Introduction
Primary nephrotic syndrome (PNS) is among the most
common chronic kidney diseases affecting children, and the
magnitude and pattern of proteinuria remain central to diagnosis,
risk evaluation, treatment selection and monitoring of response
(1,2). As is known, 24-h urine protein
(24hUP) excretion has long been the gold standard for quantifying
protein loss and is a strong predictor of renal outcomes across
glomerular diseases, underscoring the clinical need for accurate
24hUP estimates in routine care, beyond diagnosis alone (1,2).
Current guidelines permit the diagnosis of nephrotic-range
proteinuria using either a urine protein-creatinine ratio (UPCR)
≥200 mg/mmol or a 24hUP ≥40 mg/m2/h (2,3).
However, these diagnostic cut-offs primarily establish disease
presence and do not provide the granularity required for decisions,
such as initiating thromboprophylaxis, triaging for inpatient
monitoring, or tailoring follow-up intensity during the acute phase
(3,4).
In pediatrics, 24-h urine collections are frequently
impractical due to poor cooperation, incomplete or contaminated
samples, and the logistical burden on families and health systems,
which compromises reliability and feasibility in routine practice
(5). A single-voided UPCR is
therefore widely recommended as an accessible surrogate,
physiologically normalizing protein to creatinine and approximating
24hUP (2,3). Although pediatric studies have
demonstrated a positive association between UPCR and 24hUP,
residual heterogeneity persists due to age- and sex-related
differences in creatinine generation, edema and hydration status,
sampling time, assay methodology and laboratory quality control
(6,7). Moreover, the association between UPCR
and 24hUP is often multiplicative and heteroscedastic, predisposing
to proportional bias at the lower and upper extremes on the raw
scale and limiting confidence in directly inferring very high
proteinuria from a single UPCR value, precisely when clinicians
seek to avoid missing high-risk presentations (8,9).
Population-specific kidney equations have recently been derived in
Vietnamese patients, underscoring that nephrology prediction tools
often require local calibration rather than direct adoption of
Western formulas (10).
Key evidence gaps include the lack of externally
validated UPCR-to-24hUP equations in pediatric PNS, the limited
assessment of clinical interchangeability beyond simple correlation
and the sparse representation of Asian cohorts. The present study
aimed to: i) Derive a pediatric UPCR-to-24hUP equation in PNS; ii)
externally validate its predictive performance, including
calibration and agreement; and iii) translate predictions into
clinically actionable gray-zone thresholds, using
prediction-interval (PI)-guided classification and decision-curve
analysis (DCA) to compare net clinical benefit with threshold-based
care.
Patients and methods
Study design and setting
The present study conducted a single-center
observational, method-comparison study with prospectively defined
derivation and validation cohorts. Consecutive pediatric patients
with PNS treated at the Department of Nephrology, Can Tho
Children's Hospital (Can Tho, Vietnam) between January, 2023 and
October, 2025 were included. The cohort split was time-based and
non-overlapping: Derivation (January, 2023 to January, 2025) and
validation (February, 2025 to October, 2025). No individual
contributed data to both cohorts. For the derivation cohort,
encounters from January 1, 2023 to December 17, 2024 constituted a
retrospective segment. No additional specimens were collected for
research; all laboratory results and clinical information were
routinely generated as part of standard care and were already
available within the same clinical site (Can Tho Children's
Hospital) in the medical records and the laboratory information
system. The study was approved by the Institutional Review Board
(IRB) of Can Tho University of Medicine and Pharmacy (IRB No.
4618/QĐ-ĐHYDCT/24.KY.60), approval date December 17, 2024 and
received permission to conduct the study at Can Tho Children's
Hospital. The retrospective component was granted a waiver of
informed consent on the basis of minimal risk, impracticability of
contacting all prior patients, and adequate privacy safeguards
(coded identifiers, restricted access). The prospective component
obtained written informed consent from a parent/legal guardian and
age-appropriate child assent when applicable. Following IRB
approval on December 17, 2024, study-specific data
abstraction/extraction for the retrospective segment began on
December 18, 2024, and data verification/cleaning continued until
March 30, 2025. Data from December 18, 2024 to January 31, 2025
represented the prospective segment of the derivation cohort and
were collected according to the IRB-approved protocol.
Patient selection
Children aged 1-16 years with PNS, defined per
contemporary pediatric guidance (2,3),
were eligible if, within 48 h of admission, a complete 24-h urine
collection was available to quantify 24hUP and 24-hour urine
creatinine (24hUCr), and a spot urine sample from the same episode
was obtained for UPCR. Patients were excluded from the study for
the following reasons: i) Secondary nephrotic syndrome; ii)
decreased renal function with estimated glomerular filtration rate
(eGFR) <60 ml/min/1.73 m2; iii) known primary muscle
disease or medications expected to materially alter creatinine
generation; iv) inadequate 24-h collection, predefined as 24hUCr
per body weight below the site-specific threshold (11); and (v) missing core variables after
attempted retrieval.
