Introduction
The increase of the adipogenic cell population and
accumulation of lipid may lead to obesity. Obesity and related
diseases, including type 2 diabetes, stroke, hypertension and
cardiovascular disease, have become major medical problems in
modern society (1,2). To control obesity and related
diseases, it is crucial to understand and monitor cellular
adipogenic lineage differentiation and development.
Embryonic stem cells (ESCs) (3), mesenchymal stem cells (MSCs)
(4) and preadipocytes (5) have been used to study cellular
adipogenic differentiation in vitro. Key factors regulating
adipogenesis have been extensively investigated at the gene and
protein levels in recent years. Adipogenesis may be divided into
two stages, namely commitment and terminal differentiation.
Commitment has been defined as the process leading from multipotent
stem cells to preadipocytes (4).
Certain factors, including bone morphogenetic proteins 2, 4, Wnt
and retinoic acid (RA), have been identified as activators of the
commitment pathway (4,6,7),
whereas sonic hedgehog inhibits this process (8,9).
RA was found to play an important role in the early differentiation
of ESCs, and RA-pretreated embryoid bodies (EBs) displayed a high
level of adipogenesis (3).
Terminal differentiation has been defined as cell differentiation
from preadipocytes to adipocytes. In vitro, terminal
differentiation may be induced by a cocktail containing
insulin-like growth factor-1 (IGF-1) (10), glucocorticoid and cyclic AMP
(cAMP) (11). During the process
of terminal differentiation, the expression of several key
transcription factor families, such as CCATT/enhancer-binding
protein (c/ebp) family (12) and
the peroxisome proliferator-activated receptor (ppar) family, was
found to be upregulated. The coordination of these transcription
factors produces the adipocyte phenotype and maintains the
expression of adipocyte genes (13).
Protein glycosylation is an enzymatic process that
produces glycosidic linkages of glycans to proteins, and it is a
ubiquitous post-translational modification. Approximately 70% of
human proteins contain ≥1 glycan chains (14) and 1% of the human genome is
involved in glycan production and modification (15). Protein glycosylation plays a
crucial role in cell growth and development (15,16), cell-to-cell communication
(17,18), tumour growth and metastasis
(19,20). Indeed, several stemness markers,
such as stage-specific embryonic antigen (SSEA)-1 and -3, are
glycoproteins (21). Although
adipogenesis has been studied at the gene and protein level, it
remains unknown how protein glycosylation is altered during this
process, particularly in parthenogenetic ESCs (pESCs), which are
obtained from parthenogenetic blastocysts, exhibit low
immunogenicity and are assosiated with no ethical issues regarding
their use.
Lectin microarray is a technology for screening the
structure of the glycome, where a large number of lectins with
various glycan-binding specificities are printed on a microarray.
This technology enables rapid and high-throughput profiling of
protein glycosylation without liberation of glycans from proteins.
In addition, altered glycan patterns may be visualised and located
in tissues and cells through lectin histochemistry (22). Lectin mircoarray has been applied
mainly to detect glycans as disease-related biomarkers (23,24). Tateno et al compared
protein glycosylation conditions of human ESCs and somatic cells,
and found that rBC2LCN [binding for Fucα1-2Galβ1-3GlcNAc
(GalNAc)-containing glycans] signaling only existed in
undifferentiated ESCs, but not in differentiated somatic cells.
This result indicated that rBC2LCN was the optimal lectin probe for
evaluating pluripotency, and that the presence of
Fucα1-2Galβ1-3GlcNAc (GalNAc)-containing glycans was a biomarker of
pluripotency.
In the present study, the alterations of J1 ESCs and
pESCs during adipogenesis were analysed using a lectin microarray.
The basic properties and differentiation potential of the two types
of ESCs were first compared, and it was observed that J1 ESCs and
pESCs have different adipogenic potentials. The alterations of
protein glycosylation in J1 ESCs and pESCs during this process were
then compared. This experiment is not only helpful for the study of
the mechanism of adipogenesis, but may also indicate new markers
for monitoring adipogenic development.
Materials and methods
Cell cultivation and
characterization
J1 ESCs and pESCs (from C57BL/6J mice, a kind gift
from Professor Jinlian Hua, the Northwest A&F University,
Shaanxi, China) were cultured on a feeder layer of mitomycin C
(Sigma-Aldrich, Merck KGaA, St. Louis, MO, USA)-treated mouse
embryonic fibroblasts with ESGRO Complete PLUS Clonal Grade Medium
(Millipore, Billerica, MA, USA) (25) in a 37°C/5% CO2
incubator. The cells were passaged every 4–5 days using Accutase
(Millipore) at a ratio of 1:3–1:6 and observed under a phase
contrast microscope (Nikon, Tokyo, Japan).
To assess cellular pluripotency in vitro, J1
ESCs and pESCs grown on glass coverslips were fixed with 4% cold
paraformaldehyde (Sigma-Aldrich, Merck KGaA) in phosphate-buffered
saline (PBS) for 30 min, and then permeabilized with 0.1% Triton
X-100 (Sigma-Aldrich, Merck KGaA) in PBS for 8 min. Fixed cells
were blocked with 5% bovine serum albumin (BSA; Sigma-Aldrich,
Merck KGaA) in PBS for 1 h and incubated overnight at 4°C with
primary antibodies including rabbit anti-Nanog (M-149, sc-33760),
rabbit anti-Oct4 (H-134, sc-9081) and mouse anti-SSEA-1 (480,
sc-21702) (dilution 1:200; all from Santa Cruz Biotechnology, Inc.,
Santa Cruz, CA, USA). Secondary antibodies (A-11034 and A-21044,
dilution 1:300; Invitrogen Life Technologies, Thermo Fisher
Scientific, Carlsbad, CA, USA) were added and allowed to incubate
for 1 h at 37°C. The nuclei were counterstained with
4′,6-diamidino-2-phenylindole (DAPI; Sigma-Aldrich, Merck KGaA).
The cells were washed thoroughly with PBS after each incubation
step. The images were captured using a FV1000 confocal microscope
(Olympus, Tokyo, Japan).
Animals
Five-week-old nude mice were obtained from the
Experimental Animal Center of the Fourth Military Medical
University. All the animal studies and protocols were processed
according to the guidelines of the Animal Holding Unit of Northwest
University. To test pluripotency in vivo, J1 ESCs and pESCs
were dispersed using Accutase. Cells (1×106) in 50
μl Dulbecco's modified Eagle's medium (DMEM; Hyclone, Logan,
UT, USA) were subcutaneously injected into nude mice. The animals
were sacrificed and the formed teratomas were harvested 4 weeks
after the injections. The specimens were fixed with 4%
paraformaldehyde, embedded in paraffin, sectioned at 4 μm,
and stained with hematoxylin and eosin.
J1 ESC and pESC proliferation was analyzed using the
cell proliferation reagent
4-[3-(4-iodophenyl)-2-(4-nitrophenyl)-2H-5-tetrazolio]-1,3-benzene
disulfonate (WST-1) assay (Roche Diagnostics, Mannheim, Germany).
For this procedure, 2,000 cells in 100 μl medium were seeded
into 96-well plates in triplicate. WST-1 (10 μl) was added
to each well and incubated for up to 3 days. The absorbance was
measured every 24 h with a microplate reader (Synergy HT; BioTek
Instruments, Inc., Winooski, VT, USA) at 450 nm, according to the
manufacturer's instructions.
