Introduction
Head and neck squamous cell carcinoma (HNSCC) is one
of the most prevalent types of cancer, accounting for ~3.6% of all
new cancer cases in the United States (1). Advances in the treatment of HNSCC have
been made over the past few decades, but the mortality rate has
remained essentially unchanged (2,3). HNSCC
encompasses a diverse range of tumors, including squamous cell
carcinoma (SCC) in the oral cavity, pharynx, larynx, nasal cavity
and paranasal sinuses and salivary glands (4). Human papillomavirus (HPV) is a widely
recognized risk factor for carcinogenesis and prognosis in patients
with oropharyngeal squamous cell carcinoma (OPSCC). HPV-positive
OPSCC is associated with a more favorable prognosis, whereas
HPV-negative OPSCC is associated with a poor prognosis.
Furthermore, high cell proliferation also emerges as a firmly
established adverse prognostic factor. A meta-analysis revealed a
correlation between a high proliferation index and lower survival
in patients with HNSCC (5).
Angiopoietin-like 4 (ANGPTL4) is a member of the
angiopoietin-related family, which has been reported to play a
crucial role in regulating glucose and lipid metabolism (6). In the field of oncology, it has been
previously demonstrated that ANGPTL4 has been reported in various
cancers, including HNSCC (7).
ANGPTL4 has been shown to play multiple roles in cancer
progression, including tumor growth, anoikis resistance,
angiogenesis, tumor invasion and metastasis. Most previous studies
have described that the upregulation of ANGPTL4 is associated with
the promotion of tumor growth, progression, angiogenesis, invasion
and metastasis and reduces overall survival (OS) (8–14). By
contrast, ANGPTL4 has also been reported to have an inhibitory
function against tumor growth, angiogenesis, and vascular
leakiness, prevents metastasis and is associated with an improved
prognosis (15–18). Hsieh et al (19) reported that ANGPTL4 had both
oncogenic and tumor-suppressing roles in urothelial carcinoma.
These conflicting results indicate that ANGPTL4 could be both
promotive and inhibitory for tumorigenesis, depending on tissue
context and organ site. Few studies have reported on the roles of
ANGPTL4 in HNSCC. In the present study, it was investigated whether
ANGPTL4 expression is associated with pro- or anti-tumorigenic
effects in OPSCC.
Materials and methods
Patient and tissue specimens
The data of 146 patients with OPSCC who underwent
initial surgery or biopsy at the Department of Otolaryngology, Head
and Neck Surgery, Keio University School of Medicine between April
2005 and September 2018 were retrospectively reviewed. A total of
three patients who had received chemotherapy and one patient who
had received radiotherapy for the head and neck region before the
initial surgery or biopsy were excluded. A total of five patients
who underwent biopsy only and were treated at other hospitals were
also excluded. Consequently, 137 patients were enrolled in the
present study. The cohort included 116 men and 21 women. Their
median age was 64 years (range, 37–87 years). Their characteristics
(age, sex, smoking and alcohol status, tumor subsite, TNM
classification, pathologic characteristics, and follow-up
examination findings) were extracted from their medical records.
The TNM classification was determined based on the eighth edition
of the American Joint Committee on Cancer staging manual (20). The histopathological diagnoses were
based on the World Health Organization criteria (21). Tissue samples were obtained from the
hospital tissue bank.
The treatment strategy for OPSCC in the present
study was as follows. Early T-stage (T1 and T2) tumors were treated
with transoral resection or radiotherapy alone. Advanced T-stage
(T3 and T4) tumors were treated with concurrent chemoradiotherapy
(chemotherapy: Cisplatin 80 mg/m2, administered 2–3
times every 3 weeks; radiotherapy: 2.0 Gy/fraction, administered
five times a week for a total dose of 60–66 Gy) or radical tumor
resection with functional reconstruction. The patients with
multiple LN metastases underwent neck dissection before
radiotherapy or concurrently with primary surgery. The patients
with resectable locoregional recurrences or neck metastases
detected during follow-up underwent additional resections
immediately. They received chemotherapy as palliative treatment for
persistent disease or distant metastases if they were amenable to
treatment according to the NCCN guidelines (4).
Immunohistochemistry (IHC)
The paraffin sections were sliced at 5 µm each,
dewaxed in xylene, rehydrated in ethanol, and washed in water.
Heat-induced antigen retrieval was performed using the Decloaking
Chamber NxGen® (Biocare Medical, LLC) with Dako Target
Retrieval Solution (cat. no. S1700; Dako; Agilent Technologies,
Inc.) at 95°C for 1 h. Endogenous peroxidase activity was quenched
by treatment with 3% hydrogen peroxide solution for 10 min.
Non-specific binding of the primary antibodies was blocked by
treatment with normal goat serum (cat. no. IHR-8136;
ImmunoBioScience/IBC) in phosphate-buffered saline (PBS) containing
TWEEN®20 for 1 h at room temperature. The slides were
stained with the primary antibody (1:500) overnight at 4°C. They
were washed and incubated with SignalStain® Boost IHC
Reagent (HRP; Rabbit; cat. no. 8114; Cell Signaling Technology,
Inc.) for 30 min at room temperature. Peroxidase activity was
visualized using a DAB Substrate Kit (cat. no. SK-4100; Vector
Laboratories, Inc.). The sections were counterstained and sealed
with hematoxylin solution. The stained slides were imaged using a
light microscope. The tissue images were imported, and the
proportions of ANGPTL4-positive tumor cells were determined using
Tissue Studio® (Definiens). The tumorous area was
selected as the region of interest for each slide. The following
parameters were set: Hematoxylin threshold of 0.2, typical nucleus
size of 23 µm2, maximum cell growth of 5, and
classification of 0.15. ANGPTL4 expression was automatically
calculated as the ratio of the number of ANGPTL4-positive tumor
cells to the total number of tumor cells.
Cell line
FaDu, Detroit 562, OSC-19, HSC-2 and HSC-4 (human
HNSCC cell lines) were obtained from the American Type Culture
Collection.
Cell culture
The cell lines were cultured in Eagle's Minimum
Essential Medium (Sigma-Aldrich; Merck KGaA) supplemented with 10%
fetal bovine serum (FBS) and 1% penicillin-streptomycin (solution
stabilized; Sigma-Aldrich; Merck KGaA). They were incubated in a
humidified incubator at 37°C in a 5% carbon dioxide environment.
The cells were sub-cultured continuously in accordance with the
American Type Culture Collection protocol.
Reverse transcription-quantitative
polymerase chain reaction (RT-qPCR)
RNA was extracted using the RNeasy Mini Kit (Qiagen
GmbH), and its quality was assessed using the Nanodrop 1000 (Thermo
Fisher Scientific, Inc.). Complementary DNA synthesis was performed
using the Superscript III First-Strand Synthesis System
(Invitrogen; Thermo Fisher Scientific, Inc.) according to the
manufacturer's instructions. RT-qPCR was conducted using the
StepOnePlus Real-Time PCR system and software (Applied Biosystems;
Thermo Fisher Scientific, Inc.) following the manufacturer's
protocol. The primers and probes were procured from Applied
Biosystems (TaqMan® Gene Expression Assays) and had the
following IDs: ANGPTL4 (Hs01101127_m1), ACTB
(Hs01060665_g1), MKI67 (Hs04260396_g1), BAX
(Hs0018269_m1), BCL2 (Hs00608023_m1), and CASP3
(Hs00234387_m1). According to the manufacturer's product
information, the sequences for these pre-designed assays are not
publicly disclosed. PCR amplification involved initial denaturation
at 95°C for 20 sec, followed by 40 cycles of denaturation at 95°C
for 3 sec and annealing at 60°C for 30 sec. The relative mRNA
expression levels were determined using the 2−ΔΔCq
method and were compared with those of ACTB, which served as
the endogenous control (22).
