Introduction
Lung cancer, a malignancy that poses a major threat
to human health, is generally classified into two main subtypes: i)
Small cell lung cancer; and ii) non-small cell lung cancer (NSCLC)
(1), with the latter accounting for
~85% of all cases (2). Although
notable progress has been achieved in the treatment of lung cancer,
through advances in surgical techniques, refinement of
chemoradiotherapy regimens, and the introduction of targeted and
immune-based therapies (3), overall
survival (OS) remains unsatisfactory, particularly among patients
with advanced disease (4).
Metastasis is a principal cause of therapeutic failure and
mortality in lung cancer (5).
Epithelial-mesenchymal transition (EMT) refers to a process in
which epithelial cells, under specific physiological or
pathological conditions, lose their epithelial characteristics and
acquire the phenotype and functional properties of mesenchymal
cells (6). During EMT, the
expression of epithelial markers, such as E-cadherin, is markedly
reduced, whereas mesenchymal markers, including N-cadherin and
Vimentin, are upregulated, thereby enhancing cellular migratory and
invasive capacities (7,8). Increasing evidence indicates that EMT
serves a pivotal role in the initiation, progression, invasion and
metastasis of lung cancer. Multiple signaling pathways have been
implicated in regulating EMT in lung cancer cells, including the
TGF-β pathway (9), Notch pathway
(10) and Wnt pathway (11,12).
EMT-related transcription factors, such as Snail (13), Twist (14) and zinc-finger E homeobox-binding
family members (15), are
frequently upregulated in lung cancer tissues (16). These factors directly repress
E-cadherin expression while inducing the expression of mesenchymal
markers such as N-cadherin and Vimentin, thereby driving EMT
(17). Despite advances in the
understanding of the relationship between lung cancer and EMT,
numerous mechanistic questions remain to be addressed.
The sodium channel β4 subunit (SCN4B) gene
encodes a crucial subunit of the voltage-gated sodium channel,
serving a key role in cellular electrophysiological functions
(18,19). Previous studies have primarily
concentrated on its involvement in the nervous system, particularly
in relation to neuromuscular disorders (20,21).
However, emerging evidence indicates that SCN4B may also
contribute to pathologies beyond the nervous system, notably in
cancer, a major threat to human health. For instance, alterations
in SCN4B expression have been implicated in modulating tumor
cell proliferation and metastasis in colorectal cancer (22). In lung cancer, SCN4B
expression levels have been reported to be closely associated with
tumor aggressiveness and patient prognosis, underscoring its
potential influence on disease progression (23). In summary, SCN4B was proposed
to act as a tumor suppressor by restraining the excessive
activation of tumor cell migration.
In the present study, the differential expression of
SCN4B in lung adenocarcinoma (LUAD) was analyzed and its
prognostic significance was evaluated. Based on these findings,
Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes
(KEGG) pathway enrichment analyses were performed on
SCN4B-associated differentially expressed genes, alongside
immune infiltration profiling. To further elucidate the role of
SCN4B in suppressing LUAD cell migration and invasion,
SCN4B-overexpressing cell lines were established. The effect
of SCN4B on tumor progression was assessed by examining
changes in cell viability, migration and invasion. Finally, the
effect of SCN4B on the EMT process was investigated through
the evaluation of EMT-related marker protein expression levels.
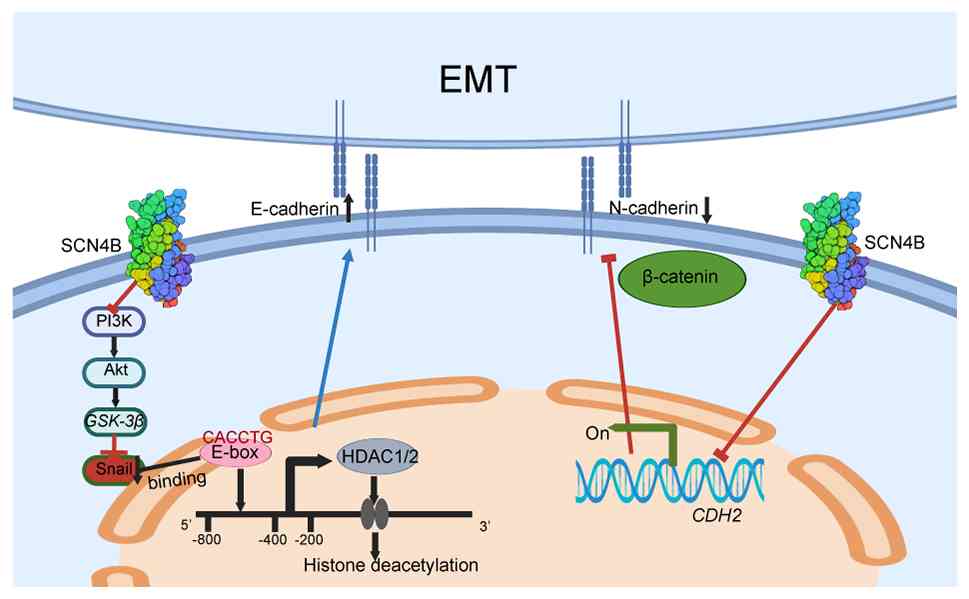
Based on previous reports (24,25),
we hypothesized that SCN4B may be involved in EMT regulation in
LUAD, and the working hypothesis is summarized in Fig. 1.
Materials and methods
Data acquisition and
preprocessing
Transcriptomic data for The Cancer Genome Atlas
(TCGA)-LUAD cohort were obtained from TCGA portal (https://portal.gdc.cancer.gov) using the Spliced
Transcripts Alignment to a Reference processing pipeline.
Expression profiles were extracted in the transcripts per million
format, and paired tumor-adjacent and tumor tissue samples were
identified based on matched patient identifiers. In total, 598
samples were included in the analysis, comprising 59 normal lung
tissues and 539 LUAD specimens.
Differential expression analysis of
SCN4B
Bioinformatics and statistical analyses were
performed in R (v4.5.2; R Core Team; http://www.R-project.org/) (26). The expression levels of SCN4B
in LUAD and adjacent normal tissues were compared using the ‘limma’
R package (v3.66.0; http://bioconductor.org/packages/release/bioc/html/limma.html).
The log2 fold change (log2FC) and adjusted
P-value (adj. P.Val) were calculated for all genes, and those with
|log2FC|>1 and adj. P.Val <0.05 were considered
significantly differentially expressed. Receiver operating
characteristic (ROC) curve analysis was performed using the ‘pROC’
package (v1.19.0.1; http://cran.r-project.org/package=pROC), and the
results were visualized with ‘ggplot2’ (v4.0.1; http://cran.r-project.org/package=ggplot2) (27).
Visualization of gene expression and
correlation analysis
Boxplots were generated using the ‘ggplot2’ package
to display SCN4B expression differences between tumor and
normal lung tissues. Volcano plots were generated using the
‘ggplot2’ package to visualize SCN4B-associated
differentially expressed genes between the SCN4B high- and
low-expression groups within the TCGA-LUAD tumor cohort.
Correlation heatmaps illustrating the relationships between
SCN4B and selected lung cancer-associated marker genes were
created to facilitate visualization of co-expression patterns
(28).
GO and KEGG functional enrichment
analysis
SCN4B-associated differentially expressed
genes identified between the SCN4B high- and low-expression
groups within the TCGA-LUAD tumor cohort (|log2FC|>1;
adj. P.Val <0.05) were selected for enrichment analysis. GO
annotation and KEGG pathway enrichment analysis were performed
using the R package ‘clusterProfiler’ (v4.4.4; http://bioconductor.org/packages/release/bioc/html/clusterProfiler.html)
package (29), running under R
(v4.5.2; R Core Team; http://www.R-project.org/). Bubble plots generated
with the R package ‘ggplot2’ were used to present the enrichment
results.
Prognostic evaluation
OS data for patients with LUAD were obtained from
the Gene Expression Profiling Interactive Analysis 2 platform
(v2.0; http://gepia2.cancer-pku.cn/#index), while
disease-specific survival (DSS) and progression-free interval (PFI)
information were sourced from TCGA. Kaplan-Meier survival analysis
was conducted, and statistical differences were assessed using the
log-rank test. Corresponding P-values were calculated and displayed
on the survival plots (30).
Expression validation and prognostic
analysis in the Gene Expression Omnibus (GEO; https://www.ncbi.nlm.nih.gov/geo/) dataset
Expression validation
The present study used the GSE31210 (31) (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE31210;
Affymetrix GPL570; Thermo Fisher Scientific, Inc.) series matrix
(robust multi-array average log2-transformed).
SCN4B was mapped to probe 236359_at. Samples were grouped as
tumor vs. normal (unpaired), and group differences were tested
using Welch's t-test; genome-wide analysis employed the linear
modeling framework implemented in the limma R/Bioconductor package
(version 4.4.4; http://bioconductor.org/packages/release/bioc/html/limma.html)
with Benjamini-Hochberg false discovery rate correction.
Prognostic analysis
In GSE31210 (GPL570), OS was used as the endpoint.
SCN4B expression was extracted using probe 236359_at and
patients were dichotomized at the cohort median to generate
Kaplan-Meier curves and the number at risk table. Groups were
compared using the two-sided log-rank test.
Clinical association analysis
Clinical information for patients with LUAD,
including age, sex, tumor stage and metastatic status, was
retrieved from TCGA. The association between SCN4B
expression and clinical characteristics (such as tumor stage and
metastasis) was evaluated using either the χ2 test or
Fisher's exact test (32) (Table I). Bar plots illustrating the
relationship between SCN4B expression and clinical features
were generated with the ‘ggplot2’ package (33).
 | Table I.Baseline clinical characteristics of
patients with lung adenocarcinoma with high and low SCN4B
expression. |
Table I.
Baseline clinical characteristics of
patients with lung adenocarcinoma with high and low SCN4B
expression.
|
Characteristics | Low SCN4B
expression, n (%) (n=269) | High SCN4B
expression, n (%) (n=270) | P-value |
|---|
| Pathologic T
stage |
|
| 0.003 |
| T1 | 69 (12.9) | 107 (20.0) |
|
| T2 | 166 (31.0) | 126 (23.5) |
|
| T3 | 24 (4.5) | 25 (4.7) |
|
| T4 | 9 (1.7) | 10 (1.9) |
|
| Pathologic N
stage |
|
| 0.658 |
| N0 | 172 (32.9) | 178 (34.0) |
|
| N1 | 54 (10.3) | 43 (8.2) |
|
| N2 | 40 (7.6) | 34 (6.5) |
|
| N3 | 1 (0.2) | 1 (0.2) |
|
| Pathologic M
stage |
|
| 0.734 |
| M0 | 188 (48.2) | 177 (45.4) |
|
| M1 | 12 (3.1) | 13 (3.3) |
|
| Pathologic
stage |
|
| 0.164 |
| Stage
I | 137 (25.8) | 159 (29.9) |
|
| Stage
II | 72 (13.6) | 53 (10.0) |
|
| Stage
III | 45 (8.5) | 39 (7.3) |
|
| Stage
IV | 12 (2.3) | 14 (2.6) |
|
| Sex |
|
| <0.001 |
|
Female | 124 (23.0) | 165 (30.6) |
|
|
Male | 145 (26.9) | 105 (19.5) |
|
| Age, years |
|
| 0.005 |
|
≤65 | 144 (27.7) | 113 (21.7) |
|
|
>65 | 115 (22.1) | 148 (28.5) |
|
| Smoker |
|
| 0.005 |
| No | 27 (5.1) | 50 (9.5) |
|
|
Yes | 235 (44.8) | 213 (40.6) |
|
| Anatomic neoplasm
subdivision |
|
| 0.628 |
|
Left | 100 (19.1) | 107 (20.4) |
|
|
Right | 160 (30.5) | 157 (30.0) |
|
Immune infiltration analysis
Correlations between SCN4B expression levels
and immune cell infiltration scores were assessed, and the results
were visualized using lollipop plots generated with the ‘ggplot2’
package.
