Introduction
Human adipose tissue-derived stem cells (hASCs) are
an attractive cell type for tissue engineering which may be
harvested by direct excision or liposuction from human adipose
tissue. Physiologically, hASCs are capable of differentiating into
various lineages, such as adipocytes, osteoblasts, myocytes and
chondrocytes (1,2). The ability of hASCs to undergo
multilineage differentiation has attracted increasing interest in
their use clinically and in regenerative medicine (3). A number of studies have suggested
that hASCs possess significant potential for tissue rescue in
multiple animal models, including heart failure, myocardial
infarction, bone formation and wound healing, by differentiating
into a variety of lineages (4–6).
Many factors have been reported to be involved in
the mechanisms of hASC differentiation. Nutritional and hormonal
signaling affects hASC differentiation in a negative or a positive
manner, and the molecules involved in cell-matrix or cell-cell
interactions play key roles in regulating the differentiation
process (7–9). It is well known that fibroblast
growth factor 2 (FGF2) inhibits the osteogenic
differentiation of hASCs whereas it promotes chondrogenesis
(10,11). Moreover, microRNA (miRNA or
miR)-26a has been shown to modulate the late stage of osteoblast
differentiation by targeting the transcription factor (TF) SMAD
family member 1 (SMAD1) (4). The
upregulation of miRNA-22 has been proved to promote the osteogenic
differentiation of human adipose tissue-derived mesenchymal stem
cells by suppressing histone deacetylase 6 (HDAC6)
expression (12). Furthermore,
hASCs are capable of differentiating into skeletal myocytes and
cardiomyocytes under specific conditions (incubation in myogenic
medium) (13,14). In vitro,
sphingosylphosphorylcholine and transforming growth factor-β
(TGF-β) induced the expression of smooth muscle-associated markers
including α-smooth muscle actin, calponin and SM22 in hASCs
(15,16). Numerous studies have been
performed to reveal the molecular mechanisms controlling the
differentiation of hASCs (7–16).
However, the mechanisms responsible for the regulation of myocyte
and osteocyte differentiation remain largely unknown.
Increasing evidence has proved that the conversion
of hASCs into differentiated myocytes and osteocytes involves
changes in gene expression which are mainly regulated by miRNAs and
TFs (17,18). For instance, Luzi et al
(4) showed that miR-26a
expression was increased during hASC differentiation, whereas the
expression of SMAD1 was complementary to that of miR-26a. In
addition, Kim et al (17)
reported that miR-196a regulates the differentiation and
proliferation of hASCs by modulating the levels of the HOXC8
transcription factor.
To gain further insight into the molecular
mechanisms responsible for the differentiation of hASCs into
myocytes and osteocytes, we re-analyzed the microarray data
GSE37329 through the identification of differentially expressed
genes (DEGs) in hASC-derived myocytes and osteocytes compared with
hASCs, as well as through functional annotation and protein-protein
interaction (PPI) network construction. Furthermore, TFs and miRNAs
targeting the DEGs were predicted and functionally analyzed.
Materials and methods
Gene datasets
The gene expression profile of GSE37329 was
retrieved from the Gene Expression Omnibus (GEO) database available
at http://www.ncbi.nlm.nih.gov/geo/
(19). This dataset was deposited
by Berdasco et al (19) on
October 3, 2013 and was based on GPL11532 platform (Affymetrix
Human Gene 1.1 ST array, Santa Clara, CA, USA). A total of 7
samples were available for further study, including three hASC cell
lines from healthy donors, two osteogenic lineages and two myogenic
lineages which were all obtained through the in vitro
induction of hASCs.
Data preprocessing
The raw expression data (Affymetrix CEL files) were
firstly preprocessed by the Robust Multiarray Average (RMA)
normalization approach of Bioconductor affy package in R (20) (http://www.bioconductor.org), which returned the
expression signals of each probe as log 2 scale. When different
probes were mapped to the same gene, the mean value of the probes
was considered as the gene value. Subsequently, the probe serial
numbers in the matrix were transformed into gene names using the
platform R/Bioconductor note package of the dataset chip. The
matrix consisting of 20,253 genes was finally acquired.
Screening of DEGs
To screen out the DEGs in the in
vitro-obtained osteogenic and myogenic lineages derived from
hASCs compared with the freshly isolated hASCs obtained from
healthy donors, respectively, Linear Models for Microarray Data
(Limma) package of Bioconductor (21) was applied in the comparisons
(osteogenic lineages vs. hASCs and myogenic lineages vs. hASCs).
Unadjusted P-values were calculated using the Student's t-test.
Genes with P<0.05 and log 2|FC (fold change)| ≥1 were considered
to be differentially expressed. Hierarchical cluster analysis with
the eligible DEGs was then performed in order to identify clusters
of samples and genes.
Functional annotation of the DEGs
Functional enrichment of the two sets of DEGs in the
osteogenic and the myogenic lineages in vitro-induced from
the hASCs was assessed based on the biological process (BP)
category in Gene Ontology (GO) (22) and Kyoto Encyclopedia of Genes and
Genomes (KEGG) annotation terms (23). GO and KEGG signaling pathway
analyses were performed using the GO Function package (version
1.14.0) in Bioconductor (http://www.bioconductor.org/packages/release/bioc/html/GOFunction.html)
(24), which conducted the
standard hypergeometric test. A P-value <0.05 was considered to
indicate a statistically significant difference.
PPI network construction
Search Tool for the Retrieval of Interacting Genes
(STRING; http://string-db.org/) is an online
database which is comprised of more than 1,100 completely sequenced
organisms and includes experimental as well as predicted
interaction information (25).
The up- and down-regulated genes in both sets of DEGs verified
above were directly mapped to the STRING database in order to
acquire significant PPI pairs which were previously verified by
experiments, text mining and/or co-expressed analysis,
respectively. Notable PPI pairs in which both of the genes were
differentially expressed and the medium confidence was ≥0.4 were
integrated to construct a PPI network. The network was visualized
using CytoScape (26), available
at http://www.cytoscape.org. Considering
the complexity of PPI networks, we computed the degree of each node
by measuring the numbers of links of the node in the network.
Computational identification of TFs
To determine the common mechanism responsible for
the differentiation of hASCs into myocytes and osteocytes, DEGs
shared in the osteogenic and the myogenic lineages were screened
out. KEGG pathway enrichment analysis of the shared up- and
downregulated genes was performed, respectively. P-values were
calculated using hypergeometric distribution and a P-value <0.05
was considered to indicate a significant pathway.
To further explore the molecular mechanism,
eukaryotic TFs for the shared and unshared DEGs in osteogenic and
myogenic lineages were collected based on the the Encyclopedia of
DNA Elements (ENCODE) data from the USCS Genome Browser (27) available at http://genome.ucsc.edu/. P-values were calculated
using Fisher's exact test and adjusted using the Benjamini and
Hochberg method to define the false discovery rate (FDR). Only the
results with an FDR <5.5 E-06 were considered to be
significant.
miRNAs-target gene interaction network
construction
To better understand the function of miRNAs in
regulating the differentiation of hASCs, miRNAs targeting the
shared up- and downregulated DEGs screened above were predicted
using the miRecords database (28) available at http://c1.accurascience.com/miRecords/ and the miRWalk
database (29) available at
http://zmf.umm.uni-heidelberg.de/apps/zmf/mirwalk2/.
The miRNA-target interactions that were presented in miRecords
and/or miRWalk and verified by experiment were used for the
construction of the miRNA-mRNA interaction network. The network was
visualized using CytoScape and the degree of each miRNA node was
also measured. Furthermore, the predicted miRNAs were annotated
with BP terms in the GO database. P-values were calculated using
hypergeometric distribution and GO terms with a P-value <0.05
were defined as significantly enriched.
