Introduction
Colorectal cancer (CRC) is the third most common
type of cancer in China (1). CRC
tumorigenesis occurs when mutations accumulate in critical
oncogenes and tumor suppressor genes (2,3).
Hypoxia is a common feature in numerous solid tumors and the
microenvironment is now recognized as a key factor associated with
biologically aggressive phenotypes in cancer (4). Hypoxia can induce formation of
reactive oxygen species (ROS), which may induce genotoxic effects
or cancer cell apoptosis (5). In
addition, hypoxia can induce endoplasmic reticulum (ER) stress and
activate the unfolded protein response (UPR), which is an adaptive
response that increases cancer cell survival under ER stress
(6,7). The main pathways of ER stress
include three functional components: i) Transcriptional
upregulation of specific genes to tackle unfolded and misfolded
proteins, namely, the UPR; ii) inhibition of translation to reduce
the load of client proteins; and iii) triggering apoptosis when ER
functions are severely impaired (8-10).
Thioredoxin domain containing 5 (TXNDC5), which is a
member of the disulfide isomerase family, is predominantly
expressed in the ER (11).
Accumulating evidence has suggested that TXNDC5 is induced by
hypoxia in endothelial cells and tumor endothelium (12). It has been reported that TXNDC5
facilitates proteins to fold correctly by the formation of
disulfide bonds through its thioredoxin domains (13); this can protect cells from ER
stress-induced apoptosis (12,14). In addition to upregulation in
numerous types of cancer, including hepatocellular, breast,
cervical, esophageal, liver, lung, stomach and uterine carcinoma
(15), TXNDC5 is overexpressed in
CRC (16,17) and is considered an oncogene
(18). However, the role of
TXNDC5 in CRC tumorigenesis remains unclear.
The present study aimed to explore the role of
TXNDC5 in CRC tumorigenesis in vitro and in vivo
under hypoxic and normoxic conditions.
Materials and methods
Patient samples
Tumor and paired normal mucosal tissues were
surgically removed from 102 patients with CRC at the Xiangya
Hospital of Central South University (Changsha, China) between
January 2009 and December 2010. Fresh tissues underwent reverse
transcription-quantitative polymerase chain reaction (RT-qPCR) and
western blot analysis. The sections were fixed with 4%
polyformaldehyde phosphate buffer for 12 h at 37°C. In addition,
samples were embedded in paraffin for immunohistochemistry. Normal
mucosal tissues were excised 5 cm away from the tumor. The
histomorphology of tumor and normal mucosa were confirmed by
pathologists. The present study was approved by the Medical Ethics
Committee of Xiangya Hospital and all patients provided written
informed consent.
Cell culture and treatment
The human CRC cell lines RKO and HCT-116 reportedly
express high levels of HIF-1α under hypoxia (20). These cell lines were purchased
from the American Type Culture Collection (ATCC, Manassas, VA,
USA). All cells were cultured in RPMI-1640 medium (Biological
Industries, Beit-Haemek, Israel) supplemented with 10% fetal bovine
serum (FBS; Biological Industries). Cells were cultured at 37°C in
a humidified atmosphere containing 5% CO2.
RT-qPCR analysis
Total RNA was extracted from cells (RKO and HCT-116
cells) and tissues using TRIzol® reagent (Invitrogen;
Thermo Fisher Scientific, Inc., Waltham, MA, USA), and equal
amounts of RNA underwent RT-qPCR analysis (PCR kit, SYBR-Green/ROX
Mastermix; Takara Bio, Inc., Otsu, Japan) according to the
manufacturer's protocol. Firstly, Total RNA underwent DNase
treatment with gDNA Eraser buffer and gDNA Eraser. The RNA without
DNA was then reversed into cDNA according to the manufacturer's
protocol (PrimeScript RT reagent kit with gDNA Eraser; Takara Bio,
Inc.). Finally, the cDNA, qPCR reagent (SYBR Premix Ex Taq II;
Takara Bio, Inc.) and primer were mixed together for thermocycling,
and the thermocycling conditions were according to the
manufacturer's operation guidelines as follows: cDNA 2 µl,
qPCR mix 10 µl, forward primer 0.6 µl, reverse primer
0.6 µl, ROX 0.3 µl, DEPC water, 6.5 µl, STEP1
95°C 30 sec, STEP2 95°C 5 sec, STEP3 60°C 35 min STEP4 Go To STEP2
for 30 cycles, STEP5: Melt Curve Stage, STEP6:4°C end. β-actin was
used as an internal control. The primer sequences used are as
follows: TXNDC5 (human), sense 5′-GGG TCA AGA TCG CCG AAGTA-3′,
antisense 5′-GCC TCC ACT GTG CTC ACTGA-3′; TXNDC5 (mouse), sense
5′-CGC ACT TCG TCA TGT TCT TCG-3′, antisense 5′-CAG AGC ACA CGT CGG
AATCA-3′; hypoxia-inducible factor-1α (HIF-1α) (human), sense
5′-ATC GCG GGG ACC GATT-3′, antisense 5′-CGA CGT TCA GAA CTT ATC
TTT TTCTT-3′; and β-actin, sense 5′-GCA CCA CAC CTT CTA CAA
TGAGC-3′ and antisense 5′-GGA TAG CAC AGC CTG GAT AGC AAC-3′. The
2−ΔΔCq method was used to calculate the relative
abundance of mRNA for each gene compared with β-actin expression
(19). Each experiment was perfo
rmed in triplicate.
Western blot analysis
Tissue samples and cancer cells (RKO and HCT-116
cells) were homogenized and lysed in radioimmunoprecipitation assay
buffer (Sigma-Aldrich, St. Louis, MO, USA) supplemented with
proteinase inhibitors. Equal amounts of protein (2
µg/µl) were loaded and separated by 10% SDS-PAGE. The
protein concentration was determined by BCA. Subsequently, proteins
were transferred to a polyvinylidene fluoride membrane (EMD
Millipore, Billerica, MA, USA) and the membrane was blocked in 5%
(w/v) non-fat milk. The membrane was then incubated with primary
antibodies overnight at 4°C, followed by incubation with a
secondary antibody (anti-rabbit IgG HRP-linked secondary antibody,
7074S; 1:2,000 dilution; Cell Signaling Technology, Inc., Danvers,
MA, USA) for 1 h at room temperature. Bands were visualized using
the Enhanced Chemiluminescence Advance Detection system (EMD
Millipore). Blots were semi-quantified using Image Lab V4.0
(Bio-Rad, Hercules, CA, USA). The dilution of primary antibodies
were as follows: TXNDC5 (ENT2133), 1:2,000; HIF-1α (ENT2133),
1:1,000; glucose-regulated protein 78 (GRP78; ENT2245), 1:1,000;
CCAAT-enhancer-binding protein homologous protein (CHOP; ENT0911),
1:1,000; activating transcription factor 4 (ATF4; EAP0008),
1:1,000; B-cell lymphoma 2 (Bcl-2)-associated X protein (Bax;
ENT0456), 1:1,000; Bcl-2 (ENT0469), 1:500; cleaved caspase-8
(ENC011), 1:1,000; and β-actin (4970), 1:1,000.
Immunohistochemistry
Tissue slides were prepared as follows: The tissue
sections (pruned into 7-10-mm diameter and 5-7 µm thick)
were incubated in 3 washes of xylene for 5 min each and then
incubated in 2 washes of 100% ethanol for 10 min each and in 2
washes of 95% ethanol for 10 min each. The sections were then
washed twice in dH2O for 5 minutes each and the slides
were brought to boil in 1 mM EDTA (pH 8.2) followed by 4 min at a
sub-boiling temperature. No cooling was necessary. The sections
were washed in dH2O 3 times for 5 min each and then
incubated in 3% hydrogen peroxide for 15 min. Afterwards, the
sections were washed in dH2O twice for 5 min each.
Tissue slides were deparaffinized and antigen retrieval was
performed by immersing the slides in boiling EDTA-Tris buffer (pH
8.2) for 4 min. Following incubation with 3%
H2O2 for 15 min, slides were incubated with
primary antibodies against TXNDC5 (ENT2133; 1:100) and HIF-1α
(ENT2133; 1:100) (both from Wuhan Elabscience Biotechnology Co.,
Ltd., Wuhan, China) overnight at 4°C, followed by incubation with
the secondary antibody (SA00004-2; 1:500; Proteintech; Nanjing,
China) for 30 min at room temperature. 3,3′-Diaminobenzidine
(0.05%) was used for 5-10 min at room temperature to visualize a
positive immune reaction. Nuclei were counterstained with
hematoxylin. The microscope used was from Leica (Wetzlar, Germany).
