1. Introduction
Acute myeloid leukemia (AML) is a form of cancer
that affects the blood and bone marrow, characterized by the rapid
proliferation of abnormal white blood cells (WBCs) that displace
normal cells and may become life-threatening if left untreated
(1). The accumulation of immature
myeloid cells interferes with the production of WBCs, red blood
cells (RBCs) and platelets, resulting in symptoms such as fever,
fatigue, bruising and an increased risk of acquiring infections
(2). AML occurs due to mutations
in certain genes or chromosomes, such as Ras, FLT3, c-kit,
translocation between chromosomes 8 and 21 and the inversion of
chromosome 16. It mainly affects individuals aged ≥60 years
(3). The term ‘acute’ refers to
the rapid onset and progression of the disease, including the
presence of immature myeloid cells, known as blasts, in blood and
bone marrow (2). AML comprises of
different subtypes, including acute promyelocytic leukemia (APL),
acute monocytic leukemia (AML-M5), acute megakaryocytic leukemia
(AML-M7) and other myeloid leukemia variants. These subtypes affect
blood cell development at different stages, cause distinct symptoms
and respond differently to treatment (4). Although AML includes several
biologically distinct subtypes (e.g., APL and AML-M4/M5), in the
current literature, there appears to be an absence of the
subtype-specific profiling of exosomal circular RNAs (circRNAs)
(5). The majority of studies have
assessed either intracellular circRNAs or have evaluated single
exosomal circRNAs in unstratified AML cohorts (6). To the best of our knowledge,
comprehensive analyses comparing exosomal circRNA expression across
AML subtypes have not yet been conducted, limiting conclusions on
diagnostic or prognostic specificity (7). Even though outcomes for patients with
AML are often described as ‘poor’, some patients can be cured and
survival rates can be improved. The standard chemotherapeutic
regimen for AML involves a combination of cytarabine and
anthracycline that has been used for >40 years. Recent
breakthroughs in molecular and cell biology have enhanced the
understanding of the pathophysiology of AML and the available
treatment strategies, and has improved the identification of
patients who are at a higher risk of developing leukemia (8). These advancements result in an
increased number of treatment options for numerous patients and may
provide future opportunities to reduce or prevent the progression
of AML (9).
CircRNAs comprise a diverse and stable class of RNA
molecules generated through the linkage of a downstream splice
donor to an upstream splice acceptor. Certain circRNAs function as
microRNA (miRNA/miR) sponges in the cytoplasm, while others in the
nucleus enhance the expression of their parental genes through
interactions with RNA polymerase II (10). Recently, novel peptides translated
from circRNAs have been identified; for example, the SHPRH protein
encoded by circSHPRH is expressed in normal brain tissue (11,12).
Numerous studies have demonstrated that circRNAs are deregulated in
cancer and leukemia (13,14), contributing to disease pathogenesis
and cancer hallmarks (15).
Exosomes, initially reported to contain mRNA and
miRNA, have since been shown to harbor various small non-coding RNA
(ncRNA) species, in addition to long ncRNAs (lncRNAs), mRNAs and
miRNAs (16,17). Notably, large amounts of circRNAs
have been detected in exosomes, highlighting their high enrichment
and stability primarily in tumor-derived exosomes compared to the
circRNA levels in the parent cells.
In 2019, 61,559 new AML cases were reported among
individuals aged 60-80 years, with 5,362 associated deaths and an
estimated 990,656 disability-adjusted life years lost (18). The number of annual cancer cases in
India doubled between 1990 and 2013. Of note, ~1.39 million cancer
cases were reported in India in 2020, increasing to 1.42 million
and 1.46 million in 2021 and 2022, respectively (19,20).
2. Exosomes
Extracellular vesicles (EVs) are small,
membrane-bound structures enclosed by a lipid bilayer and released
by all types of cells. These vesicles play crucial roles in
cell-to-cell communications and transport a variety of
biomolecules, including proteins, lipids and nucleic acids. EVs are
typically categorized into different types based on their origin.
Microvesicles range in size from 30 to 5,000 nanometers, while
exosomes, a type of EV, contain cargoes such as DNA, RNA, lipids
and proteins (21). Based on their
biogenesis and size, EVs are generally classified into three
groups, namely microvesicles, exosomes and apoptotic bodies
(22): i) Microvesicles bud
directly from the plasma membrane and range from 100 to 1,000 nm in
size; ii) exosomes are produced by the fusion of multivesicular
bodies with the plasma membrane and range from 30 to 150 nm in
size; and iii) apoptotic bodies are released by dying cells into
the extracellular space and range from 50 to 5,000 nm in diameter
(21,23).
In multicellular organisms, exosomes are found in a
variety of biological fluids, including cerebrospinal fluid, urine,
blood and saliva. Their composition is influenced by the
physiological state, environmental stimuli and characteristics of
the parent cells. Exosomes contain diverse proteins, such as
membrane, extracellular matrix and nuclear proteins, as well as
nucleic acids including DNA, mRNA and various ncRNAs. Cytosolic
metabolites are found in some cases (23). Notably, circRNAs are incorporated
into exosomes more than their linear analogue (24). Various studies have demonstrated
the roles of circRNAs in AML, demonstrating that they can either
promote or inhibit tumor development and regulate key malignant
behaviors, such as proliferation, metastasis, invasion and
migration (25). In cancer,
exosomes aid in the communications between tumor cells and the
surrounding microenvironment (26). However, circRNAs within exosomes
have not extensively investigated. Recent research indicates that
exosome-associated circRNAs result in cancer progression, in the
formation of premetastatic niches and metastatic spread (27). Exosomes may also participate in
tumor immune regulation and support cancer therapy (28). The comparison of exosomal circRNAs
comparison between patients with cancer and healthy individuals has
revealed substantial differences, suggesting their clinical
relevance and research potential (29). Exosomes further contribute to tumor
biology by enabling horizontal transfer, thereby transporting
various cargoes from donor cells to recipient cells located either
locally or at distant sites (30).
Communication between malignant cells and surrounding normal or
tumor-associated cells via exosomes significantly influences tumor
progression.
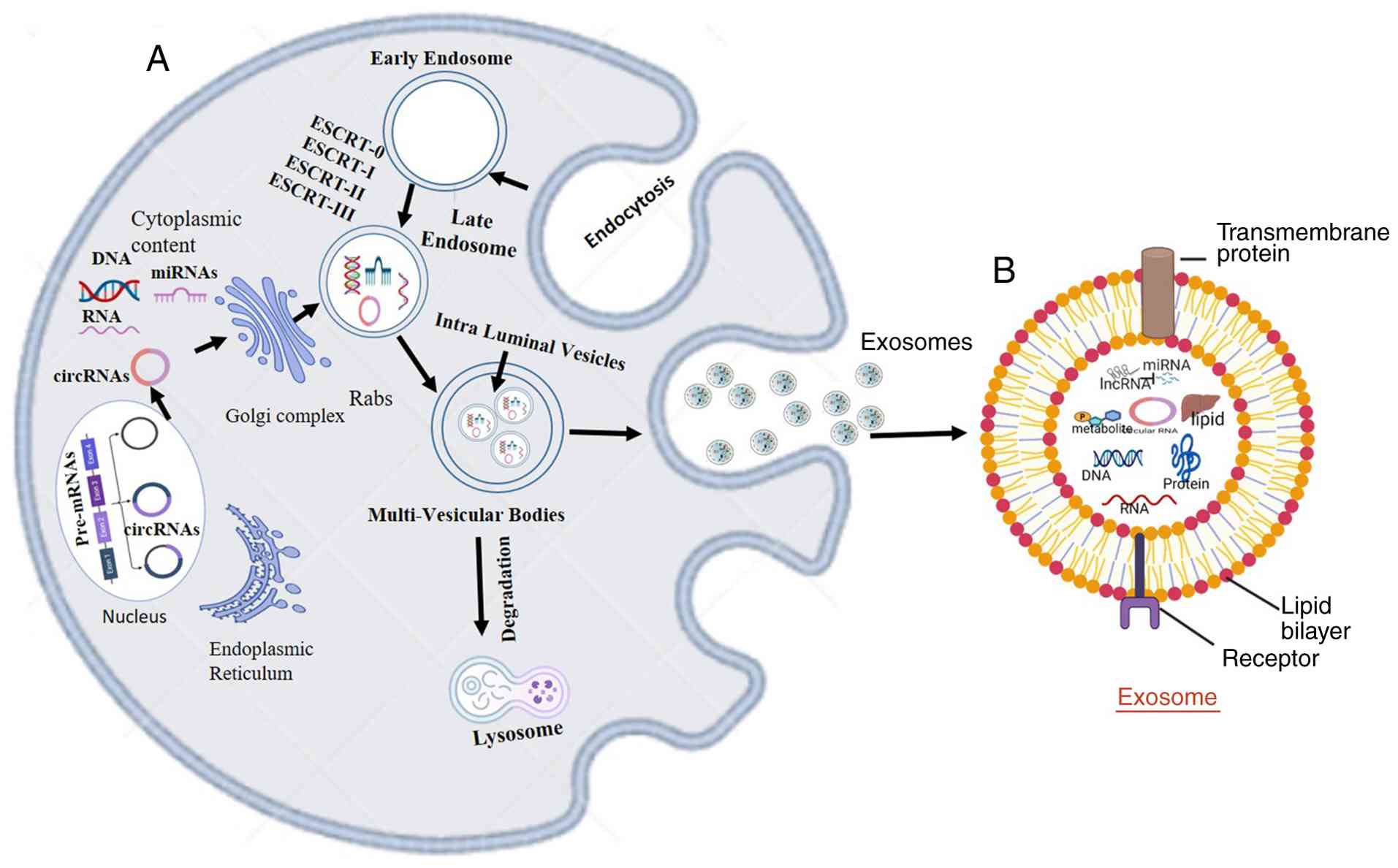
Biogenesis of exosomes
Exosome biogenesis begins with an endocytic event at
the plasma membrane. The most well-described mechanism for exosome
formation is driven by the endosomal sorting complex required for
transport (ESCRT), along with a multimolecular machinery that
operates at the endosomal membrane during the various stages of
exosome biogenesis. Although ESCRT complexes (TSG101, ALIX and VPS
proteins) regulate membrane budding and cargo selection, current
evidence indicates they do not directly recognize circRNA sequences
(31). The selective loading of
circRNAs into exosomes is predominantly mediated by RNA-binding
proteins (RBPs), such as hnRNPA2B1, YBX1, FUS and QKI, which bind
specific sequence motifs or secondary structures, including the
exosome-enrichment motif 5'-GMWGVWGRAG-3' reported in circRNAs
(32). These RBPs recruit circRNAs
to sites of intraluminal vesicle formation, while ESCRT machinery
facilitates vesicle scission (33). Emerging evidence suggests that AML
blasts exhibit an increased exosomal output and an enhanced
enrichment of oncogenic circRNAs compared to normal hematopoietic
cells, likely due to dysregulated RBP expression, altered MVB
dynamics and leukemic stress pathways (34). However, AML-specific
circRNA-sorting mechanisms remain incompletely defined and warrant
further investigation. Exosomes are synthesized from multivesicular
bodies (MVBs). The formation of MVBs can occur through
ESCRT-independent pathways and is influenced by lipid components
such as plasma-membrane-derived lipids and ceramides (35). Additionally, MVB formation is
regulated by syndecan heparan sulfate proteoglycans and the
cytoplasmic adaptor protein syntenin. Rab GTPases and soluble
N-ethylmaleimide-sensitive factor attachment protein receptors
facilitate either the fusion of MVBs with lysosomes for degradation
or their docking at the cell periphery for exosome secretion
(36). Moreover, research
indicates that the elevated expression of p53, pyruvate kinase M2
and tumor suppressor-activated pathway 6 enhances exosome secretion
in tumor cells (37). Exosome
release has also been shown to be influenced by changes in the
microenvironment, including altered pH and the accumulation of
intracellular Ca²+ ions (37). Exosomes are taken up by target
cells through several mechanisms, such as endocytosis and direct
membrane fusion (35). It has been
demonstrated that exosome entry can be inhibited under certain
conditions; for example, the internalization of exosomes by ovarian
cancer cells via multiple endocytic pathways is significantly
reduced at 4˚C (38). The
mechanisms of exosome biogenesis are illustrated in Fig. 1.