The derivation cohort was sized using Fisher's z to
target a 95% CI half-width of 0.10 around an expected log-scale
correlation r0=0.80, yielding n=60
(conservatively set to 85). For external validation, root mean
square error (RMSE) precision was based on the Chi-square
distribution of residual variance; the 95% relative half-width is
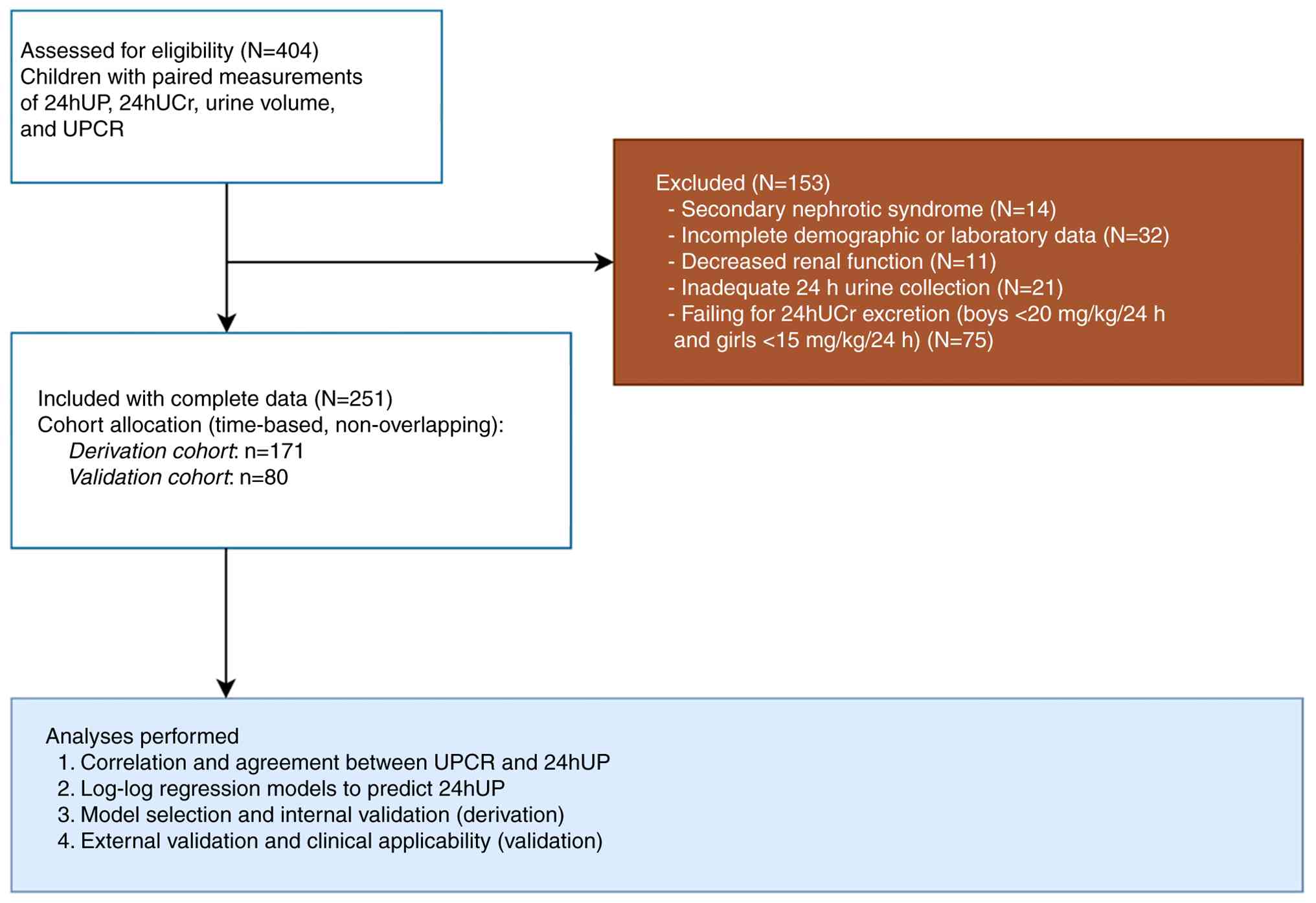
approximately z0.975 . Finally, 171 participants were
included in the derivation cohort to develop the model and perform
internal validation, whereas 80 children comprised the validation
cohort for external validation to assess clinical applicability
(Fig. 1).
Specimen collection and laboratory
measurements
Caregivers received standardized, illustrated
instructions and in-person training. After discarding the first
morning void, all urine passed over the subsequent 24 h was
collected, including the terminal void at 24 h. Immediately upon
completing the 24-h collection, an early-morning spot urine sample
was obtained from the same participant. All specimens were
delivered to the laboratory within 2 h of completion. At sampling,
we recorded age, sex, height/weight, serum albumin, serum
creatinine and current corticosteroid or antiproteinuric therapy.
Body weight was recorded as dry weight, defined as the baseline
(pre-edema) weight of the child when free of clinically apparent
edema or ascites. In the event that dry weight was unavailable, a
height-based expected weight derived from World Health Organization
(WHO) weight-for-height standards was used as a surrogate.
Total urine protein was measured by the pyrogallol
red-molybdate method on a cobas c501 analyzer (Roche Diagnostics)
using the Roche urine/CSF total protein calibrator (albumin-based),
traceable to the manufacturer's reference materials. Urine
creatinine was measured by the enzymatic (creatinase-sarcosine
oxidase) method on the same analyzer and standardized to the IDMS
reference measurement procedure. Both spot UPCR and the 24-h
reference specimens were analyzed on the same platform to ensure
methodological consistency.
Calculations. 24hUCr was estimated using three
published equations: Hellerstein (12), Ghazali-Barratt (13), and Cockcroft-Gault (14). Age is represented in years and
weight in kg; units are mg/24 h.
i) Hellerstein: For girls and boys <14 years:
UCrHellerstein= [0.17x(age)+17.1]xweight; for boys ≥14
years: UCrHellerstein= [0.39x(age)+14.9]xweight.
ii) Ghazali-Barratt:
UCrGhazali-Barratt=[0.46x(age)+15.4]x weight.
iii) Cockcroft-Gault: For boys:
UCrCockcroft-Gault=[28-(0.2x age)]xweight; for girls:
UCrCockcroft-Gault=[28-(0.2xage)]xweightx0.85.
Statistical analysis. Data
handling
Continuous variables are summarized as the mean and
standard deviation (SD) or median and interquartile range (IQR);
categorical variables are presented as frequencies (percentages).
Variables exhibiting right skew (including UPCR and 24hUP) were
analyzed on the log10-scale, with back-transformation to
the original scale where appropriate. All statistical tests were
two-sided with α=0.05; a P-value <0.05 was considered to
indicate a statistically significant difference.
Correlation and agreement. Spearman's ρ was
performed for correlations between 24hUP and: i) Measured UPCR
(mUPCR); ii) estimated UPCR (eUPCR) defined as mUPCR x 24hUCr,
using measured (m24hUCr; eUPCR0) or estimated (e24hUCr)
from Hellerstein (eUPCR1), Ghazali-Barratt
(eUPCR2) and Cockcroft-Gault (eUPCR3)
equations. A Bland-Altman analysis was performed on the
log10-scale, reporting the geometric mean ratio (bias)
and 95% limits of agreement (LoA); proportional bias was tested by
regressing log-difference on log-mean. P-values for correlations
were obtained from the two-sided Spearman's rank correlation
test.