Small interfering RNA (siRNA)
knockdown
The FaDu cells were seeded in a 6-well dish at a
density of 25,000 cells/ml and incubated in a medium containing 10%
FBS for 24 h. The medium was changed, and the cells were
transfected with 30 nM siRNA targeting ANGPTL4 (cat. no.
NM_001039667; ID: SASI_Hs01_00144609; Sigma-Aldrich; Merck KGaA)
and negative control (MISSION siRNA Universal Negative Control #1;
Sigma-Aldrich; Merck KGaA) using 4 µl of Lipofectamine 2000
Transfection Reagent (Thermo Fisher Scientific, Inc.) in 200 µl of
Opti-MEM medium (Thermo Fisher Scientific, Inc.) for 30 min at room
temperature. The Opti-MEM medium was removed, and 2 ml of fresh
culture medium was added. The cells were incubated for an
additional 24 h, scraped, and collected for analysis. The
efficiency of ANGPTL4 knockdown was evaluated using RT-qPCR.
According to the manufacturer's product information, the sequences
of the siRNAs used in the present study are not publicly
disclosed.
In addition to FaDu cells, ANGPTL4 knockdown
experiments were attempted in other HNSCC cell lines, including
Detroit 562, OSC-19, HSC-2 and HSC-4, using the same siRNA sequence
and transfection protocols.
Cell proliferation assay
(CellTiter-Glo 2.0 luminescence assay)
The FaDu cells were seeded in a 96-well plate (3.000
cells per well) and incubated overnight after transfection. Their
viability was subsequently assessed using a CellTiter-Glo 2.0
luminescence-based assay kit (cat. no. G9241; Promega Corporation).
The results were normalized to those of the negative control (set
as 1.0).
Immunofluorescence staining
The FaDu cells were seeded in slide chambers
(CHAMBER SLIDEII IWAKI, http://iwaki.atgc.co.jp) after ANGPTL4
knockdown for immunofluorescence analyses of ANGPTL4 and Ki-67. The
cells were washed extensively with PBS and fixed with 4%
paraformaldehyde for 15 min at room temperature. The samples were
washed with PBS and blocked with 10% normal goat serum in PBS for 1
h. The cells were incubated with the primary antibodies overnight
at 4°C. They were subsequently incubated with Alexa Fluor Plus 555-
and 488-conjugated secondary antibodies (cat. nos. A32732 and
A32723; Thermo Fisher Scientific, Inc.) against ANGPTL4 and Ki-67,
respectively, and observed 24 h after transfection. Hoechst 33258
(Sigma-Aldrich) was used for nuclear staining. The numbers of
ANGPTL4- and Ki-67-positive cells were determined based on counts
in four randomly selected areas at ×10 magnification using a
BZ-X710 fluorescence microscope (Keyence Corporation).
Antibody
The primary antibodies used for immunostaining were
anti-ANGPTL4 (rabbit; 1:500; cat. no. ab115798; Abcam) for
immunofluorescence staining and IHC and anti-Ki-67 (mouse; 1:100;
cat. no. ab245113; Abcam) for immunofluorescence staining.
The cancer genome atlas (TCGA)
RNA sequencing and corresponding clinical data for
HNSCC were obtained from TCGA via the University of California
Santa Cruz Cancer Browser (https://xenabrowser.net/). A total of 78 patients with
primary OPSCC and corresponding clinical data from the cohort
labeled ‘GDC TCGA Head and Neck Cancer’ were included. The patients
were divided into high and low groups based on their ANGPTL4
expression levels (FPKM-UQ). The prognoses of 128 and 116 patients
with tongue and laryngeal SCCs, respectively, were also evaluated.
The ANGPTL4 expression data for 520 patients with HNSCC were
extracted based on the subsites. Gene set enrichment analysis
(GSEA) was performed based on the ANGPTL4 expression to
explore its biological role in OPSCC. GSEA was performed using the
Hallmark gene set from the Molecular Signatures Database (version
7.5.1, http://www.gsea-msigdb.org/gsea/msigdb/index.jsp). The
enrichment score, normalized enrichment score, nominal P-value and
false discovery rate (FDR) were determined using UCSC Xena
(https://xenabrowser.net/). Statistical
significance was set at FDR <0.05. The differentially expressed
genes (DEGs) were extracted based on a fold change of >2 or
<-2 and adjusted P-value of <0.05.
Statistical analysis
The five-year OS and disease-free survival (DFS)
rates of the patients with OPSCC were determined using the
Kaplan-Meier method, and survival curves were compared using the
log-rank test. When survival curves crossed, weighted tests
(Breslow and Tarone-Ware) were additionally performed as
sensitivity analyses. The survival durations were calculated from
the date of initial treatment to the date of the event or the
latest follow-up. The variables included were age, sex, smoking and
alcohol status, T classification, N classification, M
classification, TNM stage, p16 status, initial definitive therapy
and ANGPTL4 status. To determine the optimal cut-off value for
ANGPTL4 expression, receiver operating characteristic (ROC) curve
analysis was performed using the OS. The cut-off value was defined
as the point that maximized Youden's index. The ROC curve is
provided in Fig. S1. To evaluate
the robustness of the selected cut-off value, sensitivity analyses
were performed using alternative thresholds for ANGPTL4 expression.
Tumors were reclassified using the median expression level, as well
as fixed cut-off values of 10 and 15%. The OS and DFS of the
patients in the subgroups were compared using the log-rank test
during the univariate analysis. The factors that were significant
in the univariate analysis were further analyzed using multivariate
analysis. This was performed using a Cox proportional hazards model
with backward elimination. The correlations between the variables
analyzed in the multivariate analysis were examined using Pearson's
correlation coefficient to avoid multicollinearity. The
relationships between ANGPTL4 and the other variables were
evaluated. The distributions of the categorical variables for
ANGPTL4 and the other variables were compared using the chi-squared
test. In addition, survival analyses were performed stratified by
p16 status to evaluate the prognostic impact of ANGPTL4 within each
subgroup. Furthermore, sensitivity analyses were performed using
the multivariable Cox models in patients who underwent definitive
therapy. Associations between continuous variables were analyzed
using unpaired Student's t-test. The differences in ANGPTL4
expression across the subsites were determined using one-way
analysis of variance. Post hoc tests were not performed.
Statistical analyses were performed using SPSS version 27 for Mac
(IBM Corp.). P<0.05 was considered to indicate a statistically
significant difference.