Results
Screening of DEGs
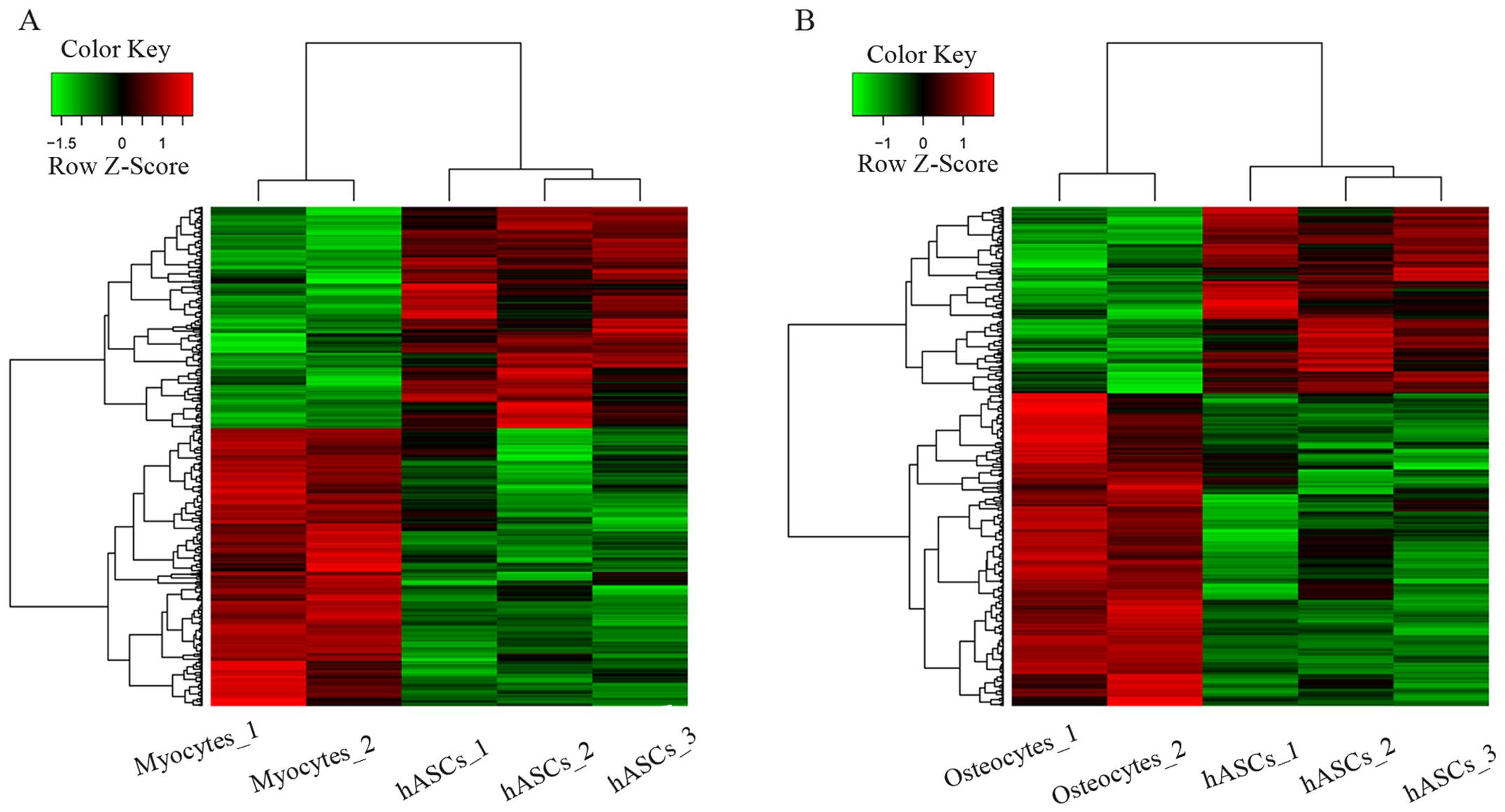
Compared with the hASCs, 665 DEGs in myogenic
lineages (370 up- and 295 downregulated genes) and 485 DEGs in
osteogenic lineages (304 up- and 181 downregulated genes) were
finally identified. The two sets of eligible DEGs were evaluated
using unsupervised hierarchical clustering. As shown in Fig. 1, DEGs were found in different
samples.
Annotating the biological functions of
DEGs
To elucidate the functions of DEGs, the up- and
downregulated genes in the in vitro-obtained myogenic and
osteogenic lineages were mapped to BP terms in the GO database, and
the top 10 GO terms are shown in Tables I and II, respectively. Briefly, the
upregulated genes identified from the myogenic lineages were mainly
involved in the regulation of multicellular organismal processes,
inflammatory responses and cellular responses to chemical stimuli,
whereas the downregulated genes were mainly involved in the
regulation of multicellular organismal processes,
single-multicellular organism processes, single-organism
developmental processes and multicellular organismal development.
On the other hand, the upregulated genes in the osteogenic lineages
were mainly associated with responses to stimuli, regulation of
multicellular organismal processes and regulation of localization,
whereas the downregulated genes were mainly associated with
anatomical structure development, system development and tissue
development.
 | Table ITop 10 enriched GO terms in the BP
category for both upregulated and downregulated differentially
expressed genes in myocytes. |
Table I
Top 10 enriched GO terms in the BP
category for both upregulated and downregulated differentially
expressed genes in myocytes.
| | GO ID | Name of BP | Count | P-value |
|---|
| Up | BP | GO:0051239 | Regulation of
multicellular organismal process | 86 | 2.28E-11 |
| | GO:0006954 | Inflammatory
response | 35 | 2.20E-09 |
| | GO:0070887 | Cellular response
to chemical stimulus | 83 | 2.25E-09 |
| | GO:0042221 | Response to
chemical | 115 | 2.87E-08 |
| | GO:0050896 | Response to
stimulus | 198 | 6.91E-08 |
| | GO:0032879 | Regulation of
localization | 70 | 7.58E-08 |
| | GO:0050727 | Regulation of
inflammatory response | 20 | 7.83E-08 |
| | GO:0006805 | Xenobiotic
metabolic process | 16 | 7.94E-08 |
| | GO:0050793 | Regulation of
developmental process | 67 | 8.64E-08 |
| | GO:0071466 | Cellular response
to xenobiotic stimulus | 16 | 8.68E-08 |
| Down | BP | GO:0001944 | Vasculature
development | 43 | 0 |
| | GO:0007275 | Multicellular
organismal development | 147 | 0 |
| | GO:0009653 | Anatomical
structure morphogenesis | 104 | 0 |
| | GO:0009888 | Tissue
development | 75 | 0 |
| | GO:0030154 | Cell
differentiation | 118 | 0 |
| | GO:0032501 | Multicellular
organismal process | 173 | 0 |
| | GO:0032502 | Developmental
process | 156 | 0 |
| | GO:0044707 |
Single-multicellular organism process | 172 | 0 |
| | GO:0044767 | Single-organism
developmental process | 153 | 0 |
| | GO:0048731 | System
development | 139 | 0 |
 | Table IITop 10 enriched GO terms in the BP
category for both upregulated and downregulated differentially
expressed genes in osteocytes. |
Table II
Top 10 enriched GO terms in the BP
category for both upregulated and downregulated differentially
expressed genes in osteocytes.
| | GO ID | Name of BP | Count | P-value |
|---|
| Up | BP | GO:0050896 | Response to
stimulus | 174 | 2.59E-10 |
| | GO:0006805 | Xenobiotic
metabolic process | 15 | 3.63E-08 |
| | GO:0071466 | Cellular response
to xenobiotic stimulus | 15 | 3.96E-08 |
| | GO:0032879 | Regulation of
localization | 61 | 5.48E-08 |
| | GO:0009410 | Response to
xenobiotic stimulus | 15 | 6.02E-08 |
| | GO:0051239 | Regulation of
multicellular organismal process | 64 | 4.22E-07 |
| | GO:0051049 | Regulation of
transport | 48 | 4.74E-07 |
| | GO:0006954 | Inflammatory
response | 27 | 5.39E-07 |
| | GO:0051046 | Regulation of
secretion | 26 | 7.18E-07 |
| | GO:1901700 | Response to
oxygen-containing compound | 42 | 1.88E-06 |
| Down | BP | GO:0072358 | Cardiovascular
system development | 33 | 4.90E-12 |
| | GO:0072359 | Circulatory system
development | 33 | 4.90E-12 |
| | GO:0014706 | Striated muscle
tissue development | 20 | 3.27E-11 |
| | GO:0060537 | Muscle tissue
development | 20 | 6.71E-11 |
| | GO:0048731 | System
development | 74 | 1.71E-10 |
| | GO:0001944 | Vasculature
development | 24 | 9.03E-10 |
| | GO:0048856 | Anatomical
structure development | 80 | 1.07E-09 |
| | GO:0009888 | Tissue
development | 42 | 3.06E-09 |
| | GO:2000026 | Regulation of
multicellular organismal development | 37 | 3.14E-09 |
| | GO:0009653 | Anatomical
structure morphogenesis | 51 | 4.97E-09 |
KEGG pathway enrichment analysis was used to further
understand the biological functions of the DEGs. Analysis of the
myogenic lineages revealed that the upregulated genes mainly
participated in neuroactive ligand-receptor interactions and drug
metabolism-cytochrome P450 pathways (Table III), which was the same as the
upregulated genes in the osteogenic lineages (Table IV). By contrast, the
downregulated genes in the myogenic lineages were mainly enriched
in pathways in cancer, ECM-receptor interactions and focal adhesion
(Table III), while the
downregulated genes in the osteogenic lineages were mainly involved
in the TGF-β signaling pathway and pathways in cancer (Table IV).