Mesoderm differentiation of J1 ESCs and
pESCs
EBs were generated using the hanging drop method
(3). Cells were dissociated with
Accutase to obtain single-cell suspensions. Hanging drops
containing 1×103 cells in 20 μl medium [DMEM
supplemented with 50 mg/ml L-proline (Sigma-Aldrich, Merck KGaA),
1% non-essential amino acids (Millipore), 1% 2-mercaptoethanol
(Sigma-Aldrich, Merck KGaA) and 20% fetal bovine serum (FBS; Gibco
Life Technologies, Thermo Fisher Scientific, Grand Island, NY,
USA)] were maintained for 2 days on the lids of non-adherent dishes
filled with PBS. The formed EBs were then transferred into
non-adherent plates in medium with or without 10−5 mM RA
(Sigma-Aldrich, Merck KGaA) for a further 3 days.
For osteogenic and chondrogenic differentiation,
non-RA-pretreated EBs were trypsinized and plated on cell culture
dishes coated with 0.1% gelatin (Sigma-Aldrich, Merck KGaA). The
cells were cultured in osteogenic induction medium [DMEM
supplemented with 50 nM dexamethasone, 10 mM β-glycerophosphate and
50 μM ascorbate-2-phosphate (all from Sigma-Aldrich, Merck
KGaA) and 10% FBS] and in chondrogenic induction medium [DMEM
supplemented with 50 μM ascorbic acid 2-phosphate, 10 ng/ml
transforming growth factor-β1 and 500 ng/ml IGF (R&D Systems,
Inc., Minneapolis, MN, USA) and 10% FBS] for 20 days. The medium
was changed every 3 days and the specimens were stained with
Alizarin Red and Safranin O, respectively. To detect osteogenic and
chondrogenic differentiation-related gene expression, total RNA was
extracted from cells after induction with RNA Isolation reagent
(Takara Bio, Inc., Tokyo, Japan) according to the manufacturer's
instructions. The extracted RNA was quantified using a GeneQuant
pro (GE Healthcare Life Sciences, Logan, UT, USA). The RevertAid™
First Strand cDNA Synthesis kit (Thermo Fisher Scientific, Inc.,
Waltham, MA, USA) was used to convert the RNA template into cDNA.
The reaction was conducted in an Eppendorf Mastercycler (Eppendorf,
Hamburg, Germany) as follows: 65°C for 5 min, 25°C for 5 min, 50°C
for 50 min and 70°C for 15 min. The amplified products were
separated with agarose gels and visualized with the assistance of
ethidium bromide and ultraviolet light. The polymerase chain
reaction (PCR) primers are listed in Table I.
 | Table IList of primers used for
RT-PCR/qPCR. |
Table I
List of primers used for
RT-PCR/qPCR.
| Genes | Primer sequences,
sense/antisense (5′-3′) | Product size
(bp) |
|---|
| fabp4 |
ATGGCCAAGCCCAACATGAT | 135 |
|
CTTCCTGTCGTCTGCGGTGAT | |
| ppar γ |
AAGGGTGCCAGTTTCGATCCGTAG | 154 |
|
ATCAGGGAGGCCAGCATCGTGTAG | |
| alp |
ACAACCTGACTGACCCTTCG | 104 |
|
AATCCTGCCTCCTTCCACC | |
| ocn |
TGGCCATCACCCTGTCTCCT | 229 |
|
GAGACCACTCCAGCACAACTCCT | |
| aggrecan |
TGTAAGGACTGTCTATCTACACGCC | 147 |
|
TGGATGGTGATGTCGTCTTCAC | |
| Type II
collagen |
AAGACGGTGAGACGGGAGCC | 287 |
|
ACCATCAGTACCAGGAGTGCCAG | |
| oct4 |
GATCACTCACATCGCCAATCA | 128 |
|
CTGTAGCCTCATACTCTTCTCGTT | |
| c/ebp α |
AAGAGGACGAGGCGAAGCAGC | 132 |
|
GCGCGATCTGGAACTGCAAGT | |
| Alg3 |
TACCTATCCCGCTCCTTTGAC | 171 |
|
CCCTGTTCTGTGCCACCTAC | |
| Alg12 |
GGAGTCCTTGGGCTTGGTGTG | 191 |
|
CAGGCTGTGAGGGCTGGTAAGAC | |
| β-actin |
CAGCCTTCCTTCTTGGGTAT | 100 |
|
TGGCATAGAGGTCTTTACGG | |
For adipogenic differentiation, RA-pretreated EBs
were trypsinized and plated onto gelatin-coated plates and cultured
in adipogenic induction medium [DMEM supplemented with 1 μM
dexamethasone, 200 μM indomethacin and 10 μg/ml
insulin (both from Sigma-Aldrich, Merck KGaA), 0.5 mM
methylisobutylxanthine (MP Biomedicals LLC, Santa Ana, CA, USA) and
10% FBS] for 20 days. The medium was changed every 3 days. At 5,
10, 15 and 20 days after induction, the specimens were processed
for quantitative PCR (qPCR) to evaluate the expression of
adipogenic differentiation-related genes, followed by
immunofluorescence staining of c/ebp α and fabp4 (dilution 1:200;
both from Santa Cruz Biotechnology, Inc.), as described above. The
specimens were further stained with Oil red O. The proportion of
cells undergoing adipogenic differentiation was calculated by
counting Oil red O-positive and -negative cells using a microscope.
Five fields of view were counted for each specimen, as previously
described (26). The qPCR primers
are listed in Table I.
Lectin microarrays and lectin
histochemistry
A lectin microarray of 37 lectins (Vector
Laboratories, Burlingame, CA, USA; Sigma-Aldrich, Merck KGaA and
Calbiochem-Merck Co., Darmstadt, Germany) was produced as
previously described (22). The
protein was extracted from specimens 20 days after adipogenic
induction with T-PER reagent (Pierce Biotechnology Inc., Rockford,
IL, USA), according to the manufacturer's instructions. The
extracted protein was labelled by a Cy3 fluorescent dye and
purified by Sephadex G-25 columns (both from GE Healthcare Life
Sciences). Subsequently, 4 μg of Cy3-labeled protein diluted
with 0.5 ml of hybridization buffer (2% BSA, 500 Mm glycine and
0.1% Tween-20 in PBS) was applied to the blocked lectin
microarrays, and the array was then incubated at 37°C at
2.6×10−3 × g for 3 h. The slides were washed three times
with PBST (0.2% Tween-20 in PBS), then washed with PBS, and dried
by centrifugation at 60 × g for 5 min. The microarrays were then
scanned with a Genepix 4000B confocal scanner (Axon Instruments,
Union City, CA, USA) using a 70% photomultiplier tube and 100%
laser power. The images were analysed at 532 nm for Cy3 detection
using the GenePix 3.0 software (Axon Instruments). The mean
background was subtracted, and the values lower than the mean
background ± 2 standard deviations (SD) were removed from each data
point. For each lectin, the median of the effective data points was
globally normalized to the sum of the medians for all the effective
data points in one block. Each sample was consistently observed in
three repeated slides. The normalized medians and the SD of each
lectin from 9 repeated blocks were averaged. The normalized data
were then compared to determine any relative changes in the protein
glycosylation levels. The original data were further analysed using
Expander 6.0 software (http://acgt.cs.tau.ac.il/expander/).
Cy3-labeled lectins were also applied to detect the
specific glycan structure present in the cells following the
protocol previously described (27). Briefly, cells grown on glass
coverslips were fixed with 4% paraformaldehyde in PBS for 30 min,
permeabilized with 0.1% Triton X-100 in PBS for 30 min, and blocked
with 5% BSA in PBS for 1 h. The cells were incubated with a
solution containing a final concentration of 100 μg/ml Cy3
fluorescein-labeled lectins and 5% (w/v) BSA for 3 h at room
temperature in the dark. Subsequently, they were counterstained
with 1 μg/ml DAPI and washed with PBS. Imaging was performed
using a FV1000 confocal microscope (Olympus).
qPCR
The qPCR primers are listed in Table I. qPCR was performed in a MyiQ
single-color detection system (Bio-Rad Laboratories, Inc.,
Richmond, CA, USA). The qPCR protocol was as follows: Hot start for
30 sec at 95°C, denaturation for 10 sec at 95°C, annealing for 15
sec at 55°C, and extension for 20 sec at 72°C for 30–35 cycles.