Results
High ANGPTL4 expression is associated
with a favorable prognosis in patients with OPSCC
The characteristics of the 137 patients with OPSCC
included in the present study are presented in Table I. In total, 40 had locally advanced
disease (T3 and T4), 101 had cervical lymph node metastases, 2 had
distant metastases, and 39 had advanced TNM stages (III and IV). A
total of ~80% of the patients were positive for p16 staining. The
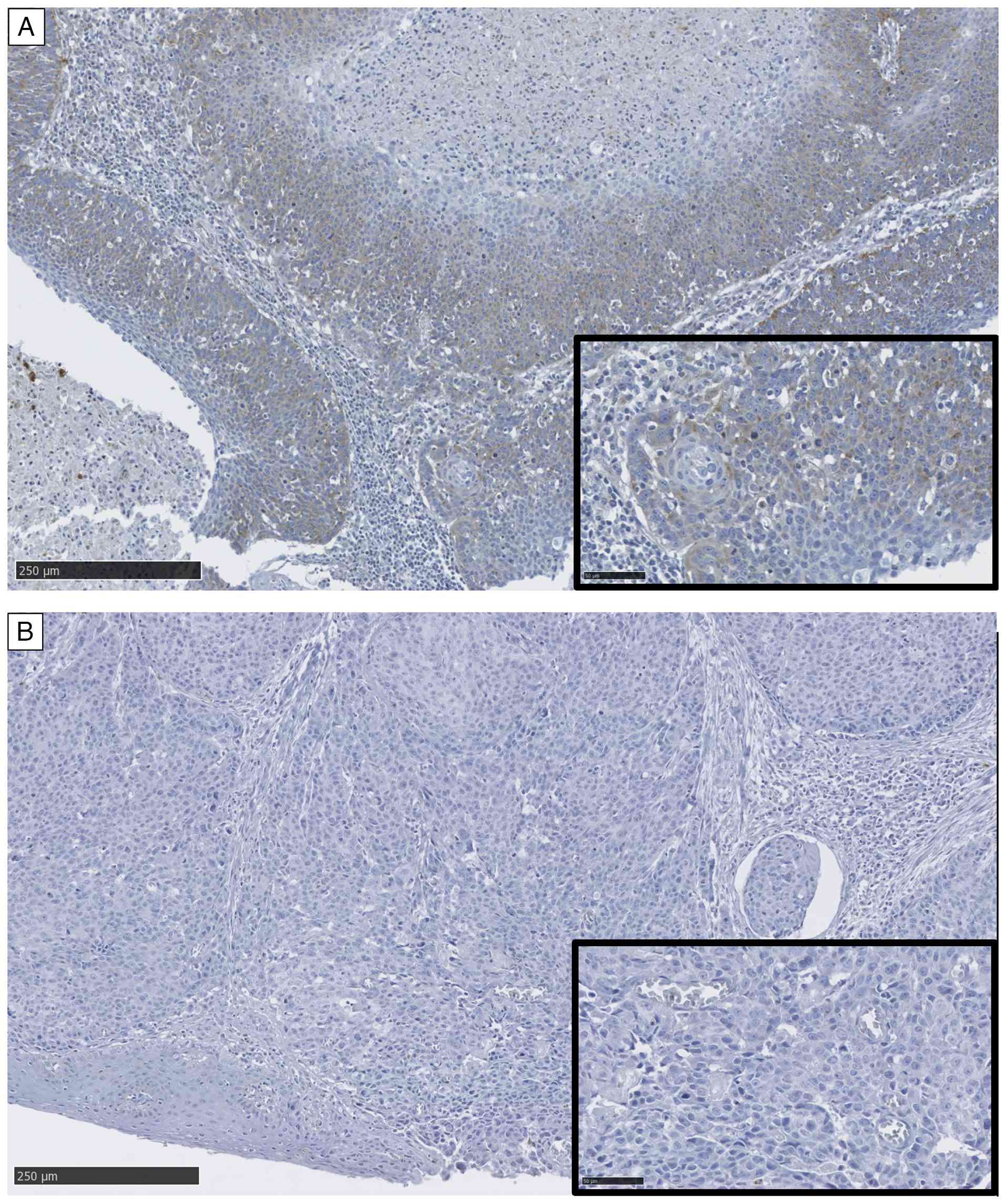
IHC staining for ANGPTL4 is shown in Fig. 1. Totally, 71 and 66 patients had
positive and negative staining for ANGPTL4, respectively. The
5-year OS and DFS rates were 75.1 and 67.6%, respectively. The
median follow-up duration was 52 months (range, 1–171 months). The
univariate analyses of OS and DFS based on the clinicopathological
factors are summarized in Table
II. T classification (T3 and T4), M classification (M1), TNM
stage (III and IV) and ANGPTL4 (<7.7%) were significantly
associated with poorer OS (P=0.001, P<0.001, P=0.034 and
P=0.002, respectively). The subsite (anterior wall), T
classification (T3 and T4), M classification (M1), TNM stage (III
and IV) and ANGPTL4 (<7.7%) were significant predictors of worse
DFS in patients with OPSCC (P=0.007, P<0.001, P<0.001,
P=0.007, and P<0.001, respectively). Multivariate analysis
revealed that T classification [hazard ratio (HR)=3.209; 95%
confidence interval (CI), 1.605–6.415; P<0.001], M
classification (HR=94.613; 95% CI, 15.385–581.837; P<0.001) and
ANGPTL4 (HR=3.676; 95% CI, 1.678–8.056; P=0.001) were independent
prognostic factors for OS. The subsite (HR=2.161; 95% CI,
1.183–3.947; P=0.012), T classification (HR=3.029; 95% CI,
1.652–5.550; P<0.001), M classification (HR=15.081; 95% CI,
3.083–73.771; P<0.001) and ANGPTL4 (HR=2.959; 95% CI,
1.533–5.713; P=0.001) were independent prognostic factors for DFS
in patients with OPSCC (Table
III). The Kaplan-Meier curves for OS and DFS stratified by
ANGPTL4 expression are shown in Fig.
2. The 5-year OS rates for patients who tested positive and
negative for ANGPTL4 expression were 88.4 and 61.6%, respectively
(P=0.002). The corresponding 5-year DFS rates were 82.7 and 52.5%,
respectively (P<0.001). Sensitivity analyses using alternative
cut-off values including median ANGPTL4 expression level, 10%, and
15% demonstrated consistent results (Table SI). In addition, baseline
characteristics stratified by p16 status are summarized in Table SII. High ANGPTL4 expression was
associated with improved OS and DFS in the p16-positive subgroup,
whereas no significant prognostic impact was observed in the
p16-negative subgroup (Table
SIII). In sensitivity analyses restricted to patients who
received definitive therapy, additional adjustment for p16 status
(model 1) and for initial definitive therapy (model 2) did not
alter the prognostic association of ANGPTL4 with the OS and DFS
(Table SIV).
 | Table I.Patient characteristics. |
Table I.
Patient characteristics.
| Variables | Cases (n=137) | Percentage, % |
|---|
| Age |
|
|
| Mean
(SD) | 64 (10) |
|
| Sex |
|
|
|
Male/Female | 116/21 | 85/15 |
| Smoking |
|
|
|
Yes/No | 112/25 | 82/18 |
| Alcohol |
|
|
|
Yes/No | 104/33 | 76/24 |
| Subsite |
|
|
|
Anterior wall | 38 | 28 |
| Lateral
wall | 83 | 61 |
|
Posterior wall | 7 | 5 |
|
Superior wall | 9 | 7 |
| T
classification |
|
|
|
T0/T1/T2/T3/T4 | 1/40/56/21/19 | 1/29/41/15/14 |
| N
classification |
|
|
|
N0/N1/N2/N3 | 36/71/27/3 | 26/52/20/2 |
| M
classification |
|
|
|
M0/M1 | 135/2 | 99/1 |
| TNM stage |
|
|
|
0/I/II/III/IV | 1/68/29/23/16 | 1/50/21/17/12 |
| p16 |
|
|
|
Positive/Negative | 109/28 | 80/20 |
| ANGPTL4 |
|
|
|
≥7.7%/<7.7% | 71/66 | 52/48 |
 | Table II.Univariate analyses of prognostic
factors for OS and DFS of patients with oropharyngeal
carcinoma. |
Table II.