Briefly, HIF-1α and TXNDC5 expression levels were scored according
to the percentage of positively stained cells and staining
intensity: (−) or 0, no staining (0-10%); (+) or 1, weak staining
(10-25%); (++) or 2, moderate staining (25-50%) and (+++) or 3,
strong staining (>50%). (−) and (+) were defined as low
expression, and (++) and (+++) were defined as high expression. The
staining results were independently evaluated by two
board-certified clinical pathologists blinded to the clinical
parameters. Any discrepancy between the two evaluators was resolved
by re-evaluation and careful discussion until agreement was
reached.
Hypoxia treatment [1% O2 or
cobalt chloride (CoCl2)]
Cells were cultured in 96-well plates, or 60- or
100-mm petri dishes. For each assay, RKO and HCT-116 cells were
cultured in RPMI-1640 containing 10% FBS for 24 h. For hypoxic
treatment, cells were exposed to either 1% O2 or to
various concentrations of the hypoxia-mimicking agent
CoCl2 (PhytoTechnology Laboratories, Lenexa, KS, USA)
for various time intervals, as previously described (21).
MTT cell proliferation assay
In vitro cell proliferation was determined
using a MTT Cell Proliferation Assay kit (ATCC) according to the
manufacturer's protocol. Briefly, cells were plated at
15×103 cells/well in 96-well tissue culture plates. At
the end of the culture period, cells were washed with PBS, and the
MTT reagents were added according to the manufacturer's protocol.
Subsequently, formazan was dissolved using DMSO and the absorbance
was then measured at 570 nm using an ELISA plate reader. Each
experiment was repeated three times in triplicate.
Colony formation assay
Cells were seeded at a density of 500 cells/6-well
plates (Corning Inc., Corning, NY, USA) in triplicate and were
cultured for 14 days. During colony growth assays, the culture
medium was replaced every 3 days. Colonies with >50 cells were
counted when visible to naked eye and stained with GIEMSA under a
light microscope.
Cell apoptosis assay
Cells were seeded at 1×105 cells/well in
96-well tissue culture plates. Cell apoptosis was measured at 24 h
using a microplate reader-based TiterTACS in situ apoptosis
detection kit (R&D Systems, Inc., Minneapolis, MN, USA)
according to the manufacturer's protocol. The percentage of
apoptotic cells was calculated.
Measurement of ROS
The intracellular total ROS Activity Assay kit
(Abnova, Walnut, CA, USA) was used to detect ROS levels, according
to the manufacturer's protocol. Briefly, the treated cells were
lysed and the amount of intracellular ROS was calculated according
to dichlorodihydrofluorescein production, which was measured using
a fluorometric plate reader at excitation and emission wavelengths
of 490 and 525 nm, respectively.
Transient transfection of small
interfering (si)RNA
Cells (3.0×105 per well) were seeded into
24-well plates at a density of 1.5×105 cells/ml, and
allowed to reacĥ50% confluence on the day of transfection. Cells
were transfected with 100 nmol/l of each siRNA duplex for 6 h at
37°C using Lipofectamine® 2000 reagent (Invitrogen;
Thermo Fisher Scientific, Inc.) according to the manufacturer's
protocol. Scrambled control siRNA (siScr), siHIF-1α
(AGCCATTTACATAATATAGAA) and siTXNDC5 (ATCGAGCTACTTCCCATAATA) were
all purchased from Guangzhou RiboBio Co., Ltd. (Guangzhou,
China).
Azoxymethane (AOM)-induced colorectal
tumorigenesis model
AOM was used to induce a sporadic colorectal
tumorigenesis model, as previously described (22). Briefly, 8 week-old male mice
[n=18; A/J strain; divided into 3 groups; weight, >18 g;
maintenance conditions: temperature, 18-29°C; relative humidity,
50-60%; free access to clean food and water; lighting for 10-14 h
(lights turned on at 8:00 every day and turned off at 18:00)] were
intraperitoneally injected with 10 mg/kg body weight AOM
(Sigma-Aldrich) six times. The mice were injected with AOM every
week from the first to the sixth week. The mice were then provided
with regular water and did not undergo any further treatment. Mice
were sacrificed at various time-points, after which the colorectum
was excised, opened longitudinally, flushed with ice-cold PBS and
fixed in 10% formalin/PBS for 24 h at room temperature.
Subsequently, macroscopic tumors were counted and morphological
evaluation of normal, adenoma and tumor tissues were performed
under a light microscope. All mice were divided into 3 groups; the
blank control group was treated with normal saline, the adenoma and
tumor groups were treated with AOM. The normal tissues were
obtained from the blank control group, and the adenoma tissues were
obtain from mice treated with AOM at the end of the 12th week. All
animals received humane care according to the Institutional Animal
Care and Treatment Committee of Central South University, and the
proof number of ethics was 201603426.
Statistical analysis
Statistical analysis of the differences in mRNA
expression levels of HIF-1α and TXNDC5 between paired tissues were
analyzed by paired t-test. The association between target gene
expression and clinicopathological factors was estimated by
χ2 test. The statistical differences among ≥2 groups
were analyzed using one-way analysis of variance followed by post
hoc pairwise comparisons (Tukey's post hoc test). The experiments
were repeated 3 times. The correlation between HIF-1α and TXNDC5
expression was determined by Spearman's correlation analysis. All
data are expressed as the means ± standard error. P<0.05 was
considered to indicate a statistically significant difference. The
statistical software programs GraphPad Prism 6.0 (GraphPad
Software, Inc., La Jolla, CA, USA) and SPSS 13.0 for Windows (SPSS,
Inc., Chicago, IL, USA) were used to analyze data.
Results
TXNDC5 upregulation is associated with
HIF-1α over-expression in human CRC specimens
The expression levels of HIF-1α and TXNDC5 were
detected in CRC tissues from patients. As shown in Fig. 1A, the mRNA expression levels of
HIF-1α and TXNDC5 were increased in CRC tissues compared with in
paired normal mucosal tissues. In addition, TXNDC5 mRNA expression
was positively correlated with HIF-1α mRNA expression in CRC
tissues (r=0.435, P<0.0001). The protein expression levels of
HIF-1α and TXNDC5 were detected in randomly selected CRC samples by
western blot analyses. As shown in Fig. 1B, CRC tissues exhibited higher
expression levels of TXNDC5 compared with in paired normal tissues.
As presented in Fig. 1C, analyses
of the staining intensity in immunohistochemistry images indicated
that the expression of TXNDC5 was highly correlated with the
expression of HIF-1α in CRC tissues (r=20.508, P<0.0001; data
not shown). In addition, strong HIF-1α and TXNDC5 staining was
revealed to be associated with advanced TNM stage (Fig. 1D). As shown in Table I, the expression levels of HIF-1α
and TXNDC5 were significantly associated with the TNM stage of CRC,
whereas no significant association was found with other
clinicopathological variables, including age, gender, tumor
location, tumor size, tumor differentiation, adjuvant therapy and
carcinoembryonic antigen levels.