Model development. Log-log linear regressions
were fit to predict log10 (24hUP):
M1=β0+β1 xlog10(mUPCR)+ε;
M2=β0+β1
xlog10(eUPCR0)+ε;
M3-M5=β0+β1
xlog10(eUPCR1-3)+ε, using three published
e24hUCr formulas: Hellerstein (12), Ghazali-Barratt (13) and Cockcroft-Gault (14). Model selection used 10-fold
cross-validation (CV), prioritizing CV-RMSE/MAE (adj. R², AIC, BIC
secondary). Standard diagnostics were applied (linearity,
residuals, influence, multicollinearity); residual normality,
heteroskedasticity and model specification were assessed using
standard tests (e.g., Shapiro-Wilk/skewness-kurtosis;
Breusch-Pagan/White/IM; Ramsey RESET; and linktest), with two-sided
P-values reported for the corresponding diagnostics. HC3 robust
standard errors were reported if heteroskedasticity was detected.
P-values for regression coefficients were obtained from two-sided
Wald t-tests using the corresponding standard errors (HC3 when
applied).
Internal validation
Bootstrap resampling (B=2000) was performed to
obtain optimism-corrected RMSE/MAE/R2 and calibration
(slope/intercept). Predictions were returned to the original scale
using Duan's smearing estimator (smearing factor reported).
External validation
Coefficients from the best model were applied
unchanged to the validation cohort. We reported RMSE, MAE,
R2 (squared Pearson correlation between observed and
predicted 24hUP), and P20/P30 (percentage of predictions within
20/30% of observed 24hUP). Calibration was evaluated by intercept,
slope and a calibration plot. For clinical interpretation, we
generated a patient-level PI for each predicted 24hUP and applied
prespecified cut-points relevant to acute pediatric PNS (100
mg/m2/h as a lower-risk threshold and 150
mg/m2/h as a higher-risk threshold). A case was labeled
‘ruled out’ if the upper PI bound was below the lower-risk
threshold, ‘ruled in’ if the lower PI bound exceeded the
higher-risk threshold, and ‘gray zone’ otherwise. Clinical utility
was then evaluated using DCA. We compared the net benefit of the
model-based classification with treat-all and treat-none
strategies, as well as with fixed guideline-style cut-offs for
nephrotic-range proteinuria, across threshold probability ranges
chosen a priori to reflect plausible willingness to intervene in
suspected very high proteinuria.
Statistical software
Analyses were performed in R (version 4.5.1, R
Foundation for Statistical Computing, Vienna, Austria).
Results
Study population characteristics
A total of 251 children were included in the present
study: A total of 171 children in the derivation cohort for model
development and 80 children in the validation cohort for external
assessment of performance and applicability. The demographic and
clinical baseline characteristics of the children are summarized in
Table I. Overall, the two cohorts
were generally comparable at baseline.
 | Table IBaseline characteristics of the
derivation and validation cohorts. |
Table I
Baseline characteristics of the
derivation and validation cohorts.
| Characteristic | Derivation
cohort(n=171) | Validation cohort
(n=80) |
|---|
| Age (mean ± SD),
years | 9.33±3.70 | 10.02±4.37 |
| Male, n (%) | 122 (71.35) | 49 (61.25) |
| Body weight (mean ±
SD), kg | 32.17±11.19 | 33.57±15.34 |
| Height (mean ± SD),
cm | 128.08±20.12 | 132.55±25.53 |
| BSA (mean ± SD),
m2 | 1.06±0.26 | 1.10±0.35 |
| BUN (median, IQR),
mg/dl | 12.88 (10.08,
16.24) | 13.33 (10.86,
17.78) |
| SCr (median, IQR),
mg/dl | 0.57 (0.48,
0.73) | 0.65 (0.52,
0.83) |
| eGFR (median, IQR),
ml/min/1.73 m2 | 93.97 (72.71,
111.62) | 83.41 (70.02,
100.13) |
| Albumin (mean ±
SD), g/l | 18.89±4.59 | 20.32±3.45 |
| Spot UPCR (median,
IQR), mg/mmol | 410.84 (287.09,
613.28) | 396.74 (281.61,
471.62) |
| 24hUP (median,
IQR), mg/m2/h | 116.81 (87.81,
160.47) | 107.06 (81.1,
133.99) |
| 24hUCr (median,
IQR), mg/24 h | 934.03 (679.36,
1159.39) | 875.49 (563.55,
1119.58) |
Correlation and agreement between
24hUP and UPCR variants in the derivation cohort
Spearman's correlation analyses demonstrated a
strong correlation between mUPCR and 24hUP (ρ=0.81, P<0.001).
Adjusting by measured m24hUCr (eUPCR0) markedly improved
the correlation (ρ=0.92). Using e24hUCr, the correlation increased
with the Hellerstein equation (eUPCR1, ρ=0.83), was
essentially unchanged with Ghazali-Barratt (eUPCR2,
ρ=0.80), and improved modestly with Cockcroft-Gault
(eUPCR3, ρ=0.88) (Fig.
S1).
Bland-Altman plots on the log10-scale
confirmed that 95% of observations lay within the LoA for all
UPCR-derived indices (Fig. S2).
Unadjusted spot UPCR substantially overestimated 24hUP [geometric
mean ratio (GMR), 3.63; Fig.
S2A]. Adjusting UPCR by m24hUCr (eUPCR0) markedly
reduced bias (GMR, 1.15; Fig.
S2B), and the Cockcroft-Gault-based adjusted UPCR
(eUPCR3) exhibited the optimal agreement overall (GMR,
0.98 with tight ratio-scale LoA; Fig.
S2E).
Development of a 24hUP prediction
model
The present study developed five
log10-linear regression models to predict 24hUP from
UPCR (denoted M1-M5). M2 yielded
the optimal performance (CV-RMSE, 21.84; CV-MAE, 16.67; Adj.
R2 0.84; AIC, -367.64; BIC, -358.21; Table SI). However, M2
requires measured 24hUCr urine, which is impractical in children
due to continuous collection, volume errors and poor adherence.
Given clinical deployability as a key criterion, M2 was
not selected. M5 (eUPCR3 via Cockcroft-Gault)
provided the optimal accuracy-feasibility balance. Adding age and
sex to M5 to form M6 further improved fit vs.
M5 alone: CV-RMSE 28.14 vs. 29.15 (Δ=-1.01; 95% CI,
-1.42 to -0.78; P<0.0001) and CV-MAE 20.25 vs. 21.22 (Δ=-0.97;
95% CI: -1.19 to -0.68; P<0.0001); adjusted R2
increased from 0.78 to 0.80, and AIC/BIC decreased.
