Introduction
Cervical cancer, a potentially preventable disease
with a high incidence, remains the second most common malignancy in
women worldwide (1). Approximately
one-third of patients who manifest invasive cervical cancer die
because of this disease (2). The
onset of this disease occurs in young individuals to a high extent
(3). Although advanced surgical
techniques and chemoradiotherapy can improve the treatment rate of
cervical cancer, mortality rate remains high because of tumor
recurrence and drug resistance in chemoradiotherapy (4). As such, novel targets required for
cervical cancer treatment should be developed.
Forkhead box protein M1 (FoxM1) is a specific
transcription factor that belongs to a family of evolutionarily
conserved proteins characterized by the presence of a DNA-binding
domain called the forkhead box (5). The aberrant expression and function
of FoxM1 have been verified in carcinoma progression and malignant
carcinomas, such as lung cancer (6), breast cancer (5), glioblastoma (7), pancreatic cancer (8), gastric cancer (9), hepatocellular carcinoma (10) and cervical cancer (11,12).
FoxM1 is also known as a dynamic cancer-associated biomarker
involved in cell cycle progression, differentiation, DNA damage
repair, angiogenesis and other biological processes (13–20).
In our previous study, the total expression of FoxM1
in cervical cancer tissues is higher than that in normal cervical
tissues, and the nuclear expression of FoxM1 is evidently
correlated with pathological stages (12). These results demonstrate that FoxM1
has been linked to tumorigenesis and progression. FoxM1 also
participates in or stimulates other biological behaviors of tumors,
such as angiogenesis, invasion and metastasis. Studies have
simultaneously demonstrated that the expression of FoxM1 is
positively correlated with urokinase-type PA (uPA), matrix
metalloproteinase (MMP)-2 and MMP-9 expressions, resulting in the
degradation of the extracellular matrix, migration and invasion of
tumor cells (5,8). Another biomarker, VEGF is involved in
angiogenesis and tumor growth; this biomarker has also been
implicated in tumor progression (17,21).
Hence, FoxM1 signalling possibly regulates tumor progression with
these essential factors in cervical cancer.
To elucidate this information, we applied RNAi
technique and evaluated the function of FoxM1 on the proliferation,
apoptosis, migration and invasion of HeLa cells as well as
tumorigenesis and angiogenesis in nude mice. The results may
provide evidence for the molecular-targeted therapy of cervical
cancer.
Materials and methods
Cell culture
Human cervical cancer cell lines, including HeLa,
SiHa and C33A, used in the present study were generously provided
by the Scientific Research Center in Zhongnan Hospital of Wuhan
University. The cells were maintained in DMEM (Hyclone, China)
containing 10% fetal bovine serum, 1% penicillin and streptomycin
in a humid 5% CO2 atmosphere at 37°C.
Plasmid and stable transfection
Human FoxM1-specific RNAi plasmid vectors that
express shRNAs or empty vectors were purchased from Shanghai
Genechem Co. Ltd. (Shanghai, China). All of the vectors were
expressed under the control of a CMV promoter. In brief,
2.0×105 HeLa cells/lane were seeded in a 6-well culture
plate and transfected with the appropriate plasmids by using
Lipofectamine 2000 (Invitrogen, USA) according to the
manufacturer’s instructions. The cells were incubated at 37°C for 6
h. Afterwards, lipid and plasmid complexes were removed and a fresh
medium was added. At 48 h after transfection, stable transfectants
were selected from 700 μg/ml G418 for four weeks. Individual clones
were isolated using pipette tips and maintained in G418 (350
μg/ml).
Cell proliferation by MTT assay and cell
apoptosis by TUNEL assay
We applied MTT assay to determine cell viability.
The number of cells was counted every 24 h for 7 days. A TUNEL
apoptosis detection kit (Promega) was used for DNA fragmentation
fluorescence staining according to the manufacturer’s protocol.
Positively stained, fluorescein-labelled cells were visualised and
counted using a fluorescence microscope (Nikon Eclipse 80i,
China).
Immunofluorescence analysis
In immunofluorescent staining, the cells were fixed
with 4% formaldehyde, blocked for 30 min in 1% BSA prepared in PBS
and incubated overnight in primary antibody at a concentration of
1:100 at 4°C. The cells were subsequently incubated in appropriate
fluorescence-labelled secondary antibody for 1 h at room
temperature. The slides were then mounted with DAPI (Beyotime,
China) to visualize the nucleus. Fluorescent photomicrographs were
obtained using a fluorescence microscope.
ELISA for uPA, MMP-2, MMP-9 and VEGF
The assays were assessed using uPA, MMP-2, MMP-9 and
VEGF ELISA kits according to the manufacturer’s protocol. The
cultivated cells were incubated in 6-well plates for 24 h. UPA,
MMP-2, MMP-9 and VEGF concentrations were measured using the
corresponding ELISA kits (Elabscience, China), afterwards, the
culture medium was collected and centrifuged to remove cell
debris.
Cell migration and invasion assays
Cell migration and invasion assays were obtained
using 24-well chambers (8-μm pore size) with or without Matrigel
according to the manufacturer’s protocol. The cells that penetrated
the membrane were determined by counting the mean cell number of
five ×20 magnification fields randomly and photographed under an
inverted phase-contrast microscope (Nikon Eclipse 80i). These
experiments were repeated in triplicate.
Xenograft experiments
Nude mice (BALB/c nu/nu, females; 4–5-week-old) were
purchased from the Laboratory Animal Center of Wuhan University and
housed under SPF conditions. The experimental protocols were
approved by the Animal Research Committee of Zhongnan Hospital of
Wuhan University. The nude mice were randomly assigned to three
groups. FoxM1-shRNA, empty vector and parental HeLa cells suspended
in PBS were inoculated subcutaneously on the right oxter with
1×107 cells. Tumor growth was measured at an interval of
6 days after injection by using a calliper, and tumor volume was
calculated according to the following formula: length ×
width2 × 0.5 (22). All
the mice were euthanized at day 35 post-inoculation. Harvested
tumor tissues were removed from each mouse, weighed and cut into
two parts. One part was placed in liquid nitrogen and then frozen
at −80°C; the remaining part was fixed in 10% buffered formalin,
embedded in paraffin, sectioned and stained.
Real-time quantitative PCR (qPCR)
assay
Total RNA was extracted from the cultured cells or
tumor xenografts by using TRIzol. The purity of the extracted RNA
was then determined by spectrophotometry. The designed premier
sequences of uPA, MMP-2, MMP-9, VEGF and β-actin genes are shown in
Table I. The first-strand cDNA was
synthesized from 4.823 μg of total purified mRNA in 96-well plates
with these primers in a total volume of 20 μl. The targeted cDNAs
were amplified using SYBR Green Master Mix. The PCR conditions of
the genes included the following: one cycle of 50°C for 2 min and
95°C for 10 min; 40 cycles of 95°C for 30 sec and 60°C for 30
sec.
 | Table IThe designed premier sequences in
qPCR. |
Table I
The designed premier sequences in
qPCR.
| Abbreviations | Forward | Reverse |
|---|
| FoxM1 |
5′-CAACTCAGCCTCCAGGACTC-3′ |
5′-CTGCCTCACCATCACAGGTC-3′ |
| uPA |
5′-CAGGCGTCTACACGAGAGTC-3′ |
5′-TGGCACAGGCAAATCCATCT-3′ |
| MMP-2 |
5′-GATAACCTGGATGCCGTCGT-3′ |
5′-CGAAGGCAGTGGAGAGGAAG-3′ |
| MMP-9 |
5′-CGACGTCTTCCAGTACCGAG-3′ |
5′-TTGTATCCGGCAAACTGGCT-3′ |
| VEGF |
5′-GGTGCCCGCTGCTGTCTAAT-3′ |
5′-GAGATCTGGTTCCCGAAACCC-3′ |
| β-actin |
5′-CACGATGGAGGGGCCGGACTCATC-3′ |
5′-TAAAGACCTCTATGCCAACACAGT-3′ |
Western blot assay
Tumor protein was extracted from cells and tumor
xenografts by using RIPA buffer. Equal amounts of protein (50
μg/lane) were electrophoresed on SDS-PAGE gels and blotted on a
PVDF membrane (Millipore). Rabbit anti-FoxM1 (1:500, Santa, China),
anti-uPA (1:500, Santa, China), anti-MMP2 (1:600, Bioworld, China),
anti-MMP9 (1:600, Bioworld), anti-VEGF (1:500, Abcam, China) and
β-actin (1:1,000, Boster, China) were used as primary antibodies.