Melt curve analyses were performed for each experiment to verify
the formation of a single, desired PCR product. GAPDH was used as
an internal control to normalize the amount of RNA. Absolute values
obtained for each sample were normalized to the GAPDH signal.
Single-peak melting profiles were obtained for the amplifications,
and the size of the PCR product was confirmed by agarose gel
electrophoresis. Each experiment was repeated three times. The ΔΔCt
method was used to calculate relative amounts of transcripts.
Statistical analysis
Data are presented as mean ± SD and compared using
unpaired Student's t-test using SPSS statistics, version 18 (SPSS,
Inc., Chicago, IL, USA). The statistical significance level was
defined as P<0.05.
Results
Basic properties of J1 ESCs and
pESCs
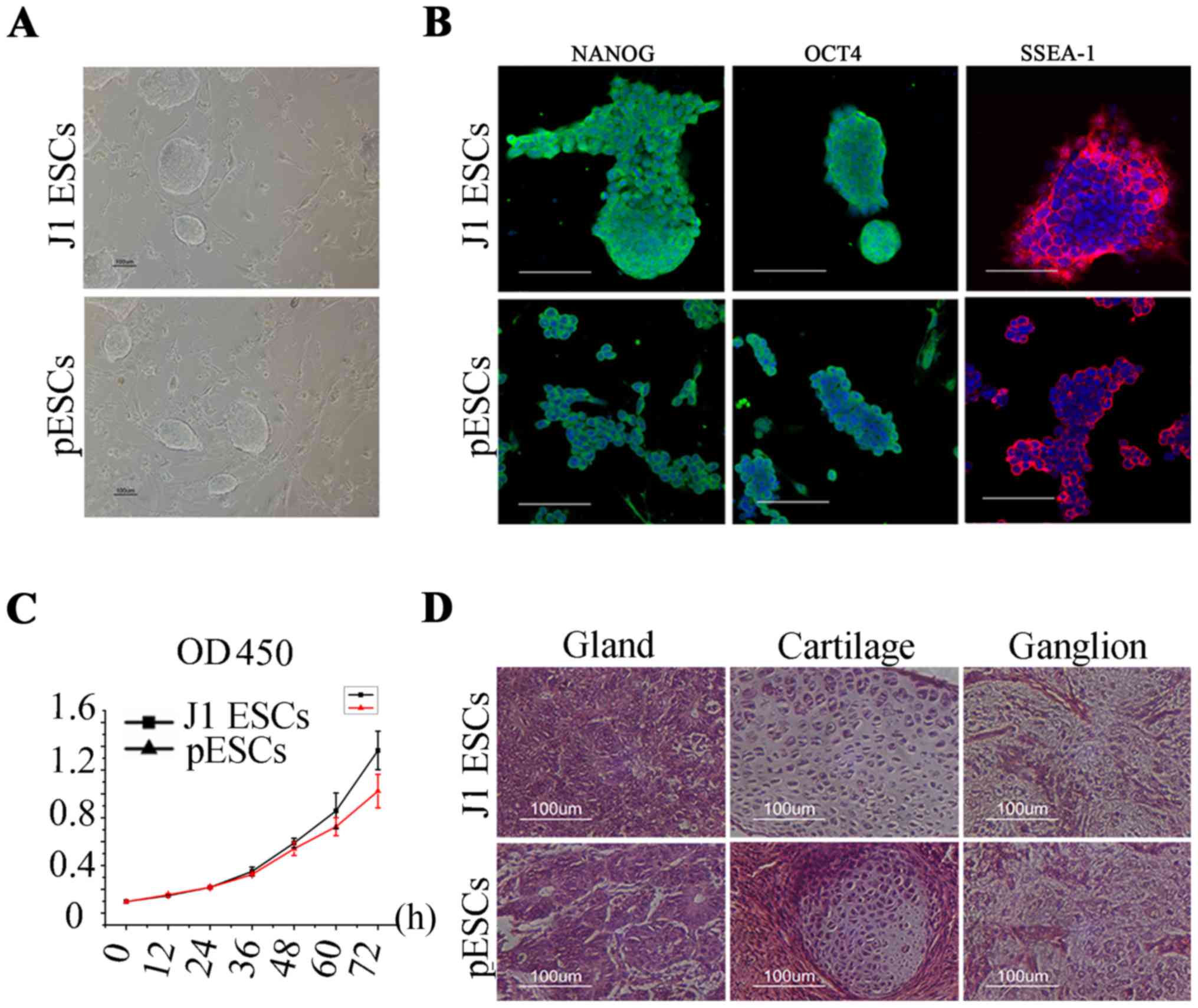
Both J1 ESCs and pESCs had a high nucleus:cytoplasm
ratio and exhibited compact colony structure during in vitro
culture and expansion (Fig. 1A).
Oct4 and Nanog are transcription factors located in the cell
nucleus. Ssea-1 is a type of glycoprotein located on the cell
membrane. According to the results presented in Fig. 1B, J1 ESCs and pESCs were strongly
positive for immunofluorescence staining of these stemness factors.
Both cell types proliferated rapidly in vitro. The growth
kinetics of the two types of ESCs were compared (Fig. 1C), and no difference in
proliferative capacity was observed between the two (P>0.05).
The capacity of teratoma formation of the two types of cells was
then examined. Histological observation revealed that both types
could form teratomas containing ganglion (ectodermal), gland
(endodermal) and cartilage (mesodermal) tissue 4 weeks after
subcutaneous injection in nude mice (Fig. 1D), indicating that these cells
possessed pluripotent differentiation potential in vivo.