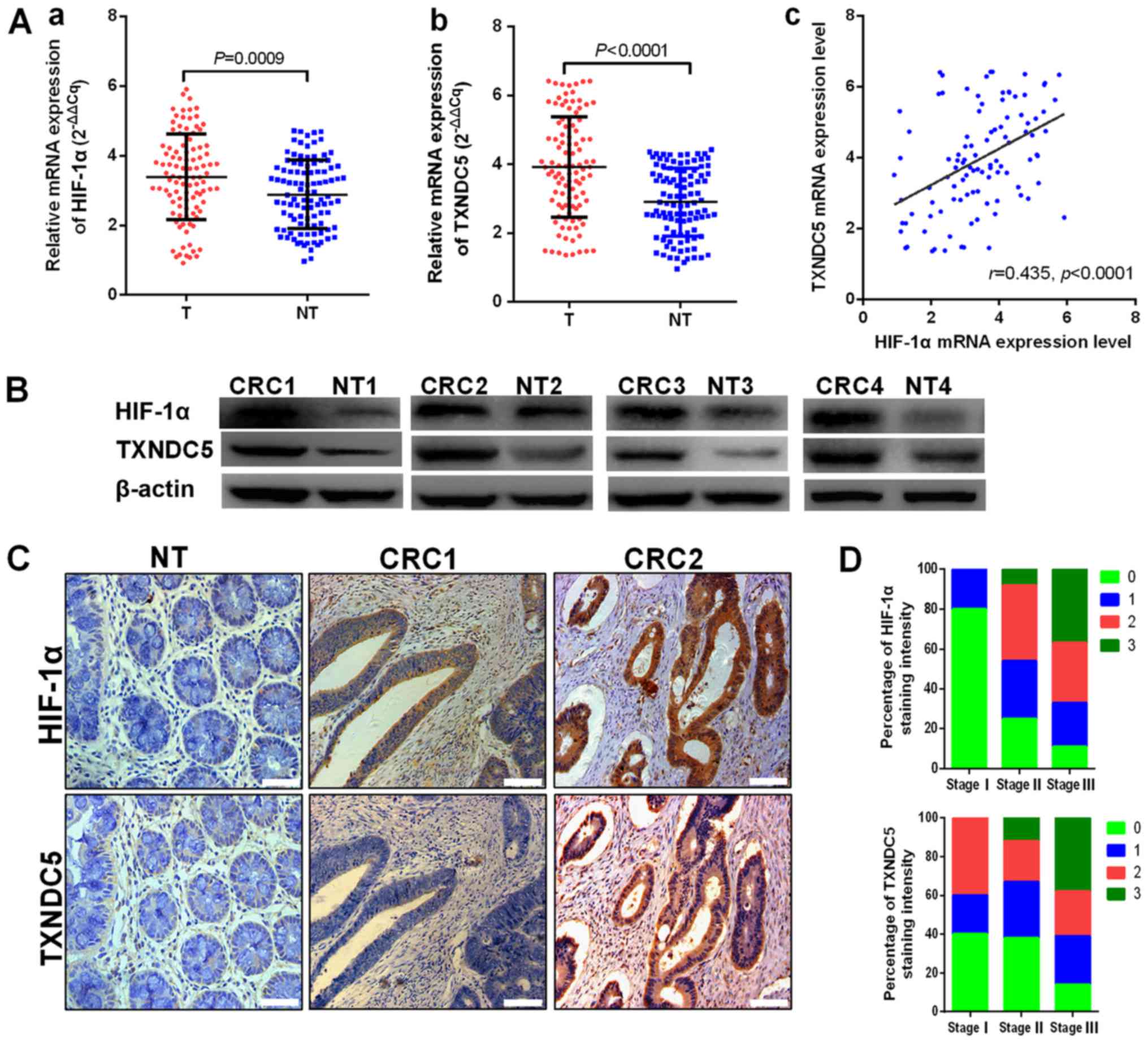 | Figure 1TXNDC5 upregulation is associated
with HIF-1α overexpression in human CRC tissues. (A) mRNA
expression levels of (a) HIF-1α and (b) TXNDC5 in T and NT were
detected by reverse transcription-quantitative polymerase chain
reaction in patients with CRC. (c) TXNDC5 mRNA expression was
positively correlated with HIF-1α mRNA expression (r=0.435,
P<0.0001). (B) Western blot analyses were used to detect the
protein expression levels of HIF-1α and TXNDC5 in randomly selected
CRC tissues and NT. (C) Expression levels of HIF-1α and TXNDC5 were
examined by immunohistochemistry in CRC tissues and NT from 102
patients with CRC. Images show negative staining of HIF-1α and
TXNDC5 in NT; negative or low staining of HIF-1α and TXNDC5 in
CRC1; and strong staining of HIF-1α and TXNDC5 in CRC2.
Magnification, ×200; scale bar, 100 µm. (D) Strong staining
of HIF-1α and TXNDC5 was associated with advanced TNM stage. Scores
indicate staining intensity: 0, negative; 1, weak staining; 2,
intermediate staining; and 3, strong staining. CRC, colorectal
cancer; HIF-1α, hypoxia-inducible factor-1α; NT, paired normal
tissues; T, tumor tissues; TXNDC5, thioredoxin domain-containing
5. |
 | Table ICorrelation between
clinicopathological features and HIF-1α or TXNDC5 protein
expression in 102 patients with CRC. |
Table I
Correlation between
clinicopathological features and HIF-1α or TXNDC5 protein
expression in 102 patients with CRC.
| Clinicopathological
variable | n | HIF-1α expression
| P-value | TXNDC5 expression
| P-value |
|---|
| Low (n=42) | High (n=60) | Low (n=48) | High (n=54) |
|---|
| Age (years) | | | | 1.000 | | | 0.099 |
| <60 | 63 | 26 (41.3%) | 37 (58.7%) | | 26 (41.3%) | 37 (58.7%) | |
| ≥60 | 39 | 16 (41.0%) | 23 (59.0%) | | 22 (56.4%) | 17 (43.6%) | |
| Gender | | | | 0.432 | | | 0.843 |
| Male | 58 | 26 (44.8%) | 32 (55.2%) | | 28 (48.3%) | 30 (51.7%) | |
| Female | 44 | 16 (36.4%) | 28 (63.6%) | | 20 (45.5%) | 24 (54.5%) | |
| Location | | | | 0.329 | | | 0.424 |
| Left colon | 38 | 19 (50.0%) | 19 (50.0%) | | 21 (55.3%) | 17 (44.7%) | |
| Right colon | 25 | 10 (40.0%) | 15 (60.0%) | | 10 (40.0%) | 15 (60.0%) | |
| Rectum | 39 | 13 (33.3%) | 26 (66.7%) | | 17 (43.6%) | 22 (56.4%) | |
| Size (cm) | | | | 0.687 | | | 1.000 |
| <5.0 | 47 | 19 (38.3%) | 28 (61.7%) | | 22 (46.8%) | 25 (53.2%) | |
| ≥5.0 | 55 | 24 (43.6%) | 31 (56.4%) | | 26 (47.3%) | 29 (52.7%) | |
|
Differentiation | | | | 0.510 | | | 0.661 |
| Well-moderate | 74 | 29 (39.2%) | 45 (60.8%) | | 36 (48.6%) | 38 (51.4%) | |
| Poor | 28 | 13 (46.4%) | 15 (53.6%) | | 12 (42.9%) | 16 (57.1%) | |
| CEA | | | | 0.071 | | | 0.824 |
| <5 ng/ml | 75 | 35 (46.7%) | 40 (53.3%) | | 36 (48.0%) | 39 (52.0%) | |