Differential expression of SCN4B in
tissues
The immunohistochemical expression patterns of SCN4B
in adjacent non-tumor and LUAD tissues were obtained from the Human
Protein Atlas (HPA) database (https://www.proteinatlas.org/). The analysis was
conducted within the ‘Tissue’ section to explore potential
biological implications of SCN4B in LUAD.
Cell culture, passaging and
cryopreservation
The human LUAD cell lines A549, NCI-H1299,
NCI-H1975, PC-9 and HCC827, and the normal human bronchial
epithelial cell line BEAS-2B (all from Wuhan Servicebio Technology
Co., Ltd.), were maintained in a humidified incubator at 37°C with
5% CO2. A549, NCI-H1299 and BEAS-2B cells were cultured
in DMEM (Wuhan Servicebio Technology Co., Ltd.), whereas NCI-H1975,
PC-9 and HCC827 cells were cultured in RPMI-1640 medium (Wuhan
Servicebio Technology Co., Ltd.). All media were supplemented with
10% FBS (Wuhan Servicebio Technology Co., Ltd.) and 1%
penicillin-streptomycin (Wuhan Servicebio Technology Co., Ltd.;
final concentration 100 U/ml penicillin and 100 µg/ml streptomycin,
100X stock solution). The culture medium was refreshed every 48 h,
and cells were passaged at 80–90% confluence (typically every 2–3
days). Cells were cryopreserved at −80°C overnight and then
transferred to liquid nitrogen for long-term storage following
standard procedures.
Plasmid transfection and experimental
grouping
SCN4B overexpression plasmid DNA was
transiently transfected into A549 and NCI-H1299 cells using
Lipofectamine® 3000 (Invitrogen; Thermo Fisher
Scientific, Inc.). The SCN4B coding sequence was cloned into
the pcDNA3.1(+) backbone at the BamHI/EcoRI sites
(Sangon Biotech Co., Ltd.). Cells were seeded in 6-well plates
(0.3–1×105 cells/well) 1 day before transfection. On the
day of transfection, 0.5 µg plasmid DNA and 1.5–2.5 µl
Lipofectamine 3000 were prepared in serum-free DMEM (Wuhan
Servicebio Technology Co., Ltd.) to form transfection complexes
according to the manufacturer's instructions, which were then added
to the cells. Cells were incubated at 37°C with 5% CO2
for 18–24 h (without antibiotics). The transfection efficiency was
validated by reverse transcription-quantitative PCR (RT-qPCR) to
assess SCN4B mRNA expression. Cells were divided into two
groups: Vector (empty plasmid) and oe-SCN4B (SCN4B
overexpression). Subsequent experiments were initiated at 48 h
post-transfection, unless otherwise stated.
Cell Counting Kit-8 (CCK-8) viability
assay
Cells were seeded into 96-well plates at a density
of 5×103 cells/well and incubated at 37°C with 5%
CO2 to allow attachment. At 48 h post-transfection
(defined as 0 h for the CCK-8 assay), and at 24, 48 and 72 h
thereafter, 10 µl CCK-8 reagent (Wuhan Servicebio Technology Co.,
Ltd.) was added to each well containing 100 µl culture medium, and
plates were incubated for 1–4 h. Absorbance was measured at 450 nm
using a microplate reader to quantify the formazan product, which
reflects cell viability.
Transwell invasion assays
For the invasion assay, Transwell inserts were
precoated with Matrigel (Wuhan Servicebio Technology Co., Ltd.) at
37°C for 30 min prior to cell seeding. Cells were harvested and
resuspended in serum-free DMEM, and a total of 5×105
cells in 50 µl were seeded into the upper chamber of each Transwell
insert, with the lower chamber filled with DMEM supplemented with
20% FBS (Wuhan Servicebio Technology Co., Ltd.). Each group was
prepared in triplicate and cells were incubated at 37°C with 5%
CO2 for 48 h. Non-invaded cells on the upper surface
were gently removed with cotton swabs. The inserts were fixed with
4% paraformaldehyde (Beyotime Biotechnology) for 15 min at room
temperature (20–25°C), stained with crystal violet for 10 min at
room temperature and washed three times with PBS. Invaded cells in
five random fields (magnification, ×100) were imaged using an
inverted light microscope and counted using ImageJ software
(v1.52a; National Institutes of Health).
Wound healing assay
A549 and H1299 cells were seeded at 5×105
cells/well in 6-well plates. Upon reaching ~97% confluence, a
scratch was made using a 200-µl pipette tip guided by a ruler to
maintain uniform width. Detached cells were removed by PBS washing,
followed by incubation in low-serum medium (1% FBS). Images were
captured at 0 and 48 h (34) at the
same wound site using an inverted light microscope (magnification,
×100). The wound closure rate (%) was calculated as: Wound closure
rate (%)=[(area at 0 h-area at 48 h)/area at 0 h] ×100. The wound
area was quantified using ImageJ software (v1.52a; National
Institutes of Health). All assays were repeated three times.
RNA extraction and RT-qPCR
Total RNA was extracted from cells using
TRIzol® reagent (Beyotime Biotechnology) following the
manufacturer's instructions. Briefly, cells at 70–80% confluence
(~1×106 cells per sample) were lysed in 1 ml TRIzol for
RNA extraction. First-strand cDNA was synthesized from 1 µg total
RNA using a first-strand cDNA synthesis kit (Beyotime
Biotechnology) according to the manufacturer's protocol. qPCR was
performed on a Rotor-Gene 3000 real-time PCR system (Gene Company,
Ltd.) using SYBR® Green qPCR Master Mix (Beyotime
Biotechnology; the fluorophore was SYBR Green I dye provided in the
master mix). Each reaction was carried out in a 20 µl volume, and
amplification was performed with the following thermocycling
conditions: 95°C for 30 sec, followed by 40 cycles of 95°C for 5
sec and 60°C for 30 sec; melt-curve analysis was performed to
verify specificity. GAPDH served as the internal control, and
relative expression levels were calculated using the
2−ΔΔCq method (35). All
qPCR reactions were performed in triplicate (technical replicates).
The primer sequences for the target genes were as follows:
SCN4B forward, 5′-TCTTCCTGCTCCCCGTAAC-3′ and reverse,
5′-AATGCGTCACTGCTGTTGTAG-3′; and GAPDH forward,
5′-CTGGGCTACACTGAGCACC-3′ and reverse,
5′-AAGTGGTCGTTGAGGGCAATG-3′.
Western blotting
Total cellular proteins were extracted from the
collected cell pellets using pre-chilled RIPA lysis buffer (Beijing
Solarbio Science & Technology Co., Ltd.) supplemented with
protease and phosphatase inhibitors. Protein concentrations were
determined via a BCA assay. A total of 30 µg protein was loaded per
lane. Equal amounts of protein samples were separated on 10%
SDS-PAGE gels and subsequently transferred onto PVDF membranes.
After blocking with 5% non-fat milk at room temperature for 1 h,
the membranes were incubated overnight at 4°C with primary
antibodies against SCN4B (cat. no. DF4512; 1:1,000),
E-cadherin (cat. no. BF0219; 1:1,000), N-cadherin (cat. no. AF5239;
1:1,000), Vimentin (cat. no. BF8006; 1:1,000), Snail (cat. no.
AF6032; 1:1,000) and GAPDH (cat. no. AF7021; 1:1,000).
Subsequently, the membranes were incubated for 2 h at room
temperature with horseradish peroxidase-conjugated secondary
antibodies corresponding to the species of the primary antibodies:
Anti-rabbit IgG (cat. no. S0001; 1:10,000) or anti-mouse IgG (cat.
no. S0002; 1:10,000). All primary and secondary antibodies were
purchased from Affinity Biosciences. Protein bands were visualized
using a SuperECL Plus chemiluminescent detection kit (LI-COR
Biosciences) and imaged with the Bio-Rad ChemiDoc MP system
(Bio-Rad Laboratories, Inc.). Band intensities were semi-quantified
using ImageJ software (v1.5.2a; National Institutes of Health).
Flow cytometry
Cells were harvested, washed twice with cold PBS and
resuspended in 1X Annexin V binding buffer (10 mM HEPES, 140 mM
NaCl, 2.5 mM CaCl2, pH 7.4) at ~1×106
cells/ml. For each sample, 100 µl of cell suspension was incubated
with 5 µl annexin V-FITC and 5 µl PI (Annexin V-FITC Apoptosis
Detection Kit; cat. no. C1062; Beyotime Biotechnology) for 15 min
at room temperature in the dark. After adding 400 µl binding
buffer, samples were analyzed within 1 h on a CytoFLEX flow
cytometer (Beckman Coulter, Inc.) using the FITC and PI channels.
Data acquisition was performed with CytExpert software (v2.4;
Beckman Coulter, Inc.), and data were analyzed with FlowJo
(v10.10.0; BD Biosciences). Single-stained and unstained controls
were used for compensation. Events were gated to exclude debris and
doublets [forward scatter (FSC)/side scatter; FSC-area vs.
FSC-height]. No fixation was performed prior to acquisition. Cells
were classified as viable (annexin V−/PI−),
early apoptotic (annexin V+/PI−), late
apoptotic (annexin V+/PI+) or necrotic
(annexin V−/PI+). The apoptosis rate (%) was
calculated as early + late apoptosis.
Statistical analysis
Unless otherwise stated, all experiments were
performed in triplicate. Data are presented as the mean ± SD from
three independent experiments (n=3), unless otherwise indicated.
For non-normally distributed continuous variables, data are
presented as the median (interquartile range). Statistical analyses
were conducted using GraphPad Prism 8.0 (Dotmatics) and SPSS 26.0
(IBM Corp.). Categorical clinicopathological variables were
compared between high and low SCN4B expression groups using
χ2 tests or Fisher's exact tests. Survival was analyzed
using the Kaplan-Meier method, and differences were evaluated using
the log-rank test. For expression analyses based on transcriptomic
data, paired data were analyzed using a paired Student's t-test,
and unpaired comparisons with unequal variances were analyzed using
Welch's t-test. Comparisons among multiple groups were performed
using the Kruskal-Wallis test followed by Dunn's multiple
comparisons test when data were not normally distributed, whereas
two-group comparisons were performed using the Wilcoxon rank-sum
test under the same conditions. Spearman's rank correlation was
used to assess correlations between SCN4B expression and the
expression of selected genes. A two-tailed unpaired Student's
t-test was used to analyze the wound healing, Transwell and flow
cytometry apoptosis assay data, as well as RT-qPCR (ΔCq values) and
western blot densitometric semi-quantification data, comparing
between the vector and oe-SCN4B groups. Baseline SCN4B
expression across multiple lung adenocarcinoma cell lines was
compared using one-way ANOVA with Dunnett's post hoc test. The
CCK-8 assay data were analyzed using two-way ANOVA with
Bonferroni's post hoc test. Correlations between SCN4B
expression and immune cell infiltration scores were evaluated using
Spearman correlation analysis, and the corresponding correlation
coefficients and P-values are shown in the figures. ROC curves were
generated to assess the diagnostic performance of SCN4B.
P<0.05 was considered to indicate a statistically
significant difference.
Results
Differential expression of SCN4B in
LUAD
In TCGA-LUAD dataset, 539 tumor samples and 59
adjacent normal tissue samples were analyzed, and 58 matched
tumor-normal pairs were available for paired analysis (Fig. 2A). The baseline clinical
characteristics of patients with LUAD with high and low
SCN4B expression are summarized in Table I. Elevated SCN4B expression
was markedly associated with earlier T stage, sex, age and smoking
status, whereas no significant differences were observed in N
stage, M stage, overall pathological stage or anatomic neoplasm
subdivision.
