Introduction
Worldwide, malignant tumors are the leading cause of
human mortality (1). In order to
successfully treat cancer, it is crucial to increase the
cytotoxicity on targeted tumor cells while reducing side-effects on
normal cells during radiotherapy. Nanotechnology combined with
biomedical techniques offers us tremendous opportunities and
challenges to develop improved cancer diagnosis and therapeutic
designs (2–4). An ideal strategy is to develop
effective nanoscale radiosensitizers specifically targeting tumor
cells. For instance, naked gold nanoparticles (GNPs) can accumulate
at tumor tissues based on passively targeting mechanisms and
experimental studies have shown promising results for the use of
GNPs as effective radiosensitizers (5).
The size of nanoparticles used in cancer treatment
should be large enough to prevent their rapid leakage from tumor
tissue into blood capillaries. On the other hand, the size should
also be small enough to avoid capture by macrophages in the
reticuloendothelial system (RES). The optimized size of
nanoparticles used in cancer treatment is suggested to be ∼50–100
nm to accumulate on tumor tissues utilizing so-called ‘enhanced
permeability and retention’ (EPR) effect (6). Sperling et al systematically
and analytically reviewed the biological applications of GNPs
recently. However, the majority of biological research has involved
small-sized GNPs (5–20 nm) (7),
which are easily prepared, but are too small to take advantage of
the EPR effect in vivo to target tumors. The mechanism of
cellular uptake of GNPs of large sizes (∼14–100 nm) has previously
been reported (8,9). However, the intracellular uptake
dynamics and the biological effects of these large-sized GNPs, in
particular functionalized GNPs such as glucose-capped GNPs
(Glu-GNPs), have never been studied in depth. Furthermore, the dose
of GNPs used in published studies is high (∼100–10,000 billion
particles per ml) (10–13), which may induce severe side-effects
and limit its clinical applications. Our aim in this study was to
systematically investigate the minimum effective dose of GNPs,
which is critical for clinical trials.
Although naked GNPs can accumulate at tumor tissues
by passive targeting mechanisms, GNPs conjugated with
tumor-specific ligands can achieve active targeting at tumor
lesions, which is more promising for tumor diagnosis and treatment.
In our previous study, we repored on the binding of GNPs with
glucose for targeted delivery (12). The size of GNPs used was 10.8 nm.
In this study, we investigated the impact of different sized GNPs
on cell uptake and distribution. We synthesized the GNPs with
larger sizes (57 and 84 nm) by a new method based on a modified
seeding technique. The potential effects on tumor cellular uptake
and the radiosensitizing effect induced by these large-sized
glucose-capped GNPs (Glu-GNPs) at lower doses were investigated
using two human cancer cell lines (HeLa and MCF-7). We systemically
investigated the cellular uptake dynamics, the subcellular location
and the internalization mechanisms of Glu-GNPs. The cancer
radiotherapy-enhancing effects of GNPs were also evaluated.
Materials and methods
Cell lines and cell growth curve
The HeLa human cervical cancer cell line and the
MCF-7 human breast cancer cell line were purchased from The
American Type Culture Collection (ATCC). Cells were cultured in
DMEM medium (Gibco Life Technologies) enriched with 10%
heat-inactivated fetal bovine serum (FBS; Gibco) plus 100 IU
penicillin G, and 100 mg/ml streptomycin (Sigma), and incubated
under standardized conditions (37°C, 5% carbon dioxide, 100%
humidity).
Cells (10×104 per dish) were seeded and
the cells kept growing for seven days under the same conditions.
MTT assay was used to determine the cell growth based on the
optical density (OD) value of the cells in each dish every day. MTT
assay was applied as described previously with slight modifications
(14). Briefly, following the
procedures mentioned above, 200 μl of the MTT dye (5 mg/ml)
were added into each dish. Three hours later, the unreactive
supernatants in each dish were carefully aspirated and were
replaced with another 1 ml of dimethyl sulfoxide (DMSO) solution to
dissolve the reactive dye. Subsequently, 200 μl solution in
each dish was then removed into a well in a 96-well plate. The OD
value of each cell sample at 490 nm was read using an automatic
multi-well spectrophotometer (Bio-Rad-Coda, Richmond, CA). The
negative control well, which contained medium only, was used to set
the absorbance to zero. The experiments were performed in
triplicate and the averaged values were used to draw the growth
curves for both HeLa and MCF-7 lines.
Synthesis and characteristics of GNPs and
Glu-GNPs Synthesis procedure for GNPs of various sizes
We synthesized the GNPs with a modified protocol
based on our previous study (12).
In brief, the mixture of 25 ml deionized (D.I.) water and 0.5 ml
aqueous solution of HAuCl4 (25 mM) was heated in an oil
bath at ∼120–140°C with stirring speed at 300 rpm. After reflux for
10 min, 2.5 ml aqueous solution of sodium citrate (34 mmol/l) was
quickly added into the reflux system. After continuous reflux for
another 10 min and cooling to room temperature, GNPs with an
average diameter of ∼21 nm were obtained, which were used as seeds
for the preparation of larger-sized GNPs.
The mixture of D.I. water (25 ml), sodium citrate
solution (34 mM, 0.5 ml) and the seed GNPs (2 ml) was heated in an
oil bath at ∼120–140°C with the stirring speed at 300 rpm. After a
10-min reflux, 0.5 ml HAuCl4 (25 mM) was quickly added
into the reflux solution, followed by continuous reflux for another
10 min and then cooling to room temperature. In this case, GNPs
with an average diameter of ∼57 nm were obtained. Alternatively, 1
ml of 21-nm GNP seeds was added to the mixture reaction system
mentioned above and then GNPs with an average diameter of 84 nm
were obtained. We functionalized the naked GNPs by adding 4 ml of
25 mM thioglucose into the previously prepared GNP solutions and
then mixed them at room temperature for 24 h to obtain Glu-GNPs.
Both naked GNPs and Glu-GNPs were dialyzed for 48 h and sterilized
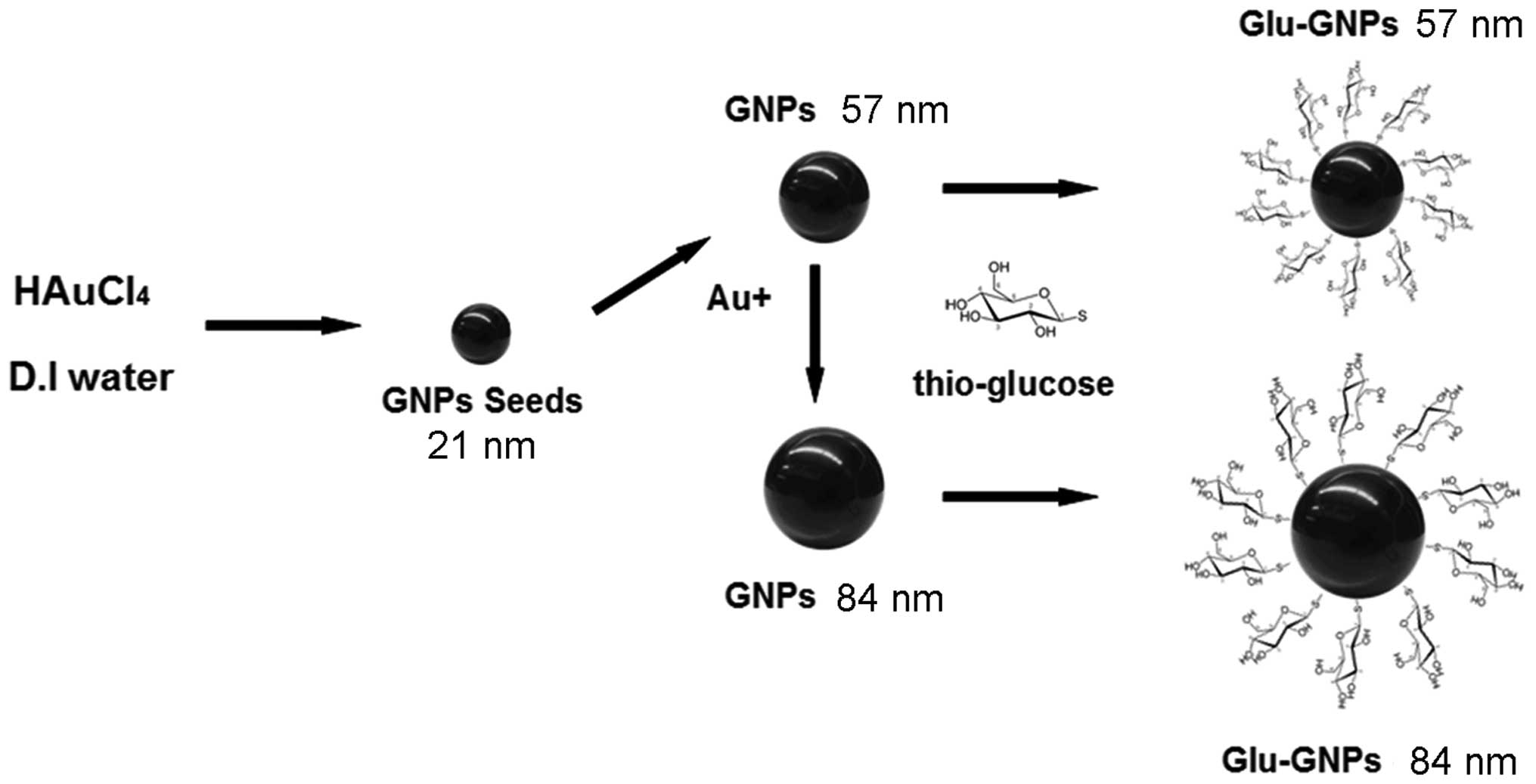
using 200 μm aperture filter before use. The schematic
diagram of the procedure for synthesizing GNPs and Glu-GNPs is
shown in Fig. 1.