 | Table IIITop 10 enriched KEGG pathways of
upregulated and downregulated differentially expressed genes in
myocytes. |
Table III
Top 10 enriched KEGG pathways of
upregulated and downregulated differentially expressed genes in
myocytes.
| KEGG ID | Name | Count | P-value |
|---|
| Up | 00982 | Drug metabolism -
cytochrome P450 | 9 | 2.36E-05 |
| 00350 | Tyrosine
metabolism | 5 | 0.001759753 |
| 05145 | Toxoplasmosis | 8 | 0.007544112 |
| 00460 | Cyanoamino acid
metabolism | 2 | 0.009086159 |
| 00071 | Fatty acid
metabolism | 4 | 0.013452544 |
| 05014 | Amyotrophic lateral
sclerosis (ALS) | 4 | 0.027079145 |
| 04080 | Neuroactive
ligand-receptor interaction | 11 | 0.032800823 |
| 00603 | Glycosphingolipid
biosynthesis - globo series | 2 | 0.035670336 |
| 00590 | Arachidonic acid
metabolism | 4 | 0.038155628 |
| 00120 | Primary bile acid
biosynthesis | 2 | 0.045739216 |
| Down | 04512 | ECM-receptor
interaction | 12 | 5.83E-08 |
| 04350 | TGF-β signaling
pathway | 11 | 4.72E-07 |
| 05323 | Rheumatoid
arthritis | 10 | 8.16E-06 |
| 04510 | Focal adhesion | 14 | 2.53E-05 |
| 04640 | Hematopoietic cell
lineage | 9 | 4.23E-05 |
| 05200 | Pathways in
cancer | 18 | 4.23E-05 |
| 04514 | Cell adhesion
molecules (CAMs) | 10 | 0.000217686 |
| 05144 | Malaria | 6 | 0.000396575 |
| 05412 | Arrhythmogenic
right ventricular cardiomyopathy (ARVC) | 7 | 0.000505663 |
| 05217 | Basal cell
carcinoma | 6 | 0.000599721 |
 | Table IVTop 10 enriched KEGG pathways of
upregulated and downregulated differentially expressed genes in
osteocytes. |
Table IV
Top 10 enriched KEGG pathways of
upregulated and downregulated differentially expressed genes in
osteocytes.
| KEGG ID | Name | Count | P-value |
|---|
| Up | 00982 | Drug metabolism -
cytochrome P450 | 10 | 5.30E-07 |
| 00350 | Tyrosine
metabolism | 6 | 7.68E-05 |
| 04080 | Neuroactive
ligand-receptor interaction | 14 | 0.000309764 |
| 04270 | Vascular smooth
muscle contraction | 8 | 0.001033472 |
| 00460 | Cyanoamino acid
metabolism | 2 | 0.006279036 |
| 00071 | Fatty acid
metabolism | 4 | 0.006987698 |
| 00260 | Glycine, serine and
threonine metabolism | 3 | 0.018947162 |
| 00590 | Arachidonic acid
metabolism | 4 | 0.020746069 |
| 00603 | Glycosphingolipid
biosynthesis - globo series | 2 | 0.025079512 |
| 00010 |
Glycolysis/gluconeogenesis | 4 | 0.028465778 |