Induced mesoderm differentiation
potential of J1 ESCs and pESCs in vitro
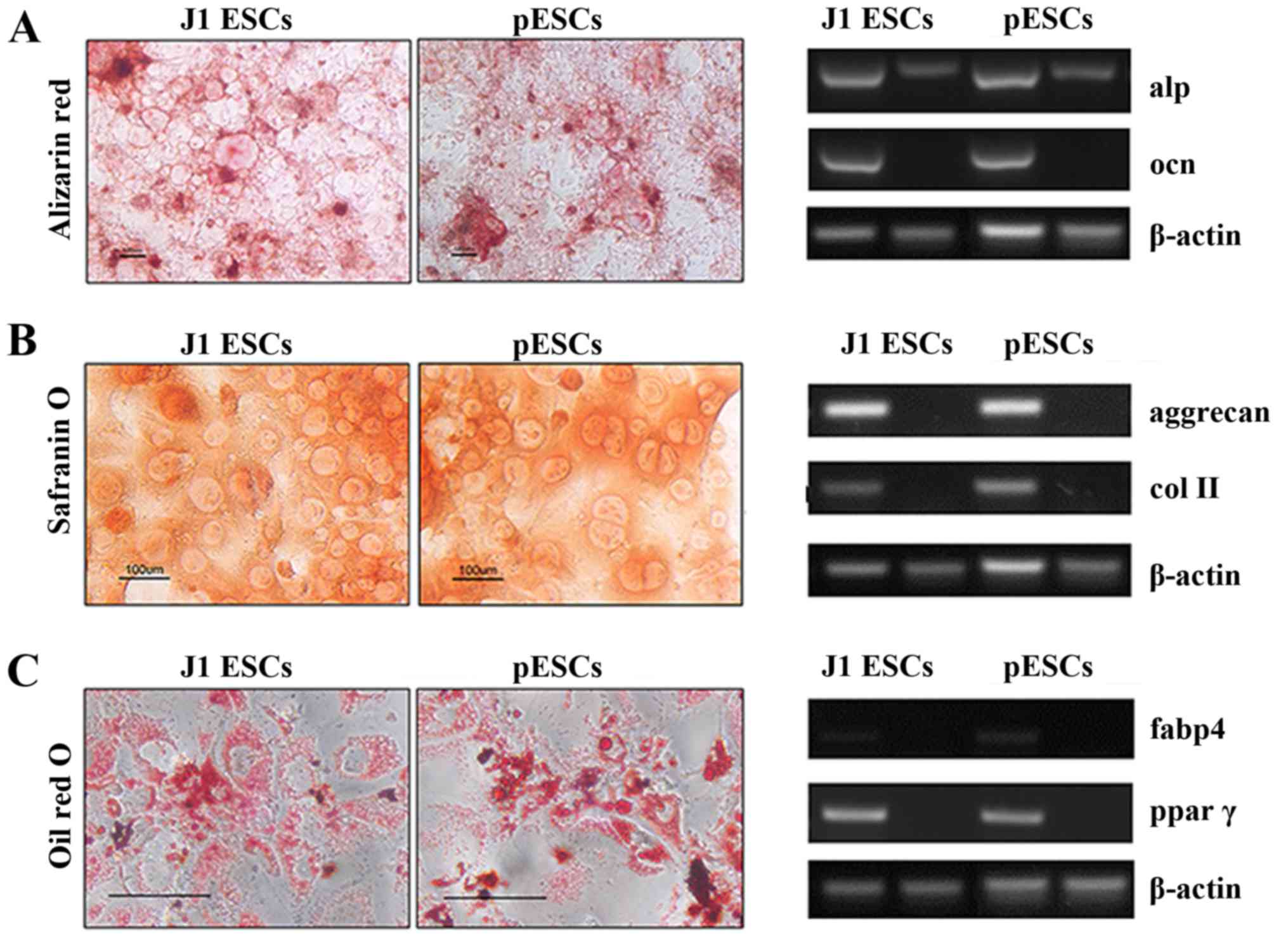
The osteogenic, chondrogenic and adipogenic
differentiation potentials were tested in J1 ESCs and pESCs. At 20
days after being cultured in osteogenic induction medium,
osteogenesis was indicated by the expression of osteocalcin and
upregulation of alkaline phosphatase. Alizarin Red S staining
further confirmed that both ESCs exhibited osteogenic potential and
triggered calcium deposition (Fig.
2A). Chondrogenic differentiation was proven by gene expression
of aggrecan and type II collagen. The strong staining by Safranin O
could be observed in J1 ESC derivatives and pESC derivatives 20
days after induction (Fig. 2B).
The adipogenic differentiation was validated by the expression of
adipogenic genes, including fatty acid-binding protein 4 (fabp4)
and peroxisome proliferator-activated receptor γ (ppar γ), and Oil
red O staining confirmed the formation of lipid droplets in the
cytoplasm (Fig. 2C). The study of
gene expression and matrix production revealed that both J1 ESCs
and pESCs were able to differentiate toward mesoderm lineages,
including osteoblasts, chondrocytes and adipocytes.
pESCs exhibited a lower adipogenic
differentiation capability comp ared with J1 ESCs
The adipogenic potential of J1 ESCs and pESCs was
further compared systematically. The scheme of adipogenic
differentiation is shown in Fig.
3A. The mRNA levels of adipogenic-specific and stemness markers
during the process were first compared using qPCR. As shown in
Fig. 3B, the expression of
adipogenic-specific genes increased gradually, and that of stemness
marker Oct4 decreased gradually during adipogenic differentiation.
At 20 days after induction, the expression of ppar γ, c/ebp α and
fabp4 was upregulated ~2,757-, 13- and 147-fold, respectively, in
J1 ESC derivatives, and 151-, 5- and 23-fold, respectively, in pESC
derivatives. The cells were then incubated with antibodies to
detect adipogenic-specific protein expression during adipogenesis.
The results demonstrated that c/ebp α and fabp4 were both gradually
upregulated during the process (Fig.