Optimal prediction equation
(M6)
The association between 24hUP and UPCR is presented
as: Log10[24hUP (mg/m2/h)]=0.291+0.880 x
Log10(adjusted UPCR) - 0.006 x age (years) + 0.057 x sex
(0=boy, 1=girl). Adjusted UPCR=mUPCR x
e24hUCrCockcroft-Gault; when applying the model, units
are harmonized so the product is expressed in mg/mm2/h
(Table II). Residuals were
approximately normal, with only borderline heteroskedasticity
(addressed using robust standard errors). Model diagnostics (RESET,
link test, low VIFs), sensitivity analyses excluding high-influence
observations or using robust regression (Table SII and Fig. S3), and component-plus-residual
plots (Fig. S4) all supported a
stable, approximately linear specification for M6
without meaningful multicollinearity or functional-form
misspecification.
 | Table IIMultivariable linear regression
analysis for Log10-transformed 24hUP excretion (final
model, M6). |
Table II
Multivariable linear regression
analysis for Log10-transformed 24hUP excretion (final
model, M6).
| | | 95% CI | |
|---|
| Predictor | β | SE (HC3) | Lower | Upper | P-value |
|---|
|
Log10(adjusted UPCR) | 0.880 | 0.036 | 0.810 | 0.951 | <0.001 |
| Age | -0.006 | 0.001 | -0.009 | -0.002 | <0.001 |
| Sex | 0.057 | 0.017 | 0.024 | 0.090 | <0.001 |
| Intercept | 0.291 | 0.076 | 0.142 | 0.440 | <0.005 |
Bootstrap optimism-corrected
performance
Internal validation with 2,000 bootstrap resamples
provided optimism-corrected estimates of performance and
calibration. Following optimism correction, prediction error on the
log10-scale remained low: RMSE increased by only 2.3%
(95% CI, 2.0 to 2.6%), MAE by 2.2% (95% CI, 2.0 to 2.4%), and
R2 decreased minimally from 0.803 to 0.796 (-0.785
percentage points; 95% CI for R2, 0.758 to 0.857).
Calibration was near-ideal (slope 0.999, intercept 0.002; Fig. 2), indicating negligible overfitting
and no need for coefficient shrinkage.
External validation of 24hUP
prediction in validation cohort
For clinical implementation, a back-transformed
model M6 from the Log10-scale used Duan's
smearing. Using the smearing factor (SF) estimated in the
derivation cohort (SF=1.022). With k=10β0=1.955, the
overall constant on the original scale is C=SF x k=1.998. The
prediction equation on the original scale is: 24hUP
(mg/m2/h)=1.998x(adjusted
UPCR)0.880x10(-0.006) agex100.057
sex. Applied unaltered to the independent validation cohort
(n=80), the model achieved an RMSE of 11.39 mg/m2/h (95%
CI, 9.92 to 12.83), an MAE of 9.26 mg/m2/h (7.87 to
10.78), and an R2 of 0.912 (95% CI, 0.866 to 0.943). P20
was 93.8% (86.0 to 97.9) and P30 was 96.3% (89.4 to 99.2),
indicating that the majority of predictions were within 20-30% of
observed 24hUP.
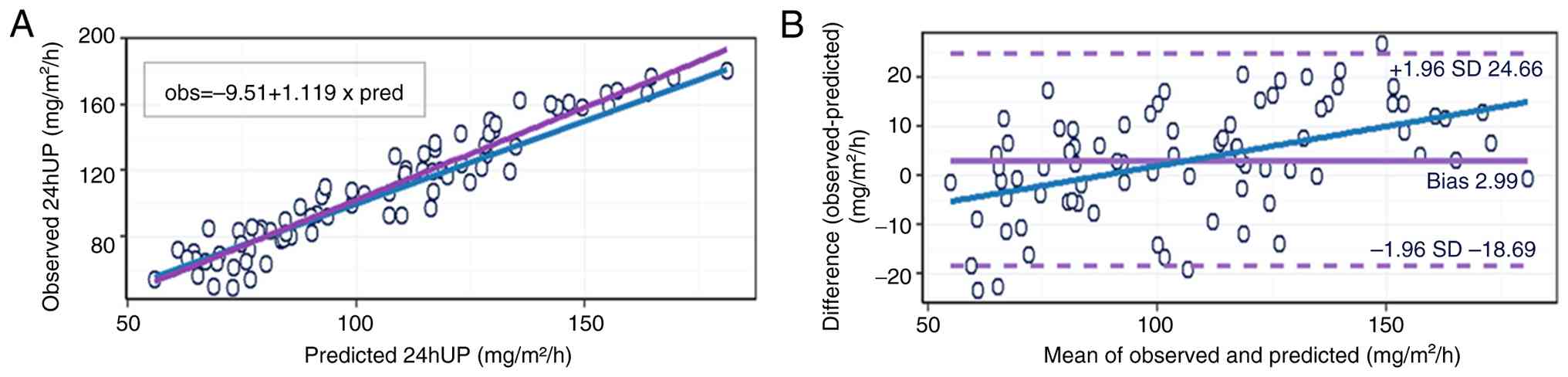
The calibration of observed vs. predicted 24hUP in
the validation cohort revealed a calibration intercept of -9.51
mg/m2/h (HC3 95% CI, -17.55 to -1.47) and slope of 1.12
(HC3 95% CI, 1.05 to 1.19), indicating mild compression of
predictions (slight overestimation at lower values and
underestimation at higher values) but overall acceptable
calibration (Fig. 3A).
Bland-Altman analysis on the original scale showed a mean bias of
2.99 mg/m2/h (observed-predicted), with LoA from -18.69
to 24.66 mg/m2/h, and evidence of proportional bias
(β=0.161; 95% CI, 0.094 to 0.229), meaning error increased at very
high proteinuria (Fig. 3B).
Using the prespecified PI three-zone rule on the
external validation cohort, the primary threshold at 150
mg/m2/h allowed omission of 75% of 24-h collections with
0% false-omits (PI-95%); 23.8% fell into the gray zone and 1.3%
were definite collections. At 100 mg/m2/h, the rule
omitted 23.8% and triggered 30% definite collections (PI-95%) with
0% false-omits (Table III).