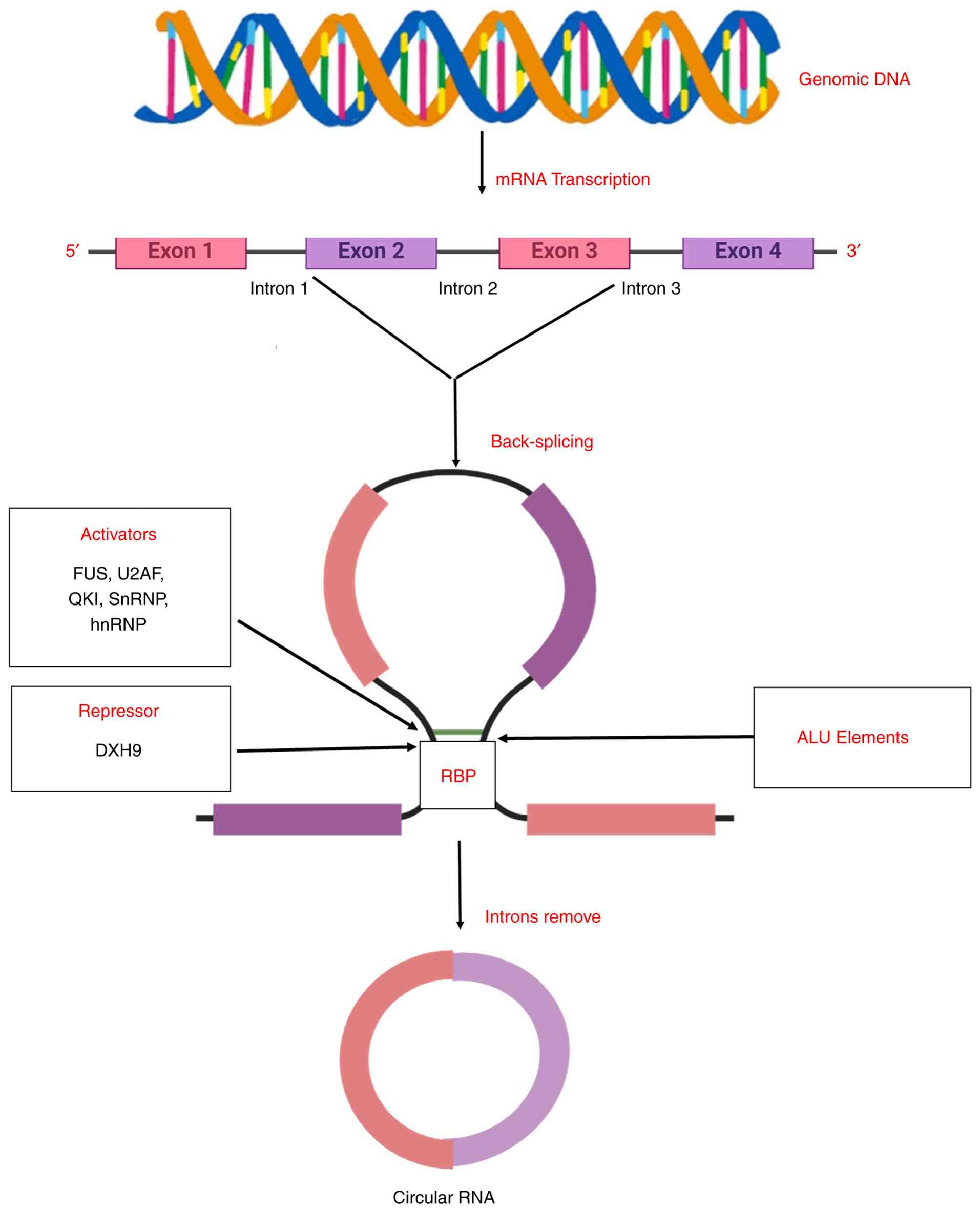
Biogenesis of circRNAs
The biogenesis of circRNAs occurs through the
back-splicing of pre-mRNA, in which a downstream splice donor site
(5'splice site) attaches to an upstream splice acceptor site
(3'splice site), and through transcription mediated by RNA
polymerase II. Various circRNAs can be synthesized by the
alternative back-splicing of the same sequence (39). However, the exact mechanisms that
determine circRNA formation have not yet been fully delineated.
CircRNAs are classified into three main categories based on their
structure and mode of circularization: Circular intronic RNAs
(ciRNAs), exonic circRNAs (ecircRNAs), and exon-intron circRNAs
(EIciRNAs). EcircRNAs consist of one or more exons and are
sometimes generated through alternative splicing that are primarily
located in the cytoplasm (40).
Several models have been proposed to explain the biosynthesis of
ecircRNAs, including RBP-mediated circularization, lariat-driven
circularization and intron-pairing-driven circularization. In the
lariat-driven model, partial RNA folding occurs during pre-mRNA
transcription, leading to exon skipping as the RNA folds. These
structural changes can produce lariat intermediates containing
initially non-adjacent exons along with their introns. CircRNAs are
generated after intron sequences are removed through splicing
within the lariat structure (41).
In the intron-pairing-driven model, reverse complementary sequences
within introns on both sides of the pre-mRNA facilitate
complementary base pairing, mediating circularization. RBPs acting
as trans-acting activators or inhibitors also play key roles in
regulating circRNA production. DHX9, an abundant nuclear RNA
helicase and ALU-element-binding protein, functions as an
antagonist of circRNA biosynthesis. The lLoss of DHX9 increases
circRNA production by allowing the formation of RNA pairs flanking
circularized exons (40).
Additionally, several splicing factors can bind specific RNA motifs
and regulate circRNA biogenesis. Quaking (QKI), a mesenchymal
splicing factor, is crtiical for generating circRNAs during
epithelial-mesenchymal transition. The RBP FUS regulates
back-splicing reactions and thus controls circRNA formation in
mouse embryonic-stem-cell-derived motor neurons. While numerous
RBPs, such as QKI, DHX9, hnRNPA2B1 and FUS regulate circRNA
circularization, AML-specific alterations in these RBPs remain
poorly defined (42). Available
transcriptomic datasets indicate that DHX9 is not downregulated in
AML, but is often upregulated; however, to date, at least to the
best of our knowledge, no study has demonstrated a direct link
between DHX9 expression and increased cellular or exosomal circRNA
production in leukemic cells (43). Similarly, although QKI is a major
enhancer of back-splicing, recurrent QKI mutations have not been
reported in AML, and changes in QKI expression across AML subtypes
have not been shown to modify circRNA biogenesis. Moreover,
although specific AML driver lesions [e.g., FLT3-ITD, NPM1
mutations, t(8;21)] have been associated with altered circRNA
expression patterns, there is currently no evidence that these
mutations directly disrupt intronic complementary sequences or
intron-pairing mechanisms that determine circRNA formation
(25,44). Thus, the influence of
AML-associated RBPs and genomic lesions on circRNA circularization
and exosomal loading remains an important but unresolved area for
future investigation (34).
EIciRNAs are primarily located in the nucleus and consist of exons
and introns that are retained during splicing. These EIciRNAs play
a crucial role in regulating the transcription of their parental
genes (45). During the formation
of ecircRNAs, the introns flanking the exons are typically spliced
out; however, when these introns are retained, EIciRNAs are formed.
Some lariat structures are produced during splicing from introns
where most of them are rapidly degraded through debranching
(40). A schematic illustration of
the biogenesis of exosomal circRNAs is presented in Fig. 2.
3. Functions of exosomal circRNAs in
AML
In recent years, circRNAs have gained attention due
to their diverse functions and regulatory roles in various
biological processes, including cancer. In AML, circRNAs are
recognized as potential biomarkers and regulators of disease
development and progression (46).
The dysregulated expression of circRNAs in patients with AML has
been linked to cellular processes associated with the pathogenesis
of AML, such as apoptosis, proliferation, cell differentiation and
drug resistance. For example, circ_0009910 has been shown to
function as a sponge for miR-20a-5p, thereby influencing the
expression of RUNX1, which promotes the progression of AML by
inhibiting apoptosis and increased cell proliferation, leading to
disease development (25). By
contrast, circ_0004277 has been found to be downregulated in AML.
Circ_0004277 upregulates its target gene, SSBP2, by sequestering
miR-134-5p and resulting in the inhibition of the PI3K/AKT
signaling pathway and the suppression of AML cell invasion and
proliferation (47). CircRNAs have
also been implicated in regulating the response of AML cells to
chemotherapy. For instance, circ_100053 upregulates TCL1A by
sponging miR-29b-3p. The increased expression of circ_100053
enhances CML cell resistance to imatinib, a commonly used
chemotherapeutic drug by promoting cell proliferation and
inhibiting apoptosis (48).
Similarly, resistance to vincristine has been shown to be
associated with the overexpression of circPVT1 in patients with AML
(49). Conversely, some circRNAs
enhance sensitivity to anti-leukemic drugs, as observed with the
knockdown of the fusion circRNA, circM9(50). Overall, circRNAs play vital roles
in the pathogenesis of AML by regulating various signaling pathways
and cellular processes (51).