HRP-conjugated goat anti-rabbit (1:50,000, Boster) was used as a
secondary antibody. The protein bands were detected on X-ray film
by using an enhanced chemiluminescence detection system.
Microvessel density test
To compare the number of capillaries that formed, we
immunohistochemically analyzed the serial sections from xenograft
tumors of each group. The sections were blocked with 3%
H2O2 at room temperature for 15 min and the
slides were incubated with primary antibody CD31, which is
considered as an endothelial cell-specific marker (1:200,
Bioworld), overnight at 4°C. Secondary antibodies biotin-labelled
anti-rabbit IgG (Bioworld) were used to visualize the specific
markers by avidin-HRP/DAB reaction. Negative controls were obtained
by replacing the primary antibody with PBS. Microvessel density
(MVD) was quantified by observing the number of vessels and
immunoreactive endometrial cells per field at ×100 high-power
magnification in four vascular ‘hot spots’, and other details were
conducted as described previously (23).
Statistical analysis
Data were expressed as mean ± SD from at least three
separate experiments. Statistical analysis was performed using
GraphPad Prism 5 software and SPSS 13.0 software. The statistical
significance of differences was determined by Student’s two-tailed
t-test in two groups and one-way ANOVA in multiple groups.
P<0.05 was considered statistically significant.
Results
Plasmid vector stably expressing FoxM1
shRNA effectively suppresses FoxM1 expression
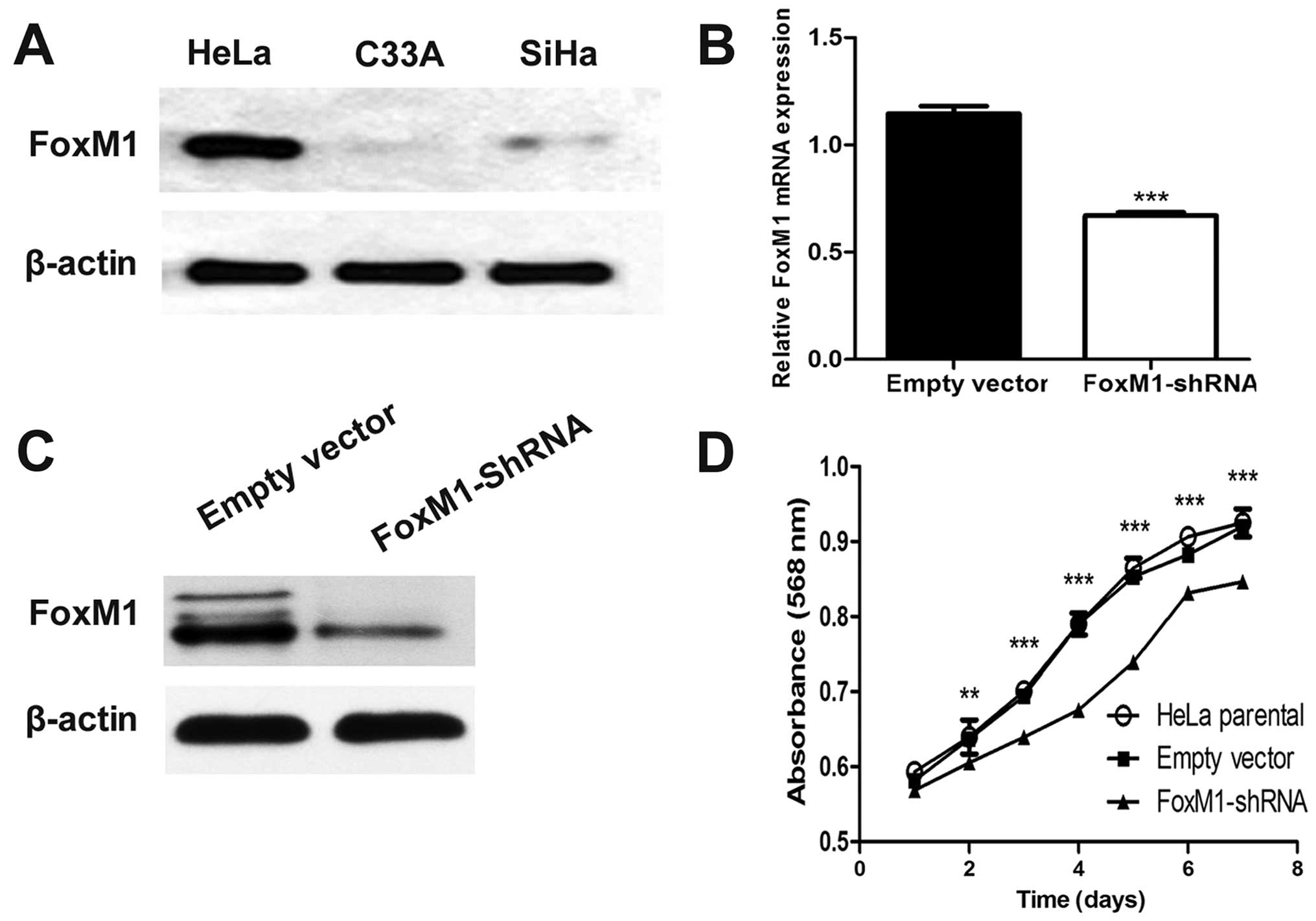
In advance, the baseline expression of FoxM1 in a
panel of cervical cells in our laboratory was determined by western
blot analysis. The highest expression of FoxM1 was observed in HeLa
cells, which were then used in our study (Fig. 1A). After FoxM1-shRNA transfection
was performed, FoxM1 expression was remarkably decreased as
revealed by qPCR and western blot analysis compared with empty
vector-transfected cells (Fig. 1B and
C). These results indicated that FoxM1 expression was
effectively suppressed by the specific shRNA of FoxM1 in HeLa
cells.
FoxM1 knockdown affects cell
proliferation in vitro
To clarify whether or not FoxM1 downregulation can
be a functional alteration, we examined cell viability by
conducting MTT assay. We found that FoxM1 knockdown significantly
inhibited cell growth from the second day (Fig. 1D).