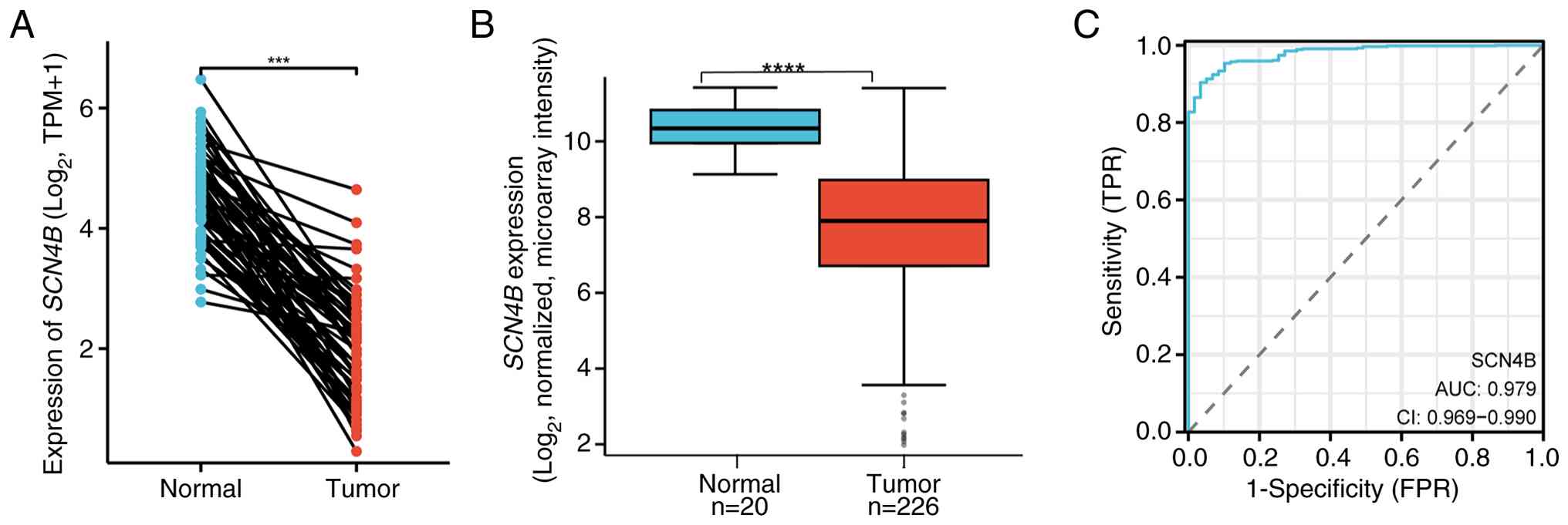
Expression profiling revealed that SCN4B was
markedly downregulated in LUAD tissues compared with adjacent
normal tissues (Fig. 2A). The
external GEO cohort (GSE31210) exhibited significant downregulation
of SCN4B in tumor vs. normal lung tissues (Fig. 2B). ROC curve analysis (Fig. 2C) indicated that SCN4B
possessed strong diagnostic potential, with an area under the curve
of 0.979 (95% CI, 0.969–0.990).
Differential analysis of SCN4B and
related genes
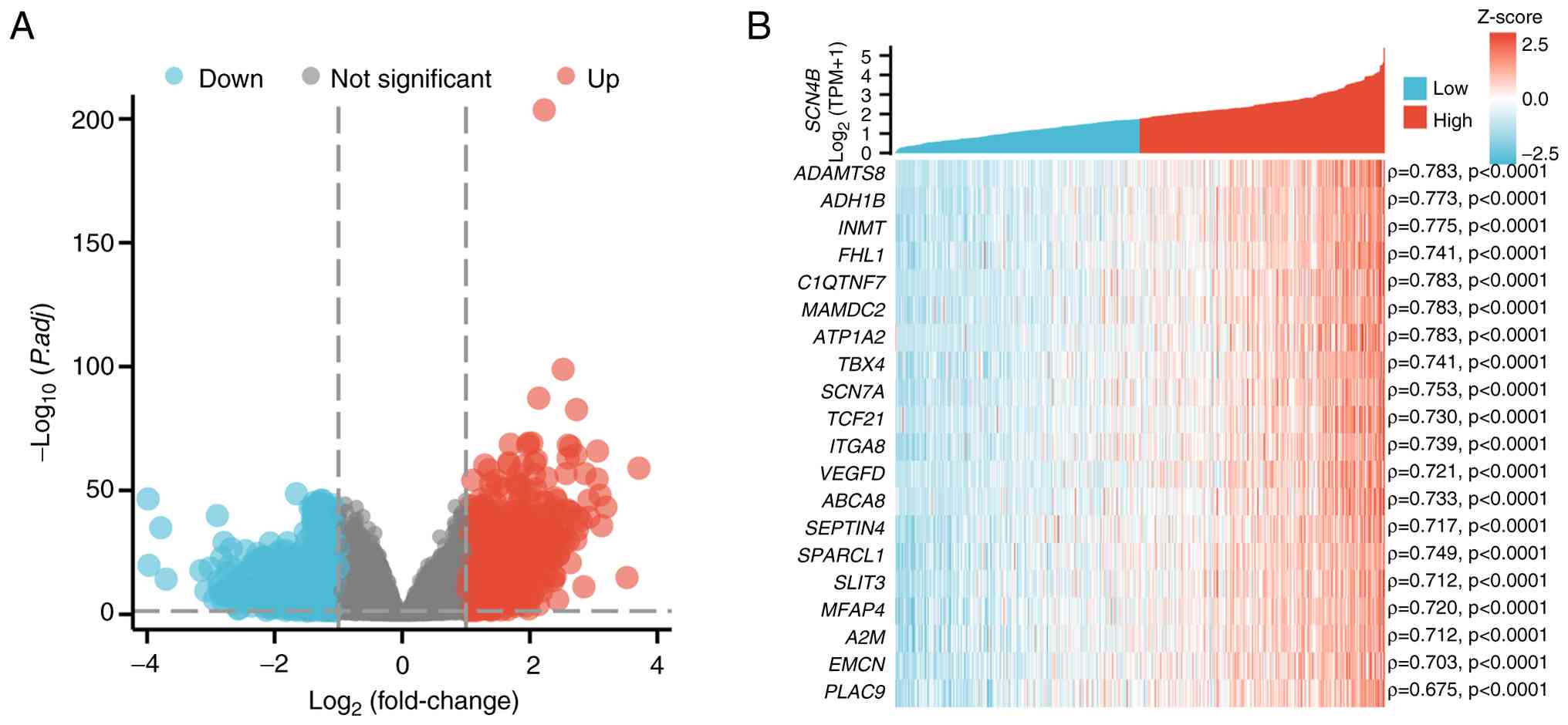
The volcano plot of SCN4B-associated
differentially expressed genes between the SCN4B high- and
low-expression groups is shown in Fig.
3A. Spearman's rank correlation analysis was performed to
assess the correlation between SCN4B and several genes,
including ADAMTS8, ADH1B, INMT, FHL1, C1QTNF7, MAMDC2, ATP1A2,
TBX4, SCN7A, TCF21 and ITGA8 (Fig. 3B). These associations suggest that
SCN4B and the identified genes may participate in common
biological processes (BPs) or signaling pathways, potentially
exerting cooperative or reciprocal regulatory effects.
GO and KEGG functional enrichment
analysis
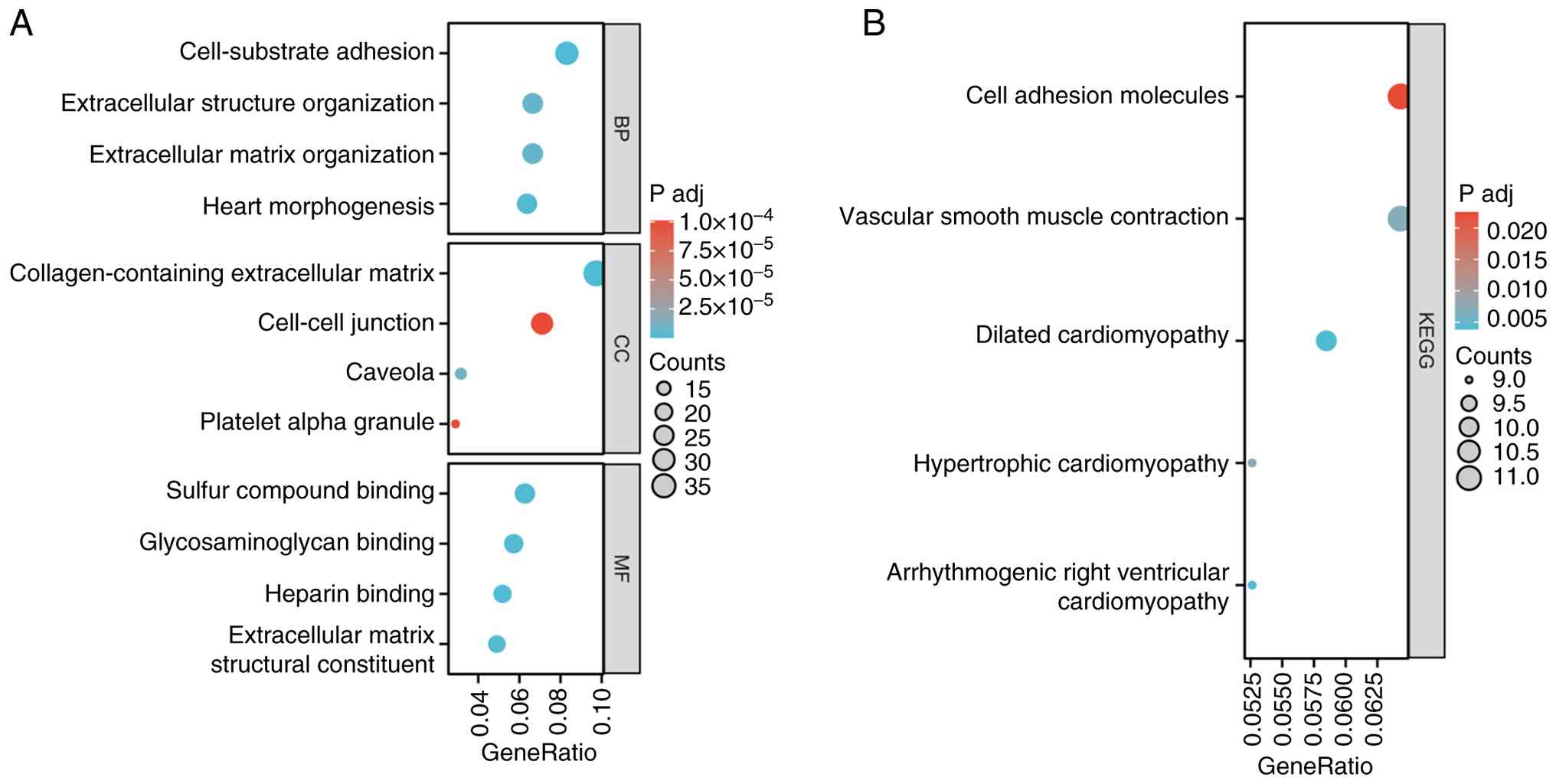
GO functional enrichment analysis indicated that the
SCN4B-associated differentially expressed genes were
significantly enriched in BP terms including ‘cell-substrate
adhesion’, ‘extracellular structure organization’, ‘extracellular
matrix organization’ and ‘heart morphogenesis’. At the cellular
component level, the SCN4B-associated differentially
expressed genes were mainly associated with ‘collagen-containing
extracellular matrix’, ‘cell-cell junction’, ‘caveola’ and
‘platelet alpha granule’. In terms of molecular functions (MFs),
the SCN4B-associated differentially expressed genes were
primarily enriched in ‘sulfur compound binding’, ‘glycosaminoglycan
binding’, ‘heparin binding’ and ‘extracellular matrix structural
constituent’ (Fig. 4A). In
addition, KEGG pathway enrichment analysis showed that the
SCN4B-associated differentially expressed genes were
enriched in pathways such as ‘cell adhesion molecules’ (CAMs),
‘vascular smooth muscle contraction’, ‘dilated cardiomyopathy’,
‘hypertrophic cardiomyopathy’ and ‘arrhythmogenic right ventricular
cardiomyopathy’ (Fig. 4B).