The sizes and shapes of the prepared nanoparticles
were measured by transmission electron microscopy (TEM). To obtain
the size distribution of nanoparticles, we used the PerkinElmer
Lambda 900 spectrometer (PerkinElmer, Santa Clara, CA) to scan the
visible absorption spectra within a ∼200–800 nm range of both
freshly prepared GNP and Glu-GNP stock solutions. Inductively,
coupled plasma mass spectrometry (ICP-Mass; Elan 6000, Perkin
Elmer) was used to measure the final concentrations of the GNP
solutions with different sizes. The surface characterization of
Glu-GNPs was carried out using X-ray photoelectron spectroscopy
(XPS) to determine the elemental composition on each Glu-GNP. Both
ICP-Mass and XPS mass measurements were repeated three times for
each sample.
Intracellular distribution of
nanoparticles
To investigate the dynamic intracellular
distribution of GNPs and Glu-GNPs, 6×105 cells were
seeded in each culture dish and allowed to adhere and acclimate for
one day. GNPs and Glu-GNPs of different sizes (57 and 84 nm) were
then applied to the cells with the final concentration of
7×108 particles/ml and incubated with the cells for
different periods of time (1, 6 and 24 h) at 37°C. Following two
washes with phosphate-buffered saline (PBS), cells were collected
and fixed with cold 2% glutaraldehyde in 0.1 M sodium cacodylate
buffer at 4°C for at least 4 h. The cells were post-fixed in 1%
osmium tetroxide in 0.2 M sodium cacodylate buffer for 1 h and then
stained with 2% aqueous uranyl acelate for 30 min at room
temperature, followed by dehydration in a graded series of ethanol.
Propylene oxide was used as a link reagent before embedding the
cells in closed, labeled gelatine capsules with fresh resin.
Ultrathin sections of the samples were cut and observed with a
Philips/FEI (Morgagni) Transmission Electron Microscope with a CCD
camera (TEM-CCD). The intracellular location of the GNPs and
Glu-GNPs was then analyzed.
GNP and Glu-GNP uptake by cells
To dynamically investigate the difference between
the cellular uptake of GNPs and that of Glu-GNPs, HeLa and MCF-7
cells were individually seeded in dishes (6×105 cells
per dish) and cultured overnight. Either GNPs or Glu-GNPs with
sizes of 57 nm were added into the cell cultures to reach the final
concentration of 2.5×109 particles/ml and incubated with
the cells for different periods of time (15 and 30 min, 1, 3, 6,
12, 24 and 48 h) at 37°C. The medium with free nanoparticles was
removed and the cells were washed with PBS twice at each
time-point. Cells were collected and the number of cells was
counted, followed by centrifugation. A total of 4 ml of 20%
HNO3 was then added into each sample to lyse the
pellets. The gold mass in the lysis solution was detected by
ICP-Mass. We calculated the number of nanoparticles via the
measurement of gold in the solution. In addition, the number of
GNPs and Glu-GNPs in the lysis solution divided by the number of
cells provided a quantitative measurement of GNPs uptaken in each
cell. Each experiment was performed in triplicate, and the average
values and standard deviation are presented. The same experiments
were performed for the 84-nm nanoparticles.
Determination of cell death
MTT assay was used to determine cell death induced
by optimal X-ray doses and time-points in the following
experiments. Irradiation was carried out at various X-ray doses (5,
10 and 15 Gy) and the cells were allowed to keep growing for 24,
48, 72, 96 and 120 h. Based on the MTT results at each time-point
(data not shown), the irradiation dose of 10 Gy and incubation time
of 96 h after X-ray treatment were selected for our cell death
experiments.
Quantitative analysis of apoptosis or necrosis of
the cells was performed by flow cytometry using the Annexin V-FITC
apoptosis kit (BioVision, Milpitas, CA) as discribled previously
(15). HeLa cells in 8 ml of 10%
FBS DMEM medium (2×105 cells/dish) were seeded and
cultured overnight at 37°C. The supernatants were removed and
replaced with 8 ml fresh glucose-free medium solution containing
GNPs and Glu-GNPs (57 nm) with the final concentrations of 10 and
20 μM, respectively, in each well. After incubation for 48
h, the medium containing the drugs was aspirated and the cells were
rinsed with PBS twice and then replaced with another 8 ml DMEM
prior to X-ray radiation. Following irradiation, the cells were
cultured for another 96 h. Approximately 1×106 cells
were collected, washed with PBS twice, and then suspended in 100
μl of Annexin V binding buffer (1X) and incubated with 10
μl Annexin V (20 μg/ml) and 5 μl propidium
iodide (PI) for 15 min at room temperature in dark. A total of 400
μl binding buffer were then added into each tube and the
cells were analyzed with a FacsCalibur flow cytometer
(Becton-Dickinison, Franklin Lakes, NJ) within 1 h. Data analysis
was performed with CellQuest software (Becton-Dickinison).
Experiments were repeated six times. The results were interpreted
as follows: cells that were Annexin V(−)/PI(−) (lower left
quadrant) were considered as living cells, the Annexin V(+)/PI(−)
cells (lower right quadrant) as apoptotic cells, Annexin V(+)/PI(+)
(upper right quadrant) as necrotic or advanced apoptotic cells, and
Annexin V(−)/PI(+) (upper left quadrant) as bare nuclei, or cells
in late necrosis, or even cellular debris. Each sample including
cells received treatment with both Annexin V and PI, blank cells
received single staining with Annexin V alone or PI alone and
non-stained cells were employed as references for setting up the
parameters for flow cytometer detection.
TUNEL assay
TUNEL staining was used to identify the morphology
features of cell death induced by either X-ray alone or irradiation
plus GNPs. HeLa cells were seeded in cell culture dishes
(1×104 cells per dish) and incubated overnight followed
by the addition of either GNPs or Glu-GNPs. The final concentration
of both GNPs was 20 mM (2.5×109 particles/ml) and 10
μM (1.25×109 particles/ml), respectively. After
incubation for 48 h, the cells were treated with X-ray irradiation
(125 kVp with the Pantak Therapax3 series) at a single dose of 10
Gy and were subsequently cultured for various periods of time (48,
72 and 96 h). At each time-point, TUNEL staining was performed
following the instructions in the manual of the DeadEnd™
Fluorometric TUNEL System (Promega Corp., Madison, WI) with slight
modifications. In brief, cells were fixed in 4% methanol-free
formaldehyde in PBS for 30 min at room temperature. After washing
with PBS twice, cells were permeabilized with 0.1% Triton X-100
solution for 5 min at 4°C. The cells were washed twice and
incubated with 100 μl of the equilibration buffer at room
temperature for 10 min. The cells were then incubated with the
TUNEL reaction mixture (Nucleotide Mix and rTdT) in a dark
humidified chamber for 60 min. The reaction was terminated with the
kit 2X SSC reagent and the cells were washed with PBS twice to
remove unincorporated fluorescein-12-dUTP. The slides were mounted
in VECTASHIELD® mounting medium with DAPI (Vector
Laboratories, Burlingame, CA) to stain the nuclei followed by the
addition of coverslips to the slides and the edge of the coverslips
was sealed by nail polish. The fluorescein-12-dUTP-labeled DNA of
apoptotic nuclei appears green when visualized by a Zeiss LSM 510
Laser Scanning Confocal Microscope (Zeiss, Germany). The cells
without any interference and the cells with X-ray treatment only
served as the controls.
Statistical analysis
The statistical analysis was performed using SPSS
14.0 for Windows. Differences between each group were analyzed by
the Student’s t-test. P<0.05 was considered to indicate a
statistically significant difference.
