Introduction
Bladder cancer (BC) is the 7th most common cancer in
men and 17th most common cancer in women worldwide. Smoking is the
most common risk factor. Exposure to aromatic amines and polycyclic
aromatic hydrocarbons is also a risk factor. Diet and environmental
pollution are not clear risk factors in BC (1).
In recent years, there has been a growing interest
in the use of naturally occurring agents for cancer prevention.
Several meta-analyses showed that allium vegetable intake is
associated with a reduced risk of several cancers (2,3).
Garlic (Allium sativum L.), one of the most ancient
medicinal plants, was first used to treat tumors in 1958 (4) and has been extensively studied in
cancer ever since.
In garlic, the major anticancer organosulfur
compounds are diallyl sulfide (DAS), diallyl disulfide (DADS), and
diallyl trisulfide (DATS). These compounds can induce cell cycle
arrest and apoptosis in cancer, especially BC, and, among them,
DATS has the most potent anticancer activity (5,6). The
association between cancer prevention and garlic is generally
under-studied and has not been studied at all in BC (5–8).
Cancer xenograft models are a useful source of
tissue for array analysis when the availability of human clinical
specimens is limited. The stromal component in mouse-derived,
primary xenograft models have been successfully used for
next-generation sequencing analysis in cancer (9).
Tissue microarray technology makes it possible to
gain comprehensive insight into molecular mechanisms and can survey
the RNA expression levels of more than 10,000 genes simultaneously
(10). ClueGo and Cytoscape can
provide a comprehensive view of a pathway or process among
candidate genes in tissue microarray analysis (11,12).
This study investigated whether garlic extract had
activity in bladder cancer prevention in the T24 BC cell xenograft
model and mechanisms were assessed by tissue microarray and gene
network analysis. Furthermore, we investigated the expression value
of selected genes in the data of 165 BC patients.
Materials and methods
Bladder tumor xenograft model
Drugs
Garlic extract, a white powder of high purity (>
Alliin 10 mg/g), was obtained from Namhaegun Blackgarlic Co., Ltd.
(Gyeongsangnam-do, Korea).
Cell line and cell culture
The T24 cell line was grown in Dulbecco's modified
Eagle's medium (DMEM, Gibco BRL, Grand Island, NY, USA)
supplemented with 10% fetal bovine serum (FBS) (Gibco BRL) and
antibiotic-antimycotic (×100) (Gibco BRL). As recommended by the
supplier, American Type Culture Collection (ATCC, Manassas, VA,
USA), cultures were incubated in a humidified atmosphere of 5%
CO2 at 37°C.
Animals
Male BALB/c-nude mice (5 weeks old, weighing 16–20
g) were used to establish the T24 xenograft tumor model and were
purchased from Daehan Biolink Co., Ltd. (Eumsung, Chungbuk, Korea).
The mice were housed in pathogen-free conditions with a constant
temperature and humidity. All animal tests and experimental
procedures were performed by EBO Co., Ltd. (Cheongju, Korea). All
animal experiments were approved by the appropriate Institutional
Review Boards of the EBO Co., Ltd. (EBOA-2015-4).
Methods
Tumor xenograft inoculation
Cells were grown in culture. After digestion with
0.25% trypsin at 37°C, the cells were washed once in
phosphate-buffered saline (PBS, Welgene Inc., Daegu, Korea) and the
cell concentration was adjusted to 3.6×107/ml with PBS.
The cell suspension was mixed (1:1) with Matrigel™ (BD Biosciences,
Franklin Lakes, NJ, USA; lot 4258010) and implanted into the
scapular region of nude mice by subcutaneous (s.c.) injection (0.2
ml). After 10 days of growth, tumor volume (TV) was assessed by
measuring two perpendicular diameters (L, length; W, width) every 3
days with an electronic caliper. TV was calculated according to the
method developed by the American National Cancer Institute: TV = (L
× W2)/2.
Diet
Mice were randomized into four groups (n=6). After
measuring body weight, the animals were administered garlic powder
every day per os (po.) in 10 ml/kg: 1) distilled water + 0 mg/kg
garlic powder (GP 0 group, negative control), 2) distilled water +
20 mg/kg garlic powder (GP 20 group), 3) distilled water + 200
mg/kg (GP 200 group), and 4) distilled water + 1000 mg/kg (GP 1000
group). After 21 days, T24 cells were implanted into each mouse.
The mice were administered garlic powder for a total of 43
days.
Tumor growth evaluation
Relative TV was assessed by dividing the TV on
different observation days by the starting TV. The mice were
sacrificed on day 23 after inoculation. The tumor growth inhibition
rate (IR) was also used as a reference test and calculated as
follows: IR (%) = (tumor weight of control group − tumor weight of
test group)/tumor weight of control group ×100%.
Statistical analysis
Results were expressed as mean ± SEM. Statistically
significant differences were assessed by Mann-Whitney U test, a
non-parametric test. The criterion for significance was
p<0.05.
Microarray and gene network analysis
RNA purification
RNA purity and integrity were evaluated using a
ND-1000 Spectrophotometer (NanoDrop, Wilmington, DE, USA) and an
Agilent 2100 Bioanalyzer (Agilent Technologies, Palo Alto, CA,
USA).
Labeling and purification
Total RNA was amplified and purified using the
TargetAmp-Nano labeling kit for the Illumina Expression BeadChip
(Epicentre, Madison, WI, USA) to yield biotinylated cRNA according
to the manufacturer's instructions. Briefly, 500 ng of total RNA
was reverse-transcribed to cDNA using a T7 oligo(dT) primer.
Second-strand cDNA was synthesized, in vitro transcribed,
and labeled with biotin-NTP. After purification, the cRNA was
quantified using the ND-1000 Spectrophotometer (NanoDrop).
