
ZG, HZ and QL contributed to the conception and
overall design of the study. ZG and HZ drafted the manuscript and
prepared the figures and tables. QL reviewed and revised the
manuscript. All authors read and approved the final version of the
manuscript. Data authentication was not applicable.
Not applicable.
Not applicable.
The authors declare that they have no competing
interests.
|
1
|
Bray F, Ferlay J, Soerjomataram I, Siegel
RL, Torre LA and Jemal A: Global cancer statistics 2018: GLOBOCAN
estimates of incidence and mortality worldwide for 36 cancers in
185 countries. CA Cancer J Clin. 68:394–424. 2018.PubMed/NCBI
|
|
2
|
GBD 2019 Colorectal Cancer Collaborators,
. Global, regional, and national burden of colorectal cancer and
its risk factors, 1990–2019: A systematic analysis for the Global
Burden of disease study 2019. Lancet Gastroenterol Hepatol.
7:627–647. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Mauri G, Patelli G, Crisafulli G, Siena S
and Bardelli A: Tumor ‘age’ in early-onset colorectal cancer. Cell.
188:589–593. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Zhou J, Yang Q, Zhao S, Sun L, Li R, Wang
J, Wang L and Wang D: Evolving landscape of colorectal cancer:
Global and regional burden, risk factor dynamics, and future
scenarios (the Global Burden of Disease 1990–2050). Ageing Res Rev.
104:1026662025. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Kim BJ and Hanna MH: Colorectal cancer in
young adults. J Surg Oncol. 127:1247–1251. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Bouvard V, Loomis D, Guyton KZ, Grosse Y,
Ghissassi FE, Benbrahim-Tallaa L, Guha N, Mattock H and Straif K;
International Agency for Research on Cancer Monograph Working
Group, : Carcinogenicity of consumption of red and processed meat.
Lancet Oncol. 16:1599–1600. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Johnson CH, Dejea CM, Edler D, Hoang LT,
Santidrian AF, Felding BH, Ivanisevic J, Cho K, Wick EC,
Hechenbleikner EM, et al: Metabolism links bacterial biofilms and
colon carcinogenesis. Cell Metab. 21:891–897. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Dejea CM, Wick EC, Hechenbleikner EM,
White JR, Mark Welch JL, Rossetti BJ, Peterson SN, Snesrud EC,
Borisy GG, Lazarev M, et al: Microbiota organization is a distinct
feature of proximal colorectal cancers. Proc Natl Acad Sci USA.
111:18321–18326. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Rosato V, Bosetti C, Levi F, Polesel J,
Zucchetto A, Negri E and La Vecchia C: Risk factors for young-onset
colorectal cancer. Cancer Cause Control. 24:335–341.
2013.PubMed/NCBI
|
|
10
|
Zhang Y, Luo J, Yang W and Ye WC: CircRNAs
in colorectal cancer: Potential biomarkers and therapeutic targets.
Cell Death Dis. 14:3532023. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Yang W, Zheng H, Lv W and Zhu Y: Current
status and prospect of immunotherapy for colorectal cancer. Int J
Colorectal Dis. 38:2662023. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Abedizadeh R, Majidi F, Khorasani HR,
Abedi H and Sabour D: Colorectal cancer: A comprehensive review of
carcinogenesis, diagnosis, and novel strategies for classified
treatments. Cancer Metastasis Rev. 43:729–753. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Arjmand B, Alavi-Moghadam S, Faraji Z,
Aghajanpoor-Pasha M, Jalaeikhoo H, Rajaeinejad M, Nikandish M,
Faridfar A, Rezazadeh-Mafi A, Rezaei-Tavirani M and Irompour A: The
potential role of intestinal stem cells and microbiota for the
treatment of colorectal cancer. Adv Exp Med Biol. 1470:115–128.
2024. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Logozzi M, Di Raimo R, Mizzoni D and Fais
S: What we know on the potential use of exosomes for nanodelivery.
Semin Cancer Biol. 86:13–25. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Lai RC, Yeo RW, Tan KH and Lim SK:
Exosomes for drug delivery-a novel application for the mesenchymal
stem cell. Biotechnol Adv. 31:543–551. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Lai RC, Chen TS and Lim SK: Mesenchymal
stem cell exosome: A novel stem cell-based therapy for
cardiovascular disease. Regener Med. 6:481–492. 2011. View Article : Google Scholar
|
|
17
|
Nabariya DK, Pallu R and Yenuganti VR:
Exosomes: The protagonists in the tale of colorectal cancer?
Biochim Biophys Acta Rev Cancer. 1874:1884262020. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Zhang M, Hu S, Liu L, Dang P, Liu Y, Sun
Z, Qiao B and Wang C: Engineered exosomes from different sources
for cancer-targeted therapy. Signal Transduct Target Ther.
8:1242023. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Guo G, Tan Z, Liu Y, Shi F and She J: The
therapeutic potential of stem cell-derived exosomes in the
ulcerative colitis and colorectal cancer. Stem Cell Res Ther.
13:1382022. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Hu JL, Wang W, Lan XL, Zeng ZC, Liang YS,
Yan YR, Song FY, Wang FF, Zhu XH, Liao WJ, et al: CAFs secreted
exosomes promote metastasis and chemotherapy resistance by
enhancing cell stemness and epithelial-mesenchymal transition in
colorectal cancer. Mol Cancer. 18:912019. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Brennan CW, Verhaak RG, McKenna A, Campos
B, Noushmehr H, Salama SR, Zheng S, Chakravarty D, Sanborn JZ,
Berman SH, et al: The somatic genomic landscape of glioblastoma.