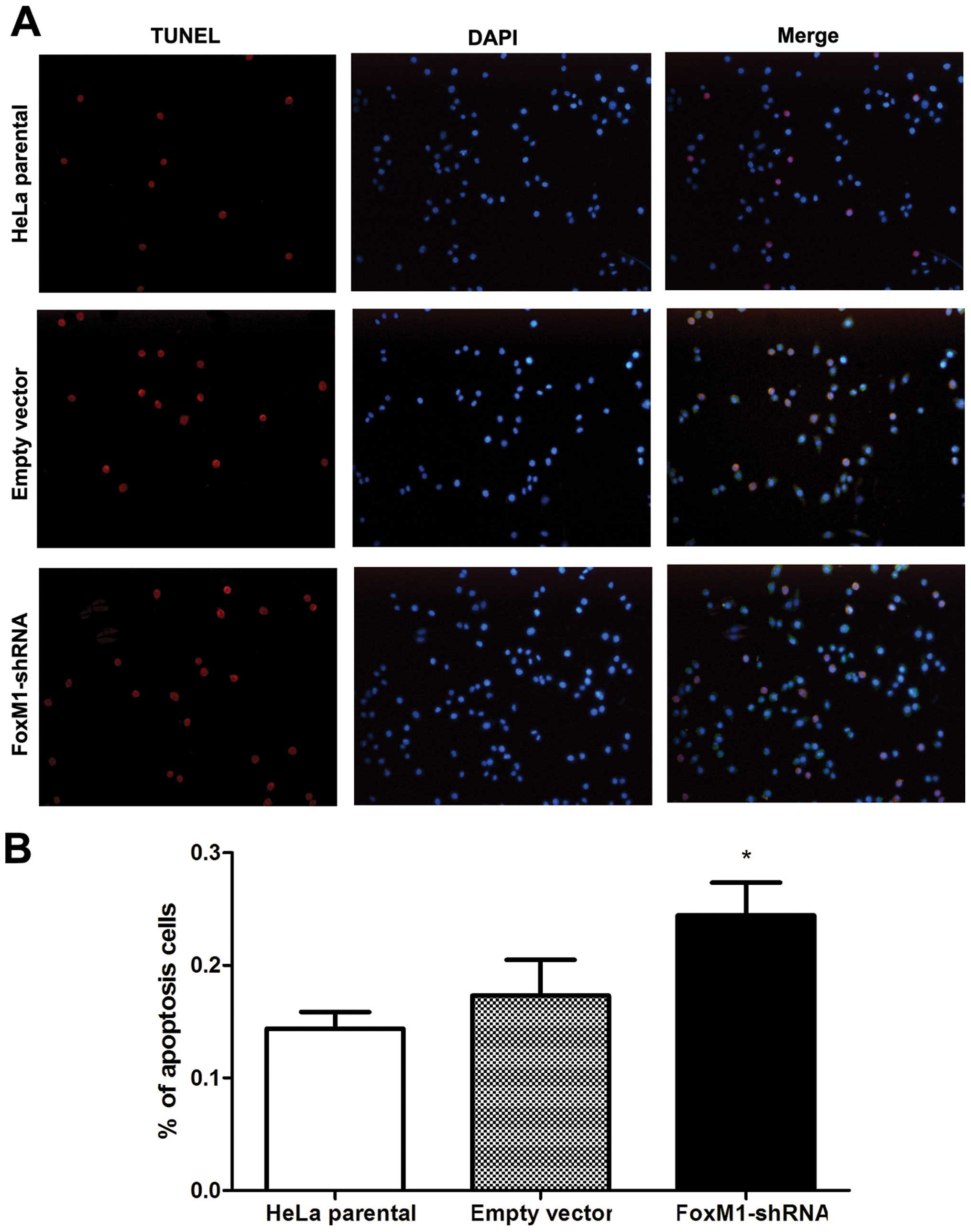
FoxM1 downregulation promotes
apoptosis
To evaluate the effect of FoxM1 knockdown on cell
apoptosis, we investigated nuclear morphology by TUNEL and DAPI
staining. In TUNEL-positive cells, nuclear condensation and
fragmentation representing apoptosis was observed. By contrast, the
normal cells only showed blue DAPI-stained nucleus (Fig. 2A). The number of TUNEL-positive
FoxM1 shRNA-transfected HeLa cells significantly increased compared
with the control cells (Fig. 2B).
Thus, FoxM1 knockdown markedly induced apoptosis of HeLa cells.
FoxM1 downregulation affects uPA, MMP-2,
MMP-9 and VEGF in vitro and in vivo
Several proteins that perform primary functions in
the invasion, migration and metastasis of cervical cancer include
uPA, MMP-2, MMP-9 and VEGF. Immunofluorescence analysis results
demonstrated that FoxM1 downregulation decreased the expressions of
uPA, MMP-2, MMP-9 and VEGF (Fig.
3A). This result is consistent with the signal differences in
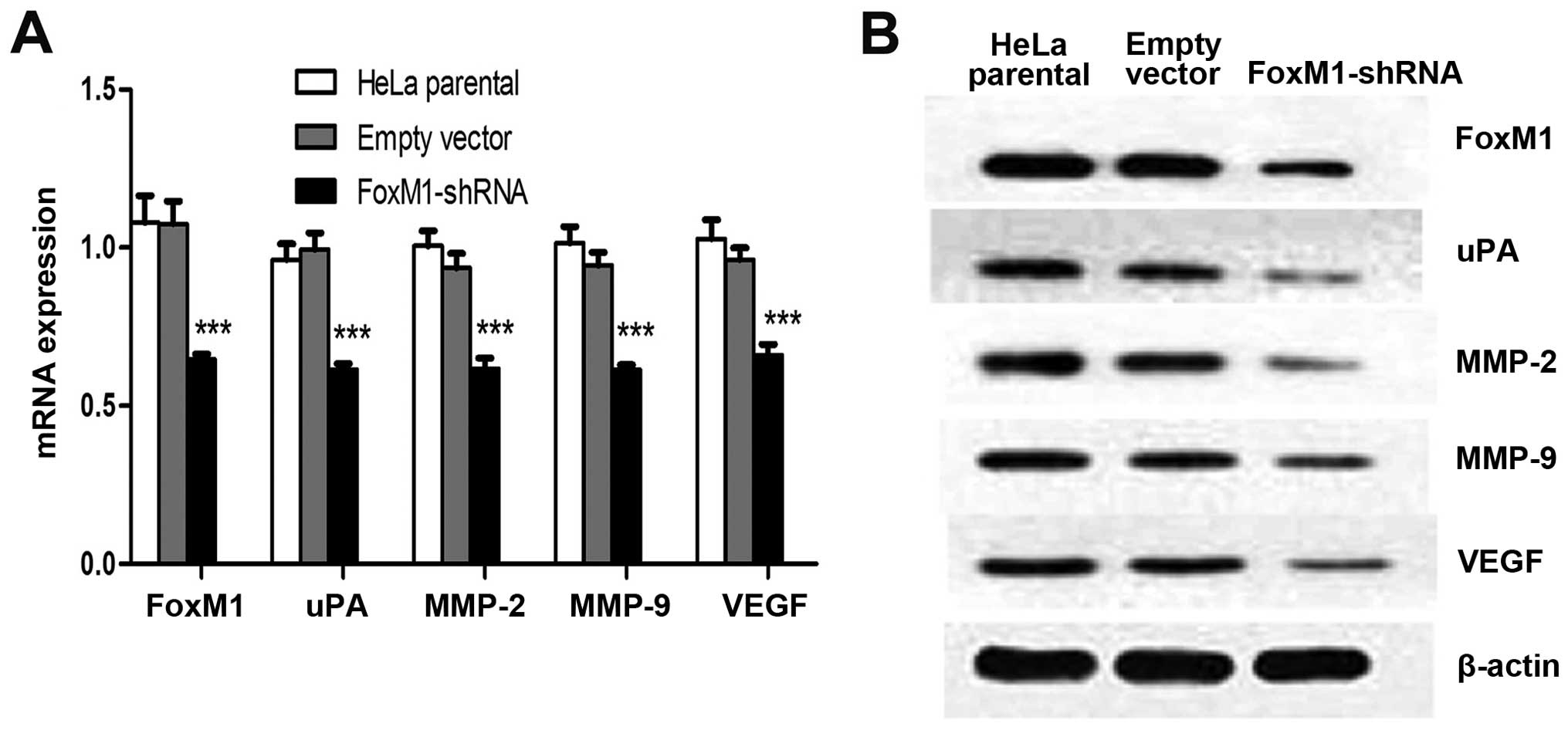
the fluorescence-labelled cells. We conducted qPCR and western blot
analysis to determine whether or not the expression levels of uPA,
MMP-2, MMP-9 and VEGF are influenced by suppressed FoxM1 in HeLa
cells. The mRNA and protein levels of uPA, MMP-2, MMP-9 and VEGF
were markedly decreased in FoxM1 shRNA-transfected cells (Fig. 3B and C). We also conducted ELISA
assays and observed similar patterns in the activities of uPA,
MMP-2, MMP-9 and VEGF in stable-transfected cells (Fig. 3D–G). The primary FoxM1-shRNA
xenografts were associated with the downregulation of these four
factors at mRNA and protein levels (Fig. 4). These results suggested that the
downregulation of FoxM1 inhibited uPA, MMP-2, MMP-9 and VEGF
expressions in vivo and in vitro and prevented
aggressive tumor invasion.