Univariate analyses of prognostic
factors for OS and DFS of patients with oropharyngeal
carcinoma.
| Variables | Cases | 5-year OS (%) | P-value | 5-year DFS (%) | P-value |
|---|
| Overall | 137 | 75.1 |
| 67.6 |
|
| Age |
|
| 0.685 |
| 0.063 |
|
<65 | 69 | 75.3 |
| 72.7 |
|
|
≥65 | 68 | 75.8 |
| 62.5 |
|
| Sex |
|
| 0.911 |
| 0.774 |
|
Male | 116 | 75.6 |
| 67.5 |
|
|
Female | 21 | 71.3 |
| 67.1 |
|
| Smoking |
|
| 0.400 |
| 0.320 |
|
Yes | 112 | 73.1 |
| 64.9 |
|
| No | 25 | 83.6 |
| 79.6 |
|
| Alcohol |
|
| 0.380 |
| 0.199 |
|
Yes | 104 | 72.5 |
| 65.0 |
|
| No | 33 | 83.6 |
| 76.4 |
|
| Subsite |
|
| 0.056 |
| 0.007 |
|
Anterior wall | 38 | 65.1 |
| 50.7 |
|
|
Others | 99 | 79.1 |
| 74.3 |
|
| T
classification |
|
| 0.001 |
| <0.001 |
|
0,1,2 | 97 | 81.9 |
| 77.1 |
|
|
3,4 | 40 | 57.1 |
| 43.6 |
|
| N
classification |
|
| 0.692 |
| 0.944 |
| 0 | 36 | 72.2 |
| 65.2 |
|
|
1,2,3 | 101 | 76.0 |
| 68.4 |
|
| M
classification |
|
| <0.001 |
| <0.001 |
| 0 | 135 | 76.2 |
| 68.6 |
|
| 1 | 2 | 0 |
| 0 |
|
| TNM stage |
|
| 0.034 |
| 0.007 |
| I,
II | 98 | 78.7 |
| 73.7 |
|
| III,
IV | 39 | 65.3 |
| 51.5 |
|
| p16 |
|
| 0.677 |
| 0.765 |
|
Positive | 109 | 74.0 |
| 67.8 |
|
|
Negative | 28 | 79.4 |
| 66.3 |
|
| Initial definitive
therapy |
|
| 0.910 |
| 0.275 |
|
Surgery | 75 | 77.3 |
| 74.7 |
|
|
Radiation or
chemoradiation | 53 | 79.2 |
| 67.9 |
|
| ANGPTL4 |
|
| 0.002 |
| <0.001 |
|
≥7.7% | 71 | 88.4 |
| 82.7 |
|
|
<7.7% | 66 | 61.6 |
| 52.5 |
|
 | Table III.Multivariate analysis of prognostic
factors for OS and DFS of patients with oropharyngeal
carcinoma. |
Table III.
Multivariate analysis of prognostic
factors for OS and DFS of patients with oropharyngeal
carcinoma.
|
| OS | DFS |
|---|
|
|
|
|
|---|
| Variables | HR | 95% CI | P-value | HR | 95% CI | P-value |
|---|
| Subsite (Anterior
wall) | - | - | - | 2.161 | 1.183–3.947 | 0.012 |
| T classification
(3, 4) | 3.209 | 1.605–6.415 | <0.001 | 3.029 | 1.652–5.550 | <0.001 |
| M classification
(1) | 94.613 | 15.385–581.837 | <0.001 | 15.081 | 3.083–73.771 | <0.001 |
| TNM stage (III,
IV) | 1.093 | 0.467–2.557 | 0.837 | 1.153 | 0.537–2.476 | 0.715 |
| ANGPTL4
(<7.7%) | 3.676 | 1.678–8.056 | 0.001 | 2.959 | 1.533–5.713 | 0.001 |
The correlations between ANGPTL4 expression and
clinicopathological factors in OPSCC are provided in Table IV. No significant correlations were
observed between ANGPTL4 expression and age, sex, subsite, stage,
or p16 expression.
 | Table IV.Univariate analysis of outcomes
stratified by ANGPTL4 expression. |
Table IV.
Univariate analysis of outcomes
stratified by ANGPTL4 expression.
|
| ANGPTL4 |
|
|---|
|
|
|
|
|---|
| Variables | ≥7.7% | <7.7% | P-value |
|---|
| Overall | 71 | 66 |
|
| Age |
|
| 0.795 |
|
<65 | 35 | 34 |
|
|
≥65 | 36 | 32 |
|
| Sex |
|
| 0.956 |
|
Male | 60 | 56 |
|
|
Female | 11 | 10 |
|
| Subsite |
|
| 0.518 |
|
Anterior wall | 18 | 20 |
|
|
Others | 53 | 46 |
|
| T
classification |
|
| 0.161 |
|
1,2 | 54 | 43 |
|
|
3,4 | 17 | 23 |
|
| N
classification |
|
| 0.602 |
| 0 | 20 | 16 |
|
|
1,2,3 | 51 | 50 |
|
| M
classification |
|
| 0.733 |
| 0 | 70 | 65 |
|
| 1 | 1 | 1 |
|
| TNM stage |
|
| 0.224 |
| I,
II | 54 | 44 |
|
| III,
IV | 17 | 22 |
|
| p16 |
|
| 0.828 |
|
Positive | 57 | 52 |
|
|
Negative | 14 | 14 |
|
ANGPTL4 expression inhibits cell
proliferation in the FaDu cell lines
The FaDu cells were transfected with siRNA targeting
ANGPTL4, and their proliferation was evaluated compared with
the negative controls. Their relative mRNA expression levels were
determined using RT-qPCR. The relative mRNA expression level of
ANGPTL4 decreased to 38%. The results of immunofluorescence
staining are provided in Fig. 3.
The knockdown cells had significantly reduced ANGPTL4 expression
and significantly increased Ki-67 expression compared with the
negative controls (P=0.021 and 0.021, respectively).
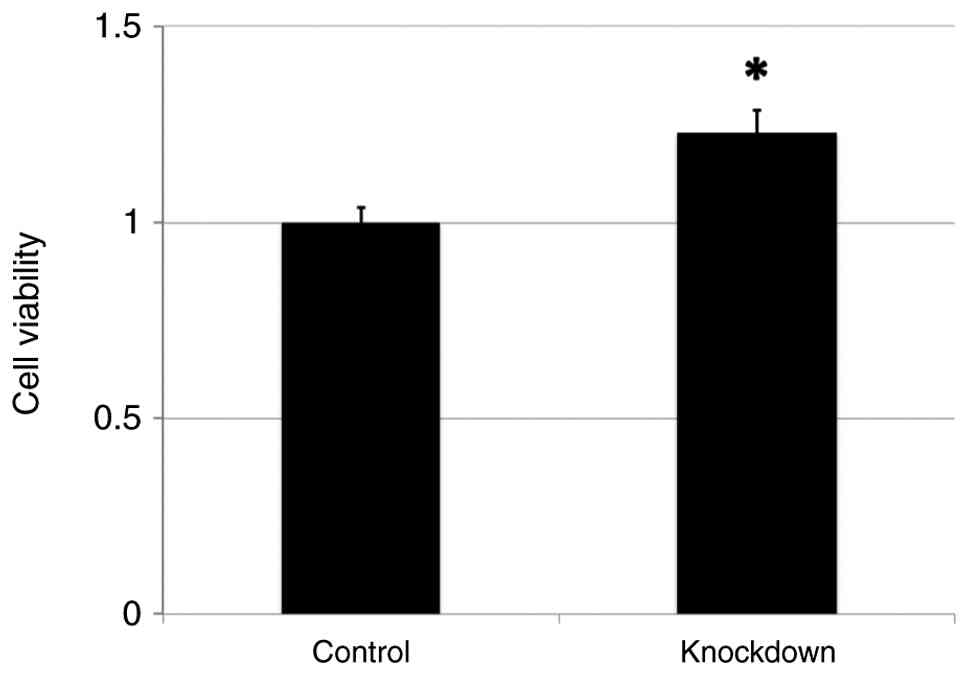
The FaDu cells with ANGPTL4 knockdown showed
significantly increased proliferation compared with the negative
controls (P=0.010) (Fig. 4).