| Down | 04350 | TGF-β signaling
pathway | 7 | 2.00E-05 |
| 04610 | Complement and
coagulation cascades | 4 | 0.005201809 |
| 05217 | Basal cell
carcinoma | 3 | 0.018290654 |
| 04916 | Melanogenesis | 4 | 0.01930188 |
| 04972 | Pancreatic
secretion | 4 | 0.01930188 |
| 04710 | Circadian rhythm -
mammal | 2 | 0.020821702 |
| 00512 | Mucin type O-Glycan
biosynthesis | 2 | 0.037222845 |
| 04360 | Axon guidance | 4 | 0.042211972 |
| 05200 | Pathways in
cancer | 7 | 0.04712619 |
| 05020 | Prion diseases | 2 | 0.049293741 |
PPI network construction
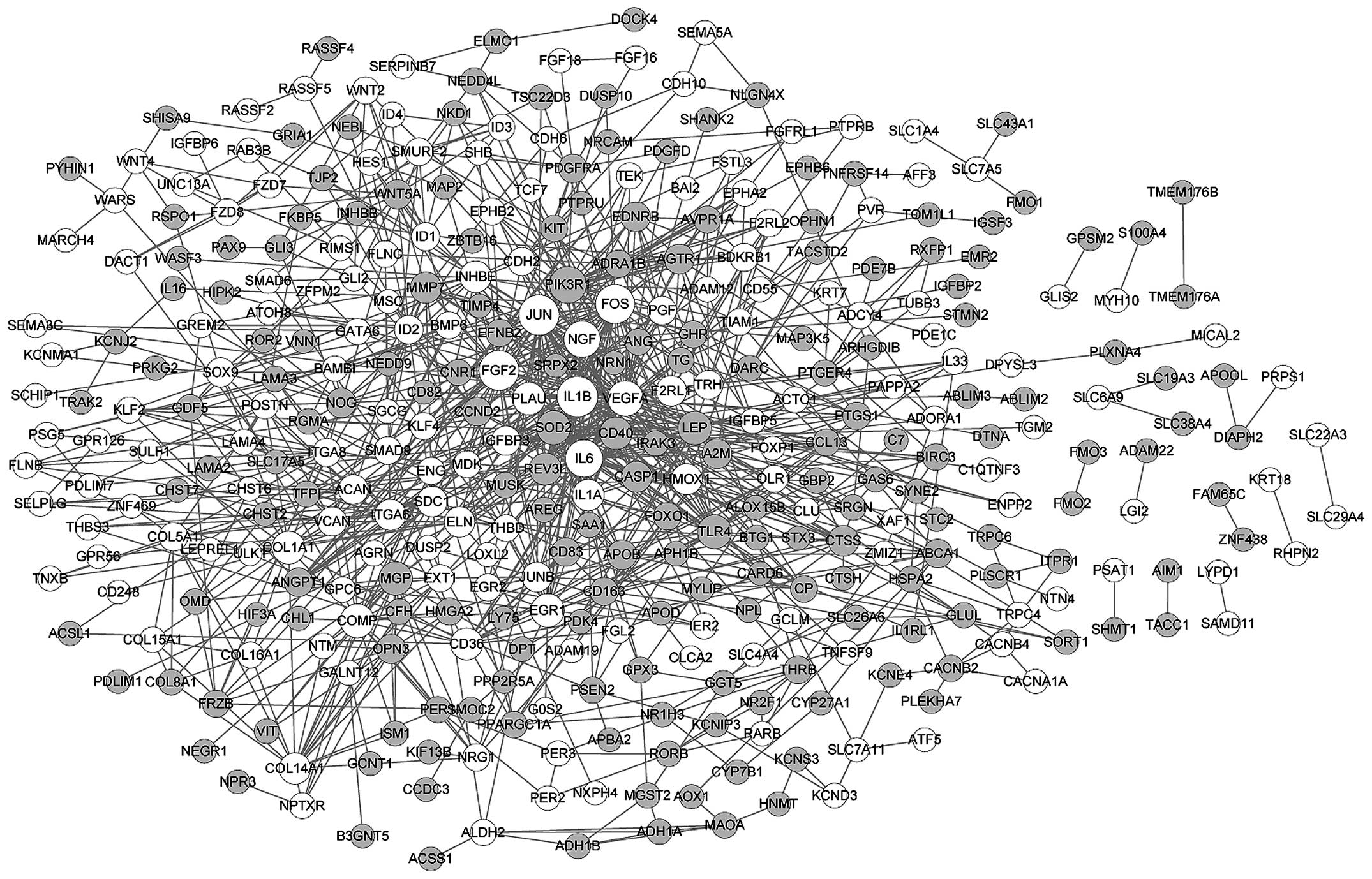
There were 363 nodes and 996 edges in the PPI
network of DEGs in myogenic lineages (Fig. 2). Based on the number of links,
the top 8 nodes were identified as vascular endothelial growth
factor A (VEGFA; degree, 57), interleukin (IL)6 (degree, 49), FBJ
murine osteosarcoma viral oncogene homolog (FOS; degree, 41), FGF2
(degree, 37), jun proto-oncogene (JUN; degree, 35), IL1B (degree,
34), phosphoinositide-3-kinase, regulatory subunit 1 (PIK3R1;
degree, 28) and nerve growth factor (NGF; degree, 27). In addition,
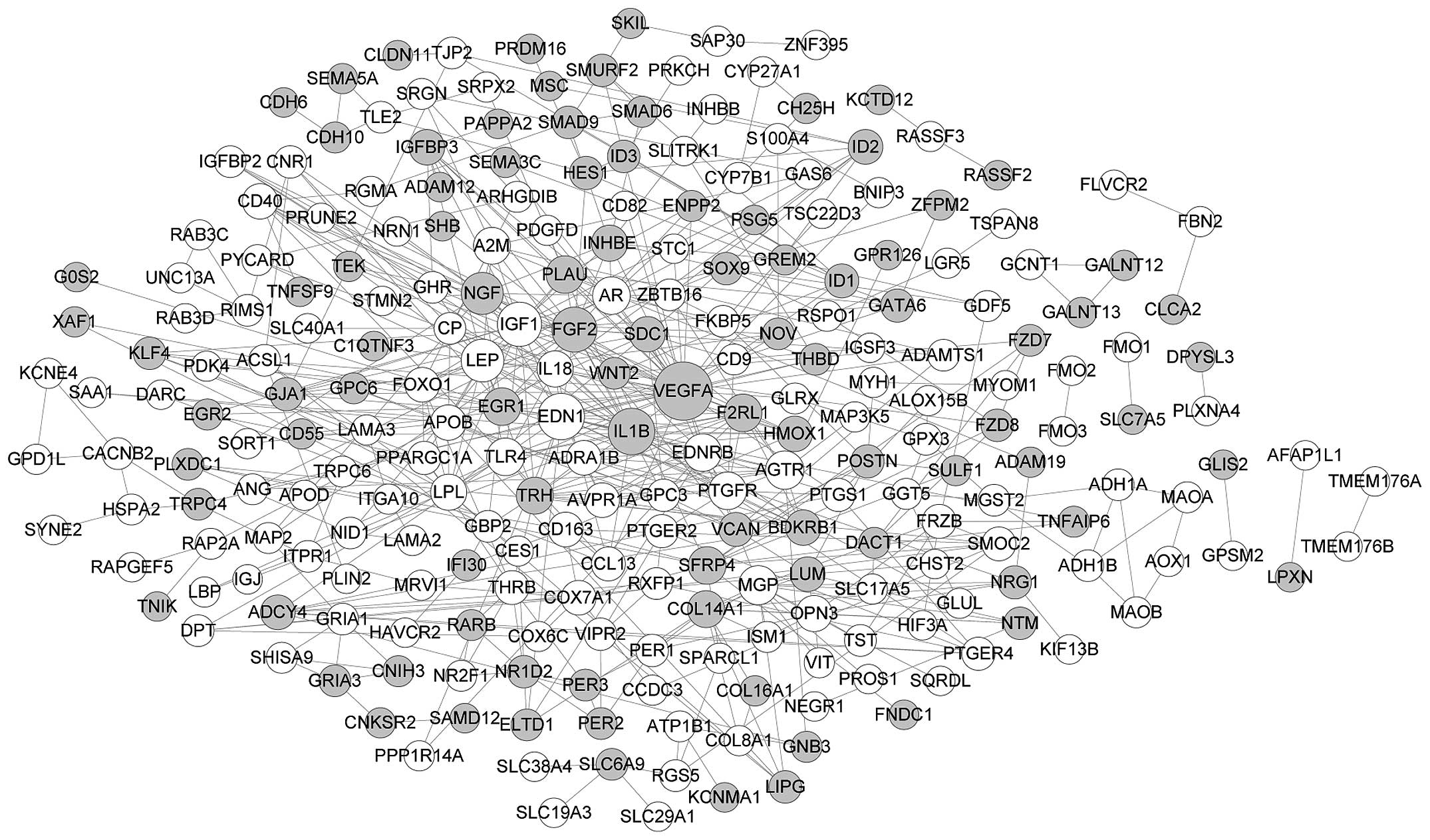
246 nodes and 520 edges constructed the PPI network of DEGs in the
osteogenic lineages (Fig. 3), and
the top 8 nodes were VEGFA (degree, 40), endothelin 1 (EDN1;
degree, 24), IL1B (degree, 24), FGF2 (degree, 22), insulin-like
growth factor 1 (IGF1; degree, 21), leptin (LEP; degree, 19), NGF
(degree, 18) and matrix Gla protein (MGP; degree, 14). Considering
the higher degree of VEGFA, IL1B, FGF2 and NGF in both networks, we
hypothesized that these four genes play similar roles in the
differentiation of hASCs into the two cell types.
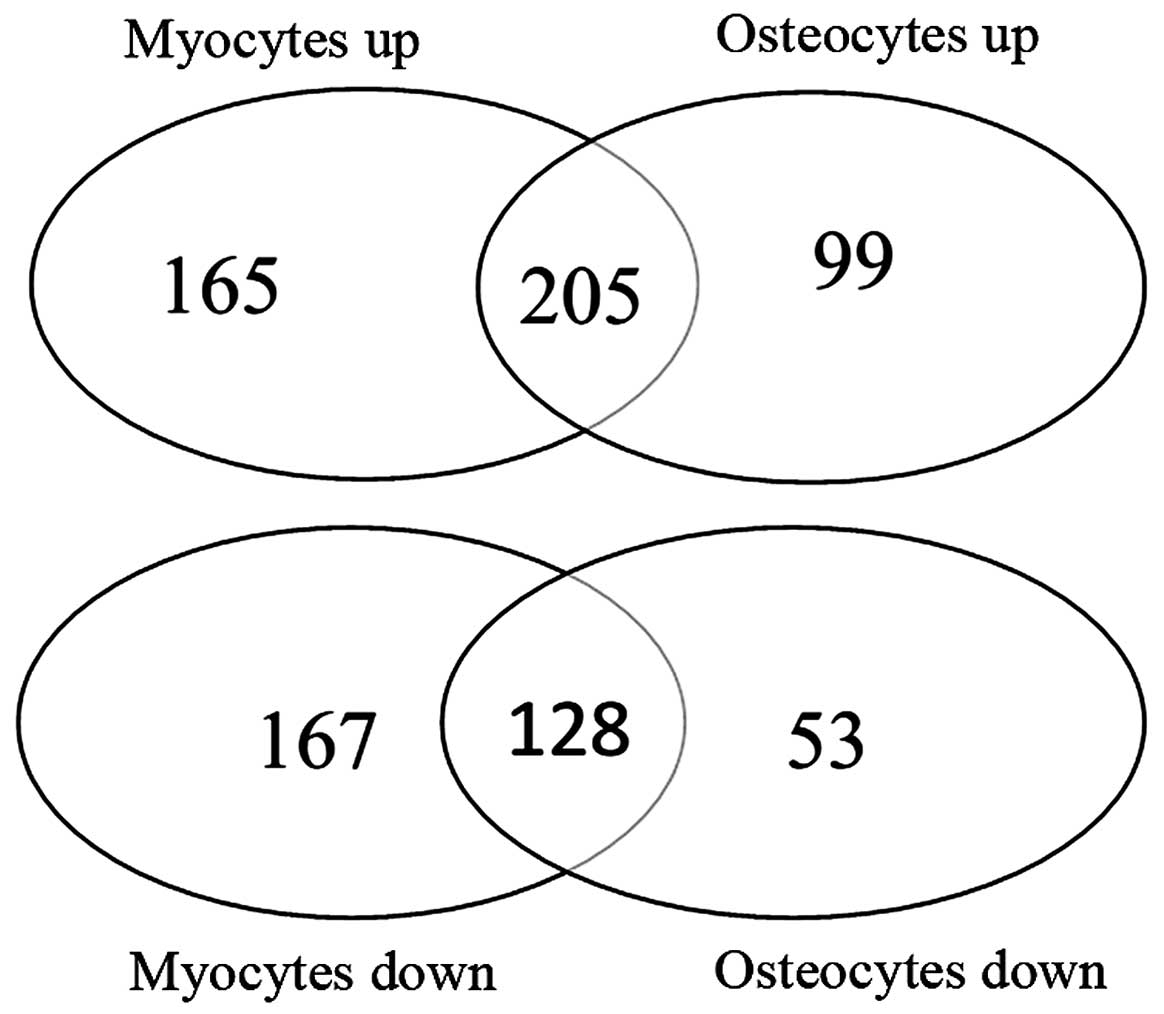
Enrichment analysis of TFs
To further explore the molecular mechanisms
responsible for the differentiation of hASCs into myocytes and
osteocytes, the shared and unshared DEGs in the in
vitro-obtained osteogenic and myogenic lineages were analyzed,
respectively (Fig. 4). The
results of the KEGG enrichment analysis revealed that 205 shared
upregulated genes were mainly involved in metabolism-related
pathways, including drug metabolism and tyrosine metabolism, and
128 shared downregulated genes were significantly enriched in the
TGF-β signaling pathway (Fig. 4
and Table V).