3C). However, the expression of c/ebp α and fabp4 could be
detected in J1 ESCs derivatives as early as 5 days after induction,
while it could only be detected in pESC derivatives 10 days after
induction. Both cell types could be induced to differentiate into
adipocytes accompanied by increased accumulation of lipids
(Fig. 3D). Cell counting revealed
that, after 20 days of induction, 60.4±6.4% and 42.2±6.9% Oil red
O-positive cells were observed in J1 ESC and pESC derivatives,
respectively (Fig. 3E;
P<0.05). Collectively, these results indicate that pESCs
exhibited a lower adipogenic potential compared with J1 ESCs.
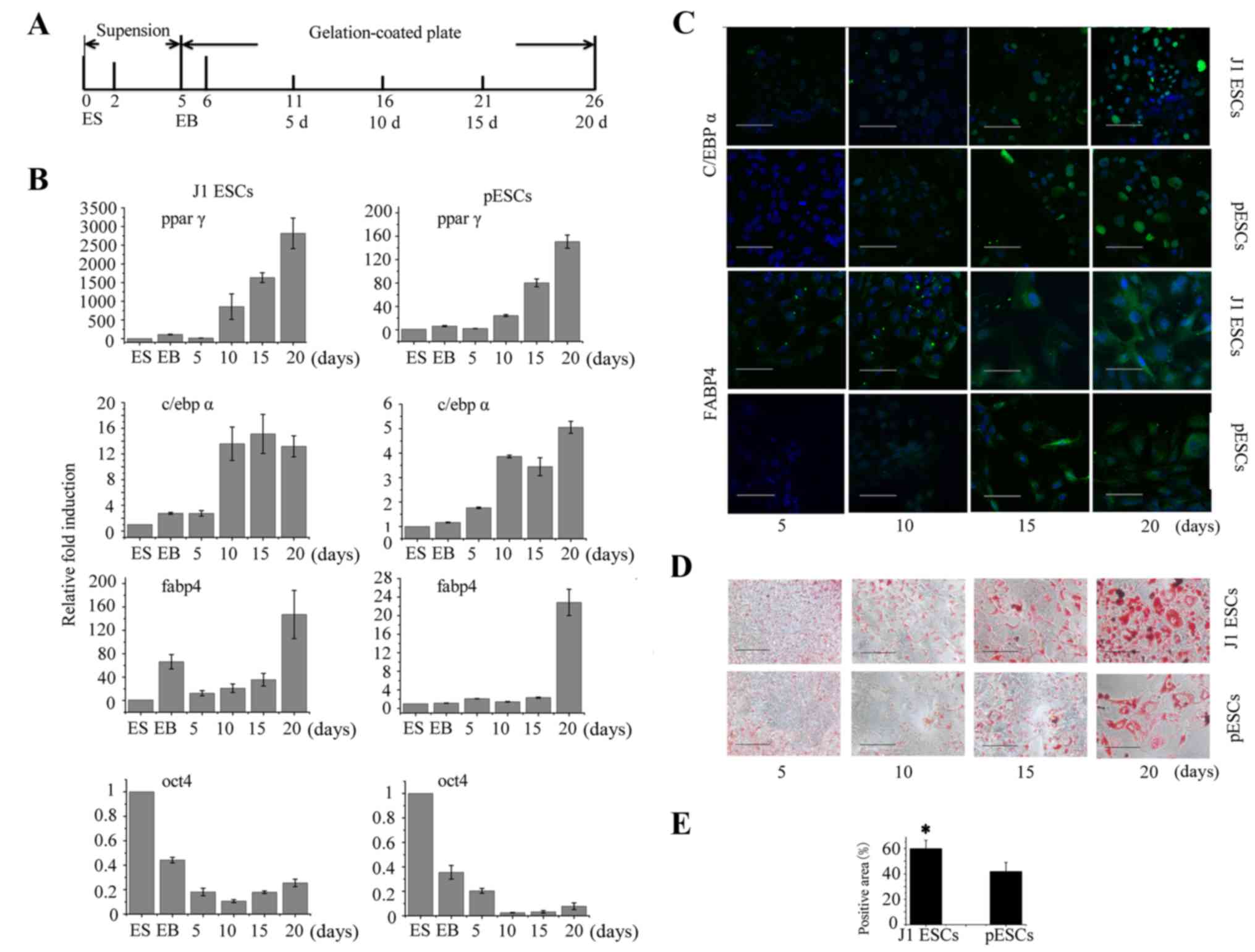 | Figure 3Adipogenesis of J1 embryonic stem
cells (ESCs) and parthenogenetic ESCs (pESCs). (A) Scheme used for
ESC differentiation into adipocytes. (B) Adipogenic differentiation
of J1 ESCs and pESCs was confirmed by upregulation of
CCATT/enhancer-binding protein α (c/ebp α), fatty acid-binding
protein 4 (fabp4) and peroxisome proliferator-activated receptor γ
(ppar γ), and downregulation of oct4 by quantitative polymerase
chain reaction at the ES and EB stages, at 5, 10, 15 and 20 days (3
repeat/timepoint; data represent means ± standard deviation). (C)
Immunofluorescence labeling of adipogenic-specific factors c/ebp α
and fabp4 during adipogenesis of J1 ESCs and pESCs in adipogenic
induction medium at 5, 10, 15 and 20 days. (D) Image of Oil red O
showing fat droplet formation at 5, 10, 15 and 20 days during
adipogenesis of J1 ESCs and pESCs. (E) Statistical analysis of Oil
red O-positive cells. Scale bar, 100 μm. ES, embryonic stem;
EB, embryoid body. |
Glycosylation alteration of J1 ESCs and
pESCs during adipogenesis
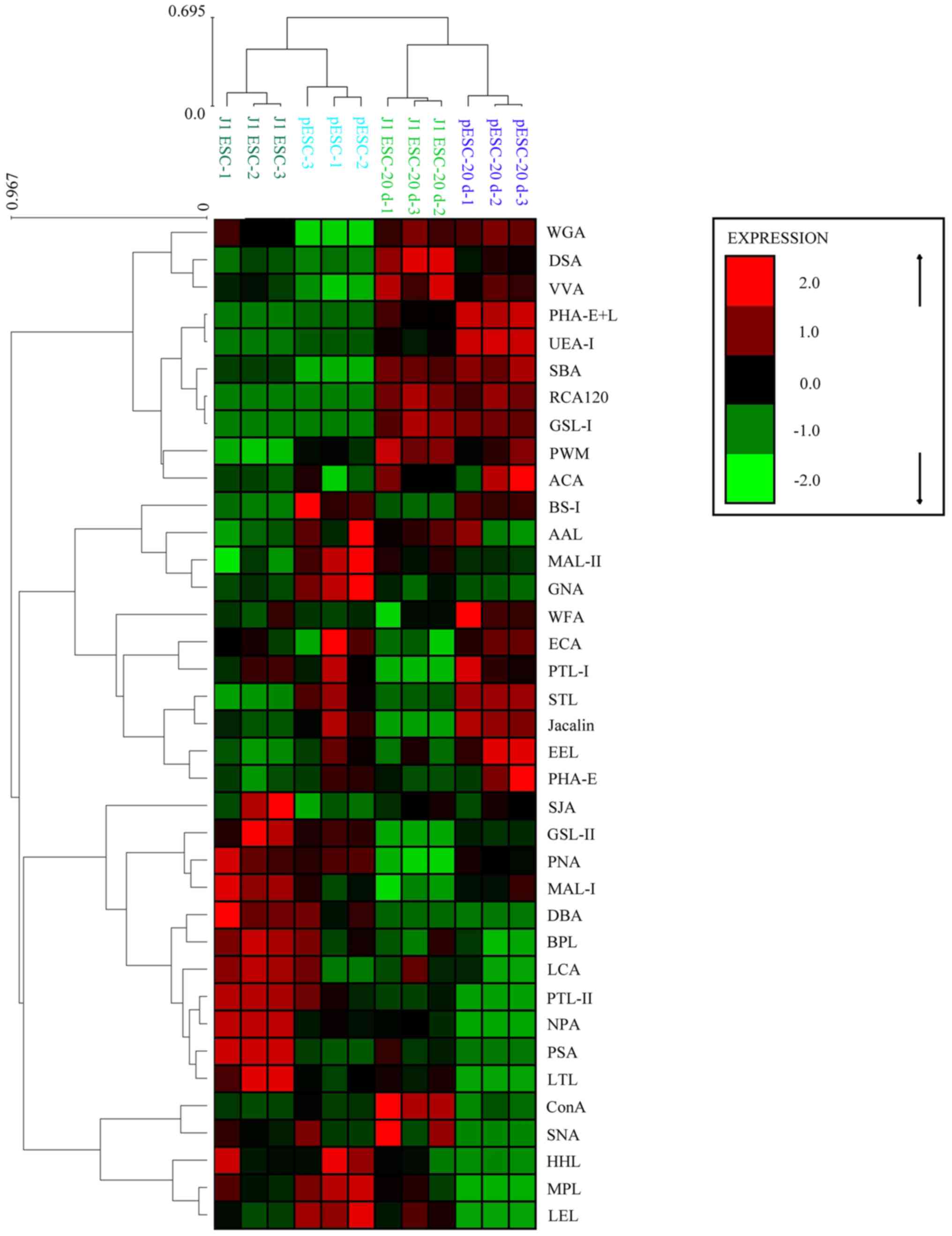
Lectin microarray chips containing 37 lectins were
used to screen the alterations of protein glycosylation of J1 ESCs
and pESCs during adipogenesis. The data were analyzed by
hierarchical clustering after being mean-normalized. As shown in
Fig. 4, J1 ESCs/pESCs and their
adipogenic lineage derivatives were clearly separated into two
large clusters, indicating that undifferentiated ESCs and their
derivatives exhibited specified glycan profiles.