ANGPTL4 knockdown was attempted in additional
HNSCC cell lines (Detroit 562, OSC-19, HSC-2 and HSC-4); however,
robust and reproducible knockdown could not be achieved. Therefore,
functional analyses were performed only in FaDu cells.
Knockdown of ANGPTL4 expression
enhances the cell proliferation signal in FaDu cells
The relative gene expression levels of various mRNAs
in the FaDu cells transfected with ANGPTL4 siRNA were
determined via RT-qPCR (Fig. 5).
The relative mRNA expression levels of ANGPTL4 decreased to
38%, and those of MKI67 increased significantly. The
expression of apoptosis-related genes was also evaluated. The
expression of BAX decreased, and that of BCL2
increased. In addition, the expression of CASP3 increased in
the ANGPTL4 knockdown cells.
ANGPTL4 expression has different
prognostic implications for the subsites of HNSCC based on TCGA
data
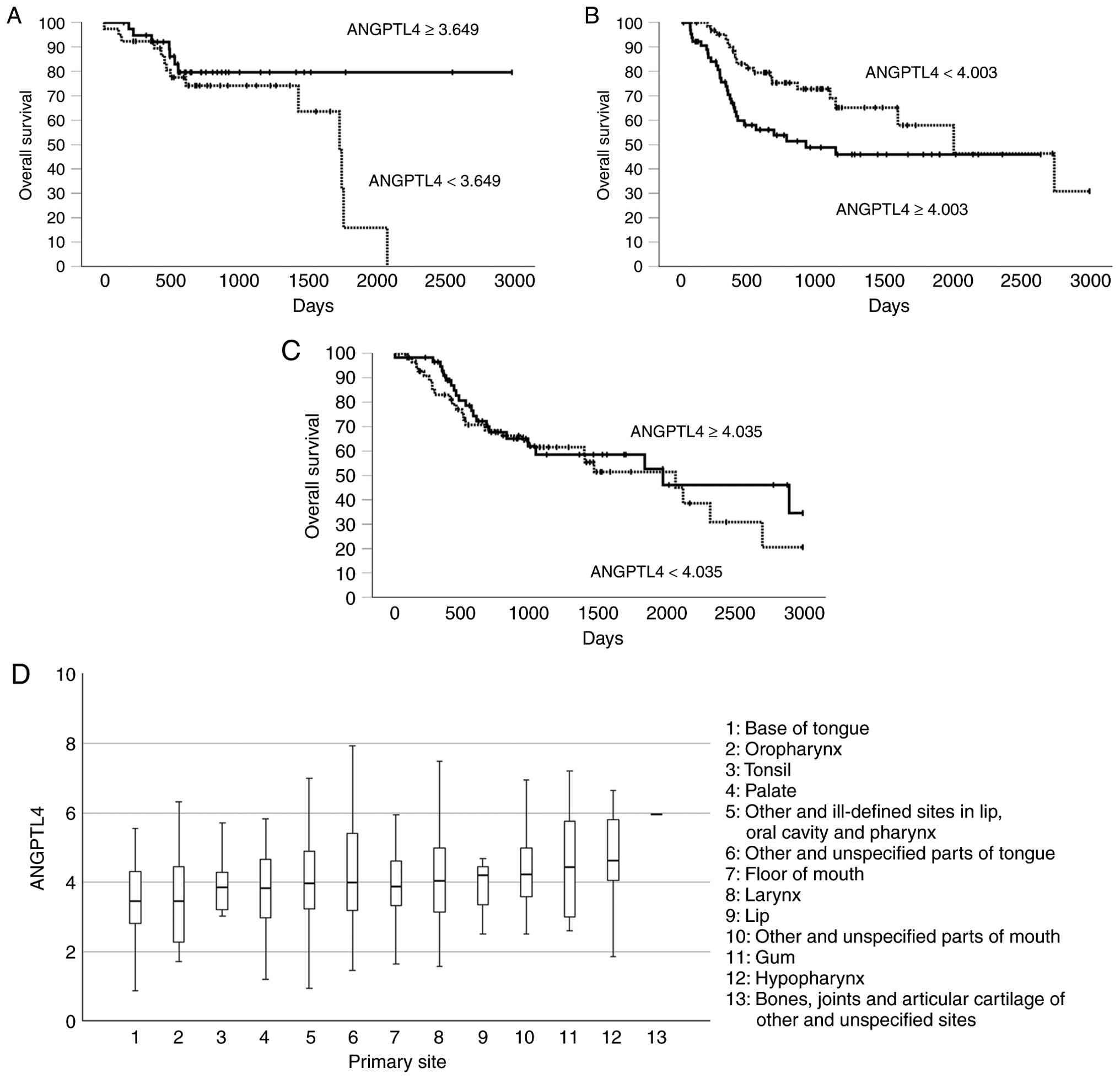
The correlation between ANGPTL4 expression
and prognosis was analyzed using TCGA data. High ANGPTL4
expression was associated with a tendency towards improved
prognosis in 78 patients with OPSCC, but the difference was not
statistically significant (P=0.09; Fig.
6A). The group with high ANGPTL4 expression had
significantly worse OS than the group with low expression in tongue
SCC (log-rank, P=0.02; Breslow, P=0.003; Tarone-Ware, P=0.005;
Fig. 6B). No significant
differences in OS were detected between the patients with laryngeal
SCC with high and low ANGPTL4 expressions (log-rank, P=0.45;
Breslow, P=0.47; Tarone-Ware, P=0.49; Fig. 6C). The mean ANGPTL4
expression differed significantly across HNSCC subsites (P=0.042;
Fig. 6D). The OPSCCs, including
those of the base of the tongue, oropharynx and tonsils, had lower
ANGPTL4 expression levels than those of other subsites.
ANGPTL4 expression is associated with
a cell proliferation signature in the GSEA
GSEA revealed significant enrichment of the
HALLMARK_E2F_TARGETS and HALLMARK_G2M_CHECKPOINT gene sets in the
ANGPTL4-low expression group (NES=6.86 and 4.42,
respectively; FDR <0.001 for both) (Table V). The
HALLMARK_TNFA_SIGNALING_VIA_NFKB,
HALLMARK_INTERFERON_GAMMA_RESPONSE, and
HALLMARK_INFLAMMATORY_RESPONSE gene sets were significantly
depleted in the ANGPTL4-low expression group. NDRG1
and CDH3 were identified as DEGs based on their fold changes
(>2 or <-2) and adjusted P-values (<0.05) (Table VI).
 | Table V.Result of Gene Set Enrichment
Analysis of in low ANGPTL4 expression using The Cancer
Genome Atlas data. |
Table V.
Result of Gene Set Enrichment
Analysis of in low ANGPTL4 expression using The Cancer
Genome Atlas data.