Cell. 155:462–477. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Grady WM and Pritchard CC: Molecular
alterations and biomarkers in colorectal cancer. Toxicol Pathol.
42:124–139. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Parsons DW, Wang TL, Samuels Y, Bardelli
A, Cummins JM, DeLong L, Silliman N, Ptak J, Szabo S, Willson JK,
et al: Colorectal cancer: mutations in a signalling pathway.
Nature. 436:7922005. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Bardelli A, Parsons DW, Silliman N, Ptak
J, Szabo S, Saha S, Markowitz S, Willson JK, Parmigiani G, Kinzler
KW, et al: Mutational analysis of the tyrosine kinome in colorectal
cancers. Science. 300:9492003. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Kuipers EJ, Grady WM, Lieberman D,
Seufferlein T, Sung JJ, Boelens PG, van de Velde CJ and Watanabe T:
Colorectal cancer. Nat Rev Dis Primers. 1:150652015. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Chargaff E and West R: The biological
significance of the thromboplastic protein of blood. J Biol Chem.
166:189–197. 1946. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Wolf P: The nature and significance of
platelet products in human plasma. Br J Haematol. 13:269–288. 1967.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Harding C, Heuser J and Stahl P:
Receptor-mediated endocytosis of transferrin and recycling of the
transferrin receptor in rat reticulocytes. J Cell Biol. 97:329–339.
1983. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Pan BT, Teng K, Wu C, Adam M and Johnstone
RM: Electron microscopic evidence for externalization of the
transferrin receptor in vesicular form in sheep reticulocytes. J
Cell Biol. 101:942–948. 1985. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Alzhrani GN, Alanazi ST, Alsharif SY,
Albalawi AM, Alsharif AA, Abdel-Maksoud MS and Elsherbiny N:
Exosomes: Isolation, characterization, and biomedical applications.
Cell Biol Int. 45:1807–1831. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Simpson RJ, Kalra H and Mathivanan S:
ExoCarta as a resource for exosomal research. J Extracell Vesicles.
1:183742012. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Mathivanan S, Lim JWE, Tauro BJ, Ji H,
Moritz RL and Simpson RJ: Proteomics analysis of A33
immunoaffinity-purified exosomes released from the human colon
tumor cell line LIM1215 reveals a tissue-specific protein
signature. Mol Cell Proteomics. 9:197–208. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Welton JL, Khanna S, Giles PJ, Brennan P,
Brewis IA, Staffurth J, Mason MD and Clayton A: Proteomics analysis
of bladder cancer exosomes. Mol Cell Proteomics. 9:1324–1338. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Runz S, Keller S, Rupp C, Stoeck A, Issa
Y, Koensgen D, Mustea A, Sehouli J, Kristiansen G and Altevogt P:
Malignant ascites-derived exosomes of ovarian carcinoma patients
contain CD24 and EpCAM. Gynecol Oncol. 107:563–571. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Mears R, Craven RA, Hanrahan S, Totty N,
Upton C, Young SL, Patel P, Selby PJ and Banks RE: Proteomic
analysis of melanoma-derived exosomes by two-dimensional
polyacrylamide gel electrophoresis and mass spectrometry.
Proteomics. 4:4019–4031. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Sivalingam AM and Sureshkumar DD: Exosomes
in regulating miRNAs for biomarkers of neurodegenerative disorders.
Mol Neurobiol. 62:7576–7596. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Gao S, Dong Y, Yan C, Yu T and Cao H: The
role of exosomes and exosomal microRNA in diabetic cardiomyopathy.
Front Endocrinol (Lausanne). 14:13274952024. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Mei R, Qin W, Zheng Y, Wan Z and Liu L:
Role of adipose tissue derived exosomes in metabolic disease. Front
Endocrinol (Lausanne). 13:8738652022. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Li Y, Tang Y and Yang GY: Therapeutic
application of exosomes in ischaemic stroke. Stroke Vasc Neurol.
6:483–495. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Lötvall J, Hill AF, Hochberg F, Buzás EI,
Di Vizio D, Gardiner C, Gho YS, Kurochkin IV, Mathivanan S,
Quesenberry P, et al: Minimal experimental requirements for
definition of extracellular vesicles and their functions: A
position statement from the International society for extracellular
vesicles. J Extracell Vesicles. 3:269132014. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Zhang Y, Bi J, Huang J, Tang Y, Du S and
Li P: Exosome: A review of its classification, isolation
techniques, storage, diagnostic and targeted therapy applications.
Int J Nanomedicine. 15:6917–6934. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Kalluri R and LeBleu VS: The biology,
function, and biomedical applications of exosomes. Science.
367:eaau69772020. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Chia BS, Low YP, Wang Q, Li P and Gao Z:
Advances in exosome quantification techniques. Trends Anal Chem.
86:93–106. 2017. View Article : Google Scholar
|
|
44
|
Langevin SM, Kuhnell D, Orr-Asman MA,
Biesiada J, Zhang X, Medvedovic M and Thomas HE: Balancing yield,
purity and practicality: A modified differential
ultracentrifugation protocol for efficient isolation of small
extracellular vesicles from human serum. RNA Biol. 16:5–12. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Gardiner C, Di Vizio D, Sahoo S, Théry C,
Witwer KW, Wauben M and Hill AF: Techniques used for the isolation
and characterization of extracellular vesicles: Results of a
worldwide survey. J Extracell Vesicles. 5:329452016. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Karimi N, Cvjetkovic A, Jang SC,
Crescitelli R, Hosseinpour Feizi MA, Nieuwland R, Lötvall J and
Lässer C: Detailed analysis of the plasma extracellular vesicle
proteome after separation from lipoproteins. Cell Mol Life Sci.