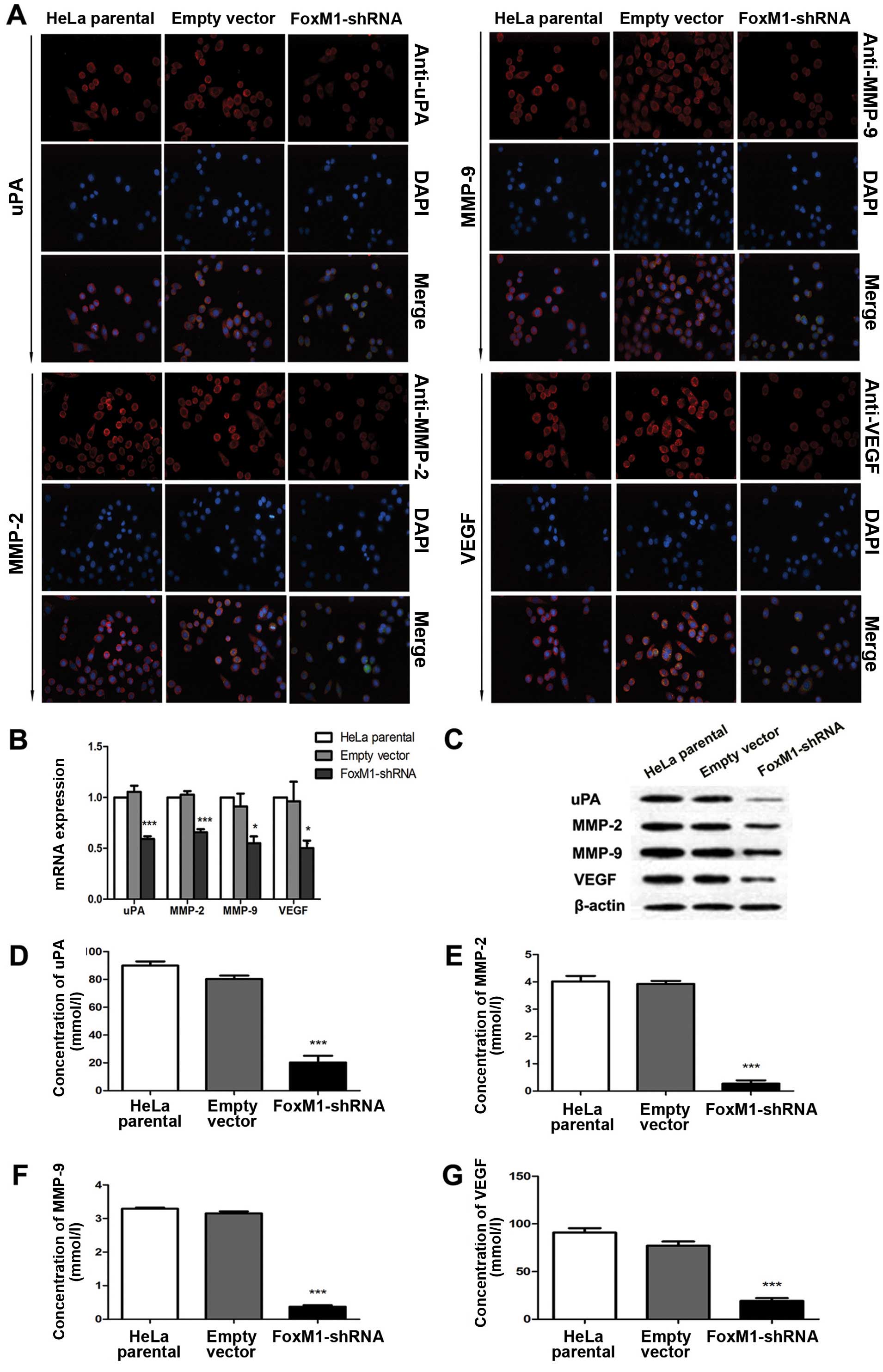 | Figure 3FoxM1 knockdown reduced the
expression levels and activities of uPA, MMP-2, MMP-9 and VEGF
in vitro. (A) The fluorescence signals of uPA, MMP-2, MMP-9
and VEGF were determined in cells after FoxM1 was downregulated.