Consistent with these findings, DCA showed positive net benefit
over treat-all and treat-none across clinically plausible threshold
probabilities (Fig. S5).
 | Table IIIClinical impact of a PI-based
three-zone rule using original M6 predictions (external validation,
n=80). |
Table III
Clinical impact of a PI-based
three-zone rule using original M6 predictions (external validation,
n=80).
| Threshold T
(mg/m2/h) | PI | No need 24hUP (n,
%) | Measure 24hUP (n,
%) | Gray zone (n,
%) | False-omit (n,
%) | False-do (n,
%) |
|---|
| 100 | 95% | 19 (23.8) | 24 (30.0) | 37 (46.2) | 0 (0.0) | 0 (0.0) |
| 150 | 95% | 60 (75.0) | 1 (1.3) | 19 (23.8) | 0 (0.0) | 0 (0.0) |
| 100 | 80% | 27 (33.8) | 33 (41.2) | 20 (25.0) | 0 (0.0) | 1 (1.3) |
| 150 | 80% | 68 (85.0) | 3 (3.8) | 9 (11.2) | 1 (1.3) | 0 (0.0) |
Discussion
Severe PNS is critical clinically not only due to
massive edema and profound hypoalbuminemia, but also due to early,
potentially life-threatening complications such as thromboembolism,
severe bacterial infection/sepsis and intravascular volume
depletion with prerenal acute kidney injury, all of which cluster
in the high-proteinuria, hypoalbuminemia phase and therefore track
with the severity of proteinuria (15-18).
Yet, there is currently no widely adopted pediatric tool that turns
a single-void UPCR into a reliable estimate of severity tier of
24hUP to support those specific bedside decisions. UPCR correlates
well with 24hUP in children; however, key gaps persist. The
majority of studies stop at simple correlations and do not assess
agreement, proportional bias, or calibration across the full
proteinuria range (6,19,20).
Numerous equations also lack external validation in independent
pediatric nephrotic cohorts, particularly in Asian children,
limiting use where 24-h urine collection is impractical. In
addition, the UPCR-24hUP association is multiplicative (log-log),
causing heteroscedastic and proportional error on the raw scale,
with underestimation at the highest protein leaks, exactly where
clinicians most need to identify children at highest risk (8,20,21).
In the present study cohort of children with PNS, a
multivariable equation was developed that converts a single-void
UPCR into an estimated 24-h protein excretion. Herein, three
creatinine-excretion approaches (Cockcroft-Gault, Hellerstein, and
Ghazali-Barratt) were compared to standardize UPCR for body-size
variability. The Cockcroft-Gault-based approach provided the
optimal overall agreement and the most favorable
accuracy-feasibility balance in the cohort; alternative pediatric
formulas did not improve performance and were therefore not
retained. The final model (M6) used a
Cockcroft-Gault-based adjusted UPCR multiplied by an age and sex
correction, reflecting physiologic variation in creatinine
generation, muscle mass and urine concentration across childhood
and adolescence, consistent with prior studies (22-25).
This approach is motivated by the known limitations of ‘raw’ UPCR:
Protein and creatinine do not scale identically with growth, edema,
hydration status and lean body mass, which introduces
heteroscedastic error and proportional bias, particularly at the
extremes of protein loss (8,21,26).
This adjustment also aligns with the broader goal of
reducing body-size-related variability when interpreting
proteinuria metrics in children. The fractional excretion of total
protein (FETP), the ratio of total protein clearance to creatinine
clearance, has been proposed as a physiologically grounded,
clearance-based index to account for body size with preserved
kidney function (27). However,
FETP typically requires synchronous blood measurements in addition
to urine protein and urine creatinine, which were not consistently
available at the time of urine sampling in the dataset in the
present study; thus, calculating FETP would have introduced
substantial missingness and potential selection bias. By contrast,
UPCR is emphasized in the majority of pediatric nephrology guides
as a practical alternative to timed urine collections for defining
and monitoring nephrotic-range proteinuria, supporting our focus on
UPCR for real-world, guideline-concordant implementation (1-3).
Future studies with standardized paired blood-urine sampling are
warranted to directly compare UPCR-based equations with
clearance-based indices such as FETP to assess any incremental
predictive or clinical utility.
The model used herein explained a large proportion
of variability in 24hUP during development and performed robustly
in a temporally independent validation cohort without refitting. In
external validation, it achieved a high coefficient of
determination (R2=0.91), low absolute prediction error
(9 mg/m2/h) and clinically tight accuracy (94% within 20
and 96% within 30% of observed 24hUP). Calibration remained
acceptable when the original coefficients were applied unaltered,
and the average bias on the original mg/m2/h scale was
minimal. These findings suggest that a single-voided urine specimen
can recover the approximate 24-h protein excretion of a child with
clinically usable precision, not merely correlation.
The present study also examined how the model
behaves at the most critical areas clinically: At the extreme upper
end of proteinuria (24,28). Bland-Altman analysis in the
validation cohort revealed narrow LoA and only modest mean bias
overall, but with proportional error at very high 24hUP values;
specifically, the model tended to slightly underpredict the most
extreme protein losses. This matters clinically as children with
extremely high-grade proteinuria have the highest short-term risk
for complications, such as intravascular volume depletion, severe
hypoalbuminemia requiring infusion, bacterial infection and
thromboembolism in the setting of marked urinary protein wasting
and hemoconcentration (15,16,18).
These are the very patients for whom pediatric nephrologists decide
on immediate inpatient monitoring and aggressive supportive care.
As proportional bias emerges at the extreme end of proteinuria, it
may be unsafe to base high-stakes decisions on a single predicted
value alone. Accordingly, when predicted values fall in the highest
range, confirmatory testing with a timed urine collection (full
24-h urine) should be considered to support clinical
decision-making. In such children, escalation decisions should
incorporate the uncertainty of the model, not only its mean
prediction.