Hybridization and data export
Labeled cRNA samples (750 ng) were hybridized to
each Human HT-12 v4.0 Expression Beadchip for 17 h at 58°C
according to the manufacturer's instructions (Illumina, Inc., San
Diego, CA, USA). Detection of array signal was carried out using
Amersham fluorolink streptavidin-Cy3 (GE Healthcare Bio-Sciences,
Little Chalfont, UK) following the bead array manual. Arrays were
scanned with an Illumina bead array reader confocal scanner
according to the manufacturer's instructions.
Raw data preparation and microarray
statistical analysis
The quality of hybridization and overall chip
performance were monitored by visual inspection of both internal
quality control checks and raw scanned data. Raw data were
extracted using the software provided by the manufacturer [Illumina
GenomeStudio v2011.1 (gene expression module v1.9.0)]. Array probes
were transformed by logarithm and normalized by the quantile
method. The statistical significance of the expression data was
determined by independent t-test and fold change; the null
hypothesis was an absence of difference between the groups. The
false discovery rate was controlled by adjusting the p-value using
the Benjamini-Hochberg algorithm. For a DEG set, hierarchical
cluster analysis was performed using complete linkage and Euclidean
distance as a measure of similarity. Gene enrichment and functional
annotation analysis was performed for significant probes using Gene
Ontology (www.geneontology.org/). All data analysis and
visualization of differentially expressed genes was conducted using
R 3.1.2 (www.r-project.org).
Gene network analysis
Gene network analysis was performed using Cytoscape
with ClueGo software.
Investigation of the expression of
selected genes in 165 BC patient data
Independent t-test was performed to identify the
difference of gene expression between normal tissues vs. BC patient
tissues. The BC patient data set is available in the NCBI Gene
Expression Omnibus public database (microarray data, GSE13507)
(13).
Results
Study overview
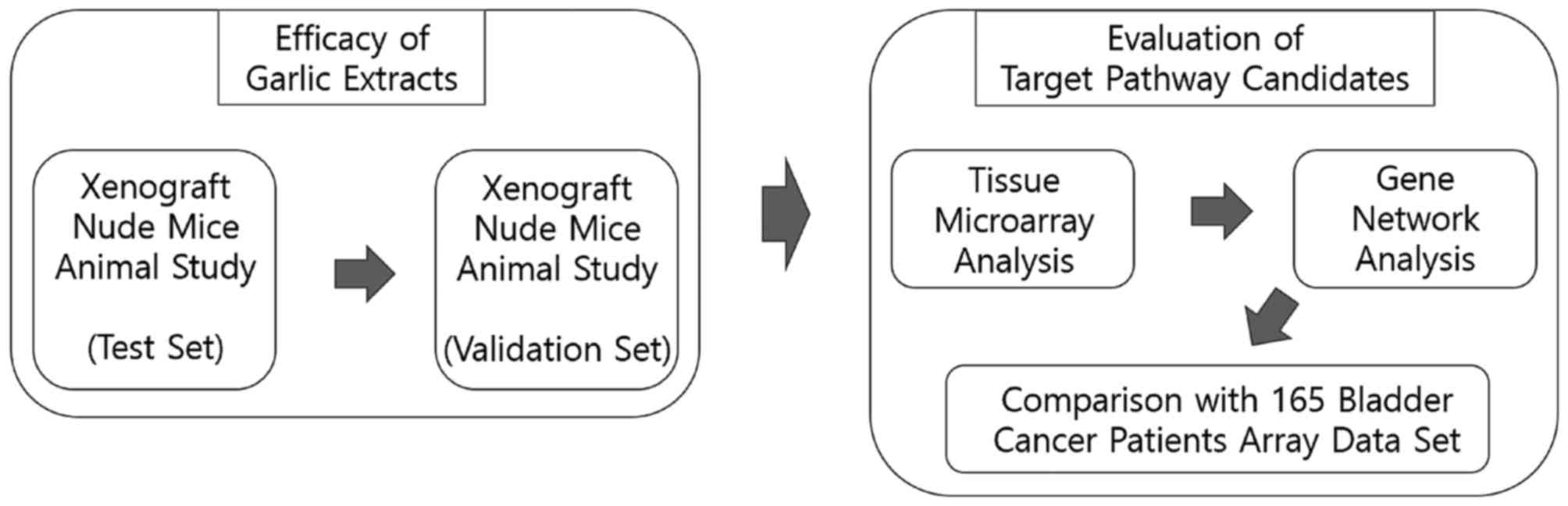
The workflow and overall study design are shown in
Fig. 1.