| Term | ES | NES | P-value | FDR |
|---|
|
HALLMARK_E2F_TARGETS | 0.452643176 | 5.244648544 |
1.57×10−7 |
7.12×10−7 |
|
HALLMARK_G2M_CHECKPOINT | 0.338413055 | 3.065174051 |
2.18×10−3 |
5.44×10−3 |
|
HALLMARK_INTERFERON_GAMMA_RESPONSE | −0.67230996 | −8.693683244 |
3.51×10−18 |
1.75×10−16 |
|
HALLMARK_TNFA_SIGNALING_VIA_NFKB | −0.658426312 | −8.483918737 |
2.18×10−17 |
5.44×10−16 |
|
HALLMARK_HYPOXIA | −0.615556414 | −7.657331309 |
1.90×10−14 |
3.16×10−13 |
|
HALLMARK_INFLAMMATORY_RESPONSE | −0.722841711 | −7.488593625 |
6.96×10−14 |
8.70×10−13 |
|
HALLMARK_INTERFERON_ALPHA_RESPONSE | −0.71989998 | −7.157175996 |
8.24×10−13 |
8.24×10−12 |
|
HALLMARK_EPITHELIAL_MESENCHYMAL_TRANSITION | −0.567963778 | −7.027159618 |
2.11×10−12 |
1.76×10−11 |
|
HALLMARK_COMPLEMENT | −0.582111115 | −6.1058472 |
1.02×10−9 |
7.30×10−9 |
|
HALLMARK_IL2_STAT5_SIGNALING | −0.573317571 | −5.768325086 |
8.01×10−9 |
5×10−8 |
|
HALLMARK_APICAL_JUNCTION | −0.540791657 | −5.611683099 |
2.00×10−8 |
1.11×10−7 |
|
HALLMARK_P53_PATHWAY | −0.480027635 | −5.561511073 |
2.67×10−8 |
1.34×10−7 |
|
HALLMARK_ESTROGEN_RESPONSE_LATE | −0.464291062 | −5.123247455 |
3.00×10−7 |
1.25×10−6 |
|
HALLMARK_GLYCOLYSIS | −0.466279034 | −5.039067076 |
4.68×10−7 |
1.80×10−6 |
|
HALLMARK_ALLOGRAFT_REJECTION | −0.497074315 | −4.905992143 |
9.3×10−7 |
3.32×10−6 |
|
HALLMARK_KRAS_SIGNALING_UP | −0.573985774 | −4.409147761 |
1.04×10−5 |
3.46×10−5 |
|
HALLMARK_ESTROGEN_RESPONSE_EARLY | −0.428017248 | −4.001047521 |
6.31×10−5 | 0.000197071 |
|
HALLMARK_APOPTOSIS | −0.386057656 | −3.582960349 | 0.000339722 | 0.000999183 |
|
HALLMARK_UV_RESPONSE_UP | −0.377520025 | −3.377546754 | 0.000731355 | 0.002031542 |
|
HALLMARK_UV_RESPONSE_DN | −0.49585039 | −3.229595402 | 0.001239655 | 0.00326225 |
|
HALLMARK_IL6_JAK_STAT3_Signaling | −0.515307214 | −2.744010577 | 0.006069358 | 0.014450853 |
 | Table VI.Significant altered gene expression
in low-angiopoietin-like 4 group based on The Cancer Genome Atlas
data. |
Table VI.
Significant altered gene expression
in low-angiopoietin-like 4 group based on The Cancer Genome Atlas
data.
|
| logFold Change | P-value | Adjusted
P-value |
|---|
| CDH3 | −1.21550025 |
1.89×10−5 | 0.030221783 |
| NDRG1 | −1.156636514 |
3.76×10−6 | 0.009006122 |
Discussion
Previous findings have revealed a critical role for
ANGPTL4 in cancer growth and progression, angiogenesis, metastasis
and anoikis resistance. ANGPTL4 has diverse roles of pro- and
anti-tumorigenesis in different cancers (7), but its roles in HNSCC have been
reported in a few studies (8,9). A
previous study reported that EGF-induced ANGPTL4 plays a role in
the regulation and progression of cancer metastasis (8); however, its inhibitory roles for tumor
progression have not been fully elucidated. The findings of the
present study suggest an inhibitory association of ANGPTL4 with
tumor progression in OPSCC, which are supported by clinical
outcomes and complementary in vitro and TCGA-based analyses.
The present study revealed that higher ANGPTL4 expression (≥7.7%)
in patients with OPSCC was significantly associated with improved
OS and DFS. This finding suggested that ANGPTL4 could act as a
tumor suppressor in OPSCC. Several studies have previously reported
the correlation between the expression level of ANGPTL4 and
survival rates in various cancers. Similar results were reported
for gastric carcinoma (16),
hepatocellular carcinoma (17) and
triple-negative breast carcinoma (18). There have also been studies of no
correlation between the expression of ANGPTL4 and prognoses of
urothelial (19) and colorectal
(23) carcinomas. Some studies
reported that high ANGPTL4 expression was associated with poor
prognoses of cervical (14),
colorectal (10) and tongue
(13) carcinomas. These variations
suggest that the role of ANGPTL4 can differ with histology or tumor
origin. The current analysis using TCGA data indicated that the
expression and prognostic impact of ANGPTL4 differed across
the subsites of HNSCC, with poorer survival in tongue SCC, whereas
no significant association was observed in laryngeal SCC. These
subsite-specific patterns may reflect differences in etiologic
exposures, oncogenic drivers and tumor microenvironmental states
across anatomic locations, as genomic, transcriptional and
microenvironmental heterogeneity has been reported among
HPV-negative HNSCC arising from different subsites (24). In addition, ANGPTL4 is increasingly
recognized as a context-dependent secreted factor whose functions
can diverge based on the tumor type and microenvironment, and may
be proteolytically processed into different forms that may have
distinct biological functions (7,25).
Accordingly, the clinical significance of ANGPTL4 may not be
uniform across HNSCC and may require subsite-specific validation as
a prognostic biomarker. On the other hand, a plausible explanation
for the discrepancy with prior HNSCC studies (8,9) is
that ANGPTL4 may exert distinct functions depending on the anatomic
subsite and molecular context. In addition, HPV-positive OPSCC
represents a distinct subtype, and HPV status may modify the
clinical relevance of ANGPTL4 expression. Consistent with this, the
current stratified analyses by p16 status demonstrated that the
favorable prognosis of high ANGPTL4 expression was evident in the
p16-positive subgroup, whereas no significant association was
observed in the p16-negative subgroup. Notably, the TCGA-based GSEA
revealed that low ANGPTL4 expression was associated with enrichment
of cell cycle-related gene sets. These observations raise the
hypothesis that the prognostic relevance of ANGPTL4 may be more
pronounced in HPV-related tumors with deregulated checkpoints;
however, this interpretation remains hypothesis-generating and
warrants further validation in larger cohorts and mechanistic
studies. On the other hand, the results of the present study
demonstrated a favorable prognosis, which differed from those of
prior studies describing a promotive role of ANGPTL4 in HNSCC
(8,9). These studies primarily used
metastasis-related in vitro assays (for example, anoikis
resistance and invasion/migration) under specific stimuli, whereas
the present study focused on clinical prognosis in OPSCC and
proliferation-related phenotypes. Therefore, these
context-dependent differences may account for the apparent
discrepancy with prior studies.
The present study demonstrated that the
downregulation of ANGPTL4 significantly increased the expression
level of Ki-67 in immunofluorescence staining with the increase of
MKI67 mRNA expression and promoted cell proliferation in the
CellTiter-Glo 2.0 assay. These findings suggested that ANGPTL4 can
have an inhibitory role in cell proliferation in the HNSCC cell
line FaDu. Previous studies have reported that ANGPTL4 has both
promoting and inhibiting roles in cell proliferation in various
cancers. Ito et al (15)
reported that ANGPTL4 inhibited cell proliferation in vitro
and in vivo by regulating angiogenesis and vascular
leakiness. Ng et al (17)
reported several mechanisms of ANGPTL4 in suppressing tumor
progression, invasion and metastasis of HCC. Overexpression of
ANGPTL4 suppressed tumor growth by enhancing tumor cell apoptosis,
indicating that suppression of ANGPTL4 in HCC may be a way to
escape from apoptosis (17). In
addition, treatment with ANGPTL4-overexpressing adenovirus via
portal vein significantly suppressed the formation of new vessels
in the tumor by repressing the expression of vascular endothelial
growth factor and suppressing the activation of Raf-MEK-Erk
signaling pathway, suggesting an anti-angiogenic effect of ANGPTL4
on HCC. These in vitro findings were consistent with an
antiproliferative association of ANGPTL4. However, the current
functional assessments were limited to proliferation endpoints
(CellTiter-Glo 2.0 assay and Ki-67), and other malignant
phenotypes, including cell cycle progression, apoptosis, migration
and invasion, were not examined. Furthermore, these functional
observations were derived from siRNA-mediated knockdown without
complementary rescue or overexpression experiments; therefore,
off-target effects cannot be fully excluded. On the other hand,
RT-qPCR revealed altered expression of apoptosis-related genes
after ANGPTL4 knockdown in FaDu cells. However, because
apoptosis was not directly assessed using dedicated functional
assays, these RT-qPCR findings should be interpreted as
hypothesis-generating rather than as functional evidence of altered
apoptosis. In OPSCC, further studies are needed to clarify the
mechanism of the inhibitory role of ANGPTL4 for tumor cell
proliferation.