To address this issue, in the present study, model
output was operationalized into a three-zone decision rule based on
the PI of each child, rather than relying solely on the point
estimate. Of note, two clinically relevant thresholds were
prespecified for 24hUP (100 mg/m2/h as a lower-risk
cut-point, and 150 mg/m2/h as a higher-risk cut-point)
consistent with levels at which clinicians begin to escalate
surveillance, consider albumin/diuretic management, or are
concerned about the thrombotic risk (1-3,29). For each patient, the
following questions were raised: Even in a worst-plausible scenario
consistent with model error (95% PI), is this child clearly below
the cut-point, clearly above it, or uncertain (‘gray zone’)? Using
this PI-based strategy at the 150 mg/m2/h threshold, a
full 24-h urine collection could have been avoided in 75% of
children in the validation cohort, while directing only 1% straight
to ‘must collect now’, and leaving 24% in a gray zone for clinician
review, without falsely ruling out any truly very-high-proteinuria
cases (0% false-omits). At the 100 mg/m2/h threshold,
the same framework again separated ‘safe to defer collection’,
‘must collect’, and ‘gray zone’ with no missed high-proteinuria
cases. In practice, gray-zone results (i.e., the PI overlaps the
cut-point) are most effectively handled with an iterative
confirmation strategy rather than immediate escalation. The authors
suggest the following: i) Repeating a spot UPCR after stabilization
(e.g., next void or within 12-24 h) to reduce random variability;
ii) integrating contemporaneous clinical context (edema burden,
hemodynamics, urine output) and key laboratory tests (serum albumin
and creatinine/eGFR); and iii) obtaining confirmatory timed urine
collection (full 24-h urine) when uncertainty persists or
management decisions are high-stakes. Conceptually, this functions
as a triage layer on top of the numeric prediction: It concentrates
limited nursing and laboratory effort on the few children whose
proteinuria is most concerning, and it reassures clinicians that
avoiding immediate 24-h collection in the others is unlikely to
miss a dangerous presentation.
This approach is responsive to how pediatric
nephrotic syndrome is actually managed. A tool that i) estimates
the likely magnitude of ongoing protein loss from a single spot
sample and ii) classifies the child into ‘clearly below’, ‘clearly
above’, or ‘uncertain relative to clinically relevant danger
zones’, directly mirrors bedside thinking. DCA in the external
validation cohort supports this interpretation: Across plausible
intervention thresholds (reflecting the tolerance of the clinician
to act on suspected very high proteinuria), a model-guided strategy
provided higher net clinical benefit than treating all children as
high risk (i.e. obtaining 24-h collections on everyone) or treating
none as high-risk. This indicates that the model is not only
statistically well calibrated, but also improves how efficiently
attention and resources would be allocated. This ‘equation → triage
rule → net benefit’ chain is largely absent from prior pediatric
work in nephrotic syndrome and, to the best of our knowledge, has
not yet been reported in an Asian pediatric cohort.
The present study has several strengths. It uses
consecutive real-world pediatric nephrotic admissions, including
children with substantial hypoalbuminemia and heavy protein loss,
rather than an artificially ‘clean’ subset. The validation cohort
was acquired later in time, reducing optimism from overfitting. The
present study reports discrimination-like metrics (R2,
P20/P30), absolute error (MAE, RMSE), calibration
(slope/intercept), clinical agreement on the original scale and
decision safety at actionable thresholds. This prediction model
study is reported in accordance with the TRIPOD statement (30). Of note, the PI-based gray-zone rule
achieved 0% false-omits for very high proteinuria at 150
mg/m2/h, while suggesting that the majority of children
could safely avoid a 24-h urine collection. That directly targets
the operational bottleneck in pediatric nephrotic care: Deciding
who truly needs an immediate timed collection as opposed to who can
be managed using spot data plus close observation.
However, certain limitations of the present study
should also be acknowledged. First, although the temporal split
provides external validation, both cohorts were drawn from a single
center. Accordingly, external validity is limited, and the equation
should be externally validated in independent, multicenter and
multi-ethnic pediatric cohorts prior to widespread clinical
adoption. Variation in anthropometry, nutritional status, steroid
exposure, supportive care practices (e.g., albumin infusion
thresholds, thromboprophylaxis norms), and laboratory calibration
across institutions may affect transportability (31,32).
Local recalibration, particularly of the smearing factor and
potentially the intercept, may be required before routine
deployment elsewhere. Second, the most extreme levels of
proteinuria were relatively uncommon; proportional underestimation
at the very high end means clinicians should interpret predictions
conservatively in profoundly edematous, hemodynamically tenuous, or
thrombosis-suspect presentations. Third, the prespecified
thresholds (100 and 150 mg/m2/h) and the resulting
‘rule-out/gray zone/rule-in’ logic are clinically reasoned, but not
yet guideline-endorsed. They should be viewed as an implementation
proposal rather than a definitive standard. Fourth, the primary
endpoint was proteinuria itself, not downstream hard outcomes, such
as thromboembolism, albumin-requiring hypovolemia, pediatric
intensive care unit transfer, or readmission. Linking the model and
the gray-zone rule to those outcomes will be essential to determine
whether triage truly improves safety and resource use. Finally,
this model is intended for children with preserved renal function
(eGFR, ≥60 ml/min/1.73 m2); performance in renal
impairment requires dedicated validation. Although the equation
uses spot UPCR as input, it was validated against measured 24-h
protein excretion in children able to complete timed collections,
and transportability to outpatient settings or to children unable
to complete 24-h urine remains to be established. Future studies
are thus required to focus on multicenter validation, recalibration
in other populations and outcome-linked impact evaluation. In
conclusion, if confirmed, this approach could reduce the burden of
timed urine collection in hospitalized children, while preserving
attention for those at greatest risk.
Supplementary Material
Correlations between 24hUP and UPCR in
the derivation cohort. (A) 24hUP vs. mUPCR; (B) 24hUP vs. UPCR
adjusted by m24hUCr (eUPCR0=mUPCR x m24hUCr); (C-E)
24hUP vs. UPCR adjusted by e24hUCr using the Hellerstein
(eUPCR1), Ghazali-Barratt (eUPCR2) and
Cockcroft-Gault (eUPCR3) equations. UPCR, urine
protein-creatinine ratio; 24hUP, 24-h urine protein; 24hUCr, 24-h
urine creatinine; m, measured; e, estimated.
Bland-Altman agreements between 24hUP
and UPCR in the derivation cohort. (A) 24hUP vs. mUPCR; (B) 24hUP
vs. UPCR adjusted by m24hUCr (eUPCR0=mUPCR x m24hUCr);
(C-E) 24hUP vs. UPCR adjusted by e24hUCr using the Hellerstein
(eUPCR1), Ghazali-Barratt (eUPCR2), and
Cockcroft-Gault (eUPCR3) equations. All values are
log10-transformed. UPCR, urine protein-creatinine ratio;
24hUP, 24-h urine protein; 24hUCr, 24-h urine creatinine; m,
measured; e, estimated.