| ≥5 ng/ml | 27 | 7 (25.9%) | 20 (74.1%) | | 12 (44.4%) | 15 (55.6%) | |
| Adjuvant
therapy | | | | 0.384 | | | 0.667 |
| Yes | 31 | 15 (48.4%) | 16 (51.6%) | | 16 (51.6%) | 15 (48.4%) | |
| No | 71 | 27 (38.0%) | 44 (62.0%) | | 32 (45.1%) | 39 (54.9%) | |
| TNM stage | | | | 0.008a | | | 0.027a |
| I-II | 29 | 18 (62.1%) | 11 (37.9%) | | 19 (65.5%) | 10 (34.5%) | |
| III | 73 | 24 (32.9%) | 49 (67.1%) | | 29 (39.7%) | 44 (60.3%) | |
| T stage | | | | 0.036a | | | 0.035a |
| T1-T2 | 13 | 9 (69.2%) | 4 (30.8%) | | 10 (76.9%) | 3 (23.1%) | |
| T3-T4 | 89 | 33 (37.1%) | 56 (62.9%) | | 38 (42.7%) | 51 (57.3%) | |
| N stage | | | | 0.023a | | | 0.045a |
| N0 | 28 | 17 (60.7%) | 11 (39.3%) | | 18 (64.3%) | 10 (35.7%) | |
| N1-N2 | 74 | 25 (33.8%) | 49 (66.2%) | | 30 (40.5%) | 44 (59.5%) | |
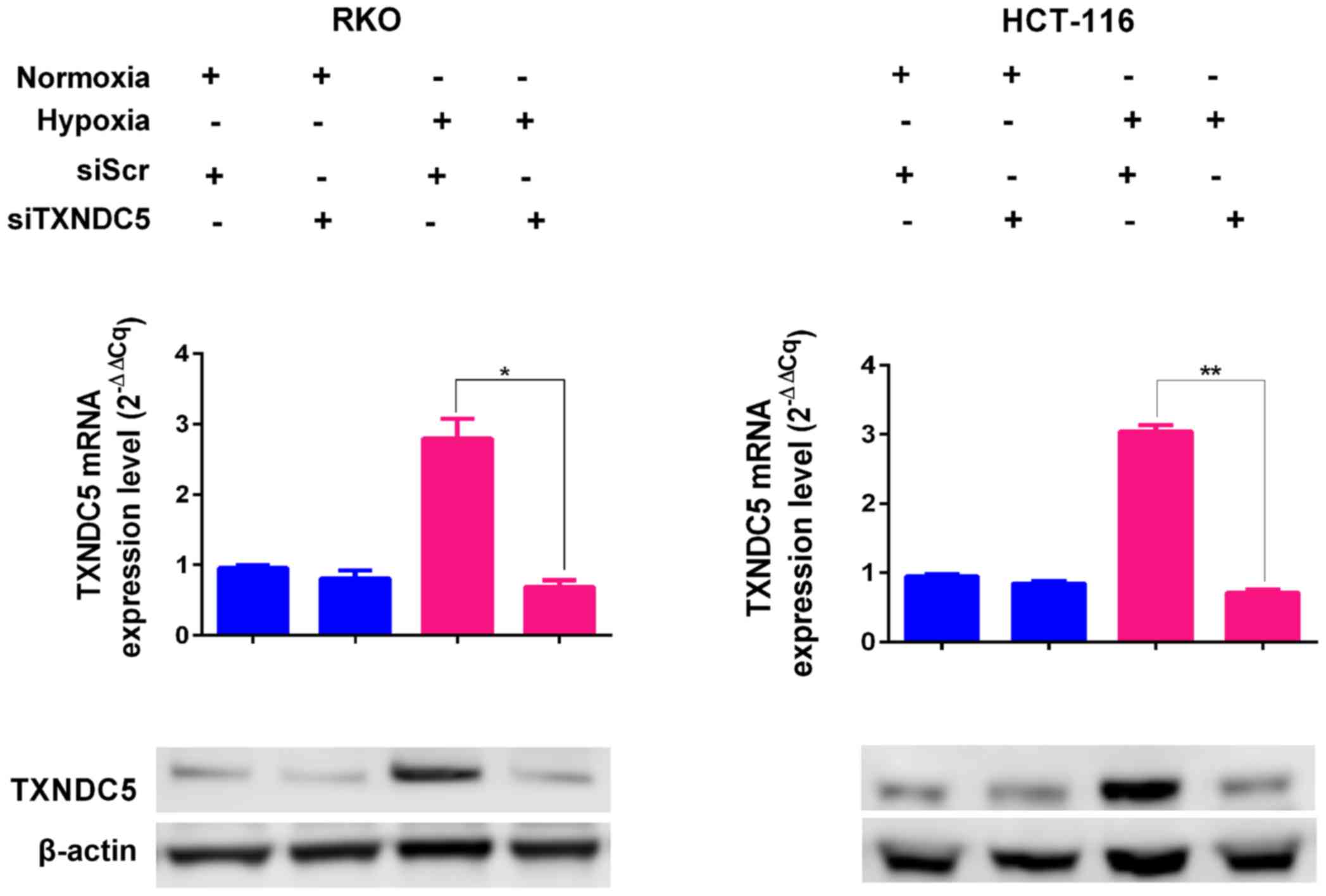
Hypoxia induces the expression of TXNDC5
via upregulating HIF-1α in CRC cell lines
To examine whether HIF-1α induced the expression of
TXNDC5, RKO and HCT-116 human CRC cells were cultured under hypoxic
conditions (1% O2) in a sealed hypoxic chamber (23). As shown in Fig. 2A, the expression levels of HIF-1α
and TXNDC5 reached a peak after 24 h under hypoxia. The cells were
also treated with various concentrations of CoCl2, which
is a HIF prolyl hydroxylase antagonist (24) that was used as a positive control
for HIF-1α induction. As shown in Fig. 2B, 1 µM CoCl2 induced the
highest expression of TXNDC5; similar results were observed with
regards to the expression of HIF-1α. Subsequently, a specific siRNA
(siHIF-1α) was used to knockdown HIF-1α by ~65% in RKO cells and
~55% in HCT-116 cells under hypoxic conditions. As shown in
Fig. 3, knockdown of HIF-1α
abolished the hypoxia-induced expression of TXNDC5 in RKO and
HCT-116 cells.
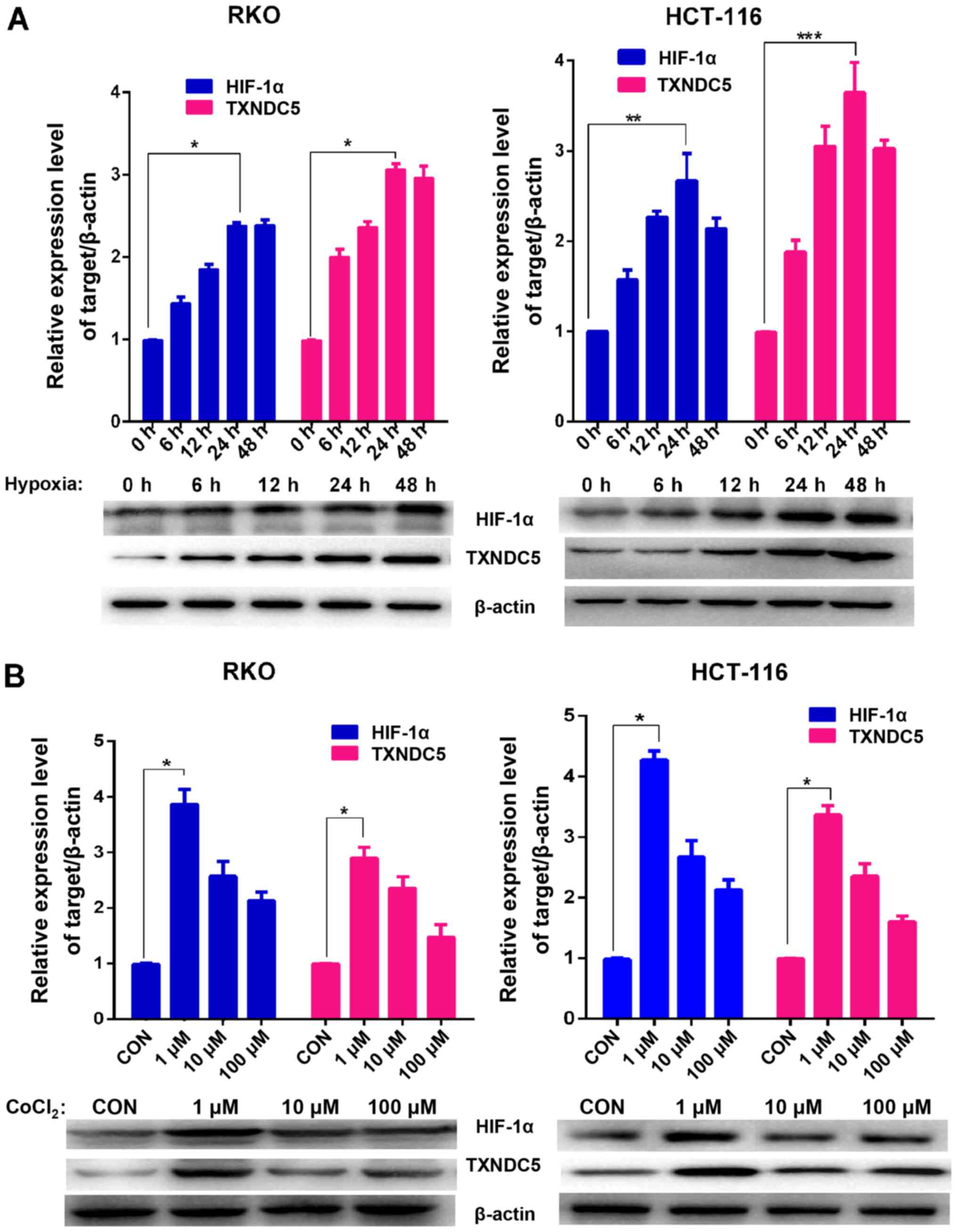 | Figure 2TXNDC5 expression is increased in CRC
cells during hypoxia. (A) Western blot analyses were used to detect
the protein expression levels of HIF-1α and TXNDC5 in RKO and
HCT-116 CRC cells under hypoxia (1% O2). β-actin was
used as a loading control. Density of the HIF-1α or TXNDC5 blot was
normalized against that of the β-actin blot, in order to obtain
relative blot density, which was expressed as a fold change to that
of the control (0 h) (designated as 1). *P<0.0001;
**P=0.0007 and ***P=0.0002. (B) RKO and
HCT-116 cells were treated with 1, 10 or 100 µM
CoCl2 for 48 h. Western blot analyses were used to
detect the protein expression levels of HIF-1α and TXNDC5. β-actin
was used as a loading control. Density of the HIF-1α or TXNDC5 blot
was normalized against that of the β-actin blot, in order to obtain
relative blot density, which was expressed as a fold change to that
of the control (untreated cells) (designated as 1).