Garlic extract inhibits the growth of
xenograft tumors
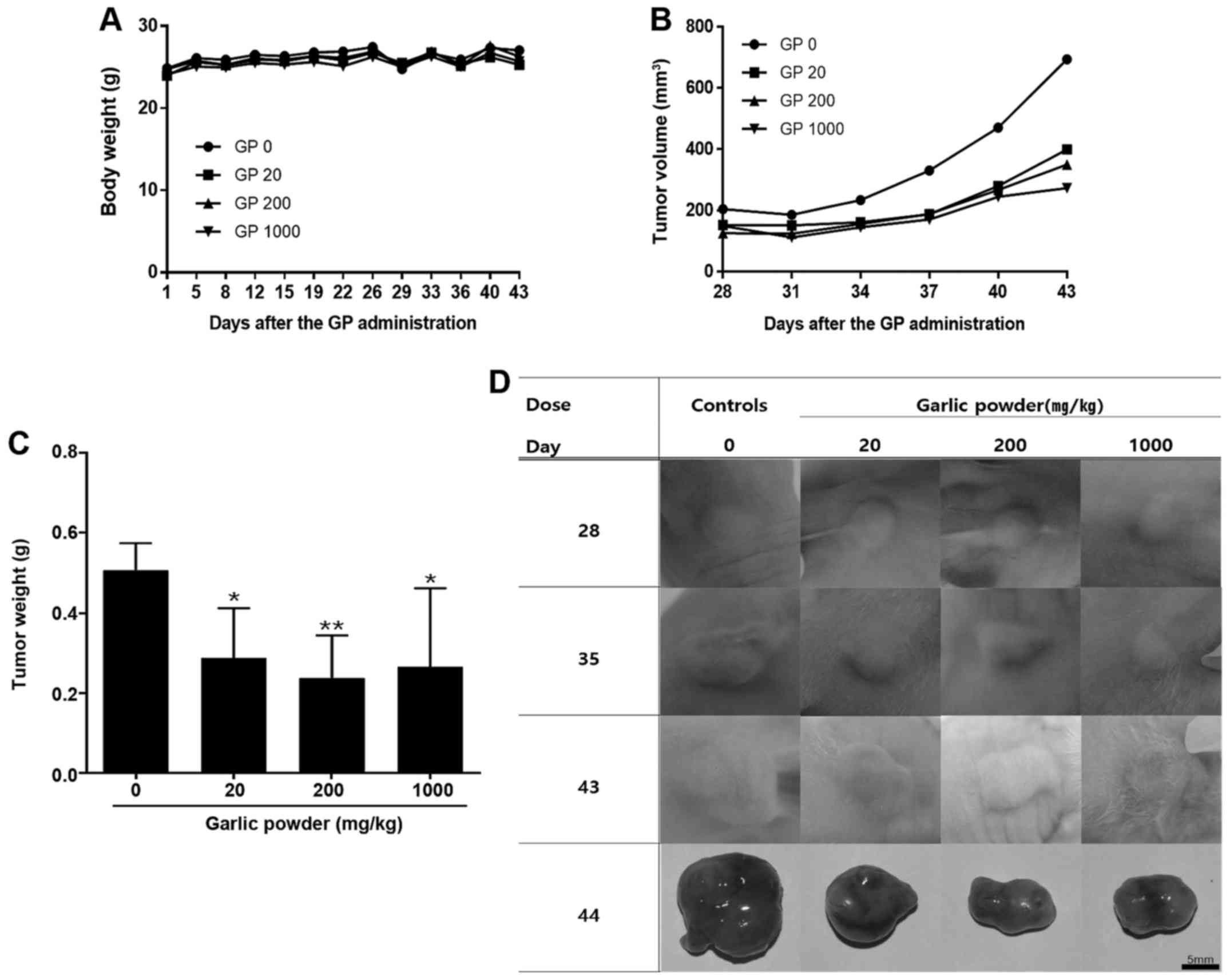
There was no significant difference in body weight
between the garlic extract groups (20, 200, and 1000 mg/kg) and
control group (p>0.05, Fig.
2A). Compared to the control group, significant differences in
TV and tumor weight were observed in the 20 mg/kg (p<0.05), 200
mg/kg and 1000 mg/kg (p<0.01) garlic extract groups (Fig. 2B and C). In addition, TV decreased
in a garlic extract concentration-dependent manner (Fig. 2D).
Validation of xenograft animal study
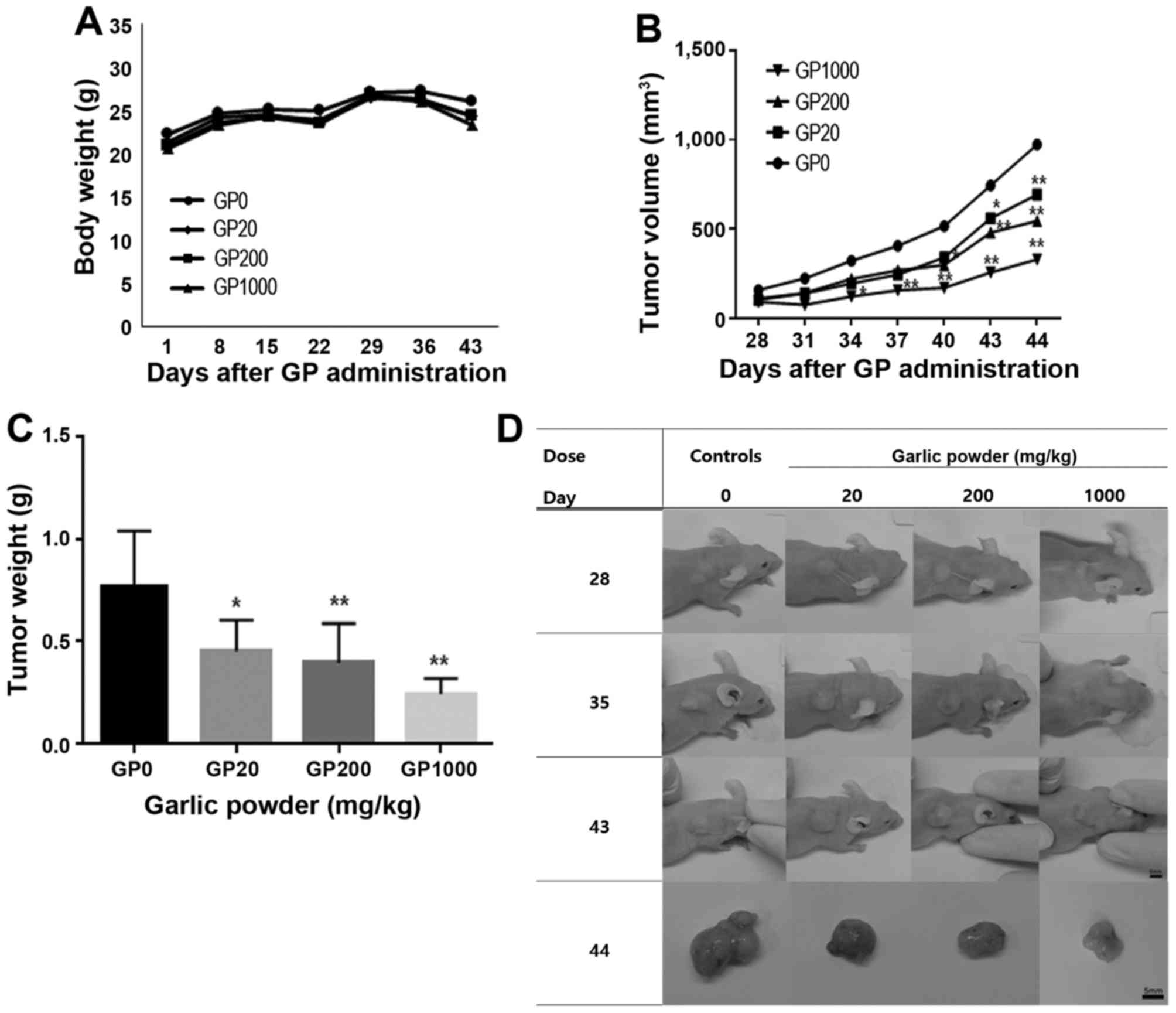
To confirm the preventive effect of garlic extract,
a second xenograft animal study was performed. In this validation
study, a similar effect was observed (Fig. 3).