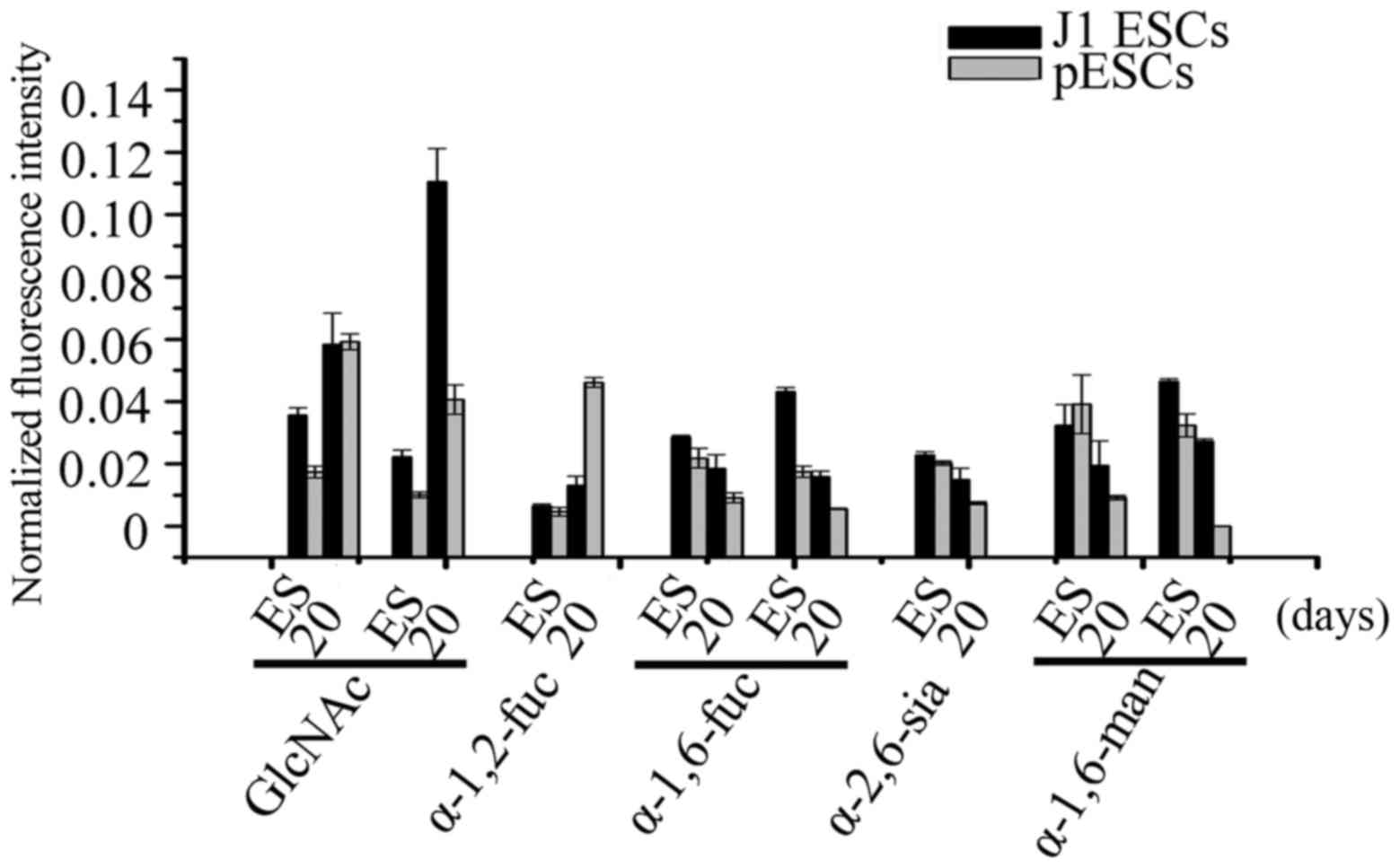
After 20 days of adipogenic differentiation, 21 of
the 37 lectins used in the lectin microarray exhibited a similar
trend of alteration in J1 ESCs and pESCs, among which the signal of
7 lectins increased, such as α-1-2-fucose-binding lectin Ulex
europaeus agglutinin-1 (UEA-1), GlcNAc-binding lectins wheat
germ agglutinin (WGA) and Datura stramonium agglutinin
(DSA). The signal of another 12 lectins decreased, including
α-1-6-fucose-binding lectins Lens culinaris agglutinin (LCA)
and Pisum sativum agglutinin (PSA),
α-2-6-sialic-acid-binding lectin Sambucus nigra agglutinin
(SNA), α-1-6-mannose-binding lectins Narcissus
pseudonarcissus (NPA) and Hippeastrum hybrid lectin
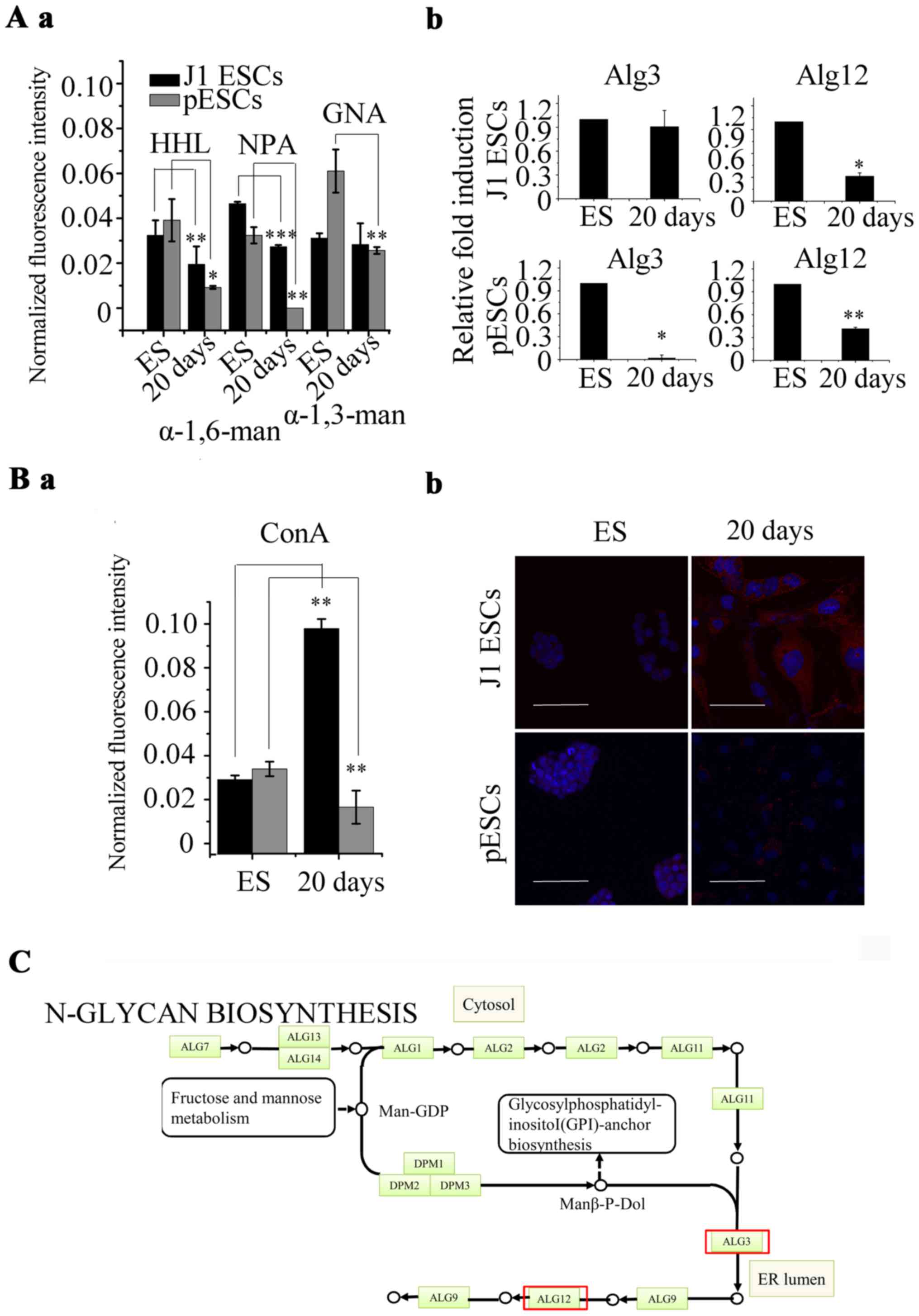
(HHL) (Fig. 5). In addition, the
signals of all 4 mannose-binding lectins we used [NPA, HHL,
Galanthus nivalis agglutinin (GNA) and concanavalin A
(ConA)] were decreased in pESCs. The signals of
α-1-6-mannose-binding lectins (NPA and HHL) were decreased, and the
signal of α-1-3-mannose-binding lectin (GNA) exhibited no
significant alteration in J1 ESCs (Fig. 6A-a). The signal of ConA was
increased in J1 ESCs (Fig. 6B-a).
The signals of the other GlcNAc lectins (identified with DSA) were
largely increased during adipogenesis in J1 ESCs (4.95-fold) and
pESCs (4.02-fold) (Fig. 5).