Immune infiltration analysis of
SCN4B
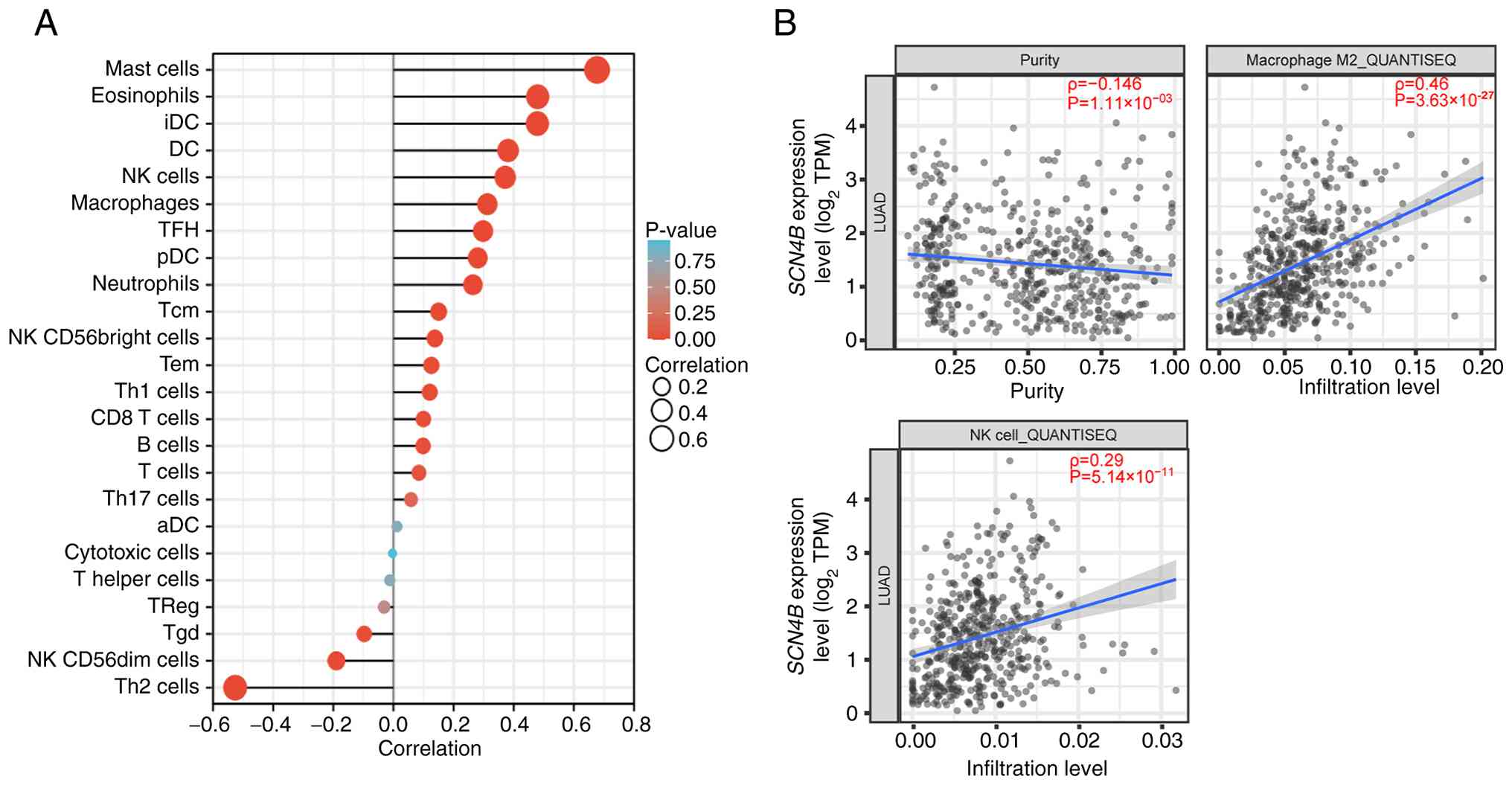
SCN4B expression was positively associated
with mast cells, eosinophils and immature dendritic cells (iDCs).
By contrast, SCN4B expression showed a negative correlation
with type 2 helper T cells (Th2 cells), natural killer (NK)
CD56dim cells, γδ T cells (Tgds) and regulatory T cells
(Tregs) (Fig. 5A).
As shown in Fig. 5B,
SCN4B expression was negatively associated with tumor
purity, and weak positive correlations were observed with M2
macrophage and NK cell infiltration levels estimated by
quantification of the tumor immune contexture from RNA
sequencing.
SCN4B expression in LUAD and adjacent
normal tissues
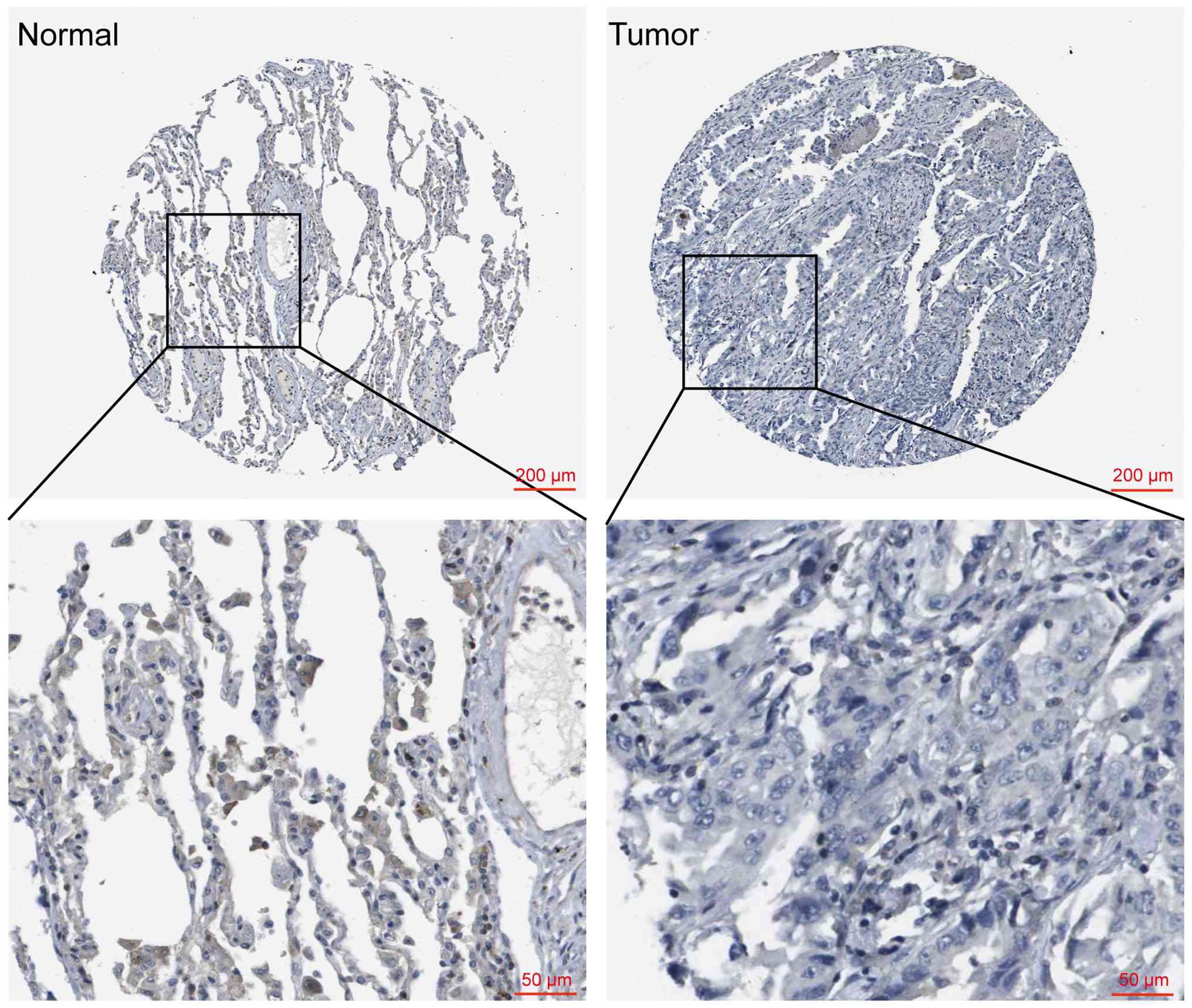
Analysis using the HPA database revealed that
SCN4B expression was nearly absent in tumor tissues, whereas
detectable expression was observed in adjacent non-tumor tissues
(Fig. 6).
Prognostic and clinicopathological
association analysis of SCN4B
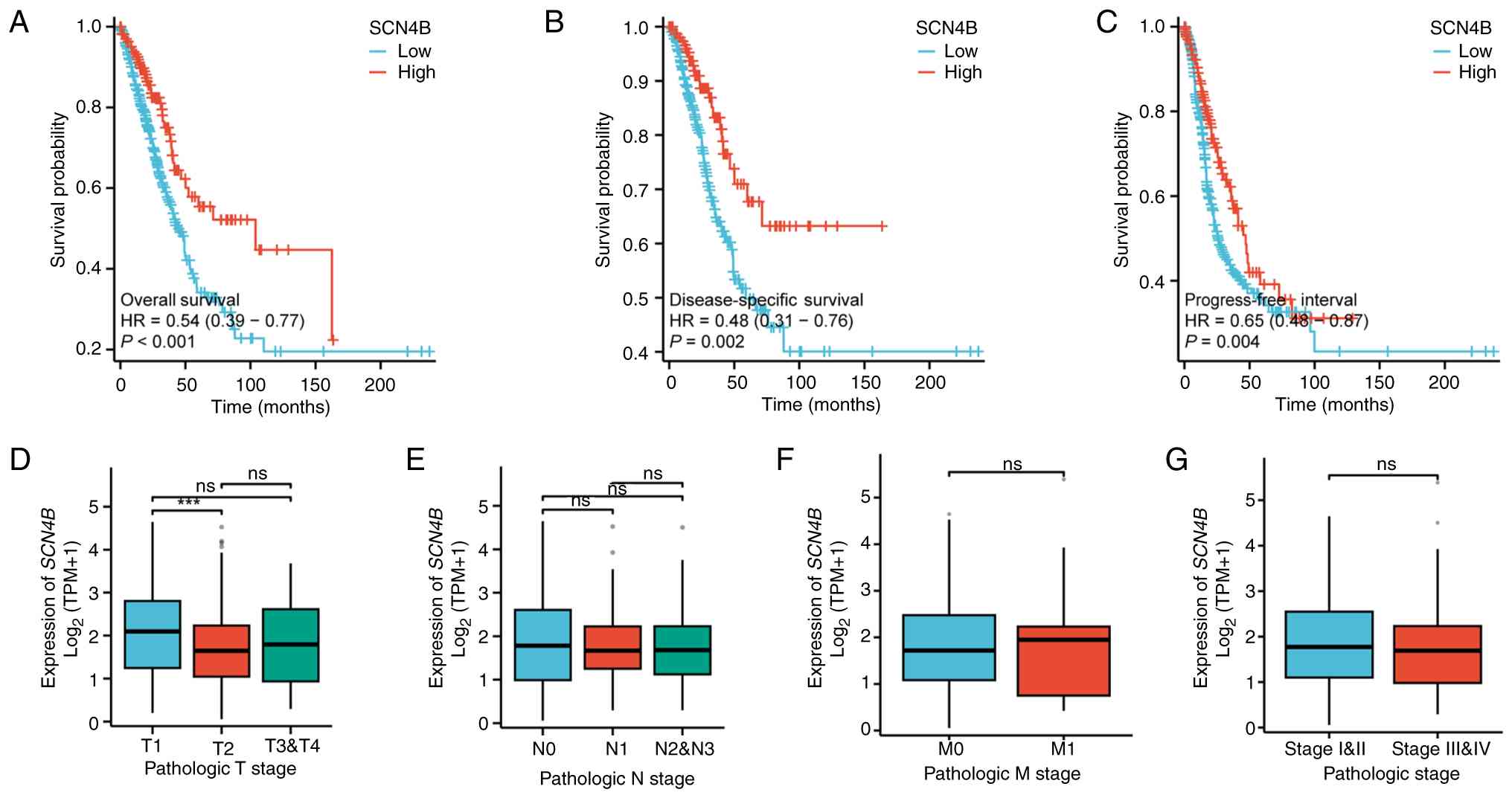
Patients with high SCN4B expression exhibited
longer OS and DSS than those with low expression (Fig. 7A and B). High SCN4B
expression was associated with a longer PFI (Fig. 7C).