75:2873–2886. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Navajas R, Corrales FJ and Paradela A:
Serum exosome isolation by size-exclusion chromatography for the
discovery and validation of preeclampsia-associated biomarkers.
Methods Mol Biol. 1959:39–50. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Witwer KW, Buzás EI, Bemis LT, Bora A,
Lässer C, Lötvall J, Nolte-'t Hoen EN, Piper MG, Sivaraman S, Skog
J, et al: Standardization of sample collection, isolation and
analysis methods in extracellular vesicle research. J Extracell
Vesicles. 2:2013. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Cheng H, Fang H, Xu RD, Fu MQ, Chen L,
Song XY, Qian JY, Zou YZ, Ma JY and Ge JB: Development of a rinsing
separation method for exosome isolation and comparison to
conventional methods. Eur Rev Med Pharmacol Sci. 23:5074–5083.
2019.PubMed/NCBI
|
|
50
|
Soares Martins T, Catita J, Martins Rosa
I, A B da Cruz E Silva O and Henriques AG: Exosome isolation from
distinct biofluids using precipitation and column-based approaches.
PLoS One. 13:e01988202018. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
García-Romero N, Madurga R, Rackov G,
Palacín-Aliana I, Núñez-Torres R, Asensi-Puig A, Carrión-Navarro J,
Esteban-Rubio S, Peinado H, González-Neira A, et al: Polyethylene
glycol improves current methods for circulating extracellular
vesicle-derived DNA isolation. J Transl Med. 17:752019. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
He L, Zhu D, Wang J and Wu X: A highly
efficient method for isolating urinary exosomes. Int J Mol Med.
43:83–90. 2019.PubMed/NCBI
|
|
53
|
Konoshenko MY, Lekchnov EA, Vlassov AV and
Laktionov PP: Isolation of extracellular vesicles: General
methodologies and latest trends. Biomed Res Int. 2018:85453472018.
View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Lobb RJ, Becker M, Wen SW, Wong CSF,
Wiegmans AP, Leimgruber A and Möller A: Optimized exosome isolation
protocol for cell culture supernatant and human plasma. J Extracell
Vesicles. 4:270312015. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Fitzgerald J, Leonard P, Darcy E, Sharma S
and O'Kennedy R: Immunoaffinity Chromatography: Concepts and
Applications. Methods Mol Biol. 1485:27–51. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Li P, Kaslan M, Lee SH, Yao J and Gao Z:
Progress in exosome isolation techniques. Theranostics. 7:789–804.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Zarovni N, Corrado A, Guazzi P, Zocco D,
Lari E, Radano G, Muhhina J, Fondelli C, Gavrilova J and Chiesi A:
Integrated isolation and quantitative analysis of exosome shuttled
proteins and nucleic acids using immunocapture approaches. Methods.
87:46–58. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Jackson EL and Lu H: Advances in
microfluidic cell separation and manipulation. Curr Opin Chem Eng.
2:398–404. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
59
|
Gholizadeh S, Shehata Draz M, Zarghooni M,
Sanati-Nezhad A, Ghavami S, Shafiee H and Akbari M: Microfluidic
approaches for isolation, detection, and characterization of
extracellular vesicles: Current status and future directions.
Biosens Bioelectron. 91:588–605. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
60
|
Johnstone RM, Bianchini A and Teng K:
Reticulocyte maturation and exosome release: Transferrin receptor
containing exosomes shows multiple plasma membrane functions.
Blood. 74:1844–1851. 1989. View Article : Google Scholar : PubMed/NCBI
|
|
61
|
Johnstone RM: Revisiting the road to the
discovery of exosomes. Blood Cells Mol Dis. 34:214–219. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
62
|
Lathe GH and Ruthven CR: The separation of
substances on the basis of their molecular weights, using columns
of starch and water. Biochem J. 60:xxxiv1955.PubMed/NCBI
|
|
63
|
Gámez-Valero A, Monguió-Tortajada M,
Carreras-Planella L, Franquesa MI, Beyer K and Borràs FE:
Size-Exclusion Chromatography-based isolation minimally alters
Extracellular Vesicles' characteristics compared to precipitating
agents. Sci Rep. 6:336412016. View Article : Google Scholar : PubMed/NCBI
|
|
64
|
Yu LL, Zhu J, Liu JX, Jiang F, Ni WK, Qu
LS, Ni RZ, Lu CH and Xiao MB: A comparison of traditional and novel
methods for the separation of exosomes from human samples. Biomed
Res Int. 2018:36345632018. View Article : Google Scholar : PubMed/NCBI
|
|
65
|
Greening DW, Xu R, Ji H, Tauro BJ and
Simpson RJ: A protocol for exosome isolation and characterization:
evaluation of ultracentrifugation, density-gradient separation, and
immunoaffinity capture methods. Methods Mol Biol. 1295:179–209.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
66
|
Salieb-Beugelaar GB, Simone G, Arora A,
Philippi A and Manz A: Latest developments in microfluidic cell
biology and analysis systems. Anal Chem. 82:4848–4864. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
67
|
Koh YQ, Almughlliq FB, Vaswani K, Peiris
HN and Mitchell MD: Exosome enrichment by ultracentrifugation and
size exclusion chromatography. Front Biosci (Landmark Ed).