Merged images are the overlays of uPA, MMP-2, MMP-9 and VEGF red
signals and nuclear staining by DAPI (blue). (B) Statistical
diagrams showing the mRNA levels of uPA, MMP-2, MMP-9 and VEGF. (C)
Protein levels of uPA, MMP-2, MMP-9 and VEGF in FoxM1-shRNA stably
transfected cells, empty vector-transfected cells and HeLa parental
cells were inspected by western blot analysis. (D–G) ELISA assay
results showing that FoxM1 knockdown in HeLa cells inhibits the
activities of uPA, MMP-2, MMP-9 and VEGF. Data are shown as mean ±
SEM from three independent experiments. ***P<0.001
compared with HeLa parental and/or empty vector-transfected
cells. |
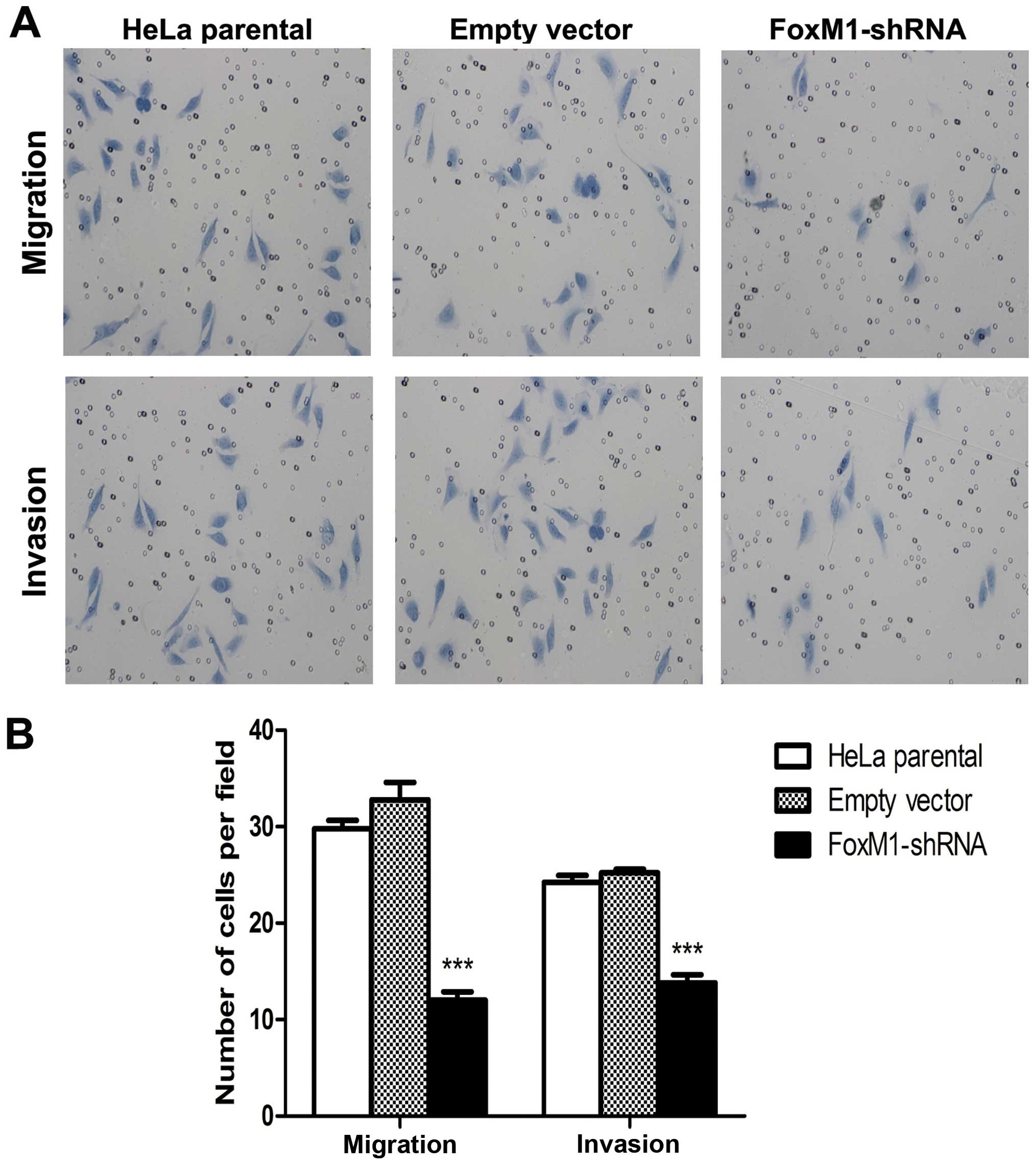
FoxM1 knockdown suppresses cell migration
and invasion
FoxM1 downregulation inhibited the expression and
impaired the activity of several important factors involved in
tumor cell migration and invasion. We further examined whether or
not FoxM1 downregulation affects cell invasion and migration
ability by using a transwell system. The FoxM1 shRNA-transfected
cells showed a low level of penetration into the membrane with
(invasion) or without (migration) Matrigel compared with the
control cells (Fig. 5). These
results showed that FoxM1 downregulation notably suppressed cell
migration and invasion.
FoxM1-shRNA group affects tumor growth in
vivo
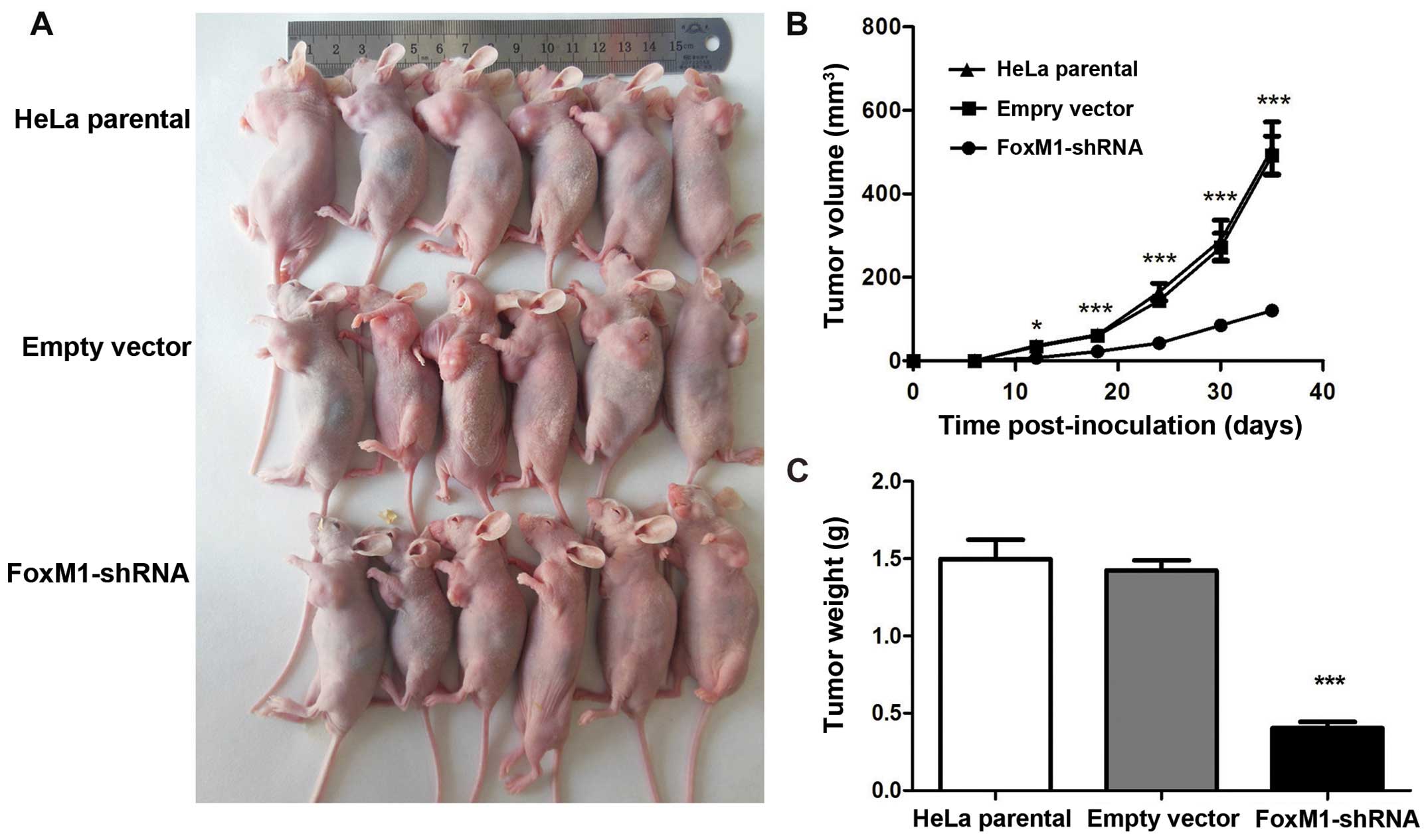
As expected, all of the three groups of cells
developed tumors (Fig. 6A). The
tumor growth curves showed that the growth pattern in the
shRNA-FoxM1 group was significantly slower than that in the two
control groups (Fig. 6B). The mice
were sacrificed at day 35 after injection. Tumor weight in the
shRNA-FoxM1 group were significantly smaller than those in the
parental HeLa group and the empty vector group (Fig. 6C). Tumor inhibitory rate in
shRNA-FoxM1 group reached 72.76% compared with the empty vector
group. No differences were observed between the two control groups
in terms of growth pattern, tumor size and weight. These data
indicated that the suppressed FoxM1 expression of HeLa cells could
inhibit tumorigenicity in nude mice.