 | Table VEnriched KEGG pathways of shared
genes between two groups (myocytes vs. hASCs and osteocytes vs.
hASCs). |
Table V
Enriched KEGG pathways of shared
genes between two groups (myocytes vs. hASCs and osteocytes vs.
hASCs).
| KEGG ID | Name of
pathway | Count | P-value |
|---|
| Shared up | 00982 | Drug metabolism -
cytochrome P450 | 9 | 2.80E-07 |
| 00350 | Tyrosine
metabolism | 5 | 0.000155935 |
| 04080 | Neuroactive
ligand-receptor interaction | 11 | 0.000601223 |
| 00071 | Fatty acid
metabolism | 4 | 0.002082258 |
| 00460 | Cyanoamino acid
metabolism | 2 | 0.003246415 |
| Shared down | 04350 | TGF-β signaling
pathway | 7 | 3.78E-06 |
| 04610 | Complement and
coagulation cascades | 4 | 0.002133141 |
| 04916 | Melanogenesis | 4 | 0.008361484 |
| 04972 | Pancreatic
secretion | 4 | 0.008361484 |
The relationship between TFs and DEGs may aid in
defining regulatory controls. Finally, a total of 27 TFs targeting
the shared upregulated genes were predicted. In addition, 11 TFs,
which are all involved in the targeting of the shared upregulated
genes, were predicted to target the shared downregulated genes,
including RAD21, zinc finger protein 263 (ZNF263), signal
transducer and activator of transcription 3 (STAT3), RE1-silencing
transcription factor (REST, also known as NRSF), tripartite motif
containing 28 (TRIM28, also known as KAP1), GATA binding protein 2
(GATA2), CCCTC-binding factor (CTCF), E1A binding protein p300
(EP300), early growth response 1 (EGR1), CCAAT/enhancer binding
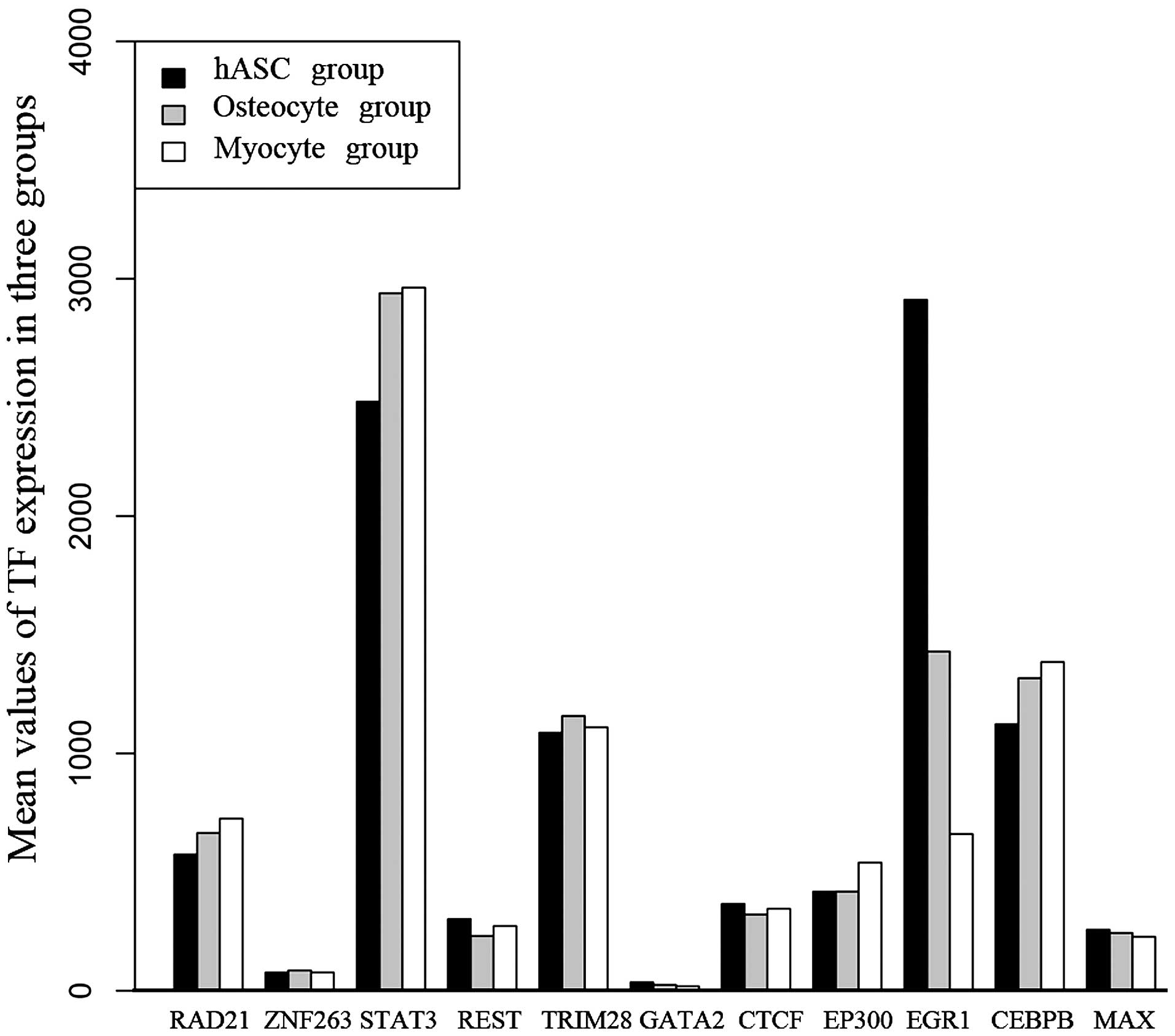
protein (C/EBP), beta (CEBPB) and MYC-associated factor X (MAX).
The expression of these 11 TFs in the three sample types is shown
in Fig. 5. The results revealed
that the expression of EGR1 was significantly higher in the hASCs
than in the osteogenic and the myogenic lineages. Conversely, the
expression of STAT3 was significantly lower in the hASCs than in
the osteogenic and the myogenic lineages. Differential expression
of the other 9 TFs among the three cell types was not found.
In addition, 26 and 21 TFs were predicted to
regulate the unshared up- and downregulated genes in the myogenic
lineages, respectively. In the osteogenic lineages, 11 TFs were
predicted to target the upregulated genes whereas only RAD21 was
found to regulate the downregulated genes. Moreover, RAD21 was also
included among the TFs regulating unshared downregulated genes in
the myogenic lineages, including VEGFA and SMAD family
member 6 (SMAD6).