The expression of glycosyltransferases was
consistent with the results of the lectin microarray. The
expression of α-1-6-mannosyltransferase (Alg12) was decreased in J1
ESCs (0.35-fold) and pESCs (0.42-fold). The expression of
α-1-3-mannosyltransferase (Alg3) was decreased in pESCs
(0.02-fold), while there was no significant alteration in J1 ESCs
(Fig. 6A-b). The lectin
histochemistry of ConA was consistent with the results of the
lectin microarray. ConA displayed strong binding to the central
and/or perinuclear cytoplasm in J1 ESCs and pESCs. After 20 days of
adipogenic differentiation, the binding intensified in J1 ESC
derivatives. However, ConA exhibited a similar binding intensity in
pESC derivatives compared with undifferentiated pESCs (Fig. 6B-b). Alg12 was decreased during
adipogenesis of ESCs, while Alg3 was decreased in pESCs. Alg3 and
Alg12 are involved in the biosynthesis of the N-glycan precursor,
suggesting that the synthesis of N-glycans may decrease (Fig. 6C).
Discussion
In the present study, the alterations in the protein
glycosylation of ESCs during adipogenesis were investigated. The
level of GlcNAc residue and α-1-2-fucosylation increased,
α-1-6-fucosylation, α-2-6-sialylation and α-1-6-mannosylation
decreased in J1 ESCs and pESCs, whereas α-1-3-mannosylation
decreased in pESCs, while there was no significant alteration in J1
ESCs during adipogenesis. The protein glycosylation of ESCs was
significantly altered during adipogenesis. J1 ESCs and pESCs
exhibited obvious differences in protein glycosylation during
adipogenesis. To the best of our knowledge, this is the first study
to investigate alterations of protein glycosylation in pESCs during
adipogenesis.
Protein glycosylation, a ubiquitous
post-translational modification, exerts crucial biological effects
on protein function (28).
Protein glycosylation often modulates protein function to affect
several biological processes, including development,
differentiation, cell adhesion and cell-cell recognition. Indeed,
heparin sulfate proteoglycans have been reported as regulatory
factors of ESCs (29).
ESCs may differentiate into adipocytes using an
adipogenic cocktail containing IGF-1, glucocorticoids and cAMP.
Over the past few decades, adipogenic differentiation has been
extensively investigated (3–5).
However, the alterations in protein glycosylation during this
process remain unknown. In the present study, we analysed the
alterations in the protein glycosylation of J1 ESCs and pESCs
during adipogenesis using lectin microarray. Among the 37 lectins,
21 exhibited the same alterations in J1 ESCs and pESCs.
α-2-6-Sialic acid (identified by SNA) was decreased, which was in
agreement with a previous study reporting that the expression of
2-6-sialylated glycan is higher in undifferentiated cells (30). It was also observed that
α-1-6-mannose (identified by HHL and NPA) was decreased during
adipogenesis. All these data indicate that J1 ESCs and pESCs
exxhibited similar alterations in protein glycosylation during
adipogenesis. All these glycan structures may be used as promising
stem cell biomarkers.
Mammalian oocytes may be parthenogenetically
activated under appropriate stimuli, leading to the development of
diploid non-embryonic blastocysts (31). The pluripotent stem cells from
this type of blastocoel inner cell mass are defined as pESCs, which
lack paternal imprints. pESCs are well histocompatible (32) and their use is not associated with
ethical limitations. In the present study, J1 ESCs and pESCs
exhibited similar fundamental biological properties. Both cells
were able to form teratomas with three germ layers in vivo
and differentiate into multiple mesenchymal tissues, including
bone, cartilage and fat. These data were in agreement with the
findings reported by previous studies (26,33) and suggested that pESCs may be an
attractive source for cell therapy. Chen et al used
pESC-derived MSCs for adipogenic differentiation (26). After 3 weeks of adipogenic
induction, approximately 10% and <1% Oil red O-positive cells
were observed in the ESC-derived and pESC-derived cells,
respectively. In the present study, the proportion of Oil red
O-positive cells was up to 60±6.4% in J1 ESC derivatives and
42±6.9% in pESC derivatives, indicating that pESCs have a lower
adipogenic potential. In addition, 20 days after induction, the
upregulated level of adipocyte genes in pESCs (ppar γ, 151-fold;
c/ebp α, 5-fold; and fabp4, 23-fold) was lower compared with that
in J1 ESCs (ppar γ, 2,757-fold; c/ebp α, 13-fold; and fabP4,
147-fold); c/ebp α and fabp4 could not be detected in pESC
derivatives by immunofluorescence analysis 5 days after
induction.
Of the 37 lectins used in the lectin microarray, the
majority exhibited the same alterations in J1 ESCs and pESCs during
adipogenesis. The signal of WGA (identiying multivalent sialic acid
and GlcNAc residue) was increased in J1 ESCs and pESCs during
adipogenesis, whereas the signal of ConA (identiying GlcNAc residue
and α-mannose) was increased in J1 ESCs and decreased in pESCs. The
level of GlcNAc residue (identified by DSA) was increased in J1
ESCs and pESCs during adipogenesis. This demonstrated that level of
α-mannose identified by ConA in pESCs was decreased. Four types of
mannose-binding lectins were differentially altered in J1 ESCs and
pESCs during adipogenesis. The signals of NPA and HHL (identiying
α-1-6 mannose) were decreased in J1 ESCs and pESCs. The signal of
GNA (identiying α-1-3 mannose) was only decreased in pESCs, but
exhibited no significant alterations in J1 ESCs (P>0.05). These
data suggested that the increased signal of ConA was caused by the
increased level of GlcNAc residue but not α-mannose. These findings
were further confirmed by the expression profiles of
glycosyltransferases. The expression of Alg12 (α-1-6
mannosyltransferase) was upregulated in J1 ESCs and pESCs. The
expression of Alg3 (α-1-3 mannosyltransferase) was only
downregulated in pESCs, but exhibited no significant alterations in
J1 ESCs. The decreased expression of Alg3 and 12 are involved in
the biosynthesis of the N-glycan precursor. This may be associated
with the process of adipogenesis.
In conclusion, in the present study, it was observed
that protein glycosylation in ESCs was altered similarly during
adipogenesis. The similar alterations in J1 ESCs and pESCs may be
used as glycobiology markers of ESCs during adipogenesis.
Furthermore, J1 ESCs and pESCs exhibited obvious differences in
protein glycosylation during adipogenesis. In conclusion, the
precise protein glycosylation alterations associated with ESC
adipogenesis was analyzed and validated, which may enable a better
understanding of adipogenesis in ESCs, and may also act as markers
for monitoring adipogenic development.
Acknowledgments
This study was supported by the National Natural
Science Foundation, China (grant nos. 31270126 and 31300797) and
the Natural Science Basic Research Plan in Shaanxi Province of
China (grant no. 2013JC2-03).
Glossary
Abbreviations
Abbreviations:
|
pESCs
|
parthenogenetic embryonic stem
cells
|
|
ESCs
|
embryonic stem cells
|
|
WGA
|
wheat germ agglutinin
|
|
ConA
|
concanavalin A
|
|
NPA
|
Narcissus pseudonarcissus
|
|
HHL
|
hippeastrum hybrid lectin
|
|
GNA
|
Galanthus nivalis
|
|
MSCs
|
pluripotent mesenchymal stem cells
|
|
RA
|
retinoic acid
|
|
c/ebp
|
CCATT/enhancer-binding protein
|
|
ppar γ
|
peroxisome proliferator-activated
receptor γ
|
|
SSEA
|
stage-specific embryonic antigen
|
References
|
1
|
Shepherd PR, Gnudi L, Tozzo E, Yang H,
Leach F and Kahn BB: Adipose cell hyperplasia and enhanced glucose
disposal in transgenic mice overexpressing GLUT4 selectively in
adipose tissue. J Biol Chem. 268:22243–22246. 1993.PubMed/NCBI
|
|
2
|
Wolfgang MJ and Lane MD: Control of energy
homeostasis: role of enzymes and intermediates of fatty acid
metabolism in the central nervous system. Annu Rev Nutr. 26:23–44.