Gene expression signature associated with
the effect of garlic extract
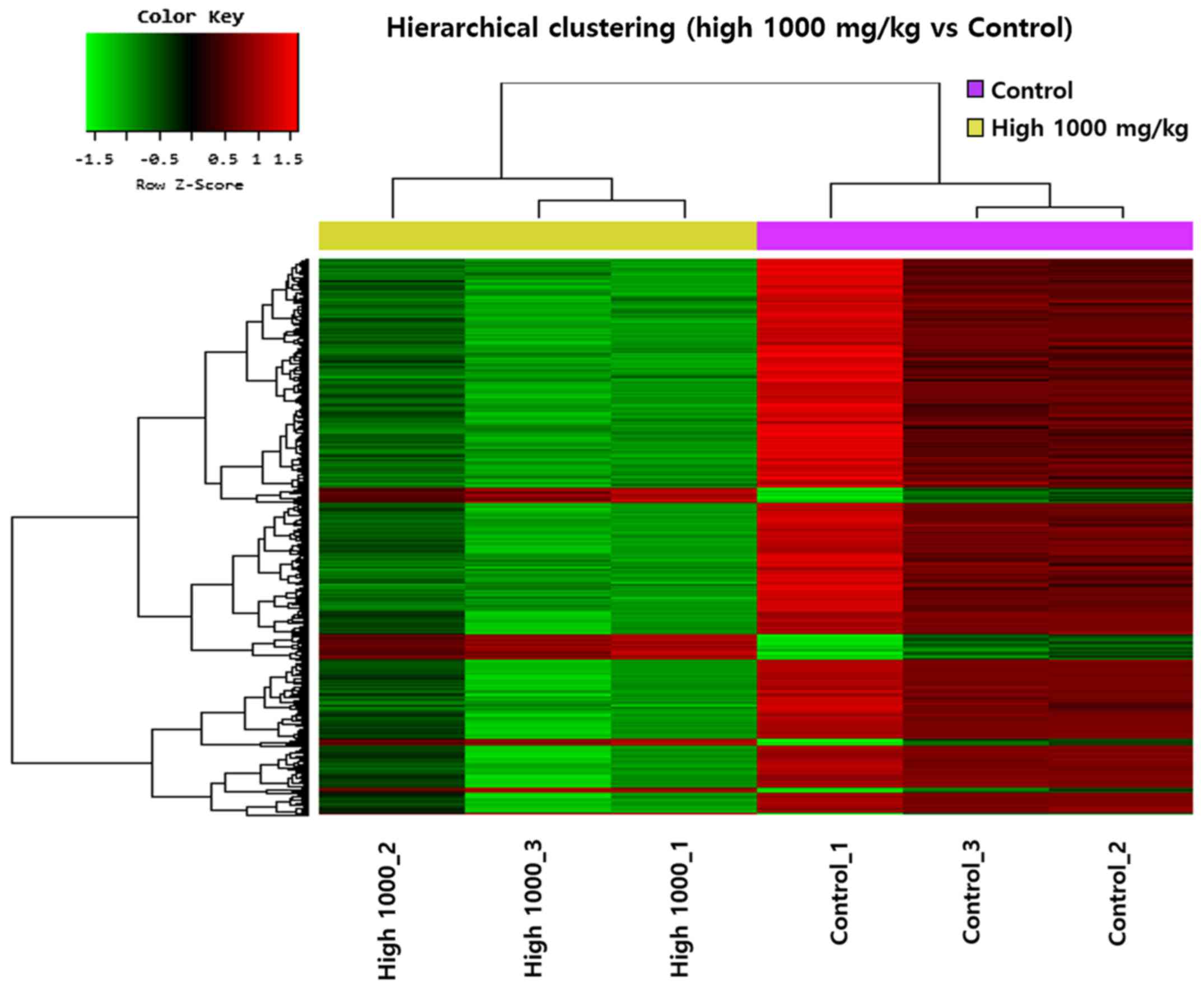
To select cancer prevention-related genes, 19,805
genes were selected by logarithm and normalization among 47,323
genes showing changes in expression levels in the 1000 mg/kg garlic
extract group compared to control. Further, 645 genes were selected
based on fold change and independent t-test analysis (fold change
>2 and p<0.05). Cluster analysis was performed and a
hierarchical clustering heatmap is shown in Fig. 4.
Interpretation of the garlic extract
cancer prevention gene signature by gene network analysis
To identify predominant signaling networks in the
cancer prevention effect of garlic extract, 279 genes were selected
based on p<0.01. Gene network analysis of these 279 genes was
carried out using Cytoscape with ClueGo software. Of the 279 genes,
36 genes and 37 gene ontologies were mapped to gene networks
defined by this tool. The eight highest scoring networks are
candidate mechanisms of action (Fig.
5 and Table I).