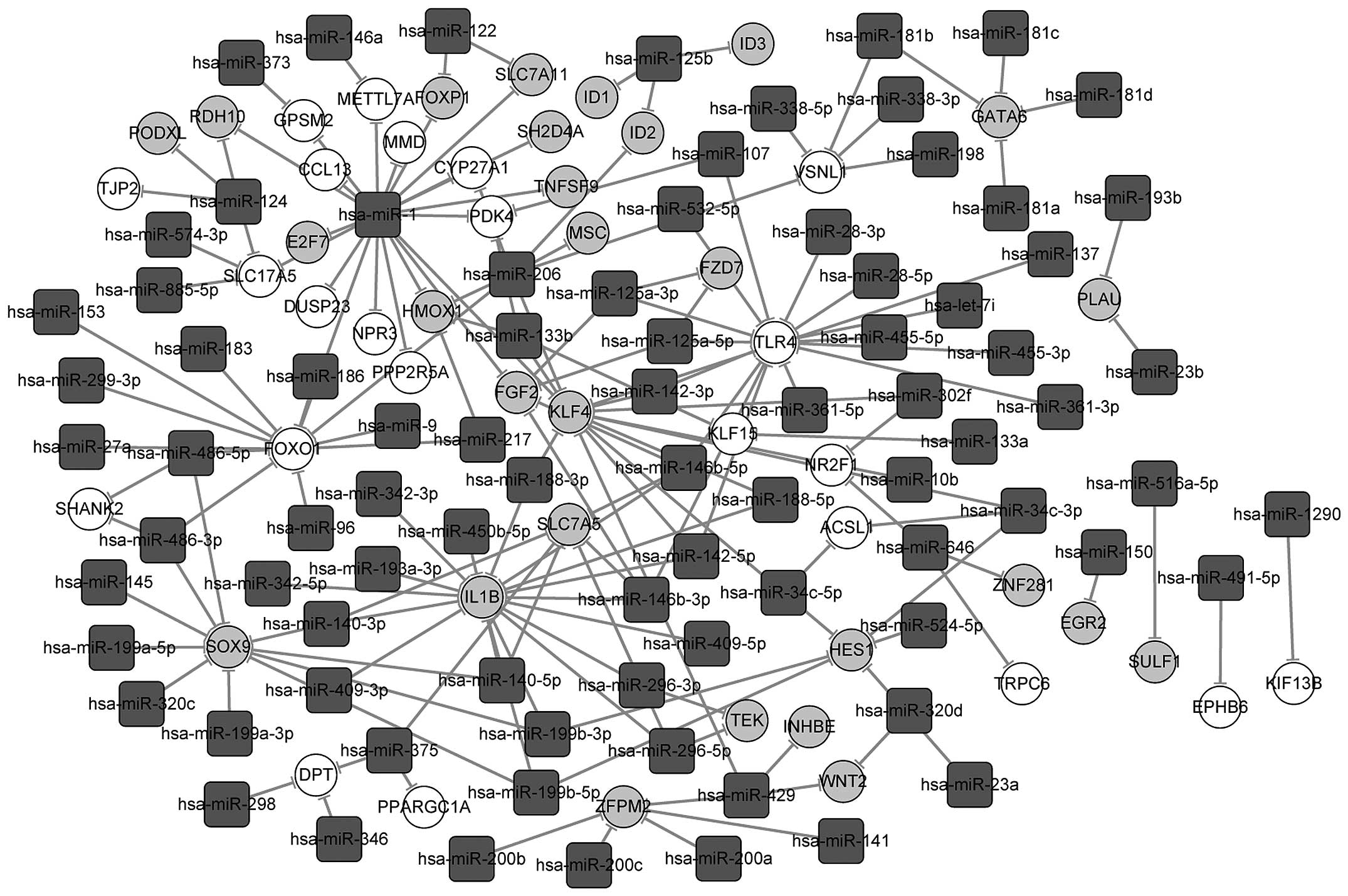
MiRNA-DEG interaction analysis
A total of 66 and 98 miRNA-mRNA pairs were finally
screened out for the shared up- and downregulated genes in the
osteogenic and the myogenic lineages to construct an miRNA-target
gene interaction network, respectively (Fig. 6). In the network, hsa-miR-1, with
the highest degree, regulated 20 common genes differentially
expressed in the two cell types, including Forkhead box P1
(FOXP1), E2F transcription factor 7 (E2F7), chemokine
(C-C motif) ligand 13 (CCL13), monocyte to macrophage
differentiation-associated (MMD) and pyruvate dehydrogenase
kinase, isozyme 4 (PDK4). Moreover, the shared upregulated
genes FOXO1, TLR4 and downregulated gene IL1B
were regulated by >9 miRNAs during the differentiation of hASCs,
and shared downregulated GATA6 was regulated by four
hsa-miR-181 family members namely miR-181a, miR-181b, miR-181c and
miR-181d.
Further, functional annotation revealed that the
shared upregulated genes targeted by the predicted miRNAs were
mainly involved in immune response-related BPs, including detection
of fungus, and host defense responses. By contrast, the shared
downregulated genes were significantly enriched in response to
ozone, smooth muscle adaptation and regulation of myosin light
chain kinase activity (Table
VI).
 | Table VITop 7 enriched GO terms in the BP
category for target genes of miRNAs. |
Table VI
Top 7 enriched GO terms in the BP
category for target genes of miRNAs.
| | GO ID | Name of BP | Count | P-value |
|---|
| miRNA-gene-up | BP | GO:0016046 | Detection of
fungus | 16 | 3.05E-14 |
| GO:0052031 | Modulation by
symbiont of host defense response | 16 | 8.69E-14 |
| GO:0052033 | Pathogen-associated
molecular pattern dependent induction by symbiont of host innate
immune response | 16 | 8.69E-14 |
| GO:0052166 | Positive regulation
by symbiont of host innate immune response | 16 | 8.69E-14 |
| GO:0052167 | Modulation by
symbiont of host innate immune response | 16 | 8.69E-14 |
| GO:0052169 | Pathogen-associated
molecular pattern dependent modulation by symbiont of host innate
immune response | 16 | 8.69E-14 |
| GO:0052255 | Modulation by
organism of defense response of other organism involved in
symbiotic interaction | 16 | 8.69E-14 |
| miRNA-
gene-down | BP | GO:0010193 | Response to
ozone | 17 | 0 |
| GO:0014805 | Smooth muscle
adaptation | 21 | 0 |
| GO:0035504 | Regulation of
myosin light chain kinase activity | 17 | 0 |
| GO:0035505 | Positive regulation
of myosin light chain kinase activity | 17 | 0 |
| GO:0060352 | Cell adhesion
molecule production | 17 | 0 |
| GO:0060353 | Regulation of cell
adhesion molecule production | 17 | 0 |
| GO:0060355 | Positive regulation
of cell adhesion molecule production | 17 | 0 |
Discussion
In the present study, we aimed to extend our
understanding of the molecular mechanisms responsible for the
differentiation of hASCs into myocytes and osteocytes. We found
that four proteins encoded by VEGFA, FGF2, NGF
and IL1B were differentially expressed in the myogenic and
the osteogenic lineages and presented in the PPI network at
relatively high degrees. Moreover, the TF RAD21 was predicted to
target both shared up- and downregulated genes as well as specific
downregulated genes in the myogenic and the osteogenic lineages. In
addition, miRNA-DEG interaction analysis revealed that hsa-miR-1
regulated the most shared DEGs in the two lineages, such as
FOXP1 and CCL13.
Previous findings have suggested that hASCs secrete
significant numbers of angiogenic factors, including VEGFA
(30). VEGFA is known to promote
both angiogenesis and osteogenesis (31,32). More recently, VEGFA has been
proved to play an integral role in the crosstalk between
endothelial cells and osteoblasts and is also considered as being
of great importance for vascularization (33). VEGFA has been found to increase
bone formation, promote osteoblast differentiation and inhibit the
apoptosis of osteoblasts (32,34). In addition, Song et al have
identified VEGF as a critical factor in cardiomyogenesis in hASCs
(35). FGF2, a member of the FGF
family, has been identified as a major candidates for the
regulation of self-renewal in human embryonic stem cells (36,37). FGF2 may also be important in
increasing the lifespan of bone marrow stromal cells and for
supporting proliferation as well as the chondrogenic and osteogenic
differentiation potential (38,39). Moreover, previous studies have
shown that the exposure of hASCs to FGF2 led to the enhancement of
chondrogenic lineage differentiation and the inhibition of
osteogenic lineage differentiation, as well as the stimulation of
adipogenic differentiation (10,40,41). Notably, IL1B, which encodes an
inflammatory cytokine, has been shown to be suppressed by
mesenchymal stem cell (MSC) transplantation at the transcriptional
and the post-transcriptional levels in myocardial infarction
(42). NGF is also reported to be
associated with many pathologic and physiologic processes, such as
differentiation of stem cells (43). In this study, VEGFA, FGF2, IL1B
and NGF were found to be downregulated in the myogenic and
osteogenic lineages compared with hASCs and connected with
relatively more DEGs in the PPI networks, which supports the
hypothesis that there may be a correlation between these genes and
the differentiation of hASCs.