Dysregulated circRNAs may provide novel therapeutic strategies for
the treatment of AML. However, further studies are required to
fully understand the functions and biogenesis of circRNAs in AML
and to explore their potential clinical applications as biomarkers
(52). Various types of circRNAs
and their roles in AML are presented in Table I.
 | Table IDifferent types of exosomal circRNAs
in AML and their roles and pathways. |
Table I
Different types of exosomal circRNAs
in AML and their roles and pathways.
| Exosomal
circRNA | Roles | Pathways | (Refs.) |
|---|
|
circRNA-0004913 | Regulates AML
progression and chemo resistance | Likely involves
pathways related to cell proliferation, survival, and drug
resistance | (117) |
|
circRNA-300445163 | Associated with AML
diagnosis and prognosis | Pathways related to
leukemogenesis, survival and disease. | (118) |
|
circRNA-0012152 | Regulates AML cell
proliferation and invasion | Signaling pathways
related to cell proliferation, Invasion, and apoptosis | (119,120) |
|
circRNA-0025202 | Involved in AML
pathogenesis and progression | Pathways associated
with leukemogenesis, cell survival, and proliferation | (121) |
|
circRNA-0033386 | Acts as a molecular
sponge for miR-1200 | Regulatory pathways
modulated by miR-1200, impacting cell proliferation and
survival | (117) |
|
circRNA-0042853 | Reduces AML cell
proliferation and apoptosis | Pathways regulating
cell cycle development, apoptosis, and leukemogenesis | (75) |
Exosomal circRNAs in the
progression/development of AML
Several studies have demonstrated that circRNAs
function in AML by either promoting or inhibiting tumor
progression, and modulating the biological behavior of malignant
cells, including proliferation, invasion, migration, and metastasis
(53,54). Exosomal circRNAs perform all of the
aforementioned roles in cancer due to their ability to mediate
intercellular communication. When exosomal circRNAs are taken up by
target cells, they influence the physiological and pathological
states of those cells (55).
Numerous studies have examined the mechanisms through which
exosomal circRNAs contribute to cancer progression and development.
Exosomal circRNAs are circular RNAs enclosed within exosomes where
small extracellular vesicles released by cells into bodily fluids
such as saliva, blood and urine. These exosomal circRNAs
participate in cell-cell communications, other biological processes
and disease progression (56).
Circ-ITCH has been shown to promote the progression of
hepatocellular carcinoma by sponging miR-214. By sequestering
miR-20b-5p, circ-ITCH reduces its inhibitory effect on the target
gene PTEN, leading to activation of the PTEN/PI3K/AKT signaling
pathway, which enhances cell proliferation and inhibits apoptosis
(57). Exosomal circRNAs further
promote cell proliferation and survival by modulating the
expression of genes involved in cell-cycle progression and
apoptosis. CircRNA_0014130, for instance, has been identified as an
oncogenic driver in AML by sponging miR-139-5p/miR-141-3p and
activating the β-catenin signaling pathway, thereby contributing to
the development of AML (25).
Similarly, circRNA_100290 has been shown to be transferred from AML
cells to bone marrow stromal cells, where it upregulates CXCL16
expression. CXCL16, in turn, supports the progression and invasion
of AML cells within the bone marrow microenvironment, promoting
disease development (58).
Exosomal circ_0004136 was among the first circRNAs implicated in
AML: The knockdown of circ_0004136 was shown to reduce AML cell
viability, migration, invasion and promoted apoptosis, whereas its
overexpression was shown to enhance proliferation and invasive
behavior in vitro (59).
Mechanistic studies link circ_0004136 to the miR-570-3p/TSPAN3
axis. As circ_0004136 was detected in AML serum exosomes, it is
considered a promising non-invasive biomarker and a potential
therapeutic target (60). While
earlier studies focused on intracellular circ_0009910(61), more recent research indicates that
it is enriched in AML-derived exosomes: Exosomal circ_0009910
promotes the proliferation, reduces the apoptosis and regulates the
cell cycle of AML cell lines (61). The functional mechanism involves
the sponging of miR-5195-3p, leading to the upregulation of GRB10,
which drives leukemic cell survival and proliferation. These data
indicate that exosomal circ_0009910 may contribute to AML
pathogenesis via intercellular signaling through EVs (51). Exosomal circRNAs affect the
antitumor immune microenvironment and promote immune evasion and
thus regulate immune response in AML (62). CircRNA_0000190 upregulates CD96 by
sponging miR-142-3p, which suppresses proliferation and the
cytotoxic activity of natural killer cells (61). Certain circRNAs activate
pro-inflammatory signaling pathways or promote cytokine and
chemokine secretion by immune cells or stomal cells. These
inflammatory mediators provide a tumor-promoting environment that
enhances AML cell survival and proliferation.
Stromal and AML cells communicate through exosomal
circRNAs in the bone marrow microenvironment (63). This intercellular crosstalk
influences disease progression, drug resistance and immune evasion
crucial for the development of AML. Exosomes containing specific
circRNAs that are secreted by AML cells can be transferred to
neighboring stromal cells. For example, circRNA_100290 has been
shown to be transferred from AML cells to bone marrow stromal
cells. Once internalized, circRNA_100290 promotes the expression of
CXCL16, which is involved in cell migration and invasion. Exosomal
circRNAs adapt to the stromal-cell environment that influence their
interactions with AML cells and modify the surrounding
microenvironment (64). By
regulating the expression or activation of specific genes or
signaling pathways within stromal cells, circRNA contribute to
shaping conditions that support leukemic growth (Fig. 3, top panel). In a 2023 study
analyzing public AML circRNA microarray datasets (GSE94591 and
GSE163386), circZBTB46 was among the most consistently upregulated
circRNAs in patients with AML compared to healthy controls.
Although not yet functionally characterized in vitro or
in vivo, its overexpression suggests potential relevance for
AML biology, warranting further studies (65).
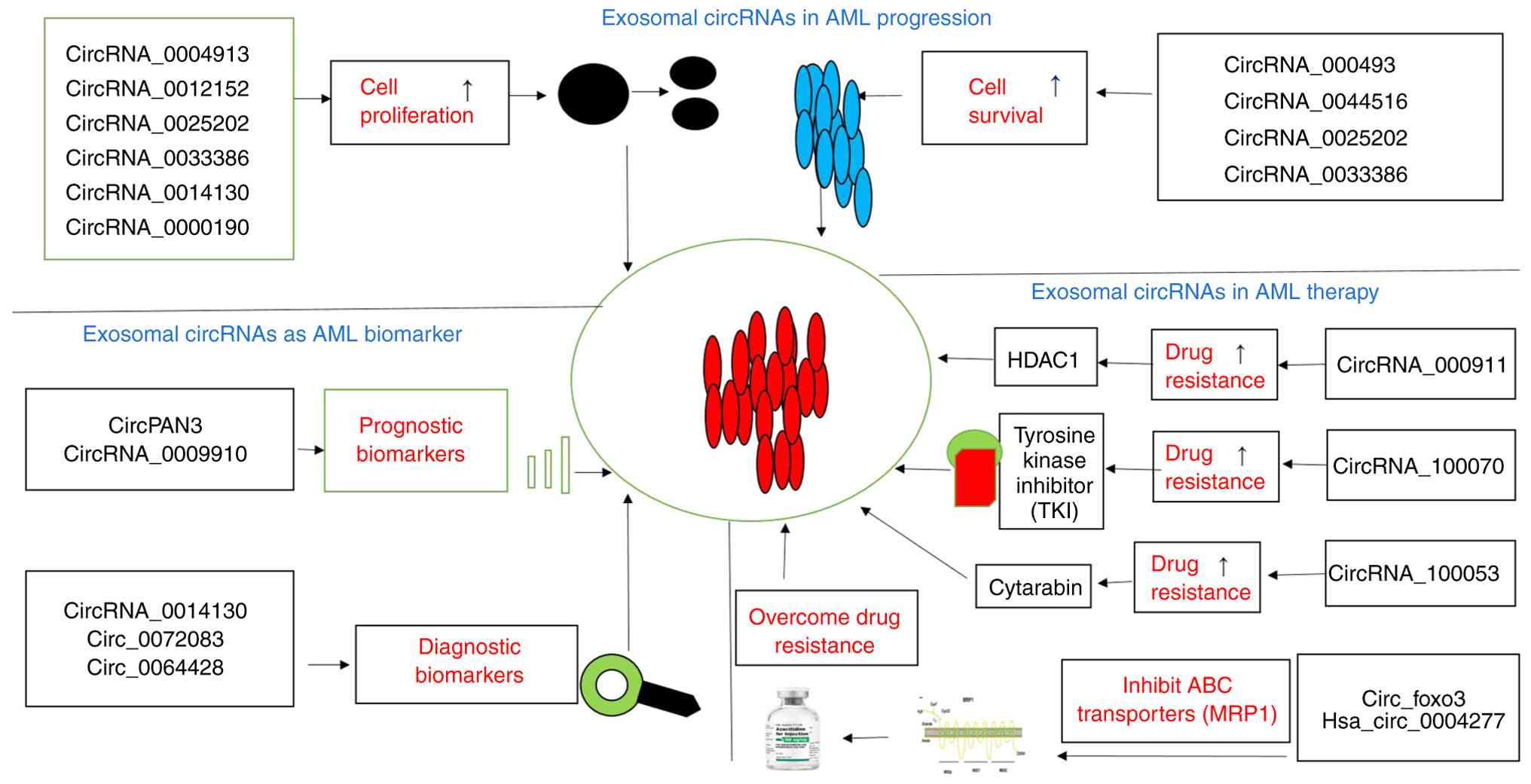 | Figure 3(Top panel) Different examples of
exosomal circular RNAs which play a crucial role in AML
progression. CircRNA_0004913, CircRNA_00012152, CircRNA_0025202,
CircRNA_0033386, CircRNA_0014130 and CircRNA_0000190 are involved
in cell proliferation, while CircRNA_000493, CircRNA_0044516,
CircRNA_0025202 and CircRNA_0033386 are involved in cell survival.
(Bottom left panel) Exosomal circular RNA as AML biomarkers. The
following circRNAs, i.e., Circ_PAN3 and CircRNA_0009910 are
prognostic biomarkers and CircRNA_0014130, Circ_0072083 and
Circ_00064428 are diagnostic biomarkers. (Bottom right panel)
Different exosomal circular RNAs involved in AML therapy. These
includes CircRNA_000911, CircRNA_100070, and CircRNA_100053 which
facilitates drug resistance while Circ_foxo3 and Has_circ_0004277
which overcome drug resistance by inhibiting ABC transporters. AML,
acute myeloid leukemia. |
Exosomal circRNAs as tumor biomarkers
in AML
In recent years, the diagnosis of cancer has been a
major focus of scientific research worldwide. Various accessible
methods for the diagnosis of cancers have continued to emerge
(61,66), using samples collected from tears,
urine (67), semen, ascites,
synovial fluid, cerebrospinal fluid, amniotic fluid (68) and feces. These sample types are
easier and less invasive to obtain, and once biomarkers are
detected, biopsies can be performed to analyze tumor tissues. This
highlights the advantages of using exosomes as potential
biomarkers. Moreover, circRNAs, as part of exosomal cargo exhibit
characteristics such as structural diversity, conservation,
stability and specificity in expression and localization (69).
Exosomal circRNAs have emerged as promising
biomarkers as they offer several advantages, such as abundance,
stability and disease specificity (69). They are considered as strong
diagnostic candidates due to their differential expression in
diseased AML and healthy individuals. For example, the expression
of circRNA_0014130 has been shown to be high in patients with AML
compared with healthy controls (25). CircRNA_0014130 has been reported to
exhibit better diagnostic accuracy, with more than 90% sensitivity
and specificity (25). The
identification of exosomal circRNAs may help in early detection of
AML resulting in timely intervention and improved patient outcomes.