Residual diagnostics for model
M6 (Log10-scale). (A) Residuals vs. fitted
(log10-scale). Points scatter around 0; the LOESS curve
is essentially flat with slight curvature near the center,
indicating no obvious model misspecification and small residual
amplitude. (B) Normal Q-Q (standardized residuals). Points lie
close to the 45˚ line with only mild tail departures, approximately
normal. For conservative inference, HC3 robust standard errors are
used. (C) Scale-location. decreases slightly and then stabilizes
across fitted values, suggesting roughly constant variance; no
clear ‘fan’ pattern, if heteroskedasticity exists, it is
borderline. (D) Residuals vs. leverage (hat values). Most
observations have leverage <2 k/n=0.047; a few exhibit higher
leverage (~0.06) but small residuals, indicating no unduly
influential observations. LOESS, locally weighted smoothing; SE
(HC3), heteroskedasticity-consistent standard errors (type
HC3).
Component-plus-Residual (partial
residual) plots for model M6. (A)
Log10(adjusted UPCR): Linear pattern, LOESS overlaps the
fitted line. (B) Age: Mild downward linear trend, no systematic
curvature. (C) Sex: Upward shift for girls and no evidence of
sex-specific heteroskedasticity. LOESS, locally weighted smoothing;
UPCR, urine protein/creatinine ratio; 24hUP, 24-h urine
protein.
Decision curve analysis for the
original M6 (external validation). (A) Threshold T=100
mg/m2/h; (B) Threshold T=150 mg/m2/h. Events
were defined as 24hUP ≥ T, with predicted risk derived from the
original M6 model. Curves display net benefit per
patient of the model, compared with treat-all and treat-none
strategies, across a range of risk thresholds (pt=0.05-0.80). The
model consistently provided higher net benefit over both reference
strategies across clinically relevant thresholds, supporting its
potential clinical utility for guiding 24h urine collection
decisions. DCA, decision-curve analysis; pt, risk threshold; 24hUP,
24-h urine protein.
Performance of log-log linear
models (M1-M6) predicting 24hUP from UPCR.
Diagnostics for model M6
(Log10-scale).
Acknowledgements
The authors would like to express their sincere
appreciation to Can Tho University of Medicine and Pharmacy (Can
Tho, Vietnam) for institutional support, and to the physicians and
nursing staff of Can Tho Children's Hospital (Can Tho, Vietnam) for
their essential assistance with patient recruitment, urine
collection and data acquisition.
Funding
Funding: No funding was received.
Availability of data and materials
The data generated in the present study may be
requested from the corresponding author.
Authors' contributions
QCN and HDT were responsible for the
conceptualization of the study. QCN, UHAN, TTH, LTKD and PHPN
performed data curation. QCN and HDT developed the study
methodology. Formal statistical analyses were conducted by QCN,
HDT, UHAN, TTH, LTKD and PHPN. The original draft of the manuscript
was written by QCN, TTH, LTKD and PHPN. QCN, HDT, UHAN, TTH, LTKD
and PHPN reviewed and edited the manuscript. HDT and QCN supervised
the study. QCN and UHAN confirm the authenticity of all raw data.
All authors have read and approved the final manuscript and agree
to be accountable for all aspects of the work.
Ethics approval and consent to
participate
The study was approved by the Institutional Review
Board (IRB) of Can Tho University of Medicine and Pharmacy (IRB No.
4618/QĐ-ĐHYDCT/24.KY.60), approval date, December 17, 2024 and
received permission to conduct the study at Can Tho Children's
Hospital. All procedures adhered to the Declaration of Helsinki of
1975, as revised in 2000. The retrospective component was granted a
waiver of informed consent on the basis of minimal risk,
impracticability of contacting all prior patients, and adequate
privacy safeguards (coded identifiers, restricted access). The
prospective component obtained written informed consent from a
parent/legal guardian and age-appropriate child assent when
applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
Use of artificial intelligence tools
During the preparation of this work, AI tools were
used to improve the readability and language of the manuscript or
to generate images, and subsequently, the authors revised and
edited the content produced by the AI tools as necessary, taking
full responsibility for the ultimate content of the present
manuscript.
References
|
1
|
Kidney Disease: Improving Global Outcomes
(KDIGO) Glomerular Diseases Work Group: KDIGO 2021 clinical
practice guideline for the management of glomerular diseases.
Kidney Int. 100:S1–S276. 2021.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Trautmann A, Boyer O, Hodson E, Bagga A,
Gipson DS, Samuel S, Wetzels J, Alhasan K, Banerjee S, Bhimma R, et
al: IPNA clinical practice recommendations for the diagnosis and
management of children with steroid-sensitive nephrotic syndrome.
Pediatr Nephrol. 38:877–919. 2023.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Floege J, Gibson KL, Vivarelli M, Liew A,
Radhakrishnan J, Rovin BH and Kidney Disease: Improving Global
Outcomes (KDIGO) Nephrotic Syndrome In Children Work Group: KDIGO
2025 clinical practice guideline for the management of nephrotic
syndrome in children. Kidney Int. 107:S241–S289. 2025.
|
|
4
|
Ngo QC, Tran HD, Nguyen NBT, Nguyen THT
and Mai TT: Hypomagnesemia as a reversible complication of
cyclosporine in pediatric nephrotic syndrome: A case report. Child
Kidney Dis. 29:142–148. 2025.
|
|
5
|
Leung AK, Wong AH and Barg SS: Proteinuria
in children: Evaluation and differential diagnosis. Am Fam
Physician. 95:248–254. 2017.PubMed/NCBI
|
|
6
|
Huang Y, Yang X, Zhang Y, Yue S, Mei X, Bi
L, Zhai W, Ren X, Ding Y, Zhang S, Deng Z and Sun Y: Correlation of
urine protein/creatinine ratios to 24-h urinary protein for
quantitating proteinuria in children. Pediatr Nephrol. 35:463–468.
2020.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Singh R, Bhalla K, Nanda S, Gupta A and
Mehra S: Correlation of spot urinary protein/creatinine ratio and
quantitative proteinuria in pediatric patients with nephrotic
syndrome. J Family Med Prim Care. 8:2343–2346. 2019.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Lane C, Brown M, Dunsmuir W, Kelly J and
Mangos G: Can spot urine protein/creatinine ratio replace 24 h
urine protein in usual clinical nephrology? Nephrology (Carlton).