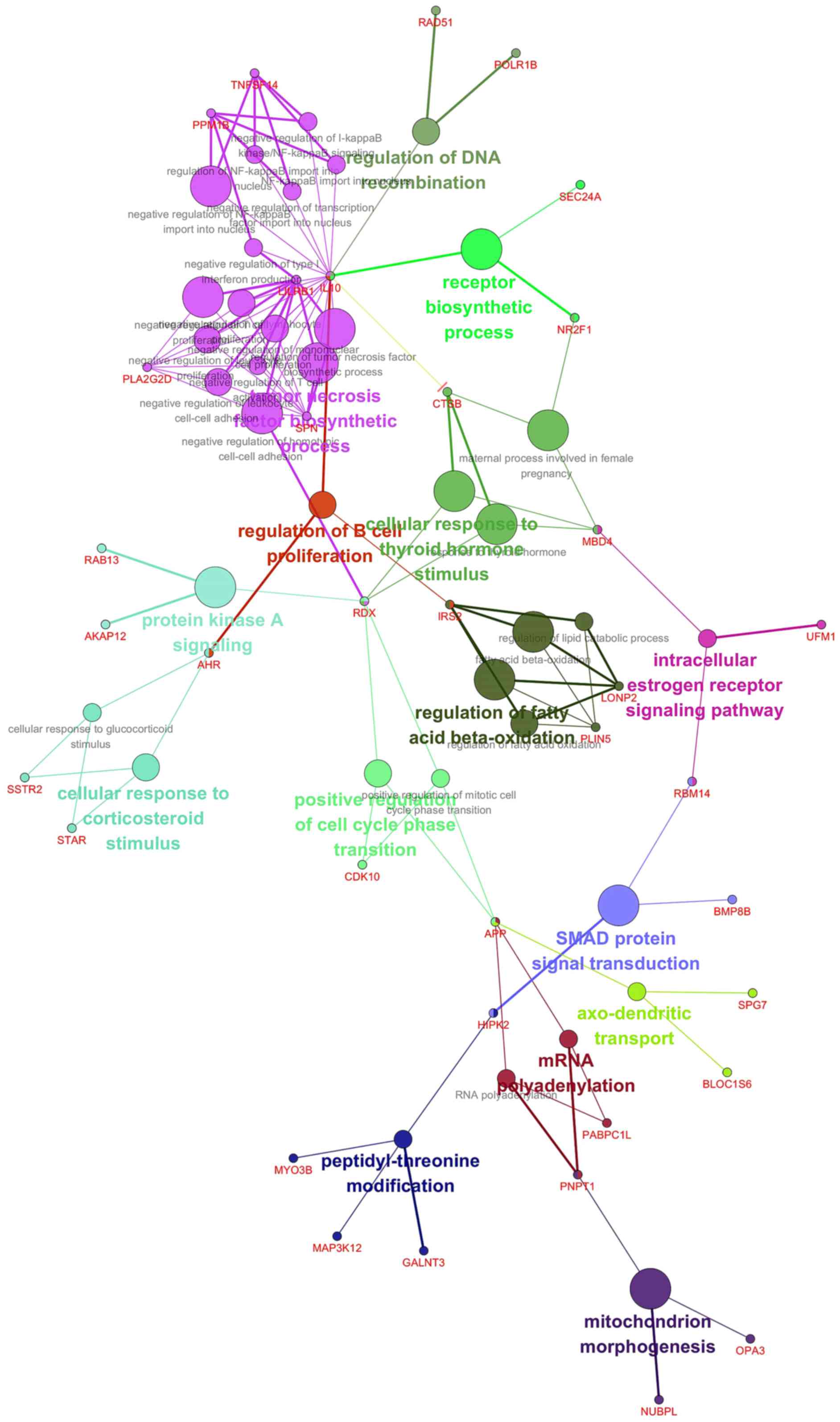 | Figure 5Gene network analysis of the
candidate target pathways of garlic compounds. By gene network
analysis, tumor necrosis factor biosynthetic process, regulation of
B cell proliferation, regulation of DNA recombination, receptor
biosynthetic process, cellular response to thyroid hormone
stimulus, protein kinase A signaling, regulation of fatty acid
β-oxidation, intracellular estrogen receptor signaling pathway,
cellular response to corticosteroid stimulus, positive regulation
of cell cycle phase transition, SMAD protein signal transduction,
axo-dendritic transport, mRNA polyadenylation, peptidyl-threonine
modification, and mitochondrion morphogenesis were possible
candidate pathways. The networks were generated through the use of
Cytoscape with ClueGo (www.cytoscape.org). |
 | Table IGene ontology and associated genes by
gene network analysis using ClueGO. |
Table I
Gene ontology and associated genes by
gene network analysis using ClueGO.