CircRNA_0014130 has been detected in the plasma exosomes of AML
patients at early stages of disease, suggesting it to be an early
diagnostic biomarker (25). These
results are promising; however, multi-center validation and larger
cohorts are required to confirm reproducibility and clinical
applicability (70). Exosomal
circ_0006896 has been shown to be upregulated in AML, and promotes
proliferation, chemoresistance and immune-evasive behavior in
preclinical models (71). Recent
reviews and meta-analyses have provided the increasing evidence
that exosomal cargo, such as circRNAs, modulates immune checkpoints
and that engineered exosomes are being developed to augment immune
checkpoint inhibitor and adoptive-cell therapies (72).
Studies have indicated that exosomal circRNAs may be
used to monitor disease development and treatment response in
patients with AML. Variations in the expression levels of specific
exosomal circRNAs throughout the course of therapy can provide key
insight into therapeutic efficacy and disease status (73). Monitoring circRNA levels enables
the assessment of treatment response and the detection of disease
recurrence (51). The isolation
and detection of exosomal circRNAs from peripheral blood samples
offer a minimally invasive approach for AML diagnosis and
monitoring (74). Certain exosomal
circRNAs exhibit prognostic value as they are associated with
survival outcomes in AML patients (75). For example, circRNA_0009910
correlates with poor prognosis as higher expression levels are
linked to reduced overall survival and ehanced risk of relapse
(76). Exosomal circRNAs may also
help in determining the treatment response and clinical outcomes in
patients undergoing treatment. Expression levels of certain
exosomal circRNAs is associated with disease advancement and
treatment resistance in AML patients providing information related
to disease severity (25).
Exosomal circRNAs can also be used to monitor minimal residual
disease in patients with AML following therapy, which provides
information related to disease recurrence and treatment response.
Patients with hepatocellular carcinoma exhibiting complete
remission following chemotherapy exhibit decreased levels of
circ-ITCH, suggesting its role as a predictive biomarker of the
treatment response (77).
It has been demonstrated that the expression levels
of certain circRNAs before treatment may determine the response to
targeted therapy thereby acting as a predictive biomarker for
targeted therapies. For example, it has been shown that higher
levels of Circ_0059706 indicate a poor response to tyrosine kinase
inhibitor (TKI) therapy and a shorter progression-free survival in
AML (78,79). Exosomal circRNAs exhibit diagnostic
and prognostic potential across various types of cancer. For
example, the study conducted by Zheng et al confirmed that
exosomal circRNAs derived from the serum of patients with gastric
cancer served as diagnostic markers for gastric cancer (80). Exosomal circ_0072083 expression was
previously shown to be upregulated in patients with glioma compared
to healthy controls (81).
Similarly, the dysregulated expression of exosomal hsa_circ_0064428
in hepatocellular carcinoma is associated with clinical
characteristics, such as WBC count and bone marrow blast percentage
(82). Elevated levels of circPAN3
have been shown to be associated with poor survival rates of
patients with AML, exhibiting its prognostic significance (83). Likewise, Yan and Yan (84) reported that the overexpression of
exosomal circRNA 100199 indicates severe clinical outcomes,
including resistance to chemotherapy and a shorter survival
duration. It has also been shown that the inhibition of exosomal
circ_0072083 suppresses AML cell proliferation and induces
apoptosis, highlighting its potential as a therapeutic target
(29). Thus, targeting exosomal
circRNAs holds considerable therapeutic promise in the management
of AML (Fig. 3, bottom right
panel).
Exosomal-based circular RNA delivery
for AML
CircRNAs play critical roles in cancer development,
including breast cancer, gastric cancer and AML. Exosomes are used
as delivery vehicles for circRNAs, which exhibits a promising
therapeutic strategy for the treatment of AML. Exosomes are
generated through the endosomal pathway and contain biomolecules,
including lipids, proteins and nucleic acids (DNA and RNA).
Specific RNA motifs help in the loading of circRNAs into exosomes
(73). Using motif prediction
software, circRNAs with motif 5'-GMWGVWGRAG-3' were predicted to be
enriched in exosomes, observed by circRNA-seq data from human blood
exosomes (85). CircRNAs
contribute to the pathogenesis of AML by regulating key signaling
pathways and gene expression. Delivering specific circRNAs via
exosomes is expected to regulate these pathways and reduce leukemic
cell proliferation and survival. Exosome isolation can be achieved
through several techniques, such as ultracentrifugation,
size-exclusion chromatography and precipitation. Exosomes are
characterized on the basis of morphology, size and protein markers,
including CD9, CD63 and CD81 along with their molecular contents
(86). Preclinical studies with
AML animal models were used to evaluate the efficacy of exosomal
circRNA delivery. These studies involve systemic administration of
exosomes loaded with specific circRNAs, followed by monitoring
leukemic burden, survival outcomes, and potential adverse effects.
Although the therapeutic application of exosomal circRNA delivery
in AML is promising, several challenges remain, including the
scalability of exosome production, optimization of loading
efficiency, safety issues associated with systemic administration
and the need to harness specific properties of exosomes to deliver
circRNAs effectively Addressing these limitations may lead to the
development of AML therapies capable of overcoming current
treatment barriers and improving patient outcomes (87).
CircRNAs can be loaded into exosomes either by
transfecting donor cells (e.g., mesenchymal stem cells) with
circRNA-expressing plasmids or through direct loading methods such
as electroporation. Transfection-based approaches allow the
efficient loading of circRNAs into exosomes during their natural
biogenesis within donor cells. Direct loading methods offer
simplicity and flexibility, but may exhibit lower loading
efficiency and can potentially alter exosomal properties. Overall,
different loading techniques vary in efficiency and may have
distinct downstream effects. Electroporation and donor-cell
transfection are the most commonly used approaches: Optimized
electroporation protocols report apparent loading efficiencies on
the order of 20-35% (compared with passive incubation); however,
electroporation can produce RNA precipitation artifacts and may
require rigorous controls (88).
Donor-cell engineering (expressing circRNA constructs or
RNA-binding domain fusions) often yields higher functional loading
and preserves exosome integrity (89). Once loaded with circRNAs, exosomes
can be administered systemically or locally to patients with AML
(90). After any loading method,
EV quality control (particle count/size, TEM morphology and
tetraspanin markers, such as CD9/CD63) and immune-safety testing
(PBMC cytokine release, in vivo cytokine panels) are
essential as cargo and surface modifications can alter
immunogenicity (91). Exosomes
possess inherent tropism toward tumor cells, including AML cells,
due to the presence of specific surface proteins and ligands.
Additionally, the modification of exosomal surface proteins or
incorporation of targeting peptides can further enhance their
specificity for AML cells. Once internalized, exosomes release
their cargo including circRNAs into the cytoplasm of AML cells.
These circRNAs can then exert regulatory effects; for example,
circRNAs functioning as miRNA sponges can sequester oncogenic
miRNAs involved in the progression of AML, while other circRNAs may
modulate the expression of key AML-associated genes or signaling
pathways (92). For targeted
delivery to AML blasts, surface functionalization approaches
directed at AML antigens (e.g., CD33) have been reported and are
promising; however, preclinical data testing exosomal circRNA
delivery together with standard AML drugs (cytarabine, FLT3
inhibitors) are currently limited, and combination experiments are
an important future direction (93).
Exosomal circRNAs in AML
metastasis
Exosomes facilitate communications between AML cells
and various other cells within the microenvironment, including
endothelial cells, immune cells and bone marrow stromal cells
(26,92). For instance, AML-derived exosomes
enriched with miR-155 have been shown to promote angiogenesis by
targeting endothelial cells (94).
These exosomes can enhance angiogenesis, a crucial process for
tumor growth and metastasis by transporting pro-angiogenic factors,
such as VEGF and FGF-2, which stimulate endothelial cell
proliferation and blood vessel formation. Exosomes also play a key
role in immune evasion by suppressing anti-tumor immune responses.
AML-derived exosomes can carry immunosuppressive molecules,
including TGF-β and PD-L1, which inhibit cytotoxic T-cell activity
and thereby promote immune escape (95).
These exosomes can enhance angiogenesis, a process
crucial for tumor growth and metastasis by transporting
pro-angiogenic factors, such as VEGF and FGF-2, which stimulate
endothelial cell proliferation and blood vessel formation. Exosomes
aid in the suppression of antitumor immune responses and thus play
a crucial role in immune evasion. AML-derived exosomes can carry
immunosuppressive molecules, such as TGF-β and PD-L1 inhibiting
cytotoxic T-cell activity and thereby promoting immune escape
(96). Among these, exosomal
circRNA_0056616 was shown to promote AML cell migration and
proliferation by regulating the miR-625-5p/HOXA13 axis (97). The prevalence and clinical
significance of these individual axes in metastatic AML remain
unquantified. The majority of supporting data are derived from
in vitro experiments, small discovery datasets, or from
non-AML tumor studies; no cohort-level studies that report the
proportion of patients with metastatic AML with the dysregulation
of circRNA_0056616 axis have been identified (98). Metastatic AML typically signifies
an advanced disease stage and a higher leukemic burden, both of
which contribute to treatment resistance and reduced survival rates
(99). The extramedullary
manifestations of AML can also pose diagnostic challenges as their
symptoms and clinical presentations may resemble those of other
conditions or primary malignancies affecting the same organs
(100). The accurate diagnosis of
AML metastasis requires comprehensive evaluation, including imaging
studies (e.g., CT scans, MRI), bone marrow biopsy, and, in some
cases, tissue biopsy of the involved extramedullary site.
Metastatic AML often necessitates tailored treatment approaches
based on the extent and location of disease involvement. Apart from
standard chemotherapy, patients with central nervous system
metastasis may require intrathecal chemotherapy or cranial
irradiation to control AML. Similarly, extramedullary AML lesions
may require treatments, such as radiation therapy or surgical
removal in combination with systemic therapy (101). AML metastasis increase the risk
of post-treatment relapse, including metastasis to the central
nervous system. During remission in the bone marrow, residual
leukemic cells present in extramedullary sites may serve as a
reservoir for relapse. Therefore, to reduce the risk of relapse
strategies are to be designed to remove of prevent extramedullary
disease (102). Patients with
metastatic AML require close monitoring and surveillance to detect
disease recurrence or progression, especially in extramedullary
locations. Regular imaging and clinical assessments are critical
for early relapse detection and timely intervention. Furthermore,
the presence of metastatic disease may influence both the frequency
and intensity of post-remission monitoring strategies. Effective
symptom management and supportive care targeted toward alleviating
pain, addressing neurological deficits and managing organ
dysfunction remain critical components of comprehensive AML care in
the metastatic setting (100).
Roles of exosomal circRNAs in AML
therapy
The advantage of exosome-based therapy is their
ability to mediate intercellular communication and deliver
therapeutic agents to specific targets. Due to the small size of
exosomes, their content is protected and thus they act as efficient
therapeutic markers (103). In
several studies, exosomes have shown improved therapeutic benefits
in cancer treatment compared with the direct use of
chemotherapeutic agents (104).