*P<0.0001. CoCl2, cobalt chloride; CON,
control; CRC, colorectal cancer; HIF-1α, hypoxia-inducible
factor-1α; TXNDC5, thioredoxin domain-containing 5. |
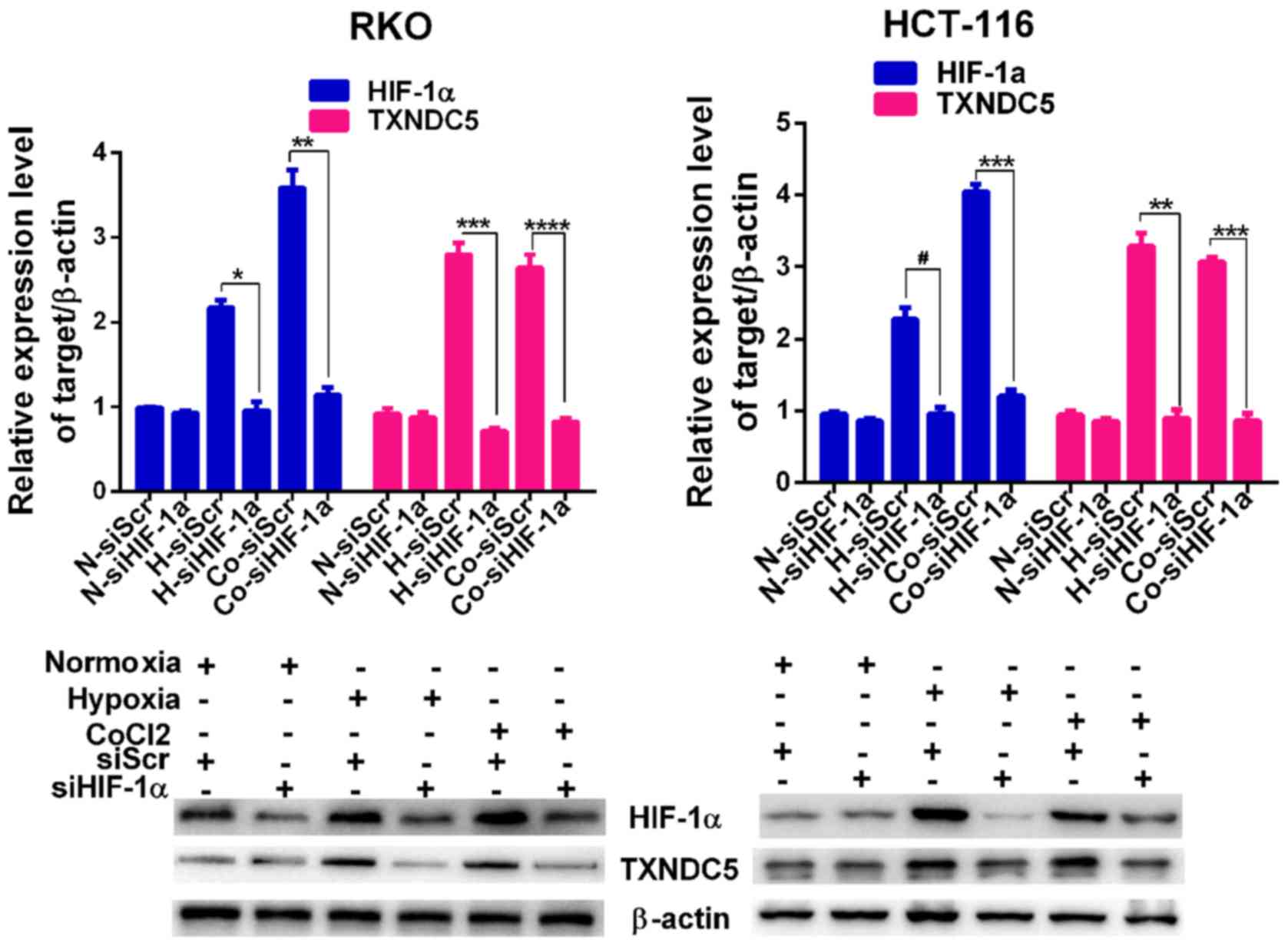 | Figure 3TXNDC5 expression is regulated by
HIF-1α in CRC cells. RKO and HCT-116 cells were transfected with a
specific siRNA (siHIF-1α) to knockdown HIF-1α. Cells transfected
with siScr were used as a control. The expression levels of HIF-1α
and TXNDC5 were detected by western blotting in cells under N, 24 h
H and CoCl2 treatment (1 µM; 48 h). β-actin was
used as a loading control. Density of the HIF-1α or TXNDC5 blot was
normalized against that of the β-actin blot, in order to obtain
relative blot density, which was expressed as a fold change to that
of cells transfected with siScr under normoxia (N-siScr)
(designated as 1). *P=0.0009; **P=0.0004;
***P= 0.0001; ****P=0.0004;
#P=0.0019. CoCl2, cobalt chloride; CRC,
colorectal cancer; H, hypoxia; HIF-1α, hypoxia-inducible factor-1α;
N, normoxia; siRNA/si, small interfering RNA; siScr, scrambled
control siRNA; TXNDC5, thioredoxin domain-containing 5. |
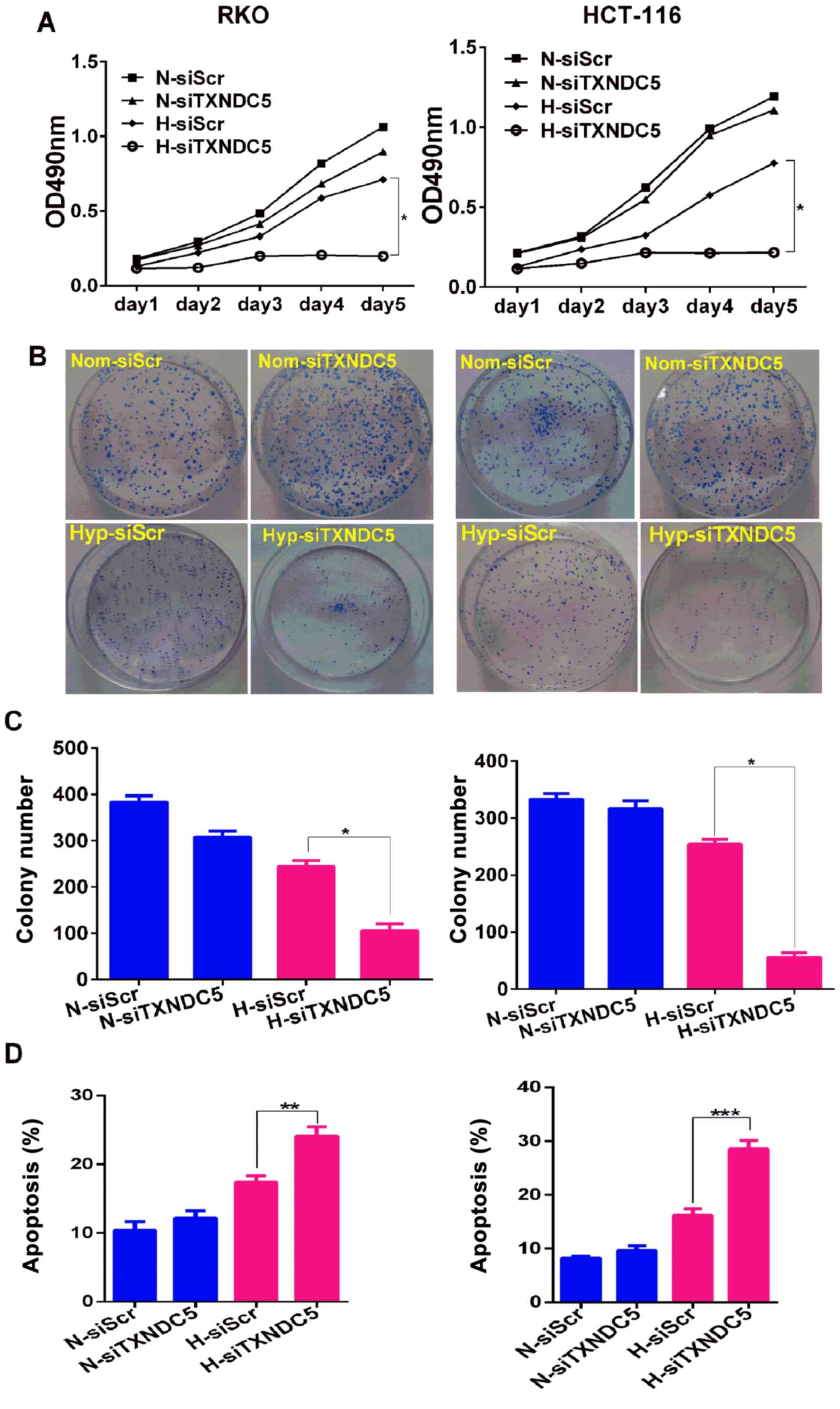
TXNDC5 is critical for CRC tumorigenesis
under hypoxia
To determine the function of TXNDC5, a specific
siRNA (siTXNDC5) was used to knockdown TXNDC5 expression by ~60% in
RKO and HCT-116 cells under hypoxic conditions (Fig. 4). To determine the effects of
TXNDC5 on cell proliferation, the MTT assay was performed on cells
transfected with siTXNDC5 or siScr. As shown in Fig. 5A, TXNDC5 knockdown resulted in a
significant reduction in the proliferation of both cell lines under
hypoxia, but not under normoxia. Furthermore, cells transfected
with siTXNDC5 exhibited a ~75% reduction in colony-forming capacity
compared with in those transfected with siScr under hypoxic
conditions, but not normoxic conditions (Fig. 5B and C). These findings suggested
that TXNDC5 was critical for CRC cell proliferation under hypoxia.