| Gene ontology
term | No. genes | % Associated
genes | Associated
genes |
|---|
| Tumor necrosis
factor biosynthetic process | 3 | 13.636 | IL10, LILRB1,
SPN |
| Regulation of tumor
necrosis factor biosynthetic process | 3 | 13.636 | IL10, LILRB1,
SPN |
| Negative
regulation of leukocyte proliferation | 4 | 5.479 | IL10, LILRB1,
PLA2G2D, SPN |
| Negative
regulation of type I interferon production | 3 | 6.383 | IL10, LILRB1,
PPM1B |
| Negative
regulation of mononuclear cell proliferation | 4 | 5.797 | IL10, LILRB1,
PLA2G2D, SPN |
| Negative
regulation of homotypic cell-cell adhesion | 5 | 4.808 | IL10, LILRB1,
PLA2G2D, RDX, SPN |
| Negative
regulation of leukocyte cell-cell adhesion | 4 | 4.082 | IL10, LILRB1,
PLA2G2D, SPN |
| Negative regulation
of I-κB kinase/NF-κB signaling | 3 | 5.172 | IL10, PPM1B,
TNFSF14 |
| Negative
regulation of lymphocyte proliferation | 4 | 5.797 | IL10, LILRB1,
PLA2G2D, SPN |
| NF-κB import into
nucleus | 3 | 6 | IL10, PPM1B,
TNFSF14 |
| Negative
regulation of T-cell activation | 4 | 4.396 | IL10, LILRB1,
PLA2G2D, SPN |
| Regulation of
NF-κB import into nucleus | 3 | 6 | IL10, PPM1B,
TNFSF14 |
| Negative
regulation of transcription factor import into nucleus | 3 | 6.122 | IL10, PPM1B,
TNFSF14 |
| Negative
regulation of T-cell proliferation | 4 | 7.407 | IL10, LILRB1,
PLA2G2D, SPN |
| Negative
regulation of NF-κB import into nucleus | 3 | 12 | IL10, PPM1B,
TNFSF14 |
| Regulation of
B-cell proliferation | 3 | 4.762 | AHR, IL10,
IRS2 |
| Receptor
biosynthetic process | 3 | 10.714 | IL10, NR2F1,
SEC24A |
| Regulation of
DNA recombination | 3 | 4.68 | IL10, POLR1B,
RAD51 |
| Regulation of
fatty acid β-oxidation | 3 | 16.667 | IRS2, LONP2,
PLIN5 |
| Regulation of fatty
acid oxidation | 3 | 9.667 | IRS2, LONP2,
PLIN5 |
| Fatty acid
beta-oxidation | 3 | 4 | IRS2, LONP2,
PLIN5 |
| Regulation of lipid
catabolic process | 3 | 5.263 | IRS2, LONP2,
PLIN5 |
| Positive
regulation of cell cycle phase transition | 3 | 4.762 | APP, CDK10,
RDX |
| Positive
regulation of mitotic cell cycle phase transition | 3 | 5.172 | APP, CDK10,
RDX |
| mRNA
polyadenylation | 3 | 7.143 | APP, PABPC1L,
PNPT1 |
| RNA
polyadenylation | 3 | 6.977 | APP, PABPC1L,
PNPT1 |
| Cellular
response to thyroid hormone stimulus | 3 | 21.429 | CTSB, MBD4,
RDX |
| Response to
thyroid hormone | 3 | 13.043 | CTSB, MBD4,
RDX |
| Maternal process
involved in female pregnancy | 3 | 4 | CTSB, MBD4,
NR2F1 |
| Cellular
response to corticosteroid stimulus | 3 | 4.615 | AHR, SSTR2,
STAR |
| Cellular response
to glucocorticoid stimulus | 3 | 4.839 | AHR, SSTR2,
STAR |
| Axo-dendritic
transport | 3 | 6.818 | APP, BLOC1S6,
SPG7 |
| Mitochondrion
morphogenesis | 3 | 14.286 | NUBPL, OPA3,
PNPT1 |
| Intracellular
estrogen receptor signaling pathway | 3 | 5.882 | MBD4, RBM14,
UFM1 |
| SMAD protein
signal transduction | 3 | 4.478 | BMP8B, HIPK2,
RBM14 |
|
Peptidyl-threonine
modification | 4 | 4.651 | GALNT3, HIPK2,
MAP3K12, MYO3B |
| Protein kinase A
signaling | 3 | 10.345 | AKAP12, RAB13,
RDX |
Expression value of the selected genes
with 165 BC patient data sets
Of the selected 36 genes, 4 genes (AKAP12, IRS2,
RDX, and UFM1) were significantly increased in garlic feeding
groups, but decreased in BC patients (Table II). Ten genes (RAB13, PLA2G2D,
OPA3, POLR1B, SSTR2, SPG7, RBM14, BMP8B, RAD51, and CDK10) were
significantly decreased in garlic feeding groups, but increased in
BC patients. AKAP12, RDX, and RAB13 are the associated genes with
PKA signaling pathway.
 | Table IIComparisons of gene expression values
between garlic animal model and data of 165 BC patients. |
Table II
Comparisons of gene expression values
between garlic animal model and data of 165 BC patients.
| Gene | Garlic animal model
| 165 BC patients
|
|---|
| Garlic
feeding/controls | p-value | BC
patients/normal | p-value |
|---|
| AKAP12 | Up | 0.003 | Down | <0.001 |
| IRS2 | Up | 0.001 | Down | <0.001 |
| RDX | Up | 0.004 | Down | <0.001 |
| UFM1 | Up | 0.003 | Down | <0.001 |
| RAB13 | Down | 0.007 | Up | 0.017 |
| PLA2G2D | Down | 0.003 | Up | 0.01 |
| OPA3 | Down | 0.007 | Up | <0.001 |
| POLR1B | Down | 0.002 | Up | <0.001 |
| SSTR2 | Down | 0.002 | Up | 0.004 |
| SPG7 | Down | 0.006 | Up | 0.022 |
| RBM14 | Down | 0.006 | Up | <0.001 |
| BMP8B | Down | 0.006 | Up | 0.023 |
| RAD51 | Down | 0.005 | Up | <0.001 |
| CDK10 | Down | 0.008 | Up | <0.001 |
Discussion
The present study identified bladder cancer (BC)
preventive effects in garlic extract. This study is the first of
its kind to investigate the cancer preventive effect of garlic
extract using a BC xenograft model in BALB/c-nude mice. Tissue
microarray analysis and gene network analysis were performed.
Candidate mechanisms of action were identified, including protein
kinase A (PKA) signaling process especially increasing AKAP12 and
RDX, and decreasing RAB13.
In BC, most of the cancer prevention studies testing
garlic have been conducted in vitro using BC cells (5–7).
Several cancer prevention studies tested garlic in ovarian,
pancreatic, esophageal, and hepatocellular cancers (14–17).
Until the present study, garlic in BC prevention has not been
tested using xenograft models. The advantage of xenograft models is
their use of human tumor cells featuring the complexity of the
genetic and epigenetic abnormalities that exist in human tumors and
their suitability to test individualized molecular therapeutic
approaches (18).