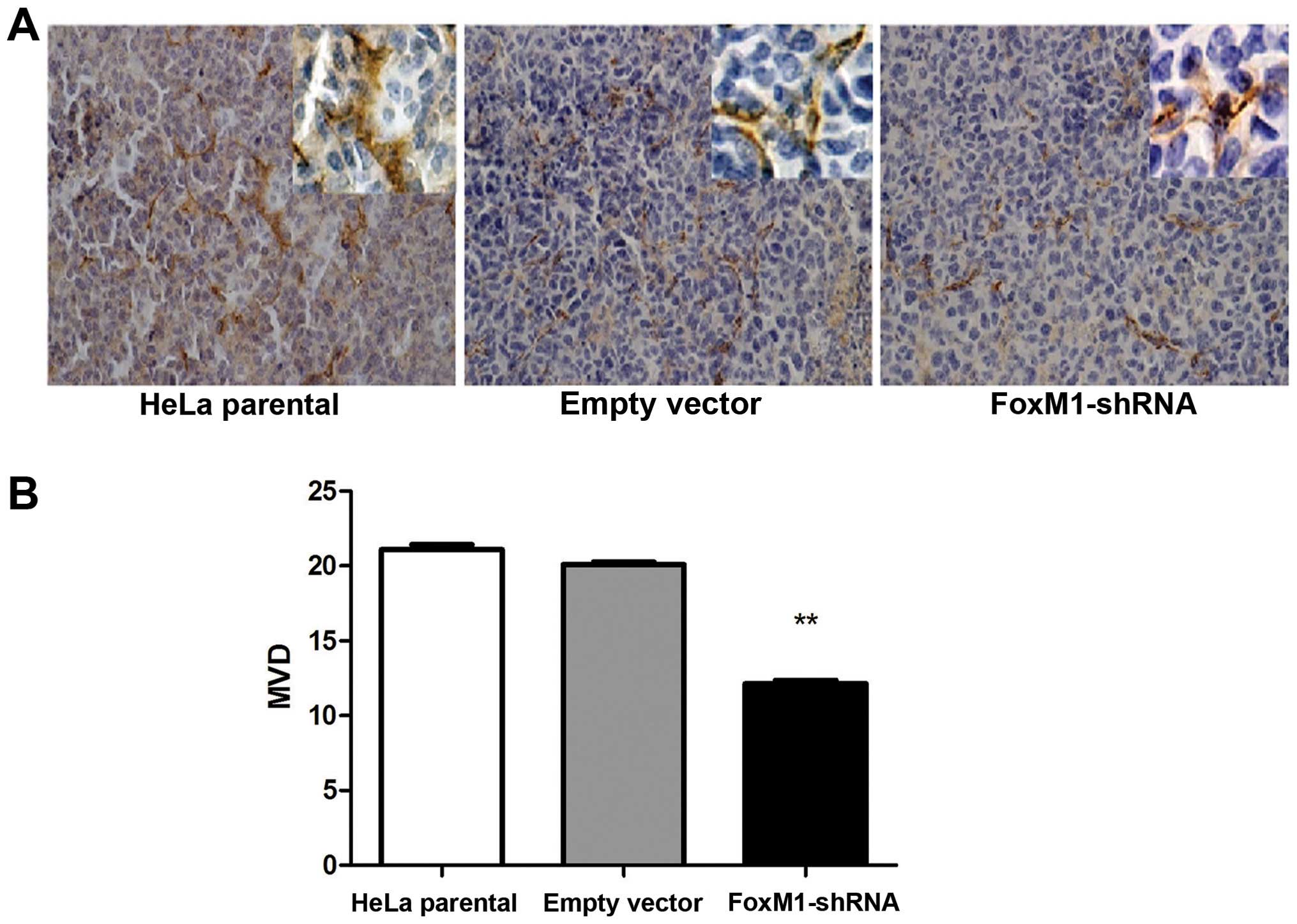
FoxM1 downregulation affects angiogenesis
in xenograft tumors
Tumor induces angiogenesis to maintain the flow of
nutrients for the increasing number of cells. To gain insights into
tumor angiogenesis, we performed the MVD test. These results
indicated that the number of capillary was significantly lower in
the FoxM1-shRNA group compared with control group (Fig. 7). This result indicated a decrease
in the angiogenic potential compared with the control groups.
Discussion
Tumorigenesis occurs as a result of excessive
proliferation combined with reduced apoptosis (24). This factor is the particular target
of FoxM1, a specific transcription factor. FoxM1 has been
considered not only as a major regulatory factor in cell
proliferation (25) but also as a
potential inhibitor of cell apoptosis in malignancies (26,27).
In a previous study, FoxM1 was shown to bind and regulate a group
of genes, which mainly participate in the control of late cell
cycle events in G2 and M phases; binding is manipulated by the
components of the homologous region of the gene involved in the
cell cycle (28). In another
study, DNA damage-induced apoptosis was induced by FoxM1 knockdown
with RNAi or specific proteasome inhibitors (29). To elucidate the function of FoxM1
in the tumorigenesis of cervical cancers, we constructed specific
shRNA and downregulated the expression of FoxM1 in HeLa cells with
a relatively high endogenous expression of FoxM1. MTT assay and
TUNEL assay results showed HeLa cells with attenuated proliferation
and induced apoptosis. Considering the biological nature of these
cells, we also found a similar pattern in our constructed nude
mouse model. This result indicated that the volume and weight of
the tumors were significantly decreased.
Angiogenesis is known as a pre-requisite process in
tumor growth. Nevertheless, angiogenesis is based on the
degradation of ECM components, including basement membrane
collagen, and the release and/or activation of growth factors in
MMPs, particularly MMP-2 and -9 (gelatinases A and B). In a
multitude of malignancies, enhanced MMP-2 and -9 mRNA levels have
been detected (30,31). These factors directly participate
in angiogenesis and metastasis as well as in clinical outcome and
prognosis (32,33). Other studies have revealed that a
latent link is present between FoxM1 and MMPs (6,8,26,34).
For instance, Dai et al (35) found that FoxM1 promotes cellular
invasiveness of glioma cells by upregulating MMP-2 and the relevant
molecular mechanism involves the binding of FoxM1 and activating
the promoter of the MMP-2 gene. The inhibition of MMP-2 and -9
expressions are attributed to the downregulation of FoxM1 in
pancreatic cancer cells and vice versa (8). In our study, FoxM1 downregulation
inhibited expression of MMP-2 and -9 at mRNA and protein levels
in vitro and in vivo. This process also inhibited the
activity of MMP-2 and -9 in the culture medium of HeLa cells. This
result suggested a positive relationship between FoxM1 and MMPs
(MMP-2 and -9).
The invasive ability of tumor cells consists of
various aspects such as the uPA system, which consists of uPA and
its specific cell surface receptor uPAR. The uPA level was
upregulated in pancreatic cancer cells. In addition, the
suppression of uPA-uPAR system results in the downregulation of
angiogenin and decrease in angiogenic potential in vitro and
in vivo (36). In
glioblastoma, a similar pattern can be observed in angiogenesis
after uPA and uPAR are inhibited (37). In other studies, cell viability and
cell invasion ability are significantly reduced after instantaneous
uPA silencing is conducted. Furthermore, decreased tumor growth and
survival rates are observed in an orthotopic mouse prostate cancer
model (38). However, the
information describing the mechanism by which FoxM1 interacts with
uPA in cervical cancer cells remains incomplete. In the present
study, immunofluorescence analysis, real-time PCR, western blot
analysis and ELISA assay results showed that the levels and
activities of uPA decreased after FoxM1 was downregulated compared
with the control cells.