Furthermore, the role of siTXNDC5 was determined in hypoxia-induced
CRC cell apoptosis. As shown in Fig.
5D, knockdown of TXNDC5 significantly increased cell apoptosis
under hypoxia, but not under normoxia, thus suggesting a protective
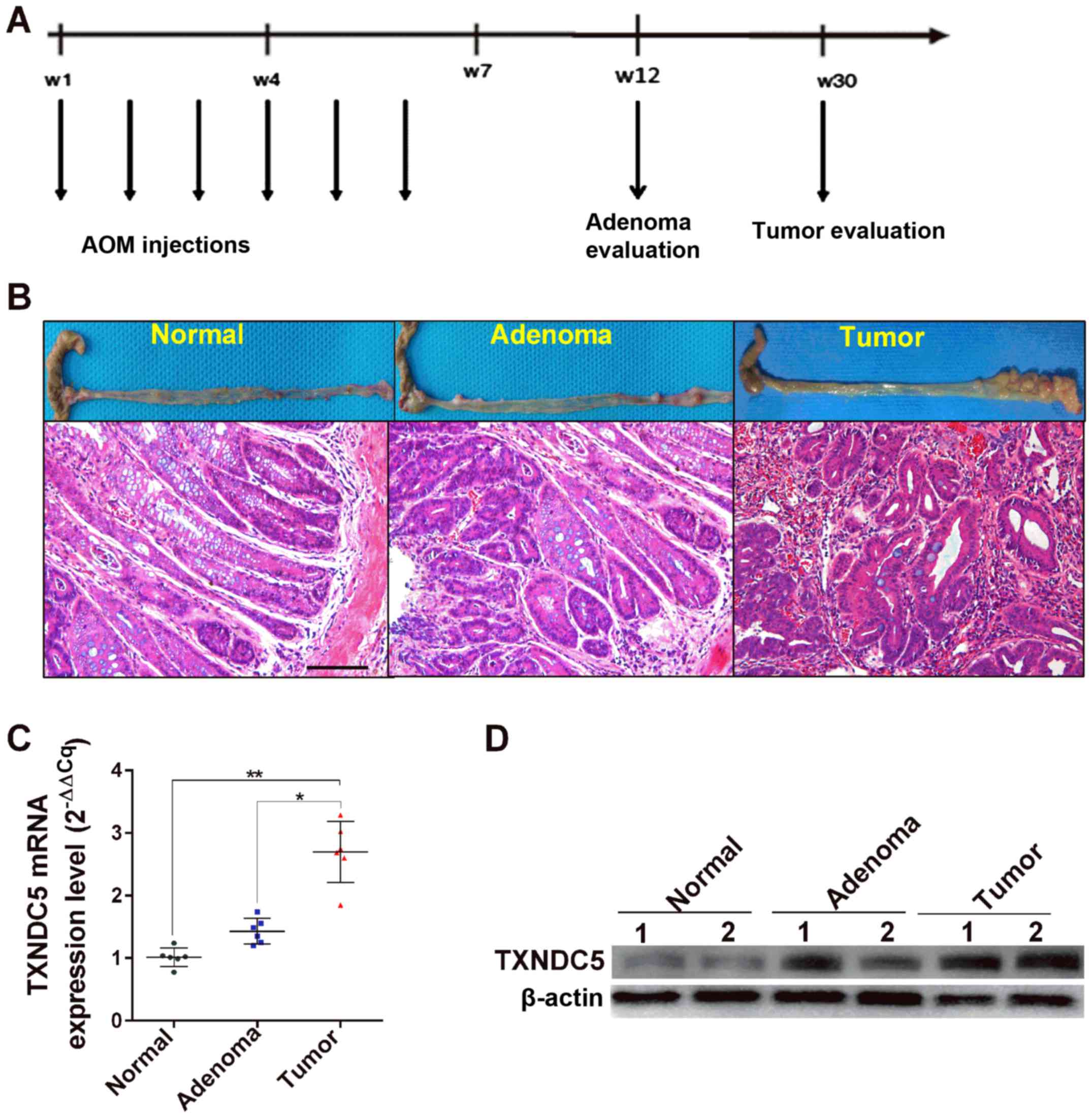
role of TXNDC5 in hypoxia-induced CRC cell apoptosis. The present
study also examined the expression levels of TXNDC5 in an
AOM-induced mouse model of CRC tumorigenesis, which reportedly
exhibits similar pathological and pathophysiological manifestations
to human sporadic colorectal tumorigenesis (22). As shown in Fig. 6, the mRNA and protein expression
levels of TXNDC5 were significantly increased in tumor tissues
compared with in adenoma and normal tissues, confirming the
important role of TXNDC5 in CRC tumorigenesis.
Knockdown of TXNDC5 aggravates ER stress
and induces apoptosis in CRC cell lines
Hypoxia can induce the formation of ROS, which are
critical inducers of ER stress (5-7).
Therefore, the present study examined the effects of TXNDC5
knockdown on the production of ROS in RKO and HCT-116 cells. As
shown in Fig. 7A, cells
transfected with siTXNDC5 exhibited signi ficantly increased ROS
production compared with those transfected with siScr under
hypoxic, but not normoxic conditions. As shown in Fig. 7B, hypoxia significantly induced
the expression of ER stress (CHOP, GRP78 and ATF4), apoptosis (Bax
and cleaved caspase-8) and survival (Bcl-2) markers in RKO and
HCT-116 cells, whereas knockdown of TXNDC5 further increased the
expression of ER stress (CHOP, GRP78 and ATF4) and apoptosis (Bax
and cleaved caspase-8) markers under hypoxic, but not normoxic
conditions. Conversely, knockdown of TXNDC5 significantly decreased
hypoxia-induced expression of the survival marker Bcl-2 under
hypoxia, but not under normoxia.
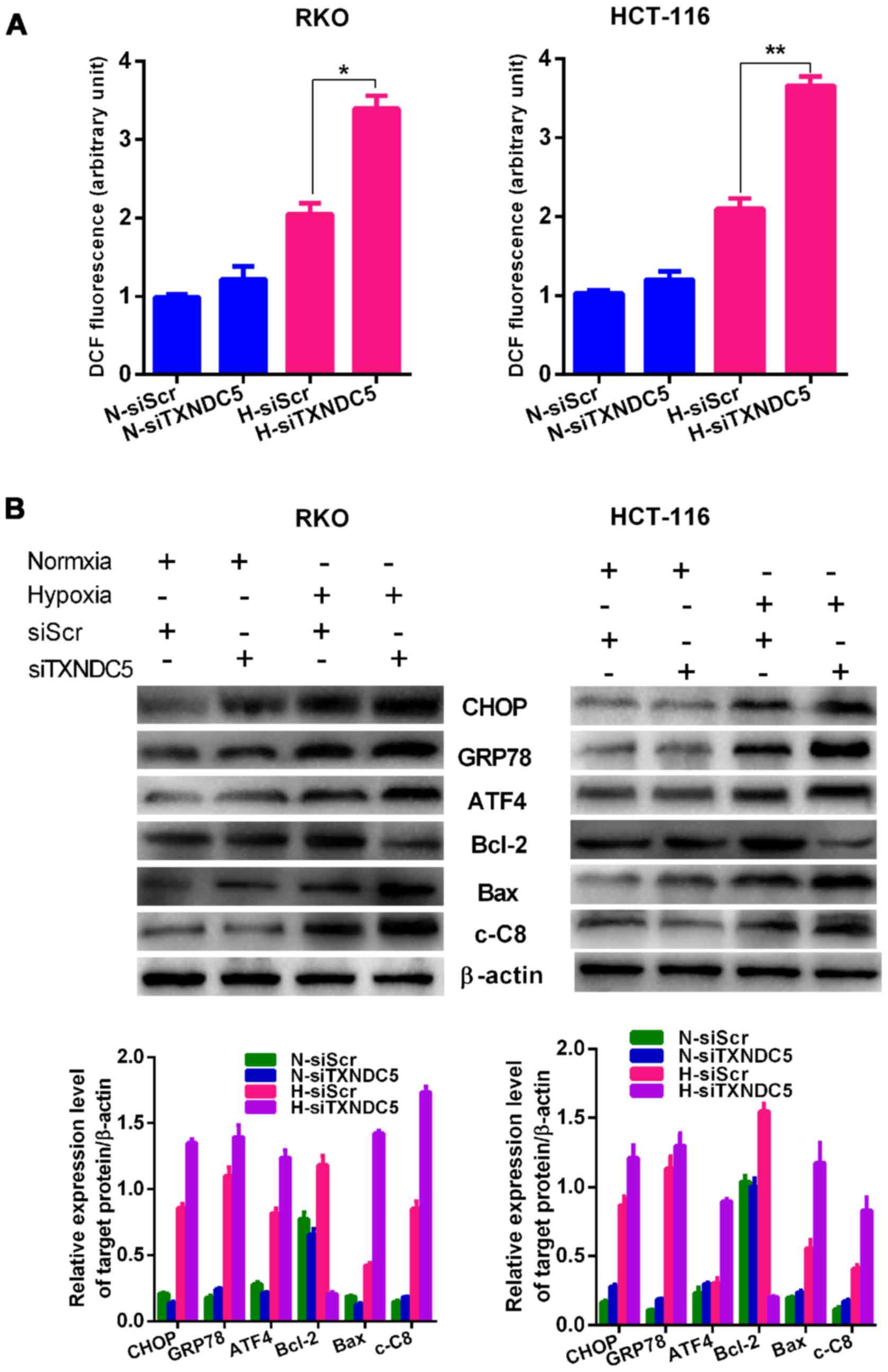 | Figure 7TXNDC5 increases the expression of ER
stress and apoptotic markers. (A) RKO and HCT-116 cells were
transfected with a specific siRNA (siTXNDC5) to knockdown TXNDC5.