The present in vivo study was performed with
and without garlic extract, and tissue microarray analysis using
tumor tissues was employed to identify candidate cancer preventive
mechanisms. The advantage of tissue microarray analysis after
giving garlic extract per os is the simultaneous evaluation of the
expression of more than 10,000 genes (19). Since cancer is a very complicated
disease with multiple heterogeneous genetic and epigenetic changes,
it is very difficult to evaluate the efficacy of garlic extract.
Using tissue microarray analysis, several important key mechanisms
of garlic extract were identified. Future studies will focus on
these candidate mechanisms.
Several reports suggest that garlic extract induces
cell cycle arrest and apoptosis and has antimutagenic properties in
numerous cancer cells (20,21).
Milner suggested that garlic could suppress carcinogen formation,
carcinogen bioactivation, and tumor proliferation (20). Further, garlic extract induced a
caspase-independent apoptotic pathway mediated by mitochondrial
release of AIF and PKA in human epithelial carcinoma cells
(22). PKA belongs to a family of
cyclic AMP-dependent holoenzymes and is involved in cell
proliferation (23). This study
suggested that the major candidate mechanisms of garlic for cancer
prevention were PKA signaling mechanisms. AKAP12, RDX, and RAB13
appear to be key genes mediating the cancer prevention effect of
garlic extract. Especially upregulation of AKAP12 and down
regulation of RAB13 are important pathways for cancer prevention by
garlic extracts.
AKAP12 is a member of the A-kinase anchoring protein
family and was first isolated from the serum of myasthenia gravis
patients (24). In cancer, AKAP12
has been down-regulated in cancer tissue by promoter
hypermethylation and might be involved in the suppression of cancer
(25). In our BC data, AKAP12 was
also downregulated in BC. Yoon et al reported that AKAP12
induces cell cycle arrest and subsequent apoptosis in cancer cells
and also suggested that AKAP12 could antagonize cancer progression
effectively (26). Several studies
strongly suggested that AKAP12 gene acts as a tumor suppressor
(25,26). In our study, upregulated AKAP12 was
found in garlic feeding group. Accordingly, garlic extracts might
restore the activity of tumor suppressor of AKAP12.
The members of the RAB (Ras-related in brain) family
of proteins are known as master regulators of vesicle trafficking.
RAB13 is now recognized as an important driver of cancer
progression (27). Consistent with
a role for RAB13 in cancer progression, RAB13 is upregulated in
many cancers (28,29). In our BC data, RAB13 was also
upregulated in BC. Ioannou et al (30) reported that RAB13 knockdown reduces
cancer cell migration, invasion, and spread in vitro and
in vivo study and showed that RAB13 plays an important role
in promoting tumorigenicity. In our study, downregulated RAB13 was
found in garlic feeding group, garlic extracts might inhibit the
activity of cancer progression of RAB13.
RDX and its roles in cancer remain unclear. However,
RDX decreased in BC patients and increased in garlic feeding group.
Probably, RDX has a tumor suppressive role like AKAP12 in cancer.
PCR validation and in vitro functional validation of these
data will be performed in future studies.
In conclusion, garlic extract intake has strong
cancer prevention activity in vivo and a suitable safety
profile. Using tissue microarray and gene network analysis, PKA
signaling process, especially increasing AKAP12 and RDX and
decreasing RAB13, appear as possible mechanisms underlying this
cancer prevention effect. Further functional in vitro
studies and large human clinical trials are necessary to validate
this study.
Acknowledgments
This work was supported by a National Research
Foundation of Korea (NRF) grant funded by the Korean government
(MSP) (no. 2014R1A2A2A04007036) and by the Functional Districts of
the Science Belt support program, Ministry of Science, ICT and
Future Planning (2016K000297). We thank Bioedit services for
assistance with language editing.
References
|
1
|
Burger M, Catto JW, Dalbagni G, Grossman
HB, Herr H, Karakiewicz P, Kassouf W, Kiemeney LA, La Vecchia C,
Shariat S, et al: Epidemiology and risk factors of urothelial
bladder cancer. Eur Urol. 63:234–241. 2013. View Article : Google Scholar
|
|
2
|
Kodali RT and Eslick GD: Meta-analysis:
Does garlic intake reduce risk of gastric cancer? Nutr Cancer.
67:1–11. 2015. View Article : Google Scholar
|
|
3
|
Guercio V, Turati F, La Vecchia C, Galeone
C and Tavani A: Allium vegetables and upper aerodigestive tract
cancers: A meta-analysis of observational studies. Mol Nutr Food
Res. 60:212–222. 2016. View Article : Google Scholar
|
|
4
|
Butt MS, Sultan MT, Butt MS and Iqbal J:
Garlic: Nature's protection against physiological threats. Crit Rev
Food Sci Nutr. 49:538–551. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Shin DY, Kim GY, Hwang HJ, Kim WJ and Choi
YH: Diallyl trisulfide-induced apoptosis of bladder cancer cells is
caspase-dependent and regulated by PI3K/Akt and JNK pathways.
Environ Toxicol Pharmacol. 37:74–83. 2014. View Article : Google Scholar
|
|
6
|
Wang YB, Qin J, Zheng XY, Bai Y, Yang K
and Xie LP: Diallyl trisulfide induces Bcl-2 and
caspase-3-dependent apoptosis via downregulation of Akt
phosphorylation in human T24 bladder cancer cells. Phytomedicine.
17:363–368. 2010. View Article : Google Scholar
|
|
7
|
Shin DY, Cha HJ, Kim GY, Kim WJ and Choi
YH: Inhibiting invasion into human bladder carcinoma 5637 cells
with diallyl trisulfide by inhibiting matrix metalloproteinase
activities and tightening tight junctions. Int J Mol Sci.