11:245–249. 2006.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Kobayashi S, Amano H, Terawaki H, Ogura M,
Kawaguchi Y and Yokoo T: Spot urine protein/creatinine ratio as a
reliable estimate of 24-hour proteinuria in patients with
immunoglobulin A nephropathy, but not membranous nephropathy. BMC
Nephrol. 20(306)2019.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Huynh BA, Ho NHH, Bui TTA, Hoang KC and
Tran TTT: New equation for estimating glomerular filtration rate in
Vietnamese kidney transplant recipients. Int Urol Nephrol.
57:2657–2667. 2025.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Leslie SW, Rathi BM, Sharif S and Rout P:
24-hour urine collection and analysis. In: StatPearls. Treasure
Island (FL): StatPearls Publishing, 2025.
|
|
12
|
Hellerstein S, Simon SD, Berenbom M, Erwin
P and Nickell E: Creatinine excretion rates for renal clearance
studies. Pediatr Nephrol. 16:637–643. 2001.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Ghazali S and Barratt TM: Urinary
excretion of calcium and magnesium in children. Arch Dis Child.
49:97–101. 1974.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Cockcroft DW and Gault MH: Prediction of
creatinine clearance from serum creatinine. Nephron. 16:31–41.
1976.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Eddy AA and Symons JM: Nephrotic syndrome
in childhood. Lancet. 362:629–639. 2003.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Kerlin BA, Haworth K and Smoyer WE: Venous
thromboembolism in pediatric nephrotic syndrome. Pediatr Nephrol.
29:989–997. 2014.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Ellis D: Pathophysiology, evaluation, and
management of edema in childhood nephrotic syndrome. Front Pediatr.
3(111)2016.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Parker K, Ragy O, Hamilton P, Thachil J
and Kanigicherla D: Thromboembolism in nephrotic syndrome:
Controversies and uncertainties. Res Pract Thromb Haemost.
7(102162)2023.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Sun LY, Xu K, Yao Y, Xiao HJ, Liu XY, Su
BG, Zhong XH, Guan N, Zhang HW, Ding J and Wang F: Suitability of
estimated urine protein using different estimated 24 h urine
creatinine equations in children with glomerular diseases. Zhonghua
Er Ke Za Zhi. 60:1178–1184. 2022.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Zhai P, Huang Y, Yue S, Yang X, Luo J,
Zhang Y, Mei X, Bi L, Zhai W, Ren X, et al: Diagnostic efficacy and
influencing factors of urinary protein/creatinine ratio replacing
24-h urine protein as an evaluator of proteinuria in children. Int
Urol Nephrol. 54:1409–1416. 2022.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Yang CY, Chen FA, Chen CF, Liu WS, Shih
CJ, Ou SM, Yang WC, Lin CC and Yang AH: Diagnostic accuracy of
urine protein/creatinine ratio is influenced by urine
concentration. PLoS One. 10(e0137460)2015.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Chuang GT, Tsai IJ and Tsau YK: Serum
creatinine reference limits in pediatric population: A
single-center electronic health record-based database in Taiwan.
Front Pediatr. 9(793446)2021.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Suwanrungroj S, Pattarapanitchai P,
Chomean S and Kaset C: Establishing age- and gender-specific serum
creatinine reference ranges for the Thai pediatric population. PLoS
One. 19(e0300369)2024.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Filler G and Huang SHS: Spot urine protein
to creatinine ratio. Pediatr Nephrol. 32:917–919. 2017.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Yang F, Shi JS, Gong SW, Xu XD and Le W:
An equation to estimate 24-hour total urine protein excretion rate
in patients who underwent urine protein testing. BMC Nephrol.
23(49)2022.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Patil P, Shah V and Shah B: Comparison of
spot urine protein/creatinine ratio with 24-hour urine protein for
estimation of proteinuria. J Assoc Physicians India. 62:406–410.
2014.PubMed/NCBI
|
|
27
|
Shimizu S, Saito H, Takahashi S, Morohashi
T, Hamada R, Hataya H, Kondo Y and Morioka I: Fractional excretion
of total protein in patients with nephrotic syndrome. Pediatr Int.
66(e15722)2024.PubMed/NCBI View Article : Google Scholar
|
|
28
|
Hogan MC, Reich HN, Nelson PJ, Adler SG,
Cattran DC, Appel GB, Gipson DS, Kretzler M, Troost JP and Lieske
JC: The relatively poor correlation between random and 24-hour
urine protein excretion in patients with biopsy-proven glomerular
diseases. Kidney Int. 90:1080–1089. 2016.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Kapur G, Valentini RP, Imam AA and Mattoo
TK: Treatment of severe edema in children with nephrotic syndrome
with diuretics alone: A prospective study. Clin J Am Soc Nephrol.
4:907–913. 2009.PubMed/NCBI View Article : Google Scholar
|
|
30
|
Collins GS, Reitsma JB, Altman DG and
Moons KGM: Transparent reporting of a multivariable prediction
model for individual prognosis or diagnosis (TRIPOD): The TRIPOD
statement. Ann Intern Med. 162:55–63. 2015.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Moons KGM, Altman DG, Reitsma JB,
Ioannidis JP, Macaskill P, Steyerberg EW, Vickers AJ, Ransohoff DF
and Collins GS: Transparent reporting of a multivariable prediction
model for individual prognosis or diagnosis (TRIPOD): Explanation
and elaboration. Ann Intern Med. 162:W1–W73. 2015.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Wolff RF, Moons KGM, Riley RD, Whiting PF,
Westwood M, Collins GS, Reitsma JB, Kleijnen J and Mallett S:
PROBAST: A tool to assess the risk of bias and applicability of
prediction model studies. Ann Intern Med. 170:51–58.
2019.PubMed/NCBI View Article : Google Scholar
|















![Calibration for model M6:
Observed Log10(24hUP) vs. predicted
Log10(24hUP). The 45˚ line is the ideal reference. The
solid line shows the apparent fit; the dashed line shows the
optimism-corrected fit from bootstrap (B=2,000). The two lines
nearly overlap (slope=0.999; intercept=0.002). Model M6:
Log10[24hUP (mg/m2/h)]=0.291 + 0.880 x
Log10 (adjusted UPCR)-0.006xage (years)+0.057xsex
(0=boy, 1=girl). Adjusted
UPCR=mUPCRxe24hUCrCockcroft-Gault; n=171; adjusted
R2=0.80. 24hUP, 24-h urine protein; lm, linear model;
OP-corr, optimism-corrected.](/article_images/wasj/8/2/wasj-08-02-00448-g02.jpg)