Cells transfected with siScr were used as a control. Reactive
oxygen species production was measured in cells under H and N for
24 h. *P=0.0004; **P=0.0010. (B) In RKO and
HCT-116 cells transfected with siTXNDC5 or siScr under H and N,
western blot analyses were used to detect the protein expression
levels of ER stress (CHOP, GRP78 and ATF4), apoptosis (Bax and
cleaved caspase-8) and survival (Bcl-2) markers. β-actin was used
as a loading control. Density of the blots was normalized against
that of the β-actin blot, in order to obtain relative blot density.
*P<0.05. ATF4, activating transcription factor 4;
Bax, Bcl-2-associated X protein; Bcl-2, B-cell lymphoma 2; c-C8,
caspase-8; CHOP, CCAAT-enhancer-binding protein homologous protein;
GRP78, glucose-regulated protein 78; DCF,
dichlorodihydrofluorescein; ER, endoplasmic reticulum; H, hypoxia;
N, normoxia; siRNA/si, small interfering RNA; siScr, scrambled
control siRNA; TXNDC5, thioredoxin domain-containing 5. |
Discussion
Numerous studies, including our previous comparative
proteomic study, demonstrated that TXNDC5 is overexpressed in CRC
(15-17). The present study provided in
vivo and in vitro evidence to sugge st that TXNDC5
serves an important role in the tumorigenesis of CRC, particularly
under hypoxia. The present in vivo data indicated a positive
association between TXNDC5 and the TNM stage of CRC in human
samples; this was corroborated by the sporadic CRC animal model,
which exhibited an ascending tendency in the expression levels of
TXNDC5 from normal to CRC tissues. The in vitro data
indicated that TXNDC5 could promote proliferation and survival of
CRC cells under hypoxia, likely by regulating ER stress.
It has previously been reported that TXNDC5 promotes
cell proliferation under hypoxic conditions, including in
fibroblast-like cells, synovial fluid and blood (25). In addition, hypoxia-induced TXNDC5
has been demonstrated to protect endothelial cells and tumor
endothelium from hypoxia-induced apoptosis (12); however, these previous studies did
not indicate whether TXNDC5 was regulated by hypoxia. In the
present study, in CRC cells, hypoxia induced the expression of
TXNDC5 via HIF-1α; HIF-1α knockdown abolished hypoxia-induced
expression of TXNDC5. A previous study reported that hypoxia
exerted no detectable effects on TXNDC5 expression in non-small
cell lung cancer cell lines under hypoxia, whereas HIF-1α was
upregulated (26). This
discrepancy may be attributed to the fact that the cells were
obtained from different organs with varying genetic
backgrounds.
Previous studies have demonstrated that hypoxia
regulates the expression of genes involved in angiogenesis,
anaerobic glycolysis, cell proliferation and survival (27,28). The present study revealed that
TXNDC5 knockdown decreased CRC cell proliferation and colony
formation, and augmented apoptosis under hypoxia, thus suggesting
that TXNDC5 is critical for CRC cell proliferation and survival
under hypoxic conditions. Previous studies have also demonstrated
that TXNDC5 regulates proliferation and survival of gastric and
pancreatic cancer cells under normoxia (29,30). However, in the present study,
knockdown of TXNDC5 only affected CRC cell proliferation and
survival under hypoxia, not normoxia. These findings suggested that
TXNDC5 is more likely a hypoxia-induced stress survival factor
rather than a common oncogene in CRC.
Hypoxia induces ROS production and ER stress
(10,31,32). ROS directly or indirectly affects
ER homeostasis and protein folding, which triggers ER stress and
may induce cell apoptosis in the case of excessive ER stress
(32,33). Accumulating evidence has suggested
that TXNDC5 is involved in ROS production and ER stress (30,34). It has previously been reported
that inhibiting the expression of TXNDC5 via knockdown of the
orphan nuclear receptor, nuclear receptor subfamily 4 group A
member 1, induces ROS and ER stress in pancreatic cancer cells
(30); conversely, increasing the
expression of TXNDC5 in lipid endothelial cells effectively
decreases ROS production and protects cells (34). The present study demonstrated
that, in CRC cells, knockdown of TXNDC5 markedly increased
hypoxia-induced ROS production, and the expression of
hypoxia-induced ER stress and apoptotic markers. This finding is in
agreement with a previous study, which indicated that
hypoxia-induced TXNDC5 was involved in helping proteins to fold
correctly via its disulfide isomerase activity (13). Taken together, these findings
suggested that TXNDC5 functions as a hypoxia-induced survival
factor to regulate hypoxia-induced ROS/ER stress signaling, thereby
maintaining the tumorigenesis of CRC cells under hypoxia and
oxidative stress.
In conclusion, the present in vivo data
demonstrated that TXNDC5 is overexpressed in CRC tissues, and this
overexpression is associated with unfavorable clinicopathological
features. Furthermore, the in vitro results indicated that
hypoxia induces TXNDC5 expression via upregulating HIF-1α; this
effect may promote CRC cell proliferation and survival under
hypoxia, likely through inhibiting hypoxia-induced ROS/ER stress
signaling. These findings suggested that TXNDC5 functions as an
important stress survival factor to maintain the tumorigenesis of
CRC cells under hypoxia by regulating hypoxia-induced ROS/ER stress
signaling. The present study provided novel insights into the role
of TXNDC5 in the tumorigenesis of CRC.
Acknowledgments
Not applicable.
Funding
The present study was funded by the Science and
Technology Plan Fund of Hunan Province, P.R. China (grant nos.
2015WK3011; 2015SK20201) and the Beijing CSCO Fund (grant no.
Y-MX2014-002).
Availability of data and materials
The datasets used or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
FT and HP conceived and designed the experiments. FT
and HZ wrote the article. XH and NY prepared the patient samples.
FT and NY performed the experiments. HX, XZ and XH collected and
analyzed the data. All authors read and approved the final
manuscript.
Ethics approval and consent to
participate
The present study was approved by the Medical Ethics
Committee of Xiangya Hospital (Changsha, China) and all patients
provided written informed consent. All experimental procedures were
conducted in conformity with institutional guidelines for the care
and use of laboratory animals, and protocols were approved by the
Institutional Animal Care and Treatment Committee of Central South
University.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Zheng R, Zeng H, Zhang S, Chen T and Chen
W: National estimates of cancer prevalence in China, 2011. Cancer
Lett. 370:33–38. 2016. View Article : Google Scholar
|
|
2
|
Pesson M, Volant A, Uguen A, Trillet K, De
La Grange P, Aubry M, Daoulas M, Robaszkiewicz M, Le Gac G, Morel
A, et al: A gene expression and pre-mRNA splicing signature that
marks the adenoma-adenocarcinoma progression in colorectal cancer.