In the analysis of SCN4B expression across
TNM stages, a significant difference was observed between T1 and T2
stages, with markedly reduced expression in T2 tumors. SCN4B
expression differed between stages T1 and T2. However, no
statistically significant differences were observed in stage T3-T4
compared with stages T1 or T2 (Fig.
7D). Furthermore, SCN4B expression was not markedly
associated with lymph node metastasis, distant metastasis or
overall tumor stage (Fig.
7E-G).
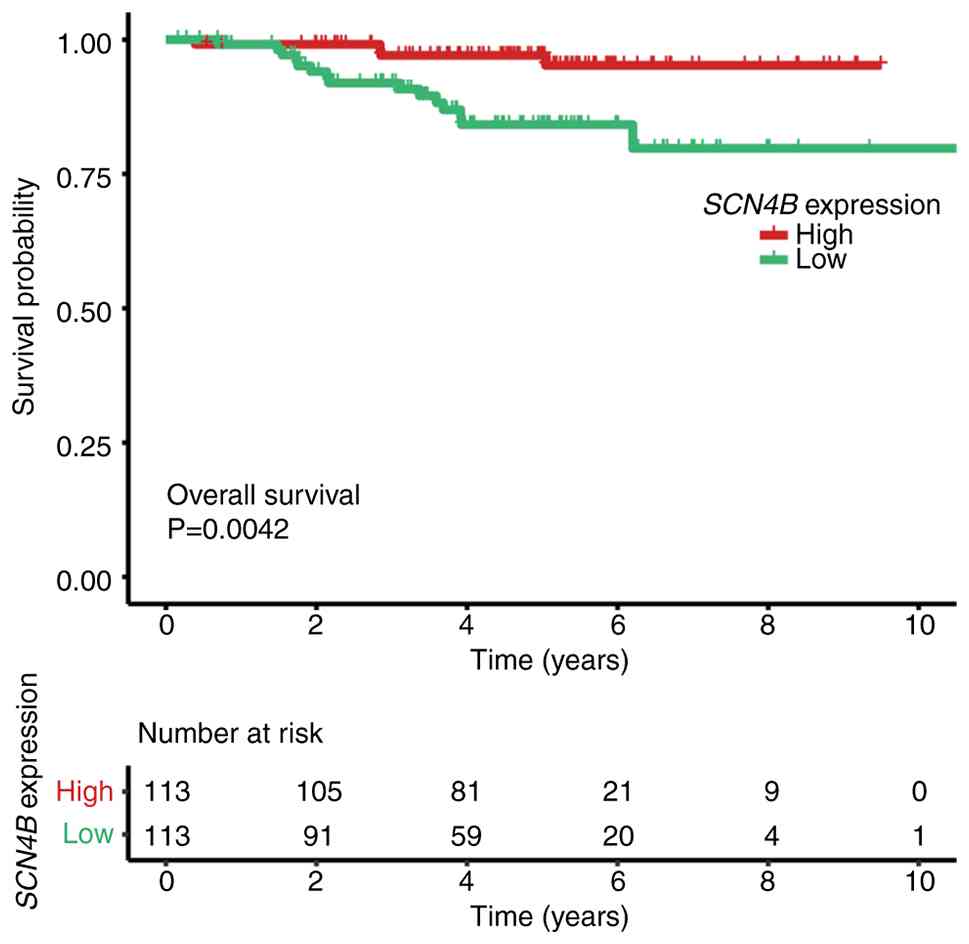
External prognostic validation
In the external LUAD cohort GSE31210, high
SCN4B expression was associated with improved survival.
After median dichotomization, Kaplan-Meier curves and the ‘number
at risk’ table displayed below the plot showed a significant
difference between groups (Fig.
8).
Baseline SCN4B profiling and model
selection
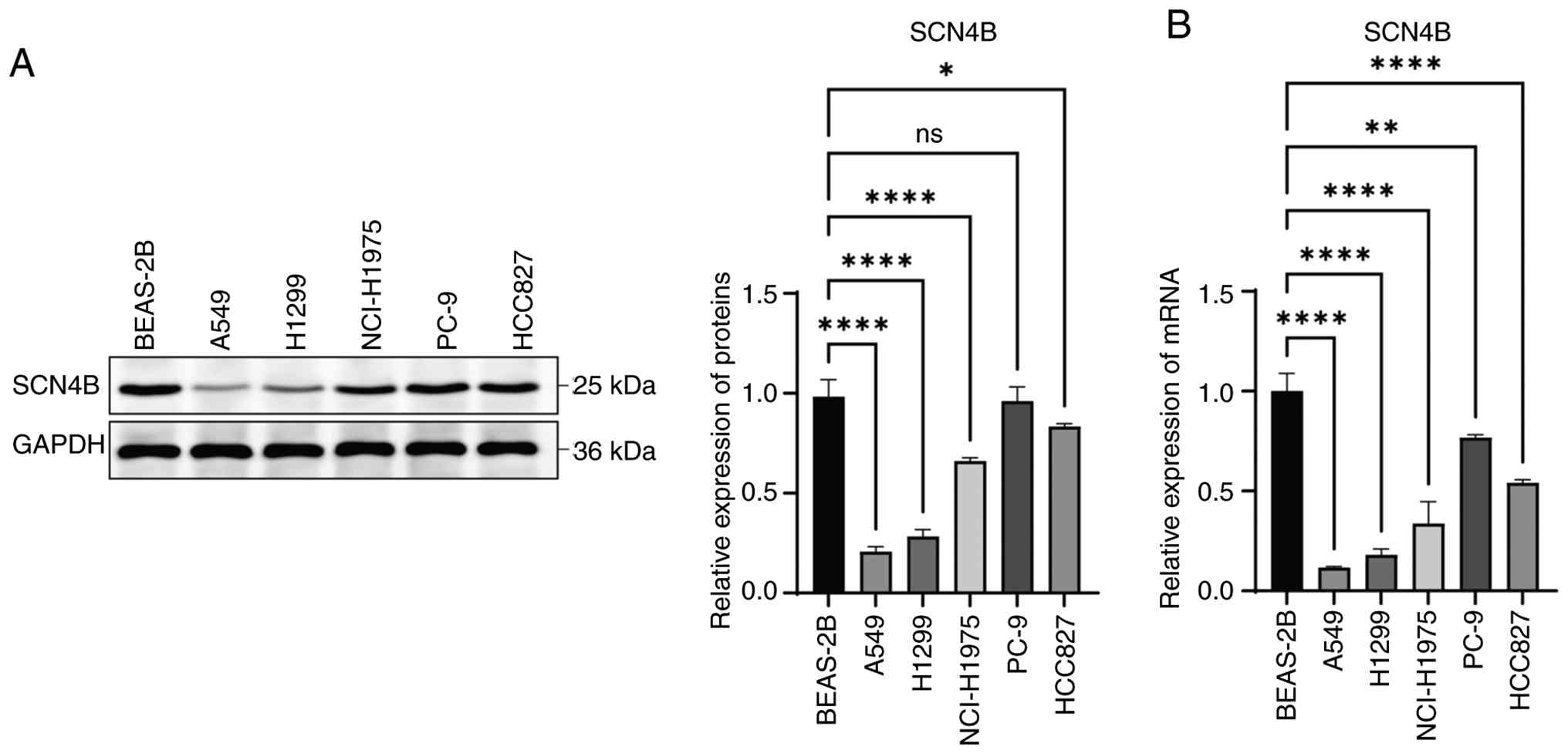
First, SCN4B protein and mRNA levels were screened
by western blotting and RT-qPCR in a normal lung-derived cell line
and multiple lung cancer cell lines. Using the normal cell line as
the reference and normalizing to GAPDH, most cancer cell lines
exhibited a marked reduction in SCN4B signal. The decrease
was most consistent and pronounced in A549 and H1299 cells, which
were therefore selected for subsequent functional assays (Fig. 9).
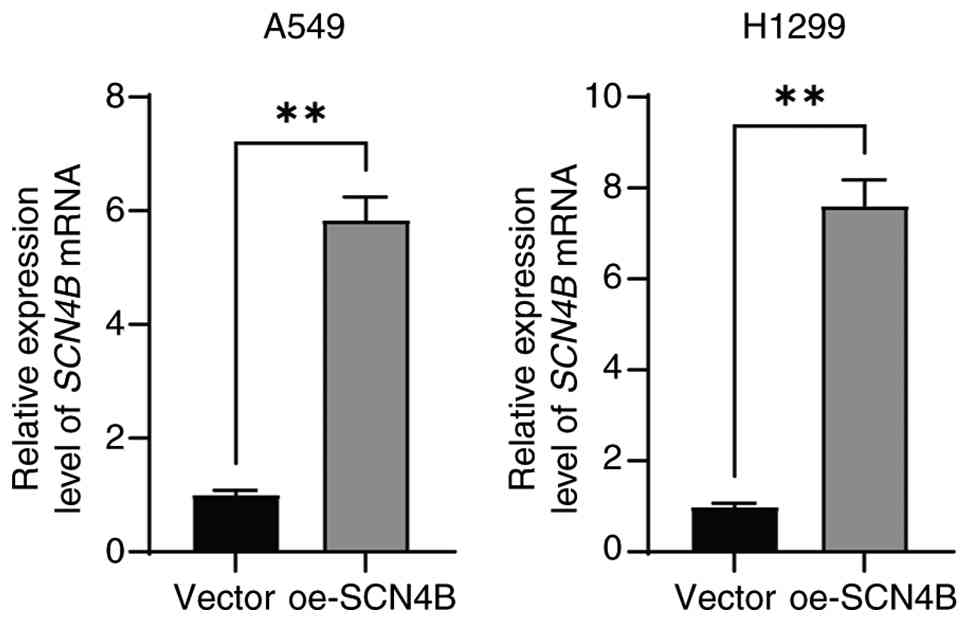
Construction of SCN4B overexpression
cell models
SCN4B overexpression cell lines were
established in A549 and H1299 cells by transfection with an
overexpression plasmid. The results showed that the mRNA expression
levels of SCN4B were markedly increased in the
oe-SCN4B group compared with the vector group (Fig. 10).