2006. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Dani C, Smith AG, Dessolin S, Leroy P,
Staccini L, Villageois P, Darimont C and Ailhaud G: Differentiation
of embryonic stem cells into adipocytes in vitro. J Cell Sci.
110:1279–1285. 1997.PubMed/NCBI
|
|
4
|
Tang QQ, Otto TC and Lane MD: Commitment
of C3H10T1/2 pluripotent stem cells to the adipocyte lineage. Proc
Natl Acad Sci USA. 101:9607–9611. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Serrero G and Lepak NM: Prostaglandin
F2alpha receptor (FP receptor) agonists are potent adipose
differentiation inhibitors for primary culture of adipocyte
precursors in defined medium. Biochem Biophys Res Commun.
233:200–202. 1997. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Bowers RR, Kim JW, Otto TC and Lane MD:
Stable stem cell commitment to the adipocyte lineage by inhibition
of DNA methylation: role of the BMP-4 gene. Proc Natl Acad Sci USA.
103:13022–13027. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Bowers RR and Lane MD: Wnt signaling and
adipocyte lineage commitment. Cell Cycle. 7:1191–1196. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Spinella-Jaegle S, Rawadi G, Kawai S,
Gallea S, Faucheu C, Mollat P, Courtois B, Bergaud B, Ramez V,
Blanchet AM, et al: Sonic hedgehog increases the commitment of
pluripotent mesenchymal cells into the osteoblastic lineage and
abolishes adipocytic differentiation. J Cell Sci. 114:2085–2094.
2001.PubMed/NCBI
|
|
9
|
van der Horst G, Farih-Sips H, Löwik CW
and Karperien M: Hedgehog stimulates only osteoblastic
differentiation of undifferentiated KS483 cells. Bone. 33:899–910.
2003. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
MacDougald OA and Lane MD: Transcriptional
regulation of gene expression during adipocyte differentiation.
Annu Rev Biochem. 64:345–373. 1995. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Student AK, Hsu RY and Lane MD: Induction
of fatty acid synthetase synthesis in differentiating 3T3-L1
preadipocytes. J Biol Chem. 255:4745–4750. 1980.PubMed/NCBI
|
|
12
|
Darlington GJ, Ross SE and MacDougald OA:
The role of C/EBP genes in adipocyte differentiation. J Biol Chem.
273:30057–30060. 1998. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Ailhaud G, Grimaldi P and Négrel R:
Cellular and molecular aspects of adipose tissue development. Annu
Rev Nutr. 12:207–233. 1992. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Apweiler R, Hermjakob H and Sharon N: On
the frequency of protein glycosylation, as deduced from analysis of
the SWISS-PROT database. Biochim Biophys Acta. 1473:4–8. 1999.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Lowe JB and Marth JD: A genetic approach
to mammalian glycan function. Annu Rev Biochem. 72:643–691. 2003.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Hwang HY, Olson SK, Esko JD and Horvitz
HR: Caenorhabditis elegans early embryogenesis and vulval
morphogenesis require chondroitin biosynthesis. Nature.
423:439–443. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Crocker PR: Siglecs: sialic-acid-binding
immunoglobulin-like lectins in cell-cell interactions and
signalling. Curr Opin Struct Biol. 12:609–615. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Collins BE and Paulson JC: Cell surface
biology mediated by low affinity multivalent protein-glycan
interactions. Curr Opin Chem Biol. 8:617–625. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Liu D, Shriver Z, Venkataraman G, El
Shabrawi Y and Sasisekharan R: Tumor cell surface heparan sulfate
as cryptic promoters or inhibitors of tumor growth and metastasis.
Proc Natl Acad Sci USA. 99:568–573. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Sasisekharan R, Shriver Z, Venkataraman G
and Narayanasami U: Roles of heparan-sulphate glycosaminoglycans in
cancer. Nat Rev Cancer. 2:521–528. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Muramatsu T and Muramatsu H: Carbohydrate
antigens expressed on stem cells and early embryonic cells.
Glycoconj J. 21:41–45. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Qin Y, Zhong Y, Dang L, Zhu M, Yu H, Chen
W, Cui J, Bian H and Li Z: Alteration of protein glycosylation in
human hepatic stellate cells activated with transforming growth
factor-β1. J Proteomics. 75:4114–4123. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Ito H, Kuno A, Sawaki H, Sogabe M, Ozaki
H, Tanaka Y, Mizokami M, Shoda J, Angata T, Sato T, et al: Strategy
for glycoproteomics: identification of glyco-alteration using
multiple glycan profiling tools. J Proteome Res. 8:1358–1367. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Yu H, Zhu M, Qin Y, Zhong Y, Yan H, Wang
Q, Bian H and Li Z: Analysis of glycan-related genes expression and
glycan profiles in mice with liver fibrosis. J Proteome Res.
11:5277–5285. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Hwang NS, Kim MS, Sampattavanich S, Baek
JH, Zhang Z and Elisseeff J: Effects of three-dimensional culture
and growth factors on the chondrogenic differentiation of murine
embryonic stem cells. Stem Cells. 24:284–291. 2006. View Article : Google Scholar
|
|
26
|
Chen Y, Ai A, Tang ZY, Zhou GD, Liu W, Cao
Y and Zhang WJ: Mesenchymal-like stem cells derived from human
parthenogenetic embryonic stem cells. Stem Cells Dev. 21:143–151.
2012. View Article : Google Scholar
|
|
27
|
Scillitani G, Zizza S, Liquori GE and
Ferri D: Lectin histochemistry of gastrointestinal glycoconjugates
in the greater horseshoe bat, Rhinolophus ferrumequinum (Schreber,
1774). Acta Histochem. 109:347–357. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Moremen KW, Tiemeyer M and Nairn AV:
Vertebrate protein glycosylation: diversity, synthesis and
function. Nat Rev Mol Cell Biol. 13:448–462. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Sasaki N, Okishio K, Ui-Tei K, Saigo K,
Kinoshita-Toyoda A, Toyoda H, Nishimura T, Suda Y, Hayasaka M,
Hanaoka K, et al: Heparan sulfate regulates self-renewal and
pluripotency of embryonic stem cells. J Biol Chem. 283:3594–3606.
2008. View Article : Google Scholar
|
|
30
|
Satomaa T, Heiskanen A, Mikkola M, Olsson
C, Blomqvist M, Tiittanen M, Jaatinen T, Aitio O, Olonen A, Helin
J, et al: The N-glycome of human embryonic stem cells. BMC Cell
Biol. 10:422009. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Paffoni A, Brevini TA, Gandolfi F and
Ragni G: Parthenogenetic activation: biology and applications in
the ART laboratory. Placenta. 29(Suppl B): 121–125. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Kim K, Lerou P, Yabuuchi A, Lengerke C, Ng
K, West J, Kirby A, Daly MJ and Daley GQ: Histocompatible embryonic
stem cells by parthenogenesis. Science. 315:482–486. 2007.
View Article : Google Scholar
|
|
33
|
Didié M, Christalla P, Rubart M, Muppala
V, Döker S, Unsöld B, El-Armouche A, Rau T, Eschenhagen T,
Schwoerer AP, et al: Parthenogenetic stem cells for
tissue-engineered heart repair. J Clin Invest. 123:1285–1298. 2013.
View Article : Google Scholar : PubMed/NCBI
|