Exosomes also offer potential solutions to limitations associated
with current drug delivery approaches, particularly those involving
programmable RNAs, which often suffer from low uptake efficiency
and substantial cytotoxicity, rendering them unsuitable for
clinical application (105). To
address these challenges, researchers have demonstrated and
validated a novel strategy for generating large quantities of
RBC-derived EVs intended for RNA drug delivery (106). Furthermore, gold
nanoparticle-based targeting systems have been used to engineer
specific exosome types capable of precise and selective elimination
of cancer cells (107-109).
While exosomes containing endogenous circRNAs may contribute to
cancer progression, exosomes loaded with therapeutic circRNAs or
custom-engineered siRNAs against pathogenic circRNAs hold
considerable promise for inhibiting tumor growth. Delivering
circRNAs that suppress cancer hallmarks and promote apoptosis may
further enhance the development of precise, effective therapeutic
strategies (110).
Exosomal circRNAs affect treatment response by
regulating processes such as drug resistance, apoptosis and immune
evasion and thus have emerged as important regulators of
therapeutic responses in AML. CircRNA_000911 has been implicated in
chemoresistance in by sponging miR-449a and upregulating HDAC1,
which reduces apoptosis and promotes cell survival (111). CircRNAs may also regulate drug
efflux pump encoding genes that leads to alteration in the
intracellular concentration of chemotherapeutic agents and thus
contributes to resistance. Exosomal circRNAs can also regulate
apoptosis-related pathways affecting sensitivity of AML cells to
treatment-induced cell death. Moreover, exosomal circRNAs play
critical roles in the response to targeted therapies in AML,
including genetically targeted drugs and TKIs. By regulating the
expression of pro- or anti-apoptotic genes, circRNAs can shift the
balance between cell survival and apoptosis (68). For instance, circRNA_0004015
promotes resistance to TKI therapy by upregulating anti-apoptotic
proteins and suppressing chemotherapy-induced apoptosis (112). The overexpression of such
circRNAs can enhance immune evasion and tumor progression by
inhibiting T-cell activation and cytokine secretion (76).
Exosomal circRNAs can regulate molecular pathways
that affect drug metabolism, cell survival and apoptosis thereby
influencing the efficacy of targeted therapies (29). They also modulate the expression of
genes encoding drug targets or signaling molecules involved in
precision therapy (75).
Additionally, exosomal circRNAs can alter drug metabolism and
pharmacokinetics by affecting intracellular uptake and
bioavailability. For example, circPAN3 has been shown to be
associated with development of AML drug resistance doxorubicin
through regulating autophagy (83). Exosomal circRNAs may serve as
predictive biomarkers of treatment resistance and disease relapse.
Reduced intracellular drug accumulation and decreased sensitivity
to chemotherapeutic agents (25),
combined with treatment-induced changes in circRNA expression, can
indicate the emergence of resistant clones and the need for
alternative therapeutic strategies. Integrating exosomal circRNA
profiles with clinical parameters, such as cytogenetics, molecular
mutations and patient demographics significantly enhances their
predictive value. Multivariate analyses incorporating circRNA
levels and conventional prognostic factors have yielded improved
risk stratification and more accurate assessments of the treatment
response (113). For instance,
the increased expression of circ_100053 enhances CML cell
resistance to cytarabine by promoting proliferation and inhibiting
apoptosis (48).
Exosomal circRNAs may also regulate gene expression
programs involved in AML progression and therapy response (75). Certain circRNAs, such as circ-foxo3
and circ-0004277, exhibit an altered expression in de novo
AML and are associated with disease characteristics (51). Their stability renders exosomal
circRNAs attractive candidates for reliable diagnostic biomarkers
that support early detection and disease monitoring. Identifying
exosomal circRNAs involved in the pathogenesis of AML may also
reveal therapeutic opportunities aimed at targeting their
expression or function. Furthermore, exosomal circRNAs can exert
immunomodulatory effects, influencing interactions between AML
cells and the immune system. Understanding these associations may
have key implications for AML immunotherapy strategies (116) (Fig.
3, bottom left panel).
4. Prospects and challenges of exosomal
circRNA in AML
Advanced molecular profiling that identifies the
various mutations and molecular aberrations driving AML
pathogenesis has paved the way for targeted therapies. Specific
drugs that target key mutations, such as FLT3 inhibitors (e.g.,
gilteritinib and midostaurin) and isocitrate dehydrogenase (IDH)
inhibitors have already demonstrated promising outcomes in clinical
trials. Immunotherapeutic approaches, combined with the integration
of transcriptomic, genomic, and proteomic data, are enabling more
individualized treatment strategies in AML. Therapies tailored to
the mutational profile and risk stratification of each patient hold
significant potential for optimizing treatment outcomes. Ongoing
research into novel agents, including differentiation-inducing
compounds, epigenetic modifiers, and emerging targeted therapies,
continues to open new avenues for the management of AML.
Combination therapies targeting multiple pathways simultaneously
are also being explored to overcome drug resistance and improve
response rates (115).
One of the major challenges in AML is the
development of treatment resistance, which contributes to disease
relapse and poor outcomes. Resistance mechanisms, such as clonal
evolution, tumor heterogeneity and the activation of alternative
signaling pathways require the development of innovative
therapeutic strategies. Although targeted therapies have shown
benefit in specific AML subgroups, a number of patients either lack
actionable mutations or eventually develop resistance (116). Identifying additional druggable
targets and developing rational combination therapies are essential
for improving treatment responses in these patients. The
significant molecular and phenotypic heterogeneity observed among
patients with AML contributes to variable therapeutic responses and
clinical outcomes. evolution and the emergence of subclones with
distinct genetic alterations further complicate treatment planning
and disease monitoring. Moreover, intensive chemotherapy regimens
commonly used in AML are associated with substantial toxicity,
including myelosuppression, infection, and organ damage. Balancing
therapeutic efficacy with effective toxicity management therefore
remains a major clinical challenge (116).
5. Conclusions and future perspectives
Exosomal circRNAs have emerged as key players in the
pathogenesis of AML. Encapsulated within exosomes, these circRNA
molecules exhibit regulatory functions in AML progression and hold
potential implications for diagnosis, prognosis, and therapy.
Exosomal circRNAs participate in various cellular processes and
play vital roles in the Clonal development of AML, including
proliferation, apoptosis evasion, drug resistance and metastasis.
They also serve as non-invasive biomarkers for AML diagnosis. Their
stability in body fluids, such as urine and blood, along with their
dysregulated expression patterns in patients with AML, renders them
attractive candidates for early detection and disease monitoring.
Clinical studies have linked specific exosomal circRNA expression
profiles with patient outcomes. High expression levels of
particular circRNAs are associated with adverse prognostic factors,
including chemotherapy resistance, disease relapse and a poor
overall survival. Targeting exosomal circRNAs represents a novel
therapeutic strategy in AML management; modulating their expression
could potentially restore normal cellular processes, overcome drug
resistance and improve treatment efficacy. Despite their promise,
the clinical application of exosomal circRNAs faces several
challenges, including the need for standardized isolation and
detection techniques, clarification of their precise molecular
mechanisms, and validation of their therapeutic utility in large
patient cohorts. The selection of the exosome isolation method
substantially affects downstream circRNA recovery and assay
specificity. For plasma-based AML biomarker work, it is generally
recommended to use ultrafiltration (UF) followed by size-exclusion
chromatography (SEC) as SEC provides superior removal of abundant
plasma protein and lipoprotein contaminants while preserving intact
vesicles and RNA suitable for qPCR and RNA-seq. Single-step
precipitation methods yield higher apparent particle/RNA yield but
poor purity and are not suitable for clinical validation. Future
research is warranted to focus on addressing the challenges that
limit the full potential of exosomal circRNAs in AML diagnosis and
therapy. All clinical validation studies should follow MISEV2018
reporting, include pre-analytical SOPs (double-spin plasma,
hemolysis checks, storage at -80˚C), and report EV QC metrics
(NTA/TEM, CD9/CD63/CD81, negative markers) (59).
To address existing gaps in the literature, the
present review primarily focused on the role of circRNAs in AML and
discussed their potential in disease management. This includes
their contributions to leukemogenesis and drug resistance, as well
as their promise as novel diagnostic, therapeutic and prognostic
biomarkers. The present review also discussed the involvement of
exosomal circRNAs, providing a deeper perspective on the genetic
mechanisms underlying AML, which may open new avenues for targeted
therapy. Furthermore, future studies are warranted to translate
these findings into clinical applications in order to improve the
diagnosis and management of patients with AML.
Acknowledgements
The authors would like to thank Sharda University
(Greater Noida, India) for providing all the resources, such as
journals accessibility, and lab space.
Funding
Funding: No funding was received.
Availability of data and materials
Not applicable.
Authors' contributions
SK conceptualized the study. THA, PKT, MS, SKM and
AMSU were involved in the writing and preparation of the original
draft of the manuscript. THA, RM and MS were involved in the
writing, review and editing of the manuscript. PKT, THA and SK were
involved in visualization. SK supervised the study. All authors
have read and agreed to the published version of the manuscript.
Data authentication is not applicable.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Vakiti A and Mewawalla P: Acute Myeloid
Leukemia. In: StatPearls. StatPearls Publishing, Treasure Island
(FL), 2024.
|
|
2
|
Chennamadhavuni A, Lyengar V, Mukkamalla
SKR and Shimanovsky A: Leukemia. In: StatPearls. StatPearls
Publishing, Treasure Island (FL), 2024.
|
|
3
|
Estey EH: Acute myeloid leukemia: 2019
update on risk-stratification and management. Am J Hematol.
93:1267–1291. 2018.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Ferrara F and Schiffer CA: Acute myeloid
leukaemia in adults. Lancet. 381:484–495. 2013.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Liu Y, Cheng Z, Pang Y, Cui L, Qian T,
Quan L, Zhao H, Shi J, Ke X and Fu L: Role of microRNAs, circRNAs
and long noncoding RNAs in acute myeloid leukemia. J Hematol Oncol.
12(51)2019.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Papaioannou D, Volinia S, Nicolet D,
Świerniak M, Petri A, Mrózek K, Bill M, Pepe F, Walker CJ, Walker
AE, et al: Clinical and functional significance of circular RNAs in
cytogenetically normal AML. Blood Adv. 4:239–251. 2020.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Mirazimi Y, Aghayan AH, Atashi A,
Mohammadi D and Rafiee M: Prognostic value of circular RNAs
expression and their correlation with clinicopathological features
in acute myeloid leukemia: A systematic review and meta-analysis.
Ann Hematol. 104:2095–2124. 2025.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Ishii H and Yano S: New therapeutic
strategies for adult acute myeloid leukemia. Cancers (Basel).
14(2806)2022.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Cingam SR and Koshy NV: Acute
Promyelocytic Leukemia. In: StatPearls. StatPearls Publishing,
Treasure Island (FL), 2024.
|
|
10
|
Li Z, Huang C, Bao C, Chen L, Lin M, Wang
X, Zhong G, Yu B, Hu W, Dai L, et al: Exon-intron circular RNAs
regulate transcription in the nucleus. Nat Struct Mol Biol.