14:19911–19922. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Hu H, Zhang XP, Wang YL, Chua CW, Luk SU,
Wong YC, Ling MT, Wang XF and Xu KX: Identification of a novel
function of Id-1 in mediating the anticancer responses of SAMC, a
water-soluble garlic derivative, in human bladder cancer cells. Mol
Med Rep. 4:9–16. 2011.PubMed/NCBI
|
|
9
|
Rossello FJ, Tothill RW, Britt K, Marini
KD, Falzon J, Thomas DM, Peacock CD, Marchionni L, Li J, Bennett S,
et al: Next-generation sequence analysis of cancer xenograft
models. PLoS One. 8:e744322013. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Quackenbush J: Microarray analysis and
tumor classification. N Engl J Med. 354:2463–2472. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Bindea G, Mlecnik B, Hackl H, Charoentong
P, Tosolini M, Kirilovsky A, Fridman WH, Pagès F, Trajanoski Z and
Galon J: ClueGO: A Cytoscape plug-in to decipher functionally
grouped gene ontology and pathway annotation networks.
Bioinformatics. 25:1091–1093. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Bindea G, Galon J and Mlecnik B: CluePedia
Cytoscape plugin: Pathway insights using integrated experimental
and in silico data. Bioinformatics. 29:661–663. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Kim WJ, Kim EJ, Kim SK, Kim YJ, Ha YS,
Jeong P, Kim MJ, Yun SJ, Lee KM, Moon SK, et al: Predictive value
of progression-related gene classifier in primary non-muscle
invasive bladder cancer. Mol Cancer. 9:32010. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Wu J, Zhao S, Zhang J, Qu X, Jiang S,
Zhong Z, Zhang F, Wong Y and Chen H: Over-expression of survivin is
a factor responsible for differential responses of ovarian cancer
cells to S-allylmercaptocysteine (SAMC). Exp Mol Pathol.
100:294–302. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Wang W, Cheng J and Zhu Y: The JNK
signaling pathway is a novel molecular target for
S-propargyl-L-cysteine, a naturally-occurring garlic derivatives:
link to its anticancer activity in pancreatic cancer in vitro and
in vivo. Curr Cancer Drug Targets. 15:613–623. 2015. View Article : Google Scholar
|
|
16
|
Yin X, Zhang J, Li X, Liu D, Feng C, Liang
R, Zhuang K, Cai C, Xue X, Jing F, et al: DADS suppresses human
esophageal xenograft tumors through RAF/MEK/ERK and
mitochondria-dependent pathways. Int J Mol Sci. 15:12422–12441.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Ng KT, Guo DY, Cheng Q, Geng W, Ling CC,
Li CX, Liu XB, Ma YY, Lo CM, Poon RT, et al: A garlic derivative,
S-allylcysteine (SAC), suppresses proliferation and metastasis of
hepatocellular carcinoma. PLoS One. 7:e316552012. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Richmond A and Su Y: Mouse xenograft
models vs. GEM models for human cancer therapeutics. Dis Model
Mech. 1:78–82. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Russo G, Zegar C and Giordano A:
Advantages and limitations of microarray technology in human
cancer. Oncogene. 22:6497–6507. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Milner JA: A historical perspective on
garlic and cancer. J Nutr. 131:S1027–S1031. 2001.
|
|
21
|
Tsai CW, Chen HW, Yang JJ, Sheen LY and
Lii CK: Diallyl disulfide and diallyl trisulfide up-regulate the
expression of the pi class of glutathione S-transferase via an
AP-1-dependent pathway. J Agric Food Chem. 55:1019–1026. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Park SY, Cho SJ, Kwon HC, Lee KR, Rhee DK
and Pyo S: Caspase-independent cell death by allicin in human
epithelial carcinoma cells: Involvement of PKA. Cancer Lett.
224:123–132. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Cho-Chung YS: Role of cyclic AMP receptor
proteins in growth, differentiation, and suppression of malignancy:
New approaches to therapy. Cancer Res. 50:7093–7100.
1990.PubMed/NCBI
|
|
24
|
Gordon T, Grove B, Loftus JC, O'Toole T,
McMillan R, Lindstrom J and Ginsberg MH: Molecular cloning and
preliminary characterization of a novel cytoplasmic antigen
recognized by myasthenia gravis sera. J Clin Invest. 90:992–999.
1992. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Hayashi M, Nomoto S, Kanda M, Okamura Y,
Nishikawa Y, Yamada S, Fujii T, Sugimoto H, Takeda S and Kodera Y:
Identification of the A kinase anchor protein 12 (AKAP12) gene as a
candidate tumor suppressor of hepatocellular carcinoma. J Surg
Oncol. 105:381–386. 2012. View Article : Google Scholar
|
|
26
|
Yoon DK, Jeong CH, Jun HO, Chun KH, Cha
JH, Seo JH, Lee HY, Choi YK, Ahn BJ, Lee SK, et al: AKAP12 induces
apoptotic cell death in human fibrosarcoma cells by regulating
CDKI-cyclin D1 and caspase-3 activity. Cancer Lett. 254:111–118.
2007. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Ioannou MS and McPherson PS: Regulation of
cancer cell behavior by the small GTPase Rab13. J Biol Chem.
291:9929–9937. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Mahadevan D, Spier C, Della Croce K,
Miller S, George B, Riley C, Warner S, Grogan TM and Miller TP:
Transcript profiling in peripheral T-cell lymphoma, not otherwise
specified, and diffuse large B-cell lymphoma identifies distinct
tumor profile signatures. Mol Cancer Ther. 4:1867–1879. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Li W, Li K, Zhao L and Zou H:
Bioinformatics analysis reveals disturbance mechanism of MAPK
signaling pathway and cell cycle in glioblastoma multiforme. Gene.
547:346–350. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Ioannou MS, Bell ES, Girard M, Chaineau M,
Hamlin JN, Daubaras M, Monast A, Park M, Hodgson L and McPherson
PS: DENND2B activates Rab13 at the leading edge of migrating cells
and promotes metastatic behavior. J Cell Biol. 208:629–648. 2015.
View Article : Google Scholar : PubMed/NCBI
|