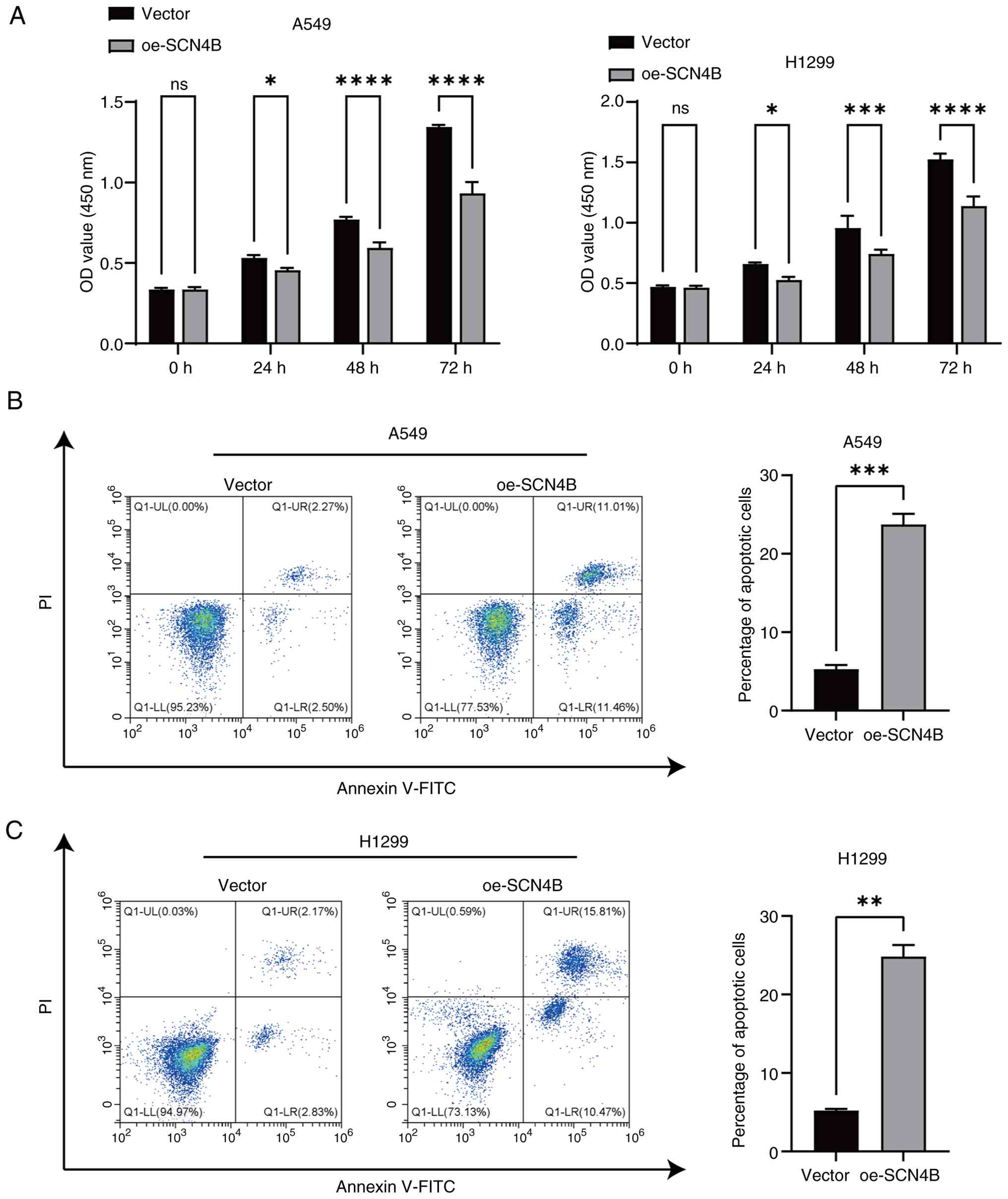
SCN4B reduces lung cancer cell
viability and induces apoptosis in LUAD cells
The CCK-8 assay results demonstrated that
SCN4B overexpression markedly reduced the viability of A549
and H1299 cells in a time-dependent manner, with prolonged exposure
leading to stronger inhibitory effects (Fig. 11A).
Flow cytometry analysis using annexin V-FITC/PI
double staining revealed that SCN4B overexpression markedly
increased the proportion of apoptotic cells (early apoptosis + late
apoptosis) in A549 and H1299 LUAD cells (Fig. 11B and C).
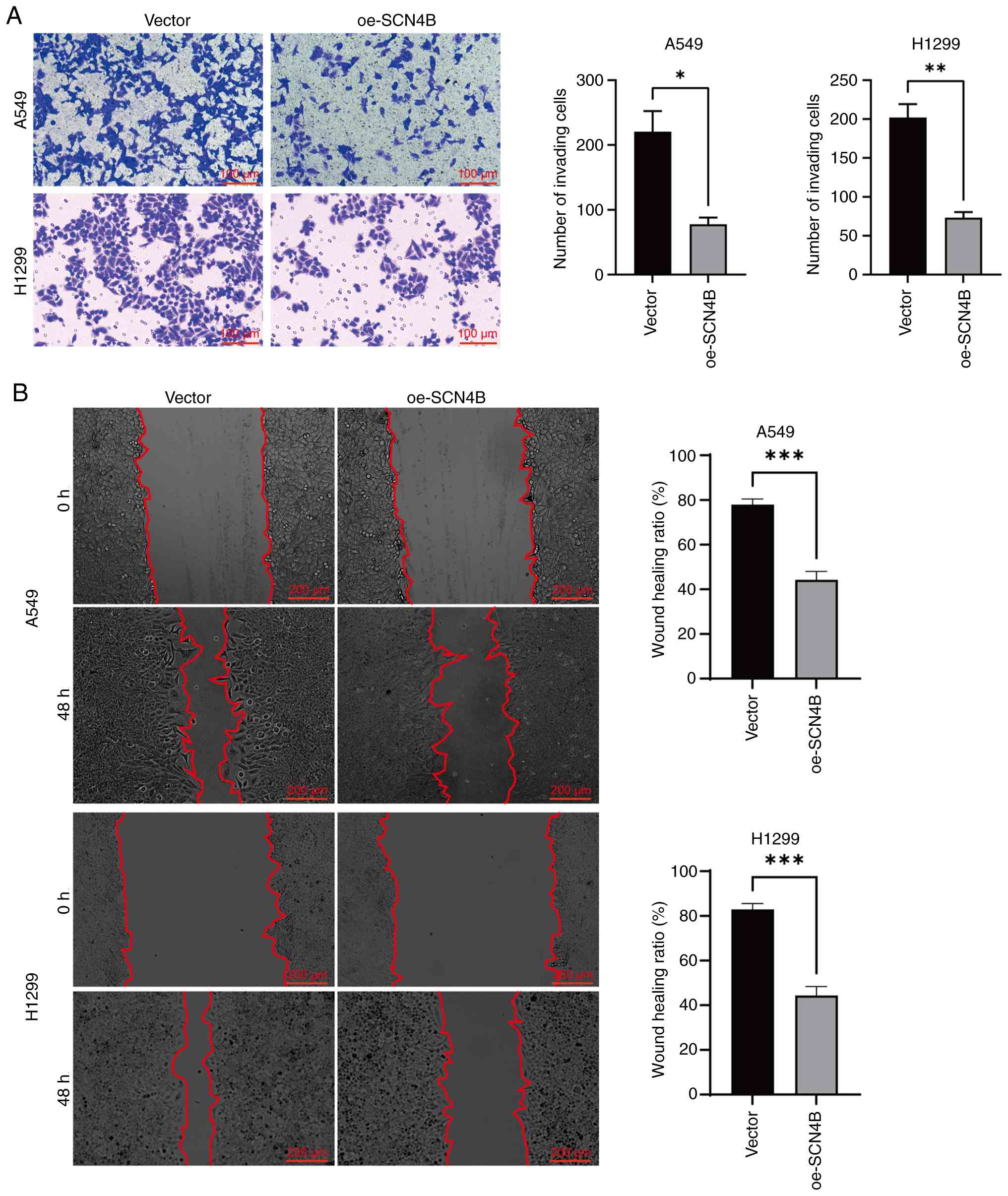
SCN4B suppresses invasion and
migration of LUAD cells
Transwell assay results showed that the number of
invasive cells was significantly decreased following SCN4B
overexpression (Fig. 12A). In
addition, wound healing assay results indicated that SCN4B
overexpression markedly slowed the wound closure rate and markedly
reduced the healing area compared with the vector group (Fig. 12B).
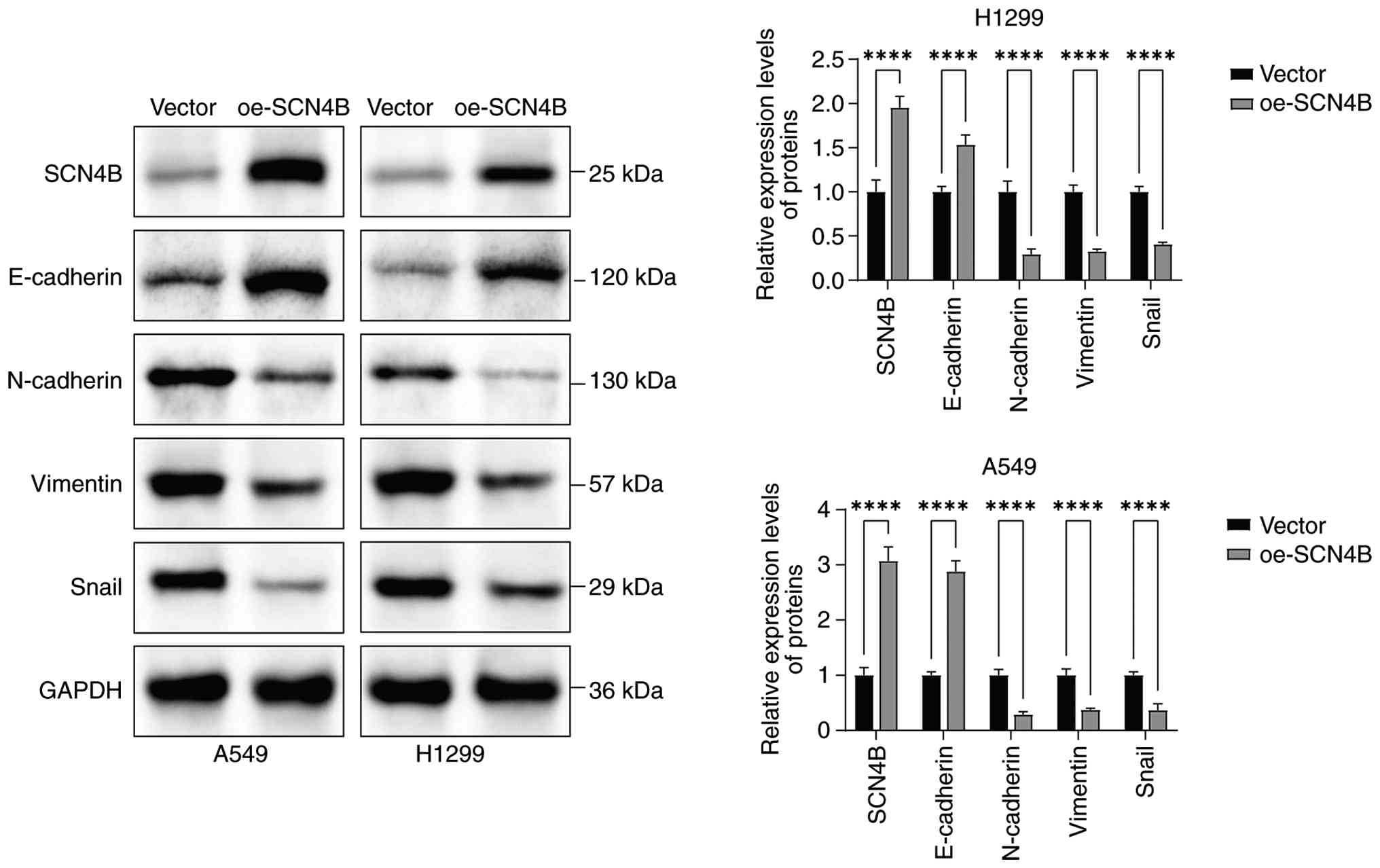
Effect of SCN4B on EMT-related marker
proteins
Western blot analysis revealed that SCN4B
overexpression markedly increased the relative expression levels of
E-cadherin, while suppressing the relative expression levels of
N-cadherin, Vimentin and Snail (Fig.
13).
Discussion
LUAD is one of the leading causes of tumor-related
mortality worldwide, with its high lethality primarily attributed
to late-stage diagnosis and highly aggressive biological behavior
(36). Among the various mechanisms
driving LUAD progression, EMT has been shown to be a central
process underlying metastasis and recurrence (37). Therefore, an improved understanding
of the molecular regulation of EMT in LUAD is of considerable
clinical value.
The present bioinformatics analysis revealed marked
downregulation of SCN4B expression in LUAD tissues, with low
expression markedly associated with unfavorable prognosis. This
observation aligns with a previous large-scale analysis reporting a
prognostic link between reduced SCN4B levels and poor lung
cancer outcomes (38).