22:256–264. 2015.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Pamudurti NR, Bartok O, Jens M,
Ashwal-Fluss R, Stottmeister C, Ruhe L, Hanan M, Wyler E,
Perez-Hernandez D, Ramberger E, et al: Translation of CircRNAs. Mol
Cell. 66:9–21.e7. 2017.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Legnini I, Di Timoteo G, Rossi F, Morlando
M, Briganti F, Sthandier O, Fatica A, Santini T, Andronache A, Wade
M, et al: Circ-ZNF609 is a circular RNA that can be translated and
functions in myogenesis. Mol Cell. 66:22–37.e9. 2017.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Hirsch S, Blätte TJ, Grasedieck S,
Cocciardi S, Rouhi A, Jongen-Lavrencic M, Paschka P, Krönke J,
Gaidzik VI, Döhner H, et al: Circular RNAs of the nucleophosmin
(NPM1) gene in acute myeloid leukemia. Haematologica.
102:2039–2047. 2017.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Buratin A, Paganin M, Gaffo E, Dal Molin
A, Roels J, Germano G, Siddi MT, Serafin V, De Decker M, Gachet S,
et al: Large-scale circular RNA deregulation in T-ALL: Unlocking
unique ectopic expression of molecular subtypes. Blood Adv.
4:5902–5914. 2020.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Huang Z, Shan R, Wen W, Li J, Zeng X and
Wan R: The emerging roles of Circ-ABCB10 in cancer. Front Cell Dev
Biol. 10(782938)2022.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Li Y, Zheng Q, Bao C, Li S, Guo W, Zhao J,
Chen D, Gu J, He X and Huang S: Circular RNA is enriched and stable
in exosomes: A promising biomarker for cancer diagnosis. Cell Res.
25:981–984. 2015.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Shao N, Xue L, Wang R, Luo K, Zhi F and
Lan Q: miR-454-3p is an exosomal biomarker and functions as a tumor
suppressor in glioma. Mol Cancer Ther. 18:459–469. 2019.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Chen P, Liu X, Zhao Y, Hu Y, Guo J and
Wang H: Global, national, and regional burden of acute myeloid
leukemia among 60-89 years-old individuals: Insights from a study
covering the period 1990 to 2019. Front Public Health.
11(1329529)2024.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Philip C, George B, Ganapule A, Korula A,
Jain P, Alex AA, Lakshmi KM, Sitaram U, Abubacker FN, Abraham A, et
al: Acute myeloid leukaemia: Challenges and real world data from
India. Br J Haematol. 170:110–117. 2015.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Nalage DN, Kudnar PS, Langhe R, Ughade B,
Sontakke T and Biradar A: Leukemia in india: Insights into
incidence, prevalence, mortality, and Disability-adjusted life
years. Cureus. 16(e62557)2024.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Kalluri R and LeBleu VS: The biology,
function, and biomedical applications of exosomes. Science.
367(eaau6977)2020.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Tiwari P, Shanmugam P, Karn V, Gupta S,
Mishra R, Rustagi S, Chouhan M, Verma D, Jha NK and Kumar S:
Extracellular vesicular miRNA in pancreatic cancer: From lab to
therapy. Cancers (Basel). 16(2179)2024.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Lu Y, Huang W, Li M and Zheng A:
Exosome-based carrier for RNA delivery: Progress and challenges.
Pharmaceutics. 15(598)2023.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Dou Y, Cha DJ, Franklin JL, Higginbotham
JN, Jeppesen DK, Weaver AM, Prasad N, Levy S, Coffey RJ, Patton JG
and Zhang B: Circular RNAs are down-regulated in KRAS mutant colon
cancer cells and can be transferred to exosomes. Sci Rep.
6(37982)2016.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Singh V, Uddin MH, Zonder JA, Azmi AS and
Balasubramanian SK: Circular RNAs in acute myeloid leukemia. Mol
Cancer. 20(149)2021.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Jin Y, Xing J, Xu K, Liu D and Zhuo Y:
Exosomes in the tumor microenvironment: Promoting cancer
progression. Front Immunol. 13(1025218)2022.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Li J, Li Z, Jiang P, Peng M, Zhang X, Chen
K, Liu H, Bi H, Liu X and Li X: Circular RNA IARS (circ-IARS)
secreted by pancreatic cancer cells and located within exosomes
regulates endothelial monolayer permeability to promote tumor
metastasis. J Exp Clin Cancer Res. 37(177)2018.PubMed/NCBI View Article : Google Scholar
|
|
28
|
Bai H, Lei K, Huang F, Jiang Z and Zhou X:
Exo-circRNAs: A new paradigm for anticancer therapy. Mol Cancer.
18(56)2019.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Yi Q, Yue J, Liu Y, Shi H and Sun W, Feng
J and Sun W: Recent advances of exosomal circRNAs in cancer and
their potential clinical applications. J Transl Med.
21(516)2023.PubMed/NCBI View Article : Google Scholar
|
|
30
|
Kim KM, Abdelmohsen K, Mustapic M,
Kapogiannis D and Gorospe M: RNA in extracellular vesicles. Wiley
Interdiscip Rev RNA. 8(10)2017.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Chen Y, Zhao Y, Yin Y, Jia X and Mao L:
Mechanism of cargo sorting into small extracellular vesicles.
Bioengineered. 12:8186–8201. 2021.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Qiu Y, Li P, Zhang Z and Wu M: Insights
into exosomal Non-coding RNAs sorting mechanism and clinical
application. Front Oncol. 11(664904)2021.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Fabbiano F, Corsi J, Gurrieri E, Trevisan
C, Notarangelo M and D'Agostino VG: RNA packaging into
extracellular vesicles: An orchestra of RNA-binding proteins? J
Extracell Vesicles. 10(e12043)2020.PubMed/NCBI View Article : Google Scholar
|
|
34
|
Liu S, Wan X, Gou Y, Yang W, Xu W, Du Y,
Peng X, Wang X and Zhang X: The emerging functions and clinical
implications of circRNAs in acute myeloid leukaemia. Cancer Cell
Int. 25(167)2025.PubMed/NCBI View Article : Google Scholar
|
|
35
|
Guo W, Gao Y, Li N, Shao F, Wang C, Wang
P, Yang Z, Li R and He J: Exosomes: New players in cancer (review).
Oncol Rep. 38:665–675. 2017.PubMed/NCBI View Article : Google Scholar
|
|
36
|
Głuszko A, Szczepański MJ, Ludwig N, Mirza
SM and Olejarz W: Exosomes in cancer: Circulating Immune-related
biomarkers. Biomed Res Int. 2019(1628029)2019.PubMed/NCBI View Article : Google Scholar
|
|
37
|
Zahra K, Dey T, Ashish Mishra SP and
Pandey U: Pyruvate kinase M2 and cancer: The role of PKM2 in
promoting tumorigenesis. Front Oncol. 10(159)2020.PubMed/NCBI View Article : Google Scholar
|
|
38
|
Escrevente C, Keller S, Altevogt P and
Costa J: Interaction and uptake of exosomes by ovarian cancer
cells. BMC Cancer. 11(108)2011.PubMed/NCBI View Article : Google Scholar
|
|
39
|
Obi P and Chen YG: The design and
synthesis of circular RNAs. Methods. 196:85–103. 2021.PubMed/NCBI View Article : Google Scholar
|
|
40
|
Zeng Y, Zou Y, Gao G, Zheng S, Wu S, Xie X
and Tang H: The biogenesis, function and clinical significance of
circular RNAs in breast cancer. Cancer Biol Med. 19:14–29.
2022.PubMed/NCBI View Article : Google Scholar
|
|
41
|
Zhang XO, Wang HB, Zhang Y, Lu X, Chen LL
and Yang L: Complementary sequence-mediated exon circularization.
Cell. 159:134–147. 2014.PubMed/NCBI View Article : Google Scholar
|
|
42
|
Li L, Wei C, Xie Y, Su Y, Liu C, Qiu G,
Liu W, Liang Y, Zhao X, Huang D and Wu D: Expanded insights into
the mechanisms of RNA-binding protein regulation of circRNA
generation and function in cancer biology and therapy. Genes Dis.
12(101383)2025.PubMed/NCBI View Article : Google Scholar
|
|
43
|
Xiong Y, Chen Y, Luo H, Shi M, Wang C, Hu
X and Zhu H: DHX9 as a prognostic biomarker and its biological
roles in acute myeloid leukemia. BMC Cancer.
25(1464)2025.PubMed/NCBI View Article : Google Scholar
|
|
44
|
Nong T, Mehra S and Taylor J: Common
driver mutations in AML: Biological impact, clinical
considerations, and treatment strategies. Cells.
13(1392)2024.PubMed/NCBI View Article : Google Scholar
|
|
45
|
Das A, Sinha T, Shyamal S and Panda AC:
Emerging role of circular RNA-protein interactions. Noncoding RNA.
7(48)2021.PubMed/NCBI View Article : Google Scholar
|
|
46
|
Huang Y, Zhang C, Xiong J and Ren H:
Emerging important roles of circRNAs in human cancer and other
diseases. Genes Dis. 8:412–423. 2020.PubMed/NCBI View Article : Google Scholar
|
|
47
|
Liu Y, Chen X, Liu J, Jin Y and Wang W:
Circular RNA circ_0004277 inhibits acute myeloid leukemia
progression through MicroRNA-134-5p/Single stranded DNA binding
protein 2. Bioengineered. 13:9662–9673. 2022.PubMed/NCBI View Article : Google Scholar
|
|
48
|
Ping L, Jian-Jun C, Chu-Shu L, Guang-Hua L
and Ming Z: High circ_100053 predicts a poor outcome for chronic
myeloid leukemia and is involved in imatinib resistance. Oncol Res:
Feb 14, 2019 doi: 10.3727/096504018X15412701483326 (Epub ahead of
print).
|
|
49
|
Ren X, Yan H, Zhou M, Wu F, Zhang Q, Ding
Y, Li Q, Wang H, Xu C, Wang J, et al: Vincristine overcomes
cytarabine resistance in acute myeloid leukemia by suppression of
ARHGAP18-mediated cellular senescence. Blood. 144(2754)2024.
|
|
50
|
Jamal M, Song T, Chen B, Faisal M, Hong Z,
Xie T, Wu Y, Pan S, Yin Q, Shao L and Zhang Q: Recent progress on
circular RNA research in acute myeloid leukemia. Front Oncol.