Various growth factors, which have been identified
as critical regulators of angiogenesis and tumor invasion, are
observed in the degradation of ECM. VEGF is considered as the main
angiogenic activator produced and secreted by cancer cells
(39). The autocrine/paracrine
action of VEGF can also promote tumor growth independent of
angiogenesis (40). Angiogenesis
is a process by which new blood vessels grow and can be observed in
physiological and pathological events (41). Angiogenesis occurs when tumor
reaches a diameter ranging from 1 to 2 mm, thereby leading to tumor
growth and metastasis (42). Zhang
et al (43) demonstrated
that FoxM1 functions as an angiogenic switch in tumors by
transcriptionally activating VEGF expression and directly binding
to the Forkhead binding elements (FHRE) of VEGF promoter in glioma
cells. A PB-Cre/Foxm1 fl/fl /TRAMP transgenic mouse model in which
FoxM1 is efficiently deleted has been established, indicating a
marked decrease in the mRNA of VEGF-A as shown by real-time PCR.
The siRNA-mediated depletion of FoxM1 is also observed in TRAMP C2
mouse prostate adenocarcinoma cells. However, aberrant angiogenesis
does not occur (44). In clear
cell renal cell carcinoma, the overexpression of FoxM1 was
determined at mRNA and protein levels. The aberrant expression and
activity of VEGF and angiogenesis are detected after FoxM1 is
downregulated (34). Similar
outcomes can be observed in gastric cancer cells (17). This result is consistent with our
study, in which the reduced levels and activities of VEGF as well
as the number of microvessels in the xenograft tumors induced FoxM1
expression.
We observed that FoxM1 downregulation resulted in
reduced expressions and activities of uPA, MMP-2, MMP-9 and VEGF.
Considering this result, we evaluated the effects of FoxM1
downregulation on the migration and invasion of HeLa cells. We
found that the ability of FoxM1-shRNA transfected cells to migrate
and invade the Matrigel was remarkably weakened compared with that
of the control cells. Therefore, these results indicated that the
knockdown of FoxM1 inhibits the aggressiveness of cervical cancer
possibly by regulating uPA, MMP-2, MMP-9 and VEGF.
Previous studies demonstrated the associations among
uPA, MMP-2, MMP-9 and VEGF. For example, a significant association
between uPA and MMP-2 expression (P=0.028) is found in cervical
intraepithelial neoplasia, which is regarded as pre-cancerous
lesions of cervical cancers (45).
Fang et al (46) verified
that the inhibition of specific MMP-2 results in a decrease in
angiogenic and proteolytic activities of tumor nodules and
restricts tumor growth by 70% in vivo. Fang et al
(46) also demonstrated that the
activated MMP-2 on the cell surface of endothelial cells can
interact with integrin αvβ3, which is highly expressed in melanoma
metastases, to promote angiogenesis and vice versa (47). Tumor cell-induced angiogenesis is
also observed in Ad-MMP-2-infected lung cancer cells, and this
abrogation of MMP-2 resulted in the reduced tumor growth and
formation of lung nodules in mice (48). In another study, the inactivation
of MMP-2 transcriptional level reduces integrin αvβ3,
PI3K/AKT-induced VEGF expression, thereby decreasing tumor
cell-induced angiogenesis (49).
Raghu et al (37) observed
the uPA and uPAR shRNA-mediated inhibition of angiogenesis; this
process can be attributed to the enhanced SVEGFR1 secretion
independent of GM-CSF but dependent on TIMP-1 in endothelial and
glioblastoma cells. Although, He et al (50) has reported the aggressive function
of FOXM1 in cervical cancer via MMP-2/9 and relative signal
pathways, we have demonstrated the function of FoxM1 in
proliferation, apoptosis, migration, invasion and angiogenesis more
systematically. While the detailed mechanisms by which FoxM1 acts
on these cells and the reciprocity among these four factors in
cervical cancers should be further investigated.
References
|
1
|
Siegel R, Ma J, Zou Z and Jemal A: Cancer
Statistics, 2014. CA Cancer J Clin. 64:9–29. 2014. View Article : Google Scholar
|
|
2
|
Pectasides D, Kamposioras K, Papaxoinis G
and Pectasides E: Chemotherapy for recurrent cervical cancer.
Cancer Treat Rev. 34:603–613. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Li S, Hu T, Lv W, et al: Changes in
prevalence and clinical characteristics of cervical cancer in the
People’s Republic of China: a study of 10,012 cases from a
nationwide working group. Oncologist. 18:1101–1107. 2013.
|
|
4
|
Yee GP, de Souza P and Khachigian LM:
Current and potential treatments for cervical cancer. Curr Cancer
Drug Targets. 13:205–220. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Ahmad A, Wang Z, Kong D, et al: FoxM1
down-regulation leads to inhibition of proliferation, migration and
invasion of breast cancer cells through the modulation of
extra-cellular matrix degrading factors. Breast Cancer Res Treat.
122:337–346. 2010. View Article : Google Scholar
|
|
6
|
Kim IM, Ackerson T, Ramakrishna S, et al:
The Forkhead Box m1 transcription factor stimulates the
proliferation of tumor cells during development of lung cancer.
Cancer Res. 66:2153–2161. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Liu M, Dai B, Kang SH, et al: FoxM1B is
overexpressed in human glioblastomas and critically regulates the
tumorigenicity of glioma cells. Cancer Res. 66:3593–3602. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Wang Z, Banerjee S, Kong D, Li Y and
Sarkar FH: Downregulation of Forkhead Box M1 transcription factor
leads to the inhibition of invasion and angiogenesis of pancreatic
cancer cells. Cancer Res. 67:8293–8300. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Feng Y, Wang L, Zeng J, et al: FoxM1 is
overexpressed in Helicobacter pylori-induced gastric
carcinogenesis and is negatively regulated by miR-370. Mol Cancer
Res. 11:834–844. 2013.
|
|
10
|
Kalinichenko VV, Major ML, Wang X, et al:
Foxm1b transcription factor is essential for development of
hepatocellular carcinomas and is negatively regulated by the
p19ARF tumor suppressor. Genes Dev. 18:830–850. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Chan DW, Yu SY, Chiu PM, et al:
Over-expression of FOXM1 transcription factor is associated with
cervical cancer progression and pathogenesis. J Pathol.
215:245–252. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Guan P, Chen H, Li HJ, Duan J and Chen JY:
Expression and significance of FOXM1 in human cervical cancer: a
tissue micro-array study. Clin Invest Med. 34:E1–E7.
2011.PubMed/NCBI
|
|
13
|
Khongkow P, Karunarathna U, Khongkow M, et
al: FOXM1 targets NBS1 to regulate DNA damage-induced senescence
and epirubicin resistance. Oncogene. Oct 21–2013.(Epub ahead of
print). View Article : Google Scholar
|
|
14
|
Raychaudhuri P and Park HJ: FoxM1: a
master regulator of tumor metastasis. Cancer Res. 71:4329–4333.
2011. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Li Y, Ligr M, McCarron JP, et al:
Natura-alpha targets forkhead box m1 and inhibits
androgen-dependent and -independent prostate cancer growth and
invasion. Clin Cancer Res. 17:4414–4424. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Sun H, Teng M, Liu J, et al: FOXM1
expression predicts the prognosis in hepatocellular carcinoma
patients after orthotopic liver transplantation combined with the
Milan criteria. Cancer Lett. 306:214–222. 2011. View Article : Google Scholar
|
|
17
|
Li Q, Zhang N, Jia Z, et al: Critical role
and regulation of transcription factor FoxM1 in human gastric
cancer angiogenesis and progression. Cancer Res. 69:3501–3509.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Wierstra I and Alves J: FOXM1, a typical
proliferation-associated transcription factor. Biol Chem.