Results
Characteristics of GNPs
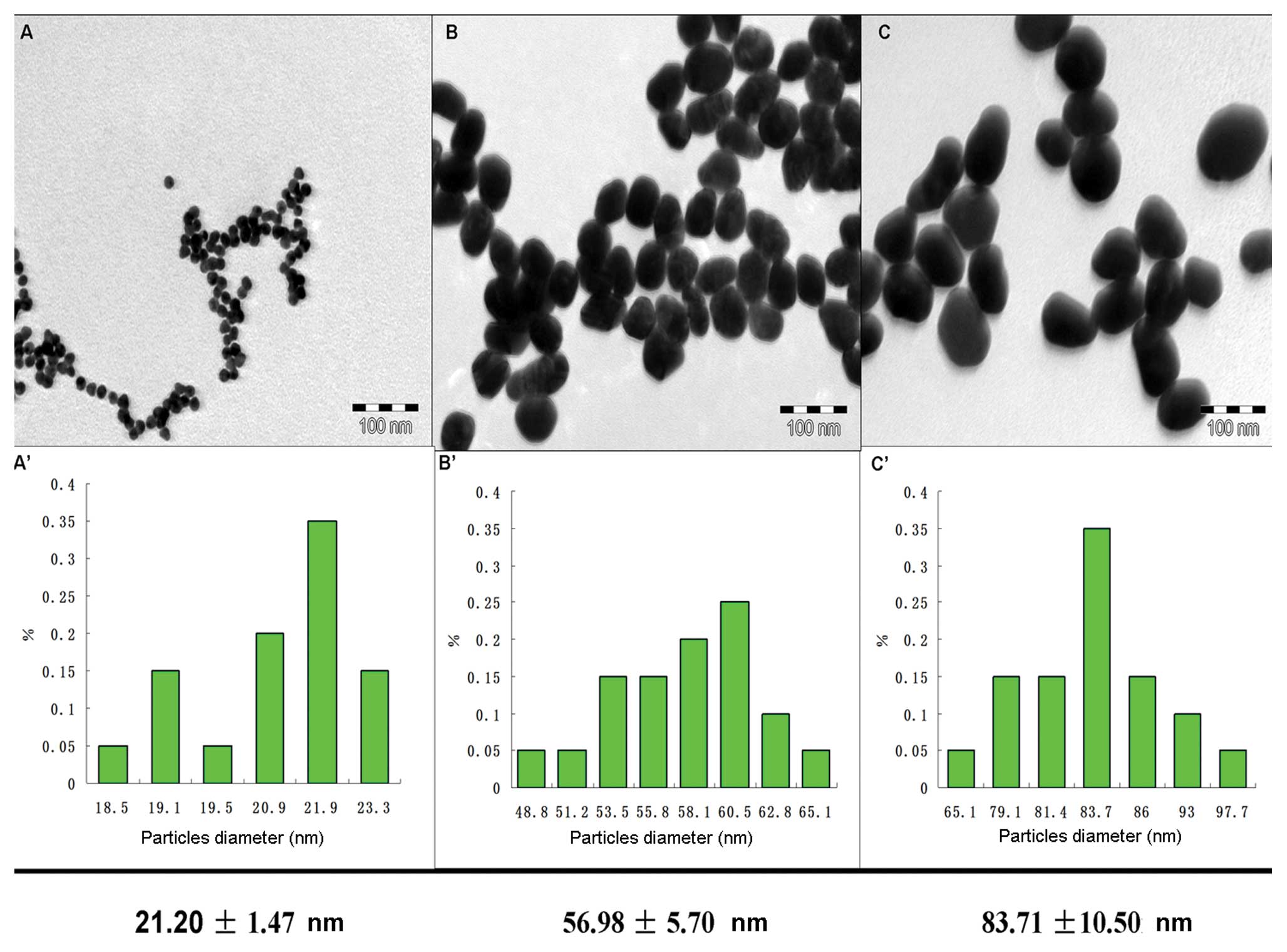
Fig. 2 shows the
TEM images of different sized GNPs. Based on the TEM results, three
sizes of GNPs with average diameters of 21.20±1.47 nm (A),
56.98±5.7 nm (B) and 83.71±10.5 nm (C), are presented. GNPs with a
size of 21.20±1.47 nm served as the seeds. GNPs of 54 and 84 nm
were obtained by letting gold molecules gradually grow on the
surface of these seed GNPs in the reaction solutions until
nanoparticles with the expected size was achieved. Using this seed
method, we can easily prepare GNPs with uniform sizes and shapes.
The sulfur-to-gold atom ratio was 1.02, 5.56 and 3.70 for 21, 57
and 84 nm Glu-GNPs, respectively. The results indicated that more
thio-glucose molecules conjugated on the surface of the 57-nm GNPs
particles. In other words, given the same amounts of glucose
molecules, more Au atoms will be uptaken by the cells using 57-nm
Glu-GNPs than the two other sized nanoparticles.
Distribution and internalization
mechanisms of GNPs and Glu-GNPs in cells
More than 300 images were taken for this experiment.
The distribution of GNPs and Glu-GNPs in the HeLa and MCF-7 cells
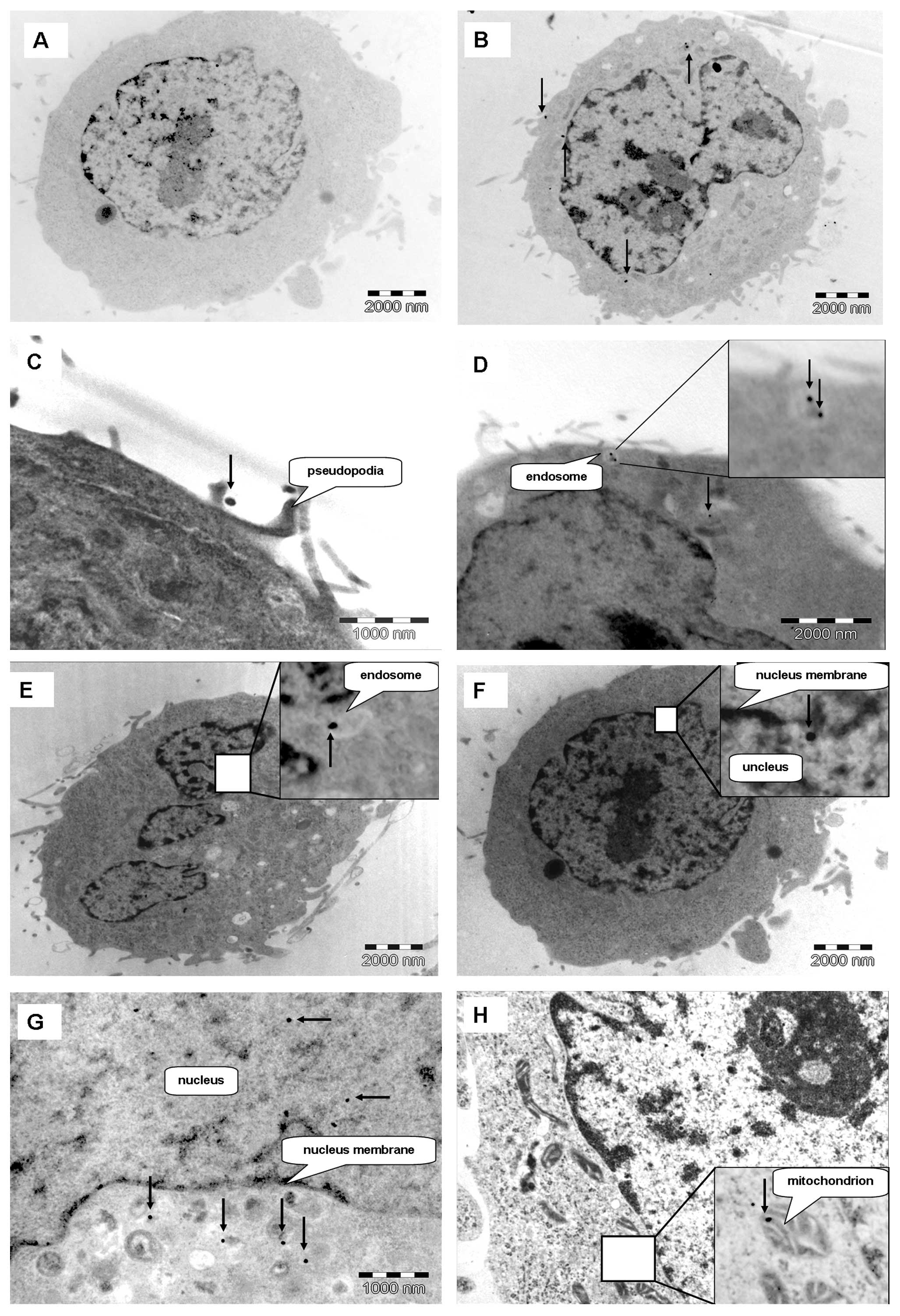
was determined by TEM. Fig. 3
shows HeLa cells treated with either GNPs or Glu-GNPs (57 nm) at
different incubation times (1, 6 and 24 h). The GNPs were mainly
located at the cytoplasm (Fig.
3B). However, several gold nanoparticles appeared in the area
of the nucleus (Fig. 3F and G).
More importantly, Fig. 3H shows
the GNPs that were located inside the mitochondrion. These results
regarding the detailed GNP location inside cells at the subcellular
level have not been reported previously. GNPs and Glu-GNPs were
rapidly internalized into the cells after the GNPs were added. TEM
images showed that after 1 h of GNP administration, the GNPs
reached areas around nucleus, while the cells continued to uptake
GNPs. The distribution of GNPs and Glu-GNPs in the MCF-7 cells was
the same as in the HeLa cells regardless of the sizes of the
particles.
Fig. 3D shows a
typical endosome containing two groups of GNPs near the cell
membrane. The endosome is formed and moves inwards of the cell. The
typical structure of the endosome then disappears, which suggests
that a typical endocytosis procedure has occurred. Fig. 3C shows that the cell protrudes its
pseudopodia to catch GNPs. All these images indicate that
endocytosis is the mechanism behind the internalization of GNPs and
Glu-GNPs.
Uptake of GNPs and Glu-GNPs by cells
The numbers of GNPs and Glu-GNPs uptaken by the HeLa
and MCF-7 cells in cell lysates was quantified using ICP-Mass.