GSEA using TCGA data revealed that both the
HALLMARK_E2F_TARGETS and HALLMARK_G2M_CHECKPOINT pathways were
significantly enriched in the low ANGPTL4 group in OPSCC.
These gene sets represent crucial regulators of cell cycle
progression, particularly in the transition through the G1/S phase
(E2F targets) and the G2/M checkpoint (26,27).
High E2F activity increases the expression of genes involved in DNA
replication and cell cycle progression, directly promoting
proliferation (26).
HALLMARK_G2M_CHECKPOINT includes genes that control the G2/M
transition, ensuring cells only divide when DNA is correctly
replicated. Activation of this checkpoint signature reflects
increased cell cycle activity and is often observed in rapidly
proliferating tumor cells (27).
The present results of GSEA suggest that low ANGPTL4 expression is
associated with enrichment of cell cycle-related gene sets in
OPSCC. Because these transcriptomic findings are correlative, the
mechanistic interpretation should be considered
hypothesis-generating and will require dedicated validation,
including flow cytometric cell-cycle analysis and protein-level
assessment of key regulators. Furthermore, GSEA showed that
inflammation- and immune-related signatures such as
HALLMARK_TNFA_SIGNALING_VIA_NFKB,HALLMARK_INTERFERON_GAMMA_RESPONSE,
and HALLMARK_INFLAMMATORY_RESPONSE were strongly suppressed. It is
hypothesized that the decreased ANGPTL4 expression may be involved
in the suppression of inflammatory signaling and may have some
effect on the tumor microenvironment.
In the analysis of DEGs, CDH3 and
NDRG1 were significantly upregulated in the high
ANGPTL4 expression group. CDH3 is a classical cadherin that
plays a crucial role in maintaining epithelial cell-cell adhesion,
contributing to the integrity of epithelial structures in normal
tissues (28). In HNSCC, the high
CDH3 expression was associated with poor prognosis and an advanced
T stage (29,30). On the other hand, NDRG1 is
recognized as a tumor suppressor, which was reported to be
associated with epithelial-mesenchymal transformation (31,32). A
decrease in NDRG1 expression is associated with the promotion of
tumor proliferation (33). In oral
SCC, knockdown of NDRG1 using short hairpin RNA
significantly promoted cell proliferation, while overexpression of
NDRG1 caused cell cycle arrest at the S phase and suppressed
proliferation. This suggested that NDRG1 acts to inhibit cell
proliferation (33). In addition,
knockdown of NDRG1 enhanced cell proliferation, migration
and invasion in nasopharyngeal carcinoma and increased tumor
formation in mice (32). The
significant reduction of these two genes (NDRG1 and
CDH3) in the low ANGPTL4 expression group using TCGA
data suggests that the expression of these genes is closely related
to the impact of ANGPTL4 on tumor proliferation. To further analyze
the function of the ANGPTL4, a larger case study is
required.
The present study has certain limitations. First,
the IHC analysis was retrospective and conducted at a single
institution. Second, the cutoff value for ANGPTL4 positivity in IHC
was relatively low. The evaluation was performed by a single
evaluator, and formal reproducibility metrics (for example, Cohen's
kappa) were not assessed; therefore, scoring variability cannot be
excluded. Third, functional validation was limited to a single cell
line, which was not derived from OPSCC and may not fully
recapitulate OPSCC biology. In addition, in vivo experiments
were not performed; therefore, the present study was unable to
establish a causal role of ANGPTL4 in tumor growth. Further studies
are needed to elucidate the mechanisms by which ANGPTL4 regulates
cancer development.
The present study demonstrated significant
associations of ANGPTL4 expression with prognosis in patients with
OPSCC and showed that ANGPTL4 inhibited cell proliferation in the
HNSCC cell line FaDu. ANGPTL4 may serve as a prognostic biomarker
in OPSCC. Further in vivo studies are warranted to establish
causality and clarify its clinical relevance.
Supplementary Material
Supporting Data
Supporting Data
Acknowledgements
The authors would like to thank Dr Yuichi Ikari, Dr
Makoto Hosoya, Dr Shintaro Nakamura and Dr Chika Saegusa and Dr
Fuyuki Miya of Keio University School of Medicine (Tokyo, Japan)
for technical assistance.
Funding
The present study was supported by JSPS KAKENHI (grant no.
JP25K12772).
Availability of data and materials
The data generated in the present study may be
requested from the corresponding author.
Authors' contributions
TM conceptualized the study, curated data, conducted
formal analysis and investigation, developed methodology, performed
project administration, provided resources, visualized data, wrote
the original draft of the manuscript, and wrote, reviewed and
edited the manuscript. NN conceptualized the study, conducted
formal analysis and investigation, provided resources, and wrote,
reviewed and edited the manuscript. MS, SS, KY RN and TK
conceptualized the study, provided resources, and wrote, reviewed
and edited the manuscript. HO conceptualized the study, curated
data, conducted formal analysis, performed investigation, developed
methodology, conducted project administration, provided resources,
visualized data, supervised the study, wrote the original draft of
the manuscript, and wrote, reviewed and edited the manuscript. TM
and HO confirm the authenticity of all the raw data. All authors
read and approved the final version of the manuscript.
Ethics approval and consent to
participate
The present study adhered to the principles of the
Declaration of Helsinki and current ethical guidelines and was
approved (approval no. 20100013) by the Institutional Review Board
and Research Ethics Committee of Keio University School of Medicine
(Tokyo, Japan). Informed consent was obtained using the opt-out
option on the website and information provided at the hospital.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
Glossary
Abbreviations
Abbreviations:
|
ANGPTL4
|
angiopoietin-like 4
|
|
CI
|
confidence interval
|
|
DEG
|
differentially expressed gene
|
|
DFS
|
disease-free survival
|
|
FDR
|
false discovery rate
|
|
GSEA
|
gene set enrichment analysis
|
|
HNSCC
|
head and neck squamous cell
carcinoma
|
|
HPV
|
human papillomavirus
|
|
HR
|
hazard ratio
|
|
OPSCC
|
oropharyngeal squamous cell
carcinoma
|
|
OS
|
overall survival
|
|
PBS
|
phosphate-buffered saline
|
|
PCR
|
polymerase chain reaction
|
|
TCGA
|
The Cancer Genome Atlas
|
References
|
1
|
Siegel RL, Miller KD and Jemal A: Cancer
statistics, 2020. CA Cancer J Clin. 70:7–30. 2020.PubMed/NCBI
|
|
2
|
Carvalho AL, Nishimoto IN, Califano JA and
Kowalski LP: Trends in incidence and prognosis for head and neck
cancer in the United States: A site-specific analysis of the SEER
database. Int J Cancer. 114:806–816. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Janz TA, Graboyes EM, Nguyen SA, Ellis MA,
Neskey DM, Harruff EE and Lentsch EJ: A comparison of the NCDB and
SEER database for research involving head and neck cancer.