9(1108)2019.PubMed/NCBI View Article : Google Scholar
|
|
51
|
Rahmati A, Mafi A, Soleymani F, Babaei
Aghdam Z, Masihipour N, Ghezelbash B, Asemi R, Aschner M, Vakili O,
Homayoonfal M, et al: Circular RNAs: Pivotal role in the
leukemogenesis and novel indicators for the diagnosis and prognosis
of acute myeloid leukemia. Front Oncol. 13(1149187)2023.PubMed/NCBI View Article : Google Scholar
|
|
52
|
Gu A, Jaijyan DK, Yang S, Zeng M, Pei S
and Zhu H: Functions of circular RNA in human diseases and
illnesses. Noncoding RNA. 9(38)2023.PubMed/NCBI View Article : Google Scholar
|
|
53
|
Dong W, Bi J, Liu H, Yan D, He Q, Zhou Q,
Wang Q, Xie R, Su Y, Yang M, et al: Circular RNA ACVR2A suppresses
bladder cancer cells proliferation and metastasis through
miR-626/EYA4 axis. Mol Cancer. 18(95)2019.PubMed/NCBI View Article : Google Scholar
|
|
54
|
Meng Q, Li S, Liu Y, Zhang S, Jin J, Zhang
Y, Guo C, Liu B and Sun Y: Circular RNA circSCAF11 accelerates the
glioma tumorigenesis through the miR-421/SP1/VEGFA axis. Mol Ther
Nucleic Acids. 17:669–677. 2019.PubMed/NCBI View Article : Google Scholar
|
|
55
|
Ma P, Pan Y, Li W, Sun C, Liu J, Xu T and
Shu Y: Extracellular vesicles-mediated noncoding RNAs transfer in
cancer. J Hematol Oncol. 10(57)2017.PubMed/NCBI View Article : Google Scholar
|
|
56
|
Liu J, Chen W, Zhang H, Liu T and Zhao L:
miR-214 targets the PTEN-mediated PI3K/Akt signaling pathway and
regulates cell proliferation and apoptosis in ovarian cancer. Oncol
Lett. 14:5711–5718. 2017.PubMed/NCBI View Article : Google Scholar
|
|
57
|
Li X, Yin X, Bao H and Liu C: Circular RNA
ITCH increases sorafenib-sensitivity in hepatocellular carcinoma
via sequestering miR-20b-5p and modulating the downstream
PTEN-PI3K/Akt pathway. Mol Cell Probes. 67(101877)2023.PubMed/NCBI View Article : Google Scholar
|
|
58
|
Pimenta DB, Varela VA, Datoguia TS,
Caraciolo VB, Lopes GH and Pereira WO: The bone marrow
microenvironment mechanisms in acute myeloid leukemia. Front Cell
Dev Biol. 9(764698)2021.PubMed/NCBI View Article : Google Scholar
|
|
59
|
Bi J, Pu Y and Yu X: Exosomal circ_0004136
enhances the progression of pediatric acute myeloid leukemia
depending on the regulation of miR-570-3p/TSPAN3 axis. Anticancer
Drugs. 32:802–811. 2021.PubMed/NCBI View Article : Google Scholar
|
|
60
|
Zhou Q, Xie D, Wang R, Liu L, Yu Y, Tang
X, Hu Y and Cui D: The emerging landscape of exosomal CircRNAs in
solid cancers and hematological malignancies. Biomark Res.
10(28)2022.PubMed/NCBI View Article : Google Scholar
|
|
61
|
Wang D, Ming X, Xu J and Xiao Y:
Circ_0009910 shuttled by exosomes regulates proliferation, cell
cycle and apoptosis of acute myeloid leukemia cells by regulating
miR-5195-3p/GRB10 axis. Hematol Oncol. 39:390–400. 2021.PubMed/NCBI View Article : Google Scholar
|
|
62
|
Du A, Yang Q, Sun X and Zhao Q: Exosomal
circRNA-001264 promotes AML immunosuppression through induction of
M2-like macrophages and PD-L1 overexpression. Int Immunopharmacol.
124(110868)2023.PubMed/NCBI View Article : Google Scholar
|
|
63
|
Lyu T, Wang Y, Li D, Yang H, Qin B, Zhang
W, Li Z, Cheng C, Zhang B, Guo R and Song Y: Exosomes from BM-MSCs
promote acute myeloid leukemia cell proliferation, invasion and
chemoresistance via upregulation of S100A4. Exp Hematol Oncol.
10(24)2021.PubMed/NCBI View Article : Google Scholar
|
|
64
|
Fan H, Li Y, Liu C, Liu Y, Bai J and Li W:
Circular RNA-100290 promotes cell proliferation and inhibits
apoptosis in acute myeloid leukemia cells via sponging miR-203.
Biochem Biophys Res Commun. 507:178–184. 2018.PubMed/NCBI View Article : Google Scholar
|
|
65
|
Long F, Lin Z, Long Q, Lu Z, Zhu K, Zhao M
and Yang M: CircZBTB46 protects acute myeloid leukemia cells from
ferroptotic cell death by upregulating SCD. Cancers (Basel).
15(459)2023.PubMed/NCBI View Article : Google Scholar
|
|
66
|
Ye D, Gong M, Deng Y, Fang S, Cao Y, Xiang
Y and Shen Z: Roles and clinical application of exosomal circRNAs
in the diagnosis and treatment of malignant tumors. J Transl Med.
20(161)2022.PubMed/NCBI View Article : Google Scholar
|
|
67
|
McKiernan J, Donovan MJ, O'Neill V,
Bentink S, Noerholm M, Belzer S, Skog J, Kattan MW, Partin A,
Andriole G, et al: A novel urine exosome gene expression assay to
predict High-grade prostate cancer at initial biopsy. JAMA Oncol.
2:882–889. 2016.PubMed/NCBI View Article : Google Scholar
|
|
68
|
Dixon CL, Sheller-Miller S, Saade GR,
Fortunato SJ, Lai A, Palma C, Guanzon D, Salomon C and Menon R:
Amniotic fluid exosome proteomic profile exhibits unique pathways
of term and preterm labor. Endocrinology. 159:2229–2240.
2018.PubMed/NCBI View Article : Google Scholar
|
|
69
|
Tang BJ, Sun B, Chen L, Xiao J, Huang ST
and Xu P: The landscape of exosome-derived non-coding RNA in
leukemia. Front Pharmacol. 13(912303)2022.PubMed/NCBI View Article : Google Scholar
|
|
70
|
Das MK: Multicenter studies: Relevance,
design and implementation. Indian Pediatr. 59:571–579.
2022.PubMed/NCBI
|
|
71
|
Can C, Yang X, Jia H, Wu H, Guo X, Wei Y,
Jia Z, Liu W, Zhang A and He N: Exosomal circ_0006896 promotes AML
progression via interaction with HDAC1 and restriction of antitumor
immunity. Mol Cancer. 24(4)2025.PubMed/NCBI View Article : Google Scholar
|
|
72
|
Li J, Zhou W, Wang H, Huang M and Deng H:
Exosomal circular RNAs in tumor microenvironment: An emphasis on
signaling pathways and clinical opportunities. MedComm (2020).
5(e70019)2024.PubMed/NCBI View Article : Google Scholar
|
|
73
|
Xu Y, Kong S, Qin S, Shen X and Ju S:
Exosomal circRNAs: Sorting mechanisms, roles and clinical
applications in tumors. Front Cell Dev Biol.
8(581558)2020.PubMed/NCBI View Article : Google Scholar
|
|
74
|
Wang S, Dong Y, Gong A, Kong H, Gao J, Hao
X, Liu Y, Wang Z, Fan Y, Liu C and Xu W: Exosomal circRNAs as novel
cancer biomarkers: Challenges and opportunities. Int J Biol Sci.
17:562–573. 2021.PubMed/NCBI View Article : Google Scholar
|
|
75
|
Zhou M, Gao X, Zheng X and Luo J:
Functions and clinical significance of circular RNAs in acute
myeloid leukemia. Front Pharmacol. 13(1010579)2022.PubMed/NCBI View Article : Google Scholar
|
|
76
|
Liu W and Cheng F: Circular RNA circCRKL
inhibits the proliferation of acute myeloid leukemia cells via the
miR-196a-5p/miR-196b-5p/p27 axis. Bioengineered.
12(7704)2021.PubMed/NCBI View Article : Google Scholar
|
|
77
|
Motomura K, Kuwano A, Tanaka K, Koga Y,
Masumoto A and Yada M: Potential predictive biomarkers of systemic
drug therapy for hepatocellular carcinoma: Anticipated usefulness
in clinical practice. Cancers (Basel). 15(4345)2023.PubMed/NCBI View Article : Google Scholar
|
|
78
|
Ma Y, Niu X, Yan S, Liu Y, Dong R and Li
Y: Circular RNA profiling facilitates the diagnosis and prognostic
monitoring of breast cancer: A pair-wise meta-analysis. J Clin Lab
Anal. 35(e23575)2020.PubMed/NCBI View Article : Google Scholar
|
|
79
|
Ma J, Wen X, Xu Z, Xia P, Jin Y, Lin J and
Qian J: Predicting the influence of Circ_0059706 expression on
prognosis in patients with acute myeloid leukemia using classical
statistics and machine learning. Front Genet.
13(961142)2022.PubMed/NCBI View Article : Google Scholar
|
|
80
|
Zheng P, Gao H, Xie X and Lu P: Plasma
exosomal hsa_circ_0015286 as a potential diagnostic and prognostic
biomarker for gastric cancer. Pathol Oncol Res.
28(1610446)2022.PubMed/NCBI View Article : Google Scholar
|
|
81
|
Ding C, Yi X, Chen X, Wu Z, You H, Chen X,
Zhang G, Sun Y, Bu X, Wu X, et al: Warburg effect-promoted exosomal
circ_0072083 releasing up-regulates NANGO expression through
multiple pathways and enhances temozolomide resistance in glioma. J
Exp Clin Cancer Res. 40(164)2021.PubMed/NCBI View Article : Google Scholar
|
|
82
|
Weng Q, Chen M, Li M, Zheng YF, Shao G,
Fan W, Xu XM and Ji J: Global microarray profiling identified
hsa_circ_0064428 as a potential immune-associated prognosis
biomarker for hepatocellular carcinoma. J Med Genet. 56:32–38.
2019.PubMed/NCBI View Article : Google Scholar
|
|
83
|
Shang J, Chen WM, Liu S, Wang ZH, Wei TN,
Chen ZZ and Wu WB: CircPAN3 contributes to drug resistance in acute
myeloid leukemia through regulation of autophagy. Leuk Res.
85(106198)2019.PubMed/NCBI View Article : Google Scholar
|
|
84
|
Yan L and Yan Q: Serum circRNA_100199 is a
prognostic biomarker in acute myeloid leukemia. Int J Gen Med.
16:4661–4668. 2023.PubMed/NCBI View Article : Google Scholar
|
|
85
|
Zhang J, Zhang X, Li C, Yue L, Ding N,
Riordan T, Yang L, Li Y, Jen C, Lin S, et al: Circular RNA
profiling provides insights into their subcellular distribution and
molecular characteristics in HepG2 cells. RNA Biol. 16:220–232.
2019.PubMed/NCBI View Article : Google Scholar
|
|
86
|
Théry C, Witwer KW, Aikawa E, Alcaraz MJ,
Anderson JD, Andriantsitohaina R, Antoniou A, Arab T, Archer F,
Atkin-Smith GK, et al: Minimal information for studies of
extracellular vesicles 2018 (MISEV2018): A position statement of
the international society for extracellular vesicles and update of
the MISEV2014 guidelines. J Extracell Vesicles.