Additionally, TFs and miRNAs are essential
regulatory molecules after DNA replication involved in the
differentiation of hASCs. The TF RAD21 has been proved to be
associated with the maintenance of embryonic stem cell identity
through association with the pluripotency transcriptional network
(44). Consistent with our
analysis, chromatin immunoprecipitation analysis was used in a
previous study to confirm that VEGFA and SMAD6
expression is regulated by RAD21 (45). SMAD6, an inhibitory SMAD, has been
reported to inhibit the TGF-β signaling pathway that suppresses
osteoblast and myogenic differentiation (46). The data from the present study
revealed that RAD21 mediates the differentiation of hASCs by
regulating the expression of VEGFA and SMAD6.
In a previous study, miR-1 was shown to strongly
enhance myogenesis following the transfection of myoblasts with
hsa-miR-1 by modulating skeletal muscle proliferation and
differentiation (47). More
importantly, hsa-miR-1 is required for smooth muscle cell lineage
differentiation from embryonic stem cells by binding with the 3′
untranslated region of the gene encoding Kruppel-like factor 4
(48). Following the construction
of an miRNA-target gene interaction network, we found that miR-1
targeted FOXP1 in the differentiation of hASCs into osteocytes and
myocytes, which is in agreement with the results of a previous
study (49). Additionally, it was
demonstrated that knockdown of FOXP1 suppressed the self-renewal
capacity of MSCs and reduced the osteogenic potential (50). In the hASC-derived myocytes and
osteocytes, CCL13 was upregulated which is consistent with
the findings of a previous study revealing a 12-fold change after
culturing hASCs with proinflammatory cytokines (51). Our results suggest that miR-1
modulates the differentiation of hASCs into myocytes and osteocytes
by regulating FOXP1 and CCL13.
In conclusion, we performed a comprehensive
bioinformatics analysis of the expression profiles of in
vitro-induced osteogenic and myogenic lineages and hASC cell
lines from healthy donors. There may be a correlation between four
shared downregulated genes in the two lineages, VEGFA,
FGF2, IL1B and NGF, and the differentiation of
hASCs. Notably, the TF RAD21 and hsa-miR-1 may play important roles
in regulating the expression of differentiation-associated genes.
This study may provide new insight into the underlying molecular
mechanisms of hASC differentiation, which may help to repair and
reconstruct damaged organs. However, further studies are warranted
to confirm these results and to clarify their roles in the
differentiation of hASCs.
Acknowledgments
The present study was supported by the Liaoning
Province Science and Technology Research Project (no.
2013225220).
References
|
1
|
Zuk PA, Zhu M, Mizuno H, Huang J, Futrell
JW, Katz AJ, Benhaim P, Lorenz HP and Hedrick MH: Multilineage
cells from human adipose tissue: implications for cell-based
therapies. Tissue Eng. 7:211–228. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Halvorsen YD, Bond A, Sen A, Franklin DM,
Lea-Currie YR, Sujkowski D, Ellis PN, Wilkison WO and Gimble JM:
Thiazolidinediones and glucocorticoids synergistically induce
differentiation of human adipose tissue stromal cells: biochemical,
cellular, and molecular analysis. Metabolism. 50:407–413. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Zuk PA: The adipose-derived stem cell:
looking back and looking ahead. Mol Biol Cell. 21:1783–1787. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Luzi E, Marini F, Sala SC, Tognarini I,
Galli G and Brandi ML: Osteogenic differentiation of human adipose
tissue-derived stem cells is modulated by the miR-26a targeting of
the SMAD1 transcription factor. J Bone Miner Res. 23:287–295. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Cai L, Johnstone BH, Cook TG, Tan J,
Fishbein MC, Chen PS and March KL: IFATS collection: human adipose
tissue-derived stem cells induce angiogenesis and nerve sprouting
following myocardial infarction, in conjunction with potent
preservation of cardiac function. Stem Cells. 27:230–237. 2009.
View Article : Google Scholar
|
|
6
|
Nambu M, Ishihara M, Nakamura S, Mizuno H,
Yanagibayashi S, Kanatani Y, Hattori H, Takase B, Ishizuka T,
Kishimoto S, et al: Enhanced healing of mitomycin C-treated wounds
in rats using inbred adipose tissue-derived stromal cells within an
atelocollagen matrix. Wound Repair Regen. 15:505–510. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Wilson A and Trumpp A: Bone-marrow
haematopoietic-stem-cell niches. Nat Rev Immunol. 6:93–106. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Luu YK, Capilla E, Rosen CJ, Gilsanz V,
Pessin JE, Judex S and Rubin CT: Mechanical stimulation of
mesenchymal stem cell proliferation and differentiation promotes
osteogenesis while preventing dietary-induced obesity. J Bone Miner
Res. 24:50–61. 2009. View Article : Google Scholar :
|
|
9
|
Lodish H, Flygare J and Chou S: From stem
cell to erythroblast: regulation of red cell production at multiple
levels by multiple hormones. IUBMB Life. 62:492–496. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Kakudo N, Shimotsuma A and Kusumoto K:
Fibroblast growth factor-2 stimulates adipogenic differentiation of
human adipose-derived stem cells. Biochem Biophys Res Commun.
359:239–244. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Stewart AA, Byron CR, Pondenis H and
Stewart MC: Effect of fibroblast growth factor-2 on equine
mesenchymal stem cell monolayer expansion and chondrogenesis. Am J
Vet Res. 68:941–945. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Huang S, Wang S, Bian C, Yang Z, Zhou H,
Zeng Y, Li H, Han Q and Zhao RC: Upregulation of miR-22 promotes
osteogenic differentiation and inhibits adipogenic differentiation
of human adipose tissue-derived mesenchymal stem cells by
repressing HDAC6 protein expression. Stem Cells Dev. 21:2531–2540.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Mizuno H, Zuk PA, Zhu M, Lorenz HP,
Benhaim P and Hedrick MH: Myogenic differentiation by human
processed lipoaspirate cells. Plast Reconstr Surg. 109:199–111.
2002. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Planat-Bénard V, Menard C, André M, Puceat
M, Perez A, Garcia-Verdugo JM, Pénicaud L and Casteilla L:
Spontaneous cardiomyocyte differentiation from adipose tissue
stroma cells. Circ Res. 94:223–229. 2004. View Article : Google Scholar
|
|
15
|
Jeon ES, Moon HJ, Lee MJ, Song HY, Kim YM,
Bae YC, Jung JS and Kim JH: Sphingosylphosphorylcholine induces
differentiation of human mesenchymal stem cells into
smooth-muscle-like cells through a TGF-beta-dependent mechanism. J
Cell Sci. 119:4994–5005. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Lee WC, Rubin JP and Marra KG: Regulation
of alpha-smooth muscle actin protein expression in adipose-derived
stem cells. Cells Tissues Organs. 183:80–86. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Kim YJ, Bae SW, Yu SS, Bae YC and Jung JS:
miR-196a regulates proliferation and osteogenic differentiation in
mesenchymal stem cells derived from human adipose tissue. J Bone
Miner Res. 24:816–825. 2009. View Article : Google Scholar
|
|
18
|
Maroni P, Brini AT, Arrigoni E, de
Girolamo L, Niada S, Matteucci E, Bendinelli P and Desiderio MA:
Chemical and genetic blockade of HDACs enhances osteogenic
differentiation of human adipose tissue-derived stem cells by
oppositely affecting osteogenic and adipogenic transcription
factors. Biochem Biophys Res Commun. 428:271–277. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Berdasco M, Melguizo C, Prados J, Gómez A,
Alaminos M, Pujana MA, Lopez M, Setien F, Ortiz R, Zafra I, et al:
DNA methylation plasticity of human adipose-derived stem cells in
lineage commitment. Am J Pathol. 181:2079–2093. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Irizarry RA, Hobbs B, Collin F,
Beazer-Barclay YD, Antonellis KJ, Scherf U and Speed TP:
Exploration, normalization, and summaries of high density
oligonucleotide array probe level data. Biostatistics. 4:249–264.