23:865–874. 2018. View
Article : Google Scholar : PubMed/NCBI
|
|
68
|
Ryu KJ, Lee JY, Park C, Cho D and Kim SJ:
Isolation of small extracellular vesicles from human serum using a
combination of ultracentrifugation with polymer-based
precipitation. Ann Lab Med. 40:253–258. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
69
|
Zhang S, Duan Z, Liu F, Wu Q, Sun X and Ma
H: The impact of exosomes derived from distinct sources on
rheumatoid arthritis. Front Immunol. 14:12407472023. View Article : Google Scholar : PubMed/NCBI
|
|
70
|
Xu F, Zhang Q, Liu Y, Tang R, Li H, Yang H
and Lin L: The role of exosomes derived from various sources in
facilitating the healing of chronic refractory wounds. Pharmacol
Res. 216:1077532025. View Article : Google Scholar : PubMed/NCBI
|
|
71
|
Si C, Gao J and Ma X: Natural killer
cell-derived exosome-based cancer therapy: From biological roles to
clinical significance and implications. Mol Cancer. 23:1342024.
View Article : Google Scholar : PubMed/NCBI
|
|
72
|
Dai X, Ye Y and He F: Emerging innovations
on exosome-based onco-therapeutics. Front Immunol. 13:8652452022.
View Article : Google Scholar : PubMed/NCBI
|
|
73
|
Liang B, Peng P, Chen S, Li L, Zhang M,
Cao D, Yang J, Li H, Gui T, Li X and Shen K: Characterization and
proteomic analysis of ovarian cancer-derived exosomes. J
Proteomics. 80:171–182. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
74
|
Panfoli I, Granata S, Candiano G, Verlato
A, Lombardi G, Bruschi M and Zaza G: Analysis of urinary exosomes
applications for rare kidney disorders. Expert Rev Proteomics.
17:735–749. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
75
|
Di SJ, Cui XW, Liu TJ and Shi YY:
Therapeutic potential of human breast milk-derived exosomes in
necrotizing enterocolitis. Mol Med. 30:2432024. View Article : Google Scholar : PubMed/NCBI
|
|
76
|
Cai H, Pang Y, Wang Q, Qin W, Wei C, Li Y,
Li T, Li F, Wang Q, Li Y, et al: Proteomic profiling of circulating
plasma exosomes reveals novel biomarkers of Alzheimer's disease.
Alzheimers Res Ther. 14:1812022. View Article : Google Scholar : PubMed/NCBI
|
|
77
|
Muller L, Hong CS, Stolz DB, Watkins SC
and Whiteside TL: Isolation of biologically-active exosomes from
human plasma. J Immunol Methods. 411:55–65. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
78
|
Ibsen SD, Wright J, Lewis JM, Kim S, Ko
SY, Ong J, Manouchehri S, Vyas A, Akers J, Chen CC, et al: Rapid
isolation and detection of exosomes and associated biomarkers from
plasma. ACS Nano. 11:6641–6651. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
79
|
Kim NH, Kim J, Lee JY, Bae HA and Kim CY:
Application of milk exosomes for musculoskeletal health: talking
points in recent outcomes. Nutrients. 15:46452023. View Article : Google Scholar : PubMed/NCBI
|
|
80
|
Karabay AZ, Barar J, Hekmatshoar Y and
Rahbar Saadat Y: Multifaceted therapeutic potential of
plant-derived exosomes: Immunomodulation, anticancer, anti-aging,
anti-melanogenesis, detoxification, and drug delivery.
Biomolecules. 15:3942025. View Article : Google Scholar : PubMed/NCBI
|
|
81
|
Cao M, Diao N, Cai X, Chen X, Xiao Y, Guo
C, Chen D and Zhang X: Plant exosome nanovesicles (PENs): Green
delivery platforms. Mater Horiz. 10:3879–3894. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
82
|
Yi Q, Xu Z, Thakur A, Zhang K, Liang Q,
Liu Y and Yan Y: Current understanding of plant-derived
exosome-like nanoparticles in regulating the inflammatory response
and immune system microenvironment. Pharmacol Res. 190:1067332023.
View Article : Google Scholar : PubMed/NCBI
|
|
83
|
Li Q, Zhang Y, Shi B, Lin C, Feng Q, Zhou
H, Hao M, Ding Y, Ma C, Mu J and Wang D: Galangin exosomes induce
cell apoptosis through miR-10b/P53 axis in gastric cancer. Sci Rep.