Otolaryngol Head Neck Surg. 160:284–294. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
NCCN Clinical Practice Guidelines in
Oncology (NCCN Guidelines®), . Head and neck cancers, version
2.2025. https://www.nccn.org/professionals/physician_gls/pdf/head-and-neck.pdfNovember
30–2025
|
|
5
|
Pich A, Chiusa L and Navone R: Prognostic
relevance of cell proliferation in head and neck tumors. Ann Oncol.
15:1319–1329. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Oike Y, Akao M, Kubota Y and Suda T:
Angiopoietin-like proteins: Potential new targets for metabolic
syndrome therapy. Trends Mol Med. 11:473–479. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Tan MJ, Teo Z, Sng MK, Zhu P and Tan NS:
Emerging roles of angiopoietin-like 4 in human cancer. Mol Cancer
Res. 10:677–688. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Liao YH, Chiang KH, Shieh JM, Huang CR,
Shen CJ, Huang WC and Chen BK: Epidermal growth factor-induced
ANGPTL4 enhances anoikis resistance and tumour metastasis in head
and neck squamous cell carcinoma. Oncogene. 36:2228–2242. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Chiang KH, Shieh JM, Shen CJ, Chang TW, Wu
PT, Hsu JY, Tsai JP, Chang WC and Chen BK: Epidermal growth
factor-induced COX-2 regulates metastasis of head and neck squamous
cell carcinoma through upregulation of angiopoietin-like 4. Cancer
Sci. 111:2004–2015. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Kim SH, Park YY, Kim SW, Lee JS, Wang D
and DuBois RN: ANGPTL4 induction by prostaglandin E2 under hypoxic
conditions promotes colorectal cancer progression. Cancer Res.
71:7010–7020. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Zhu X, Guo X, Wu S and Wei L: ANGPTL4
correlates with NSCLC progression and regulates
epithelial-mesenchymal transition via ERK pathway. Lung.
194:637–646. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Li X, Chen T, Shi Q, Li J, Cai S, Zhou P,
Zhong Y and Yao L: Angiopoietin-like 4 enhances metastasis and
inhibits apoptosis via inducing bone morphogenetic protein 7 in
colorectal cancer cells. Biochem Biophys Res Commun. 467:128–134.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Huang Z, Xie J, Lin S, Li S, Huang Z, Wang
Y and Ye J: The downregulation of ANGPTL4 inhibits the migration
and proliferation of tongue squamous cell carcinoma. Arch Oral
Biol. 71:144–149. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Nie D, Zheng Q, Liu L, Mao X and Li Z:
Up-regulated of angiopoietin-like protein 4 predicts poor prognosis
in cervical cancer. J Cancer. 10:1896–1901. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Ito Y, Oike Y, Yasunaga K, Hamada K,
Miyata K, Matsumoto S, Sugano S, Tanihara H, Masuho Y and Suda T:
Inhibition of angiogenesis and vascular leakiness by
angiopoietin-related protein 4. Cancer Res. 63:6651–6657.
2003.PubMed/NCBI
|
|
16
|
Kubo H, Kitajima Y, Kai K, Nakamura J,
Miyake S, Yanagihara K, Morito K, Tanaka T, Shida M and Noshiro H:
Regulation and clinical significance of the hypoxia-induced
expression of ANGPTL4 in gastric cancer. Oncol Lett. 11:1026–1034.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Ng KTP, Xu A, Cheng Q, Guo DY, Lim ZX, Sun
CK, Fung JH, Poon RT, Fan ST, Lo CM and Man K: Clinical relevance
and therapeutic potential of angiopoietin-like protein 4 in
hepatocellular carcinoma. Mol Cancer. 13:1962014. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Cai YC, Yang H, Wang KF, Chen TH, Jiang WQ
and Shi YX: ANGPTL4 overexpression inhibits tumor cell adhesion and
migration and predicts favorable prognosis of triple-negative
breast cancer. BMC Cancer. 20:8782020. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Hsieh HY, Jou YC, Tung CL, Tsai YS, Wang
YH, Chi CL, Lin RI, Hung SK, Chuang YM, Wu SF, et al: Epigenetic
silencing of the dual-role signal mediator, ANGPTL4 in tumor
tissues and its overexpression in the urothelial carcinoma
microenvironment. Oncogene. 37:673–686. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Edge SB, Byrd DR, Compton CC, Fritx AG,
Greene FL and Trotti A: AJCC cancer staging manual. 8th edition.
Springer; New York, NY: 2018
|
|
21
|
El-Naggar AK, JKC C, Grandis JR, Takata T,
Grandis J and Slootweg P: WHO classification of head and neck
tumours. 4th edition. IARC; Lyon: 2017
|
|
22
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Nakayama T, Hirakawa H, Shibata K, Nazneen
A, Abe K, Nagayasu T and Taguchi T: Expression of angiopoietin-like
4 (ANGPTL4) in human colorectal cancer: ANGPTL4 promotes venous
invasion and distant metastasis. Oncol Rep. 25:929–935. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Kim HAJ, Zeng PYF, Shaikh MH, Mundi N,
Ghasemi F, Di Gravio E, Khan H, MacNeil D, Khan MI, Patel K, et al:
All HPV-negative head and neck cancers are not the same: Analysis
of the TCGA dataset reveals that anatomical sites have distinct
mutation, transcriptome, hypoxia, and tumor microenvironment
profiles. Oral Oncol. 116:1052602021. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Adhikary T, Brandt DT, Kaddatz K, Stockert
J, Naruhn S, Meissner W, Finkernagel F, Obert J, Lieber S, Scharfe
M, et al: Inverse PPARβ/δ agonists suppress oncogenic signaling to
the ANGPTL4 gene and inhibit cancer cell invasion. Oncogene.
32:5241–5252. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Kent LN and Leone G: The broken cycle: E2F
dysfunction in cancer. Nat Rev Cancer. 19:326–338. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Oshi M, Takahashi H, Tokumaru Y, Yan L,
Rashid OM, Nagahashi M, Matsuyama R, Endo I and Takabe K: The E2F
pathway score as a predictive biomarker of response to neoadjuvant
therapy in ER+/HER2-breast cancer. Cells. 9:16432020. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Sun L, Hu H, Peng L, Zhou Z, Zhao X, Pan
J, Sun L, Yang Z and Ran Y: P-cadherin promotes liver metastasis
and is associated with poor prognosis in colon cancer. Am J Pathol.
179:380–390. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Liu L, Oh C, Lim MA, Zheng S, Piao Y, Ohm
S, Shan Y, Piao S, Shen S, Kim YI, et al: Dual blockage of
P-cadherin and c-Met synergistically inhibits the growth of head
and neck cancer. Cell Oncol (Dordr). 48:1019–1033. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Wang H, Yu T and Mao L:
Placental-Cadherin, a biomarker for local immune status and poor
prognosis among patients with tongue squamous cell carcinoma. Eur
Arch Otorhinolaryngol. 279:3597–3609. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Dong X, Hong Y, Sun H, Chen C, Zhao X and
Sun B: NDRG1 suppresses vasculogenic mimicry and tumor
aggressiveness in gastric carcinoma. Oncol Lett. 18:3003–3016.
2019.PubMed/NCBI
|
|
32
|
Hu ZY, Xie WB, Yang F, Xiao LW, Wang XY,
Chen SY and Li ZG: NDRG1 attenuates epithelial-mesenchymal
transition of nasopharyngeal cancer cells via blocking Smad2
signaling. Biochim Biophys Acta. 1852:1876–1886. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Lee JC, Chung LC, Chen YJ, Feng TH and
Juang HH: N-myc downstream-regulated gene 1 downregulates cell
proliferation, invasiveness, and tumorigenesis in human oral
squamous cell carcinoma. Cancer Lett. 355:242–252. 2014. View Article : Google Scholar : PubMed/NCBI
|