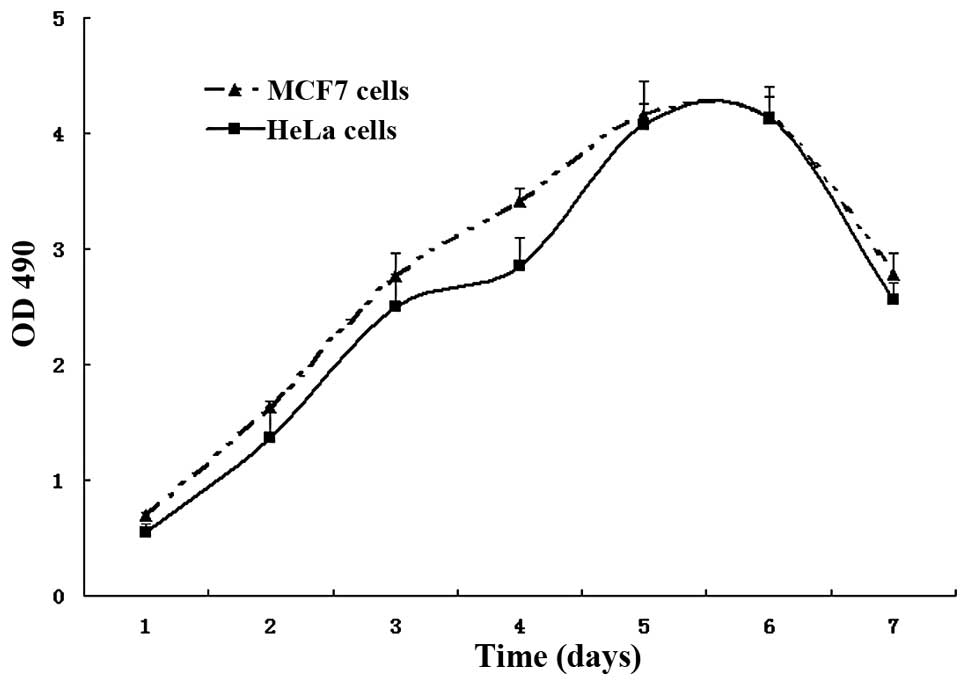
Fig. 4 shows the growth curve of
both HeLa and MCF-7 cells, which indicates that the growth cycle is
similar for both cells. However, HeLa cells grew much slower than
the MCF-7 cells during the first five days. The average number of
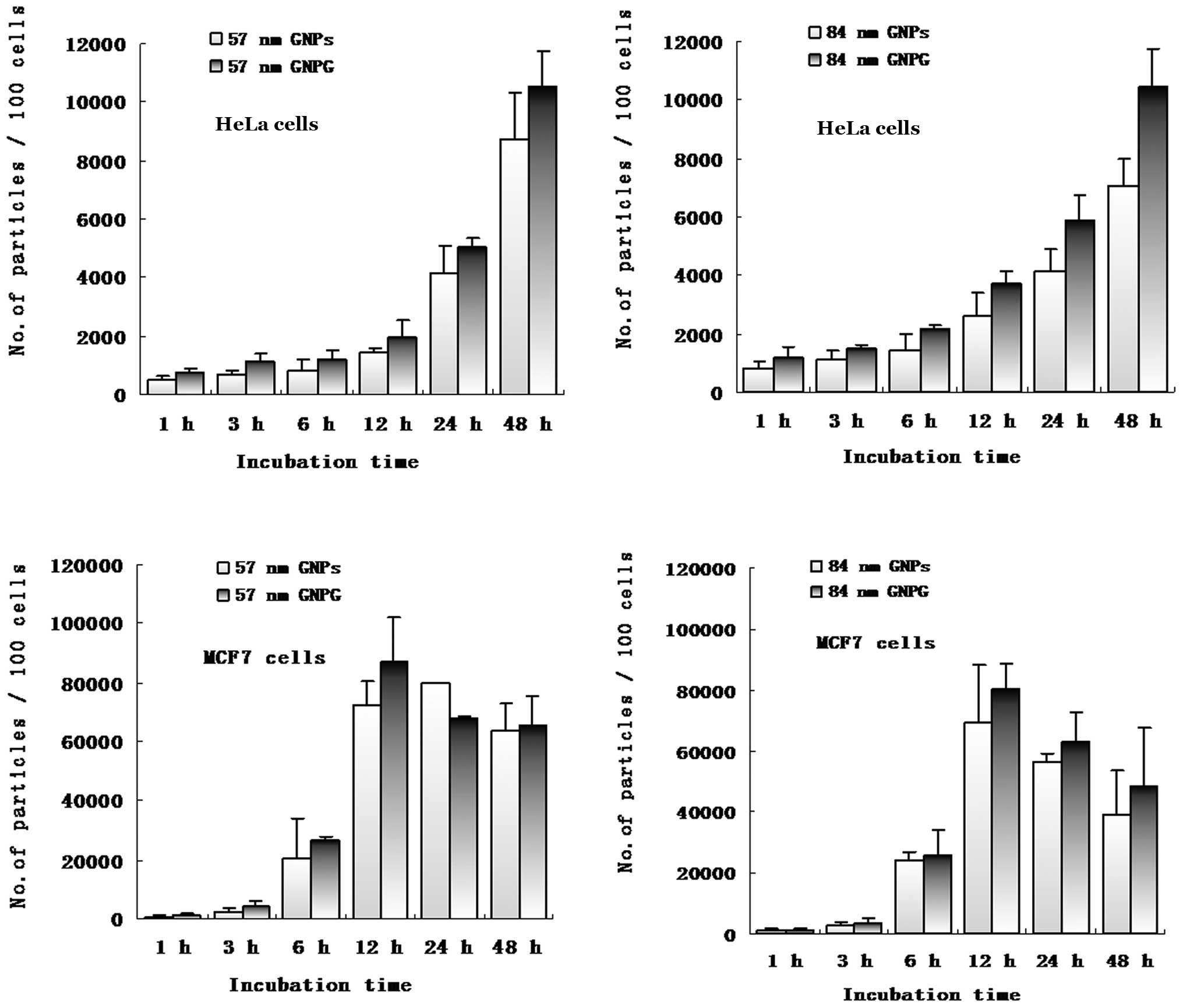
GNPs and Glu-GNPs in each cell is shown at Fig. 5, which confirms that both HeLa and
MCF-7 cells took up more Glu-GNPs than GNPs. However, MCF-7 cells
took up more GNPs and Glu-GNPs than HeLa cells. MCF-7 cells reached
the peak uptake value earlier than the HeLa cells (12 vs. 48
h).
The average number of size 57 nm nanoparticles
within every 100 HeLa cells was 8.71×103 for the naked
GNPs and 10.51×103 for the Glu-GNPs after 48 h of
incubation (P=0.043). Similarly, more Glu-GNPs were taken up by the
cells than naked GNPs with a size of 84 nm after 48 h of incubation
(10.45×103 vs. 7.08×103, P=0.025) as shown in
Table I. MCF-7 cells took up more
naked GNPs (6.34×104 vs. 8.71×103, P=0.034)
with size a of 57 nm than the HeLa cells after 48 h.
 | Table I.The average number of nanoparticles
uptaken by HeLa cells after 48 h of incubation. |
Table I.
The average number of nanoparticles
uptaken by HeLa cells after 48 h of incubation.
| Size
|
|---|
| 57 nm | 84 nm |
|---|
| The average number
of nanoparticles uptaken by 100 HeLa cells |
8.71×103
Naked GNPs |
7.08×103
Naked GNPs |
| The average number
of nanoparticles uptaken by 100 HeLa cells |
1.051×104
Glu-GNPs |
1.045×104
Glu-GNPs |
| P-value | 0.043 | 0.025 |
Both the HeLa cells and MCF-7 cells took up more
numbers of 57-nm GNPs/Glu-GNPs than 84-nm nanoparticles. For
instance, more 57-nm Glu-GNPs were taken up than 84-nm Glu-GNPs by
HeLa cells (8.71×103 vs. 7.08×103, P=0.011),
as well as by MCF-7 cells (6.58×104 vs.
4.83×104, P=0.05) at 48 h. Our results revealed that the
uptake of nanoparticles was related to the surface characteristics
of nanoparticles (GNPs vs. Glu-GNPs), the type of cells (HeLa vs.
MCF-7), the size of nanoparticles (57 nm vs. 84 nm) and incubation
time. The size-dependent uptake result was in line with our
expectations, that is Glu-GNPs components result in more glucose
being conjugated on the surface of 57-nm particles (S/Au = 10.71)
than 84-nm particles (S/Au = 3.06). The time-point of the cell
uptake peak is the optimal time for irradiation. Based on the
results we obtained for the 57-nm GNPs and Glu-GNPs, the following
experiments were designed. HeLa cells were used in our experiments
as HeLa cells take up less nanoparticles than MCF-7 cells. If the
treatment proved successful for HeLa cells, the same treatment
would then be expected to be even more effective for MCF-7
cells.
GNP enhanced radiotherapy
To reduce the side-effects induced by GNPs and
Glu-GNPs, lower nanoparticle doses (2.5×109 GNPs/ml, 20
μM and 1.25×109 GNPs/ml, 10 μM) were used
in our radiosensitivity study. The cells in left low quadrant are
characterized by Annexin V(−)/PI(−), which represent live cells.