15:128762025. View Article : Google Scholar : PubMed/NCBI
|
|
84
|
Zhang Q, Deng T, Zhang H, Zuo D, Zhu Q,
Bai M, Liu R, Ning T, Zhang L, Yu Z, et al: Adipocyte-derived
exosomal MTTP suppresses ferroptosis and promotes chemoresistance
in colorectal cancer. Adv Sci (Weinh). 9:e22033572022. View Article : Google Scholar : PubMed/NCBI
|
|
85
|
Lv X, Li Z, Dai Y, Xiao Y, Shen F, Wang J,
Cao J, Wang L, Peng Q and Jiao Y: The mir-199b-5p encapsulated in
adipocyte-derived exosomes mediates radioresistance of colorectal
cancer cells by targeting JAG1. Heliyon. 10:e244122024. View Article : Google Scholar : PubMed/NCBI
|
|
86
|
Wang Y and Lin C: Exosomes miR-22-3p
derived from mesenchymal stem cells suppress colorectal cancer cell
proliferation and invasion by regulating RAP2B and PI3K/AKT
pathway. J Oncol. 2021:38744782021.PubMed/NCBI
|
|
87
|
Xu Y, Shen L, Li F, Yang J, Wan X and
Ouyang M: microRNA-16-5p-containing exosomes derived from bone
marrow-derived mesenchymal stem cells inhibit proliferation,
migration, and invasion, while promoting apoptosis of colorectal
cancer cells by downregulating ITGA2. J Cell Physiol.
234:21380–21394. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
88
|
Ning S, Chen Y, Li S, Liu M, Liu H, Ye M,
Wang C, Pan J, Wei W, Li J and Zhang L: Exosomal miR-99b-5p
secreted from mesenchymal stem cells can retard the progression of
colorectal cancer by targeting FGFR3. Stem Cell Rev Rep.
19:2901–2917. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
89
|
Li T, Wan Y, Su Z, Li J, Han M and Zhou C:
Mesenchymal stem cell-derived exosomal microRNA-3940-5p inhibits
colorectal cancer metastasis by targeting integrin α6. Dig Dis Sci.
66:1916–1927. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
90
|
Yu S, Liao R, Bai L, Guo M, Zhang Y, Zhang
Y, Yang Q, Song Y, Li Z, Meng Q, et al: Anticancer effect of
hUC-MSC-derived exosome-mediated delivery of PMO-miR-146b-5p in
colorectal cancer. Drug Deliv Transl Res. 14:1352–1369. 2024.
View Article : Google Scholar : PubMed/NCBI
|
|
91
|
Han Y, Zheng W, Zhang Y, Tong C, Song P,
Qi Z and Zhang S: Oxaliplatin-loaded natural killer cell-derived
exosomes for a safe and efficient chemoimmunotherapy of colorectal
cancer. J Pharm Sci. 114:1037832025. View Article : Google Scholar : PubMed/NCBI
|
|
92
|
Xu M, Chen Q, Li J, Peng L and Ding L:
Dendritic cell-derived exosome-entrapped fluorouracil can enhance
its anti-colon cancer effect. J BUON. 25:1413–1422. 2020.PubMed/NCBI
|
|
93
|
Jiang J, Wang W, Zhu L, Shi B, Chen Y, Xia
Y, Feng W, Yao W, Lu A and Zhang H: Unveiling the role of hypoxic
macrophage-derived exosomes in driving colorectal cancer
progression. Front Immunol. 14:12606382023. View Article : Google Scholar : PubMed/NCBI
|
|
94
|
Liu F, Ai F, Tang A, Yang Z, Li Z and Liu
S: Macrophage-derived exosomes promoted the development and
stemness of inflammatory bowel disease-related colorectal cancer
via nuclear paraspeckle assembly transcript 1-mediated
miRNA-34a-5p/phosphoprotein enriched in astrocytes 15 axis. Inflamm
Bowel Dis. 31:524–538. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
95
|
Zhao S, Mi Y, Guan B, Zheng B, Wei P, Gu
Y, Zhang Z, Cai S, Xu Y, Li X, et al: Tumor-derived exosomal
miR-934 induces macrophage M2 polarization to promote liver
metastasis of colorectal cancer. J Hematol Oncol. 13:1562020.
View Article : Google Scholar : PubMed/NCBI
|
|
96
|
Wu Y, Xiao Y, Ding Y, Ran R, Wei K, Tao S,
Mao H, Wang J, Pang S, Shi J, et al: Colorectal cancer cell-derived
exosomal miRNA-372-5p induces immune escape from colorectal cancer
via PTEN/AKT/NF-κB/PD-L1 pathway. Int Immunopharmacol.
143:1132612024. View Article : Google Scholar : PubMed/NCBI
|
|
97
|
Zhang C, Wang XY, Zhang P, He TC, Han JH,
Zhang R, Lin J, Fan J, Lu L, Zhu WW, et al: Cancer-derived exosomal
HSPC111 promotes colorectal cancer liver metastasis by
reprogramming lipid metabolism in cancer-associated fibroblasts.
Cell Death Dis. 13:572022. View Article : Google Scholar : PubMed/NCBI
|
|
98
|
Meng Q, Xiang H, Wang Y, Hu K, Luo X, Wang
J, Chen E, Zhang W, Chen J, Chen X, et al: Exosomes containing
circSCP2 in colorectal cancer promote metastasis via sponging
miR-92a-1-5p and interacting with PTBP1 to stabilize IGF2BP1. Biol
Direct. 19:1302024. View Article : Google Scholar : PubMed/NCBI
|
|
99
|
Li T, Li T, Liang Y, Yuan Y, Liu Y, Yao Y
and Lei X: Colorectal cancer cells-derived exosomal miR-188-3p
promotes liver metastasis by creating a pre-metastatic niche via
activation of hepatic stellate cells. J Transl Med. 23:3692025.
View Article : Google Scholar : PubMed/NCBI
|
|
100
|
Feng CZ, Zhong SQ, Ye SW, Zheng Z, Sun H
and Zhou SH: Tumor-derived exosomal miR-425-5p and miR-135b-3p
enhance colorectal cancer progression through immune suppression
and vascular permeability promotion. World J Gastrointest Oncol.