7(1535750)2018.PubMed/NCBI View Article : Google Scholar
|
|
87
|
Ghaffari K, Moradi-Hasanabad A,
Sobhani-Nasab A, Javaheri J and Ghasemi A: Application of
cell-derived exosomes in the hematological malignancies therapy.
Front Pharmacol. 14(1263834)2023.PubMed/NCBI View Article : Google Scholar
|
|
88
|
Amiri Yekta A, Dalman A, Sanati MH, Fatemi
N, Vazirinasab H, Zomorodipour A, Chehrazi M and Gourabi H:
Optimization of the electroporation conditions for transfection of
human factor IX into the goat fetal fibroblasts. Cell J.
14(270)2013.PubMed/NCBI
|
|
89
|
Pomatto MAC, Bussolati B, D'Antico S,
Ghiotto S, Tetta C, Brizzi MF and Camussi G: Improved loading of
Plasma-derived extracellular vesicles to encapsulate antitumor
miRNAs. Mol Ther Methods Clin Dev. 13:133–144. 2019.PubMed/NCBI View Article : Google Scholar
|
|
90
|
Amin AH, Sharifi LMA, Kakhharov AJ,
Opulencia MJC, Alsaikhan F, Bokov DO, Majdi HS, Jawad MA, Hammid
AT, Shalaby MN, et al: Role of acute myeloid leukemia (AML)-derived
exosomes in tumor progression and survival. Biomed Pharmacother.
150(113009)2022.PubMed/NCBI View Article : Google Scholar
|
|
91
|
Chen C, Wang J, Sun M, Li J and Wang HMD:
Toward the next-generation phyto-nanomedicines: Cell-derived
nanovesicles (CDNs) for natural product delivery. Biomed
Pharmacother. 145(112416)2022.PubMed/NCBI View Article : Google Scholar
|
|
92
|
Wang W, Wu X, Zheng J, Yin R, Li Y, Wu X,
Xu L and Jin Z: Utilizing exosomes as sparking clinical biomarkers
and therapeutic response in acute myeloid leukemia. Front Immunol.
14(1315453)2024.PubMed/NCBI View Article : Google Scholar
|
|
93
|
Chen H, Jayasinghe MK, Yeo EYM, Wu Z,
Pirisinu M, Usman WM, Pham TT, Lim KW, Tran NV, Leung AYH, et al:
CD33-targeting extracellular vesicles deliver antisense
oligonucleotides against FLT3-ITD and miR-125b for specific
treatment of acute myeloid leukaemia. Cell Prolif.
55(e13255)2022.PubMed/NCBI View Article : Google Scholar
|
|
94
|
Nehrbas J, Butler JT, Chen DW and Kurre P:
Extracellular vesicles and chemotherapy resistance in the AML
microenvironment. Front Oncol. 10(90)2020.PubMed/NCBI View Article : Google Scholar
|
|
95
|
Hong CS, Muller L, Boyiadzis M and
Whiteside TL: Isolation and characterization of CD34+ blast-derived
exosomes in acute myeloid leukemia. PLoS One.
9(e103310)2014.PubMed/NCBI View Article : Google Scholar
|
|
96
|
Percival ME, Lai C, Estey E and Hourigan
CS: Bone marrow evaluation for diagnosis and monitoring of acute
myeloid leukemia. Blood Rev. 31:185–192. 2017.PubMed/NCBI View Article : Google Scholar
|
|
97
|
Genç EE, Saraç İS, Arslan H and Eşkazan
AE: Diagnostic and treatment obstacles in acute myeloid leukemia:
Social, operational, and financial. Oncol Ther. 11:145–152.
2023.PubMed/NCBI View Article : Google Scholar
|
|
98
|
Shimony S, Stahl M and Stone RM: Acute
myeloid leukemia: 2025 update on diagnosis, Risk-stratification,
and management. Am J Hematol. 100:860–891. 2025.PubMed/NCBI View Article : Google Scholar
|
|
99
|
Floren M and Gillette JM: Acute myeloid
leukemia: Therapy resistance and a potential role for tetraspanin
membrane scaffolds. Int J Biochem Cell Biol.
137(106029)2021.PubMed/NCBI View Article : Google Scholar
|
|
100
|
Fianchi L, Quattrone M, Criscuolo M,
Bellesi S, Dragonetti G, Maraglino AME, Bonanni M, Chiusolo P, Sica
S and Pagano L: Extramedullary involvement in acute myeloid
leukemia. A single center ten years' Experience. Mediterr J Hematol
Infect Dis. 13(e2021030)2021.PubMed/NCBI View Article : Google Scholar
|
|
101
|
Döhner H, Wei AH, Appelbaum FR, Craddock
C, DiNardo CD, Dombret H, Ebert BL, Fenaux P, Godley LA, Hasserjian
RP, et al: Diagnosis and management of AML in adults: 2022
recommendations from an international expert panel on behalf of the
ELN. Blood. 140:1345–1377. 2022.PubMed/NCBI View Article : Google Scholar
|
|
102
|
Leotta S, Condorelli A, Sciortino R,
Milone GA, Bellofiore C, Garibaldi B, Schininà G, Spadaro A, Cupri
A and Milone G: Prevention and treatment of acute myeloid leukemia
relapse after hematopoietic stem cell transplantation: The state of
the art and future perspectives. J Clin Med. 11(253)2022.PubMed/NCBI View Article : Google Scholar
|
|
103
|
Li J, Wang J and Chen Z: Emerging role of
exosomes in cancer therapy: Progress and challenges. Mol Cancer.
24(13)2025.PubMed/NCBI View Article : Google Scholar
|
|
104
|
Kim MS, Haney MJ, Zhao Y, Yuan D, Deygen
I, Klyachko NL, Kabanov AV and Batrakova EV: Engineering
macrophage-derived exosomes for targeted paclitaxel delivery to
pulmonary metastases: In vitro and in vivo evaluations.
Nanomedicine. 14:195–204. 2018.PubMed/NCBI View Article : Google Scholar
|
|
105
|
Aqil F, Munagala R, Jeyabalan J, Agrawal
AK, Kyakulaga AH, Wilcher SA and Gupta RC: Milk exosomes-Natural
nanoparticles for siRNA delivery. Cancer Lett. 449:186–195.
2019.PubMed/NCBI View Article : Google Scholar
|
|
106
|
Usman WM, Pham TC, Kwok YY, Vu LT, Ma V,
Peng B, Chan YS, Wei L, Chin SM, Azad A, et al: Efficient RNA drug
delivery using red blood cell extracellular vesicles. Nat Commun.
9(2359)2018.PubMed/NCBI View Article : Google Scholar
|
|
107
|
Kamerkar S, LeBleu VS, Sugimoto H, Yang S,
Ruivo CF, Melo SA, Lee JJ and Kalluri R: Exosomes facilitate
therapeutic targeting of oncogenic kras in pancreatic cancer.
Nature. 546:498–503. 2017.PubMed/NCBI View Article : Google Scholar
|
|
108
|
Kim SM, Yang Y, Oh SJ, Hong Y, Seo M and
Jang M: Cancer-derived exosomes as a delivery platform of
CRISPR/Cas9 confer cancer cell tropism-dependent targeting. J
Control Release. 266:8–16. 2017.PubMed/NCBI View Article : Google Scholar
|
|
109
|
Qi H, Liu C, Long L, Ren Y, Zhang S, Chang
X, Qian X, Jia H, Zhao J, Sun J, et al: Blood Exosomes endowed with
magnetic and targeting properties for cancer therapy. ACS Nano.
10:3323–3333. 2016.PubMed/NCBI View Article : Google Scholar
|
|
110
|
Liu X, Zhang Y, Zhou S, Dain L, Mei L and
Zhu G: Circular RNA: An emerging frontier in RNA therapeutic
targets, RNA therapeutics, and mRNA vaccines. J Control Release.
348:84–94. 2022.PubMed/NCBI View Article : Google Scholar
|
|
111
|
Wang H, Xiao Y, Wu L and Ma D:
Comprehensive circular RNA profiling reveals the regulatory role of
the circRNA-000911/miR-449a pathway in breast carcinogenesis. Int J
Oncol. 52:743–754. 2018.PubMed/NCBI View Article : Google Scholar
|
|
112
|
Zhou Y, Zheng X, Xu B, Chen L, Wang Q,
Deng H and Jiang J: Circular RNA hsa_circ_0004015 regulates the
proliferation, invasion, and TKI drug resistance of non-small cell
lung cancer by miR-1183/PDPK1 signaling pathway. Biochem Biophys
Res Commun. 508:527–535. 2019.PubMed/NCBI View Article : Google Scholar
|
|
113
|
Zhang F, Jiang J, Qian H, Yan Y and Xu W:
Exosomal circRNA: Emerging insights into cancer progression and
clinical application potential. J Hematol Oncol.
16(67)2023.PubMed/NCBI View Article : Google Scholar
|
|
114
|
Xie R, Zhang Y, Zhang J, Li J and Zhou X:
The role of circular RNAs in Immune-related diseases. Front
Immunol. 11(545)2020.PubMed/NCBI View Article : Google Scholar
|
|
115
|
Patnaik E, Madu C and Lu Y: Epigenetic
modulators as therapeutic agents in cancer. Int J Mol Sci.
24(14964)2023.PubMed/NCBI View Article : Google Scholar
|
|
116
|
Zhang J, Gu Y and Chen B: Mechanisms of
drug resistance in acute myeloid leukemia. Onco Targets Ther.
12:1937–1945. 2019.PubMed/NCBI View Article : Google Scholar
|
|
117
|
Han B, Chao J and Yao H: Circular RNA and
its mechanisms in disease: From the bench to the clinic. Pharmacol
Ther. 187:31–44. 2018.PubMed/NCBI View Article : Google Scholar
|
|
118
|
Sun H, Xie Y, Wu X, Hu W, Chen X, Wu K,
Wang H, Zhao S, Shi Q, Wang X, et al: circRNAs as prognostic
markers in pediatric acute myeloid leukemia. Cancer Lett.
591(216880)2024.PubMed/NCBI View Article : Google Scholar
|
|
119
|
Guo S, Li B, Chen Y, Zou D, Yang S, Zhang
Y, Wu N, Sheng L, Huang H, Ouyang G and Mu Q: Hsa_circ_0012152 and
Hsa_circ_0001857 accurately discriminate acute lymphoblastic
leukemia from acute myeloid leukemia. Front Oncol.
10(1655)2020.PubMed/NCBI View Article : Google Scholar
|
|
120
|
Hu L, Zheng B, Yang Y, Chen C and Hu M:
Construction of circRNA-miRNA-mRNA network Reveal functional
circRNAs and key genes in acute myeloid leukemia. Int J Gen Med.
16:1491–1504. 2023.PubMed/NCBI View Article : Google Scholar
|
|
121
|
Li Q, Ren X, Wang Y and Xin X: CircRNA: A
rising star in leukemia. PeerJ. 11(e15577)2023.PubMed/NCBI View Article : Google Scholar
|