Kaplan-Meier survival analysis demonstrated that
patients with low SCN4B expression had a substantially
shorter median survival time, while ROC curve analysis indicated
that SCN4B possessed promising diagnostic value in LUAD.
Furthermore, analysis by pathological T stage showed that
SCN4B expression was significantly lower in stage T2 than in
stage T1, whereas no statistically significant differences were
observed between stage T3-T4 and either stage T1 or stage T2. This
trend may indicate that changes in SCN4B expression are closely
related to local tumor expansion or primary tumor burden. However,
SCN4B expression did not significantly differ between groups
based on other staging parameters (such as N stage, M stage or
pathological stage), which may be due to the limited sample size or
may indicate that SCN4B primarily participates in
early-stage local invasion and proliferation (39). To the best of our knowledge, to
date, the biological basis for this stage-specific pattern remains
unexplored.
Dysregulated cell proliferation and inhibition of
apoptosis are considered key molecular events driving tumor
initiation and progression (40,41).
Under normal conditions, cell growth is tightly regulated by
extracellular cues. However, cancer cells frequently acquire
autocrine growth signaling, bypass cell-cycle checkpoints and
maintain uncontrolled proliferation (42,43).
In the present study, SCN4B overexpression in LUAD cell
lines (A549 and H1299) markedly reduced the proliferative capacity
in a time-dependent manner, as determined by CCK-8 assays. The
results showed that SCN4B overexpression markedly decreased
cell viability in both cell lines, with a time-dependent inhibitory
effect. Furthermore, considering that apoptosis evasion is an
important mechanism for tumor survival and immune escape (44), annexin V/PI dual-staining flow
cytometry was performed to evaluate apoptosis. The results
demonstrated that SCN4B overexpression markedly induced both
early and late apoptosis in LUAD cells. These results suggested
that SCN4B exerts a tumor-suppressive effect through
simultaneous inhibition of proliferation and promotion of
apoptosis.
In LUAD, EMT not only drives metastasis but also
contributes to chemoresistance and stemness acquisition (45). The Snail family of transcription
factors, including Snail1 and Snail2, are key EMT regulators
(46). By binding to the E-box
elements within the E-cadherin promoter region, they suppress
E-cadherin expression, thereby promoting EMT (47). N-cadherin, a mesenchymal cell
marker, is upregulated during EMT and is associated with enhanced
invasive and migratory abilities (48,49).
Vimentin, an intermediate filament protein, is another mesenchymal
marker whose increased expression during EMT is associated with
enhanced motility and invasiveness (50).
The functional assays in the present study
demonstrated that SCN4B overexpression markedly impaired
LUAD cell migration and invasion. Western blot analysis showed that
SCN4B elevated E-cadherin expression, while reducing N-cadherin,
Vimentin and Snail levels, suggesting a suppressive effect on EMT,
potentially via the Snail/E-cadherin axis. Previous work has shown
that Snail recruits histone deacetylases to the E-cadherin
promoter, modulating H3/H4 acetylation and silencing transcription,
providing a plausible mechanism for SCN4B-mediated EMT
inhibition (41). Taken together,
the present results and previous reports (24,25)
supported the proposed schematic model whereby SCN4B may regulate
EMT in LUAD (Fig. 1).
In the LUAD cell models used in the present study,
SCN4B overexpression attenuated EMT and invasive behavior,
suggesting that its downstream effects may involve several key
signaling pathways. Among the classical EMT-inducing cascades, the
TGF-β/Smad axis is a major driver of epithelial plasticity,
invasive growth and treatment resistance in NSCLC. TGF-β receptor
signaling promotes the disassembly of tight junctions,
reorganization of the actin cytoskeleton and induction of
EMT-related transcription factors such as Snail (51). Previous studies have indicated that
SCN4B participates in the regulation of cell adhesion and
cytoskeletal dynamics (52,53). In the present study, SCN4B
overexpression increased E-cadherin expression, while reducing
N-cadherin, Vimentin and Snail expression. These findings support
the hypothesis that SCN4B may dampen TGF-β-induced EMT in
LUAD cells. Biologically, such an effect would be expected to
reduce EMT plasticity and local invasiveness of tumor cells and to
alleviate the development of an immunosuppressive,
therapy-resistant tumor microenvironment driven by chronic TGF-β
signaling, thereby collectively constraining tumor progression.
The Wnt/β-catenin pathway represents another
critical signaling cascade that promotes EMT, maintenance of
stem-like properties and metastatic spread in lung cancer. Under
physiological conditions, β-catenin is sequestered at adherens
junctions through binding to E-cadherin. When E-cadherin is lost,
β-catenin accumulates in the nucleus and activates Wnt target genes
(54). In TCGA-LUAD, enrichment
analysis of SCN4B-associated differentially expressed genes
highlighted adhesion-related processes, including the KEGG ‘cell
adhesion molecules’ pathway, and in the in vitro
experiments, SCN4B overexpression increased E-cadherin expression,
suggesting that SCN4B may indirectly limit sustained
activation of canonical Wnt/β-catenin signaling by stabilizing the
membrane E-cadherin/β-catenin complex. Integrating these
observations, a unified and testable working model in which
SCN4B functions as a membrane-associated ‘brake’ was
proposed. On the one hand, it attenuates TGF-β/Smad signaling by
restricting TGF-β1-induced Smad2/3 phosphorylation and the
upregulation of EMT transcription factors (52). On the other hand, it suppresses
canonical Wnt/β-catenin signaling by stabilizing epithelial
cell-cell junctions, reducing nuclear β-catenin accumulation and
lowering T-cell factor/lymphoid enhancer factor reporter activity
(55). This integrated model can be
evaluated by combining TGF-β1 stimulation and pharmacologic
modulation of the Wnt pathway in SCN4B-manipulated LUAD
cells, together with TCF/LEF-dependent luciferase reporter assays
using the TOPflash reporter (containing wild-type TCF/LEF-binding
sites) and the FOPflash (containing mutated TCF/LEF-binding sites)
mutant control, and analyses of Smad2/3, EMT markers and β-catenin
expression and subcellular localization (56). Given that both TGF-β and
Wnt/β-catenin pathways are tightly linked to EMT plasticity,
stem-like tumor cell subpopulations, metastatic colonization and
resistance to systemic therapy, the regulation of these pathways by
SCN4B may have important implications for the biological
aggressiveness and clinical behavior of LUAD (56,57).
GO and KEGG enrichment analyses of
SCN4B-associated differentially expressed genes between
SCN4B-high and SCN4B-low LUAD samples were performed
to investigate the potential role of SCN4B in
post-transcriptional regulation and in modulating cellular
structure and function during the EMT process. In TCGA-LUAD
dataset, GO enrichment analysis revealed that
SCN4B-associated differentially expressed genes were
markedly enriched in BP terms including ‘cell-substrate adhesion’,
‘extracellular structure organization’ and ‘extracellular matrix
organization’. KEGG pathway analysis further demonstrated that
these genes were markedly enriched in the ‘cell adhesion molecules’
pathway. The dynamic balance between microtubule polymerization and
depolymerization is a crucial mechanism for maintaining
cytoskeletal stability and regulating the migratory capacity of
cancer cells (58). The enrichment
of SCN4B-associated differentially expressed genes in the
‘cell adhesion molecules’ pathway, involving key adhesion molecules
such as E-cadherin and integrins, suggests that SCN4B may
interfere with the EMT process by modulating intercellular junction
structures (such as adherens junctions and desmosomes), thereby
inhibiting the invasion and metastasis of LUAD cells (59,60).
In addition, marked enrichment of
‘glycosaminoglycan binding’ in the MF category was observed in the
present study. Glycosaminoglycans, covalently linked to proteins to
form proteoglycans, are ubiquitous across mammalian cells, and are
present on membranes, in the intracellular space and within the
extracellular matrix (ECM). By interacting with a wide range of
ligands, they participate in multiple physiological and
pathological events, including cancer (61). In the present study, the MF term
‘glycosaminoglycan binding’ was markedly enriched, suggesting that
SCN4B may influence the remodeling of the tumor
microenvironment and regulate pro-tumorigenic signaling pathways by
modulating ECM components.
Immune profiling revealed a positive association
between SCN4B expression and mast cells, eosinophils and
iDCs, but an inverse correlation with Th2 cells, NK
CD56dim cells and Tgds. Such patterns suggest that
SCN4B may act as an immune microenvironment regulator by
shaping immune cell infiltration profiles. Given that the tumor
microenvironment is integral to cancer initiation, progression,
metastasis and therapeutic response, the infiltration landscape of
immune cells has direct implications for prognosis (62,63).
Despite the promising findings, the present study
has several limitations that must be acknowledged. First, most
conclusions were drawn from association analyses based on publicly
available transcriptomic datasets, which may introduce selection
bias and cannot fully reflect protein-level regulation. Second, the
in vitro validation was limited to a small number of LUAD
cell lines, and thus, may not capture the heterogeneity of the
disease. Third, in vivo evidence from animal models is still
lacking, which restricts the translational interpretation of
SCN4B function in lung tumor progression. Future research
should therefore include validation in larger independent clinical
cohorts, incorporate proteomic and in vivo functional
assays, and explore the underlying molecular mechanisms by which
SCN4B regulates EMT and tumor metastasis. Addressing these
aspects will provide a more comprehensive understanding of
SCN4B and strengthen its potential as a prognostic biomarker
and therapeutic target in LUAD.
In conclusion, low SCN4B expression was
closely associated with poor prognosis, whereas its high expression
may suppress LUAD cell proliferation, migration and invasion by
negatively regulating the EMT process.
Acknowledgements
Not applicable.
Funding
Funding: No funding was received.
Availability of data and materials
The data generated in the present study are
included in the figures and/or tables of this article.
Authors' contributions
MG performed the experiments, collected the data
and drafted the manuscript. HL contributed to preliminary research
and data analysis. ZZ and YW contributed to the conception and
methodology of the study, provided technical input for experimental
design and troubleshooting, and critically revised the manuscript
for important intellectual content. JT and BZ participated in
experimental implementation. YZ conceived and designed the study,
and was responsible for overall supervision. MG and HL confirm the
authenticity of all the raw data. All authors read and approved the
final manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Zheng M: Classification and pathology of
lung cancer. Surg Oncol Clin N Am. 25:447–468. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Mithoowani H and Febbraro M:
Non-small-cell lung cancer in 2022: A review for general
practitioners in oncology. Curr Oncol. 29:1828–1839. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Hirsch FR, Scagliotti GV, Mulshine JL,
Kwon R, Curran WJ, Wu YL and Paz-Ares L: Lung cancer: Current
therapies and new targeted treatments. Lancet. 389:299–311. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Jonna S and Subramaniam DS: Molecular
diagnostics and targeted therapies in non-small cell lung cancer
(NSCLC): An update. Discov Med. 27:167–170. 2019.PubMed/NCBI
|
|
5
|
Zhang Y, Chen M, Fang X, Han Y and Li Y:
Progression and metastasis of lung cancer: Clinical features,
molecular mechanisms, and clinical managements. Medcomm.
6:e704772025. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Manfioletti G and Fedele M:
Epithelial-mesenchymal transition (EMT). Int J Mol Sci.
24:113862023. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Manfioletti G and Fedele M:
Epithelial-mesenchymal transition (EMT) 2021. Int J Mol Sci.