17:1061612025. View Article : Google Scholar : PubMed/NCBI
|
|
101
|
Li K, Xue W, Lu Z, Wang S, Zheng J, Lu K,
Li M, Zong Y, Xu F, Dai J, et al: Tumor-derived exosomal ADAM17
promotes pre-metastatic niche formation by enhancing vascular
permeability in colorectal cancer. J Exp Clin Cancer Res.
43:592024. View Article : Google Scholar : PubMed/NCBI
|
|
102
|
Liang Y, Li J, Yuan Y, Ju H, Liao H, Li M,
Liu Y, Yao Y, Yang L, Li T and Lei X: Exosomal miR-106a-5p from
highly metastatic colorectal cancer cells drives liver metastasis
by inducing macrophage M2 polarization in the tumor
microenvironment. J Exp Clin Cancer Res. 43:2812024. View Article : Google Scholar : PubMed/NCBI
|
|
103
|
Li S, Fu X, Ning D, Liu Q, Zhao J, Cheng
Q, Chen X and Jiang L: Colon cancer exosome-associated HSP90B1
initiates pre-metastatic niche formation in the liver by polarizing
M1 macrophage into M2 phenotype. Biol Direct. 20:522025. View Article : Google Scholar : PubMed/NCBI
|
|
104
|
Sun J, Luo J, Liu J, Wu H, Li Y, Xu Y, Liu
L, Liu X and Zhang Q: Cancer-secreted exosomal miR-1825 induces
angiogenesis to promote colorectal cancer metastasis. Cancer Cell
Int. 25:632025. View Article : Google Scholar : PubMed/NCBI
|
|
105
|
Yan YY, Deng ZF, Wu XT, Lu Y, Zhu ZY, Wen
Q, Zhang W, Zhang HY, Chen XZ, Wu YS, et al: Low miR-224-5p in
exosomes confers colorectal cancer 5-FU resistance by upregulating
S100A4. Drug Resist Updat. 79:1012112025. View Article : Google Scholar : PubMed/NCBI
|
|
106
|
Ren H, Wang M, Ma X, An L, Guo Y and Ma H:
METTL3 in cancer-associated fibroblasts-derived exosomes promotes
the proliferation and metastasis and suppresses ferroptosis in
colorectal cancer by eliciting ACSL3 m6A modification. Biol Direct.
19:682024. View Article : Google Scholar : PubMed/NCBI
|
|
107
|
Yang C, Zhang Y, Yan M, Wang J, Wang J,
Wang M, Xuan Y, Cheng H, Ma J, Chai C, et al: Exosomes derived from
cancer-associated fibroblasts promote tumorigenesis, metastasis and
chemoresistance of colorectal cancer by upregulating circ_0067557
to target Lin28. BMC Cancer. 24:642024. View Article : Google Scholar : PubMed/NCBI
|
|
108
|
Liu Y, Wang Y, Yu Z and Wang Z: Impacts of
TP53TG1 in cancer-associated fibroblasts-derived exosomes on
epithelial-mesenchymal transition capacity of colorectal carcinoma
cells by targeting miR-330-3p. Heliyon. 10:e303012024. View Article : Google Scholar : PubMed/NCBI
|
|
109
|
Xu H, Lan Q, Huang Y, Zhang Y, Zeng Y, Su
P and Chu Z, Lai W and Chu Z: The mechanisms of colorectal cancer
cell mesenchymal-epithelial transition induced by hepatocyte
exosome-derived miR-203a-3p. BMC Cancer. 21:7182021. View Article : Google Scholar : PubMed/NCBI
|
|
110
|
Yu Q, Zhang Y, Tian Y, Peng A, Cui X, Ding
B, Yang L, Liu Y, Ju Y and Gao C: Exosomal Circ_FMN2 derived from
the serum of colorectal cancer patients promotes cancer progression
by miR-338-3p/MSI1 Axis. Appl Biochem Biotechnol. 195:7322–7337.
2023. View Article : Google Scholar : PubMed/NCBI
|
|
111
|
Qiao XX, Shi HB and Xiao L: Serum exosomal
hsa-circ-0004771 modulates the resistance of colorectal cancer to
5-fluorouracil via regulating miR-653/ZEB2 signaling pathway.
Cancer Cell Int. 23:2432023. View Article : Google Scholar : PubMed/NCBI
|
|
112
|
Karbasi S, Erfanian N, Dehghan H, Zarban
A, Namaei MH, Hanafi-Bojd MY and Nasseri S: Assessment of the
anti-cancer effects of camel milk exosomes (CMEXOs) on murine
colorectal cancer cell line (CT-26). Iran J Allergy Asthma Immunol.