PLoS One. 9:e877612014. View Article : Google Scholar :
|
|
3
|
Schell MJ, Yang M, Missiaglia E, Delorenzi
M, Soneson C, Yue B, Nebozhyn MV, Loboda A, Bloom G and Yeatman TJ:
A composite gene expression signature optimizes prediction of
colorectal cancer metastasis and outcome. Clin Cancer Res.
22:734–745. 2016. View Article : Google Scholar :
|
|
4
|
Keith B and Simon MC: Hypoxia-inducible
factors, stem cells, and cancer. Cell. 129:465–472. 2007.
View Article : Google Scholar
|
|
5
|
Saito S, Lin YC, Tsai MH, Lin CS, Murayama
Y, Sato R and Yokoyama KK: Emerging roles of hypoxia-inducible
factors and reactive oxygen species in cancer and pluripotent stem
cells. Kaohsiung J Med Sci. 31:279–286. 2015. View Article : Google Scholar
|
|
6
|
Hotokezaka Y, van Leyen K, Lo EH, Beatrix
B, Katayama I, Jin G and Nakamura T: AlphaNAC depletion as an
initiator of ER stress-induced apoptosis in hypoxia. Cell Death
Differ. 16:1505–1514. 2009. View Article : Google Scholar
|
|
7
|
Pereira ER, Frudd K, Awad W and Hendershot
LM: Endoplasmic reticulum (ER) stress and hypoxia response pathways
interact to potentiate hypoxia-inducible factor 1 (HIF-1)
transcriptional activity on targets like vascular endothelial
growth factor (VEGF). J Biol Chem. 289:3352–3364. 2014. View Article : Google Scholar :
|
|
8
|
Chevet E, Hetz C and Samali A: Endoplasmic
reticulum stress-activated cell reprogramming in oncogenesis.
Cancer Discov. 5:586–597. 2015. View Article : Google Scholar
|
|
9
|
Yadav RK, Chae SW, Kim HR and Chae HJ:
Endoplasmic reticulum stress and cancer. J Cancer Prev. 19:75–88.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Clarke HJ, Chambers JE, Liniker E and
Marciniak SJ: Endoplasmic reticulum stress in malignancy. Cancer
Cell. 25:563–573. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Horna-Terrón E, Pradilla-Dieste A,
Sánchez-de-Diego C and Osada J: TXNDC5, a newly discovered
disulfide isomerase with a key role in cell physiology and
pathology. Int J Mol Sci. 15:23501–23518. 2014. View Article : Google Scholar
|
|
12
|
Sullivan DC, Huminiecki L, Moore JW, Boyle
JJ, Poulsom R, Creamer D, Barker J and Bicknell R: EndoPDI, a novel
protein-disulfide isomerase-like protein that is preferentially
expressed in endothelial cells acts as a stress survival factor. J
Biol Chem. 278:47079–47088. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Kojima R, Okumura M, Masui S, Kanemura S,
Inoue M, Saiki M, Yamaguchi H, Hikima T, Suzuki M, Akiyama S, et
al: Radically different thioredoxin domain arrangement of ERp46, an
efficient disulfide bond introducer of the mammalian PDI family.
Structure. 22:431–443. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Funkner A, Parthier C, Schutkowski M,
Zerweck J, Lilie H, Gyrych N, Fischer G, Stubbs MT and Ferrari DM:
Peptide binding by catalytic domains of the protein disulfide
isomerase-related protein ERp46. J Mol Biol. 425:1340–1362. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Chang X, Xu B, Wang L, Wang Y, Wang Y and
Yan S: Investigating a pathogenic role for TXNDC5 in tumors. Int J
Oncol. 43:1871–1884. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Wang Y, Ma Y, Lü B, Xu E, Huang Q and Lai
M: Differential expression of mimecan and thioredoxin
domain-containing protein 5 in colorectal adenoma and cancer: a
proteomic study. Exp Biol Med (Maywood). 232:1152–1159. 2007.
View Article : Google Scholar
|
|
17
|
Shen H, Huang J, Pei H, Zeng S, Tao Y,
Shen L, Zeng L and Zhu H: Comparative proteomic study for profiling
differentially expressed proteins between Chinese left- and
right-sided colon cancers. Cancer Sci. 104:135–141. 2013.
View Article : Google Scholar
|
|
18
|
Affer M, Chesi M, Chen WD, Keats JJ,
Demchenko YN, Tamizhmani K, Garbitt VM, Riggs DL, Brents LA,
Roschke AV, et al: Promiscuous MYC locus rearrangements hijack
enhancers but mostly super-enhancers to dysregulate MYC expression
in multiple myeloma. Leukemia. 28:1725–1735. 2014. View Article : Google Scholar :
|
|
19
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-ΔΔC(T)) method. Methods. 25:402–408. 2001. View Article : Google Scholar
|
|
20
|
Dang DT, Chen F, Gardner LB, Cummins JM,
Rago C, Bunz F, Kantsevoy SV and Dang LH: Hypoxia-inducible
factor-1alpha promotes nonhypoxia-mediated proliferation in colon
cancer cells and xenografts. Cancer Res. 66:1684–1936. 2006.
View Article : Google Scholar
|
|
21
|
Shweta, Mishra KP, Chanda S, Singh SB and
Ganju L: A comparative immunological analysis of CoCl2
treated cells with in vitro hypoxic exposure. Biometals.
28:175–185. 2015. View Article : Google Scholar
|
|
22
|
Neufert C, Becker C and Neurath MF: An
inducible mouse model of colon carcinogenesis for the analysis of
sporadic and inflammation-driven tumor progression. Nat Protoc.
2:1998–2004. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
George AL, Rajoria S, Suriano R, Mittleman
A and Tiwari RK: Hypoxia and estrogen are functionally equivalent
in breast cancer-endothelial cell interdependence. Mol Cancer.
11:802012. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Wang L, Zheng Y, Xu H, Yan X and Chang X:
Investigate pathogenic mechanism of TXNDC5 in rheumatoid arthritis.
PLoS One. 8:e533012013. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Chang X, Cui Y, Zong M, Zhao Y, Yan X,
Chen Y and Han J: Identification of proteins with increased
expression in rheumatoid arthritis synovial tissues. J Rheumatol.
36:872–880. 2009. View Article : Google Scholar
|
|
26
|
Vincent EE, Elder DJ, Phillips L, Heesom
KJ, Pawade J, Luckett M, Sohail M, May MT, Hetzel MR and Tavaré JM:
Overexpression of the TXNDC5 protein in non-small cell lung
carcinoma. Anticancer Res. 31:1577–1582. 2011.PubMed/NCBI
|
|
27
|
Mizukami Y, Kohgo Y and Chung DC: Hypoxia
inducible factor-1 independent pathways in tumor angiogenesis. Clin
Cancer Res. 13:5670–5674. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Nagaraju GP, Bramhachari PV, Raghu G and
El-Rayes BF: Hypoxia inducible factor-1α: its role in colorectal
carcinogenesis and metastasis. Cancer Lett. 366:11–18. 2015.
View Article : Google Scholar
|
|
29
|
Zhang L, Hou Y, Li N, Wu K and Zhai J: The
influence of TXNDC5 gene on gastric cancer cell. J Cancer Res Clin
Oncol. 136:1497–1505. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Lee SO, Jin UH, Kang JH, Kim SB, Guthrie
AS, Sreevalsan S, Lee JS and Safe S: The orphan nuclear receptor
NR4A1 (Nur77) regulates oxidative and endoplasmic reticulum stress
in pancreatic cancer cells. Mol Cancer Res. 12:527–538. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Malhotra JD and Kaufman RJ: Endoplasmic
reticulum stress and oxidative stress: a vicious cycle or a
double-edged sword. Antioxid Redox Signal. 9:2277–2293. 2007.
View Article : Google Scholar
|
|
32
|
Clarke R, Cook KL, Hu R, Facey CO,
Tavassoly I, Schwartz JL, Baumann WT, Tyson JJ, Xuan J, Wang Y, et
al: Endoplasmic reticulum stress, the unfolded protein response,
autophagy, and the integrated regulation of breast cancer cell
fate. Cancer Res. 72:1321–1331. 2012.
|
|
33
|
Wang WA, Groenendyk J and Michalak M:
Endoplasmic reticulum stress associated responses in cancer.
Biochim Biophys Acta. 1843:2143–2149. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Gu MX, Fu Y, Sun XL, Ding YZ, Li CH, Pang
W, Pan S and Zhu Y: Proteomic analysis of endothelial lipid rafts
reveals a novel role of statins in antioxidation. J Proteome Res.
11:2365–2373. 2012. View Article : Google Scholar
|