While cells in the other three quadrants are dead cells. Both GNPs
and Glu-GNPs can increase the death rate by X-rays in HeLa cells
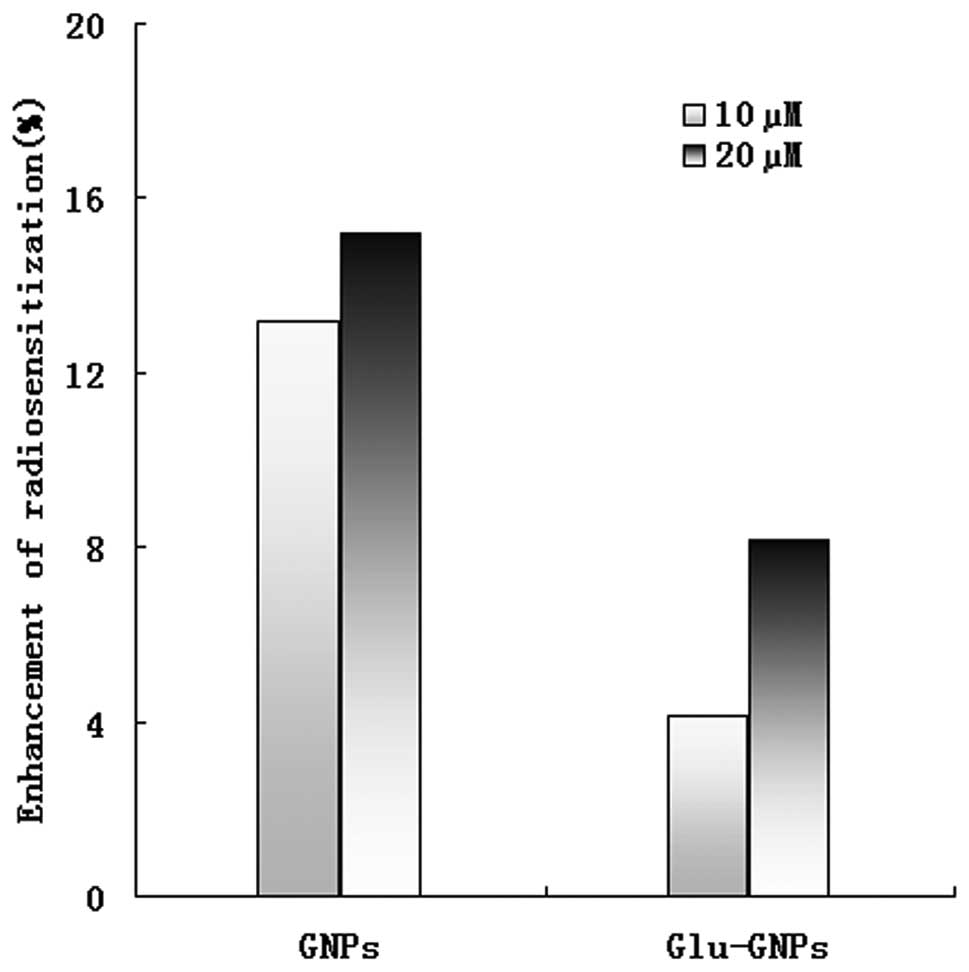
even at a lower dose. The relative radiosensitizing effect reached
15% when 20 μM GNPs were used. However, the radiosensitizing
effect of Glu-GNPs was not as effective as GNPs alone, 8% vs. 15%
killing effects for 20 μM and 4% vs. 13% killing effects for
10 μM (P=0.053) even though more Glu-GNPs than GNPs were
uptaken by HeLa cells (please refer to our explanations in the
Discussion) as shown in Fig.
6.
Apoptosis detection
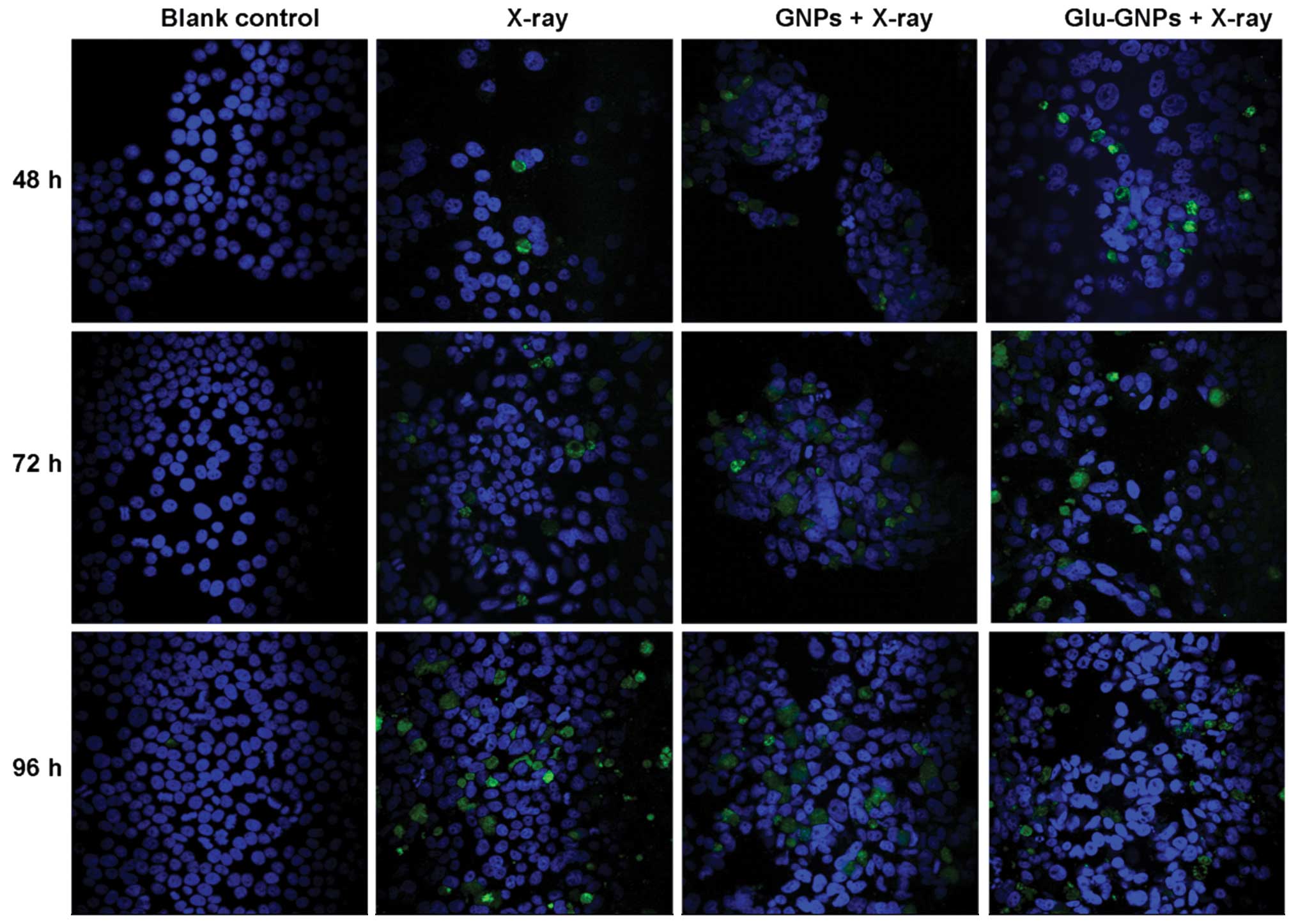
The TUNEL method can be used to label the fragmented
DNAs inside cells, which is a characteristic method for determining
apoptosis. As shown in Fig. 7,
X-rays clearly induced apoptosis compared to the blank controls.
The size of TUNEL-positive nuclei tended to increase as incubation
time increased (48, 72 and 96 h) after X-ray treatment. There was
no significant difference among cells only treated with X-ray alone
or nanoparticles without X-ray irradiation. The results are in line
with our flow cytometry analysis results.
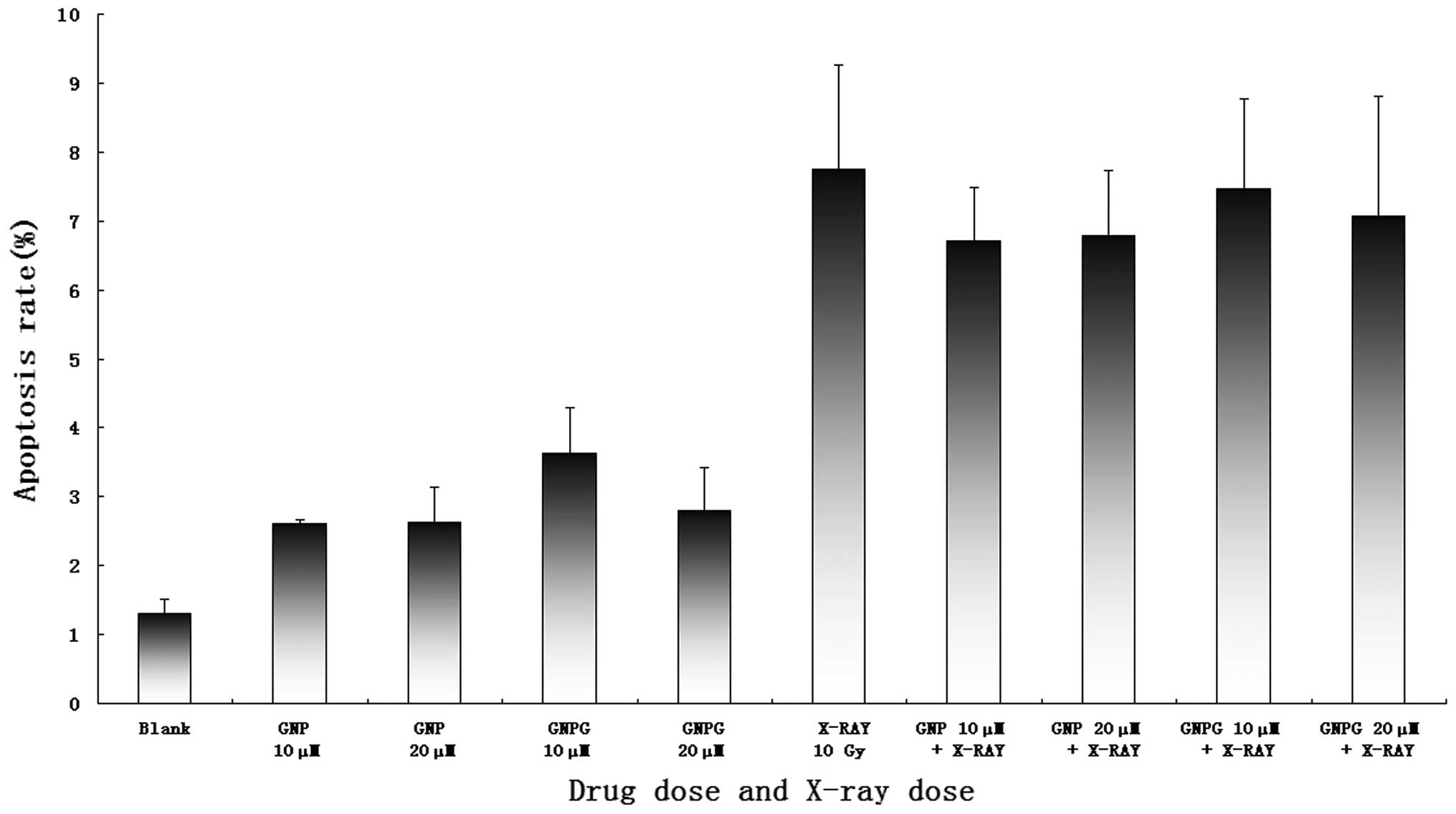
Dual staining of cells with Annexin V and PI can be
used to quantitatively distinguish apoptotic from necrotic cells.