23:321–329. 2024.PubMed/NCBI
|
|
113
|
Babaker MA, Aljoud FA, Alkhilaiwi F,
Algarni A, Ahmed A, Khan MI, Saadeldin IM and Alzahrani FA: The
therapeutic potential of milk extracellular vesicles on colorectal
cancer. Int J Mol Sci. 23:68122022. View Article : Google Scholar : PubMed/NCBI
|
|
114
|
Martino E, Balestrieri A, Mele L, Sardu C,
Marfella R, D'Onofrio N, Campanile G and Balestrieri ML: Milk
exosomal miR-27b worsen endoplasmic reticulum stress mediated
colorectal cancer cell death. Nutrients. 14:50812022. View Article : Google Scholar : PubMed/NCBI
|
|
115
|
Lu Y, Zhou H, Han C, Gong Y, Li Y, Xia Y,
Liang B, Yang H and Wang Z: Enhanced therapeutic impact of
Shikonin-encapsulated exosomes in the inhibition of colorectal
cancer progression. Nanotechnology. 35:2024. View Article : Google Scholar
|
|
116
|
Liu Y, Cheng DH, Su ZY, Lv JH, Wang L,
Deng YY and Li L: Effects of total coumarins from Pileostegia
tomentella on exosomal miRNA expression and angiogenesis in
colorectal cancer cells. Pharm Biol. 62:153–161. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
117
|
Jin Y, Sun L, Chen Y and Lu Y: The
homologous tumor-derived-exosomes loaded with miR-1270 selectively
enhanced the suppression effect for colorectal cancer cells. Cancer
Med. 13:e69362024. View Article : Google Scholar : PubMed/NCBI
|
|
118
|
Liu M, Li TZ and Xu C: The role of
tumor-associated fibroblast-derived exosomes in chemotherapy
resistance of colorectal cancer and its application prospect.
Biochim Biophys Acta Gen Subj. 1869:1307962025. View Article : Google Scholar : PubMed/NCBI
|
|
119
|
Wang Z, Kim SY, Tu W, Kim J, Xu A, Yang
YM, Matsuda M, Reolizo L, Tsuchiya T, Billet S, et al:
Extracellular vesicles in fatty liver promote a metastatic tumor
microenvironment. Cell Metab. 35:1209–1226.e13. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
120
|
Sung H, Ferlay J, Siegel RL, Laversanne M,
Soerjomataram I, Jemal A and Bray F: Global cancer statistics 2020:
GLOBOCAN estimates of incidence and mortality worldwide for 36
cancers in 185 countries. CA Cancer J Clin. 71:209–249.
2021.PubMed/NCBI
|
|
121
|
Arnold M, Sierra MS, Laversanne M,
Soerjomataram I, Jemal A and Bray F: Global patterns and trends in
colorectal cancer incidence and mortality. Gut. 66:683–691. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
122
|
Liu SC and Zhang H: Early diagnostic
strategies for colorectal cancer. World J Gastroenterol.
30:3818–3822. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
123
|
Yin H, Xie J, Xing S, Lu X, Yu Y, Ren Y,
Tao J, He G, Zhang L, Yuan X, et al: Machine learning-based
analysis identifies and validates serum exosomal proteomic
signatures for the diagnosis of colorectal cancer. Cell Rep Med.
5:1016892024. View Article : Google Scholar : PubMed/NCBI
|
|
124
|
Hui J, Zhou M, An G, Zhang H, Lu Y, Wang X
and Zhao X: Regulatory role of exosomes in colorectal cancer
progression and potential as biomarkers. Cancer Biol Med.
20:575–598. 2023.PubMed/NCBI
|
|
125
|
Lin WC, Lin CC, Lin YY, Yang WH, Twu YC,
Teng HW and Hwang WL: Molecular actions of exosomes and their
theragnostics in colorectal cancer: Current findings and
limitations. Cell Oncol (Dordr). 45:1043–1052. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
126
|
Rizk NI, Kassem DH, Abulsoud AI,
AbdelHalim S, Yasser MB, Kamal MM and Hamdy NM: Revealing the role
of serum exosomal novel long non-coding RNA NAMPT-AS as a promising
diagnostic/prognostic biomarker in colorectal cancer patients. Life
Sci. 352:1228502024. View Article : Google Scholar : PubMed/NCBI
|
|
127
|
Zhao Y, Du T, Du L, Li P, Li J, Duan W,
Wang Y and Wang C: Long noncoding RNA LINC02418 regulates MELK
expression by acting as a ceRNA and may serve as a diagnostic
marker for colorectal cancer. Cell Death Dis. 10:5682019.
View Article : Google Scholar : PubMed/NCBI
|
|
128
|
Fabijanec M, Hulina-Tomašković A,
Štefanović M, Verbanac D, Ćelap I, Somborac-Bačura A, Grdić
Rajković M, Demirović A, Ramić S, Krušlin B, et al:
MicroRNA-193a-3p as a valuable biomarker for discriminating between
colorectal cancer and colorectal adenoma patients. Int J Mol Sci.
25:81562024. View Article : Google Scholar : PubMed/NCBI
|
|
129
|
Xu Y and Zhu M: Novel exosomal miR-46146
transfer oxaliplatin chemoresistance in colorectal cancer. Clin
Transl Oncol. 22:1105–1116. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
130
|
Zhao Y, Zhao Y, Liu L, Li G, Wu Y, Cui Y
and Xie L: Tumor-exosomal miR-205-5p as a diagnostic biomarker for
colorectal cancer. Clin Transl Oncol. 27:1185–1197. 2025.