2003. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Smyth GK: Limma: linear models for
microarray data. Bioinformatics and Computational Biology Solutions
Using R and Bioconductor. Springer; New York: pp. 397–420. 2005,
View Article : Google Scholar
|
|
22
|
Ashburner M, Ball CA, Blake JA, Botstein
D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT,
et al: Gene ontology: Tool for the unification of biology. The Gene
Ontology Consortium Nat Genet. 25:25–29. 2000.
|
|
23
|
Kanehisa M and Goto S: KEGG: Kyoto
encyclopedia of genes and genomes. Nucleic Acids Res. 28:27–30.
2000. View Article : Google Scholar
|
|
24
|
Wang J, Zhou X, Zhu J, Gu Y, Zhao W, Zou J
and Guo Z: GO-function: deriving biologically relevant functions
from statistically significant functions. Brief Bioinform.
13:216–227. 2012. View Article : Google Scholar
|
|
25
|
Franceschini A, Szklarczyk D, Frankild S,
Kuhn M, Simonovic M, Roth A, Lin J, Minguez P, Bork P, von Mering C
and Jensen LJ: STRING v9.1: protein-protein interaction networks,
with increased coverage and integration. Nucleic Acids Res.
41:D808–D815. 2013. View Article : Google Scholar :
|
|
26
|
Kohl M, Wiese S and Warscheid B:
Cytoscape: software for visualization and analysis of biological
networks. Methods Mol Biol. 696:296–303. 2011.
|
|
27
|
Meyer LR, Zweig AS, Hinrichs AS, Karolchik
D, Kuhn RM, Wong M, Sloan CA, Rosenbloom KR, Roe G, Rhead B, et al:
The UCSC Genome Browser database: extensions and updates 2013.
Nucleic Acids Res. 41:D64–D69. 2013. View Article : Google Scholar :
|
|
28
|
Xiao F, Zuo Z, Cai G, Kang S, Gao X and Li
T: miRecords: an integrated resource for microRNA-target
interactions. Nucleic Acids Res. 37:D105–D110. 2009. View Article : Google Scholar
|
|
29
|
Dweep H, Sticht C, Pandey P and Gretz N:
miRWalk - database: prediction of possible miRNA binding sites by
'walking' the genes of three genomes. J Biomed Inform. 44:839–847.
2011. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Rehman J, Traktuev D, Li J, Merfeld-Clauss
S, Temm-Grove CJ, Bovenkerk JE, Pell CL, Johnstone BH, Considine RV
and March KL: Secretion of angiogenic and antiapoptotic factors by
human adipose stromal cells. Circulation. 109:1292–1298. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Olsson AK, Dimberg A, Kreuger J and
Claesson-Welsh L: VEGF receptor signalling - in control of vascular
function. Nat Rev Mol Cell Biol. 7:359–371. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Street J, Bao M, deGuzman L, Bunting S,
Peale FV Jr, Ferrara N, Steinmetz H, Hoeffel J, Cleland JL,
Daugherty A, et al: Vascular endothelial growth factor stimulates
bone repair by promoting angiogenesis and bone turnover. Proc Natl
Acad Sci USA. 99:9656–9661. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Clarkin CE, Emery RJ, Pitsillides AA and
Wheeler-Jones CP: Evaluation of VEGF-mediated signaling in primary
human cells reveals a paracrine action for VEGF in
osteoblast-mediated crosstalk to endothelial cells. J Cell Physiol.
214:537–544. 2008. View Article : Google Scholar
|
|
34
|
Street J and Lenehan B: Vascular
endothelial growth factor regulates osteoblast survival - evidence
for an autocrine feedback mechanism. J Orthop Surg. 4:192009.
View Article : Google Scholar
|
|
35
|
Song YH, Gehmert S, Sadat S, Pinkernell K,
Bai X, Matthias N and Alt E: VEGF is critical for spontaneous
differentiation of stem cells into cardiomyocytes. Biochem Biophys
Res Commun. 354:999–1003. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Xu C, Rosler E, Jiang J, Lebkowski JS,
Gold JD, O'Sullivan C, Delavan-Boorsma K, Mok M, Bronstein A and
Carpenter MK: Basic fibroblast growth factor supports
undifferentiated human embryonic stem cell growth without
conditioned medium. Stem Cells. 23:315–323. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Dvorak P, Dvorakova D, Koskova S, Vodinska
M, Najvirtova M, Krekac D and Hampl A: Expression and potential
role of fibroblast growth factor 2 and its receptors in human
embryonic stem cells. Stem Cells. 23:1200–1211. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Martin I, Muraglia A, Campanile G,
Cancedda R and Quarto R: Fibroblast growth factor-2 supports ex
vivo expansion and maintenance of osteogenic precursors from human
bone marrow. Endocrinology. 138:4456–4462. 1997.PubMed/NCBI
|
|
39
|
Solchaga LA, Penick K, Porter JD, Goldberg
VM, Caplan AI and Welter JF: FGF-2 enhances the mitotic and
chondrogenic potentials of human adult bone marrow-derived
mesenchymal stem cells. J Cell Physiol. 203:398–409. 2005.
View Article : Google Scholar
|
|
40
|
Quarto N and Longaker MT: FGF-2 inhibits
osteogenesis in mouse adipose tissue-derived stromal cells and
sustains their proliferative and osteogenic potential state. Tissue
Eng. 12:1405–1418. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Chiou M, Xu Y and Longaker MT: Mitogenic
and chondrogenic effects of fibroblast growth factor-2 in
adipose-derived mesenchymal cells. Biochem Biophys Res Commun.
343:644–652. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Guo J, Lin GS, Bao CY, Hu ZM and Hu MY:
Anti-inflammation role for mesenchymal stem cells transplantation
in myocardial infarction. Inflammation. 30:97–104. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Sariola H: The neurotrophic factors in
non-neuronal tissues. Cell Mol Life Sci. 58:1061–1066. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Nitzsche A, Paszkowski-Rogacz M, Mata rese
F, Janssen-Megens EM, Hubner NC, Schulz H, de Vries I, Ding L,
Huebner N, Mann M, et al: RAD21 cooperates with pluripotency
transcription factors in the maintenance of embryonic stem cell
identity. PLoS One. 6:e194702011. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Tang M, Chen B, Lin T, Li Z, Pardo C,
Pampo C, Chen J, Lien CL, Wu L, Ai L, et al: Restraint of
angiogenesis by zinc finger transcription factor CTCF-dependent
chromatin insulation. Proc Natl Acad Sci USA. 108:15231–15236.
2011. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Roelen BA and Dijke P: Controlling
mesenchymal stem cell differentiation by TGFBeta family members. J
Orthop Sci. 8:740–748. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Chen JF, Mandel EM, Thomson JM, Wu Q,
Callis TE, Hammond SM, Conlon FL and Wang DZ: The role of
microRNA-1 and microRNA-133 in skeletal muscle proliferation and
differentiation. Nat Genet. 38:228–233. 2006. View Article : Google Scholar
|
|
48
|
Xie C, Huang H, Sun X, Guo Y, Hamblin M,
Ritchie RP, Garcia-Barrio MT, Zhang J and Chen YE: MicroRNA-1
regulates smooth muscle cell differentiation by repressing
Kruppel-like factor 4. Stem Cells Dev. 20:205–210. 2011. View Article : Google Scholar :
|
|
49
|
Datta J, Kutay H, Nasser MW, Nuovo GJ,
Wang B, Majumder S, Liu CG, Volinia S, Croce CM, Schmittgen TD, et
al: Methylation mediated silencing of MicroRNA-1 gene and its role
in hepatocellular carcinogenesis. Cancer Res. 68:5049–5058. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Kubo H, Shimizu M, Taya Y, Kawamoto T,
Michida M, Kaneko E, Igarashi A, Nishimura M, Segoshi K, Shimazu Y,
et al: Identification of mesenchymal stem cell (MSC)-transcription
factors by microarray and knockdown analyses, and signature
molecule-marked MSC in bone marrow by immunohistochemistry. Genes
Cells. 14:407–424. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Crop MJ, Baan CC, Korevaar SS, Ijzermans
JN, Pescatori M, Stubbs AP, van Ijcken WF, Dahlke MH, Eggenhofer E,
Weimar W and Hoogduijn MJ: Inflammatory conditions affect gene
expression and function of human adipose tissue-derived mesenchymal
stem cells. Clin Exp Immunol. 162:474–486. 2010. View Article : Google Scholar : PubMed/NCBI
|