388:1257–1274. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Wang IC, Chen YJ, Hughes D, et al:
Forkhead box M1 regulates the transcriptional network of genes
essential for mitotic progression and genes encoding the SCF
(Skp2-Cks1) ubiquitin ligase. Mol Cell Biol. 25:10875–10894. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Laoukili J, Kooistra MR, Bras A, et al:
FoxM1 is required for execution of the mitotic programme and
chromosome stability. Nat Cell Biol. 7:126–136. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Cao Y, EG, Wang E, et al: VEGF exerts an
angiogenesis-independent function in cancer cells to promote their
malignant progression. Cancer Res. 72:3912–3918. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Naito S, von Eschenbach AC, Giavazzi R and
Fidler IJ: Growth and metastasis of tumor cells isolated from a
human renal cell carcinoma implanted into different organs of nude
mice. Cancer Res. 46:4109–4115. 1986.PubMed/NCBI
|
|
23
|
Kim YH, Kim MA, Park IA, et al: VEGF
polymorphisms in early cervical cancer susceptibility,
angiogenesis, and survival. Gynecol Oncol. 119:232–236. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Green DR and Evan GI: A matter of life and
death. Cancer Cell. 1:19–30. 2002. View Article : Google Scholar
|
|
25
|
Katoh M and Katoh M: Human FOX gene family
(Review). Int J Oncol. 25:1495–1500. 2004.PubMed/NCBI
|
|
26
|
Ahmed M, Uddin S, Hussain AR, et al: FoxM1
and its association with matrix metalloproteinases (MMP) signaling
pathway in papillary thyroid carcinoma. J Clin Endocrinol Metab.
97:E1–E13. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Uddin S, Ahmed M, Hussain A, et al:
Genome-wide expression analysis of Middle Eastern colorectal cancer
reveals FOXM1 as a novel target for cancer therapy. Am J Pathol.
178:537–547. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Chen X, Muller GA, Quaas M, et al: The
forkhead transcription factor FOXM1 controls cell cycle-dependent
gene expression through an atypical chromatin binding mechanism.
Mol Cell Biol. 33:227–236. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Halasi M and Gartel AL: Suppression of
FOXM1 sensitizes human cancer cells to cell death induced by
DNA-damage. PLoS One. 7:e317612012. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Kallakury BV, Karikehalli S, Haholu A,
Sheehan CE, Azumi N and Ross JS: Increased expression of matrix
metalloproteinases 2 and 9 and tissue inhibitors of
metalloproteinases 1 and 2 correlate with poor prognostic variables
in renal cell carcinoma. Clin Cancer Res. 7:3113–3119.
2001.PubMed/NCBI
|
|
31
|
Maatta M, Soini Y, Liakka A and
Autio-Harmainen H: Differential expression of matrix
metalloproteinase (MMP)-2, MMP-9, and membrane type 1-MMP in
hepatocellular and pancreatic adenocarcinoma: implications for
tumor progression and clinical prognosis. Clin Cancer Res.
6:2726–2734. 2000.
|
|
32
|
Klein G, Vellenga E, Fraaije MW, Kamps WA
and de Bont ES: The possible role of matrix metalloproteinase
(MMP)-2 and MMP-9 in cancer, e.g acute leukemia. Crit Rev Oncol
Hematol. 50:87–100. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Passlick B, Sienel W, Seen-Hibler R, et
al: Overexpression of matrix metalloproteinase 2 predicts
unfavorable outcome in early-stage non-small cell lung cancer. Clin
Cancer Res. 6:3944–3948. 2000.PubMed/NCBI
|
|
34
|
Xue YJ, Xiao RH, Long DZ, et al:
Overexpression of FoxM1 is associated with tumor progression in
patients with clear cell renal cell carcinoma. J Transl Med.
10:2002012. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Dai B, Kang SH, Gong W, et al: Aberrant
FoxM1B expression increases matrix metalloproteinase-2
transcription and enhances the invasion of glioma cells. Oncogene.
26:6212–6219. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Gorantla B, Asuthkar S, Rao JS, Patel J
and Gondi CS: Suppression of the uPAR-uPA system retards
angiogenesis, invasion, and in vivo tumor development in pancreatic
cancer cells. Mol Cancer Res. 9:377–389. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Raghu H, Nalla AK, Gondi CS, Gujrati M,
Dinh DH and Rao JS: uPA and uPAR shRNA inhibit angiogenesis via
enhanced secretion of SVEGFR1 independent of GM-CSF but dependent
on TIMP-1 in endothelial and glioblastoma cells. Mol Oncol.
6:33–47. 2012. View Article : Google Scholar
|
|
38
|
Pulukuri SM, Gondi CS, Lakka SS, et al:
RNA interference-directed knockdown of urokinase plasminogen
activator and urokinase plasminogen activator receptor inhibits
prostate cancer cell invasion, survival, and tumorigenicity in
vivo. J Biol Chem. 280:36529–36540. 2005. View Article : Google Scholar
|
|
39
|
Zhao R, Liu XQ, Wu XP, et al: Vascular
endothelial growth factor (VEGF) enhances gastric carcinoma
invasiveness via integrin alpha(v)beta6. Cancer Lett. 287:150–156.
2010. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Lee J, Lee J, Yu H, Choi K and Choi C:
Differential dependency of human cancer cells on vascular
endothelial growth factor-mediated autocrine growth and survival.
Cancer Lett. 309:145–150. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Risau W: Mechanisms of angiogenesis.
Nature. 386:671–674. 1997. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Folkman J: Angiogenesis. Annu Rev Med.
57:1–18. 2006. View Article : Google Scholar
|
|
43
|
Zhang Y, Zhang N, Dai B, et al: FoxM1B
transcriptionally regulates vascular endothelial growth factor
expression and promotes the angiogenesis and growth of glioma
cells. Cancer Res. 68:8733–8742. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Cai Y, Balli D, Ustiyan V, et al: Foxm1
expression in prostate epithelial cells is essential for prostate
carcinogenesis. J Biol Chem. 288:22527–22541. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
No JH, Jo H, Kim SH, et al: Expression of
MMP-2, MMP-9, and urokinase-type plasminogen activator in cervical
intraepithelial neoplasia. Ann NY Acad Sci. 1171:100–104. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Fang J, Shing Y, Wiederschain D, et al:
Matrix metalloproteinase-2 is required for the switch to the
angiogenic phenotype in a tumor model. Proc Natl Acad Sci USA.
97:3884–3889. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Silletti S, Kessler T, Goldberg J, Boger
DL and Cheresh DA: Disruption of matrix metalloproteinase 2 binding
to integrin alpha vbeta 3 by an organic molecule inhibits
angiogenesis and tumor growth in vivo. Proc Natl Acad Sci USA.
98:119–124. 2001.PubMed/NCBI
|
|
48
|
Chetty C, Bhoopathi P, Joseph P,
Chittivelu S, Rao JS and Lakka S: Adenovirus-mediated small
interfering RNA against matrix metalloproteinase-2 suppresses tumor
growth and lung metastasis in mice. Mol Cancer Ther. 5:2289–2299.
2006. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Chetty C, Lakka SS, Bhoopathi P and Rao
JS: MMP-2 alters VEGF expression via alphaVbeta3 integrin-mediated
PI3K/AKT signaling in A549 lung cancer cells. Int J Cancer.
127:1081–1095. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
He SY, Shen HW, Xu L, et al: FOXM1
promotes tumor cell invasion and correlates with poor prognosis in
early-stage cervical cancer. Gynecol Oncol. 127:601–610. 2012.
View Article : Google Scholar : PubMed/NCBI
|