View Article : Google Scholar : PubMed/NCBI
|
|
131
|
Yan S, Jiang Y, Liang C, Cheng M, Jin C,
Duan Q, Xu D, Yang L, Zhang X, Ren B and Jin P: Exosomal
miR-6803-5p as potential diagnostic and prognostic marker in
colorectal cancer. J Cell Biochem. 119:4113–4119. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
132
|
Liu X, Pan B, Sun L, Chen X, Zeng K, Hu X,
Xu T, Xu M and Wang S: Circulating exosomal miR-27a and miR-130a
act as novel diagnostic and prognostic biomarkers of colorectal
cancer. Cancer Epidemiol Biomarkers Prev. 27:746–754. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
133
|
Wang L, Song X, Yu M, Niu L, Zhao Y, Tang
Y, Zheng B, Song X and Xie L: Serum exosomal miR-377-3p and
miR-381-3p as diagnostic biomarkers in colorectal cancer. Future
Oncol. 18:793–805. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
134
|
Liu W, Yang D, Chen L, Liu Q, Wang W, Yang
Z, Shang A, Quan W and Li D: Plasma exosomal miRNA-139-3p is a
novel biomarker of colorectal cancer. J Cancer. 11:4899–4906. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
135
|
Zhao YJ, Song X, Niu L, Tang Y, Song X and
Xie L: Circulating exosomal miR-150-5p and miR-99b-5p as diagnostic
biomarkers for colorectal cancer. Front Oncol. 9:11292019.
View Article : Google Scholar : PubMed/NCBI
|
|
136
|
Wang J, Yan F, Zhao Q, Zhan F, Wang R,
Wang L, Zhang Y and Huang X: Circulating exosomal miR-125a-3p as a
novel biomarker for early-stage colon cancer. Sci Rep. 7:41502017.
View Article : Google Scholar : PubMed/NCBI
|
|
137
|
Li T, Zhou T, Wu J, Lv H, Zhou H, Du M,
Zhang X, Wu N, Gong S, Ren Z, et al: Plasma exosome-derived
circGAPVD1 as a potential diagnostic marker for colorectal cancer.
Transl Oncol. 31:1016522023. View Article : Google Scholar : PubMed/NCBI
|
|
138
|
Xie Y, Li J, Li P, Li N, Zhang Y, Binang
H, Zhao Y, Duan W, Chen Y, Wang Y, et al: RNA-Seq profiling of
serum exosomal circular RNAs reveals circ-PNN as a potential
biomarker for human colorectal cancer. Front Oncol. 10:9822020.
View Article : Google Scholar : PubMed/NCBI
|
|
139
|
Choi DS, Park JO, Jang SC, Yoon YJ, Jung
JW, Choi DY, Kim JW, Kang JS, Park J, Hwang D, et al: Proteomic
analysis of microvesicles derived from human colorectal cancer
ascites. Proteomics. 11:2745–2751. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
140
|
Erozenci LA, Böttger F, Bijnsdorp IV and
Jimenez CR: Urinary exosomal proteins as (pan-)cancer biomarkers:
Insights from the proteome. FEBS Lett. 593:1580–1597. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
141
|
Ma L, Yu H, Zhu Y, Xu K, Zhao A, Ding L,
Gao H and Zhang M: Isolation and proteomic profiling of urinary
exosomes from patients with colorectal cancer. Proteome Sci.
21:32023. View Article : Google Scholar : PubMed/NCBI
|
|
142
|
Yao J, Chen Y and Lin Z: Exosomes:
Mediators in microenvironment of colorectal cancer. Int J Cancer.
153:904–917. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
143
|
Huang C, Zhou Y, Feng X, Wang J, Li Y and
Yao X: Delivery of engineered primary tumor-derived exosomes
effectively suppressed the colorectal cancer chemoresistance and
liver metastasis. ACS Nano. 17:10313–10326. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
144
|
Shekh R, Ahmad A, Tiwari RK, Saeed M,
Shukla R, Al-Thubiani WS, Ansari IA, Ashfaque M and Bajpai P: High
therapeutic efficacy of 5-fluorouracil-loaded exosomes against
colon cancer cells. Chem Biol Drug Des. 101:962–976. 2023.
View Article : Google Scholar : PubMed/NCBI
|
|
145
|
Li Y, Gao Y, Gong C, Wang Z, Xia Q, Gu F,
Hu C, Zhang L, Guo H and Gao S: A33 antibody-functionalized
exosomes for targeted delivery of doxorubicin against colorectal
cancer. Nanomedicine. 14:1973–1985. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
146
|
Liu T, Zhang X, Du L, Wang Y, Liu X, Tian
H, Wang L, Li P, Zhao Y, Duan W, et al: Correction to:
Exosome-transmitted miR-128-3p increase chemosensitivity of
oxaliplatin-resistant colorectal cancer. Mol Cancer. 19:892020.
View Article : Google Scholar : PubMed/NCBI
|
|
147
|
Bagheri E, Abnous K, Farzad SA, Taghdisi
SM, Ramezani M and Alibolandi M: Targeted doxorubicin-loaded
mesenchymal stem cells-derived exosomes as a versatile platform for
fighting against colorectal cancer. Life Sci. 261:1183692020.
View Article : Google Scholar : PubMed/NCBI
|
|
148
|
Wu S, Yun J, Tang W, Familiari G,
Relucenti M, Wu J, Li X, Chen H and Chen R: Therapeutic m6A Eraser
ALKBH5 mRNA-loaded exosome-liposome hybrid nanoparticles inhibit
progression of colorectal cancer in preclinical tumor models. ACS
Nano. 17:11838–11854. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
149
|
Shakerian N, Darzi-Eslam E, Afsharnoori F,
Bana N, Noorabad Ghahroodi F, Tarin M, Mard-Soltani M, Khalesi B,
Hashemi ZS and Khalili S: Therapeutic and diagnostic applications
of exosomes in colorectal cancer. Med Oncol. 41:2032024. View Article : Google Scholar : PubMed/NCBI
|