23:58482022. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Jiang Y, Han Q, Zhao H and Zhang J:
Promotion of epithelial-mesenchymal transformation by
hepatocellular carcinoma-educated macrophages through
Wnt2b/β-catenin/c-Myc signaling and reprogramming glycolysis. J Exp
Clin Canc Res. 40:132021. View Article : Google Scholar
|
|
9
|
Lin YT and Wu KJ: Epigenetic regulation of
epithelial-mesenchymal transition: Focusing on hypoxia and TGF-β
signaling. J Biomed Sci. 27:392020. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Shi Q, Xue C, Zeng Y, Yuan X, Chu Q, Jiang
S, Wang J, Zhang Y, Zhu D and Li L: Notch signaling pathway in
cancer: From mechanistic insights to targeted therapies. Signal
Transduct Tar. 9:1282024. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Ang HL, Mohan CD, Shanmugam MK, Leong HC,
Makvandi P, Rangappa KS, Bishayee A, Kumar AP and Sethi G:
Mechanism of Epithelial-mesenchymal transition in cancer and its
regulation by natural compounds. Med Res Rev. 43:1141–1200. 2023.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Xue W, Yang L, Chen C, Ashrafizadeh M,
Tian Y and Sun R: Wnt/β-catenin-driven EMT regulation in human
cancers. Cell Mol Life Sci. 81:792024. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Wei P, Zhang N, Wang Y, Li D, Wang L, Sun
X, Shen C, Yang Y, Zhou X and Du X: FOXM1 promotes lung
adenocarcinoma invasion and metastasis by upregulating SNAIL. Int J
Biol Sci. 11:186–198. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Pan J, Fang S, Tian H, Zhou C, Zhao X,
Tian H, He J, Shen W, Meng X, Jin X and Gong Z: lncRNA
JPX/miR-33a-5p/Twist1 axis regulates tumorigenesis and metastasis
of lung cancer by activating Wnt/β-catenin signaling. Mol Cancer.
19:92020. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Clarhaut J, Gemmill RM, Potiron VA,
Ait-Si-Ali S, Imbert J, Drabkin HA and Roche J: ZEB-1, a repressor
of the semaphorin 3F tumor suppressor gene in lung cancer cells.
Neoplasia. 11:157–166. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Wang X, Li M, Yin J, Fang J, Ying Y, Ye T,
Zhang F, Ma S, Qin H and Liu X: Emetine dihydrochloride alleviated
radiation-induced lung injury through inhibiting EMT. J Cell Mol
Med. 27:3839–3850. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Chanvorachote P, Petsri K and Thongsom S:
Epithelial to mesenchymal transition in lung cancer: Potential
EMT-Targeting natural product-derived compounds. Anticancer Res.
42:4237–4246. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Al-Ward H, Liu CY, Liu N, Shaher F,
Al-Nusaif M, Mao J and Xu H: Voltage-gated sodium channel β1 Gene:
An overview. Hum Hered. 85:101–109. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Lewis AH and Raman IM: Resurgent current
of voltage-gated Na(+) channels. J Physiol. 592:4825–4838. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Ślęczkowska M, Almomani R, Marchi M, Salvi
E, de Greef B, Sopacua M, Hoeijmakers J, Lindsey P, Waxman SG,
Lauria G, et al: Peripheral ion channel genes screening in painful
small fiber neuropathy. Int J Mol Sci. 23:140952022. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Lu Y, Yu W, Xi Z, Xiao Z, Kou X and Wang
XF: Mutational analysis of SCN2B, SCN3B and SCN4B in a large
Chinese Han family with generalized tonic-clonic seizure. Neurol
Sci. 31:675–677. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Dai W, Zhou J, Wang H, Zhang M, Yang X and
Song W: miR-424-5p promotes the proliferation and metastasis of
colorectal cancer by directly targeting SCN4B. Pathol Res Pract.
216:1527312020. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Li X, Chen W, Jiang S, Zhang L, Huang H,
Ji Y, Ni Q and Ling C: Low expression of SCN4B predicts poor
prognosis in Non-small cell lung cancer. Curr Cancer Drug Targets.
25:445–466. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Peinado H, Ballestar E, Esteller M and
Cano A: Snail mediates E-cadherin repression by the recruitment of
the Sin3A/histone deacetylase 1 (HDAC1)/HDAC2 complex. Mol Cell
Biol. 24:306–319. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Ma Q, Ye S, Liu H, Zhao Y, Mao Y and Zhang
W: HMGA2 promotes cancer metastasis by regulating
epithelial-mesenchymal transition. Front Oncol. 14:13208872024.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
R Core Team R, . A language and
environment for statistical computing. R Foundation for Statistical
Computing; Vienna, Austria: 2025, URL. http://www.R-project.org/
|
|
27
|
Hughes G, Kopetzky J and McRoberts N:
Mutual information as a performance measure for binary predictors
characterized by both ROC curve and PROC curve analysis. Entropy
(Basel). 22:9382020. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Gustavsson EK, Zhang D, Reynolds RH,
Garcia-Ruiz S and Ryten M: Ggtranscript: An R package for the
visualization and interpretation of transcript isoforms using
ggplot2. Bioinformatics. 38:3844–3846. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Wu X, Sui Z, Zhang H, Wang Y and Yu Z:
Integrated analysis of lncRNA-mediated ceRNA network in lung
adenocarcinoma. Front Oncol. 10:5547592020. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Hess AS and Hess JR: Kaplan-Meier survival
curves. Transfusion. 60:670–672. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Yamauchi M, Yamaguchi R, Nakata A, Kohno
T, Nagasaki M, Shimamura T, Imoto S, Saito A, Ueno K, Hatanaka Y,
et al: Epidermal growth factor receptor tyrosine kinase defines
critical prognostic genes of stage I lung adenocarcinoma. PLoS One.
7:e439232012. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Feng X, Zahed H, Onwuka J, Callister M,
Johansson M, Etzioni R and Robbins HA: Cancer stage compared with
mortality as end points in randomized clinical trials of cancer
screening: A systematic review and Meta-analysis. JAMA.
331:1910–1917. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Qin H, Lu H, Qin C, Huang X, Peng K, Li Y,
Lan C, Bi A, Huang Z, Wei Y, et al: Pan-cancer analysis suggests
that LY6H is a potential biomarker of diagnosis,
immunoinfiltration, and prognosis. J Cancer. 15:5515–5539. 2024.
View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Wang Y, Wang H, Pan T, Li L, Li J and Yang
H: STIM1 silencing inhibits the migration and invasion of A549
cells. Mol Med Rep. 16:3283–3289. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Albuquerque C, Manguinhas R, Costa JG, Gil
N, Codony-Servat J, Castro M, Miranda JP, Fernandes AS, Rosell R
and Oliveira NG: A narrative review of the migration and invasion
features of non-small cell lung cancer cells upon xenobiotic
exposure: Insights from in vitro studies. Transl Lung Cancer Res.
10:2698–2714. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Guo Q, Yu W, Tan J, Zhang J, Chen J, Rao
S, Guo X and Cai K: Remodelin delays non-small cell lung cancer
progression by inhibiting NAT10 via the EMT pathway. Cancer Med.
13:e72832024. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Liu X, Chen Y, Li Y, Petersen RB and Huang
K: Targeting mitosis exit: A brake for cancer cell proliferation.
Biochim Biophys Acta Rev Cancer. 1871:179–191. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Venugopal N, Yeh J, Kodeboyina SK, Lee TJ,
Sharma S, Patel N and Sharma A: Differences in the early stage gene
expression profiles of lung adenocarcinoma and lung squamous cell
carcinoma. Oncol Lett. 18:6572–6582. 2019.PubMed/NCBI
|
|
40
|
Seetharaman S and Etienne-Manneville S:
Cytoskeletal crosstalk in cell migration. Trends Cell Biol.
30:720–735. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Biswas KH: Molecular Mobility-mediated
regulation of E-cadherin adhesion. Trends Biochem Sci. 45:163–173.
2020. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Morana O, Wood W and Gregory CD: The
apoptosis paradox in cancer. Int J Mol Sci. 23:13282022. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Moyer A, Tanaka K and Cheng EH: Apoptosis
in cancer biology and therapy. Annu Rev Pathol. 20:303–328. 2025.
View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Yang S, Zhang D, Sun Q, Nie H, Zhang Y,
Wang X, Huang Y and Sun Y: Single-cell and spatial transcriptome
profiling identifies the transcription factor BHLHE40 as a driver
of EMT in metastatic colorectal cancer. Cancer Res. 84:2202–2217.
2024. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Corso G, Figueiredo J, De Angelis SP,
Corso F, Girardi A, Pereira J, Seruca R, Bonanni B, Carneiro P,
Pravettoni G, et al: E-cadherin deregulation in breast cancer. J
Cell Mol Med. 24:5930–5936. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Na TY, Schecterson L, Mendonsa AM and
Gumbiner BM: The functional activity of E-cadherin controls tumor
cell metastasis at multiple steps. Proc Natl Acad Sci USA.
117:5931–5937. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Osuka S, Zhu D, Zhang Z, Li C, Stackhouse
CT, Sampetrean O, Olson JJ, Gillespie GY, Saya H, Willey CD and Van
Meir EG: N-cadherin upregulation mediates adaptive radioresistance
in glioblastoma. J Clin Invest. 131:e1360982021. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Gerber TS, Ridder DA, Goeppert B, Brobeil
A, Stenzel P, Zimmer S, Jakel J, Metzig MO, Schwab R, Martin SZ, et
al: N-cadherin: A diagnostic marker to help discriminate primary
liver carcinomas from extrahepatic carcinomas. Int J Cancer.
154:1857–1868. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Ridge KM, Eriksson JE, Pekny M and Goldman
RD: Roles of vimentin in health and disease. Gene Dev. 36:391–407.
2022. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Cui H, Huang J, Lei Y, Chen Q, Hu Z, Niu
J, Wei R, Yang K, Li H, Lu T, et al: Design and synthesis of dual
inhibitors targeting snail and histone deacetylase for the
treatment of solid tumour cancer. Eur J Med Chem. 229:1140822022.
View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Huang Y, Hong W and Wei X: The molecular
mechanisms and therapeutic strategies of EMT in tumor progression
and metastasis. J Hematol Oncol. 15:1292022. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Hao Y, Baker D and Ten DP: TGF-β-Mediated
Epithelial-mesenchymal transition and cancer metastasis. Int J Mol
Sci. 20:27672019. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Brackenbury WJ and Isom LL: Na Channel β
Subunits: Overachievers of the ion channel family. Front Pharmacol.
2:532011. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Peng L, Gao R, Han L, Fu T and Bian C:
Roles of β-catenin protein in non-small cell lung cancer (Review).
Oncol Lett. 30:5652025. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Yu F, Yu C, Li F, Zuo Y, Wang Y, Yao L, Wu
C, Wang C and Ye L: Wnt/β-catenin signaling in cancers and targeted
therapies. Signal Transduct Tar. 6:3072021. View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Fontana R, Mestre-Farrera A and Yang J:
Update on Epithelial-mesenchymal plasticity in cancer progression.
Annu Rev Pathol. 19:133–156. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Haddadin L and Sun X: Stem cells in
cancer: From mechanisms to therapeutic strategies. Cells.
14:5382025. View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Hu Q, Saleem K, Pandey J, Charania AN,
Zhou Y and He C: Cell adhesion molecules in fibrotic diseases.
Biomedicines. 11:19952023. View Article : Google Scholar : PubMed/NCBI
|
|
59
|
Lin W, Fang J, Wei S, He G, Liu J, Li X,
Peng X, Li D, Yang S, Li X, et al: Extracellular vesicle-cell
adhesion molecules in tumours: Biofunctions and clinical
applications. Cell Commun Signal. 21:2462023. View Article : Google Scholar : PubMed/NCBI
|
|
60
|
Morla S: Glycosaminoglycans and
glycosaminoglycan mimetics in cancer and inflammation. Int J Mol
Sci. 20:19362019. View Article : Google Scholar
|
|
61
|
Xiao Y and Yu D: Tumor microenvironment as
a therapeutic target in cancer. Pharmacol Ther. 221:1077532021.
View Article : Google Scholar : PubMed/NCBI
|
|
62
|
Du T, Gao J, Li P, Wang Y, Qi Q, Liu X, Li
J, Wang C and Du L: Pyroptosis, metabolism, and tumor immune
microenvironment. Clin Transl Med. 11:e4922021. View Article : Google Scholar : PubMed/NCBI
|
|
63
|
Elhanani O, Ben-Uri R and Keren L: Spatial
profiling technologies illuminate the tumor microenvironment.
Cancer Cell. 41:404–420. 2023. View Article : Google Scholar : PubMed/NCBI
|