The apoptotic rate of the X-ray-treated group was significantly
higher than the blank control (7.75±1.51 vs. 1.31±.27%, P=0.004).
Compared to the cells treated with X-ray alone, the cells that
received GNPs or Glu-GNPs plus X-ray treatment showed no
significant difference on apoptosis induction. As shown in Fig. 8, the apoptotic rate in the X-ray
alone group was slightly higher than in the other groups, but with
no significant statistical difference (7.75±1.51 vs. 6.70±0.78 and
6.79±0.95 vs. 7.46±1.31 vs. 7.07±1.74%, P>0.05).
Discussion
GNPs are promising novel nanoscale drug carriers,
radiosensitizers and bio-imaging contractors for cancer diagnosis
and therapy. The biological functions of GNPs have been extensively
investigated in various types of malignant tumor. All the
applications are based on the active chemistry and physical
properties of gold atoms and the size-dependent passive tumor
targeting mechanism of GNPs (16–19).
Nanoparticles can selectively accumulate on tumor tissues due to
their small sizes and the defective tumor vasculature. The diameter
of the endothelial gap junction around fenestrated capillaries in
normal human tissues is 60–80 nm (20). However, the size of the gap
junction between endothelial cells of the incomplete cancer
vasculature may vary from 100–600 nm 21,22). As a result,
nanoparticles with sizes of 50–100 nm have the ability to enter and
accumulate around tumor tissue. The non-functioning developed
lymphatic drainage system of cancer cannot recollect these
nanoparticle back into blood circulation, which is so-called EPR
effect. However, it is difficult to control the size and shape of
large GNPs during synthesis. The majority of GNP studies thus far
have used small-sized nanoparticles with diameters ranging from
1.8–30 nm. To investigate the effect of larger-sized GNPs, we
synthesized larger and high quality GNPs using a modified seeding
technique based on the methods described in our previous study.
Compared with a popular ‘citrate reduction’ method, our method can
easily generate GNPs of a more uniform size and shape, which is
important for assessing the GNP cytotoxic effects. TEM images
confirmed that the size and shape of the resulting nanoparticles
was uniform with an average diameter of 57 and 84 nm.
Our XPS results revealed that many glucose molecules
bound to the surface of GNPs. The ratio of the atom numbers for S
to Au in a single 57 Glu-GNPs particle was 5.56, which was the
highest among the three different sizes of nanoparticles we
obtained. This suggests that, for glucose-coated GNPs, a single
glucose molecule can carry more Au atoms into cells using a 57-nm
nanoparticle; this type of nanoparticles is more cost-effective for
treatment. The ICP-Mass results showed that, as the size increased,
the nanoparticle concentration decreased accordingly, given the
same mass concentration. The choice of optimal nanoparticle
concentration is therefore essential as more nanoparticles mean
more toxicity to the human body. Based on these observations and
our uptake study results, more 57-nm nanoparticles were taken up by
the cells. GNPs and Glu-GNPs with a size of 57-nm are desirable for
future clinical studies.
There is an elevated glucose consumption and
overexpression of glucose transporters (GLUTs) in cancers (23–25).
It has long been recognized that cancer cells have increased rates
of glucose metabolism compared with healthy cells. Gulcose can
serve as an excellent tumor-targeting tracer. The most successful
illustration of this theory is the widespread clinical application
of PET scanning based on the selective uptake of
[18F]fluorodeoxyglucose (FDG), a glucose analog that competes for
glucose transport sites on the membrane by cancer cells (26). To target cancer cells more
specifically, we modified the surface of naked GNPs using glucose
so that active targeting to malignant tumor cells could be
achieved. By doing so, a reasonably rapid enhancement of GNP uptake
by both HeLa and MCF-7 carcinoma cell lines was clearly observed in
our experiments by comparing the uptake of Glu-GNPs vs. naked GNPs.
Therefore, when cancer cells uptake more glucose, more GNPs can
also be internalized into the cells.
In this study, more 57-nm particles were taken up by
the HeLa and MCF-7 cells than 84-nm nanoparticles regardless of
whether glucose was capped on the GNPs or not. This result is
partially in line with the results of the study by Chithrani et
al, and the result confirms that 50 nm is the best choice for
gold-based nanoparticles for tumor diagnosis and treatment
(8,9). However, we are the first group to
illustrate that the Glu-GNP uptake by cells is also size-dependent
and that the optimal size is 57 nm instead of 84 nm. Based on our
results, the nanoparticle size is critical for biomedical
applications regardless of whether the surface of the nanoparticles
is modified or not.
Another important factor for GNP-enhanced cell
killing is when radiotherapy is applied after GNPs are
administrated, and the application time depends on when the highest
uptake is reached. Our results showed different cellular uptake
dynamics for the HeLa cells and MCF-7 cells. The MCF-7 cells
reached peak uptake much earlier than the HeLa cells (12 h vs. 48
h) and more GNPs/Glu-GNPs were taken up by the MCF-7 cells than the
HeLa cells (approximately a 10-fold increase). Therefore, the best
time to apply irradiation following the GNP application is 48 h for
HeLa cells and 12 h for MCF-7 cells. The growth curve confirmed
that the HeLa cells grew much slower than the MCF-7 cells at days
two to four, when the GNPs and Glu-GNPs had just been administered
to the cells. That may be one of the reasons why MCF-7 cells uptake
more particles than HeLa cells. In conclusion, the uptake of
gold-based nanoparticles depends on the surface characters of the
nanoparticles, the size of the nanoparticles and the type of cells
used. All the factors are important for the success of future
clinical trials.
The mechanism of GNP-based radiosensitization is
reported due to the generation of free radicals from gold atoms
after they are bombarded by irradiation, such as X-ray. Free
radicals can further generate reactive oxygen species (ROS) that
are toxic and damage proteins and genetic materials (27,28).
However, the diffusion distance of free radicals is very short
(29). For example, the diffusion
range of 1O2 is limited to approximately 10
nm in cells (30–32). Since the diameter of human cells
ranges from approximately 10–100 μm, the cancer killing
effect depends on the site where the primary
1O2 is generated. In other words, where the
gold-based naoparticles locates consequently determines which
subcellular structures are affected. Therefore, it is not
surprising that the type of the response triggered by the
activation of the GNPs depends on their intracellular localization.
The subcellular location of nanoparticles is also essential for us
to gain in-depth understanding of the toxic effect of GNPs on tumor
cells. According to TEM images, we dynamically monitored the
locations of both naked GNPs and Glu-GNPs in the HeLa and MCF-7
cells after 1, 6 and 24 h. GNPs and Glu-GNPs were mainly located in
the cell cytoplasms regardless of the cell type and incubation
time. That means subcellular organelles nearby the nanoparticles
inside cytoplasms are potential targets for gold-based X-ray
treatment. Fig. 3H shows that a
group of nanoparticles is located in the mitochondrion; this is
important as mitochondria-mediated apoptosis plays an essential
role in cell death, which is one of the two apoptotic pathways in
cells. Fig. 3F shows that several
groups of nanoparticles inside the nucleus after a 24-h
administration of 84-nm GNPs. Fig.
3G shows that a group of nanoparticles is located inside the
nucleus close to the nuclear membrane after a 6-h administration of
84-nm GNPs. Although further evidence is required to support our
discovery, it is reasonable to postulate that nanoparticles can
enter the nucleus. The diameter of nuclear pore complexs (NPCs) is
approximately 80–120 nm. It has been reported that small
nanoparticles are able to pass through the NPCs by passive
diffusion. Larger particles are also able to pass through the large
diameter of the pore, but at almost negligible rates (33). The detailed distribution of GNPs at
subcellular organelles has not been reported previously. The
majority of studies only describe the location in the cytoplasm.
Chang et al recently confirmed the subcellular location of
GNPs on the endoplasmic reticulum (ER) and Golgi apparatus using a
double staining technique (11).
Based on their results and ours, it is possible for GNPs to attach
to the surface or even enter the subcellular organelles. The
sucellular organelle-targeted radiotherapy using GNPs can thus be
achieved.
Endocytosis has thus far been suggested to be the
internalization mechanism for naked GNPs. Our TEM images clearly
showed that the endosome contained GNPs, which supports our cell
endocytosis postulation. Chithrani et al(8,9)
demonstrated the receptor-mediated endocytosis process on three
different cell lines by transferring coated GNPs. However,
different ligands coated on the GNPs may lead to different results.
However, how glucose-coated GNPs enter the cell has never been
explored. Glucose can enter the cells by two ways. Facilitated
diffusion and energy-dependent active transport are both mediated
by GLUTs. With facilitated diffusion, GLUTs transport glucose
molecules from higher to lower concentrations. Active transport
occurs with glucose being carried into the cell against a
concentration gradient (34). In
order to determine the internalization mechanism of Glu-GNPs, two
cancer cell lines with GLUT1 overpression were employed (35,36).
Both types of cells were first cultured in glucose-contained DMEM
and then the medium was replaced with glucose-free medium, which
means zero glucose concentration outside the cells. Subsequently,
both the naked GNPs and Glu-GNPs were added into the medium. At
various time-points, the uptake concentrations of the nanoparticles
by the cells were measured by ICP-Mass. As mentioned above, more
glucose-bounded GNPs than naked GNPs were taken by the cells. These
results suggest that GLUTs play an important role in the uptake
process. This process is an energy-dependent transport as the
glucose concentration is lower in the medium. Six functional GLUTs
have been identified (GLUT 1–5,7) thus far. Among them, GLUT-1 is
most important. With the highest glucose affinity, GLUT-1 can
transport glucose molecules even in a lower concentration
environment (37). Based on the
results from our experiments and results from other reports, we can
postulate that the procedure of Glu-GNP internalization is as
follows: glucose molecules on the surface of GNPs first bind with
GLUTs and then the endocytosis process is initiated (the size of
GNPs is too large compared with the inner diameter of GLUT and thus
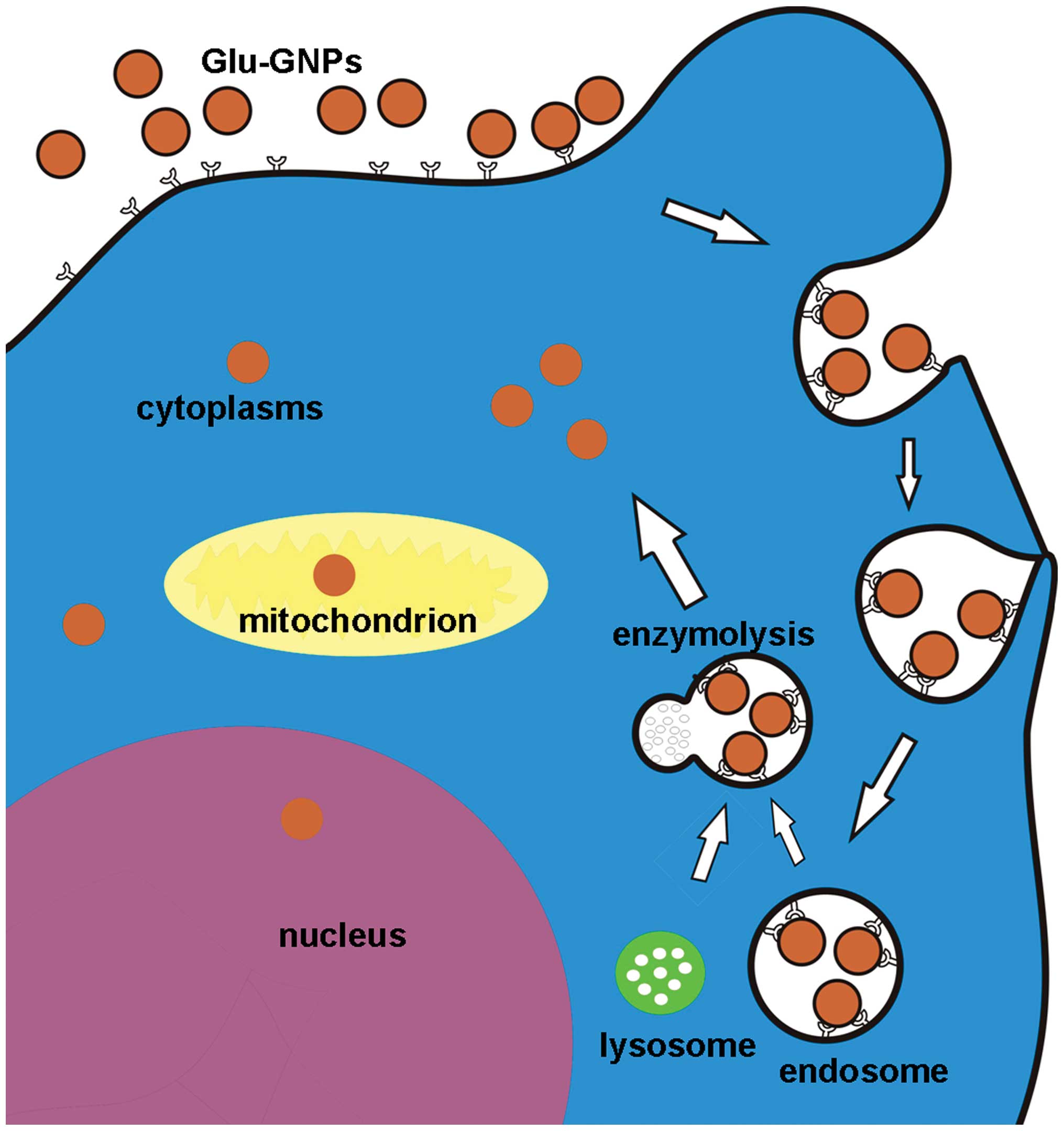
impossible to directly pass through). The postulated schematic
procedure for Glu-GNPs internalization and their subcellular
location is shown in Fig. 9.
Combining nanotechnology and biology offers us
tremendous opportunities to develop improved cancer diagnosis and
therapeutic designs. However, there are also many challenges.
Although nanoparticles can lead to biomedical breakthroughs, they
may also cause toxic side-effects. The majority of studies have
only focused on the positive effects of nanoparticles and have
ignored their negative effects. Nanoparticles, in particular those
made of undegradable non-biological components, can induce
inflammation, affect immune reaction and cause platelet aggregation
(38). GNPs are commonly
considered safe for biomedical applications (39). However, the scientific evidence for
the possible side-effects of GNPs is still lacking. One of the
strategies is to limit the use of GNPs as much as possible before
we completely understand the side-effects involved. In this study,
the nanoparticle concentrations used were low, only
2.5×109 nanoparticles/ml and 1.25×109
nanoparticles/ml, or the dose was much lower than the doses used in
other reports. Our aim was to determine the minimal dose at which
the GNP-radiosensitizing effects can be achieved. According to our
MTT results, the dose of 10 Gy X-ray was the most effective and 96
h of incubation after irradiation was most effective based on our
quantitative analysis results obtained from flow cytometry. At 96 h
after radiotherapy, even at a low dose, GNPs and Glu-GNPs still led
to enhanced irradiation cytotoxicity. The radiosensitivity reached
15% when 20 μM GNPs were used. However, it should be noted
that the radiosensitizing effect of the Glu-GNPs was not
significantly different from that of the GNPs alone (15% vs. 8%,
P=0.054), even though more Glu-GNPs were taken up by the HeLa
cells. We postulated that radicals were generated when the external
radiation source hit the GNPs or Glu-GNPs, which can induce cancer
cell damage. When the surface of the GNPs was coated by glucose,
this coated layer reduced the number of radicals coming out from
the gold surface. On the other hand, evidence has shown a link
between glucose metabolism and radiation resistance (40). Kunkel et al reported that
the overexpression of GLUT-1 is also associated with cancer
resistance to radiotherapy (41).
These studies support the hypothesis that cellular glucose
metabolism directly affects cellular radiosensitivity. In our
study, the cells were exposed in glucose-free medium for 48 h after
either naked GNPs or Glu-GNPs were added. The cells treated with
naked GNPs did not uptake any glucose molecules under these
conditions. However, the cells treated with glucose-capped GNPs
still utilized the glucose attached on the surface of the GNPs.
Therefore, the cells treated with naked GNPs should be more
susceptible to X-rays. We also found in this study that the GNPs
and Glu-GNPs had no effect on X-ray-induced apoptosis (P>0.05).
Generally, the type of cell death through necrosis and/or apoptosis
depends on the drug property, the treatment conditions, the type of
cells involved and the treatment dose (42). X-rays can induce cell death mainly
by cell cycle arrest and partly by apoptosis (43). In this study, X-ray irradiation
induced more apoptosis compared with the controls (P=0.004).
However, TUNEL staining and flow cytometry confirmed that the
apoptotic rate was not altered when the GNPs or Glu-GNPs were
added. Chang et al(11)
demonstrated that GNPs in conjunction with ionizing radiation
significantly retarded tumor growth and induced apoptosis in B16F10
melanoma tumor-bearing mice; these results are not consistent with
our results. Apoptosis is a complex procedure and many factors can
affect its development. We used different cell lines and the dose
of GNPs/Glu-GNPs was relatively low in our in vitro
experiments.
Based on our novel seeding techniques, we
successfully developed high-quality GNPs and Glu-GNPs with larger
sizes (57 and 84 nm). Both types of gold-based nanoparticles were
mainly located in the cytoplasm; however, some nanoparticles
appeared inside the nucleus and on the surface of mitochondrion.
This finding is original. The uptake curve of the GNPs and Glu-GNPs
demonstrated a size- and cell-dependent uptake. TEM studies
confirmed that endocytosis was the internalization mechanism of
both GNPs and Glu-GNPs. Due to glucose being bound to the surface
of the GNPs, cancer cells took up more Glu-GNPs than naked GNPs
through glucose receptor-mediated endocytosis, which is very useful
for targeted delivery in vivo. Furthermore, lower doses of
GNPs and Glu-GNPs can still enhance the killing effect using X-ray
irradiation, although the apoptotic rate is not altered. The data
given in our study may provide useful information as to the
application of GNPs and their modified derivatives in clinical
trials.
Acknowledgements
This study was supported by grants
from the National Natural Science Foundation of China (nos.
81172488 and 30700897), the Doctor Foundation of Education Ministry
(20090131120063) and the Canadian Breast Cancer Foundation.
References
|
1.
|
American cancer society: Cancer Facts and
Figures 2010.
|
|
2.
|
O’Neal DP, Hirsch LR, Halas NJ, Payne JD
and West JL: Photothermal tumor ablation in mice using near
infrared-absorbing nanoparticles. Cancer Lett. 209:171–176.
2004.PubMed/NCBI
|
|
3.
|
Hirsch LR, Stafford RJ, Bankson JA, et al:
Nanoshell-mediated near-infrared thermal therapy of tumors under
magnetic resonance guidance. Proc Natl Acad Sci USA.
100:13549–13554. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
4.
|
Loo C, Lowery A, Halas N, West J and
Drezek R: Immunotargeted nanoshells for integrated cancer imaging
and therapy. Nano Lett. 5:709–711. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
5.
|
Hainfeld JF, Dilmanian FA, Slatkin DN and
Smilowitz HM: Radiotherapy enhancement with gold nanoparticles. J
Pharm Pharmacol. 60:977–985. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
6.
|
Greish K: Enhanced permeability and
retention of macromolecular drugs in solid tumors: a royal gate for
targeted anticancer nanomedicines. J Drug Target. 15:457–464. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
7.
|
Sperling RA, Rivera Gil P, Zhang F,
Zanella M and Parak WJ: Biological applications of gold
nanoparticles. Chem Soc Rev. 37:1896–1908. 2008. View Article : Google Scholar
|
|
8.
|
Chithrani BD, Ghazani AA and Chan WC:
Determining the size and shape dependence of gold nanoparticle
uptake into mammalian cells. Nano Lett. 6:662–668. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
9.
|
Chithrani BD and Chan WC: Elucidating the
mechanism of cellular uptake and removal of protein-coated gold
nanoparticles of different sizes and shapes. Nano Lett.
7:1542–1550. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
10.
|
Zhang X, Xing JZ, Chen J, et al: Enhanced
radiation sensitivity in prostate cancer by gold-nanoparticles.
Clin Invest Med. 31:E160–E167. 2008.PubMed/NCBI
|
|
11.
|
Chang MY, Shiau AL, Chen YH, Chang CJ,
Chen HH and Wu CL: Increased apoptotic potential and dose-enhancing
effect of gold nanoparticles in combination with single-dose
clinical electron beams on tumor-bearing mice. Cancer Sci.
99:1479–1484. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
12.
|
Kong T, Zeng J, Wang X, et al: Enhancement
of radiation cytotoxicity in breast-cancer cells by localized
attachment of gold nanoparticles. Small. 4:1537–1543. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
13.
|
Hainfeld JF, Slatkin DN and Smilowitz HM:
The use of gold nanoparticles to enhance radiotherapy in mice. Phys
Med Biol. 49:309–315. 2004. View Article : Google Scholar
|
|
14.
|
Song K, Kong B, Qu X, Li L and Yang Q:
Phototoxicity of hemoporfin to ovarian cancer. Biochem Biophys Res
Commun. 337:127–132. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
15.
|
Song K, Kong B, Li L, Yang Q, Wei Y and Qu
X: Intraperitoneal photodynamic therapy for an ovarian cancer
ascite model in Fischer 344 rat using hematoporphyrin monomethyl
ether. Cancer Sci. 98:1959–1964. 2007. View Article : Google Scholar
|
|
16.
|
Paciotti GF, Myer L, Weinreich D, et al:
Colloidal gold: a novel nanoparticle vector for tumor directed drug
delivery. Drug Deliv. 11:169–183. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
17.
|
Ghosh P, Han G, De M, Kim CK and Rotello
VM: Gold nanoparticles in delivery applications. Adv Drug Deliv
Rev. 60:1307–1315. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
18.
|
Han G, Ghosh P and Rotello VM:
Functionalized gold nanoparticles for drug delivery. Nanomed.
2:113–123. 2007. View Article : Google Scholar
|
|
19.
|
Han G, Ghosh P and Rotello VM:
Multi-functional gold nanoparticles for drug delivery. Adv Exp Med
Biol. 620:48–56. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
20.
|
Rostgaard J and Qvortrup K: Electron
microscopic demonstrations of filamentous molecular sieve plugs in
capillary fenestrae. Microvasc Res. 53:1–13. 1997. View Article : Google Scholar : PubMed/NCBI
|
|
21.
|
Yuan F, Dellian M, Fukumura D, et al:
Vascular permeability in a human tumor xenograft: molecular size
dependence and cutoff size. Cancer Res. 55:3752–3756.
1995.PubMed/NCBI
|
|
22.
|
Moghimi SM and Szebeni J: Stealth
liposomes and long circulating nanoparticles: critical issues in
pharmacokinetics, opsonization and protein-binding properties. Prog
Lipid Res. 42:463–478. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
23.
|
Hatanaka M: Transport of sugars in tumor
cell membranes. Biochem Bio Phys Acta. 355:77–104. 1974.PubMed/NCBI
|
|
24.
|
Flieri S, Mueckler MM, Usher P and Lodish
HF: Elevated levels of glucose transportand transporter messenger
RNA are induced by ras or src oncogenes. Science. 235:1492–1495.
1987. View Article : Google Scholar : PubMed/NCBI
|
|
25.
|
Yamamoto T, Seino Y, Fukumoto H, et al:
Over-expression of facilitative glucose transporter genes in human
cancer. Biochem Biophys Res Commun. 170:223–230. 1990. View Article : Google Scholar : PubMed/NCBI
|
|
26.
|
Fukuda H, Matsuzawa T, Abe Y, et al:
Experimental study for cancer diagnosis with positron-labeled
fluorinated glucose analogs: [18F]-2-fluoro-2-deoxy-D-mannose: a
new tracer for cancer detection. Eur J Nucl Med. 7:294–297.
1982.PubMed/NCBI
|
|
27.
|
Herold DM, Das IJ, Stobbe CC, Iyer RV and
Chapman JD: Gold microspheres: a selective technique for producing
biologically effective dose enhancement. Int J Radiat Biol.
76:1357–1364. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
28.
|
Nel A, Xia T, Mädler L and Li N: Toxic
potential of materials at the nano level. Science. 311:622–627.
2006. View Article : Google Scholar : PubMed/NCBI
|
|
29.
|
Bhattacharjee SB and Das Gupta NN:
Diffusion distance of the radicals produced by ionizing radiations
in cells. Radiat Res. 16:773–782. 1962. View Article : Google Scholar : PubMed/NCBI
|
|
30.
|
Sobolev AS, Jans DA and Rosenkranz AA:
Targeted intracellular delivery of photosensitizers. Prog Biophys
Mol Biol. 73:51–90. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
31.
|
Li H, Jacque A, Wang F and Byrnes RW:
Diffusion distances of known iron complexes in model systems. Free
Radic Biol Med. 26:61–72. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
32.
|
Elgohary WG, Sidhu S, Krezoski SO,
Petering DH and Byrnes RW: Protection of DNA in HL-60 cells from
damage generated by hydroxyl radicals produced by reaction of
H2O2 with cell iron by zinc-metallothionein.
Chem Biol Interact. 115:85–107. 1998. View Article : Google Scholar : PubMed/NCBI
|
|
33.
|
Rodriguez M, Dargemont C and Stutz F:
Nuclear export of RNA. Biol Cell. 96:639–655. 2004. View Article : Google Scholar
|
|
34.
|
Kahn BB: Dietary regulation of glucose
transporter gene expression: tissue specific effects in adipose
cells and muscle. J Nutr. 124(Suppl 8): S1289–S1295.
1994.PubMed/NCBI
|
|
35.
|
Mendez LE, Manci N, Cantuaria G, et al:
Expression of glucose transporter-1 in cervical cancer and its
precursors. Gynecol Oncol. 86:138–143. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
36.
|
Brown RS and Wahl RL: Overexpression of
Glut-1 glucose transporter in human breast cancer. An
immunohistochemical study. Cancer. 72:2979–2985. 1993. View Article : Google Scholar : PubMed/NCBI
|
|
37.
|
Purrello F, Buscema M and Vetri M: Glucose
regulates both glucose transport and the glucose transporter gene
expression in a hamster-derived pancreatic beta-cell line (HIT).
Diabetologia. 34:366–369. 1991. View Article : Google Scholar : PubMed/NCBI
|
|
38.
|
De Jong WH and Borm PJ: Drug delivery and
nanoparticles: applications and hazards. Int J Nanomed. 3:133–149.
2008.PubMed/NCBI
|
|
39.
|
Lai MK, Chang CY, Lien YW and Tsiang RC:
Application of gold nanoparticles to microencapsulation of
thioridazine. J Control Release. 111:352–361. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
40.
|
Kubota H, Suzuki T, Lu J, et al: Increased
expression of GRP94 protein is associated with decreased
sensitivity to X-rays in cervical cancer cell lines. Int J Radiat
Biol. 81:701–709. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
41.
|
Kunkel M, Moergel M, Stockinger M, et al:
Overexpression of GLUT-1 is associated with resistance to
radiotherapy and adverse prognosis in squamous cell carcinoma of
the oral cavity. Oral Oncol. 43:796–803. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
42.
|
Wyld L, Reed MW and Brown NJ: Differential
cell death response to photodynamic therapy is dependent on dose
and cell type. Br J Cancer. 84:1384–1386. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
43.
|
Ling CC, Guo M, Chen CH and Deloherey T:
Radiation-induced apoptosis: effects of cell age and dose
fractionation. Cancer Res. 55:5207–5212. 1995.PubMed/NCBI
|